Abstract
Acute myeloid leukemia (AML) is a complex, aggressive myeloid neoplasm characterized by frequent somatic mutations that influence different functional categories’ genes, resulting in maturational arrest and clonal expansion. AML can arise de novo (dn-AML) or can be secondary AML (s-AML) refers to a leukemic process which may arise from an antecedent hematologic disorder (AHD-AML), mostly from a myelodysplastic syndrome (MDS) or myeloproliferative neoplasm (MPN) or can be the result of an antecedent cytotoxic chemotherapy or radiation therapy (therapy-related AML, t-AML). Clinical and biological features in secondary and therapy-related AML are distinct from de novo AML. Secondary and therapy-related AML occurs mainly in the elderly population and responds worse to therapy with higher relapse rates due to resistance to cytotoxic chemotherapy. Over the last decade, advances in molecular genetics have disclosed the sub-clonal architecture of secondary and therapy-related AML. Recent investigations have revealed that cytogenetic abnormalities and underlying genetic aberrations (mutations) are likely to be significant factors dictating prognosis and critical impacts on treatment outcome. Secondary and therapy-related AML have a poorer outcome with adverse cytogenetic abnormalities and higher recurrences of unfavorable mutations compared to de novo AML. In this review, we present an overview of the clinical features of secondary and therapy-related AML and address the function of genetic mutations implicated in the pathogenesis of secondary leukemia. Detailed knowledge of the pathogenetic mechanisms gives an overview of new prognostic markers, including targetable mutations that will presumably lead to the designing and developing novel molecular targeted therapies for secondary and therapy-related AML. Despite significant advances in knowing the genetic aspect of secondary and therapy-related AML, its influence on the disease’s pathophysiology, standard treatment prospects have not significantly evolved during the past three decades. Thus, we conclude this review by summarizing the modern and developing treatment strategies in secondary and therapy-related acute myeloid leukemia.
Keywords: Secondary AML, therapy related AML, leukemia
Introduction
Acute myeloid leukemia (AML) is a complex disorder characterized by the development of malignant myeloid precursors in the blood and bone marrow [1]. AML is the most prevalent out of all acute leukemias diagnosed in adults and the second most common among all acute leukemias diagnosed in children [2]. This disease is delineated by a clonal proliferation of myeloid blasts, suppressing normal hematopoiesis and generally having a high fatality and a low cure rate, especially among older patients [3,4]. Genetic and molecular evidences suggest that AML is exceptionally varied entity with complicated repetitive genetic modifications such as chromosomal aberrations, DNA methylation patterns, and gene mutations [5-7]. These alterations accumulate and result in the blocking of normal maturation of myeloid progenitor cells and lead to clonal expansion of blast cells in the bone marrow. Despite novel breakthroughs in understanding of the molecular pathogenesis in AML still, the pathophysiology of leukemogenesis of myeloid blasts remains unclear [8,9]. Although most cases arise de novo, AML can also develop from an antecedent hematologic disorder such as myelodysplastic syndromes (MDS) or myeloproliferative neoplasms (MPN) or as a late complication of previous exposure to cytotoxic therapy including alkylating agents and topoisomerase II inhibitors typically used for a solid malignancy/lymphoma or radiation therapy for other cancers (therapy-related AML, t-AML). Secondary AML (s-AML) accounts for a large proportion of the AML population and is correlated with biological characteristics that contribute to poor outcomes, including unfavorable cytogenetics and multidrug resistance phenotype as compared with de novo AML (dn-AML) [3-7,10,11]. Genome-wide and targeted mutational analysis from next-generation sequencing has shown that the most commonly mutated genes were usually those that played a role in DNA methylation (46%), modifications of chromatin (42%), RAS signaling (42%), RNA spliceosome machinery (55%), transcriptional regulation (34%) and proteins that regulate the three-dimensional organization of chromatin in the nucleus (22%) [12]. These mutations may also help to predict the response of hypomethylating agents (HMA) and allogeneic hematopoietic stem cell transplantations (HSCT) to be used for the treatment of different patients with AML. Treating s-AML with traditional chemotherapy or hematopoietic stem cell transplantation (HSCT) is linked with poor prognosis [13-15]. HSCT refers to a medical procedure that infuses hematopoietic stem cells in order to restore bone marrow function in cancer patients whose bone marrow or immune system is damaged or defective. Nevertheless, HSCT has been proven a treatment option for patients with s-AML. It is extensively utilized in younger patients with adverse-risk cytogenetics [16-22]. HMAs such as decitabine and azacitidine are generally well-tolerated and helpful treatment options in patients with s-AML who are not eligible for intensive chemotherapy due to reduced cytotoxic profile. HMA exerts anti-tumor effects via an epigenetic modification to induce re-expression of tumor suppressor genes. The genetic mechanisms include DNA hypomethylation, chromatin reorganization, and DNA methyltransferase-I depletion. Treatment responses are seen after a few months of therapy. HMA is typically helpful in s-AML, where patients are usually elderly and frequently have TP53 alterations. They are also used to treat high-risk MDS and in AML post-transplant in high-risk patients. The median time to response is 3-4 cycles, and a minimum of 4-6 cycles should be given unless otherwise contraindicated. Median duration of response is 12-18 months. HMA is less toxic than intensive chemotherapy. The side effects seen are commonly related to myelosuppression, including infections and bleeding. Other side effects include nausea, vomiting, and occasional tumor lysis syndrome. Oral analogues have recently become available and used as maintenance therapy in AML patients after consolidation therapy. Of late, they are increasingly combined with other targeted agents, including FLT3 and IDH inhibitors. HMAs can be used in AML irrespective of the underlying genetic status. TP53 mutated AML does not behave worse compared to TP53 unmutated AML when treated with HMAs. Both Decitabine and azacitidine HMAs have proven efficacy, safety and confer an OS benefit in the absence of CR. The management of secondary AML (s-AML) is more complex compared to de novo AML as it is associated with worse clinical outcomes, less hematopoietic stem cell reserve, co-morbidities due to old age, and overall worse survival [23-28]. Hence, a better knowledge of the molecular landscape in secondary and therapy-related AML has significant implications on treatment response, prognostication, and the development of new molecular therapeutic targets. There is a necessity to introduce advanced personalized therapies for s-AML and t-AML [29]. In the light of diversified treatment options available for secondary and therapy-related AML, we also summarize the popular and developing treatment strategies in secondary and therapy-related AML.
Secondary acute myeloid leukemia (s-AML)
The majority of cases of AML are de novo AML without any antecedent hematologic disease or other cancer. Secondary leukemia can be further categorized into leukemia arising out of a prior myelodysplastic syndrome (MDS) or myeloproliferative neoplasm (MPN) or that originating from a bone marrow failure state (BMF). Secondary AML (s-AML) is a biologically complex heterogeneous disease that covers 25 to 30% of all AML cases [10,11]. About 20% of cases of AML are s-AML having an antecedent hematologic disorder (AHD-AML) involving progression from myelodysplastic syndromes (MDS), myeloproliferative neoplasms (MPN), or chronic myelomonocytic leukemia (CMML) as the antecedent disorders. Conventional prognostication depends on clinicopathological correlation and cytogenetic information. Additionally, secondary AML (s-AML) occurs more in the elderly population, responds worse to therapy, and displays worse prognosis and low response rates as compared to de novo AML [13,30,31]. Secondary AML is a distinguished unfavorable prognostic factor that confers distinct clinical and biological features characterized by adverse genetic mutations, cytogenetic alterations, and aberrations in TP53; leukemic clones are often more resistant to cytotoxic chemotherapy, contribute to a high relapse rate in s-AML [32-36]. In 2017 FDA authorized CPX-351 (liposomal formulation of cytarabine and daunorubicin), especially for secondary AML. However, current investigations show promising outcomes with high-dose cytarabine or hypomethylating agents [29]. Targeted immunotherapy is likely to evade multidrug resistance mechanisms but is unlikely to give continual remission due to a complicated and rapidly developing clonal disease profile [37]. Secondary AML is considered as an outcome of various somatic mutations that influence genes of different functional categories.
Secondary AML (s-AML), secondary to other myeloid neoplasms
Patients with s-AML developed as a result of an antecedent hematologic disorder (AHD-AML) have approximately 60% of the cases-MDS, 30% of the cases-MPN, and about 10% of the cases-CMML [30]. The development of s-AML is defined by the progressive evolution of clones [38].
Secondary AML (s-AML) Arising from Myelodysplastic Syndrome (MDS)
Secondary AML results from an antecedent hematologic condition (AHD) or previous chemotherapy/radiation therapy with distinct genomic alterations. The most prevalent cause of s-AML is myelodysplastic syndrome (MDS), a group of diverse clonal abnormalities of the hematopoietic stem cells (HSCs) characterized by morphologic dysplasia in the hematopoiesis, and bone marrow failure, resulting in cytopenias and a significant risk of evolving into secondary acute myeloid leukemia (s-AML). Nearly one-third of MDS patients progress to s-AML. MDS and its evolution into s-AML are correlated with the accumulation of various genetic mutations particularly, RUNX1 [30,39-44]. The International Prognostic Scoring System is used to evaluate the prognosis of MDS [45]. The median time to develop s-AML from MDS is about 17 months, while from a precursor MPN, it’s about 43 months. AML resulting from an antecedent MPN displays a lower response to chemotherapy and poorer overall survival (OS) than AML from antecedent MDS [30]. MDS is divided into five prognostic subgroups based on the development of dysplasia, blast counts percentage, and specific cytogenetic abnormalities. A combination of cytogenetics and clinical parameters classifies patients into prognostic risk categories using a Revised-International Prognostic Scoring System (IPSS-R), with a variable uncertainty of disease progression [46,47]. In higher-risk groups, a substantial overlap of cytogenetic aberrations and mutations was observed between MDS and s-AML [48-50]. Poor prognosis is correlated with a complex karyotype (CK) which significantly raises the risk of progression to s-AML [47,48]. Cytogenetic abnormalities linked with MDS and s-AML are usually due to copy number alterations, which are more prevalent in dn-AML. Many genetic studies have led us to classify mutations common in MDS and s-AMLs into the following main groups: spliceosome genes, epigenetic modifiers, transcription factors, and activated signaling genes [12].
Mutations in spliceosome genes (e.g., SRSF2, SF3B1, U2AF1, EZH2) are more common in AML resulting from MDS compared to dn-AMLs [51,52]. Mutations in epigenetic modifiers (e.g., TET2, DNMT3A, ASXL1) are well-known in multiple clonal hematopoiesis (e.g., CHIP, MDS, s-AML) [53]. This suggests that these mutations occur early in the pathogenesis of MDS before the development of s-AML. While mutations in transcription factors (e.g., RUNX1, GATA2, CEBPA) and signaling pathway genes (e.g., RAS, FLT3) are more frequent in s-AML, implying that these mutations are acquired later during progression to s-AML [54,55]. Mutational variation in s-AMLs and dn-AMLs is given in Figure 1. However, whole-genome sequencing reports suggest that the intrinsic pathophysiology of s-AML progression includes complex stages of clonal evolution [56]. An extensive population-based study has concluded that the risk of transformation is driven by a single genetic or epigenetic event [57].
Figure 1.
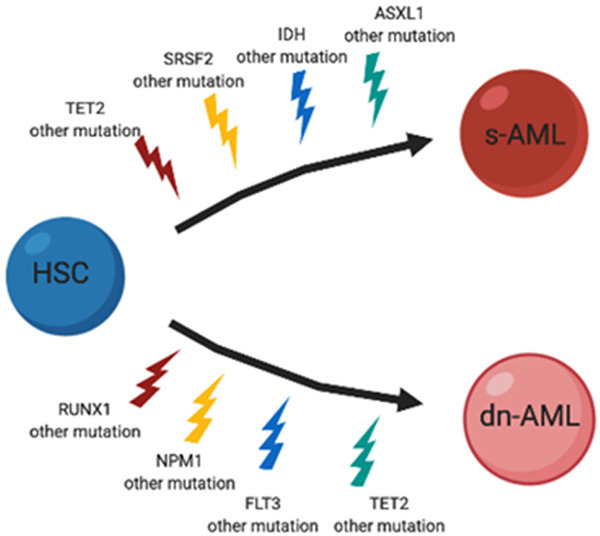
Mutational variation in de-novo and secondary acute myeloid Leukemia. HSC: Hematopoetic Stem cell, s-AML: Secondary acute myeloid leukemia, dn-AML: de novo acute myeloid leukemia.
Secondary AML (s-AML) arising from myeloproliferative neoplasms (MPN)
The myeloproliferative neoplasms (MPN) are a group of chronic clonal proliferative disorders of the myeloid series of cells. The development of MPNs in most cases is driven by various mutations, including myeloproliferative leukemia protein (MPL), Janus Kinase 2 gene (JAK2), and calreticulin (CALR) by activating the JAK2-STAT signaling pathway [58-65]. Multiple risk factors, including decreased platelet count, anemia, increasing age, circulating blast forms, have been proposed to prognosticate the risk of leukemic progression from a myeloproliferative neoplasm [66]. Chromosomal aberrations like del7p were found in cases of MPN linked with progression to secondary AML (s-AML). Furthermore, an adverse karyotype indicates a six-fold higher risk of progression to the MPN-Blast Phase [49,67]. Exposure to one or more cytoreductive treatments was seen to have enhanced the risk of progression to AML [68]. Various mutations, including SRSF2, IDH1, or IDH2, were exhibited as independent risk factors for leukemic alteration in MPNs [69,70]. Heterozygous TP53 mutation is usually found early in MPN, but evolution to leukemia correlates with the transition of TP53 from heterozygous to homozygous mutated form [71]. Post-MPN AML is linked with a poorer prognosis [72]. TP53, SRSF2, and TET2 mutations have been seen to be associated with a poor overall survival rate in secondary AML arising from MPN [73,74].
Therapy-related acute myeloid leukemia (t-AML)
Therapy-related AML in most events emerges after cytotoxic chemotherapy following treatment of other malignancies, including lymphoma, breast cancer, multiple myeloma, ovarian cancer, and testicular cancer, or may also begin after autoimmune disorders due to the mutagenic treatment agents [30-33,36,75,76]. Two common types of t-AML have previously been recognized. One type is seen after therapy with alkylating agents (e.g., cyclophosphamide, melphalan, etc.) and presents typically as therapy-related myelodysplasia with deletions or loss of chr 5 or 7. Alkylating agents begin cell necrosis by crosslinking in DNA, followed by mutations and chromosomal rearrangements [63,77-79,95]. The other type is seen after therapy with topoisomerase II inhibitors (e.g., doxorubicin, etoposide, etc.) and characteristically manifests as overt t-AML with repeated balanced chromosome abnormalities. The leukemogenic mechanism of topoisomerase II inhibitors depends on their involvement with DNA re-ligation and chromosomal damage. Topoisomerase II inhibitors prevent the DNA resealing step and inducing double-strand breaks [80]. Chromosomal abnormalities generally correlated with topoisomerase II inhibitors are balanced translocations, most usually including MLL (e.g., KMT2A) at 11q23 and RUNX1 at 21q22 [47,48,81,82]. Novel investigation intimates that these 2 common types of t-AML can now be subdivided into 8 genetic pathways with diverse etiology and different biologic characteristics, as shown in (Table 1).
Table 1.
Genetic pathways involved in pathogenesis of therapy related acute myeloid leukemia
| Pathways | Type of therapy for primary malignancy | Primary abnormality | Secondary abnormality | Ref. |
|---|---|---|---|---|
| I | Alkylating agent | del 7q/loss of chr. 7 | t(3;21), Mutation in RAS gene | [85,86] |
| II | Alkylating agent | del 5q/loss of chr. 5 cane be associated with a chr. 7 aberration | Mutation of TP53 gene, trisomy 8 | [87,88] |
| III | Epipodophyllotoxin | Balanced translocation of chr. 11q23 | TP53 gene mutation | [89-91] |
| IV | Topoisomerase II Inhibitors | t(8;16), t(8;21) and inv(16) | Additional aberrations | [93] |
| V | Topoisomerase II Inhibitors | PML-RARα fusion | Mutation in FLT3 gene | [94] |
| VI | Topoisomerase II Inhibitors | Fusion of NUP98 & HOXChr. (11p15q22) | Absent | [95,96] |
| VII | Not associated with any specific therapy | Normal karyotype | FLT3, NPM1 and MLL gene | [85,97] |
| VIII | Not associated with any specific therapy | Other aberrations | - | [98] |
Pathway I
This pathway is delineated by the loss of chromosome 7 or monosomy 7 or other chromosomal abnormalities in chromosome 7 [83-85]. These aberrations further cause secondary abnormalities such as balanced t(3; 21) [85]. This pathway is closely correlated to therapy with alkylating agents for treating any primary malignancy. It is also characterized by mutations of the rat sarcoma (RAS) pathway, which leads to aberrant proliferation of myeloid hematopoietic stem cells through activation of the signal transduction pathway [86].
Pathway II
This pathway is characterized by the loss of the long arm of chromosome 5 or monosomy 5 [87]. This may also; in addition, be associated with chromosomal aberrations in chromosome 7. Further, secondary aberrations such as mutation of TP53, trisomy 8 can occur [88]. These result in impaired differentiation and enhanced expression of cell cycle regulatory proteins due to the loss of the tumor suppressor gene [89]. This pathway is closely linked to therapy with alkylating agents for treating any primary malignancy.
Pathway III
This pathway is characterized by balanced translocation of chromosome 11q23 [89]. This aberration further causes secondary abnormalities like mutations of the TP53 gene. This pathway is firmly associated with therapy with epipodophyllotoxin for treating any primary malignancy. This mutation is most commonly seen in infants [90,91]. Patients with this mutation usually achieve a complete remission following intense antileukemic treatment, but the prognosis is terrible due to multiple relapses [91,92].
Pathway IV
This pathway is characterized by balanced translocation, including t(8;16), t(8;21), and inv(16). They are usually associated with many additional aberrations. This pathway is closely linked with therapy with topoisomerase II inhibitors for treating any primary malignancy [93]. Patients in this pathway display an excellent response to intensive antileukemic chemotherapy.
Pathway V
This pathway is characterized by chimeric rearrangement between the PML and the RARα genes. This aberration further causes secondary anomalies like mutations of the FLT3 gene. This pathway is closely correlated to therapy with topoisomerase II inhibitors for treating any primary malignancy. Patients with this pathway usually respond favorably to intense antileukemic chemotherapy when used with retinoic acid [94].
Pathway VI
This pathway is characterized by the Fusion of NUP98 and HOX-Chr.11p15q22. This pathway is closely associated with therapy with topoisomerase II inhibitors for treating any primary malignancy [95,96].
Pathway VII
This pathway is associated with normal karyotype and associated with various mutations like FLT3 and NPM1 mutations [85,97].
Pathway VIII
This pathway is uncharacteristic. They may represent incidentally identified cases of dn-AML [98].
Therapy-related acute myeloid leukemia (t-AML) following chemotherapy and radiation
In 5-10% of cases, patients present with therapy-related AML (t-AML) due to antecedent cytotoxic therapeutics or radiation treatment for a previous malignant or non-malignant disorder [99]. Since radiation therapy is typically administered along with chemotherapy as a part of multi-modality treatment, comparable to alkylating agents, radiation therapy induces double-stranded DNA breaks [66]. Consequently, most incidents of t-AML affect the proliferation of a myeloid clone with genetic variability influenced by previous chemotherapy and radiation. Chemotherapy may build an environment that allows proto-leukemia cells to harbor mutations and develop uncontrolled proliferation from local niche cells [100-102].
Mutational landscape of secondary and therapy related acute myeloid leukemia
The pathophysiology of s-AML and t-AML and its progression involve multiple genetic aberrations. The development of myeloid malignancies is triggered by somatic mutations resulting in the clonal extension and maturational arrest of HSCs. Advanced genome-wide and targeted analyses by microarrays and next-generation sequencing (NGS) have distinguished novel mutations of prognostic and therapeutic importance responsible for the development of AML. In 2008, AML was the first disease for which the entire cancer genome was sequenced. The genetic heterogeneity in AML is vast and includes structural chromosomal aberrations as well as point mutations. Conventional cytogenetics, somatic point mutations, and smaller copy number alterations plays a critical role in the pathogenesis of disease. Various unfavorable mutations were present in patients with s-AML that promote clonal development and disease progression [6,10,103-111]. The number of driver mutations in s-AML resulting from an AHD is more when compared to t-AML [112]. AML displays a lower number of coding mutations per exome at the molecular level compared to other cancers. The mutational landscape in s-AML and t-AML can be classified according to various functional categories, and the genes involved in each of the categories are: Epigenetic regulation (TET2, DNMT3A, IDH1/IDH2, EZH2, ASXL1), RNA spliceosome machinery regulators (SRSF2, SF3B1, U2AF1), Transcription regulator (TP53, RUNX1) and Signaling pathway regulators (FLT3) correlate to a mechanistic role in leukemogenesis [6,113-116]. These mutation groups correlate with clinical outcomes, underlying disease progression, and predict responses to therapy. There is a substantial overlap in the genomic landscape of the s-AML resulting from an AHD-like precursor MDS, MPN, and t-AML [10]. But, on the other hand, several specific mutations can also prove to be possible therapeutic targets [31,32]. Hence, the molecular landscape in secondary and therapy-related AML has significant involvement in prognostication, therapeutic response, and different molecular therapeutic targeting.
Targeting mutations in epigenetic regulator genes (TET2, DNMT3A, IDH1, IDH2, EZH2, ASXL1)
The epigenetic modifications lead to a change in the gene expression without actually changing the DNA sequence. Abnormal methylation of DNA, histone modifications, and RNA-mediated silencing lead to dysregulation of gene expression promotes the pathogenesis of myeloid malignancies [117,118]. In cancer, DNA hypermethylation probably leads to chromosomal instability & promoter hypermethylation has been correlated with silencing of tumor suppressor genes [119]. Genome-wide DNA methylation profiling in AML has displayed different patterns in various genetic subgroups that can categorize AML into prognostic groups [119-121]. In AMLs, epigenetic mutations, chromosomal aberrations, and somatic mutations are closely associated. Abnormal DNA methylation is a crucial mechanism to alter epigenetic regulations in secondary and therapy-related AML [11]. Commonly mutated genes related to this pathway include TET2, DNMT3A, IDH1 & IDH2, EZH2, and ASXL1.
TET2 mutations
The TET gene produces TET1, TET2, and TET3 dioxygenase proteins. TET2 is located on chromosome 4q24. TET1/2, which catalyzes the conversion of 5-methylcytosine to 5-hydroxymethylcytosine (5-hmc), likely influences the epigenetic regulation of transcription factors and thereby has a crucial function in the maintenance of hematopoietic stem cell (HSC) homeostasis [122-124]. Moreover, TET2 mutations involve frameshift, nonsense, and missense mutations seen in a diversity of myeloid neoplasms, including s-AML, MDS, and MPNs [43,123,125-127]. A higher incidence of TET2 mutations is observed in patients with s-AML (24%-32%), but these mutations occur at a lower frequency of about 20%-25% in MDS [128-131]. TET2 mutations confer a higher survival in MDS but a poorer survival in the case of AML and chronic myelomonocytic leukemia (CMML) [132-134]. Moreover, TET2 mutations may predict a more beneficial response to HMAs in patients carrying these mutations [42,110,124].
DNMT3A Mutations
The de novo methyltransferase 3 alpha (DNMT3A) belongs to the DNA methyltransferases family, which has other members, including DNMT1 and DNMT3B, which play an essential role to catalyze DNA methylation by transfer of methyl groups to specific cytosine residues of CpG islands [135-137]. They are involved in differentiation, proliferation, and apoptosis [138]. Aberrant methylation of CpG island promoters of tumor suppressor genes is an important pathogenetic mechanism for the development of many malignant tumors and suggests that DNMTs play a significant role in oncogenesis. DNMT3A mutations are associated with age, white blood cell (WBC) count, cytogenetics, prognosis, and patient’s response to chemotherapy [148]. DNMT3A is essential in normal hematopoietic stem cell differentiation and self-renewal and is highly expressed in lymphocytes [139]. DNMT3A mutations are present in 30%-35% of AML patients with normal karyotype and in approximately 10% of MDS patients [140,144,145]. The majority of DNMT3A missense mutations were found in hematological cancers at amino acid residue R882 near the carboxyl terminus of the DNMT3A protein [5,140-143]. DNMT3A R882H the mutation seems to predict a worse prognosis, has a lower overall survival (OS) rate and has higher leukemic transformation [146,147].
IDH1 AND IDH2 mutations
Isocitrate dehydrogenases 1 and 2 (IDH1 and IDH2) belong to the family of epigenetic modifiers that transform isocitrate to α-ketoglutarate (α-KG) [144,145]. Missense mutations of IDH1/2 associated with blood tumors frequently occur at specific arginine residues R132 in IDH1 and R140 or R172 in IDH2 [149]. These mutations generate D-2-hydroxyglutarate (2-HG), which is structurally similar to α-KG in that it can bind in place of α-KG and inhibits its function [150]. As a result, the overproduction of D-2-HG inhibits the α-ketoglutarate-dependent dioxygenase enzymes JumonjiC (JmjC), TET2, and cytochrome C oxidase (COX), leading to DNA hypermethylation and can block cell differentiation. These enzymes intervene with cellular metabolism, and this epigenetic regulation promotes oncogenesis [144-146,151]. In myeloid malignancies, IDH1/2 mutations can be observed in patients with cytogenetically normal AML, MDS, and MPN [144,152]. IDH1 mutations at residues R132 are found in 6%-16% of dn-AML and are associated with inferior OS rate [153,154]. Patients with IDH2 R172K mutation have an inferior prognosis, a higher relapse rate, a lower complete remission rate, and bad OS [155,156]. IDH2 mutations are seen in 4% to 12% of MDS cases and are often present together with ASXL1, SRSF2, and DNMT3A mutations and prognosticate lower OS, particularly those at codon 172 [153,157-160]. Additionally, some investigations propose that IDH1 and IDH2 mutations contribute to the progression from MDS or MPN to AML through reactive oxygen species (ROS) accumulation and DNA damage influencing the activation of HIF-1 [161-163]. IDH1/2-mutated myeloid malignancies are likely targetable by obstructing mutant enzymatic activity and their metabolic pathways [164].
EZH2 mutations
Mutations in the enhancer of zeste homolog 2 (EZH2) gene is linked with hematologic malignancies. EZH2, a histone-lysine N-methyltransferase enzyme, is responsible for the addition of methyl groups to a highly conserved histone H3 at lysine 27 (H3K27) that regulates stem cell differentiation and maturation by epigenetic modifications in the genes linked with transcriptional repression [165]. EZH2 mutations have been associated with many hematopoietic malignancies, most commonly in myelodysplastic syndromes (MDS) and myeloproliferative neoplasms (MPN) [166,167]. EZH2 mutations are observed in 6%-12% of patients with MDS, and these mutations correlate with a lower survival rate. EZH2 mutations usually co-exist with TET2 mutations, linked with a higher risk of developing secondary AML [145].
ASXL1 mutations
Additional sex comb-like 1 (ASXL1) gene is one of the most commonly mutated genes in a variety of myeloid malignancies, including in CMML (43%), MDS (20%), MPN (10%), and AML (20%) [174,175]. The ASXL1 protein is implicated in epigenetic modulation and transcription by interaction with polycomb repressive complex PRC2 and different transcriptional activators or repressors [168-171]. ASXL1 gene is present on chromosome region 20q11.21, and it controls histone modification by interacting with the PRC2. PRC2, along with ASXL1 methylates histone protein (H3K27), is a crucial regulator in malignant myeloid diseases and hematopoiesis [172,173]. ASXL1 deprivation is linked with hematopoietic alteration and enhanced self-renewal [145]. Simultaneously, the mutated ASXL1 indicates a poor prognosis for AML and MDS patients, resulting in an inferior OS rate linked to a higher risk of secondary AML [176,177].
Targeting mutations in the RNA spliceosome machinery genes (SRSF2, SF3B1, U2AF1)
RNA spliceosome machinery genes code for the spliceosome components, including small nuclear RNAs (sn-RNA) and proteins that catalyze the process of splicing by which mature RNA is developed from pre-mRNA through intron replacement and exon splicing. Components of pre-mRNA splicing machinery are generally mutated in myeloid malignancies, including SRSF2, SF3B1, and U2AF1 [94,178].
SRSF2 mutations
The Serine/Arginine-rich Splicing Factor 2 (SRSF2) gene is positioned on chromosome 17q25.1 and encodes pre-mRNA, which is required for spliceosome assembly, and alternative splicing [160,178]. SRSF2 mutations may lead to disease progression in patients with MDS [179]. SRSF2 mutations are observed in 11%-15% of patients with MDS, it has been seen that SRSF2 mutations generally co-exist with RUNX1, ASXL1, IDH1, and IDH2 mutations. In CMML, SRSF2 mutation is seen in nearly 28-47% of cases, closely associated with TET2 mutations [110,178]. SRSF2 mutations are linked with more inferior OS and an enhanced risk for AML progression [178,180,181].
SF3B1 mutations
The Splicing Factor 3B Subunit 1 (SF3B1) gene (chromosome 2q33.1) encodes the splicing factor 3B crucial for forming a ribonucleoprotein complex known as U2 snRNP [182]. Clonal and in vitro investigations of human leukemia cells have revealed that SF3B1 mutations initiate events that occur in rare lymphoid-myeloid hematopoietic stem cells (HSCs) [183,184]. SF3B1 mutations have been observed in 6%-18% of patients with MDS [185]. SF3B1 mutations are linked with cytopenias, thrombocytosis, and lower blasts percentage in MDS-Ring Sideroblast (RS) or MDS/MPN-RS. SF3B1 mutation is an indicator of good prognosis and prolonged survival [186].
U2AF1 mutations
The U2-complex auxiliary factor 1 (U2AF1) gene, located on chromosome 21q22.3, encodes the U2 spliceosome that promotes dinucleotide recognition in pre-mRNA introns. Repetitive mutations in U2AF1 at codon Q157 and S34 have been observed in 11% of cases with MDS and approximately 4% of patients with AML [178,187]. U2AF1 mutations affect the generation of neomorphic phenotypes by changing splicing patterns for multiple RNA downstream genes. U2AF1 modifications seem to affect erythroid progenitors’ division and consequently alter the differentiation of leukocytes [188]. The prognostic impact of U2AF1 mutations remains with a lower remission rate and poorer survival linked to an increased risk of AML transformation [178].
Targeting mutations in transcriptional regulatory genes (TP53, RUNX1)
Transcriptional regulatory genes are usually involved in leukemic progression across secondary and therapy-related AML includes TP53, RUNX1.
TP53 mutation
Tumor protein 53 (TP53) is a tumor suppressor gene found on chromosome 17p13.1. TP53 controls apoptosis, senescence, cell cycle arrest, and DNA repair [189]. More than 70% of patients have been seen to harbor TP53 mutations in t-AML cases [12,190,191]. TP53 gene mutation may benefit the malignant clone that expedites the development of a malignant tumor [192,193]. The vast majority of TP53 mutations arise in two mutational hot spots: residues R273 and R248 encode the DNA-binding domain [192,194]. TP53 mutations are linked with a high risk of leukemic transformation, resistance to chemotherapy, inferior OS, and DFS [110,192,194].
RUNX1 mutations
Runt-related transcription factor 1 (RUNX1), also known as acute myeloid leukemia 1 protein (AML1) or core-binding factor subunit alpha-2 (CBFA2) found on chromosome 21q22.12, is a crucial regulator of myeloid differentiation that strongly drives leukemic transformation [195]. Co-expression of RUNX1 and ASXL1 mutations contribute to leukemic change by preventing myeloid stem cells’ differentiation and improving self-renewal activity by transcriptional activation of hypoxia-inducible factor 1 (HIF1-α) [198,199]. RUNX1 mutations are observed in 24% of patients with secondary AML and independently confer inferior prognosis [110,196,197].
Targeting mutations in activated signaling pathway
Tyrosine kinase and RAS pathways have been seen to get affected in s-AML [119]. FLT3 is the most commonly mutated gene associated with this pathway [204].
FLT3 mutation
Fms-like tyrosine kinase-3 (FLT3) is a receptor tyrosine kinase highly expressed in hematopoietic stem cells [200-203]. The activation of the FLT3-mediated signaling accelerates PI3K/AKT and RAS/MAPK pathways, which leads to an uncontrolled proliferation of the hematopoietic stem cells (HSCs). FLT3 ITD mutation, which is observed in about 30% of AML cases, is linked with higher relapse rates and poorer survival [200,204].
Treatment strategies and novel therapeutics approach in secondary and therapy related AML
Secondary AML is a complex disease with high incidence and mortality. The standard of care for the treatment of s-AML and t-AML has remained consistent for decades. Hence, novel treatment strategies are needed to combat s-AML [29]. The therapeutic armamentarium of AML has progressively evolved in the past several years due to recent advancements in sequencing technologies and experimentation methodologies that improve our knowledge insight of the functioning of multiple genomic abnormalities. These genomic abnormalities play an impactful role in the pathogenesis of s-AML and t-AML as well as on mechanisms facilitating resistance to conventional therapies [205,206]. With plenty of novel therapeutic modalities available, the patient’s treatment will depend on a specific combination of the personalized genomics. s-AML has generally been treated with intense chemotherapy 7 + 3 regimens, comprised of 7 days consecutive intravenous cytarabine infusions and 3-day anthracycline infusions (i e, daunorubicin or idarubicin) is considered a standard of care for induction which significantly diminishing the leukemia burden in patients. Complete remission (CR) varies between 30 to 60%, and overall survival (OS) is nearly nine months for cases with s-AML treated when treated with standard 7 + 3 induction chemotherapy with cytarabine and an anthracycline [30,207-209]. Usually, older patients diagnosed with s-AML are treated with low-dose cytarabine (LDAC) or treatment with a hypomethylating agent (HMA) such as 5-azacitidine, which can influence remission and increase OS [210-212]. Despite the effectiveness of chemotherapy regimens, the prognosis of patients with sAML remains poor, particularly for older patients. Allogeneic hematopoietic stem cell transplant (allo-HSCT) is a potential therapeutic treatment and possibly offers the most promising long-term remission in s-AML [213,214]. Due to the absence of sustained response to traditional induction chemotherapy, patients with s-AML who attain CR, belonging to intermediate or adverse risk groups, will be referred for allo-HSCT [215]. In preclinical investigations, several novel agents in combination with conventional chemotherapeutic medications have been conferred to overcome drug resistance in AML [216-218]. A few pathway-specific agents and chemotherapy with a new drug delivery system being approved by the US FDA to treat AML, including CPX-351, venetoclax, gemtuzumab ozogamicin (GO), glasdegib, and various agents targeting FLT3 or IDH such as midostaurin, gilteritinib, enasidenib, ivosidenib. A few of these new drugs have been on trial s-AML cases. A p-glycoprotein expression is responsible for poor outcomes with anthracycline-based regimens in patients with s-AML [219]. CPX351 therapy with a cytarabine and daunorubicin liposomal formulation has been considered to enhanced overall survival (OS) in s-AML and overcome resistance mechanisms such as p-glycoprotein efflux [220,221]. CPX-351, a dual-drug liposomal co-encapsulation of cytarabine and daunorubicin, has been approved to treat patients diagnosed with s-AML and t-AML. A nanoliposomal carrier of the drug improves intracellular uptake of the drug [222-224]. In contrast with the traditional 7 + 3 regimen, CPX-351 medication was linked with significantly increased OS, EFS, CR in newly diagnosed s-AML patients [225,226]. Regimens with high-dose cytarabine (HiDAC) have shown an added advantage in cases with s-AML [227]. AML cells typically depend on anti-apoptotic Bcl-2. Bcl-2 is usually overexpressed in AML and associated with cell survival, chemoresistance, and lower OS in AML patients [228]. BCL-2 inhibitor Venetoclax, combined with hypomethylating agents such as azacitidine, decitabine exhibit promising alternative therapeutic medications for high-risks s-AMLs that may be preferred in older patients who cannot tolerate the intense and likely critically unfavorable consequences of intensive chemotherapy [229,230]. HMA exerts anti-tumor outcomes primarily through an epigenetic alteration to facilitate tumor suppressor genes re-expression [231-236]. Therefore, venetoclax, a small-molecule inhibitor of Bcl-2, mainly targets AML cells, leading to cell death [237,238]. CD33 antigen is a transmembrane receptor, and blasts express myeloid differentiation marker in most cases with AML. Using the benefit of the particular expression of CD33 by blasts, Gemtuzumab ozogamicin (GO) was developed as a conjugate of an anti-CD33 antibody. After binding to the antigen on the surface of leukemic blasts, the antibody is internalized and binds to DNA leading to double-strand break and cell death [239]. In a randomized phase III trial, GO in combination with daunorubicin and cytarabine has improved EFS, OS, and RFS in s-AML cases [240-244]. The hedgehog pathway performs a crucial function in developing leukemic stem cells and procuring a drug-resistant phenotype in AML. Glasdegib (Daurismo™, Pfizer) drug acts by repressing the hedgehog pathway in leukemic stem cells. In phase 2 multicenter randomized investigation, the combination of glasdegib with low dose cytarabine (LDAC) increased OS. It attained a higher CR rate than LDAC alone in newly diagnosed AML patients who were unfit for intense chemotherapy [245]. This led to its approval by US FDA in 2018 as a treatment for adults with newly diagnosed AML [246,247]. Various drugs that can repress FLT3 kinases have been tested in AML. In April 2017, the US FDA approved Midostaurin, a small-molecule inhibitor of FMS-like tyrosine kinase 3 (FLT3), which works by inhibiting multiple receptor tyrosine kinases. In a randomized phase 3 trial, midostaurin was administered in addition to conventional chemotherapy extended OS, and EFS was significantly higher than placebo. Due to its survival advantage, the US FDA has approved its usage in combination with cytarabine and daunorubicin chemotherapy to treat AML patients with FLT3- mutation-positive [248,249]. Gilteritinib is a profoundly effective inhibitor approved by the FDA in 2018 for relapsed AML patients with FLT3-mutation. This drug as a monotherapy conclusively enhanced survival in relapsed FLT3-mutated AML patients contrasted with conventional chemotherapy [250,251]. Ivosidenib and enasidenib were approved by the FDA in 2018 for treatment of relapsed AML with IDH mutation, which raises myeloid differentiation and decreases blast counts by repressing IDH1 and IDH2 mutant, respectively. Enasidenib has been shown to target the IDH2 mutation selectively and gave an overall response rate (ORR) of 40.3% with survival advantages [252,253]. On the other hand, Ivosidenib inhibited the IDH1 mutant enzyme and showed an ORR of 42% and CR of 22% in relapsed AML patients [254]. The possible advantage of emerging therapies such as chimeric antigen receptor T cells therapy targeting surface marker of AML, e.g., CD33, CD123, and FLT3, or agents targeting different pathways such as DNA repair, epigenetic modification, TP53 pathway assessed in patients with s-AML in future clinical trials [255]. MHC differentially expresses multiple leukemia-associated antigens on tumor cells. In clinical investigations, they have been examined to be used as a vaccine or dendritic cell-based therapy [256]. Immune checkpoint inhibitors are also of great interest and need to be investigated, especially in s-AML treatment [257]. While the genomic complexity and the interaction of the various complex molecular aberrations in AML pose an immense challenge in specific risk stratification and targeted therapeutics [258,259]. Several novel agents, such as immune checkpoint inhibitors, monoclonal or bispecific antibodies, have been developed as a new therapy for AML [260,261]. The availability of various personlised inhibitors gives a better option of treatment to the patients. The genomic signature, age, co-morbid conditions and afforadability are some of the major driving factors for the treatment of choice. The maitainance of balance between the toxicity and the therapeutic effects has always been a crtical phenomenon. The prior exposure to chemotherapy and the illhealth created due to toxicity also plays a major role for the therapy selection in s-AML and t-AML.
Conclusion
The treatment of secondary and therapy-related acute myeloid leukemia remains one of the most formidable challenges in oncology today. Transformative molecular technologies have led to a more in-depth insight into the molecular and genomic landscape of driver mutations has unraveled the subclonal lineage of secondary acute myeloid leukemia. A better understanding of the molecular and genomic landscape of secondary and therapy-related AML will undoubtedly be the key to improved prognostication and monitoring. These advances have also paved the way for developing novel targeted and personalized therapeutics for individuals harboring the specific genetic signature. These advances also provide a rationale for drug development and repurposing the available drugs in combination with current treatment options.
Disclosure of conflict of interest
None.
References
- 1.Vardiman JW, Thiele J, Arber DA, Brunning RD, Borowitz MJ, Porwit A, Harris NL, Le Beau MM, Hellström-Lindberg E, Tefferi A, Bloomfield CD. The 2008 revision of the World Health Organization (WHO) classification of myeloid neoplasms and acute leukemia: rationale and important changes. Blood. 2009;114:937–951. doi: 10.1182/blood-2009-03-209262. [DOI] [PubMed] [Google Scholar]
- 2.Sakamoto KM, Grant S, Saleiro D, Crispino JD, Hijiya N, Giles F, Platanias L, Eklund EA. Targeting novel signaling pathways for resistant acute myeloid leukemia. Mol Genet Metab. 2015;114:397–402. doi: 10.1016/j.ymgme.2014.11.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Khwaja A, Bjorkholm M, Gale RE, Levine RL, Jordan CT, Ehninger G, Bloomfield CD, Estey E, Burnett A, Cornelissen JJ, Scheinberg DA, Bouscary D, Linch DC. Acute myeloid leukaemia. Nat Rev Dis Primers. 2016;2:16010. doi: 10.1038/nrdp.2016.10. [DOI] [PubMed] [Google Scholar]
- 4.Juliusson G, Antunovic P, Derolf A, Lehmann S, Mollgard L, Stockelberg D, Tidefelt U, Wahlin A, Hoglund M. Age and acute myeloid leukemia: real world data on decision to treat and outcomes from the Swedish Acute Leukemia Registry. Blood. 2009;113:4179–4187. doi: 10.1182/blood-2008-07-172007. [DOI] [PubMed] [Google Scholar]
- 5.Nguyen CH, Gluxam T, Schlerka A, Bauer K, Grandits AM, Hackl H, Dovey O, Zochbauer-Muller S, Cooper JL, Vassiliou GS, Stoiber D, Wieser R, Heller G. SOCS2 is part of a highly prognostic 4-gene signature in AML and promotes disease aggressiveness. Sci Rep. 2019;9:9139. doi: 10.1038/s41598-019-45579-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Cancer Genome Atlas Research Network. Ley TJ, Miller C, Ding L, Raphael BJ, Mungall AJ, Robertson A, Hoadley K, Triche TJ Jr, Laird PW, Baty JD, Fulton LL, Fulton R, Heath SE, Kalicki-Veizer J, Kandoth C, Klco JM, Koboldt DC, Kanchi KL, Kulkarni S, Lamprecht TL, Larson DE, Lin L, Lu C, McLellan MD, McMichael JF, Payton J, Schmidt H, Spencer DH, Tomasson MH, Wallis JW, Wartman LD, Watson MA, Welch J, Wendl MC, Ally A, Balasundaram M, Birol I, Butterfield Y, Chiu R, Chu A, Chuah E, Chun HJ, Corbett R, Dhalla N, Guin R, He A, Hirst C, Hirst M, Holt RA, Jones S, Karsan A, Lee D, Li HI, Marra MA, Mayo M, Moore RA, Mungall K, Parker J, Pleasance E, Plettner P, Schein J, Stoll D, Swanson L, Tam A, Thiessen N, Varhol R, Wye N, Zhao Y, Gabriel S, Getz G, Sougnez C, Zou L, Leiserson MD, Vandin F, Wu HT, Applebaum F, Baylin SB, Akbani R, Broom BM, Chen K, Motter TC, Nguyen K, Weinstein JN, Zhang N, Ferguson ML, Adams C, Black A, Bowen J, Gastier-Foster J, Grossman T, Lichtenberg T, Wise L, Davidsen T, Demchok JA, Shaw KR, Sheth M, Sofia HJ, Yang L, Downing JR, Eley G. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med. 2013;368:2059–2074. doi: 10.1056/NEJMoa1301689. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Kirtonia A, Pandya G, Sethi G, Pandey AK, Das BC, Garg MA. Comprehensive review of genetic alterations and molecular targeted therapies for the implementation of personalized medicine in acute myeloid leukemia. J Mol Med (Berl) 2020;98:1069–1091. doi: 10.1007/s00109-020-01944-5. [DOI] [PubMed] [Google Scholar]
- 8.Watts J, Nimer S. Recent advances in the understanding and treatment of acute myeloid leukemia. F1000Res. 2018;7:F1000 Faculty Rev-1196. [Google Scholar]
- 9.DiNardo CD, Perl AE. Advances in patient care through increasingly individualized therapy. Nat Rev Clin Oncol. 2019;16:73–74. doi: 10.1038/s41571-018-0156-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Higgins A, Shah MV. Genetic and genomic landscape of secondary and therapy-related acute myeloid Leukemia. Genes (Basel) 2020;11:749. doi: 10.3390/genes11070749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Hulegardh E, Nilsson C, Lazarevic V, Garelius H, Antunovic P, Rangert Derolf A, Mollgard L, Uggla B, Wennstrom L, Wahlin A, Hoglund M, Juliusson G, Stockelberg D, Lehmann S. Characterization and prognostic features of secondary acute myeloid leukemia in a population-based setting: a report from the Swedish Acute Leukemia Registry. Am J Hematol. 2015;90:208–214. doi: 10.1002/ajh.23908. [DOI] [PubMed] [Google Scholar]
- 12.Lindsley RC, Mar BG, Mazzola E, Grauman PV, Shareef S, Allen SL, Pigneux A, Wetzler M, Stuart RK, Erba HP, Damon LE, Powell BL, Lindeman N, Steensma DP, Wadleigh M, DeAngelo DJ, Neuberg D, Stone RM, Ebert BL. Acute myeloid leukemia ontogeny is defined by distinct somatic mutations. Blood. 2015;125:1367–1376. doi: 10.1182/blood-2014-11-610543. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kayser S, Dohner K, Krauter J, Kohne CH, Horst HA, Held G, von Lilienfeld-Toal M, Wilhelm S, Kundgen A, Gotze K, Rummel M, Nachbaur D, Schlegelberger B, Gohring G, Spath D, Morlok C, Zucknick M, Ganser A, Dohner H, Schlenk RF German-Austrian AMLSG. The impact of therapy-related acute myeloid leukemia (AML) on outcome in 2853 adult patients with newly diagnosed AML. Blood. 2011;117:2137–2145. doi: 10.1182/blood-2010-08-301713. [DOI] [PubMed] [Google Scholar]
- 14.Shin SH, Yahng SA, Yoon JH, Lee SE, Cho BS, Eom KS, Lee S, Min CK, Kim HJ, Cho SG, Kim DW, Lee JW, Min WS, Park CW, Kim YJ. Survival benefits with transplantation in secondary AML evolving from myelodysplastic syndrome with hypomethylating treatment failure. Bone Marrow Transplant. 2013;48:678–683. doi: 10.1038/bmt.2012.214. [DOI] [PubMed] [Google Scholar]
- 15.de Witte T, Suciu S, Peetermans M, Fenaux P, Strijckmans P, Hayat M, Jaksic B, Selleslag D, Zittoun R, Dardenne M. Intensive chemotherapy for poor prognosis myelodysplasia (MDS) and secondary acute myeloid leukemia (sAML) following MDS of more than 6 months duration. A pilot study by the leukemia cooperative group of the European organisation for research and treatment in cancer (EORTC-LCG) Leukemia. 1995;9:1805–1811. [PubMed] [Google Scholar]
- 16.Østgård LSG, Lund JL, Nørgaard JM, Nørgaard M, Medeiros BC, Nielsen B, Nielsen OJ, Overgaard UM, Kallenbach M, Marcher CW, Riis AH, Sengeløv H. Impact of allogeneic stem cell transplantation in first complete remission in acute myeloid leukemia: a national population-based cohort study. Biol Blood Marrow Transplant. 2018;24:314–323. doi: 10.1016/j.bbmt.2017.10.019. [DOI] [PubMed] [Google Scholar]
- 17.Stelljes M, Krug U, Beelen DW, Braess J, Sauerland MC, Heinecke A, Ligges S, Sauer T, Tschanter P, Thoennissen GB, Berning B, Kolb HJ, Reichle A, Holler E, Schwerdtfeger R, Arnold R, Scheid C, Müller-Tidow C, Woermann BJ, Hiddemann W, Berdel WE, Buchner T. Allogeneic transplantation versus chemotherapy as postremission therapy for acute myeloid leukemia: a prospective matched pairs analysis. J. Clin. Oncol. 2014;32:288–296. doi: 10.1200/JCO.2013.50.5768. [DOI] [PubMed] [Google Scholar]
- 18.Koreth J, Schlenk R, Kopecky KJ, Honda S, Sierra J, Djulbegovic BJ, Wadleigh M, DeAngelo DJ, Stone RM, Sakamaki H, Appelbaum FR, Dohner H, Antin JH, Soiffer RJ, Cutler C. Allogeneic stem cell transplantation for acute myeloid leukemia in first complete remission: systematic review and meta-analysis of prospective clinical trials. JAMA. 2009;301:2349–2361. doi: 10.1001/jama.2009.813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Lim Z, Brand R, Martino R, van Biezen A, Finke J, Bacigalupo A, Beelen D, Devergie A, Alessandrino E, Willemze R, Ruutu T, Boogaerts M, Falda M, Jouet JP, Niederwieser D, Kroger N, Mufti GJ, De Witte TM. Allogeneic hematopoietic stem-cell transplantation for patients 50 years or older with myelodysplastic syndromes or secondary acute myeloid leukemia. J. Clin. Oncol. 2010;28:405–411. doi: 10.1200/JCO.2009.21.8073. [DOI] [PubMed] [Google Scholar]
- 20.Litzow MR, Tarima S, Pérez WS, Bolwell BJ, Cairo MS, Camitta BM, Cutler CS, de Lima M, Dipersio JF, Gale RP, Keating A, Lazarus HM, Luger S, Marks DI, Maziarz RT, McCarthy PL, Pasquini MC, Phillips GL, Rizzo JD, Sierra J, Tallman MS, Weisdorf DJ. Allogeneic transplantation for therapy-related myelodysplastic syndrome and acute myeloid leukemia. Blood. 2010;115:1850–1857. doi: 10.1182/blood-2009-10-249128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Sengsayadeth S, Labopin M, Boumendil A, Finke J, Ganser A, Stelljes M, Ehninger G, Beelen D, Niederwieser D, Blaise D, Dreger P, Mufti G, Chevallier P, Mailhol A, Gatwood KS, Gorin N, Esteve J, Ciceri F, Baron F, Schmid C, Giebel S, Mohty M, Savani BN, Nagler A. Transplant outcomes for secondary acute myeloid leukemia: acute leukemia working party of the European society for blood and bone marrow transplantation study. Biol Blood Marrow Transplant. 2018;24:1406–1414. doi: 10.1016/j.bbmt.2018.04.008. [DOI] [PubMed] [Google Scholar]
- 22.Yakoub-Agha I, de La Salmonière P, Ribaud P, Sutton L, Wattel E, Kuentz M, Jouet JP, Marit G, Milpied N, Deconinck E, Gratecos N, Leporrier M, Chabbert I, Caillot D, Damaj G, Dauriac C, Dreyfus F, François S, Molina L, Tanguy ML, Chevret S, Gluckman E. Allogeneic bone marrow transplantation for therapy-related myelodysplastic syndrome and acute myeloid leukemia: a long-term study of 70 patients-report of the French society of bone marrow transplantation. J. Clin. Oncol. 2000;18:963–971. doi: 10.1200/JCO.2000.18.5.963. [DOI] [PubMed] [Google Scholar]
- 23.Preiss BS, Bergmann OJ, Friis LS, Sorensen AG, Frederiksen M, Gadeberg OV, Mourits-Andersen T, Oestergaard B, Kerndrup GB. Cytogenetic findings in adult secondary acute myeloid leukemia (AML): frequency of favorable and adverse chromosomal aberrations do not differ from adult de novo AML. Cancer Genet Cytogenet. 2010;202:108–122. doi: 10.1016/j.cancergencyto.2010.06.013. [DOI] [PubMed] [Google Scholar]
- 24.Santini V, Ossenkoppele GJ. Hypomethylating agents in the treatment of acute myeloid leukemia: a guide to optimal use. Crit Rev Oncol Hematol. 2019;140:1–7. doi: 10.1016/j.critrevonc.2019.05.013. [DOI] [PubMed] [Google Scholar]
- 25.Hollenbach PW, Nguyen AN, Brady H, Williams M, Ning Y, Richard N, Krushel L, Aukerman SL, Heise C, MacBeth KJ. A comparison of azacitidine and decitabine activities in acute myeloid leukemia cell lines. PLoS One. 2010;5:e9001. doi: 10.1371/journal.pone.0009001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kim TK, Gore SD, Zeidan AM. Epigenetic therapy in acute myeloid leukemia: current and future directions. Semin Hematol. 2015;52:172–183. doi: 10.1053/j.seminhematol.2015.04.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Richardson DR, Green SD, Foster MC, Zeidner JF. Secondary AML emerging after therapy with hypomethylating agents: outcomes, prognostic factors, and treatment options. Curr Hematol Malig Rep. 2021;16:97–111. doi: 10.1007/s11899-021-00608-6. [DOI] [PubMed] [Google Scholar]
- 28.He J, Xiu L, De Porre P, Dass R, Thomas X. Decitabine reduces transfusion dependence in older patients with acute myeloid leukemia: results from a post hoc analysis of a randomized phase III study. Leuk Lymphoma. 2015;56:1033–1042. doi: 10.3109/10428194.2014.951845. [DOI] [PubMed] [Google Scholar]
- 29.Cheung E, Perissinotti AJ, Bixby DL, Burke PW, Pettit KM, Benitez LL, Brown J, Scappaticci GB, Marini BL. The leukemia strikes back: a review of pathogenesis and treatment of secondary AML. Ann Hematol. 2019;98:541–559. doi: 10.1007/s00277-019-03606-0. [DOI] [PubMed] [Google Scholar]
- 30.Granfeldt Ostgard LS, Medeiros BC, Sengelov H, Norgaard M, Andersen MK, Dufva IH, Friis LS, Kjeldsen E, Marcher CW, Preiss B, Severinsen M, Norgaard JM. Epidemiology and clinical significance of secondary and therapy-related acute myeloid leukemia: a national population-based cohort study. J. Clin. Oncol. 2015;33:3641–3649. doi: 10.1200/JCO.2014.60.0890. [DOI] [PubMed] [Google Scholar]
- 31.Borthakur G, Lin E, Jain N, Estey EE, Cortes JE, O’Brien S, Faderl S, Ravandi F, Pierce S, Kantarjian H. Survival is poorer in patients with secondary core-binding factor acute myelogenous leukemia compared with de novo core-binding factor leukemia. Cancer. 2009;115:3217–3221. doi: 10.1002/cncr.24367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Leone G, Mele L, Pulsoni A, Equitani F, Pagano L. The incidence of secondary leukemias. Haematologica. 1999;84:937–945. [PubMed] [Google Scholar]
- 33.Smith SM, Le Beau MM, Huo D, Karrison T, Sobecks RM, Anastasi J, Vardiman JW, Rowley JD, Larson RA. Clinical-cytogenetic associations in 306 patients with therapy-related myelodysplasia and myeloid leukemia: the University of Chicago series. Blood. 2003;102:43–52. doi: 10.1182/blood-2002-11-3343. [DOI] [PubMed] [Google Scholar]
- 34.Rollison DE, Howlader N, Smith MT, Strom SS, Merritt WD, Ries LA, Edwards BK, List AF. Epidemiology of myelodysplastic syndromes and chronic myeloproliferative disorders in the United States, 2001-2004, using data from the NAACCR and SEER programs. Blood. 2008;112:45–52. doi: 10.1182/blood-2008-01-134858. [DOI] [PubMed] [Google Scholar]
- 35.Swerdlow SH, Harris NL, Jaffa ES, Pileri SA, Stein H, Thiele J. Lyon: IARC; 2017. WHO classification of tumours of haematopoietic and lymphoid tissues (revised 4th edition) [Google Scholar]
- 36.Larson RA. Is secondary leukemia an independent poor prognostic factor in acute myeloid leukemia? Best Pract Res Clin Haematol. 2007;20:29–37. doi: 10.1016/j.beha.2006.10.006. [DOI] [PubMed] [Google Scholar]
- 37.Oliai C, Schiller G. How to address second and therapy-related acute myelogenous leukaemia. Br J Haematol. 2020;188:116–128. doi: 10.1111/bjh.16354. [DOI] [PubMed] [Google Scholar]
- 38.Lichtman MA. Distinguishing clonal evolution from so-called secondary acute myelogenous leukemia: adhering to unifying concepts of the genetic basis of leukemogenesis. Blood Cells Mol Dis. 2015;55:1–2. doi: 10.1016/j.bcmd.2015.03.002. [DOI] [PubMed] [Google Scholar]
- 39.Blau O, Hofmann WK, Baldus CD, Thiel G, Serbent V, Schumann E, Thiel E, Blau IW. Chromosomal aberrations in bone marrow mesenchymal stroma cells from patients with myelodysplastic syndrome and acute myeloblastic leukemia. Exp Hematol. 2007;35:221–229. doi: 10.1016/j.exphem.2006.10.012. [DOI] [PubMed] [Google Scholar]
- 40.Raza A, Gezer S, Mundle S, Gao XZ, Alvi S, Borok R, Rifkin S, Iftikhar A, Shetty V, Parcharidou A. Apoptosis in bone marrow biopsy samples involving stromal and hematopoietic cells in 50 patients with myelodysplastic syndromes. Blood. 1995;86:268–276. [PubMed] [Google Scholar]
- 41.Valent P, Horny HP, Bennett JM, Fonatsch C, Germing U, Greenberg P, Haferlach T, Haase D, Kolb HJ, Krieger O, Loken M, van de Loosdrecht A, Ogata K, Orfao A, Pfeilstocker M, Rüter B, Sperr WR, Stauder R, Wells DA. Definitions and standards in the diagnosis and treatment of the myelodysplastic syndromes: Consensus statements and report from a working conference. Leuk Res. 2007;31:727–736. doi: 10.1016/j.leukres.2006.11.009. [DOI] [PubMed] [Google Scholar]
- 42.Ades L, Itzykson R, Fenaux P. Myelodysplastic syndromes. Lancet. 2014;383:2239–2252. doi: 10.1016/S0140-6736(13)61901-7. [DOI] [PubMed] [Google Scholar]
- 43.Tefferi A, Vardiman JW. Myelodysplastic syndromes. N Engl J Med. 2009;361:1872–1885. doi: 10.1056/NEJMra0902908. [DOI] [PubMed] [Google Scholar]
- 44.Dicker F, Haferlach C, Sundermann J, Wendland N, Weiss T, Kern W, Haferlach T, Schnittger S. Mutation analysis for RUNX1, MLL-PTD, FLT3-ITD, NPM1 and NRAS in 269 patients with MDS or secondary AML. Leukemia. 2010;24:1528–1532. doi: 10.1038/leu.2010.124. [DOI] [PubMed] [Google Scholar]
- 45.Greenberg P, Cox C, LeBeau MM, Fenaux P, Morel P, Sanz G, Sanz M, Vallespi T, Hamblin T, Oscier D, Ohyashiki K, Toyama K, Aul C, Mufti G, Bennett J. International scoring system for evaluating prognosis in myelodysplastic syndromes. Blood. 1997;89:2079–2088. [PubMed] [Google Scholar]
- 46.Greenberg PL, Tuechler H, Schanz J, Sanz G, Garcia-Manero G, Sole F, Bennett JM, Bowen D, Fenaux P, Dreyfus F, Kantarjian H, Kuendgen A, Levis A, Malcovati L, Cazzola M, Cermak J, Fonatsch C, Le Beau MM, Slovak ML, Krieger O, Luebbert M, Maciejewski J, Magalhaes SM, Miyazaki Y, Pfeilstöcker M, Sekeres M, Sperr WR, Stauder R, Tauro S, Valent P, Vallespi T, van de Loosdrecht AA, Germing U, Haase D. Revised international prognostic scoring system for myelodysplastic syndromes. Blood. 2012;120:2454–2465. doi: 10.1182/blood-2012-03-420489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Schanz J, Tuchler H, Sole F, Mallo M, Luno E, Cervera J, Granada I, Hildebrandt B, Slovak ML, Ohyashiki K, Steidl C, Fonatsch C, Pfeilstocker M, Nosslinger T, Valent P, Giagounidis A, Aul C, Lübbert M, Stauder R, Krieger O, Garcia-Manero G, Faderl S, Pierce S, Le Beau MM, Bennett JM, Greenberg P, Germing U, Haase D. New comprehensive cytogenetic scoring system for primary myelodysplastic syndromes (MDS) and oligoblastic acute myeloid leukemia after MDS derived from an international database merge. J. Clin. Oncol. 2012;30:820–829. doi: 10.1200/JCO.2011.35.6394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Takahashi K, Jabbour E, Wang X, Luthra R, Bueso-Ramos C, Patel K, Pierce S, Yang H, Wei Y, Daver N, Faderl S, Ravandi F, Estrov Z, Cortes J, Kantarjian H, Garcia-Manero G. Dynamic acquisition of FLT3 or RAS alterations drive a subset of patients with lower risk MDS to secondary AML. Leukemia. 2013;27:2081–2083. doi: 10.1038/leu.2013.165. [DOI] [PubMed] [Google Scholar]
- 49.Jager R, Gisslinger H, Passamonti F, Rumi E, Berg T, Gisslinger B, Pietra D, Harutyunyan A, Klampfl T, Olcaydu D, Cazzola M, Kralovics R. Deletions of the transcription factor Ikaros in myeloproliferative neoplasms. Leukemia. 2010;24:1290–1298. doi: 10.1038/leu.2010.99. [DOI] [PubMed] [Google Scholar]
- 50.Bjorkholm M, Derolf AR, Hultcrantz M, Kristinsson SY, Ekstrand C, Goldin LR, Andreasson B, Birgegard G, Linder O, Malm C, Markevarn B, Nilsson L, Samuelsson J, Granath F, Landgren O. Treatment-related risk factors for transformation to acute myeloid leukemia and myelodysplastic syndromes in myeloproliferative neoplasms. J. Clin. Oncol. 2011;29:2410–2415. doi: 10.1200/JCO.2011.34.7542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Haferlach T, Nagata Y, Grossmann V, Okuno Y, Bacher U, Nagae G, Schnittger S, Sanada M, Kon A, Alpermann T, Yoshida K, Roller A, Nadarajah N, Shiraishi Y, Shiozawa Y, Chiba K, Tanaka H, Koeffler HP, Klein HU, Dugas M, Aburatani H, Kohlmann A, Miyano S, Haferlach C, Kern W, Ogawa S. Landscape of genetic lesions in 944 patients with myelodysplastic syndromes. Leukemia. 2014;28:241–247. doi: 10.1038/leu.2013.336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Saez B, Walter MJ, Graubert TA. Splicing factor gene mutations in hematologic malignancies. Blood. 2017;129:1260–1269. doi: 10.1182/blood-2016-10-692400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Sperling AS, Gibson CJ, Ebert BL. The genetics of myelodysplastic syndrome: from clonal haematopoiesis to secondary leukaemia. Nat Rev Cancer. 2017;17:5–19. doi: 10.1038/nrc.2016.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Corces-Zimmerman MR, Hong WJ, Weissman IL, Medeiros BC, Majeti R. Preleukemic mutations in human acute myeloid leukemia affect epigenetic regulators and persist in remission. Proc Natl Acad Sci U S A. 2014;111:2548–2553. doi: 10.1073/pnas.1324297111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Kim T, Tyndel MS, Kim HJ, Ahn JS, Choi SH, Park HJ, Kim YK, Yang DH, Lee JJ, Jung SH, Kim SY, Min YH, Cheong JW, Sohn SK, Moon JH, Choi M, Lee M, Zhang Z, Kim DDH. The clonal origins of leukemic progression of myelodysplasia. Leukemia. 2017;31:1928–1935. doi: 10.1038/leu.2017.17. [DOI] [PubMed] [Google Scholar]
- 56.Walter MJ, Shen D, Ding L, Shao J, Koboldt DC, Chen K, Larson DE, McLellan MD, Dooling D, Abbott R, Fulton R, Magrini V, Schmidt H, Kalicki-Veizer J, O’Laughlin M, Fan X, Grillot M, Witowski S, Heath S, Frater JL, Eades W, Tomasson M, Westervelt P, DiPersio JF, Link DC, Mardis ER, Ley TJ, Wilson RK, Graubert TA. Clonal architecture of secondary acute myeloid leukemia. N Engl J Med. 2012;366:1090–1098. doi: 10.1056/NEJMoa1106968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Shukron O, Vainstein V, Kundgen A, Germing U, Agur Z. Analyzing transformation of myelodysplastic syndrome to secondary acute myeloid leukemia using a large patient database. Am J Hematol. 2012;87:853–860. doi: 10.1002/ajh.23257. [DOI] [PubMed] [Google Scholar]
- 58.Spivak JL. Myeloproliferative neoplasms. N Engl J Med. 2017;377:895–896. doi: 10.1056/NEJMc1708485. [DOI] [PubMed] [Google Scholar]
- 59.Arber DA, Orazi A, Hasserjian R, Thiele J, Borowitz MJ, Le Beau MM, Bloomfield CD, Cazzola M, Vardiman JW. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood. 2016;127:2391–2405. doi: 10.1182/blood-2016-03-643544. [DOI] [PubMed] [Google Scholar]
- 60.Goel H, Mathur R, Syeda S, Shrivastava A, Jha AK. Promoter hypermethylation of LATS1 gene in oral squamous cell carcinoma (OSCC) among North Indian population. Asian Pac J Cancer Prev. 2021;22:977–982. doi: 10.31557/APJCP.2021.22.3.977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Nangalia J, Massie CE, Baxter EJ, Nice FL, Gundem G, Wedge DC, Avezov E, Li J, Kollmann K, Kent DG, Aziz A, Godfrey AL, Hinton J, Martincorena I, Van Loo P, Jones AV, Guglielmelli P, Tarpey P, Harding HP, Fitzpatrick JD, Goudie CT, Ortmann CA, Loughran SJ, Raine K, Jones DR, Butler AP, Teague JW, O’Meara S, McLaren S, Bianchi M, Silber Y, Dimitropoulou D, Bloxham D, Mudie L, Maddison M, Robinson B, Keohane C, Maclean C, Hill K, Orchard K, Tauro S, Du MQ, Greaves M, Bowen D, Huntly BJP, Harrison CN, Cross NCP, Ron D, Vannucchi AM, Papaemmanuil E, Campbell PJ, Green AR. Somatic CALR mutations in myeloproliferative neoplasms with nonmutated JAK2. N Engl J Med. 2013;369:2391–2405. doi: 10.1056/NEJMoa1312542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Lundberg P, Karow A, Nienhold R, Looser R, Hao-Shen H, Nissen I, Girsberger S, Lehmann T, Passweg J, Stern M, Beisel C, Kralovics R, Skoda RC. Clonal evolution and clinical correlates of somatic mutations in myeloproliferative neoplasms. Blood. 2014;123:2220–2228. doi: 10.1182/blood-2013-11-537167. [DOI] [PubMed] [Google Scholar]
- 63.Vannucchi AM, Lasho TL, Guglielmelli P, Biamonte F, Pardanani A, Pereira A, Finke C, Score J, Gangat N, Mannarelli C, Ketterling RP, Rotunno G, Knudson RA, Susini MC, Laborde RR, Spolverini A, Pancrazzi A, Pieri L, Manfredini R, Tagliafico E, Zini R, Jones A, Zoi K, Reiter A, Duncombe A, Pietra D, Rumi E, Cervantes F, Barosi G, Cazzola M, Cross NC, Tefferi A. Mutations and prognosis in primary myelofibrosis. Leukemia. 2013;27:1861–1869. doi: 10.1038/leu.2013.119. [DOI] [PubMed] [Google Scholar]
- 64.Tefferi A, Lasho TL, Guglielmelli P, Finke CM, Rotunno G, Elala Y, Pacilli A, Hanson CA, Pancrazzi A, Ketterling RP, Mannarelli C, Barraco D, Fanelli T, Pardanani A, Gangat N, Vannucchi AM. Targeted deep sequencing in polycythemia vera and essential thrombocythemia. Blood Adv. 2016;1:21–30. doi: 10.1182/bloodadvances.2016000216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Tefferi A, Vannucchi AM. Genetic risk assessment in myeloproliferative neoplasms. Mayo Clin Proc. 2017;92:1283–1290. doi: 10.1016/j.mayocp.2017.06.002. [DOI] [PubMed] [Google Scholar]
- 66.Vallapureddy RR, Mudireddy M, Penna D, Lasho TL, Finke CM, Hanson CA, Ketterling RP, Begna KH, Gangat N, Pardanani A, Tefferi A. Leukemic transformation among 1306 patients with primary myelofibrosis: risk factors and development of a predictive model. Blood Cancer J. 2019;9:12. doi: 10.1038/s41408-019-0175-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Gangat N, Caramazza D, Vaidya R, George G, Begna K, Schwager S, Van Dyke D, Hanson C, Wu W, Pardanani A, Cervantes F, Passamonti F, Tefferi A. DIPSS plus: a refined dynamic international prognostic scoring system for primary myelofibrosis that incorporates prognostic information from karyotype, platelet count, and transfusion status. J. Clin. Oncol. 2011;29:392–397. doi: 10.1200/JCO.2010.32.2446. [DOI] [PubMed] [Google Scholar]
- 68.Yogarajah M, Tefferi A. Leukemic transformation in myeloproliferative neoplasms: a literature review on risk, characteristics, and outcome. Mayo Clin Proc. 2017;92:1118–1128. doi: 10.1016/j.mayocp.2017.05.010. [DOI] [PubMed] [Google Scholar]
- 69.Yonal-Hindilerden I, Daglar-Aday A, Hindilerden F, Akadam-Teker B, Yilmaz C, Nalcaci M, Yavuz AS, Sargin D. The clinical significance of IDH mutations in essential thrombocythemia and primary myelofibrosis. J Clin Med Res. 2016;8:29–39. doi: 10.14740/jocmr2405w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Tefferi A, Lasho TL, Finke CM, Elala Y, Hanson CA, Ketterling RP, Gangat N, Pardanani A. Targeted deep sequencing in primary myelofibrosis. Blood Adv. 2016;1:105–111. doi: 10.1182/bloodadvances.2016000208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Harutyunyan A, Klampfl T, Cazzola M, Kralovics R. p53 lesions in leukemic transformation. N Engl J Med. 2011;364:488–490. doi: 10.1056/NEJMc1012718. [DOI] [PubMed] [Google Scholar]
- 72.Mascarenhas J. A concise update on risk factors, therapy, and outcome of leukemic transformation of myeloproliferative neoplasms. Clin Lymphoma Myeloma Leuk. 2016;16:124–129. doi: 10.1016/j.clml.2016.02.016. [DOI] [PubMed] [Google Scholar]
- 73.Venton G, Courtier F, Charbonnier A, Dincan E, Saillard C, Mohty B, Mozziconacci MJ, Birnbaum D, Murati A, Vey N, Rey J. Impact of gene mutations on treatment response and prognosis of acute myeloid leukemia secondary to myeloproliferative neoplasms. Am J Hematol. 2018;93:330–338. doi: 10.1002/ajh.24973. [DOI] [PubMed] [Google Scholar]
- 74.Courtier F, Carbuccia N, Garnier S, Guille A, Adelaide J, Cervera N, Gelsi-Boyer V, Mozziconacci MJ, Rey J, Vey N, Chaffanet M, Birnbaum D, Murati A. Genomic analysis of myeloproliferative neoplasms in chronic and acute phases. Haematologica. 2017;102:e11–e14. doi: 10.3324/haematol.2016.152363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Ramadan SM, Fouad TM, Summa V, Hasan SKh, Lo-Coco F. Acute myeloid leukemia developing in patients with autoimmune diseases. Haematologica. 2012;97:805–817. doi: 10.3324/haematol.2011.056283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Mauritzson N, Albin M, Rylander L, Billstrom R, Ahlgren T, Mikoczy Z, Bjork J, Stromberg U, Nilsson PG, Mitelman F, Hagmar L, Johansson B. Pooled analysis of clinical and cytogenetic features in treatment-related and de novo adult acute myeloid leukemia and myelodysplastic syndromes based on a consecutive series of 761 patients analyzed 1976-1993 and on 5098 unselected cases reported in the literature 1974-2001. Leukemia. 2002;16:2366–2378. doi: 10.1038/sj.leu.2402713. [DOI] [PubMed] [Google Scholar]
- 77.Morton LM, Gibson TM, Clarke CA, Lynch CF, Anderson LA, Pfeiffer R, Landgren O, Weisenburger DD, Engels EA. Risk of myeloid neoplasms after solid organ transplantation. Leukemia. 2014;28:2317–2323. doi: 10.1038/leu.2014.132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Offman J, Opelz G, Doehler B, Cummins D, Halil O, Banner NR, Burke MM, Sullivan D, Macpherson P, Karran P. Defective DNA mismatch repair in acute myeloid leukemia/myelodysplastic syndrome after organ transplantation. Blood. 2004;104:822–828. doi: 10.1182/blood-2003-11-3938. [DOI] [PubMed] [Google Scholar]
- 79.Fu D, Calvo JA, Samson LD. Balancing repair and tolerance of DNA damage caused by alkylating agents. Nat Rev Cancer. 2012;12:104–120. doi: 10.1038/nrc3185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Nitiss JL. Targeting DNA topoisomerase II in cancer chemotherapy. Nat Rev Cancer. 2009;9:338–350. doi: 10.1038/nrc2607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Pedersen-Bjergaard J, Andersen MK, Christiansen DH, Nerlov C. Genetic pathways in therapy-related myelodysplasia and acute myeloid leukemia. Blood. 2002;99:1909–1912. doi: 10.1182/blood.v99.6.1909. [DOI] [PubMed] [Google Scholar]
- 82.Pedersen-Bjergaard J, Philip P. Balanced translocations involving chromosome bands 11q23 and 21q22 are highly characteristic of myelodysplasia and leukemia following therapy with cytostatic agents targeting at DNA-topoisomerase II. Blood. 1991;78:1147–1148. [PubMed] [Google Scholar]
- 83.Le Beau MM, Albain KS, Larson RA, Vardiman JW, Davis EM, Blough RR, Golomb HM, Rowley JD. Clinical and cytogenetic correlations in 63 patients with therapy-related myelodysplastic syndromes and acute nonlymphocytic leukemia: further evidence for characteristic abnormalities of chromosomes no. 5 and 7. J. Clin. Oncol. 1986;4:325–345. doi: 10.1200/JCO.1986.4.3.325. [DOI] [PubMed] [Google Scholar]
- 84.Johansson B, Mertens F, Heim S, Kristoffersson U, Mitelman F. Cytogenetics of secondary myelodysplasia (sMDS) and acute nonlymphocytic leukemia (sANLL) Eur J Haematol. 1991;47:17–27. doi: 10.1111/j.1600-0609.1991.tb00556.x. [DOI] [PubMed] [Google Scholar]
- 85.Pedersen-Bjergaard J, Pedersen M, Roulston D, Philip P. Different genetic pathways in leukemogenesis for patients presenting with therapy-related myelodysplasia and therapy-related acute myeloid leukemia. Blood. 1995;86:3542–3552. [PubMed] [Google Scholar]
- 86.Stephenson J, Lizhen H, Mufti GJ. Possible co-existence of RAS activation and monosomy 7 in the leukaemic transformation of myelodysplastic syndromes. Leuk Res. 1995;19:741–748. doi: 10.1016/0145-2126(95)00056-t. [DOI] [PubMed] [Google Scholar]
- 87.Joslin JM, Fernald AA, Tennant TR, Davis EM, Kogan SC, Anastasi J, Crispino JD, Le Beau MM. Haploinsufficiency of EGR1, a candidate gene in the del(5q), leads to the development of myeloid disorders. Blood. 2007;110:719–726. doi: 10.1182/blood-2007-01-068809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Horiike S, Misawa S, Kaneko H, Sasai Y, Kobayashi M, Fujii H, Tanaka S, Yagita M, Abe T, Kashima K, Taniwaki M. Distinct genetic involvement of the TP53 gene in therapy-related leukemia and myelodysplasia with chromosomal losses of Nos 5 and/or 7 and its possible relationship to replication error phenotype. Leukemia. 1999;13:1235–1242. doi: 10.1038/sj.leu.2401466. [DOI] [PubMed] [Google Scholar]
- 89.Mitelman F, Johansson B, Mertens F. The impact of translocations and gene fusions on cancer causation. Nat Rev Cancer. 2007;7:233–245. doi: 10.1038/nrc2091. [DOI] [PubMed] [Google Scholar]
- 90.Pui CH, Behm FG, Raimondi SC, Dodge RK, George SL, Rivera GK, Mirro J Jr, Kalwinsky DK, Dahl GV, Murphy SB. Secondary acute myeloid leukemia in children treated for acute lymphoid leukemia. N Engl J Med. 1989;321:136–142. doi: 10.1056/NEJM198907203210302. [DOI] [PubMed] [Google Scholar]
- 91.Pui CH, Relling MV, Rivera GK, Hancock ML, Raimondi SC, Heslop HE, Santana VM, Ribeiro RC, Sandlund JT, Mahmoud HH. Epipodophyllotoxin-related acute myeloid leukemia: a study of 35 cases. Leukemia. 1995;9:1990–1996. [PubMed] [Google Scholar]
- 92.Faller BA, Robu VG, Borghaei H. Therapy-related acute myelogenous leukemia with an 11q23/MLL translocation following adjuvant cisplatin and vinorelbine for non-small-cell lung cancer. Clin Lung Cancer. 2009;10:438–40. doi: 10.3816/CLC.2009.n.082. [DOI] [PubMed] [Google Scholar]
- 93.Quesnel B, Kantarjian H, Bjergaard JP, Brault P, Estey E, Lai JL, Tilly H, Stoppa AM, Archimbaud E, Harousseau JL. Therapy-related acute myeloid leukemia with t(8;21), inv(16), and t(8;16): a report on 25 cases and review of the literature. J. Clin. Oncol. 1993;11:2370–2379. doi: 10.1200/JCO.1993.11.12.2370. [DOI] [PubMed] [Google Scholar]
- 94.Smith KA, Cowell IG, Zhang Y, Sondka Z, Austin CA. The role of topoisomerase II beta on breakage and proximity of RUNX1 to partner alleles RUNX1T1 and EVI1. Genes Chromosomes Cancer. 2014;53:117–128. doi: 10.1002/gcc.22124. [DOI] [PubMed] [Google Scholar]
- 95.Godley LA, Larson RA. Therapy-related myeloid leukemia. Semin Oncol. 2008;35:418–429. doi: 10.1053/j.seminoncol.2008.04.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Raza-Egilmez SZ, Jani-Sait SN, Grossi M, Higgins MJ, Shows TB, Aplan PD. NUP98-HOXD13 gene fusion in therapy-related acute myelogenous leukemia. Cancer Res. 1998;58:4269–4273. [PubMed] [Google Scholar]
- 97.Christiansen DH, Pedersen-Bjergaard J. Internal tandem duplications of the FLT3 and MLL genes are mainly observed in atypical cases of therapy-related acute myeloid leukemia with a normal karyotype and are unrelated to type of previous therapy. Leukemia. 2001;15:1848–1851. doi: 10.1038/sj.leu.2402246. [DOI] [PubMed] [Google Scholar]
- 98.Pedersen-Bjergaard J, Christiansen DH, Desta F, Andersen MK. Alternative genetic pathways and cooperating genetic abnormalities in the pathogenesis of therapy-related myelodysplasia and acute myeloid leukemia. Leukemia. 2006;20:1943–1949. doi: 10.1038/sj.leu.2404381. [DOI] [PubMed] [Google Scholar]
- 99.Schroeder T, Kuendgen A, Kayser S, Kroger N, Braulke F, Platzbecker U, Klarner V, Zohren F, Haase D, Stadler M, Schlenk R, Czibere AG, Bruns I Fenk R, Gattermann N, Haas R, Kobbe G, Germing U. Therapy-related myeloid neoplasms following treatment with radioiodine. Haematologica. 2012;97:206–212. doi: 10.3324/haematol.2011.049114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Bondar T, Medzhitov R. p53-mediated hematopoietic stem and progenitor cell competition. Cell Stem Cell. 2010;6:309–322. doi: 10.1016/j.stem.2010.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Marusyk A, Porter CC, Zaberezhnyy V, DeGregori J. Irradiation selects for p53-deficient hematopoietic progenitors. PLoS Biol. 2010;8:e1000324. doi: 10.1371/journal.pbio.1000324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Takahashi K, Wang F, Kantarjian H, Doss D, Khanna K, Thompson E, Zhao L, Patel K, Neelapu S, Gumbs C, Bueso-Ramos C, DiNardo CD, Colla S, Ravandi F, Zhang J, Huang X, Wu X, Samaniego F, Garcia-Manero G, Futreal PA. Preleukaemic clonal haemopoiesis and risk of therapy-related myeloid neoplasms: a case-control study. Lancet Oncol. 2017;18:100–111. doi: 10.1016/S1470-2045(16)30626-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Anderson LA, Pfeiffer RM, Landgren O, Gadalla S, Berndt SI, Engels EA. Risks of myeloid malignancies in patients with autoimmune conditions. Br J Cancer. 2009;100:822–828. doi: 10.1038/sj.bjc.6604935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Kristinsson SY, Bjorkholm M, Hultcrantz M, Derolf AR, Landgren O, Goldin LR. Chronic immune stimulation might act as a trigger for the development of acute myeloid leukemia or myelodysplastic syndromes. J. Clin. Oncol. 2011;29:2897–2903. doi: 10.1200/JCO.2011.34.8540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Østgård LSG, Nørgaard M, Pedersen L, Østgård RD, Medeiros BC, Overgaard UM, Schöllkopf C, Severinsen M, Marcher CW, Jensen MK. Autoimmune diseases, infections, use of antibiotics and the risk of acute myeloid leukaemia: a national population-based case-control study. Br J Haematol. 2018;181:205–214. doi: 10.1111/bjh.15163. [DOI] [PubMed] [Google Scholar]
- 106.Ertz-Archambault N, Kosiorek H, Taylor GE, Kelemen K, Dueck A, Castro J, Marino R, Gauthier S, Finn L, Sproat LZ, Palmer J, Mesa RA, Al-Kali A, Foran J, Tibes R. Association of therapy for autoimmune disease with myelodysplastic syndromes and acute myeloid leukemia. JAMA Oncol. 2017;3:936–943. doi: 10.1001/jamaoncol.2016.6435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Uehara E, Takeuchi S, Tasaka T, Matsuhashi Y, Yang Y, Fujita M, Tamura T, Nagai M, Koeffler HP. Aberrant methylation in promoter-associated CpG islands of multiple genes in therapy-related leukemia. Int J Oncol. 2003;23:693–696. [PubMed] [Google Scholar]
- 108.Ley TJ, Mardis ER, Ding L, Fulton B, McLellan MD, Chen K, Dooling D, Dunford-Shore BH, McGrath S, Hickenbotham M, Cook L, Abbott R, Larson DE, Koboldt DC, Pohl C, Smith S, Hawkins A, Abbott S, Locke D, Hillier LW, Miner T, Fulton L, Magrini V, Wylie T, Glasscock J, Conyers J, Sander N, Shi X, Osborne JR, Minx P, Gordon D, Chinwalla A, Zhao Y, Ries RE, Payton JE, Westervelt P, Tomasson MH, Watson M, Baty J, Ivanovich J, Heath S, Shannon WD, Nagarajan R, Walter MJ, Link DC, Graubert TA, DiPersio JF, Wilson RK. DNA sequencing of a cytogenetically normal acute myeloid leukaemia genome. Nature. 2008;456:66–72. doi: 10.1038/nature07485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Roloff GW, Griffiths EA. When to obtain genomic data in acute myeloid leukemia (AML) and which mutations matter. Blood Adv. 2018;2:3070–3080. doi: 10.1182/bloodadvances.2018020206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Zhang L, Padron E, Lancet J. The molecular basis and clinical significance of genetic mutations identified in myelodysplastic syndromes. Leuk Res. 2015;39:6–17. doi: 10.1016/j.leukres.2014.10.006. [DOI] [PubMed] [Google Scholar]
- 111.Bejar R, Steensma DP. Recent developments in myelodysplastic syndromes. Blood. 2014;124:2793–2803. doi: 10.1182/blood-2014-04-522136. [DOI] [PubMed] [Google Scholar]
- 112.Metzeler KH, Herold T, Rothenberg-Thurley M, Amler S, Sauerland MC, Görlich D, Schneider S, Konstandin NP, Dufour A, Bräundl K, Ksienzyk B, Zellmeier E, Hartmann L, Greif PA, Fiegl M, Subklewe M, Bohlander SK, Krug U, Faldum A, Berdel WE, Wörmann B, Büchner T, Hiddemann W, Braess J, Spiekermann K AMLCG Study Group. Spectrum and prognostic relevance of driver gene mutations in acute myeloid leukemia. Blood. 2016;128:686–98. doi: 10.1182/blood-2016-01-693879. [DOI] [PubMed] [Google Scholar]
- 113.Ding L, Ley TJ, Larson DE, Miller CA, Koboldt DC, Welch JS, Ritchey JK, Young MA, Lamprecht T, McLellan MD, McMichael JF, Wallis JW, Lu C, Shen D, Harris CC, Dooling DJ, Fulton RS, Fulton LL, Chen K, Schmidt H, Kalicki-Veizer J, Magrini VJ, Cook L, McGrath SD, Vickery TL, Wendl MC, Heath S, Watson MA, Link DC, Tomasson MH, Shannon WD, Payton JE, Kulkarni S, Westervelt P, Walter MJ, Graubert TA, Mardis ER, Wilson RK, DiPersio JF. Clonal evolution in relapsed acute myeloid leukaemia revealed by whole-genome sequencing. Nature. 2012;481:506–510. doi: 10.1038/nature10738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Welch JS, Ley TJ, Link DC, Miller CA, Larson DE, Koboldt DC, Wartman LD, Lamprecht TL, Liu F, Xia J, Kandoth C, Fulton RS, McLellan MD, Dooling DJ, Wallis JW, Chen K, Harris CC, Schmidt HK, Kalicki-Veizer JM, Lu C, Zhang Q, Lin L, O’Laughlin MD, McMichael JF, Delehaunty KD, Fulton LA, Magrini VJ, McGrath SD, Demeter RT, Vickery TL, Hundal J, Cook LL, Swift GW, Reed JP, Alldredge PA, Wylie TN, Walker JR, Watson MA, Heath SE, Shannon WD, Varghese N, Nagarajan R, Payton JE, Baty JD, Kulkarni S, Klco JM, Tomasson MH, Westervelt P, Walter MJ, Graubert TA, DiPersio JF, Ding L, Mardis ER, Wilson RK. The origin and evolution of mutations in acute myeloid leukemia. Cell. 2012;150:264–278. doi: 10.1016/j.cell.2012.06.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Shlush LI, Zandi S, Mitchell A, Chen WC, Brandwein JM, Gupta V, Kennedy JA, Schimmer AD, Schuh AC, Yee KW, McLeod JL, Doedens M, Medeiros JJ, Marke R, Kim HJ, Lee K, McPherson JD, Hudson TJ HALT Pan-Leukemia Gene Panel Consortium. Brown AM, Yousif F, Trinh QM, Stein LD, Minden MD, Wang JC, Dick JE. Identification of pre-leukaemic haematopoietic stem cells in acute leukaemia. Nature. 2014;506:328–333. doi: 10.1038/nature13038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Gill H, Leung AY, Kwong YL. Molecular and cellular mechanisms of myelodysplastic syndrome: implications on targeted therapy. Int J Mol Sci. 2016;17:440. doi: 10.3390/ijms17040440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Goel H, Singhal S, Mathur R, Syeda S, Gupta RK, Kumar A, Shrivastava A, Jha AK. Promoter hypermethylation of LATS2 gene in oral squamous cell carcinoma (OSCC) among North Indian population. Asian Pac J Cancer Prev. 2020;21:1283–1287. doi: 10.31557/APJCP.2020.21.5.1283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Goodell MA, Godley LA. Perspectives and future directions for epigenetics in hematology. Blood. 2013;121:5131–5137. doi: 10.1182/blood-2013-04-427724. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Mehdipour P, Santoro F, Minucci S. Epigenetic alterations in acute myeloid leukemias. FEBS J. 2015;282:1786–1800. doi: 10.1111/febs.13142. [DOI] [PubMed] [Google Scholar]
- 120.Deneberg S, Guardiola P, Lennartsson A, Qu Y, Gaidzik V, Blanchet O, Karimi M, Bengtzen S, Nahi H, Uggla B, Tidefelt U, Hoglund M, Paul C, Ekwall K, Dohner K, Lehmann S. Prognostic DNA methylation patterns in cytogenetically normal acute myeloid leukemia are predefined by stem cell chromatin marks. Blood. 2011;118:5573–5582. doi: 10.1182/blood-2011-01-332353. [DOI] [PubMed] [Google Scholar]
- 121.Figueroa ME, Lugthart S, Li Y, Erpelinck-Verschueren C, Deng X, Christos PJ, Schifano E, Booth J, van Putten W, Skrabanek L, Campagne F, Mazumdar M, Greally JM, Valk PJ, Lowenberg B, Delwel R, Melnick A. DNA methylation signatures identify biologically distinct subtypes in acute myeloid leukemia. Cancer Cell. 2010;17:13–27. doi: 10.1016/j.ccr.2009.11.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Tahiliani M, Koh KP, Shen Y, Pastor WA, Bandukwala H, Brudno Y, Agarwal S, Iyer LM, Liu DR, Aravind L, Rao A. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science. 2009;324:930–935. doi: 10.1126/science.1170116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Shide K, Kameda T, Shimoda H, Yamaji T, Abe H, Kamiunten A, Sekine M, Hidaka T, Katayose K, Kubuki Y, Yamamoto S, Miike T, Iwakiri H, Hasuike S, Nagata K, Marutsuka K, Iwama A, Matsuda T, Kitanaka A, Shimoda K. TET2 is essential for survival and hematopoietic stem cell homeostasis. Leukemia. 2012;26:2216–2223. doi: 10.1038/leu.2012.94. [DOI] [PubMed] [Google Scholar]
- 124.Koh KP, Yabuuchi A, Rao S, Huang Y, Cunniff K, Nardone J, Laiho A, Tahiliani M, Sommer CA, Mostoslavsky G, Lahesmaa R, Orkin SH, Rodig SJ, Daley GQ, Rao A. Tet1 and Tet2 regulate 5-hydroxymethylcytosine production and cell lineage specification in mouse embryonic stem cells. Cell Stem Cell. 2011;8:200–213. doi: 10.1016/j.stem.2011.01.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Delhommeau F, Dupont S, Della Valle V, James C, Trannoy S, Masse A, Kosmider O, Le Couedic JP, Robert F, Alberdi A, Lecluse Y, Plo I, Dreyfus FJ, Marzac C, Casadevall N, Lacombe C, Romana SP, Dessen P, Soulier J, Viguie F, Fontenay M, Vainchenker W, Bernard OA. Mutation in TET2 in myeloid cancers. N Engl J Med. 2009;360:2289–2301. doi: 10.1056/NEJMoa0810069. [DOI] [PubMed] [Google Scholar]
- 126.Goel H, Rahul E, Gupta AK, Meena JP, Chopra A, Ranjan A, Hussain S, Rath GK, Tanwar P. Molecular update on biology of Wilms tumor 1 gene and its applications in acute myeloid leukemia. Am J Blood Res. 2020;10:151–160. [PMC free article] [PubMed] [Google Scholar]
- 127.Nibourel O, Kosmider O, Cheok M, Boissel N, Renneville A, Philippe N, Dombret H, Dreyfus F, Quesnel B, Geffroy S, Quentin S, Roche-Lestienne C, Cayuela JM, Roumier C, Fenaux P, Vainchenker W, Bernard OA, Soulier J, Fontenay M, Preudhomme C. Incidence and prognostic value of TET2 alterations in de novo acute myeloid leukemia achieving complete remission. Blood. 2010;116:1132–1135. doi: 10.1182/blood-2009-07-234484. [DOI] [PubMed] [Google Scholar]
- 128.Bejar R, Levine R, Ebert BL. Unraveling the molecular pathophysiology of myelodysplastic syndromes. J. Clin. Oncol. 2011;29:504–515. doi: 10.1200/JCO.2010.31.1175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Jankowska AM, Szpurka H, Tiu RV, Makishima H, Afable M, Huh J, O’Keefe CL, Ganetzky R, McDevitt MA, Maciejewski JP. Loss of heterozygosity 4q24 and TET2 mutations associated with myelodysplastic/myeloproliferative neoplasms. Blood. 2009;113:6403–6410. doi: 10.1182/blood-2009-02-205690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Itzykson R, Kosmider O, Renneville A, Gelsi-Boyer V, Meggendorfer M, Morabito M, Berthon C, Ades L, Fenaux P, Beyne-Rauzy O, Vey N, Braun T, Haferlach T, Dreyfus F, Cross NC, Preudhomme C, Bernard OA, Fontenay M, Vainchenker W, Schnittger S, Birnbaum D, Droin N, Solary E. Prognostic score including gene mutations in chronic myelomonocytic leukemia. J. Clin. Oncol. 2013;31:2428–2436. doi: 10.1200/JCO.2012.47.3314. [DOI] [PubMed] [Google Scholar]
- 131.Kosmider O, Gelsi-Boyer V, Ciudad M, Racoeur C, Jooste V, Vey N, Quesnel B, Fenaux P, Bastie JN, Beyne-Rauzy O, Stamatoulas A, Dreyfus F, Ifrah N, de Botton S, Vainchenker W, Bernard OA, Birnbaum D, Fontenay M, Solary E Groupe Francophone des Myélodysplasies. TET2 gene mutation is a frequent and adverse event in chronic myelomonocytic leukemia. Haematologica. 2009;94:1676–1681. doi: 10.3324/haematol.2009.011205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Chou WC, Chou SC, Liu CY, Chen CY, Hou HA, Kuo YY, Lee MC, Ko BS, Tang JL, Yao M, Tsay W, Wu SJ, Huang SY, Hsu SC, Chen YC, Chang YC, Kuo YY, Kuo KT, Lee FY, Liu MC, Liu CW, Tseng MH, Huang CF, Tien HF. TET2 mutation is an unfavorable prognostic factor in acute myeloid leukemia patients with intermediate-risk cytogenetics. Blood. 2011;118:3803–3810. doi: 10.1182/blood-2011-02-339747. [DOI] [PubMed] [Google Scholar]
- 133.Abdel-Wahab O, Mullally A, Hedvat C, Garcia-Manero G, Patel J, Wadleigh M, Malinge S, Yao J, Kilpivaara O, Bhat R, Huberman K, Thomas S, Dolgalev I, Heguy A, Paietta E, Le Beau MM, Beran M, Tallman MS, Ebert BL, Kantarjian HM, Stone RM, Gilliland DG, Crispino JD, Levine RL. Genetic characterization of TET1, TET2, and TET3 alterations in myeloid malignancies. Blood. 2009;114:144–147. doi: 10.1182/blood-2009-03-210039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Kosmider O, Gelsi-Boyer V, Cheok M, Grabar S, Della-Valle V, Picard F, Viguie F, Quesnel B, Beyne-Rauzy O, Solary E, Vey N, Hunault-Berger M, Fenaux P, Mansat-De Mas V, Delabesse E, Guardiola P, Lacombe C, Vainchenker W, Preudhomme C, Dreyfus F, Bernard OA, Birnbaum D, Fontenay M Groupe Francophone des Myelodysplasies. TET2 mutation is an independent favorable prognostic factor in myelodysplastic syndromes (MDSs) Blood. 2009;114:3285–3291. doi: 10.1182/blood-2009-04-215814. [DOI] [PubMed] [Google Scholar]
- 135.Walter MJ, Ding L, Shen D, Shao J, Grillot M, McLellan M, Fulton R, Schmidt H, Kalicki-Veizer J, O’Laughlin M, Kandoth C, Baty J, Westervelt P, DiPersio JF, Mardis ER, Wilson RK, Ley TJ, Graubert TA. Recurrent DNMT3A mutations in patients with myelodysplastic syndromes. Leukemia. 2011;25:1153–1158. doi: 10.1038/leu.2011.44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Yuan XQ, Chen P, Du YX, Zhu KW, Zhang DY, Yan H, Liu H, Liu YL, Cao S, Zhou G, Zeng H, Chen SP, Zhao XL, Yang J, Zeng WJ, Chen XP. Influence of DNMT3A R882 mutations on AML prognosis determined by the allele ratio in Chinese patients. J Transl Med. 2019;17:220. doi: 10.1186/s12967-019-1959-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Papaemmanuil E, Gerstung M, Malcovati L, Tauro S, Gundem G, Van Loo P, Yoon CJ, Ellis P, Wedge DC, Pellagatti A, Shlien A, Groves MJ, Forbes SA, Raine K, Hinton J, Mudie LJ, McLaren S, Hardy C, Latimer C, Della Porta MG, O’Meara S, Ambaglio I, Galli A, Butler AP, Walldin G, Teague JW, Quek L, Sternberg A, Gambacorti-Passerini C, Cross NC, Green AR, Boultwood J, Vyas P, Hellstrom-Lindberg E, Bowen D, Cazzola M, Stratton MR, Campbell PJ Chronic Myeloid Disorders Working Group of the International Cancer Genome Consortium. Clinical and biological implications of driver mutations in myelodysplastic syndromes. Blood. 2013;122:3616–3627. doi: 10.1182/blood-2013-08-518886. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Jurkowska RZ, Jurkowski TP, Jeltsch A. Structure and function of mammalian DNA methyltransferases. Chembiochem. 2011;12:206–222. doi: 10.1002/cbic.201000195. [DOI] [PubMed] [Google Scholar]
- 139.Challen GA, Sun D, Jeong M, Luo M, Jelinek J, Berg JS, Bock C, Vasanthakumar A, Gu H, Xi Y, Liang S, Lu Y, Darlington GJ, Meissner A, Issa JP, Godley LA, Li W, Goodell MA. Dnmt3a is essential for hematopoietic stem cell differentiation. Nat Genet. 2011;44:23–31. doi: 10.1038/ng.1009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Ley TJ, Ding L, Walter MJ, McLellan MD, Lamprecht T, Larson DE, Kandoth C, Payton JE, Baty J, Welch J, Harris CC, Lichti CF, Townsend RR, Fulton RS, Dooling DJ, Koboldt DC, Schmidt H, Zhang Q, Osborne JR, Lin L, O’Laughlin M, McMichael JF, Delehaunty KD, McGrath SD, Fulton LA, Magrini VJ, Vickery TL, Hundal J, Cook LL, Conyers JJ, Swift GW, Reed JP, Alldredge PA, Wylie T, Walker J, Kalicki J, Watson MA, Heath S, Shannon WD, Varghese N, Nagarajan R, Westervelt P, Tomasson MH, Link DC, Graubert TA, DiPersio JF, Mardis ER, Wilson RK. DNMT3A mutations in acute myeloid leukemia. N Engl J Med. 2010;363:2424–2433. doi: 10.1056/NEJMoa1005143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Roller A, Grossmann V, Bacher U, Poetzinger F, Weissmann S, Nadarajah N, Boeck L, Kern W, Haferlach C, Schnittger S, Haferlach T, Kohlmann A. Landmark analysis of DNMT3A mutations in hematological malignancies. Leukemia. 2013;27:1573–1578. doi: 10.1038/leu.2013.65. [DOI] [PubMed] [Google Scholar]
- 142.Gale RE, Lamb K, Allen C, El-Sharkawi D, Stowe C, Jenkinson S, Tinsley S, Dickson G, Burnett AK, Hills RK, Linch DC. Simpson’s paradox and the impact of different DNMT3A mutations on outcome in younger adults with acute myeloid leukemia. J. Clin. Oncol. 2015;33:2072–2083. doi: 10.1200/JCO.2014.59.2022. [DOI] [PubMed] [Google Scholar]
- 143.Gaidzik VI, Schlenk RF, Paschka P, Stölzle A, Späth D, Kuendgen A, von Lilienfeld-Toal M, Brugger W, Derigs HG, Kremers S, Greil R, Raghavachar A, Ringhoffer M, Salih HR, Wattad M, Kirchen HG, Runde V, Heil G, Petzer AL, Girschikofsky M, Heuser M, Kayser S, Goehring G, Teleanu MV, Schlegelberger B, Ganser A, Krauter J, Bullinger L, Dohner H, Dohner K. Clinical impact of DNMT3A mutations in younger adult patients with acute myeloid leukemia: results of the AML Study Group (AMLSG) Blood. 2013;121:4769–4777. doi: 10.1182/blood-2012-10-461624. [DOI] [PubMed] [Google Scholar]
- 144.Falini B, Sportoletti P, Brunetti L, Martelli MP. Perspectives for therapeutic targeting of gene mutations in acute myeloid leukaemia with normal cytogenetics. Br J Haematol. 2015;170:305–322. doi: 10.1111/bjh.13409. [DOI] [PubMed] [Google Scholar]
- 145.Nebbioso A, Benedetti R, Conte M, Iside C, Altucci L. Genetic mutations in epigenetic modifiers as therapeutic targets in acute myeloid leukemia. Expert Opin Ther Targets. 2015;19:1187–1202. doi: 10.1517/14728222.2015.1051728. [DOI] [PubMed] [Google Scholar]
- 146.Xie M, Lu C, Wang J, McLellan MD, Johnson KJ, Wendl MC, McMichael JF, Schmidt HK, Yellapantula V, Miller CA, Ozenberger BA, Welch JS, Link DC, Walter MJ, Mardis ER, Dipersio JF, Chen F, Wilson RK, Ley TJ, Ding L. Age-related mutations associated with clonal hematopoietic expansion and malignancies. Nat Med. 2014;20:1472–1478. doi: 10.1038/nm.3733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Kolquist KA, Schultz RA, Furrow A, Brown TC, Han JY, Campbell LJ, Wall M, Slovak ML, Shaffer LG, Ballif BC. Microarray-based comparative genomic hybridization of cancer targets reveals novel, recurrent genetic aberrations in the myelodysplastic syndromes. Cancer Genet. 2011;204:603–628. doi: 10.1016/j.cancergen.2011.10.004. [DOI] [PubMed] [Google Scholar]
- 148.Li Y, Zhu B. Acute myeloid leukemia with DNMT3A mutations. Leuk Lymphoma. 2014;55:2002–2012. doi: 10.3109/10428194.2013.869802. [DOI] [PubMed] [Google Scholar]
- 149.Waitkus MS, Diplas BH, Yan H. Biological role and therapeutic potential of IDH mutations in cancer. Cancer Cell. 2018;34:186–195. doi: 10.1016/j.ccell.2018.04.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Rakheja D, Medeiros LJ, Bevan S, Chen W. The emerging role of d-2-hydroxyglutarate as an oncometabolite in hematolymphoid and central nervous system neoplasms. Front Oncol. 2013;3:169. doi: 10.3389/fonc.2013.00169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Mondesir J, Willekens C, Touat M, de Botton S. IDH1 and IDH2 mutations as novel therapeutic targets: current perspectives. J Blood Med. 2016;7:171–80. doi: 10.2147/JBM.S70716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152.Marcucci G, Maharry K, Wu YZ, Radmacher MD, Mrozek K, Margeson D, Holland KB, Whitman SP, Becker H, Schwind S, Metzeler KH, Powell BL, Carter TH, Kolitz JE, Wetzler M, Carroll AJ, Baer MR, Caligiuri MA, Larson RA, Bloomfield CD. IDH1 and IDH2 gene mutations identify novel molecular subsets within de novo cytogenetically normal acute myeloid leukemia: a cancer and leukemia group B study. J. Clin. Oncol. 2010;28:2348–2355. doi: 10.1200/JCO.2009.27.3730. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Patnaik MM, Hanson CA, Hodnefield JM, Lasho TL, Finke CM, Knudson RA, Ketterling RP, Pardanani A, Tefferi A. Differential prognostic effect of IDH1 versus IDH2 mutations in myelodysplastic syndromes: a Mayo clinic study of 277 patients. Leukemia. 2012;26:101–105. doi: 10.1038/leu.2011.298. [DOI] [PubMed] [Google Scholar]
- 154.Molenaar RJ, Thota S, Nagata Y, Patel B, Clemente M, Przychodzen B, Hirsh C, Viny AD, Hosano N, Bleeker FE, Meggendorfer M, Alpermann T, Shiraishi Y, Chiba K, Tanaka H, van Noorden CJ, Radivoyevitch T, Carraway HE, Makishima H, Miyano S, Sekeres MA, Ogawa S, Haferlach T, Maciejewski JP. Clinical and biological implications of ancestral and non-ancestral IDH1 and IDH2 mutations in myeloid neoplasms. Leukemia. 2015;29:2134–2142. doi: 10.1038/leu.2015.91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Boissel N, Nibourel O, Renneville A, Huchette P, Dombret H, Preudhomme C. Differential prognosis impact of IDH2 mutations in cytogenetically normal acute myeloid leukemia. Blood. 2011;117:3696–3697. doi: 10.1182/blood-2010-11-320937. [DOI] [PubMed] [Google Scholar]
- 156.Green CL, Evans CM, Zhao L, Hills RK, Burnett AK, Linch DC, Gale RE. The prognostic significance of IDH2 mutations in AML depends on the location of the mutation. Blood. 2011;118:409–412. doi: 10.1182/blood-2010-12-322479. [DOI] [PubMed] [Google Scholar]
- 157.Im AP, Sehgal AR, Carroll MP, Smith BD, Tefferi A, Johnson DE, Boyiadzis M. DNMT3A and IDH mutations in acute myeloid leukemia and other myeloid malignancies: associations with prognosis and potential treatment strategies. Leukemia. 2014;28:1774–1783. doi: 10.1038/leu.2014.124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Lin CC, Hou HA, Chou WC, Kuo YY, Liu CY, Chen CY, Lai YJ, Tseng MH, Huang CF, Chiang YC, Lee FY, Liu MC, Liu CW, Tang JL, Yao M, Huang SY, Ko BS, Wu SJ, Tsay W, Chen YC, Tien HF. IDH mutations are closely associated with mutations of DNMT3A, ASXL1 and SRSF2 in patients with myelodysplastic syndromes and are stable during disease evolution. Am J Hematol. 2014;89:137–144. doi: 10.1002/ajh.23596. [DOI] [PubMed] [Google Scholar]
- 159.Thol F, Weissinger EM, Krauter J, Wagner K, Damm F, Wichmann M, Gohring G, Schumann C, Bug G, Ottmann O, Hofmann WK, Schlegelberger B, Ganser A, Heuser M. IDH1 mutations in patients with myelodysplastic syndromes are associated with an unfavorable prognosis. Haematologica. 2010;95:1668–1674. doi: 10.3324/haematol.2010.025494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Yang L, Rau R, Goodell MA. DNMT3A in haematological malignancies. Nat Rev Cancer. 2015;15:152–165. doi: 10.1038/nrc3895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161.Nassereddine S, Lap CJ, Haroun F, Tabbara I. The role of mutant IDH1 and IDH2 inhibitors in the treatment of acute myeloid leukemia. Ann Hematol. 2017;96:1983–1991. doi: 10.1007/s00277-017-3161-0. [DOI] [PubMed] [Google Scholar]
- 162.Ye D, Xiong Y, Guan KL. The mechanisms of IDH mutations in tumorigenesis. Cell Res. 2012;22:1102–1104. doi: 10.1038/cr.2012.51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163.Cohen AL, Holmen SL, Colman H. IDH1 and IDH2 mutations in gliomas. Curr Neurol Neurosci Rep. 2013;13:345. doi: 10.1007/s11910-013-0345-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164.Ok CY, Loghavi S, Sui D, Wei P, Kanagal-Shamanna R, Yin CC, Zuo Z, Routbort MJ, Tang G, Tang Z, Jorgensen JL, Luthra R, Ravandi F, Kantarjian HM, DiNardo CD, Medeiros LJ, Wang SA, Patel KP. Persistent IDH1/2 mutations in remission can predict relapse in patients with acute myeloid leukemia. Haematologica. 2019;104:305–311. doi: 10.3324/haematol.2018.191148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 165.Sparmann A, van Lohuizen M. Polycomb silencers control cell fate, development and cancer. Nat Rev Cancer. 2006;6:846–856. doi: 10.1038/nrc1991. [DOI] [PubMed] [Google Scholar]
- 166.Ernst T, Chase AJ, Score J, Hidalgo-Curtis CE, Bryant C, Jones AV, Waghorn K, Zoi K, Ross FM, Reiter A, Hochhaus A, Drexler HG, Duncombe A, Cervantes F, Oscier D, Boultwood J, Grand FH, Cross NC. Inactivating mutations of the histone methyltransferase gene EZH2 in myeloid disorders. Nat Genet. 2010;42:722–726. doi: 10.1038/ng.621. [DOI] [PubMed] [Google Scholar]
- 167.Chase A, Cross NC. Aberrations of EZH2 in cancer. Clin Cancer Res. 2011;17:2613–2618. doi: 10.1158/1078-0432.CCR-10-2156. [DOI] [PubMed] [Google Scholar]
- 168.Gelsi-Boyer V, Brecqueville M, Devillier R, Murati A, Mozziconacci MJ, Birnbaum D. Mutations in ASXL1 are associated with poor prognosis across the spectrum of malignant myeloid diseases. J Hematol Oncol. 2012;5:12. doi: 10.1186/1756-8722-5-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 169.Scheuermann JC, de Ayala Alonso AG, Oktaba K, Ly-Hartig N, McGinty RK, Fraterman S, Wilm M, Muir TW, Muller J. Histone H2A deubiquitinase activity of the polycomb repressive complex PR-DUB. Nature. 2010;465:243–247. doi: 10.1038/nature08966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170.Cho YS, Kim EJ, Park UH, Sin HS, Um SJ. Additional sex comb-like 1 (ASXL1), in cooperation with SRC-1, acts as a ligand-dependent coactivator for retinoic acid receptor. J Biol Chem. 2006;281:17588–17598. doi: 10.1074/jbc.M512616200. [DOI] [PubMed] [Google Scholar]
- 171.Boultwood J, Perry J, Pellagatti A, Fernandez-Mercado M, Fernandez-Santamaria C, Calasanz MJ, Larrayoz MJ, Garcia-Delgado M, Giagounidis A, Malcovati L, Della Porta MG, Jadersten M, Killick S, Hellstrom-Lindberg E, Cazzola M, Wainscoat JS. Frequent mutation of the polycomb-associated gene ASXL1 in the myelodysplastic syndromes and in acute myeloid leukemia. Leukemia. 2010;24:1062–1065. doi: 10.1038/leu.2010.20. [DOI] [PubMed] [Google Scholar]
- 172.Abdel-Wahab O, Figueroa ME. Interpreting new molecular genetics in myelodysplastic syndromes. Hematology Am Soc Hematol Educ Program. 2012;2012:56–64. doi: 10.1182/asheducation-2012.1.56. [DOI] [PubMed] [Google Scholar]
- 173.Lee TI, Jenner RG, Boyer LA, Guenther MG, Levine SS, Kumar RM, Chevalier B, Johnstone SE, Cole MF, Isono K, Koseki H, Fuchikami T, Abe K, Murray HL, Zucker JP, Yuan B, Bell GW, Herbolsheimer E, Hannett NM, Sun K, Odom DT, Otte AP, Volkert TL, Bartel DP, Melton DA, Gifford DK, Jaenisch R, Young RA. Control of developmental regulators by Polycomb in human embryonic stem cells. Cell. 2006;125:301–313. doi: 10.1016/j.cell.2006.02.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 174.Carbuccia N, Murati A, Trouplin V, Brecqueville M, Adelaide J, Rey J, Vainchenker W, Bernard OA, Chaffanet M, Vey N, Birnbaum D, Mozziconacci MJ. Mutations of ASXL1 gene in myeloproliferative neoplasms. Leukemia. 2009;23:2183–2186. doi: 10.1038/leu.2009.141. [DOI] [PubMed] [Google Scholar]
- 175.Sheng MY, Zhou Y, Xu MJ, Yang FC. Role of ASXL1 mutation in myeloid malignancies. Zhongguo Shi Yan Xue Ye Xue Za Zhi. 2014;22:1183–1187. doi: 10.7534/j.issn.1009-2137.2014.04.057. [DOI] [PubMed] [Google Scholar]
- 176.Gelsi-Boyer V, Trouplin V, Adelaide J, Bonansea J, Cervera N, Carbuccia N, Lagarde A, Prebet T, Nezri M, Sainty D, Olschwang S, Xerri L, Chaffanet M, Mozziconacci MJ, Vey N, Birnbaum D. Mutations of polycomb-associated gene ASXL1 in myelodysplastic syndromes and chronic myelomonocytic leukaemia. Br J Haematol. 2009;145:788–800. doi: 10.1111/j.1365-2141.2009.07697.x. [DOI] [PubMed] [Google Scholar]
- 177.Gelsi-Boyer V, Trouplin V, Roquain J, Adélaïde J, Carbuccia N, Esterni B, Finetti P, Murati A, Arnoulet C, Zerazhi H, Fezoui H, Tadrist Z, Nezri M, Chaffanet M, Mozziconacci MJ, Vey N, Birnbaum D. ASXL1 mutation is associated with poor prognosis and acute transformation in chronic myelomonocytic leukaemia. Br J Haematol. 2010;151:365–375. doi: 10.1111/j.1365-2141.2010.08381.x. [DOI] [PubMed] [Google Scholar]
- 178.Visconte V, O Nakashima M, J Rogers H. Mutations in splicing factor genes in myeloid malignancies: significance and impact on clinical features. Cancers (Basel) 2019;11:1844. doi: 10.3390/cancers11121844. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 179.Makishima H, Visconte V, Sakaguchi H, Jankowska AM, Abu Kar S, Jerez A, Przychodzen B, Bupathi M, Guinta K, Afable MG, Sekeres MA, Padgett RA, Tiu RV, Maciejewski JP. Mutations in the spliceosome machinery, a novel and ubiquitous pathway in leukemogenesis. Blood. 2012;119:3203–3210. doi: 10.1182/blood-2011-12-399774. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 180.Wu SJ, Kuo YY, Hou HA, Li LY, Tseng MH, Huang CF, Lee FY, Liu MC, Liu CW, Lin CT, Chen CY, Chou WC, Yao M, Huang SY, Ko BS, Tang JL, Tsay W, Tien HF. The clinical implication of SRSF2 mutation in patients with myelodysplastic syndrome and its stability during disease evolution. Blood. 2012;120:3106–3111. doi: 10.1182/blood-2012-02-412296. [DOI] [PubMed] [Google Scholar]
- 181.Thol F, Kade S, Schlarmann C, Loffeld P, Morgan M, Krauter J, Wlodarski MW, Kolking B, Wichmann M, Gorlich K, Gohring G, Bug G, Ottmann O, Niemeyer CM, Hofmann WK, Schlegelberger B, Ganser A, Heuser M. Frequency and prognostic impact of mutations in SRSF2, U2AF1, and ZRSR2 in patients with myelodysplastic syndromes. Blood. 2012;119:3578–3584. doi: 10.1182/blood-2011-12-399337. [DOI] [PubMed] [Google Scholar]
- 182.Padgett RA. New connections between splicing and human disease. Trends Genet. 2012;28:147–154. doi: 10.1016/j.tig.2012.01.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 183.Mortera-Blanco T, Dimitriou M, Woll PS, Karimi M, Elvarsdottir E, Conte S, Tobiasson M, Jansson M, Douagi I, Moarii M, Saft L, Papaemmanuil E, Jacobsen SEW, Hellström-Lindberg E. SF3B1-initiating mutations in MDS-RSs target lymphomyeloid hematopoietic stem cells. Blood. 2017;130:881–890. doi: 10.1182/blood-2017-03-776070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 184.Mian SA, Rouault-Pierre K, Smith AE, Seidl T, Pizzitola I, Kizilors A, Kulasekararaj AG, Bonnet D, Mufti GJ. SF3B1 mutant MDS-initiating cells may arise from the haematopoietic stem cell compartment. Nat Commun. 2015;6:10004. doi: 10.1038/ncomms10004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 185.Dorsheimer L, Assmus B, Rasper T, Ortmann CA, Ecke A, Abou-El-Ardat K, Schmid T, Brune B, Wagner S, Serve H, Hoffmann J, Seeger F, Dimmeler S, Zeiher AM, Rieger MA. Association of mutations contributing to clonal hematopoiesis with prognosis in chronic ischemic heart failure. JAMA Cardiol. 2019;4:25–33. doi: 10.1001/jamacardio.2018.3965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 186.Patnaik MM, Lasho TL, Hodnefield JM, Knudson RA, Ketterling RP, Garcia-Manero G, Steensma DP, Pardanani A, Hanson CA, Tefferi A. SF3B1 mutations are prevalent in myelodysplastic syndromes with ring sideroblasts but do not hold independent prognostic value. Blood. 2012;119:569–572. doi: 10.1182/blood-2011-09-377994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 187.Graubert TA, Shen D, Ding L, Okeyo-Owuor T, Lunn CL, Shao J, Krysiak K, Harris CC, Koboldt DC, Larson DE, McLellan MD, Dooling DJ, Abbott RM, Fulton RS, Schmidt H, Kalicki-Veizer J, O’Laughlin M, Grillot M, Baty J, Heath S, Frater JL, Nasim T, Link DC, Tomasson MH, Westervelt P, DiPersio JF, Mardis ER, Ley TJ, Wilson RK, Walter MJ. Recurrent mutations in the U2AF1 splicing factor in myelodysplastic syndromes. Nat Genet. 2011;44:53–57. doi: 10.1038/ng.1031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 188.Shirai CL, Ley JN, White BS, Kim S, Tibbitts J, Shao J, Ndonwi M, Wadugu B, Duncavage EJ, Okeyo-Owuor T, Liu T, Griffith M, McGrath S, Magrini V, Fulton RS, Fronick C, O’Laughlin M, Graubert TA, Walter MJ. Mutant U2AF1 expression alters hematopoiesis and pre-mRNA splicing in vivo. Cancer Cell. 2015;27:631–643. doi: 10.1016/j.ccell.2015.04.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 189.Lane DP. Cancer. p53, guardian of the genome. Nature. 1992;358:15–16. doi: 10.1038/358015a0. [DOI] [PubMed] [Google Scholar]
- 190.Wong TN, Ramsingh G, Young AL, Miller CA, Touma W, Welch JS, Lamprecht TL, Shen D, Hundal J, Fulton RS, Heath S, Baty JD, Klco JM, Ding L, Mardis ER, Westervelt P, DiPersio JF, Walter MJ, Graubert TA, Ley TJ, Druley T, Link DC, Wilson RK. Role of TP53 mutations in the origin and evolution of therapy-related acute myeloid leukaemia. Nature. 2015;518:552–555. doi: 10.1038/nature13968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 191.Seifert H, Mohr B, Thiede C, Oelschlagel U, Schakel U, Illmer T, Soucek S, Ehninger G, Schaich M Study Alliance Leukemia (SAL) The prognostic impact of 17p (p53) deletion in 2272 adults with acute myeloid leukemia. Leukemia. 2009;23:656–663. doi: 10.1038/leu.2008.375. [DOI] [PubMed] [Google Scholar]
- 192.Kadia TM, Jain P, Ravandi F, Garcia-Manero G, Andreef M, Takahashi K, Borthakur G, Jabbour E, Konopleva M, Daver NG, Dinardo C, Pierce S, Kanagal-Shamanna R, Patel K, Estrov Z, Cortes J, Kantarjian HM. TP53 mutations in newly diagnosed acute myeloid leukemia: clinicomolecular characteristics, response to therapy, and outcomes. Cancer. 2016;122:3484–3491. doi: 10.1002/cncr.30203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 193.Prochazka KT, Pregartner G, Rücker FG, Heitzer E, Pabst G, Wolfler A, Zebisch A, Berghold A, Dohner K, Sill H. Clinical implications of subclonal TP53 mutations in acute myeloid leukemia. Haematologica. 2019;104:516–523. doi: 10.3324/haematol.2018.205013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 194.Bouaoun L, Sonkin D, Ardin M, Hollstein M, Byrnes G, Zavadil J, Olivier M. TP53 variations in human cancers: new lessons from the IARC TP53 database and genomics data. Hum Mutat. 2016;37:865–876. doi: 10.1002/humu.23035. [DOI] [PubMed] [Google Scholar]
- 195.Song WJ, Sullivan MG, Legare RD, Hutchings S, Tan X, Kufrin D, Ratajczak J, Resende IC, Haworth C, Hock R, Loh M, Felix C, Roy DC, Busque L, Kurnit D, Willman C, Gewirtz AM, Speck NA, Bushweller JH, Li FP, Gardiner K, Poncz M, Maris JM, Gilliland DG. Haploinsufficiency of CBFA2 causes familial thrombocytopenia with propensity to develop acute myelogenous leukaemia. Nat Genet. 1999;23:166–175. doi: 10.1038/13793. [DOI] [PubMed] [Google Scholar]
- 196.Harada H, Harada Y, Niimi H, Kyo T, Kimura A, Inaba T. High incidence of somatic mutations in the AML1/RUNX1 gene in myelodysplastic syndrome and low blast percentage myeloid leukemia with myelodysplasia. Blood. 2004;103:2316–2324. doi: 10.1182/blood-2003-09-3074. [DOI] [PubMed] [Google Scholar]
- 197.Steensma DP, Gibbons RJ, Mesa RA, Tefferi A, Higgs DR. Somatic point mutations in RUNX1/CBFA2/AML1 are common in high-risk myelodysplastic syndrome, but not in myelofibrosis with myeloid metaplasia. Eur J Haematol. 2005;74:47–53. doi: 10.1111/j.1600-0609.2004.00363.x. [DOI] [PubMed] [Google Scholar]
- 198.Huynh KD, Fischle W, Verdin E, Bardwell VJ. BCoR, a novel corepressor involved in BCL-6 repression. Genes Dev. 2000;14:1810–1823. [PMC free article] [PubMed] [Google Scholar]
- 199.Bera R, Chiu MC, Huang YJ, Lin TH, Kuo MC, Shih LY. RUNX1 mutations promote leukemogenesis of myeloid malignancies in ASXL1-mutated leukemia. J Hematol Oncol. 2019;12:104. doi: 10.1186/s13045-019-0789-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 200.Leung AY, Man CH, Kwong YL. FLT3 inhibition: a moving and evolving target in acute myeloid leukaemia. Leukemia. 2013;27:260–268. doi: 10.1038/leu.2012.195. [DOI] [PubMed] [Google Scholar]
- 201.He BL, Shi X, Man CH, Ma AC, Ekker SC, Chow HC, So CW, Choi WW, Zhang W, Zhang Y, Leung AY. Functions of flt3 in zebrafish hematopoiesis and its relevance to human acute myeloid leukemia. Blood. 2014;123:2518–2529. doi: 10.1182/blood-2013-02-486688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 202.Man CH, Fung TK, Ho C, Han HH, Chow HC, Ma AC, Choi WW, Lok S, Cheung AM, Eaves C, Kwong YL, Leung AY. Sorafenib treatment of FLT3-ITD(+) acute myeloid leukemia: favorable initial outcome and mechanisms of subsequent nonresponsiveness associated with the emergence of a D835 mutation. Blood. 2012;119:5133–5143. doi: 10.1182/blood-2011-06-363960. [DOI] [PubMed] [Google Scholar]
- 203.Santos FP, Jones D, Qiao W, Cortes JE, Ravandi F, Estey EE, Verma D, Kantarjian H, Borthakur G. Prognostic value of FLT3 mutations among different cytogenetic subgroups in acute myeloid leukemia. Cancer. 2011;117:2145–2155. doi: 10.1002/cncr.25670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 204.Stanchina M, Soong D, Zheng-Lin B, Watts JM, Taylor J. Advances in acute myeloid leukemia: recently approved therapies and drugs in development. Cancers (Basel) 2020;12:3225. doi: 10.3390/cancers12113225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 205.Short NJ, Konopleva M, Kadia TM, Borthakur G, Ravandi F, DiNardo CD, Daver N. Advances in the treatment of acute myeloid leukemia: new drugs and new challenges. Cancer Discov. 2020;10:506–525. doi: 10.1158/2159-8290.CD-19-1011. [DOI] [PubMed] [Google Scholar]
- 206.Yates JW, Wallace HJ Jr, Ellison RR, Holland JF. Cytosine arabinoside (NSC-63878) and daunorubicin (NSC-83142) therapy in acute nonlymphocytic leukemia. Cancer Chemother Rep. 1973;57:485–488. [PubMed] [Google Scholar]
- 207.Lowenberg B, Ossenkoppele GJ, van Putten W, Schouten HC, Graux C, Ferrant A, Sonneveld P, Maertens J, Jongen-Lavrencic M, von Lilienfeld-Toal M, Biemond BJ, Vellenga E, van Marwijk Kooy M, Verdonck LF, Beck J, Dohner H, Gratwohl A, Pabst T, Verhoef G Dutch-Belgian Cooperative Trial Group for Hemato-Oncology (HOVON); German AML Study Group (AMLSG); Swiss Group for Clinical Cancer Research (SAKK) Collaborative Group. High-dose daunorubicin in older patients with acute myeloid leukemia. N Engl J Med. 2009;361:1235–1248. doi: 10.1056/NEJMoa0901409. [DOI] [PubMed] [Google Scholar]
- 208.Boddu P, Kantarjian HM, Garcia-Manero G, Ravandi F, Verstovsek S, Jabbour E, Borthakur G, Konopleva M, Bhalla KN, Daver N, DiNardo CD, Benton CB, Takahashi K, Estrov Z, Pierce SR, Andreeff M, Cortes JE, Kadia TM. Treated secondary acute myeloid leukemia: a distinct high-risk subset of AML with adverse prognosis. Blood Adv. 2017;1:1312–1323. doi: 10.1182/bloodadvances.2017008227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 209.Tsuji K, Wang YH, Takanashi M, Odajima T, Lee GA, Sugimori H, Motoji T. Overexpression of lung resistance-related protein and P-glycoprotein and response to induction chemotherapy in acute myelogenous leukemia. Hematol Rep. 2012;4:e18. doi: 10.4081/hr.2012.e18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 210.Zeichner SB, Arellano ML. Secondary adult acute myeloid leukemia: a review of our evolving understanding of a complex disease process. Curr Treat Options Oncol. 2015;16:1–5. doi: 10.1007/s11864-015-0355-3. [DOI] [PubMed] [Google Scholar]
- 211.De Kouchkovsky I, Abdul-Hay M. Acute myeloid leukemia: a comprehensive review and 2016 update. Blood Cancer J. 2016;6:e441. doi: 10.1038/bcj.2016.50. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 212.Boddu PC, Kantarjian HM, Ravandi F, Garcia-Manero G, Verstovsek S, Jabbour EJ, Takahashi K, Bhalla K, Konopleva M, DiNardo CD, Ohanian M, Pemmaraju N, Jain N, Pierce S, Wierda WG, Cortes JE, Kadia TM. Characteristics and outcomes of older patients with secondary acute myeloid leukemia according to treatment approach. Cancer. 2017;123:3050–3060. doi: 10.1002/cncr.30704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 213.Tang FF, Huang XJ, Zhang XH, Chen H, Chen YH, Han W, Chen Y, Wang FR, Wang Y, Wang JZ, Yan CH, Sun YQ, Mo XD, Liu KY, Xu LP. Allogeneic hematopoietic cell transplantation for adult patients with treatment-related acute myeloid leukemia during first remission: Comparable to de novo acute myeloid leukemia. Leuk Res. 2016;47:8–15. doi: 10.1016/j.leukres.2016.05.005. [DOI] [PubMed] [Google Scholar]
- 214.Dohner H, Estey E, Grimwade D, Amadori S, Appelbaum FR, Buchner T, Dombret H, Ebert BL, Fenaux P, Larson RA, Levine RL, Lo-Coco F, Naoe T, Niederwieser D, Ossenkoppele GJ, Sanz M, Sierra J, Tallman MS, Tien HF, Wei AH, Lowenberg B, Bloomfield CD. Diagnosis and management of AML in adults: 2017 ELN recommendations from an international expert panel. Blood. 2017;129:424–447. doi: 10.1182/blood-2016-08-733196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 215.Li Z, Labopin M, Ciceri F, Blaise D, Tischer J, Ehninger G, Van Lint MT, Koc Y, Santarone S, Forcade E, Castagna L. Haploidentical transplantation outcomes for secondary acute myeloid leukemia: acute leukemia working party (ALWP) of the European society for blood and marrow transplantation (EBMT) study. Am J Hematol. 2018;93:769–777. doi: 10.1002/ajh.25087. [DOI] [PubMed] [Google Scholar]
- 216.Garzon R, Savona M, Baz R, Andreeff M, Gabrail N, Gutierrez M, Savoie L, Mau-Sorensen PM, Wagner-Johnston N, Yee K, Unger TJ, Saint-Martin JR, Carlson R, Rashal T, Kashyap T, Klebanov B, Shacham S, Kauffman M, Stone R. A phase 1 clinical trial of single-agent selinexor in acute myeloid leukemia. Blood. 2017;129:3165–3174. doi: 10.1182/blood-2016-11-750158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 217.Martelli MP, Gionfriddo I, Mezzasoma F, Milano F, Pierangeli S, Mulas F, Pacini R, Tabarrini A, Pettirossi V, Rossi R, Vetro C, Brunetti L, Sportoletti P, Tiacci E, Di Raimondo F, Falini B. Arsenic trioxide and all-trans retinoic acid target NPM1 mutant oncoprotein levels and induce apoptosis in NPM1-mutated AML cells. Blood. 2015;125:3455–3465. doi: 10.1182/blood-2014-11-611459. [DOI] [PubMed] [Google Scholar]
- 218.Karathedath S, Rajamani BM, Musheer Aalam SM, Abraham A, Varatharajan S, Krishnamurthy P, Mathews V, Velayudhan SR, Balasubramanian P. Role of NF-E2 related factor 2 (Nrf2) on chemotherapy resistance in acute myeloid leukemia (AML) and the effect of pharmacological inhibition of Nrf2. PLoS One. 2017;12:e0177227. doi: 10.1371/journal.pone.0177227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 219.Lancet JE, Uy GL, Cortes JE, Newell LF, Lin TL, Ritchie EK, Stuart RK, Strickland SA, Hogge D, Solomon SR, Stone RM, Bixby DL, Kolitz JE, Schiller GJ, Wieduwilt MJ, Ryan DH, Hoering A, Banerjee K, Chiarella M, Louie AC, Medeiros BC. CPX-351 (cytarabine and daunorubicin) liposome for injection versus conventional cytarabine plus daunorubicin in older patients with newly diagnosed secondary acute myeloid leukemia. J. Clin. Oncol. 2018;36:2684–2692. doi: 10.1200/JCO.2017.77.6112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 220.Brunetti C, Anelli L, Zagaria A, Specchia G, Albano F. CPX-351 in acute myeloid leukemia: can a new formulation maximize the efficacy of old compounds? Expert Rev Hematol. 2017;10:853–862. doi: 10.1080/17474086.2017.1369400. [DOI] [PubMed] [Google Scholar]
- 221.Godley LA, Njiaju UO, Green M, Weiner H, Lin S, Odenike O, Rich ES, Artz A, Van Besien K, Daugherty CK, Zhang Y, Le Beau MM, Stock W, Larson RA. Treatment of therapy-related myeloidneoplasms with high-dose cytarabine/mitoxantrone followed by hematopoietic stem cell transplant. Leuk Lymphoma. 2010;51:995–1006. doi: 10.3109/10428191003763468. [DOI] [PubMed] [Google Scholar]
- 222.Tardi P, Johnstone S, Harasym N, Xie S, Harasym T, Zisman N, Harvie P, Bermudes D, Mayer L. In vivo maintenance of synergistic cytarabine: daunorubicin ratios greatly enhances therapeutic efficacy. Leuk Res. 2009;33:129–139. doi: 10.1016/j.leukres.2008.06.028. [DOI] [PubMed] [Google Scholar]
- 223.Eloy JO, Claro de Souza M, Petrilli R, Barcellos JP, Lee RJ, Marchetti JM. Liposomes as carriers of hydrophilic small molecule drugs: strategies to enhance encapsulation and delivery. Colloids Surf B Biointerfaces. 2014;123:345–363. doi: 10.1016/j.colsurfb.2014.09.029. [DOI] [PubMed] [Google Scholar]
- 224.Mayer LD, Harasym TO, Tardi PG, Harasym NL, Shew CR, Johnstone SA, Ramsay EC, Bally MB, Janoff AS. Ratiometric dosing of anticancer drug combinations: controlling drug ratios after systemic administration regulates therapeutic activity in tumor-bearing mice. Mol Cancer Ther. 2006;5:1854–1863. doi: 10.1158/1535-7163.MCT-06-0118. [DOI] [PubMed] [Google Scholar]
- 225.Kim HP, Gerhard B, Harasym TO, Mayer LD, Hogge DE. Liposomal encapsulation of a synergistic molar ratio of cytarabine and daunorubicin enhances selective toxicity for acute myeloid leukemia progenitors as compared to analogous normal hematopoietic cells. Exp Hematol. 2011;39:741–750. doi: 10.1016/j.exphem.2011.04.001. [DOI] [PubMed] [Google Scholar]
- 226.Lim WS, Tardi PG, Dos Santos N, Xie X, Fan M, Liboiron BD, Huang X, Harasym TO, Bermudes D, Mayer LD. Leukemia-selective uptake and cytotoxicity of CPX-351, a synergistic fixed-ratio cytarabine: daunorubicin formulation, in bone marrow xenografts. Leuk Res. 2010;34:1214–1223. doi: 10.1016/j.leukres.2010.01.015. [DOI] [PubMed] [Google Scholar]
- 227.Konopleva M, Letai A. BCL-2 inhibition in AML: an unexpected bonus? Blood. 2018;132:1007–1012. doi: 10.1182/blood-2018-03-828269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 228.Konopleva M, Zhao S, Hu W, Jiang S, Snell V, Weidner D, Jackson CE, Zhang X, Champlin R, Estey E, Reed JC, Andreeff M. The anti-apoptotic genes Bcl-X(L) and Bcl-2 are over-expressed and contribute to chemoresistance of non-proliferating leukaemic CD34+ cells. Br J Haematol. 2002;118:521–534. doi: 10.1046/j.1365-2141.2002.03637.x. [DOI] [PubMed] [Google Scholar]
- 229.Konopleva M, Pollyea DA, Potluri J, Chyla B, Hogdal L, Busman T, McKeegan E, Salem AH, Zhu M, Ricker JL, Blum W, DiNardo CD, Kadia T, Dunbar M, Kirby R, Falotico N, Leverson J, Humerickhouse R, Mabry M, Stone R, Kantarjian H, Letai A. Efficacy and biological correlates of response in a phase ii study of venetoclax monotherapy in patients with acute myelogenous leukemia. Cancer Discov. 2016;6:1106–1117. doi: 10.1158/2159-8290.CD-16-0313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 230.Fenaux P, Mufti GJ, Hellström-Lindberg E, Santini V, Gattermann N, Germing U, Sanz G, List AF, Gore S, Seymour JF, Dombret H, Backstrom J, Zimmerman L, McKenzie D, Beach CL, Silverman LR. Azacitidine prolongs overall survival compared with conventional care regimens in elderly patients with low bone marrow blast count acute myeloid leukemia. J. Clin. Oncol. 2010;28:562–569. doi: 10.1200/JCO.2009.23.8329. [DOI] [PubMed] [Google Scholar]
- 231.Dombret H, Seymour JF, Butrym A, Wierzbowska A, Selleslag D, Jang JH, Kumar R, Cavenagh J, Schuh AC, Candoni A, Récher C, Sandhu I, Bernal del Castillo T, Al-Ali HK, Martinelli G, Falantes J, Noppeney R, Stone RM, Minden MD, McIntyre H, Songer S, Lucy LM, Beach CL, Dohner H. International phase 3 study of azacitidine vs conventional care regimens in older patients with newly diagnosed AML with >30% blasts. Blood. 2015;126:291–299. doi: 10.1182/blood-2015-01-621664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 232.Kantarjian HM, Thomas XG, Dmoszynska A, Wierzbowska A, Mazur G, Mayer J, Gau JP, Chou WC, Buckstein R, Cermak J, Kuo CY, Oriol A, Ravandi F, Faderl S, Delaunay J, Lysák D, Minden M, Arthur C. Multicenter, randomized, open-label, phase III trial of decitabine versus patient choice, with physician advice, of either supportive care or low-dose cytarabine for the treatment of older patients with newly diagnosed acute myeloid leukemia. J. Clin. Oncol. 2012;30:2670–2677. doi: 10.1200/JCO.2011.38.9429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 233.Pleyer L, Stauder R, Burgstaller S, Schreder M, Tinchon C, Pfeilstocker M, Steinkirchner S, Melchardt T, Mitrovic M, Girschikofsky M, Lang A, Krippl P, Sliwa T, Egle A, Linkesch W, Voskova D, Angermann H, Greil R. Azacitidine in patients with WHO-defined AML - results of 155 patients from the Austrian Azacitidine Registry of the AGMT-Study Group. J Hematol Oncol. 2013;6:32. doi: 10.1186/1756-8722-6-32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 234.Dumas PY, Bertoli S, Berard E, Mediavilla C, Yon E, Tavitian S, Leguay T, Huguet F, Forcade E, Milpied N, Sarry A, Sauvezie M, Bories P, Pigneux A, Recher C. Azacitidine or intensive chemotherapy for older patients with secondary or therapy-related acute myeloid leukemia. Oncotarget. 2017;8:79126–79136. doi: 10.18632/oncotarget.15988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 235.Welch JS, Petti AA, Miller CA, Fronick CC, O’Laughlin M, Fulton RS, Wilson RK, Baty JD, Duncavage EJ, Tandon B, Lee YS, Wartman LD, Uy GL, Ghobadi A, Tomasson MH, Pusic I, Romee R, Fehniger TA, Stockerl-Goldstein KE, Vij R, Oh ST, Abboud CN, Cashen AF, Schroeder MA, Jacoby MA, Heath SE, Luber K, Janke MR, Hantel A, Khan N, Sukhanova MJ, Knoebel RW, Stock W, Graubert TA, Walter MJ, Westervelt P, Link DC, DiPersio JF, Ley TJ. TP53 and decitabine in acute myeloid leukemia and myelodysplastic syndromes. N Engl J Med. 2016;375:2023–2036. doi: 10.1056/NEJMoa1605949. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 236.Appelbaum FR, Bernstein ID. Gemtuzumab ozogamicin for acute myeloid leukemia. Blood. 2017;130:2373–2376. doi: 10.1182/blood-2017-09-797712. [DOI] [PubMed] [Google Scholar]
- 237.Mihalyova J, Jelinek T, Growkova K, Hrdinka M, Simicek M, Hajek R. Venetoclax: a new wave in hematooncology. Exp Hematol. 2018;61:10–25. doi: 10.1016/j.exphem.2018.02.002. [DOI] [PubMed] [Google Scholar]
- 238.Saygin C, Carraway HE. Emerging therapies for acute myeloid leukemia. J Hematol Oncol. 2017;10:93. doi: 10.1186/s13045-017-0463-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 239.Petersdorf SH, Kopecky KJ, Slovak M, Willman C, Nevill T, Brandwein J, Larson RA, Erba HP, Stiff PJ, Stuart RK, Walter RB, Tallman MS, Stenke L, Appelbaum FR. A phase 3 study of gemtuzumab ozogamicin during induction and postconsolidation therapy in younger patients with acute myeloid leukemia. Blood. 2013;121:4854–4860. doi: 10.1182/blood-2013-01-466706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 240.Jen EY, Ko CW, Lee JE, Del Valle PL, Aydanian A, Jewell C, Norsworthy KJ, Przepiorka D, Nie L, Liu J, Sheth CM, Shapiro M, Farrell AT, Pazdur R. FDA approval: gemtuzumab ozogamicin for the treatment of adults with newly diagnosed CD33-positive acute myeloid leukemia. Clin Cancer Res. 2018;24:3242–3246. doi: 10.1158/1078-0432.CCR-17-3179. [DOI] [PubMed] [Google Scholar]
- 241.Burnett AK, Hills RK, Milligan D, Kjeldsen L, Kell J, Russell NH, Yin JA, Hunter A, Goldstone AH, Wheatley K. Identification of patients with acute myeloblastic leukemia who benefit from the addition of gemtuzumab ozogamicin: results of the MRC AML15 trial. J. Clin. Oncol. 2011;29:369–377. doi: 10.1200/JCO.2010.31.4310. [DOI] [PubMed] [Google Scholar]
- 242.Michelis FV, Atenafu EG, Gupta V, Kim DD, Kuruvilla J, Lipton JH, Loach D, Seftel MD, Uhm J, Alam N, Lambie A, McGillis L, Messner HA. Comparable outcomes post allogeneic hematopoietic cell transplant for patients with de novo or secondary acute myeloid leukemia in first remission. Bone Marrow Transplant. 2015;50:907–913. doi: 10.1038/bmt.2015.59. [DOI] [PubMed] [Google Scholar]
- 243.Castaigne S, Pautas C, Terré C, Raffoux E, Bordessoule D, Bastie JN, Legrand O, Thomas X, Turlure P, Reman O, de Revel T, Gastaud L, de Gunzburg N, Contentin N, Henry E, Marolleau JP, Aljijakli A, Rousselot P, Fenaux P, Preudhomme C, Chevret S, Dombret H Acute Leukemia French Association. Effect of gemtuzumab ozogamicin on survival of adult patients with de-novo acute myeloid leukaemia (ALFA-0701): a randomised, open-label, phase 3 study. Lancet. 2012;379:1508–1516. doi: 10.1016/S0140-6736(12)60485-1. [DOI] [PubMed] [Google Scholar]
- 244.Amadori S, Suciu S, Selleslag D, Aversa F, Gaidano G, Musso M, Annino L, Venditti A, Voso MT, Mazzone C, Magro D, De Fabritiis P, Muus P, Alimena G, Mancini M, Hagemeijer A, Paoloni F, Vignetti M, Fazi P, Meert L, Ramadan SM, Willemze R, de Witte T, Baron F. Gemtuzumab ozogamicin versus best supportive care in older patients with newly diagnosed acute myeloid leukemia unsuitable for intensive chemotherapy: results of the randomized phase III EORTC-GIMEMA AML-19 trial. J. Clin. Oncol. 2016;34:972–979. doi: 10.1200/JCO.2015.64.0060. [DOI] [PubMed] [Google Scholar]
- 245.Cortes JE, Heidel FH, Hellmann A, Fiedler W, Smith BD, Robak T, Montesinos P, Pollyea DA, DesJardins P, Ottmann O, Ma WW. Randomized comparison of low dose cytarabine with or without glasdegib in patients with newly diagnosed acute myeloid leukemia or high-risk myelodysplastic syndrome. Leukemia. 2019;33:379–389. doi: 10.1038/s41375-018-0312-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 246.Aberger F, Hutterer E, Sternberg C, Del Burgo PJ, Hartmann TN. Acute myeloid leukemia-strategies and challenges for targeting oncogenic Hedgehog/GLI signaling. Cell Commun Signal. 2017;15:8. doi: 10.1186/s12964-017-0163-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 247.Campbell V, Copland M. Hedgehog signaling in cancer stem cells: a focus on hematological cancers. Stem Cells Cloning. 2015;8:27. doi: 10.2147/SCCAA.S58613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 248.Stone RM, Manley PW, Larson RA, Capdeville R. Midostaurin: its odyssey from discovery to approval for treating acute myeloid leukemia and advanced systemic mastocytosis. Blood Adv. 2018;2:444–453. doi: 10.1182/bloodadvances.2017011080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 249.Stone RM, Mandrekar SJ, Sanford BL, Laumann K, Geyer S, Bloomfield CD, Thiede C, Prior TW, Döhner K, Marcucci G, Lo-Coco F, Klisovic RB, Wei A, Sierra J, Sanz MA, Brandwein JM, de Witte T, Niederwieser D, Appelbaum FR, Medeiros BC, Tallman MS, Krauter J, Schlenk RF, Ganser A, Serve H, Ehninger G, Amadori S, Larson RA, Döhner H. Midostaurin plus chemotherapy for acute myeloid leukemia with a FLT3 mutation. N Engl J Med. 2017;377:454–64. doi: 10.1056/NEJMoa1614359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 250.Perl AE, Altman JK, Cortes J, Smith C, Litzow M, Baer MR, Claxton D, Erba HP, Gill S, Goldberg S, Jurcic JG, Larson RA, Liu C, Ritchie E, Schiller G, Spira AI, Strickland SA, Tibes R, Ustun C, Wang ES, Stuart R, Röllig C, Neubauer A, Martinelli G, Bahceci E, Levis M. Selective inhibition of FLT3 by gilteritinib in relapsed or refractory acute myeloid leukaemia: a multicentre, first-in-human, open-label, phase 1-2 study. Lancet Oncol. 2017;18:1061–1075. doi: 10.1016/S1470-2045(17)30416-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 251.Marconi G, De Polo S, Martinelli G, Nanni J, Bertamini L, Talami A, Olivi M, Ragaini S, Abbenante MC, Sartor C, Ottaviani E, Bochicchio MT, Parisi S, Fontana MC, Cristiano G, Raffini M, Baldazzi C, Testoni N, Bonifazi F, Paolini S, Curti A, Cavo M, Papayannidis C. Safety profile and impact on survival of tyrosine kinase inhibitors versus conventional therapy in relapse or refractory FLT3 positive acute myeloid leukemia patients. Leuk Res. 2021;101:106497. doi: 10.1016/j.leukres.2020.106497. [DOI] [PubMed] [Google Scholar]
- 252.Stein EM, DiNardo CD, Pollyea DA, Fathi AT, Roboz GJ, Altman JK, Stone RM, DeAngelo DJ, Levine RL, Flinn IW, Kantarjian HM, Collins R, Patel MR, Frankel AE, Stein A, Sekeres MA, Swords RT, Medeiros BC, Willekens C, Vyas P, Tosolini A, Xu Q, Knight RD, Yen KE, Agresta S, de Botton S, Tallman MS. Enasidenib in mutant IDH2 relapsed or refractory acute myeloid leukemia. Blood. 2017;130:722–731. doi: 10.1182/blood-2017-04-779405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 253.Largeaud L, Berard E, Bertoli S, Dufrechou S, Prade N, Gadaud N, Tavitian S, Bories P, Luquet I, Sarry A, De Mas V, Huguet F, Delabesse E, Recher C. Outcome of AML patients with IDH2 mutations in real world before the era of IDH2 inhibitors. Leuk Res. 2019;81:82–87. doi: 10.1016/j.leukres.2019.04.010. [DOI] [PubMed] [Google Scholar]
- 254.Popovici-Muller J, Lemieux RM, Artin E, Saunders JO, Salituro FG, Travins J, Cianchetta G, Cai Z, Zhou D, Cui D, Chen P, Straley K, Tobin E, Wang F, David MD, Penard-Lacronique V, Quivoron C, Saada V, de Botton S, Gross S, Dang L, Yang H, Utley L, Chen Y, Kim H, Jin S, Gu Z, Yao G, Luo Z, Lv X, Fang C, Yan L, Olaharski A, Silverman L, Biller S, Su SM, Yen K. Discovery of AG-120 (ivosidenib): a first-in-class mutant IDH1 inhibitor for the treatment of idh1 mutant cancers. ACS Med Chem Lett. 2018;9:300–305. doi: 10.1021/acsmedchemlett.7b00421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 255.Hofmann S, Schubert ML, Wang L, He B, Neuber B, Dreger P, Müller-Tidow C, Schmitt M. Chimeric antigen receptor (CAR) T cell therapy in acute myeloid leukemia (AML) J Clin Med. 2019;8:200. doi: 10.3390/jcm8020200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 256.Pyzer AR, Avigan DE, Rosenblatt J. Clinical trials of dendritic cell-based cancer vaccines in hematologic malignancies. Hum Vaccin Immunother. 2014;10:3125–3131. doi: 10.4161/21645515.2014.982993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 257.DeStefano CB, Hourigan CS. Personalizing initial therapy in acute myeloid leukemia: incorporating novel agents into clinical practice. Ther Adv Hematol. 2018;9:109–121. doi: 10.1177/2040620718761778. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 258.Wei H, Wang Y, Zhou C, Lin D, Liu B, Liu K, Qiu S, Gong B, Li Y, Zhang G, Wei S, Gong X, Liu Y, Zhao X, Gu R, Mi Y, Wang J. Distinct genetic alteration profiles of acute myeloid leukemia between Caucasian and Eastern Asian population. J Hematol Oncol. 2018;11:18. doi: 10.1186/s13045-018-0566-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 259.Gu R, Yang X, Wei H. Molecular landscape and targeted therapy of acute myeloid leukemia. Biomark Res. 2018;6:32. doi: 10.1186/s40364-018-0146-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 260.Zhang C, Liu J, Zhong JF, Zhang X. Engineering CAR-T cells. Biomark Res. 2017;5:22. doi: 10.1186/s40364-017-0102-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 261.Liu D. Cancer biomarkers for targeted therapy. Biomark Res. 2019;7:25. doi: 10.1186/s40364-019-0178-7. [DOI] [PMC free article] [PubMed] [Google Scholar]


