Abstract
Human immunodeficiency virus-1 (HIV-1) is a retrovirus with a 10-kb single-stranded RNA genome. HIV-1 must express all of its gene products from the same primary transcript, which undergoes alternative splicing to produce diverse protein products, including structural proteins and regulatory factors1,2. Despite the critical role of alternative splicing, the mechanisms driving splice-site choice are poorly understood. Synonymous RNA mutations that lead to severe defects in splicing and viral replication indicate the presence of unknown cis-regulatory elements3. We use DMS-MaPseq to probe the structure of HIV-1 RNA in cells and develop an algorithm called Detection of RNA folding Ensembles using Expectation-Maximization (DREEM), which reveals alternative conformations assumed by the same RNA sequence. Contrary to previous models, which analyzed population averages4, our results reveal the widespread heterogeneous nature of HIV-1 RNA structure. In addition to confirming that in vitro characterized alternative structures for the HIV-1 Rev Responsive Element (RRE) exist in cells, we discover alternative conformations at critical splice sites that influence the ratio of transcript isoforms. Our simultaneous measurement of splicing and intracellular RNA structure provides evidence for the long-standing hypothesis5–7 that RNA conformation heterogeneity regulates splice site usage and viral gene expression.
Previous work on the genome-wide HIV-1 RNA structure in vitro and in virion provided a population average model, with the underlying assumption that every molecule within the population assumes the same conformation4. However, in vitro studies identified alternative conformations for the HIV-1 RRE and 5’UTR, raising the possibility that alternative structures have roles in viral RNA export from the nucleus and packaging in virions8–10. To resolve the fundamental question of whether RNA structure impacts splicing it is necessary to distinguish multiple conformations for the same sequence in cells. We developed a clustering algorithm called Detection of RNA folding Ensembles using Expectation-Maximization (DREEM) and demonstrated that we can quantitatively detect alternative structures.
DREEM starts with single molecule, chemical probing data, in our case from DMS-MaPseq11. DMS adds methyl groups to unpaired adenine and cytosines of RNA molecules (Fig. 1). The presence of a methyl adduct is read during reverse transcription using TGIRT-III, which marks these sites by incorporating random mutations in the cDNA. PCR amplifies the cDNA product and attaches sequencing adapters to the DNA, followed by massively parallel sequencing. Each resulting read is represented as a binary readout of mutations and matches, which is the input for DREEM (Extended Data Fig. 1a). As DMS-MaPseq has negligible background error11, the mutations observed on a single DNA molecule correspond to the DMS accessible bases on the parent RNA molecule. The two key challenges for detecting heterogeneity are 1) DMS modification rates are relatively low (e.g. an open base has ~2–10% probability of being modified) and 2) the rate of DMS modification per open base is sensitive to the local chemical environment such that not all open bases are equally reactive to DMS. Traditional RNA structure determination approaches combine chemical probing data into population average signal per base, obscuring any underlying heterogeneity. In contrast, DREEM groups sequencing reads issued from each structure into distinct clusters by exploiting information contained in the observation of multiple modifications on single molecules. Theoretically, if two individual bases are DMS reactive in the population average but never both mutated on a single read, it follows that at least two conformations are present. DREEM identifies patterns of DMS-induced mutations on reads and clusters in a mathematically rigorous manner using an expectation-maximization (EM) algorithm (Fig. 1a, Extended Data Fig. 1a). The DMS modification rate per base for each cluster (or structure) is determined by iteratively maximizing a log-likelihood function to find and quantify the abundance of alternative structures directly from the dataset. The binary nature of the readouts allows for the use of a multivariate Bernoulli mixture model (MBMM) to compute the log-likelihood function12. The DMS modification pattern from each cluster is used to create a secondary structure model.
Figure 1 |. Development and validation of DREEM algorithm for analysis of alternative RNA structures.
a, Schematic of combining DMS-MaPseq data with DREEM algorithm to detect alternative RNA structure. b, Structural model of in vitro transcribed and folded Structure 1 and Structure 2 as determined by DMS-MaPseq. Nucleotides are color coded by normalized DMS signal. c, DMS mutational fraction per nucleotide and quantification of Structure 1 and Structure 2 determined by DREEM clustering for a mixing ratio of 25% (Structure 1) to 75% (Structure2) prior to DMS-modification. d, Proportion of Structure 1 and Structure 2 measured by DREEM clustering after in vitro transcription, mixing, and DMS-MaPseq. The expected (E) and observed (O) ratios are shown from an n=1 experiment for each mixing proportion.
Our control experiments on denatured RNA indicated that TGIRT-III is unable to read-through mismatches located within 3nt of each other (Extended Data Fig. 1b). In order to account for this observation, we modified the standard MBMM log-likelihood function (Extended Data Fig. 1a). Upon convergence of the clustering, the DMS-signal from each cluster was used as a constraint in RNAStructure13. DREEM is unique among algorithms for RNA folding ensembles14 because DREEM directly clusters the experimental data. Clustering before secondary structure model generation allows for the discovery of novel RNA structures, in contrast to previous work15,16. Purely computational algorithms rely on suboptimal folding to create variation not captured by minimum free energy calculations. However, using experimentally derived constraints is superior to using randomly generated constraints17,18. Moreover, DREEM does not rely on thermodynamics for detecting and identifying alternative conformations, and therefore can be used on in vivo data to model RNA folding in the presence of cellular factors, whose energetic contributions to RNA structure are unknown.
To validate DREEM, we first transcribed two RNA molecules in vitro that are nearly identical in sequence but form different structures (Structure 1 and Structure 2). These sequences were designed based on the RiboSNitch in the human gene MRPS2119. We experimentally mixed the RNAs from both structures in varying proportions and generated DMS-MaPseq data. DREEM clustered the DMS data and successfully identified the two structures down to a mixing ratio of 6% (Fig. 1b–d, Extended Data Fig. 2). We also tested DREEM using in vitro transcribed and DMS-modified adenosine deaminase (add) riboswitch, which undergoes a conformational shift upon binding of adenine20,21. We found that add structures that promote translation, which are stabilized by adenine, went from 18% to 89% upon addition of 5 mM adenine (Extended Data Fig. 3).
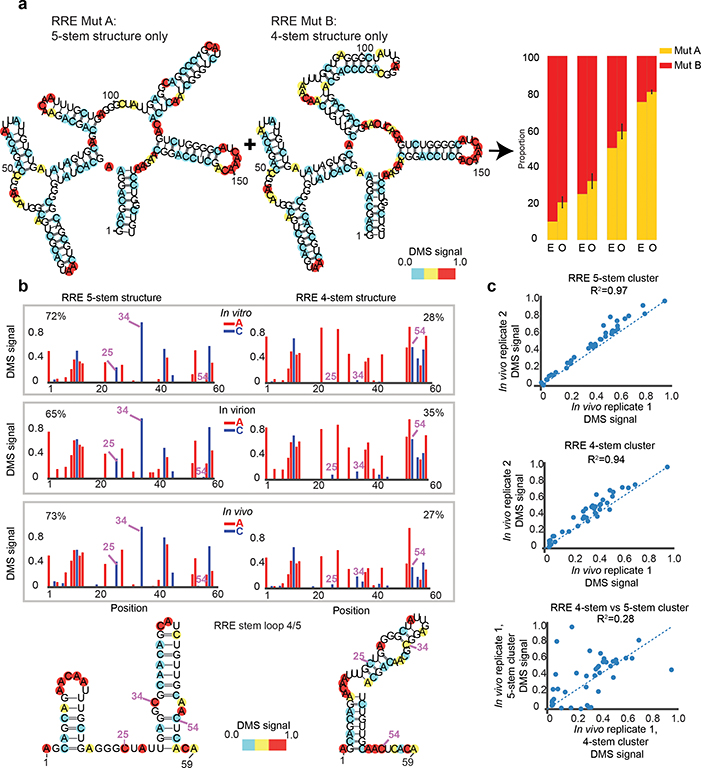
We then focused on the RRE of HIV-1NL4–3, a multi-stem structure that binds to the viral protein rev and allows for the nuclear export of unspliced and partially spliced HIV-1 RNA. Previous studies physically separated distinct RNA conformations by native gel electrophoreses and revealed two alternative structures for RRE in vitro: a 5-stem and 4-stem structure. Specific mutations stabilize either of the alternative conformations11. DREEM accurately identified the DMS signal for mixtures of 5-stem (MutA) and 4-stem (MutB) structures and robustly quantified their mixing ratios (Fig. 2a). Importantly, we found that in vitro folded wildtype RRE sequence exists in a mixture of ~27% 4-stem and ~73% 5-stem structure (Extended Data Fig. 4).
Figure 2 |. Formation of alternate structures at HIV-1 RRE is driven by intrinsic RNA thermodynamics.
a, HIV-1 RRE structural models derived from DMS-MaPseq followed by DREEM using in vitro transcribed structure locked RRE 5-stem (MutA) and 4-stem (MutB) mutants. Bar graphs represent expected (E) and observed (O) mixing ratios of 4-stem and 5-stem structures from an n=2 experiment. b, Normalized DMS signal for RRE 5-stem and 4-stem structures observed in vitro, in virion and in vivo from CD4+ T cells infected with HIV-1NL4–3 identified by DREEM clustering. The positions highlighted are examples of bases that change pairing state between the two structures, shown in both the DMS signal and the folded RNA structures of Stem 4/5. Percentages for each cluster are determined by DREEM from representative samples n=2 for in vivo and in vitro, n=1 for virion. c, Scatter plots of clustering results for n=2 biological replicates (top two plots) and the variation in DMS signal between two different clusters (4-stem vs 5-stem, bottom plot).
We next applied DREEM to the study of HIV-1 RNA structure in primary cells, which is possible as DMS is cell membrane permeable22. Activated CD4+ T cells were infected with HIV-1NL4–3. We performed chemical probing in vivo and in virions (Extended Data Fig. 5a). We discovered that the RRE sequence forms the same alternative structures regardless of the environment (in vitro, in vivo, and in virion), favoring the 5-stem fold (Fig. 2b). These results indicate that the alternative secondary structures of RRE are driven largely by intrinsic RNA thermodynamics as opposed to particular features of the cellular environment. Moreover, these results underscore the ability of DREEM to robustly identify RNA folding ensembles from in vivo data (Fig. 2b–c, Extended Data Fig. 5b) and to quantitate the abundance of the alternate conformations.
We next examined the role of RNA structure in HIV-1 splicing. Alternative splicing is the major mechanism used by HIV-1 to express all of its gene products from a single type of pre-mRNA (i.e. genomic viral RNA). Splice site usage must be regulated to produce the correct proportion of HIV-1 transcripts. HIV-1 transcripts spliced at the A3 acceptor splice site are the only source of mRNAs for the viral transcriptional activator tat1,2.
We discovered alternative structures that dictate the splicing outcome at the A3 splice site and therefore regulate tat transcript abundance. First, the structures that form for HIV-1NL4–3 A3 splice site in CD4+ T cells differ drastically from previously proposed models based on population average data4. Strikingly, the two main conformations identified by DREEM either occlude (~40%, Cluster 1) or expose (~60%, Cluster 2) the polypyrimidine tract and A3 splice site where U2AF heterodimer binds (abbreviated together as A3ss, Fig. 3a). We termed the occluded structure A3 stem-loop (A3SL). The A3SL is not specific to the HIV-1NL4–3 and forms in HIV-1NHG in HEK293t cells (Cluster 1, Fig. 3b). Notably, we detected strong heterogeneity for the A3ss folded in vitro, demonstrating that this region has an intrinsic ability to form multiple conformations, and that the A3SL is thermodynamically stable in the absence of proteins (Extended Data Fig. 6).
Figure 3 |. Alternative RNA structures at the A3 splice acceptor site regulate splice usage.
a, Structural models of the A3ss from CD4+ T cells infected with HIV-1NL4–3 made from clustering outputs of DREEM. Proportions of each cluster are a range from n=4 experiments (1 HIV-1NL4–3 and 3 HIV-1NHG). Nucleotides are color coded by normalized DMS signal. The splice site is highlighted in a blue box, a region that base-pairs to the splice site is in green. b) Scatter plots comparing alternative structures between CD4+ T cells infected with HIV-1NL4–3 and HEK293t cells transfected with HIV-1NHG in an n=2 experiment. The blue dotted line is the identity line; R2 is Pearson’s R2. c, Mutant design; all mutants were predicted to thermodynamically stabilize the A3SL. d, Splice usage fold change compared to ΔvprHIV-1NHG (n=4 for A3SLMut1,3; n=3 for A3SLMut2). The left panel represents splice usage of singly spliced transcripts and the right panel represents multiply spliced transcripts, reported as points with mean and error bars representing s.d.
To perturb the population of RNA structures and measure the effect on splicing, we took advantage of A3ss location in the vpr coding region, which is dispensable for growth in cell-culture. We used a strain with a pre-mature stop codon in vpr to ensure that observed effects were not due to loss of function of vpr (Δvpr HIV-1NHG). To test the effect of structure on splicing, we designed mutations distal from the splice site sequence, avoiding known protein-binding regions. Mutants A3SLMut1, 2, and 3 are predicted to thermodynamically stabilize A3SL and decrease splicing at A3ss (Fig. 3c). Using a deep-sequencing based HIV-1 splicing assay23, we found that all three stabilizing mutants result in lower rate usage of A3ss (Fig. 3d), significantly decreasing expression of tat transcripts relative to background strain.
In contrast, mutations in the same sequence region predicted to have little effect on the stability of A3SL, and therefore little effect on splicing, increased A3ss usage relative to the parental strain (A3SLMut4, Extended Data Fig. 7a–b). To further test the inhibitory role of A3SL, we designed a compensatory mutant to shift the population towards the A3SL in the sequence context of the A3SLMut4. Consistent with A3SL inhibiting splicing, the compensatory mutant (A3SLMut5) uses A3ss ~10-fold less frequently than Δvpr HIV-1NHG (Extended Data Fig. 7b). The percentage of the A3SL cluster for each mutant had an inverse relationship with the overall usage of the A3 splice acceptor site (Extended Data Figure 7c). To understand the origin of the increase in splicing, we probed A3SLMut1 and A3SLMut4 and found that these mutations resulted in the formation of an unanticipated alternative structure in cluster 2 of both mutants (Extended Data Fig. 8a–b). Cluster 2 was present at 35% for A3SLMut1 and 53% for A3SLMut4. This result demonstrates that thermodynamic predictions alone are incomplete. The unanticipated structures alter the accessibility of multiple nearby protein binding sites. These results indicate that the intrinsic ability of RNA to form alternative structures can regulate splicing either by directly occluding U2AF binding sites or by modifying the accessibility of nearby splicing enhancer and silencer elements, the net effect resulting in up to ~100-fold change in HIV-1 tat transcript abundance.
To test whether formation of alternative structures is a general property of the HIV-1 RNA, we prepared a genome-wide DMS-MaPseq dataset from HEK293t cells transfected with HIV-1NHG (Extended Data Fig. 9a). We used DREEM clustering on overlapping windows spanning the entire genome and applied a stringent Bayesian Information Criteria (BIC) test to determine whether the data could be separated into two distinct structure signals24. Importantly, both the RRE and A3ss match the results obtained by specific RT-PCR (Extended Data Fig. 9b–c).
Over 90% of windows with >100,000 sequencing reads coverage passed the BIC test for two clusters, indicating the presence of RNA structure heterogeneity across the entire HIV-1 genome. We quantified the extent of structure in each window using the Gini index metric, which measures the variability in reactivity of residues25. A Gini index close to zero indicates a relatively even distribution of DMS modifications, and occurs when RNA is unfolded or when RNA structure is highly heterogeneous. A Gini index close to one occurs when a subset of residues is strongly protected from DMS, and indicates a highly stable structure. We also computed a Pearson’s correlation coefficient for all windows that had alternative structures to measure how different the two structures were from each other. The low Pearson correlation (R2<0.3) and low Gini index (<0.5) indicate that that relatively unstable, alternative structures form across the entire genome (Fig. 4a), including alternative conformations for a conserved structure26 in the 4kb gag-pro-pol region (Extended Data Fig. 9d), which is present exclusively in unspliced transcripts. The smallest minor cluster that we observed was present at 20%, located in the env coding region (Extended Data Fig. 10a).
Figure 4 |. HIV-1 RNA structure heterogeneity landscape.
HIV-1 genome organization highlighting the UTR, coding regions, major splice donor and acceptor sites and RRE overlaid on structural variability plot for the library generated from HEK293t cells transfected with HIV-1NHG. Each dot represents an 80 nt window of DMS-MaPseq data used for DREEM with a maximum of 2 clusters in an n=1 experiment. The cluster for each window with a higher Gini coefficient is plotted on top in orange and the cluster with a lower Gini coefficient is plotted on bottom in blue. A heat-map comparing the Pearson’s R2 for the 2 clusters is below the Gini coefficient. Windows without sufficient coverage for clustering (<100,000 reads) are in grey. Windows that did not pass the BIC test for more than 1 cluster are in white. A Pearson’s R2 value measures the similarity in the DMS signal between each pair of clusters identified by DREEM. In dark red are most divergent clusters with R2<0.3; orange is 0.3≥ R2.
The widespread alternative structure of HIV-1 genome stood in contrast to snRNA U1 probed in vivo and U4/6 core-domain RNA probed in vitro, which exhibited minimal heterogeneity (Extended Data Fig. 10b–c). These RNAs have stable structures determined by X-ray crystallography27 and NMR28 respectively. As a control against over-clustering, we simulated reads based on the HIV-1 population average DMS-signal with no relationship between mutations. We observed no regions that passed the BIC test for two clusters (Extended Data Fig. 10d). As expected, we observed an inverse relationship between the Gini index and Shannon entropy, an alternative measure of RNA structure (Extended Data Fig. 11a–b). We used the whole genome data to identify previously validated structures such as the transcription activation region (TAR), which was detected in one conformation (Extended Data Fig. 11c). Interestingly, we found structure heterogeneity at most splice sites including A4a-c, and A5 (Extended Data Fig. 11d). Together, these results suggest splice site occlusion as a general mechanism for HIV-1 to tune alternative splicing.
In summary, our results indicate that the thermodynamic ability of RNA to form alternative conformations at critical splice sites can allow HIV-1 to express different genes from the same primary transcript. This may be necessary from an evolutionary perspective for HIV-1 to set an upper limit for splice site usage independent of splice enhancer and suppressor recognition. Splicing repression by RNA structure could ensure that a fraction of molecules remain unspliced, which is essential for packaging and transmitting the full-length HIV-1 genome. Finally, DREEM clustering opens the door to study alternative RNA structures at single nucleotide resolution in living cells. The DREEM approach has wide-range of applications including elucidating the role of RNA structure in human alternative splicing, where as little as 2-fold changes in splice site usage are associated with multiple diseases29,30.
[Methods]
DREEM clustering description
Relevant symbols and meanings
N: Total number of reads
D: Length of region of interest in the reference
X = {x1,…,xN}: Set of all observed reads
S: Set of all allowed (observable) reads
K: Number of clusters
πk: Mixing proportion of cluster k. π = {π1,…,πK} such that .
μk = (μk1,…,μkD): Mutation profile of cluster k, where μki is the mutation rate of base i in cluster k. μ = {μ1,…,μK}.
ynk: The latent Boolean variable representing the assignment of read n to cluster k.
znk: The expectation of ynk, or the probability that read n belongs to cluster k
i,α: nucleotide index
The sequencing data from a sample was mapped to the corresponding reference genome using the Bowtie2 aligner31. The data observed X consists of N reads {x1,…,xN}, each containing D nucleotides. Each read xn ∈ X represents a distinct RNA molecule that was DMS modified, reverse transcribed and amplified. The DMS modifications are read out as mutations. A read xn can then be represented as a vector of D bits (xn1,…,xnD), or a ‘bit vector’, where
As DMS modification is far from saturating (i.e. not every accessible base of a single molecule is modified), each open base in an RNA molecule has only a small probability (2–10%, depending on the DMS concentrations used) of being modified. External factors unrelated to the secondary structure, such as 3D conformation or local chemical environment, will affect this probability. As a consequence of this, a distinct mutation probability μ will be associated with each base of the read. We assume the mutation probabilities are independent from each other. This assumption allows us to consider each read as a random draw from a Bernoulli mixture model. In the event that the RNA molecules assume more than one structure, each structure will appear in the data as a collection of reads, or cluster, characterized by its own Bernoulli mixture model.
If K is the number of structures present in our sample, then the model is parameterized by:
The mutation probabilities μ = {μ1,…,μK}, where μk = (μk1,…,μkD) are the mutation probabilities of cluster k.
The mixing proportions π = {π1,…,πK} of the K clusters, where πk quantify the proportion of reads that belong to cluster k.
The EM algorithm used by DREEM for clustering assumes a Bernoulli mixture model12. Therefore, the probability of a base not being mutated in cluster k is: Pr(xni = 0|μk) = 1 − μki, while the probability of a base being mutated in cluster k is: Pr(xni = 1|μk) = μki. Hence the Bernoulli mixture model gives us the probability of observing a read xn from cluster k as:
| (1) |
We observed that in DMS-MaPseq data, reads that contain mutations within three bases next to each other are very rare and occur at a frequency close to the sequencing error rate (Extended Data Fig. 2); i.e. the bit vectors 001001000, 001010000 and 001100000 are greatly underrepresented. This is likely due to the reverse transcriptase falling off the template when encountering adjacent methylations. Truncated reads do not get amplified during PCR and therefore are not represented when sequenced. To account for this bias, we remove all rare reads containing mutations within three bases of each other and we compute S, the set of all reads with allowable mutations in {0,1}D that can be sequenced. Therefore, equation (1) is modified as follows:
In the initial step of the EM algorithm, the model parameters μ and π are randomly initialized. After the initialization of the parameters, the Expectation step and the Maximization step are executed one after the other in a loop until the log likelihood converges.
Two calculations are made in the Expectation step:
- The responsibilities of the cluster are computed, i.e. the reads are assigned probabilistically to clusters:
Here znk is the probability that read n belongs to cluster k. It can also be defined as the posterior probability, or responsibility, of cluster k given read n.
- The expected complete-data log likelihood of observing the data X and latent variables Y = {ynk} given the model parameters is computed:
In the Maximization step, the model parameters are re-estimated by maximizing the expected value of the likelihood with respect to the parameters {πk} and {μki}.
- Update mixing proportion of each cluster:
- Update mutation profile μk of each cluster by solving the following system of equations for each k:
These equations are derived by setting the derivatives of the expected complete-data log likelihood function to zero.
After the EM clustering algorithm has finished running, the reactivities of the bases in each cluster is given as input to RNAstructure13 for secondary structure prediction. The DMS signal is normalized such that the median of the top ten most reactive positions is set to 1.0. To protect from spurious outliers, we use 90% winsorization effectively capping the reactivity at 1.0. Final visualizations of RNA secondary structure were created with VARNA32.
Parameters used by the DREEM pipeline:
Minimum number of iterations of the EM algorithm to run before checking for convergence of the likelihood (num_its): 300
Number of EM algorithm runs (num_runs): 10. num_runs independent runs of the EM algorithm are carried out to ensure that the results from the algorithm are robust to the initialization of the model parameters and are repeatable.
Convergence threshold (conv_thresh): 1. The EM algorithm is stopped when log(likelihood)iteration = n+1 − log(likelihood)iteration = n < conv_thresh after num_its iterations have been completed.
- Signal threshold (sig_thresh): 0.005. Only mutation rates greater than sig_thresh are considered. All bases with a population average mutation rate less than sig_thresh are set to ‘0’ in every bit vector.
To test for over fitting the data we check whether the EM algorithm passes two clusters by using the BIC test. If BIC K = 2 > BICK = 1, the algorithm stops. Otherwise, the algorithm moves on to K = 3.
Bit vectors are filtered out if they do not satisfy one of the following four criteria:
Informative bits threshold (info_thresh): 0.05–0.2. We set xni to ‘.’ if base i is not covered by read xn and to ‘?’ if it is of low quality (defined as having a Phred Quality Score less than 20). If the fraction of non-informative bits (‘.’, ‘?’ and ‘N’) in the bit vector is greater than info_thresh, the bitvector is removed. After this filtering, all the non-informative bits are set to ‘0’ in the remaining bit vectors.
Maximum number of mutations: If the number of mutations in the bit vector is greater than 3 times the standard deviation of the mutation distribution per read, the bit vector is removed
Invalid bit vectors: rare occurrences of bit vectors with adjacent mutations (within 3nt) are considered to be part of background noise (Extended Data Fig. 2) and are filtered out.
Rare instances where a bit vector consisted of a mutation (‘1’) right next to a non-informative base such as ‘.’ and ‘?’.
Informative bases: Since DMS modifies only As and Cs, mutations at Ts and Gs are set to “0”s.
Cell Lines
HEK293t were obtained from ATCC. The cells tested negative for mycoplasma by LookOut Mycoplasma PCR Detection kit (Millipore-Sigma). The cells were maintained in Dulbecco’s Modified Eagle Medium (ThermoFisher Scientific) supplemented with 10% heat-inactivated fetal bovine serum (FBS; ThermoFisher Scientific) and 100 U/mL penicillin/streptomycin (ThermoFisher Scientific).
Plasmid Construction
HIV-1 NL4–3 Infectious Molecular Clone (pNL4–3) was obtained from the NIH AIDS Reagent program33. HIV-1NHG is a full-length HIV-1 proviral plasmid, modified to replace a non-essential gene nef with GFP (Genbank accession code JQ585717.1). A Vpr-truncated derivative (Δvpr HIV-1NHG) was constructed by generating an overlapping PCR with a C to T mutation and thus a stop codon after Vpr amino acid 20. This PCR product inserted into HIV-1NHG using AgeI and SalI. All of the A3 splice site mutants were generated via overlapping PCR and inserted into a Δvpr HIV-1NHG.
CD4+ T Cell Isolation
Apheresis leukoreduction collars, obtained from the Brigham and Women’s Hospital Crimson Core, were used to isolate peripheral blood mononuclear cell (PBMC) by Lymphocyte Separation Medium (ThermoFisher Scientific) density centrifugation. CD4+ T cells were isolated by negative selection using EasySep Human CD4+ T cell Enrichment Kit (StemCell Technologies). CD4+ T lymphocytes were cultured at a density of approximately 1 million cells/mL in RPMI-1640 (ThermoFisher Scientific) supplemented with 10% fetal bovine serum (FBS) and 100 U/mL penicillin/streptomycin.
DMS Modification of In Vitro Transcribed RNA
gBlocks were obtained from IDT for the HIV-1 RRE, RRE MutA and MutB, control Structure 1, control Structure 2 and Adeno riboswitch. HIV-1 RRE and its mutants correspond to nucleotides 7759–7990 based on HIV-1 vector pNL4–3 (Genbank accession code AF324493.1). Adenosine deaminase (add) riboswitch corresponds to nucleotides 1590535–1590663 of V.vulnificus strain (Genbank Accession code CP037932.1). The U4/6 core-domain RNA construct is based on the interface of the U4 and U6 snRNA (Genbank accession code 2N7M_X). The gblock also contain 20-nt T7 RNA polymerase promoter sequence (TTCTAATACGACTCACTATA) on the 5´ end and a 23-nt sequence (CCGGAGTCGAGTAGACTCCAACA) on the 3´ end. The region of interest was amplified by PCR with a forward primer that contained the T7 promoter sequence. The PCR product was used for T7 Megascript in vitro transcription (ThermoFisher Scientific) according to manufacturer’s instructions. 1 μL Turbo DNase I (ThermoFisher Scientific) was added to the reaction and incubated at 37°C for 15 minutes. The RNA was purified using RNA Clean and Concentrator −5 kit (Zymo). 1–2 μg of RNA was denatured at 95°C for 1 minute. Based on the DMS concentration used in the next step, 300 mM sodium cacodylate buffer (Electron Microscopy Sciences) with 6 mM MgCl2+ was added so the final volume is 100 μl. The RNA was refolded by incubating for 20 mins at 37°C. 0.25%−2.5% dimethyl sulfate (DMS; Millipore-Sigma) was added and incubated at 37°C for 5 mins while shaking at 500 rpm on a thermomixer. The DMS was neutralized by adding 60 μL β-mercaptoethanol (Millipore-Sigma). The RNA was purified using RNA Clean and Concentrator −5 kit. For in vitro transcription of add riboswitch samples, one set of samples were incubated with 5 mM Adenine during refolding stage at 37°C.
CD4+ T Cell Infection and DMS Modification
15 million CD4+ T cells were activated by treatment with culture medium containing 10 μg/mL PHA (Millipore-Sigma) and 100 U/mL IL-234 (NIH AIDS Reagent Program; discontinued) for 72 hours. The cells were pelleted and infected in a small volume with supernatant from HEK293t cells transfected with pNL4–3. After 48 hours, the supernatant was filtered with a 0.22 μM filter (Millipore-Sigma) and centrifuged at 28,000 × g for 1 hour, 4°C in order to pellet virions. The cells were washed and resuspended in 15 mL of media and placed on a thermomixer at 37°C. 200 μL DMS, or ~1.3% v/v, (Millipore-Sigma) was added and the cells were incubated for 10 minutes while shaking at 800 RPM. DMS was neutralized by adding 30 mL PBS (ThermoFisher Scientific) with 30% β-mercaptoethanol. The cells were centrifuged at 1000 × g for 5 mins, 4°C. The cells were washed twice by resuspending the pellet with 15 mL PBS with 30% β-mercaptoethanol and centrifugation to pellet. After washes, the pellet was resuspended in 1 mL Trizol (ThermoFisher Scientific) and RNA was extracted following manufacturer’s specifications. The virions were resuspended in 400 μL PBS with 10 mM Tris pH 7 and 3 mM MgCl2+. 40 μL DMS was added and the virions were incubated at 37°C on a thermomixer while shaking at 800 RPM for 10 minutes. The DMS was neutralized with 400 μL β-Mercaptoethanol and the RNA was purified using RNA Clean and Concentrator −5 kit. For unmodified RNA, 15 million CD4+ T cells were isolated and infected the same as described. 72 hours after infection, the supernatant was filtered with a 0.22 μM filter and virions were pelleted from the supernatant by centrifugation at 28,000×g for 1 hr, 4°C and resuspended in 1 mL Trizol. The cells were pelleted and resuspended in 1 mL Trizol. RNA was extracted following manufacturer’s instructions.
HEK293t Transfection and DMS Modification
0.9 million cells per well were seeded on a 6-well plate and incubated overnight. 2 μg of plasmid DNA (NL4–3, NHG or mutant) per well was transfected into the cells using X-tremeGENE 9 (Millipore-Sigma) following manufacturer’s instructions and incubated for 48 hours. After incubation, virions were collected from the supernatant and DMS modified as above. The cells were washed with PBS and 2 mL medium with ~1.3% v/v DMS was added to each well. The plates were incubated at 37°C for 4 mins. The medium containing DMS was immediately removed and replaced with PBS with 30% β-mercaptoethanol. Cells were scraped and centrifuged at 1000 × g for 5 mins, 4°C. The pellet was resuspended in PBS and centrifuged to pellet twice. The pellet was resuspended in 1 mL Trizol and RNA was extracted following manufacturer’s specifications. For unmodified RNA, HEK293t were plated and transfected with the same protocol as above. 48 hours after transfection, the supernatant was filtered with a 0.22 μM filter and virions were pelleted from the supernatant by centrifugation at 28,000×g for 1 hr, 4°C and resuspended in 1 mL Trizol. The cells were trypsinized, washed and resuspended in 1 mL Trizol. RNA was extracted following manufacturer’s instructions.
RT-PCR with DMS-modified RNA from Cells or In Vitro Transcription
1–3 ug of RNA per reaction was used as the input for rRNA subtraction. 1 μL rRNA subtraction mix (3 ug/ul) and 2.5 μL 5x hybridization buffer (1 M NaCl, 500 mM Tris-HCl pH 7.5) were added to each reaction, and final volume adjusted with water to 12.5 μL. The samples were incubated at 68°C and the temperature was reduced by 1°C/min until the reaction was at 45°C. 5 μL RNase H buffer and 2 μL Hybridase thermostable RNase H (Lucigen) were added and water was added until the final volume was 40 μL. The samples were incubated at 45°C for 30 mins. The RNA was cleaned with RNA Clean and Concentrator −5, following the manufacturer’s instructions for recovery of fragments >200 nt and eluted in 45 μL water. 5 μL Turbo DNase buffer and 1 μL Turbo DNase (ThermoFisher Scientific) were added to each reaction and incubated for 30 mins at 37°C. 5.1 μL DNase inactivation reagent (ThermoFisher Scientific) was added and incubated 5 mins at room temp with intermittent manual mixing. The RNA was cleaned with RNA Clean and Concentrator −5 following instructions for recovery of fragments >200 nt and eluted in 15 μL water. For reverse transcription, 1 μL of RNA was added to 3.5 μL water, 2 μL 5x First Strand buffer (ThermoFisher Scientific), 1 μL 10μM reverse primer, 1 μL dNTP, 0.5 μL 0.1M DTT, 0.5 μL RNaseOUT, and 0.5 μL TGIRT-III (Ingex). The RT reaction was incubated at 57°C for 1.5 hours, followed by a 5 mins at 80°C. To degrade the RNA, 1 μL RNase H (New England Biolabs) was added to the RT reaction and incubated for 20 mins at 37°C. PCR was performed to amplify the samples using either Advantage HF 2 DNA polymerase (Takara) or Phusion (NEB) for 25–30 cycles according to manufacturer’s specifications. PCR product was purified by QIAquick PCR purification (Qiagen) and sequenced either on MISeq or iSeq100 (Illumina) to produce either 100nt single-end reads or 150×150nt paired-end reads.
Library Generation with DMS-modified RNA for HIV-1 genome RNA structure
10 μg extracted DMS-modified RNA from HEK293t transfected with NHG plasmid was split into 3 reactions for the first step of RNase H-based rRNA subtraction. The steps for RNase H and DNase treatment mentioned above were followed. After DNase treatment, the three reactions were eluted in 8.5 μL water and combined. An additional rRNA subtraction step was performed using the RiboZero Human/Mouse/Rat rRNA removal kit (Illumina; discontinued) according to manufacturer’s specifications. After RiboZero, the RNA was purified with RNA Clean and Concentrator −5, following the manufacturer’s instructions for recovery of fragments >200nt and eluted in 10 μL water. The RNA was fragmented using the RNA Fragmentation kit (ThermoFisher Scientific) with a fragmentation step of 45 seconds at 70°C. The RNA was purified with RNA Clean and Concentrator −5, following the manufacturer’s instructions for recovery of all fragments and eluted in 6.5 μL water. 1 μL CutSmart buffer (New England Biolabs), 1.5 μL Shrimp Alkaline Phosphatase (New England Biolabs) and 1 μL RNaseOUT (ThermoFisher Scientific) were added and incubated at 37°C for 1 hour to dephosphorylate the RNA. 6 μL 50% PEG-800 (New England Biolabs), 2.2 μL 10x T4 RNA Ligase buffer (New England Biolabs), 2 μL T4 RNA Ligase, truncated KQ (England Biolabs) and 1 μL linker were added to the reaction and incubated for 18 hours at 22°C. The RNA was purified with RNA Clean and Concentrator −5, following the manufacturer’s instructions for recovery of all fragments and eluted in 15 μL water. Excess linker was degraded by adding 2 μL 10x RecJ buffer (Lucigen), 1μL RecJ exonuclease (Lucigen), 1 μL 5’Deadenylase (New England Biolabs) and 1 μL RNaseOUT, then incubating for 1 hour at 30°C. The RNA was purified with RNA Clean and Concentrator −5, following the manufacturer’s instructions for recovery fragments > 200nt and eluted in 11 μL water. For reverse transcription, 1 μL RT primer, 1 μL 0.1M DTT, 4 μL 5x First Strand buffer, 1 μL dNTP, 1 μL RNaseOUT and 1 μL T-GIRT III were added and the sample was incubated for 2 hours at 65°C. RNA was degraded by adding 1 μL 4N NaOH and incubating at 95°C for 3 mins. The RT product was mixed with an equal volume 2x Novex TBE-Urea sample buffer (ThermoFisher Scientific) and run on a 10% TBE-Urea gel (ThermoFisher Scientific) and the ~300–400 nt product was extracted. The purified ssDNA was circularized using the CircLigase ssDNA Ligase kit (Lucigen). 2 uL of the circularized product was used for PCR using Phusion. The sample was run for a maximum of 14 cycles. Following PCR, the product was run on an 8% TBE gel and the ~350–450nt product was gel extracted. The final PCR product was quantified by Bioanalyzer (Agilent). The product was then sequenced by Novaseq S4 (Illumina) to produce 150×150nt paired-end reads. The same library generation protocol was followed for in vitro transcribed and DMS-modified U4/6 core-domain with some modifications. The starting amount was 250 ng of RNA as part of a pool of RNA totaling 4 μg. No fragmentation and no rRNA removal were performed.
HIV-1 Splice Junction Usage Analysis
Splice analysis was performed according to the previously written protocol24. Briefly, two separate RT-PCR reactions were performed with 2 μg total unmodified RNA from HEK293t cells transfected with plasmid containing HIV-1NHG, Δvpr HIV-1NHG, and HIV-1 mutants. One reaction was designed to reverse transcribe all HIV-1 multiply spliced products with a reverse primer that spans the D4A7 splice junction. The second reaction is designed to reverse transcribe HIV-1 singly spliced mRNA with a reverse primer lies in the env intron. The forward primer used in both PCR reactions is located upstream of D1. RT was performed with SuperScript III (Thermo Fisher Scientific) at 55°C for 1 hour followed by 15 minutes at 70°C. RNA was degraded by adding 1 μL RNase H and incubating at 37°C for 20 minutes. The cDNAs were then purified with Agencourt RNACleanX beads at a ratio of 2:1 (Beckman Coulter). Two successive rounds of PCR were used to add adapters for sequencing using the KAPA robust PCR kit (KAPA Biosystems). The first PCR uses with a forward primer that is located in the shared upstream D1 sequence that also has an adapter. The second round adds the universal adapter and Illumina indexed sequencing primers. The PCR products were then sequenced by Illumina Miseq, 300×300nt paired-end reads.
Statistical Methods
Statistical analysis of DREEM clusters was quantified by Pearson’s correlation. R2 and p-values of Pearson’s correlation are reported.
Library Linker and Primers
All oligos ordered from IDT.
StemA/StemC T7 forward primer: TAATACGACTCACTATAGAAAGGATCGG
StemA/StemC T7 reverse primer: ATCCCAGCGCGTGGTGCA
StemA/StemC RT primer: ATCCCAGCGCGTGGTGCA
StemA/StemC PCR forward primer: GAAAGGATCGGAAGACTCCACAG
StemA/StemC PCR reverse primer: ATCCCAGCGCGTGGTGCA
Add riboswitch T7 forward primer
TTCTAATACGACTCACTATAGGACACGACTCGAGTAGAGTCG
Add riboswitch forward primer: GACACGACTCGAGTAGAGTCG
Add riboswitch reverse primer: TGTTGGAGTCTACTCGACTCCGGT
HIV-1 RRE T7 forward primer: TAATACGACTCACTATAGGAGCTTTGTTCC
HIV-1 RRE T7 reverse primer: GGAGCTGTTGATCCTTTAGGTATCTTTC
HIV-1 RRE RT primer: GGAGCTGTTGATCCTTTAGGTATCTTTC
HIV-1 RRE PCR forward primer: GGAGCTTTGTTCCTTGGGTTCTTGG
HIV-1 RRE PCR reverse primer: GGAGCTGTTGATCCTTTAGGTATCTTTC
HIV-1 A3 PCR forward primer: TGAAACTTACGGGGATACTTGGGCAGGA
HIV-1NL4–3 A3 PCR and RT reverse primer: GAAGCTTGATGAGTCTGACTGTTCTGATGAGC
HIV-1NHG A3 PCR and RT reverse primer: CTTCGTCGCTGTCTCCGCTTCTTCC
To generate Δvpr HIV-1NHG
NL AgeF: AGC TAG AAC TGG CAG AAA ACA GGG AGA TTC
NL SalIR: CCA TTT CTT GCT CTC CTC TGT CGA GTA ACG C
dVprS: GGA AAC TGA CAG AGG ACA GAT GGA ATA AGC CCC AGA AGA CC
dVpr AS: GGT CTT CTG GGG CTT ATT CCA TCT GTC CTC TGT CAG TTT CC
To generate A3 Splice Site Mutants
NL 5599F: CATACAATGAATGGACACTAGAGCTTTTAG
NL BamHIR: CGTCCCAGATAAGTGCCAAGGATCCGTT
A3SLMut1 STCCATTTCAGAATTGGGTGTCGAGTAAGCCTAATAGGCGTTACTCGACAGAGGA
A3SLMut1 AS TCCTCTGTCGAGTAACGCCTATTAGGCTTACTCGACACCCAATTCTGAAATGGA
A3SLMut 2 S: GAATTGGGTGTCGACAACGCCTAATAGGCGTTACTCGAC
A3SLMut2 AS: GTCGAGTAACGCCTATTAGGCGTTGTCGACACCCAATTC
A3SLMut3 S: GGTGTCGACATAGCAGAATCTGCTATACTCGACAGAGGAGAGCAA
A3SLMut3 AS: GGTGTCGACATAGCAGAATCTGCTATACTCGACAGAGGAGAGCAA
A3SLMut4 S: TCAGAATTGGGTGTCGAAACAGCGAAATAGGCGTTACTCGACAGA
A3SLMut4 AS: TCTGTCGAGTAACGCCTATTTCGCTGTTTCGACACCCAATTCTGA
A3SLMut5 S TCAGAATTGGGTGTCGAAACAGCGAAATTCGCGTGTTTCGACAGAGGAGAGCAA
A3SLMut5 AS TTGCTCTCCTCTGTCGAAACACGCGAATTTCGCTGTTTCGACACCCAATTCTGA
Library generation linker /5rApp/TCNNNNNNNNNNNNAGATCGGAAGAGCGTCGTGTAGGGAAAGA/3ddC/
Library generation RT primer
/5Phos/AGATCGGAAGAGCACACGTCTGAACTCCAG/iSp18/TCTTTCCCTACACGACGCTCTTCCGATCT
Library generation forward PCR primer
CAAGCAGAAGACGGCATACGAGATXXXXXXGTGACTGGAGTTCAGACGTGTGCTC
Library generation reverse PCR primer
AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCTC
Splice Analysis, Multiply Spliced Reverse Primer
GTGACTGGAGTTCAGACGTGTGCTCTTCCGATCTNNNNNNNNNNNNNNCAGTTCG
GGATTGGGAGGTGGGTTGC
Splice Analysis, Singly Spliced Reverse Primer GTGACTGGAGTTCAGACGTGTGCTCTTCCGATCTNNNNNNNNNNNNNNGTACTATA
GGTTGCATTACATGTACTACTTAC
Splice Analysis, PCR Round 1 Forward Primer GCCTCCCTCGCGCCATCAGAGATGTGTATAAGAGACAGNNNNTGCTGAAGCGCGC
ACGGCAAG
Splice Analysis, PCR Round 2 Reverse Primer CAAGCAGAAGACGGCATACGAGATxrefGTGACTGGAGTTCAGACGTGTGCTC
Splice Analysis, PCR Round 2 Forward Primer
AATGATACGGCGACCACCGAGATCTACACGCCTCCCTCGCGCCATCAGAGATGTG
[Data Availability]
Sequencing data can be obtained from the GEO database using accession number GSE131506.
[Software and Code Availability]
Sequence alignment: Bowtie2 2.3.4.1. For code development: python v. 3.6.7. For read trimming: TrimGalore 0.4.1. For read quality assessment: FastQC v0.11.8. For RNA secondary structure analysis: RNAstructure v6.0.1. For calculating post-mapping statistics: Picard 2.18.7. RNA secondary structure visualization: VARNA v3.93. HIV-1 splicing analysis: https://github.com/SwanstromLab/SPLICING. Splice plot creation: R version 3.5.1. For figure construction: Adobe Illustrator CC 2019. For data analysis: Microsoft Excel 2018. Plot generation: Plotly v3.2.1. DREEM clustering algorithm is available at https://codeocean.com/capsule/0380995/tree
Extended Data
Extended Data Figure 1 |. DREEM clustering pipeline for DMS-MaPseq data.
a, A read x” is represented as a series of D bits, where D is the length of the read. A base is denoted by the bit ‘1’ if it is mutated away from the reference and by ‘0’, otherwise. Let K be the number of clusters in the sample. μk = {μk1,…, μkD} is the mutation profile of cluster k and πk is the mixing proportion of cluster k such that for k = 1 to K. The model parameters μ and π are randomly initialized. In the Expectation step, reads are assigned probabilistically to clusters and the likelihood of observing the data given the model parameters is computed. In the Maximization step, the mixing proportion is calculated from the reads assignments and the mutation profiles are updated for each cluster to maximize the expectation value of the complete case likelihood. The Expectation steps alternate with the Maximization steps until the likelihood converges. The likelihood function is derived using Bernoulli mixture models modified to account for missing data in the form of the underrepresentation of reads with adjacent mutations. b, Mutational distance distribution between bases in denatured, DMS modified, total RNA. Plotted is the mutation distance verses frequency between two DMS reactive positions i.e. A or C to A or C, yellow bars, and between one DMS reactive position and a background mutation (e.g. mutation due to sequencing error) i.e. A or C to T or G, blue bars. The blue bars demonstrate the frequency of observing two mutations due to background.
Extended Data Figure 2 |. DREEM clustering identifies and quantifies individual structures from in vitro mixing experiments.
Structure 1 and Structure 2 sequences were in vitro transcribed and re-folded, mixed in different proportions, and probed with DMS-MaPseq. The region used for DREEM clustering covers nucleotides 21–135 (labeled as 1–115 on the figure), which excludes the primers used for RT-PCR (that have no DMS-induced mutations) and is identical in sequence for the two structures except for the A>C mutation at position 94. Position 94 is masked during analysis. The topmost panel shows the DMS reactivity pattern of Structure 1 by itself and Structure 2 by itself. The rest of the panels show the clustering results at specified mixing ratios (n=1).
Extended Data Figure 3 |. Secondary structure models for V. vulnificus Adenoriboswitch (add).
a, Percentages for each cluster detected in the presence or absence of 5mM adenine to the add riboswitch. b, In vitro structure models obtained from probing add using DMS-MaPseq followed by DREEM, color coded by normalized DMS signal. The apoB and apoB alternative structures represent the OFF state, which is incompetent for ligand binding. The apoA represents the ON state. Previously identified helices are boxed and labeled.
Extended Data Figure 4 |. DREEM clustering reveals an equilibrium of 4-stem and 5-stem structures for in vitro folded HIV-1 RRE.
a, Population average DMS-MaPseq data for in vitro transcribed, refolded and DMS treated or untreated samples. b, Scatter plots showing the reproducibility of the DMS signal from DREEM clustering results between 2 replicates with different DMS modification conditions. Replicate 1 was modified in 0.25% DMS and replicate 2 was modified with 2.5% DMS. R2 is Pearson’s R2. c, DREEM clustering data from b was used as constrains to generate RNA structure models. Shown are the models derived for cluster 1 and cluster 2 from replicate 1, color coded by normalized DMS signal.
Extended Data Figure 5 |. HIV-1 RRE forms two stable alternative structures in CD4+ T cells.
a, Schematic representation of DMS treatment in primary cells and isolated virions. b, DMS-MaPseq probing of intracellular HIV-1NL4–3 RRE in CD4+ T cells was used as input for DREEM clustering. Two clusters passed the BIC test and were used as RNAstructure folding constraints. Structural models are color coded by normalized DMS reactivity and bases not covered by the region of PCR are colored in gray. Data used to construct models are representative data from n=2 biologically independent experiments.
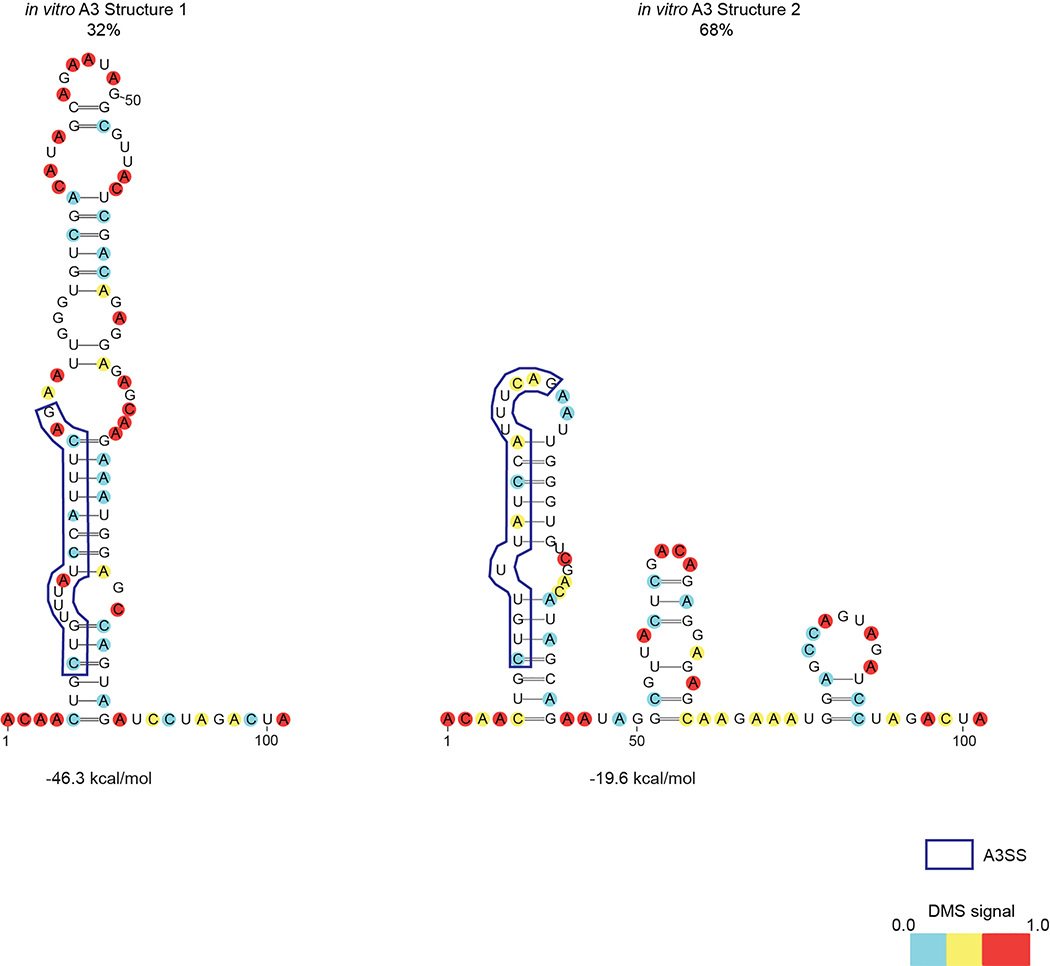
Extended Data Figure 6 |. A3 splice site forms alternative structures in vitro.
472nt A3 sequence from HIV-1NHG strain was in vitro transcribed, re-folded, and probed with DMS-MaPseq. DREEM clustering-based models for the local structures forming at A3 are shown, color coded by normalized DMS signal. Percentages of cluster 1 and 2 come from an n=1 experiment as determined by DREEM.
Extended Data Figure 7 |. Splice site usage in additional A3 mutants.
a, Structure models illustrating the mutant design for A3SLMut4 and A3SL Mut5. b, Splice usage for Mut4 and Mut5 for A1–5 reported as fold change compared to Δvpr HIV-1NHG. Central bar represents the mean and error bars indicate s.d. N=4 biologically independent experiments. c, Average fraction of transcripts using A3 compared to % cluster 1 (A3SL) as determined by DREEM (n=1) for A3SLMut1–5. Mutants are color coded. • indicates multiply spliced (MS) HIV-1 transcripts and ▲ indicates singly splice (SS) HIV-1 transcripts.
Extended Data Figure 8 |. Structural models of A3SLMut1 and A3SL Mut4.
a, Structural models for A3SLMut1 derived from n=1 experiment after DREEM clustering; pink box is the region of mutations; blue box is the splice site; Exonic Splicing Enhancer (ESE) and Exonic Splicing Silencer (ESS) binding sites are shown. b, Structural models made using DMS-MaPseq data from HEK293t cells transfected with vprHIV-1NHG A3SLMut4. The sequence of the A3 splice site is boxed in dark blue. The locations of the mutations are boxed in pink. Splice enhancer and suppressor binding sites are highlighted (ESS2p: purple, ESEtat: blue, ESE2: orange, ESS2: green). Percentages of each cluster come from an n=1 experiment.
Extended Data Figure 9 |. Genome-wide HIV-1NHG library generation quality control.
a, Coverage of HIV-1 genome with DMS-MaPseq data from HEK293t cells transfected with HIV-1NHG. b, Moving average of A and C mutational frequency in 100 nt windows after DMS-MaPseq compared to moving average T and G mutational frequency. c, DMS-MaPseq data from HEK293t cells transfected with HIV-1NHG was used as input for DREEM. Local 80nt window from Fig.4 for the RRE region was used for clustering. Percentages of cluster 1 and 2 come from an n=1 experiment. Nucleotides were color-coded based on normalized DMS signal; bases outside of the window used for clustering are colored in grey. d, The A3 splice site was analyzed using DMS-MaPseq and DREEM clustering from genome-wide data from HEK293t transfected with HIV-1NHG. Percentages of cluster 1 and 2 come from an n=1 experiment as determined by DREEM. Nucleotides were color coded with normalized DMS signal. e, A region of the HIV-1 genome in the pol coding region (nt 2000–2120 based on HIV-1NHG genomic RNA coordinates) was analyzed using DMS-MaPseq and DREEM clustering from genome-wide data from HEK293t transfected with HIV-1NHG. Two clusters passed the BIC in adjacent 80 nt windows that overlapped by 40 nt. The two 80 nt windows were combined to make the structural models. The range of proportions of each cluster come from the individual windows of n=1 experiment. Nucleotides were color coded with normalized DMS signal.
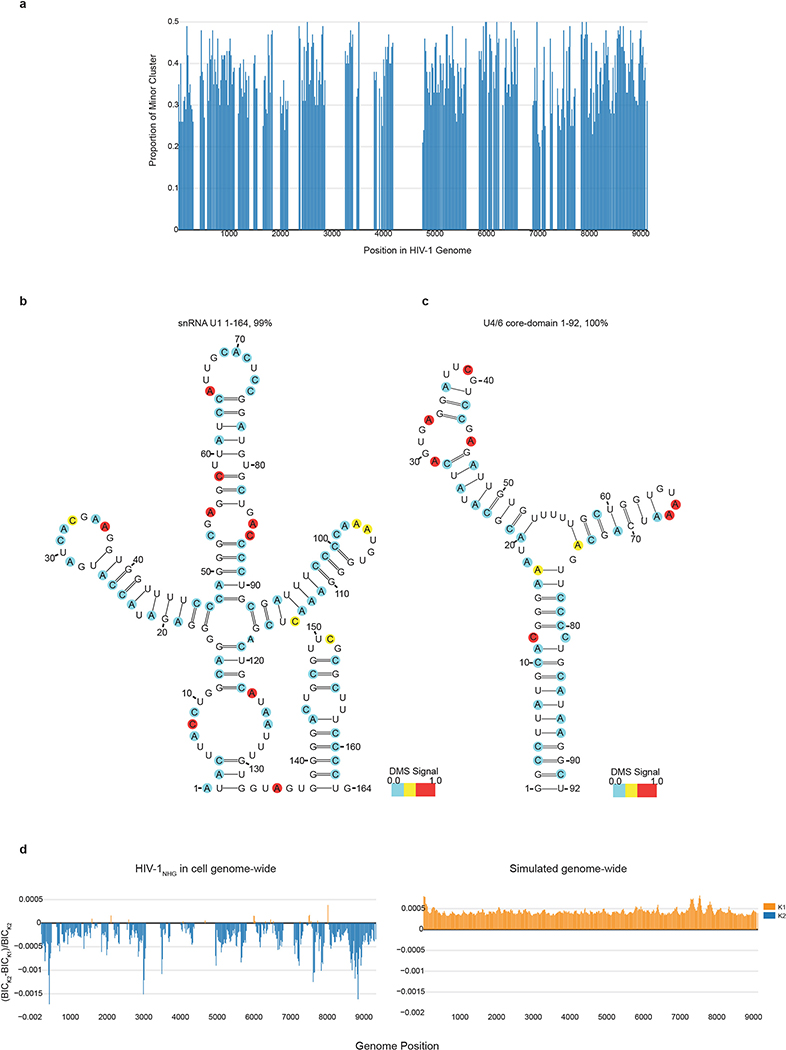
Extended Data Figure 10 |. Proportion of minor clusters across the HIV-1 genome and U1, U4/6 core-domain structural models.
a, Each bar shows the proportion of a minor cluster of an 80 nt window as a function of genome position for regions in the HIV-1NHG genome data set that are covered by at least 100,000 reads and pass 2 clusters according to the Bayesian Information Criterion test. b, U1 structural prediction from HEK293t cells transfected with HIV-1NHG. Abundance of cluster obtained from DREEM clustering. c, In vitro DMS-modified U4/6 core-domain RNA. Structure shown for population average, cluster 2 did not pass BIC. d, The left panel shows the difference in BIC test value between K=2 and K=1, normalized to the value for K=2 for the real whole-genome dataset. Each bar represents an 80 nt window across the HIV-1 genome. In orange are windows where only 1 cluster was detected according to the BIC test and in blue are clusters for which 2 clusters passed the BIC test. The right panel shows the same plot from simulated data for which the mutations were randomly distributed but had the same average number of mutations per read as the true data.
Extended Data Figure 11|. Shannon entropy across the HIV-1 genome and A4/5.
a, Overlay of the HIV-1NHG genomic organization on top of Shannon entropy plot. Each dot represents an 80 nt window in which Shannon entropy was calculated from DMS reactivity. The top plot is the major cluster and the bottom is the minor cluster. b, Scatter plot of Gini index versus Shannon entropy for the major and minor clusters (n=1). R2 is Pearson’s R2. c, Structural model of the TAR stem-loop from the genome-wide DMS-MaPseq and DREEM data. d, Structural model from 2 clusters found using the genome-wide DMS-MaPseq and DREEM data for a window containing 4 splice acceptor sites- A4a-c and A5. Splice sites are boxed. Nucleotides are color coded with normalized DMS signal.
Acknowledgments
The following reagents were obtained through the NIH AIDS Reagent Program, Division of AIDS, NIAID, NIH: Human rIL-2 from Dr. Maurice Gately, Hoffmann - La Roche Inc; HIV-1 NL4-3 Infectious Molecular Clone (pNL4-3) from Dr. Malcolm Martin (Cat# 114). This work was supported in part by the NIH (R21AI134365), the Center of HIV-1 RNA Studies (CRNA) NIH U54AI50470, the Smith Family Foundation, and the Burroughs Wellcome fund.
Footnotes
[Ethics declarations]
The authors declare no competing interests.
[References]
- 1.Purcell DF & Martin MA Alternative splicing of human immunodeficiency virus type 1 mRNA modulates viral protein expression, replication, and infectivity. J Virol 67, 6365–6378 (1993). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ocwieja KE et al. Dynamic regulation of HIV-1 mRNA populations analyzed by single-molecule enrichment and long-read sequencing. Nucleic Acids Res 40, 10345–10355, doi: 10.1093/nar/gks753 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Takata MA et al. Global synonymous mutagenesis identifies cis-acting RNA elements that regulate HIV-1 splicing and replication. PLoS Pathog 14, e1006824, doi: 10.1371/journal.ppat.1006824 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Watts JM et al. Architecture and secondary structure of an entire HIV-1 RNA genome. Nature 460, 711–716, doi: 10.1038/nature08237 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Warf MB & Berglund JA Role of RNA structure in regulating pre-mRNA splicing. Trends Biochem Sci 35, 169–178, doi: 10.1016/j.tibs.2009.10.004 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Shepard PJ & Hertel KJ Conserved RNA secondary structures promote alternative splicing. RNA 14, 1463–1469, doi: 10.1261/rna.1069408 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Singh NN, Lee BM & Singh RN Splicing regulation in spinal muscular atrophy by an RNA structure formed by long-distance interactions. Ann N Y Acad Sci 1341, 176–187, doi: 10.1111/nyas.12727 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Huthoff H & Berkhout B Two alternating structures of the HIV-1 leader RNA. RNA 7, 143–157, doi: 10.1017/s1355838201001881 (2001). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Abbink TE, Ooms M, Haasnoot PC & Berkhout B The HIV-1 leader RNA conformational switch regulates RNA dimerization but does not regulate mRNA translation. Biochemistry 44, 9058–9066, doi: 10.1021/bi0502588 (2005). [DOI] [PubMed] [Google Scholar]
- 10.Sherpa C, Rausch JW, Le Grice SF, Hammarskjold ML & Rekosh D The HIV-1 Rev response element (RRE) adopts alternative conformations that promote different rates of virus replication. Nucleic Acids Res 43, 4676–4686, doi: 10.1093/nar/gkv313 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Zubradt M et al. DMS-MaPseq for genome-wide or targeted RNA structure probing in vivo. Nat Methods 14, 75–82, doi: 10.1038/nmeth.4057 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Bishop CM Pattern Recognition and Machine Learning. Springer, New York, NY: (2006). [Google Scholar]
- 13.Reuter JS & Mathews DH RNAstructure: software for RNA secondary structure prediction and analysis. BMC Bioinformatics 11, 129, doi: 10.1186/1471-2105-11-129 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Spasic A, Assmann SM, Bevilacqua PC & Mathews DH Modeling RNA secondary structure folding ensembles using SHAPE mapping data. Nucleic Acids Res 46, 314–323, doi: 10.1093/nar/gkx1057 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Homan PJ et al. Single-molecule correlated chemical probing of RNA. Proc Natl Acad Sci U S A 111, 13858–13863, doi: 10.1073/pnas.1407306111 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Sengupta A, Rice GM & Weeks KM Single-molecule correlated chemical probing reveals large-scale structural communication in the ribosome and the mechanism of the antibiotic spectinomycin in living cells. PLoS Biol 17, e3000393, doi: 10.1371/journal.pbio.3000393 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Ding Y & Lawrence CE A statistical sampling algorithm for RNA secondary structure prediction. Nucleic Acids Res 31, 7280–7301, doi: 10.1093/nar/gkg938 (2003). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Halvorsen M, Martin JS, Broadaway S & Laederach A Disease-associated mutations that alter the RNA structural ensemble. PLoS Genet 6, e1001074, doi: 10.1371/journal.pgen.1001074 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Wan Y et al. Landscape and variation of RNA secondary structure across the human transcriptome. Nature 505, 706–709, doi: 10.1038/nature12946 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Tian S, Kladwang W & Das R Allosteric mechanism of the V. vulnificus adenine riboswitch resolved by four-dimensional chemical mapping. Elife 7, doi: 10.7554/eLife.29602 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lemay JF et al. Comparative study between transcriptionally- and translationally-acting adenine riboswitches reveals key differences in riboswitch regulatory mechanisms. PLoS Genet 7, e1001278, doi: 10.1371/journal.pgen.1001278 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Zaug AJ & Cech TR Analysis of the structure of Tetrahymena nuclear RNAs in vivo: telomerase RNA, the self-splicing rRNA intron, and U2 snRNA. RNA 1, 363–374 (1995). [PMC free article] [PubMed] [Google Scholar]
- 23.Emery A, Zhou S, Pollom E & Swanstrom R Characterizing HIV-1 Splicing by Using Next-Generation Sequencing. J Virol 91, doi: 10.1128/JVI.02515-16 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Schwartz G Estimating the dimension of a model. The Annals of Statistics 6, 461–464 (1978). [Google Scholar]
- 25.Rouskin S, Zubradt M, Washietl S, Kellis M & Weissman JS Genome-wide probing of RNA structure reveals active unfolding of mRNA structures in vivo. Nature 505, 701–705, doi: 10.1038/nature12894 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Liu Y et al. The roles of five conserved lentiviral RNA structures in HIV-1 replication. Virology 514, 1–8, doi: 10.1016/j.virol.2017.10.020 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Kondo Y, Oubridge C, van Roon AM & Nagai K Crystal structure of human U1 snRNP, a small nuclear ribonucleoprotein particle, reveals the mechanism of 5’ splice site recognition. Elife 4, doi: 10.7554/eLife.04986 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Cornilescu G et al. Structural Analysis of Multi-Helical RNAs by NMR-SAXS/WAXS: Application to the U4/U6 di-snRNA. J Mol Biol 428, 777–789, doi: 10.1016/j.jmb.2015.11.026 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Xiong HY et al. RNA splicing. The human splicing code reveals new insights into the genetic determinants of disease. Science 347, 1254806, doi: 10.1126/science.1254806 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Faustino NA & Cooper TA Pre-mRNA splicing and human disease. Genes Dev 17, 419–437, doi: 10.1101/gad.1048803 (2003). [DOI] [PubMed] [Google Scholar]
[Methods References]
- 31.Langmead B & Salzberg SL Fast gapped-read alignment with Bowtie 2. Nat Methods 9, 357–359, doi: 10.1038/nmeth.1923 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Darty K, Denise A & Ponty Y VARNA: Interactive drawing and editing of the RNA secondary structure. Bioinformatics 25, 1974–1975, doi: 10.1093/bioinformatics/btp250 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Adachi A et al. Production of acquired immunodeficiency syndrome-associated retrovirus in human and nonhuman cells transfected with an infectious molecular clone. J Virol 59, 284–291 (1986). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Lahm HW & Stein S Characterization of recombinant human IL-2 with micromethods. Journal of Chromatography 326, 357–361 (1985). [DOI] [PubMed] [Google Scholar]