Abstract
The Kerguelen Islands, located in the southern part of the Indian Ocean, are very isolated geographically. The microbial diversity and communities present on the island, especially associated to geothermal springs, have never been analyzed with high-throughput sequencing methods. In this article, we performed the first metagenomics analysis of microorganisms present in Kerguelen hot springs. From four hot springs, we assembled metagenomes and recovered 42 metagenome-assembled genomes, mostly associated with new putative taxa based on phylogenomic analyses and overall genome relatedness indices. The 42 MAGs were studied in detail and showed putative affiliations to 13 new genomic species and 6 new genera of Bacteria or Archaea according to GTDB. Functional potential of MAGs suggests the presence of thermophiles and hyperthermophiles, as well as heterotrophs and primary producers possibly involved in the sulfur cycle, notably in the oxidation of sulfur compounds. This paper focused on only four of the dozens of hot springs in the Kerguelen Islands and should be considered as a preliminary study of the microorganisms inhabiting the hot springs of these isolated islands. These results show that more efforts should be made towards characterization of Kerguelen Islands ecosystems, as they represent a reservoir of unknown microbial lineages.
Subject terms: Computational biology and bioinformatics, Ecology, Genetics, Microbiology
Introduction
Terrestrial hot springs are found all over the world, on all continents, and are abundant in areas of volcanic activity such as Iceland, Japan, Russia, Chile, Algeria or New Zealand1,2. With possible exception in the polyextreme Dallol area3, all studied geothermal environments harbor microbial cohorts. Microbial cohorts in terrestrial hot springs are often composed of Bacteria belonging to Aquificae, Chloroflexi, Deinococcus-Thermus and Thermotogae, and Archaea belonging to Desulfurococcaceae, Thermoproteaceae and Thermococcaceae (respectively referenced as follows in the Genome Taxonomy Database: Aquificota, Chloroflexota, Deinococcota, Thermotogae, Desulfurococcaceae/Ignisphaeraceae, Thermoproteaceae, and Thermococcaceae)4–6. Most of terrestrial hot springs, like those of Yellowstone (USA) or Kamchatka (Russia) areas, have been the subject of extensive microbiological investigations. These investigations include the study of the microbial community composition, the isolation and physiological characterization of microorganisms, the investigation of adaptive mechanisms of indigenous taxa, and the mining of extremophilic species for potential enzymes, activities or molecules of biotechnological interest (e.g.7–12). Studies of bacteria and archaea living in geothermal systems are essential for our knowledge of the history of life, as these environments are early Earth analogs and one of the possible cradles of life2,6,13,14. The microbial communities of hot springs in the polar regions are partly different in their composition. For example, the fumaroles of Deception Island (Antarctica) contain prokaryotic taxa belonging to Verrucomicrobia, Proteobacteria, Planctomycetes, ‘Candidatus Parcubacteria’, Firmicutes, Chloroflexi, Calditrichaeota, Bacteroidetes, Thaumarchaeota, Nanoarchaeota, Euryarchaeota and Crenarchaeota (respectively referenced as follows in the Genome Taxonomy Database: Verrucomicrobiota, Proteobacteria, Planctomycetota, Patescibacteria, Firmicutes, Chloroflexota, Calditrichota, Bacteroidota, Thermoproteota, Nanoarchaeota, Methanobacteriota and Thermoproteota)15–17. Due to their outstanding ecological and scientific values, specifically the presence of a wide diversity of rare endemic species or under-documented taxa, Deception Island and the other Antarctic geothermal ecosystems have been designated as Antarctic Special Protected Areas (ASPAs) by the Antarctic Treaty (management plan for Antarctic Specially protected area N° 175 and N° 140)18.
The volcanic Kerguelen Archipelago, which is part of the French Southern and Antarctic Lands, is situated in the southern part of the Indian Ocean (49° S, 69° E). Located at 3300 km from the nearest inhabited areas, it is amongst the most isolated islands from any continental landmass and contain a large part of the limited terrestrial habitats present at these latitudes. The Kerguelen Islands have the status of protected areas and are under the umbrella of international conventions supporting biodiversity protection (i.e., CITES, IUCN, Convention on Biological Diversity, IPBES). They represent an important UNESCO’s world heritage site and belongs to the national nature reserve (NNR) of the French Southern Lands. Such as the ASPAs, most of Kerguelen geothermal terrestrial micro-habitats are located within “strictly protected areas” (ZPI) set aside by the Nature Reserve, in order to protect biodiversity, and geological/geomorphological features. The Kerguelen Islands and the active volcanic Heard and MacDonald Islands are the only emerged entities among the vast Kerguelen oceanic Plateau. Kerguelen Archipelago is the third largest volcanic island complex in the world, after Iceland and Hawaii19. The last volcanic activity, dated 26 ± 3 Ka, took place on the Rallier du Baty (RB) Peninsula in the south-western part of the Kerguelen Islands20. Current volcanic activity, due to the Kerguelen hotspot, is evidenced by fumaroles, mud pots, hydrothermal discharges and small hot springs that rise from sea level to at least 300 m altitude. The waters of these geothermal systems contain large amounts of dissolved minerals and have wide pH ranges, from acidic to alkaline, and temperature ranges, from 35 to over 100 °C. The geochemical properties of the most accessible parts of this system have been monitored more or less regularly over the last decades21,22.
These geothermal habitats represent unique biodiversity sanctuaries in very insulated polar environments. Preliminary investigations based on 16S rRNA gene amplicon cloning and sequencing revealed a diverse collection of microbial community lineages composed of Proteobacteria, Deinococcus-Thermus, Chloroflexi, Firmicutes, Actinobacteria or Aquificae, as well as Euryarchaeota, Crenarchaeota (Thermoproteales, Desulfurococcales, Acidilobales, Sulfolobales) and Thaumarchaeota (respectively referenced as follows in the Genome Taxonomy Database: Proteobacteria, Deinococcota, Chloroflexota, Firmicutes, Actinobacteriota, Aquificota, Methanobacteriota, Thermoproteota and Thermoproteota)23,24. Some of these lineages were also found at geothermal sites in Antarctica, and others represented common taxa in geochemical environments worldwide. A small number of new species have also been isolated, enriched or highlighted through molecular approaches from these regions23,25. Apart from these few studies, this area has not been subjected to any comprehensive microbiological investigation to date. The microbial diversity hosted in these hot springs remains largely unknown, as well as its functional potential (metabolism, physiology, adaptations). Microbial communities might be shaped by the biogeographic position and the physicochemical parameters of the hot springs (temperature, pH, in situ chemistry), that probably exert a strong selective pressure on indigenous communities17. Yet, these geothermal springs represent undoubtedly unique diversity, and reservoirs of new functions and innovation.
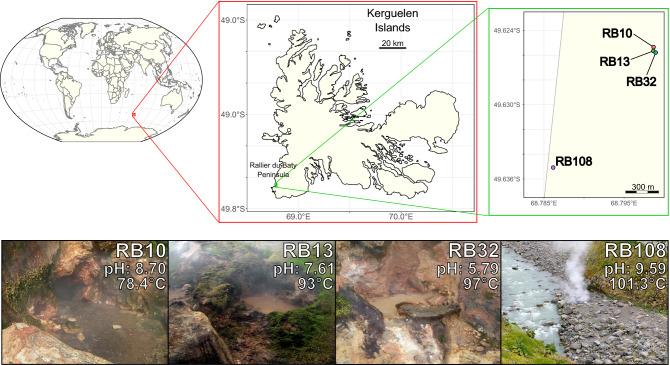
In this study, we focused on four small geothermal hot springs from the Kerguelen Islands whose microbial communities have never been studied before. We analyzed the metagenomes of the hot springs referenced as RB10, RB13 and RB32, located on the "plateau des Fumerolles" at an altitude of about 300 m on the west coast of the Rallier du Baty Peninsula, and of the ephemeral spring referenced as RB108 which flows slightly above sea level into the riverbed of the Infernet glacier (located at the base of the plateau des Fumerolles) (Fig. 1). This work was carried out from samples collected with the aim of cultivating thermophilic microorganisms, which had been stored at 4 °C for 2 years. Those storage conditions induce biases and limit the scientific discussion. Therefore, the generated data did not allow ecological or evolutionary interpretations. These samples were collected in a particularly isolated natural reserve that requires specific sampling permits, and are therefore very difficult to obtain, making them particularly valuable. This descriptive study provides the first sequenced and analyzed MAGs from these undisturbed areas and points out that more efforts must be made to characterize Kerguelen Island ecosystems. This insular area contains a reservoir of unknown microbial lineages, and thus possibly biological and genomic novelties, the study of which may prove useful in the future to discover new metabolic pathways or molecules and to understand the part of deterministic and stochastic processes underlying microbial assemblages.
Figure 1.
Sampling locations at the plateau des Fumerolles in the Rallier du Baty Peninsula (Kerguelen Islands, French Southern and Antarctic Lands), and photographs of the four hot springs studied here with their associated temperature and pH conditions. The world map and details of the Kerguelen seashore were generated using the ‘cowplot’, ‘ggplot2’, ‘lwgeom’, ‘rnaturalearth’, ‘rnaturalearthdata’, ‘rworldmap’, ‘sf’ and ‘tidyverse’ libraries implemented on the RStudio software (v1.2.5001). A 2018 map of the Kerguelen islands was retrieved from the French national portal of maritime limits (http://limitesmaritimes.gouv.fr/) according to the 2017 Etalab open license (v2.0—https://www.etalab.gouv.fr/). Pictures depicting sampling sites were taken by Marc Le Romancer. Layout was carried out using the free open source vector image editor Inkscape (v1.0).
Results and discussion
MAG binning and general features
From the four hot springs, we assembled four associated metagenomes and then binned a total of 42 MAGs. We recovered 12 MAGs from RB10 hot spring, 13 from RB13, 14 from RB32 and 3 from RB108. Out of these 42 MAGs, 7 were of high-quality, 25 of nearly-high quality, 9 of medium quality and 1 of low quality (Table 1) based on metagenomic standards26. The GC% was quite variable, ranging from 25.76 to 70.35% among all MAGs and between 32.15 and 69.21% only among the high- and near high-quality MAGs. With the exception of RB108 from which we only recovered bacterial MAGs, we retrieved both bacterial and archaeal MAGs in the other hot springs. Two thirds of the MAGs (26/42) were assigned to the domain Bacteria and the rest to the domain Archaea (16/42) (Table 2).
Table 1.
General characteristics of the 42 MAGs obtained from RB10, RB13, RB32 and RB108 samples.
In accordance with current standards, the bins were defined as high quality (HQ) (> 90% completion, < 5% contamination, presence of the 23S, 16S and 5S rRNA genes and at least 18 tRNAs), near high quality (NHQ) (> 90% completion, < 5% contamination, other criteria partially covered), medium quality (MQ) (≥ 50% completion, < 10% contamination), and low quality (LQ) (< 50% completion, < 10% contamination) MAGs. Metagenomes and MAGs accession numbers are available on ENA (Study ID: PRJEB46766).
Table 2.
Classification of the MAGs based on the taxonomic classification of GTDB-Tk software (v2.1.0) and the Genome Taxonomy Database (07-RS207 release).
Taxonomic and phylogenomic analyses of MAGs
The taxonomic affiliation of the MAGs was investigated in detail through the workflow classify of GTDB-Tk (v 2.1.0; GTDB reference tree 07-RS207) (Table 2) and through de novo phylogenomic analyses (Fig. S1a–i). We also tried to classify MAGs on the basis of overall genome relatedness indices (OGRI), which is detailed in supplementary material (Text S1, Table S2, Fig. S2).
De novo phylogenomic analyses globally confirmed the positioning of MAGs provided by GTDB-Tk, with high branching support. For Bacteria, GTDB-Tk analyses allowed us to place the MAGs in the following clades: six in the phylum Aquificota from the four different springs, comprising four MAGs belonging to the genus Hydrogenivirga (family Aquificaceae) (RB10-MAG07, RB13-MAG10, RB32-MAG07, RB108-MAG02), and two belonging to the family ‘Hydrogenobaculaceae’ (RB10-MAG12, RB32-MAG11) (Table 2, Fig. S1a). Their closest cultured relatives originated either from hot springs or from deep-sea hydrothermal vents27. Three MAGs from three geothermal springs belonged to the phylum Armatimonadota (RB10-MAG03, RB13-MAG04, RB32-MAG03) and had no close cultured relatives. Seven MAGs have been classified into the phylum Chloroflexota: three MAGs belonging to the genus Thermoflexus from three different springs (RB10-MAG04, RB13-MAG05, RB32-MAG02), one affiliating with the genus Thermomicrobium (RB32-MAG08), one falling into the family Ktedonobacteraceae (RB108-MAG03), one belonging to the class Dehalococcoidia (RB32-MAG04) and another one whose phylogenetic position is more difficult to assert because it is a MAG of medium quality (RB32-MAG14). Six MAGs from four various hot springs belonged to the phylum Deinococcota, and to the genera Thermus (RB10-MAG08, RB10-MAG11, RB13-MAG09, RB32-MAG10, RB108-MAG01) and Meiothermus (RB13-MAG13). One MAG belonged to the family ‘Sulfurifustaceae’ (RB13-MAG01), in the phylum Proteobacteria (Gamma-class). The MAG referenced as RB32-MAG13 was classified into the phylum ‘Patescibacteria’, in the class ‘Paceibacteria’, and was distantly related to MAGs originating from groundwater and from hot springs. Finally, two MAGs from two different springs belonged to the phylum WOR-3, in the Candidatus genus ‘Caldipriscus’ (RB32-MAG12, RB10-MAG09).
For Archaea, almost all the MAGs reconstructed in this study, e.g. 15 of the 16 archaeal MAGs, belonged to the phylum Thermoproteota. Among them, four belonged to the genus Ignisphaera (RB10-MAG05, RB13-MAG08, RB13-MAG11, RB32-MAG05), three to the genus Infirmifilum (RB10-MAG06, RB13-MAG03, RB32-MAG09), two to the genus Zestosphaera (RB10-MAG02, RB13-MAG06), three to the family Acidilobaceae (RB10-MAG01, RB13-MAG02, RB32-MAG01) and two to the order Geoarchaeales (RB10-MAG10, RB32-MAG06). Additionally, one belonged to the family Thermocladiaceae (RB13-MAG07). Lastly, the MAG belonging to another phylum (RB13-MAG12) was affiliated with the ‘Aenigmatarchaeota’, class ‘Aenigmatarchaeia’, and was distantly related to MAGs from hot springs and from deep-sea hydrothermal vent sediments28,29.
Out of these 42 MAGs, at least 19 MAGs corresponded to different taxa at the taxonomic rank of species or higher according to GTDB (Table 2). Eighteen of them belonged to lineages with several cultivated representatives including the species Thermus thermophilus. 13 new genomic species within the GTDB genera Hydrogenivirga, HRBIN17, Thermoflexus, SpSt-223, CADDYT01, Zestosphaera, Ignisphaera, Infirmifilum, Thermus, Thermus_A, Meiothermus_B, JAHLMO01 and Caldipriscus, and 6 putative new genomic genera belonging to the GTDB families Hydrogenobaculaceae, Acidilobaceae, WAQG01, Thermocladiaceae, Sulfurifustaceae and HR35 could be identified (Table 2). In addition, 9 MAGs belonged to lineages that are predominantly or exclusively known through environmental DNA sequences. Thus, these 42 MAGs comprised a broad phylogenetic range of Bacteria and Archaea at different levels of taxonomic organization, of which a large majority were not reported before.
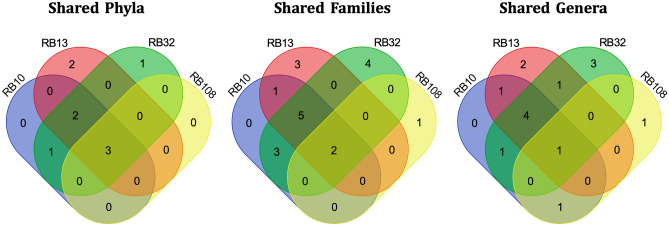
The approaches implemented here were not intended to describe the microbial diversity present in these sources in an exhaustive way or to compare them in a fine way, and cannot allow it because of a 2-year storage at 4 °C. This long storage has probably led to changes in the microbial communities and to the selective loss or enrichment of some taxa. As a result, no analysis of abundance or absence of taxa can be conducted from these metagenomes and the results are discussed taking this bias into account. However, they do provide an overview of the microbial diversity effectively present. If we compare the phylogenetic diversity of the MAGs found in the four hot springs, we can observe that 3 shared phyla (Deinococcota, Aquificota and Chloroflexota: phyla names according to GTDB), 2 shared families (Thermaceae and Aquificaceae), and one shared genus (Hydrogenivirga) were found among the four sources (Fig. 2). In addition, hot springs RB10, RB13 and RB32, that are geographically close (< 60 m), also share 2 other phyla (Thermoproteota and Armatimonadota) and 5 other families in common (Acidilobaceae, Ignisphaeraceae, Thermofilaceae, Thermoflexaceae, and HRBIN17) (Fig. 2). These phyla and families that are shared between sources are widespread lineages in terrestrial geothermal habitats (e.g.4–6,12). Phyla and families detected in the hot environments of Antarctica are also found here, such as Patescibacteria15. In summary, this metagenomic analysis highlighted the presence of bacterial and archaeal lineages commonly found in hot springs, and lineages found in hot habitats from polar areas (e.g.4–6,15,30). The microbial communities in these Kerguelen Islands hot springs samples were diverse, particularly in RB10, RB13, and RB32 hot springs. Within these lineages previously reported to occur in geothermal environments, a majority of the genomic taxa detected here were novel. Those results were obtained considering their taxonomic affiliation by GTDB-Tk, and their phylogenomic position with respect to closest relatives and the OGRI thresholds (16S rRNA gene sequence similarity, average nucleotide identity, and average amino acid identity) classically used to delineate different taxonomic ranks in cultured strains, used here as indicators of taxonomic differentiation (Table 2, Table S2).
Figure 2.
Venn diagrams showing the shared phyla, families and genera according to GTDB classification in the reconstructed MAGs from the hot springs RB10, RB13, RB32 and RB108.
Functional potential of MAGs: putative metabolisms and adaptations
A genomic characterization of the 42 MAGs has been performed to explore the possible metabolic pathways and adaptations of the microbial populations from which these MAGs originate. KEGG Decoder visualization highlighted various pathways associated with carbohydrate degradation, oxidative phosphorylation and sulfur, nitrogen, and amino-acid metabolisms, among others (Fig. 3). To confirm these initial metabolic predictions, a further annotation was performed by combining data generated by Prokka with the MetaCyc database. Efforts have been directed at studying catabolic pathways, particularly those involving inorganic electron donors and acceptors. These results are not representative of the metabolic diversity of all the hot spring ecosystems studied, but they do reflect some of the microbial catabolism likely to be used in situ to produce energy and, by assumption, the most abundant ones. Metabolic predictions are presented hereafter, at different taxonomic ranks and have been compared to the known phenotypes of the closest cultivated taxa, and in some cases to genomic content of the closest relatives.
Figure 3.
Metabolic pathway diagram of the 42 MAGs based on KEGG Decoder annotations, showing MAG classification according to GTDB-Tk and estimated genome completion.
MAGs belonging to the genus Thermoflexus (RB10-MAG04, RB13-MAG05, RB32-MAG02) encode pathways for carbon monoxide oxidation (via aerobic carbon monoxide dehydrogenase), hydrogen oxidation and nitrate respiration; the only cultivated known representative of this genus is a heterotrophic bacterium31. The same pathways, except for the nitrate reduction pathway, are encoded in the complete genome of Thermoflexus hugenholtzii (NCBI: ASM1877156v1). In contrast, the genome of T. hugenholtzii, a strain isolated from a terrestrial hot spring in Nevada31, encodes the tetrathionate reduction and thiosulfate disproportionation pathways, which are not encoded in the three Kerguelen Island MAGs. The Dehalococcoidia’s MAG (RB32-MAG04) encodes only a carbon monoxide oxidation pathway; the cultivated members of this genus are strict anaerobic hydrogenotrophic, organohalide-respiring bacteria32. In the MAG associated with the genus Thermomicrobium (RB32-MAG08), we predicted pathways for dimethylsulfide degradation, thiosulfate disproportionation and carbon monoxide oxidation; only carboxydotrophic growth has been reported in this genus and demonstrated by culture33. The same pathways are encoded in the complete genome of Thermomicrobium roseum, a strain isolated from a hot spring in Yellowstone National Park (NCBI: ASM2168v1)34. In the Chloroflexota’s MAG (RB32-MAG14) (belonging to the order Chloroflexales, Table S2), carbon monoxide oxidation and thiosulfate disproportionation pathways are present but no coding DNA sequence associated with phototrophy could be find, which may suggest a chemoorganotrophy mode of energy production35. The Ktedonobacteraceae’s MAG (RB108-MAG03) encodes enzymes for hydrogen oxidation (aerobic) pathways, carbon monoxide oxidation, dimethylsulfide degradation, selenate reduction, thiosulfate oxidation and disproportionation and finally tetrathionate oxidation; yet, the few taxa of this family isolated so far are mesophilic heterotrophic bacteria36. Within Hydrogenobaculaceae MAGs (RB10-MAG12, RB32-MAG11), we predicted a thiosulfate disproportionation pathway; most of the species within this family are capable of chemolithotrophic microaerophilic or anaerobic growth37. MAGs belonging to the genus Hydrogenivirga (RB10-MAG07, RB13-MAG10, RB32-MAG07, RB108-MAG02) possess genes encoding enzymes of aerobic respiration, thiosulfate oxidation, thiosulfate disproportionation, tetrathionate reduction, and hydrogen oxidation (aerobic and anaerobic); which is consistent with what is known about the genus (nitrate and oxygen respiration combined to hydrogen, sulfur, or thiosulfate oxidation)37. The same pathways, with the exception of the hydrogen oxidation pathway, are encoded in the genome of Hydrogenivirga caldilitoris (NCBI: ASM366400v1), a close relative isolated from a coastal hot spring in Japan27. In MAGs associated with the genus Thermus (RB10-MAG08, RB10-MAG11, RB13-MAG09, RB32-MAG10, RB108-MAG01), we predicted pathways for aerobic respiration, assimilatory sulfate reduction, hydrogen oxidation, selenate reduction, thiosulfate oxidation and thiosulfate disproportionation; cultivated species of this genus grow mainly chemoorganoheterotrophically by aerobic respiration, but some have genes coding for chemolithotrophic and anaerobic respiration enzymes38. The MAG belonging to the genus Meiothermus (RB13-MAG13) encodes pathways for carbon monoxide oxidation, hydrogen oxidation, thiosulfate oxidation and thiosulfate disproportionation; Meiothermus strains are known to grow chemoorganotrophically by oxygen or nitrate respiration39. For the RB13-MAG01 belonging to the Sulfurifustaceae, we predicted the genetic potential for aerobic respiration, ammonia oxidation, dissimilatory sulfate reduction, sulfite oxidation, sulfide oxidation (to sulfur globules), tetrathionate reduction, thiosulfate oxidation and thiosulfate disproportionation; Sulfurifustaceae (referenced as Acidiferrobacteraceae in the LPSN taxonomy) are known to be able to oxidize sulfur and iron, and the microorganism corresponding to this MAG may possess a larger panel of chemolithotrophic abilities40. For members of the Armatimonadota (RB10-MAG03, RB13-MAG04, RB32-MAG03), we predicted pathways for assimilatory sulfate reduction, carbon monoxide oxidation, selenate reduction and thiosulfate disproportionation; the members of the phylum have a phenotype of aerobic heterotrophs41. In Zestosphaera’s (RB10-MAG02, RB13-MAG06) and Ignisphaera’s (RB10-MAG05, RB13-MAG08, RB13-MAG11, RB32-MAG05) MAGs, we predicted sulfur and polysulfide reduction pathways; those MAGs could be classified as Desulfurococcaceae (LPSN taxonomy, Table S2) which are known as heterotrophs respiring sulfur species42,43. MAGs belonging to the class Thermoproteia (RB10-MAG10, RB13-MAG07, RB32-MAG06) encode dissimilatory sulfate reduction pathway; various catabolic pathways are described in this class44. In MAGs related to the genus Caldipriscus (RB10-MAG09, RB32-MAG12), phylum Patescibacteria (RB32-MAG13), family Acidilobaceae (RB10-MAG01, RB13-MAG02, RB32-MAG01), family Thermofilaceae (RB10-MAG06, RB13-MAG03, RB32-MAG09) and class Aenigmatarchaeia (RB13-MAG12), we did not predict any catabolic pathway of inorganic nutrients among those reported in the MetaCyc database. This could be explained by the low completion of the MAGs and/or the fact that only well-known pathways are documented in this database. However, all these MAGs have pathways associated with carbohydrate and protein degradation. This may indicate that these taxa are chemoheterotrophs, which has already been reported in geothermal environments and already described for relatives of some of these taxa45,46.
Sulfide oxidation may be a possible energy production pathway for 28 MAGs based on KEGG Decoder (Fig. 3), since they code for a sulfide:quinone oxidoreductase (KEGG:K17218) and a flavoprotein chain of sulfide dehydrogenase (KEGG:K17229), but this hypothesis was not confirmed by MetaCyc except for RB13-MAG01. Due to high representations of sulfur metabolisms, genes encoded in MAGs were evaluated with DiSCo, which gave similar results to those obtained when analyzed with Pathway tools. DiSCo confirmed complete dissimilatory sulfate reduction pathways for two MAGs, predicted to be associated to sulfate reduction processes (RB13-MAG07) or sulfide oxidation processes by reverse sulfate reduction pathway (RB13-MAG01). The assimilatory sulfate reduction pathway is more represented in the overall dataset formed by all MAGs than the dissimilatory pathway, which is consistent with the low sulfate concentration measured in the four hot springs (Table S1). The thiosulfate disproportionation pathway predicted by MetaCyc in many MAGs simply refers to the detection of an enzyme, the rhodanese-type thiosulfate sulfurtransferase. However, in the current state of knowledge on the disproportionation pathways of inorganic sulfur compounds47,48, this enzyme alone does not allow the implementation of this catabolic pathway. If we consider all the genes present in these MAGs, nothing indicates that the microorganisms from which these MAGs originate can achieve the disproportionation of inorganic sulfur compounds.
Additionally, no enzymes clearly associated with photosystems I and II were found. Nevertheless, it cannot be ruled out that these energy production pathways are absent in microorganisms indigenous to these sources, due to sample storage bias and low completion of some MAGs. On the other hand, our results show that these sources host chemolithoautotrophic taxa involved in the carbon and sulfur cycle, and to a lesser extent in the hydrogen and nitrogen cycles. Several taxa are likely to be involved in the primary production of these sources through chemolithoautotrophy, but in addition, heterotrophs appear to be present and diverse in the collected samples. Additional studies will be required to better understand the metabolic diversity and trophic webs of these hot springs, in order to better understand the ecology of the microbial communities of the Kerguelen hot springs.
Regarding thermophily, we found that all MAGs encode heat shock proteins, mainly associated with the HSP20 family, with the exception of RB10-MAG12 and RB13-MAG12. The absence of Hsp encoding genes in these two MAGs is possibly due to the low genome completeness of these two MAGs. Under conditions of heat stress, it has been shown that the small heat shock proteins Hsp20, protect cellular proteins from aggregation and membrane lipids from destabilization, in some thermophilic archaea49. In taxa of these geothermal sources, these proteins could help the cells to counteract the deleterious effects of environmental stress and in particular of thermal stress. In addition, reverse gyrase coding sequences were found in 29 out of the 42 MAGs; these enzymes are known to be exclusive to hyperthermophiles and involved in DNA protection and repair at high temperatures50. Only MAGs RB10-MAG04, RB10-MAG09, RB10-MAG11, RB13-MAG01, RB13-MAG05, RB13-MAG09, RB13-MAG13, RB32-MAG02, RB32-MAG04, RB32-MAG08, RB32-MAG10, RB32-MAG12 and RB108-MAG03, belonging to the phylum Chloroflexota, the family Sulfurifustaceae or the genus Caldipriscus (GTDB taxonomy), do not encode any reverse gyrase gene. These results suggest the presence of numerous thermophilic and hyperthermophilic prokaryotes in these high temperature hot springs. Further cultural and physiological investigations from samples of these Kerguelen hot springs will be necessary to confirm these statements.
In conclusion, this first metagenomic overview of the microbial diversity of Kerguelen hot springs allowed the assembly of 42 MAGs, from four hot springs. Several MAGs correspond to putative new taxa, namely 13 new putative genomic species and 6 new putative genera affiliated to Bacteria and Archaea according to GTDB. Based on their genetic potential, these taxa appear to be chemolithoautotrophs and chemoheterotrophs and thus probably involved in the carbon, sulfur, hydrogen and nitrogen cycle. Many of these MAGs are likely to be derived from populations of thermophilic/hyperthermophilic bacteria and archaea. As geographically isolated sites, the Kerguelen Islands are reservoirs of diversity and taxa of novel microorganisms that should be interesting to study the evolution of microbial life and speciation processes. It has been difficult to fully assess the microbial metabolic diversity in these geothermal pools due to the inherent limitations of MAG reconstruction and the state of knowledge of microbial pathways that remains limited. However, these geothermal ecosystems could be reservoirs of biological and genomic novelty. The physiological properties and adaptive mechanisms of microorganisms inhabiting these unique environments will deserve to be examined in detail in the future by implementing large-scale metagenomics, metatranscriptomics and cultural analyses.
Methods
Sample collection and major elements analysis in water samples
Water samples were collected from four hot springs during the 2016–2017 austral summer TALISKER field campaign (1st of December–11th of February) organized by the French Polar Institute Paul Emile Victor (https://institut-polaire.fr/en/). Water samples and water samples mixed with surficial sediments aliquots were collected. Water samples were collected in 250ML LDPE Nalgene bottles stored at 4 °C until ionic chromatography analysis. Mixed water and sediment samples were collected aseptically in sterile 50 mL Becton- Dickinson and Company-syringes, then stored anaerobically in sterile glass bottles at 4 °C. Field measurements of fluid parameters were performed using a HI9829 (Hanna instruments) multiparameter calibrated and equipped with sensors allowing the acquisition of pH, temperature (°C), alkalinity (mg/L), and electrical conductivity measurements (mS/cm) (Table S1). The major anions and cations were analyzed at LGL-TPE using ion-chromatography (Methrom ECO IC). A mixture of 3.2 mM Na2CO3 and 1 mM NaHCO3 was used as an eluent for analysis of anions and a chemical suppression module (MSM) was used to suppress the conductivity. For cations, 1.7 mM HNO3 was used as an eluent. The anions and cations were separated using analytical columns, Metrosep A Supp5 Guard/4.0 and Metrosep C4 250/4.0, respectively (Table S1).
DNA extraction and sequencing
Hot spring’s samples analyzed here were originally collected to grow thermophilic taxa. They were stored at 4 °C for 2 years before DNA was extracted. For each hot spring sample, three replicates of DNA extraction were conducted individually, and combined as a composition sample, before the sequencing. DNA was extracted with a standard PCI (Phenol:Chloroform:Isoamyl Alcohol (25:24:1)) protocol, as described elsewhere, from 10 g environmental matrix51, with the exception that 50 µM linear acrylamide were added to enhance nucleic acids precipitation (Invitrogen). One negative control was included and contained 10 mL of DNA-free sterile water. Elution of total DNA extracts was performed in 30–50 μL EB buffer (10 mM Tris–Cl, pH 8.5). Nucleic acid solution quality was determined using the NanoDrop 8000 (Thermo Scientific, Waltham, MA, USA) spectrophotometer. Double-strand DNA concentration was measured using the kit Quantifluor dsDNA system. DNA samples were sequenced by NovaSeq 6000 (2 × 150 bp, paired-end reads) technology by the Duke Center for Genomic and Computational Biology (GCB) (https://genome.duke.edu/).
Sequence processing, metagenomic assembly and binning
Metagenome sequences’ quality were controlled by FastQC (v0.11.9—https://github.com/s-andrews/FastQC) and MultiQC (v1.9—https://github.com/ewels/MultiQC). Sequences were then processed with the snakemake of Anvi’o52–54 (v7—https://github.com/merenlab/anvio), filtered by integrated minoche script (v2.8—https://github.com/merenlab/illumina-utils). Next, MetaSpades55 (v3.14.1—https://github.com/ablab/spades) was used as genome assembler and Concoct56 (v1.1.0—https://github.com/BinPro/CONCOCT) as genome binner with anvi_cluster_contigs function (“all against all” mode). Furthermore, MAGs were manually refined with anvi-refine function. Genome mapping was performed with bowtie257 (v2.4.2—https://sourceforge.net/projects/bowtie-bio/files/bowtie2/2.4.2/) and samtools58 (v1.7—https://samtools.github.io/). MAGs’ quality was estimated by Anvi’o and furthermore by CheckM59 (v1.1.3—https://ecogenomics.github.io/CheckM/), both with default parameters. Total length, number of contigs, N50, and GC% contents were extracted with anvi-summarize function.
Taxonomic and phylogenetic inference of metagenomic assemblies and MAGs
According to the standards proposed elsewhere26, bins were defined as high-quality (HQ) MAGs (> 90% completion, < 5% contamination, presence of the 23S, 16S and 5S rRNA genes and at least 18 tRNAs), nearly high-quality (NHQ) MAGs (> 90% completion, < 5% contamination, other criteria partially covered), medium-quality (MQ) MAGs (≥ 50% completion, < 10% contamination) and low-quality (LQ) MAGs (< 50% completion, < 10% contamination). MAGs often lack 16S rRNA genes due to their conserved and repetitive nature preventing their assembly60, so MAGs of near high-quality could be classified as high-quality MAGs by other authors taking this into consideration. The taxonomic affiliation of the MAGs was first investigated by placing the MAGs in a phylogenomic context. The phylogenetic reconstructions were based on 122 archaeal or 120 bacterial single copy conservative marker genes according to the Genome Taxonomy Database (07-RS207 release) and were constructed using de novo workflow implemented in GTDB-Tk (v1.4.1—https://github.com/Ecogenomics/GTDBTkk)61,62. Visualization and trees analyses were done using ARB software63.
As the taxonomy proposed by GTDB is new and does not correspond exactly to the one recognized by the International Code of Nomenclature of prokaryotes (ICNP), we also analyzed data according to the rules of the Code and its nomenclature, because at the time of writing this article, SeqCode is being implemented64. For this purpose, we implemented a combination of genomic indices classically used for the delineation of the different taxonomic ranks, namely: 16S rRNA gene sequence similarity, average nucleotide identity score (ANI) and average amino-acid identity value (AAI). The approach followed and the results are given in supplementary material (Text S1, Table S2, Fig. S2).
Metabolic profiling
MAGs were processed with KEGG Decoder script (https://github.com/bjtully/BioData/tree/master/KEGGDecoder) from Anvi’o gene calls tables generated with kegg_kofams and then plotted with R packages (ComplexHeatmap, circlize, RColorBrewer, and dplyrt) to get a general annotation with R (v.3.6.3)65,66. A more accurate annotation was performed with Prokka (v1.14.6—https://github.com/tseemann/prokka)67 and associated outputs were analyzed by using the Pathway Tools software (v.24.5)68 with the MetaCyc database (v.24.5)69 to explore in details the putative metabolisms encoded in MAGs or public available genome assemblies. Regarding sulfur metabolisms, for dsr genes, the perl script DiSCo (v.1.0.0, https://github.com/Genome-Evolution-and-Ecology-Group-GEEG/DiSCo) was used on the Prokka protein sequences outputs of each MAG to highlight the specific genes70.
Supplementary Information
Acknowledgements
We would like to thank Guillaume Lannuzel for providing assistance with molecular biology analyses. We are grateful to the Territoire des Terres Australes et Antarctiques Françaises (TAAF), the IPEV (Institut Polaire Paul-Emile Victor) and its logistical staff for logistic support for transit and fieldwork in the Kerguelen archipelago. We thank M. Jebbar and Z. Shao, co-supervisors of M.A., for letting him do this study which was not in the scope of his thesis. We acknowledge the LABGeM (CEA/Genoscope & CNRS UMR8030), the France Génomique and French Bioinformatics Institute national infrastructures (funded as part of Investissements d’Avenir program managed by Agence Nationale pour la Recherche, contracts ANR-10-INBS-09 and ANR-11-INBS-0013) are acknowledged for support within the MicroScope annotation platform. We are grateful for the financial support of IPEV provided by the TALISKER program N°1077 to D.G., the HOTVIR program N°408 to M.L.R and the SEKMET program N°1231 to K.A. and M.L.R. Metagenomic analyses were funded by the Sino-French IRP 1211 MicrobSea to K.A., by the MERLIN “Abyss” program funded by Ifremer to K.A. and by the Scientific Council of IUEM to K.A. The study was supported by two grants from the French Ministry of Higher Education and Research, and from the Région Bretagne, to M.A and S.Y.
Author contributions
Conceptualization, M.A. and K.A.; formal analysis, M.A., S.Y., A.M., V.L., D.G., J.A., J.P. and K.A.; funding acquisition, D.G., M.L.R. and K.A.; investigation, M.A., S.Y., A.M., M.C., D.G., J.A., and K.A; supervision, K.A.; writing—original draft, M.A., S.Y. and K.A.; review and editing, all coauthors. All authors have read and agreed to the published version of the manuscript.
Data availability
The metagenome bins generated and analyzed during the current study are available in the European Nucleotide Archive (ENA) (https://www.ebi.ac.uk/ena/browser/home), under the Project PRJEB46766 (Table 1).
Competing interests
The authors declare no competing interests.
Footnotes
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
The online version contains supplementary material available at 10.1038/s41598-022-26299-4.
References
- 1.Mehta D, Satyanarayana T. Thermophilic Microbes in Environmental and Industrial Biotechnology. Springer; 2013. Diversity of Hot Environments and Thermophilic Microbes; pp. 3–60. [Google Scholar]
- 2.Des Marais DJ, Walter MR. Terrestrial hot spring systems: Introduction. Astrobiology. 2019;19(12):1419–1432. doi: 10.1089/ast.2018.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Belilla J, Moreira D, Jardillier L, Reboul G, Benzerara K, López-García JM, Bertolino P, López-Archilla AI, López-García P. Hyperdiverse archaea near life limits at the polyextreme geothermal Dallol area. Nat. Ecol. Evol. 2019;3(11):1552–1561. doi: 10.1038/s41559-019-1005-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Urbieta M, Porati G, Segretín A, González-Toril E, Giaveno M, Donati E. Copahue geothermal system: A volcanic environment with rich extreme prokaryotic biodiversity. Microorganisms. 2015;3(3):344–363. doi: 10.3390/microorganisms3030344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Power JF, Carere CR, Lee CK, Wakerley GLJ, Evans DW, Button M, White D, Climo MD, Hinze AM, Morgan XC, McDonald IR, Cary SC, Stott MB. Microbial biogeography of 925 geothermal springs in New Zealand. Nat. Commun. 2018;9:1. doi: 10.1038/s41467-018-05020-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Lezcano MÁ, Moreno-Paz M, Carrizo D, Prieto-Ballesteros O, Fernández-Martínez MÁ, Sánchez-García L, Blanco Y, Puente-Sánchez F, de Diego-Castilla G, García-Villadangos M, Fairén AG, Parro V. Biomarker profiling of microbial mats in the geothermal band of Cerro Caliente, Deception Island (Antarctica): Life at the edge of heat and cold. Astrobiology. 2019;19(12):1490–1504. doi: 10.1089/ast.2018.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Brock TD, Brock KM, Belly RT, Weiss RL. Sulfolobus: A new genus of sulfur-oxidizing bacteria living at low pH and high temperature. Arch. Mikrobiol. 1972;84(1):54–68. doi: 10.1007/bf00408082. [DOI] [PubMed] [Google Scholar]
- 8.Meyer-Dombard DR, Shock EL, Amend JP. Archaeal and bacterial communities in geochemically diverse hot springs of Yellowstone National Park, USA. Geobiology. 2005;3(3):211–227. doi: 10.1111/j.1472-4669.2005.00052.x. [DOI] [Google Scholar]
- 9.Kublanov IV, Perevalova AA, Slobodkina GB, Lebedinsky AV, Bidzhieva SK, Kolganova TV, Kaliberda EN, Rumsh LD, Haertlé T, Bonch-Osmolovskaya EA. Biodiversity of thermophilic prokaryotes with hydrolytic activities in hot springs of Uzon Caldera, Kamchatka (Russia) Appl. Environ. Microbiol. 2009;75(1):286–291. doi: 10.1128/aem.00607-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Reigstad LJ, Jorgensen SL, Schleper C. Diversity and abundance of Korarchaeota in terrestrial hot springs of Iceland and Kamchatka. ISME J. 2010;4(3):346–356. doi: 10.1038/ismej.2009.126. [DOI] [PubMed] [Google Scholar]
- 11.Wemheuer B, Taube R, Akyol P, Wemheuer F, Daniel R. Microbial diversity and biochemical potential encoded by thermal spring metagenomes derived from the Kamchatka Peninsula. Archaea. 2013;2013:1–13. doi: 10.1155/2013/136714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Wilkins LGE, Ettinger CL, Jospin G, Eisen JA. Metagenome-assembled genomes provide new insight into the microbial diversity of two thermal pools in Kamchatka, Russia. Sci. Rep. 2019;9:1. doi: 10.1038/s41598-019-39576-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Deamer D, Damer B, Kompanichenko V. Hydrothermal chemistry and the origin of cellular life. Astrobiology. 2019;19(12):1523–1537. doi: 10.1089/ast.2018.1979. [DOI] [PubMed] [Google Scholar]
- 14.Van Kranendonk MJ, Deamer DW, Djokic T. Life springs. Sci. Am. 2017;317(2):28–35. doi: 10.1038/scientificamerican0817-28. [DOI] [PubMed] [Google Scholar]
- 15.Muñoz PA, Flores PA, Boehmwald FA, Blamey JM. Thermophilic bacteria present in a sample from Fumarole Bay, Deception Island. Antarct. Sci. 2011;23(6):549–555. doi: 10.1017/s0954102011000393. [DOI] [Google Scholar]
- 16.Amenábar MJ, Flores PA, Pugin B, Boehmwald FA, Blamey JM. Archaeal diversity from hydrothermal systems of Deception Island, Antarctica. Polar Biol. 2013;36(3):373–380. doi: 10.1007/s00300-012-1267-3. [DOI] [Google Scholar]
- 17.Bendia AG, Signori CN, Franco DC, Duarte RTD, Bohannan BJM, Pellizari VH. A mosaic of geothermal and marine features shapes microbial community structure on deception Island Volcano, Antarctica. Front. Microbiol. 2018;9:20. doi: 10.3389/fmicb.2018.00899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.ATCM (Antarctic Treaty Consultative Meeting) (2012). Measure 8. Management plan for Antarctic Specially Protected Area No. 140. Parts of Deception Island, South Shetland Islands. Final report of the thirty-fifth ATCM, Hobart, Australia, June 11–20. www.ats.aq.
- 19.Giret A, Gregoire M, Cottin JY, Michon G. In: Kerguelen, a Third Type Of Oceanic Island? “The Antarctic Region: Geological Evolution and Processes”. Ricci CA, editor. Terra Antarctica Publication; 1997. pp. 735–741. [Google Scholar]
- 20.Gagnevin D, Ethien R, Bonin B, Moine B, Féraud G, Gerbe MC, Cottin JY, Michon G, Tourpin S, Mamias G, Perrache C, Giret A. Open-system processes in the genesis of silica-oversaturated alkaline rocks of the Rallier-du-Baty Peninsula, Kerguelen Archipelago (Indian Ocean) J. Volcanol. Geoth. Res. 2003;123(3–4):267–300. doi: 10.1016/s0377-0273(02)00509-7. [DOI] [Google Scholar]
- 21.Parikka KJ, Jacquet S, Colombet J, Guillaume D, Le Romancer M. Abundance and observations of thermophilic microbial and viral communities in submarine and terrestrial hot fluid systems of the French Southern and Antarctic Lands. Polar Biol. 2018;41(7):1335–1352. doi: 10.1007/s00300-018-2288-3. [DOI] [Google Scholar]
- 22.Renac C, Moine B, Goudour JP, Le Romancer M, Perrache C. Stable isotope study of rainfall, river drainage and hot springs of the Kerguelen archipelago, SW Indian Ocean. Geothermics. 2020;83:101726. doi: 10.1016/j.geothermics.2019.101726. [DOI] [Google Scholar]
- 23.Postec, A., Ciobanu, M., Birrien, J.L., Prieur, D. & Le Romancer, M. (2009). Microbial biodiversity of thermophilic communities in remote geothermal springs of Saint-Paul and Kerguelen Islands. In Proceedings of “Thermophiles 2009” Conference, Beijing, China, August,16–21.
- 24.Gramain, A., Brillet, F., Birrien, J. L., & Le Romancer, M. (2011). Novel hyperthermophilic archaea flying over the Kerguelen islands: A dissemination study. In “Thermophiles 2011” Conference, Big Sky, Montana USA, September 11–16.
- 25.Cozannet M, Borrel G, Roussel E, Moalic Y, Allioux M, Sanvoisin A, Toffin L, Alain K. New insights into the ecology and physiology of Methanomassiliicoccales from terrestrial and aquatic environments. Microorganisms. 2021;9(1):30. doi: 10.3390/microorganisms9010030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Bowers RM, Kyrpides NC, Stepanauskas R, Harmon-Smith M, Doud D, Reddy TBK, Schulz F, Jarett J, Rivers AR, Eloe-Fadrosh EA, Tringe SG, Ivanova NN, Copeland A, Clum A, Becraft ED, Malmstrom RR, Birren B, Podar M, Woyke T. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea. Nat. Biotechnol. 2017;35(8):725–731. doi: 10.1038/nbt.3893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Nakagawa S, Nakamura S, Inagaki F, Takai K, Shirai N, Sako Y. Hydrogenivirga caldilitoris gen. nov., sp. Nov., a novel extremely thermophilic, hydrogen- and sulfur-oxidizing bacterium from a coastal hydrothermal field. Int. J. Syst. Evol. Microbiol. 2004;54(6):2079–2084. doi: 10.1099/ijs.0.03031-0. [DOI] [PubMed] [Google Scholar]
- 28.Seitz KW, Dombrowski N, Eme L, Spang A, Lombard J, Sieber JR, Teske AP, Ettema TJG, Baker BJ. Asgard archaea capable of anaerobic hydrocarbon cycling. Nat. Commun. 2019;10:1. doi: 10.1038/s41467-019-09364-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Zhou Z, Liu Y, Xu W, Pan J, Luo Z-H, Li M. Genome- and community-level Interaction insights into carbon utilization and element cycling functions of Hydrothermarchaeota in hydrothermal sediment. Systems. 2020;5:1. doi: 10.1128/msystems.00795-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Kochetkova TV, Toshchakov SV, Zayulina KS, Elcheninov AG, Zavarzina DG, Lavrushin VYu, Bonch-Osmolovskaya EA, Kublanov IV. Hot in cold: Microbial life in the hottest springs in permafrost. Microorganisms. 2020;8(9):1308. doi: 10.3390/microorganisms8091308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Dodsworth JA, Gevorkian J, Despujos F, Cole JK, Murugapiran SK, Ming H, Li W-J, Zhang G, Dohnalkova A, Hedlund BP. Thermoflexus hugenholtzii gen. nov., sp. nov., a thermophilic, microaerophilic, filamentous bacterium representing a novel class in the Chloroflexi, Thermoflexia classis nov., and description of Thermoflexaceae fam. nov. and Thermoflexales ord. nov. Int. J. Syst. Evol. Microbiol. 2014;64(Pt_6):2119–2127. doi: 10.1099/ijs.0.055855-0. [DOI] [PubMed] [Google Scholar]
- 32.Löffler FE, Yan J, Ritalahti KM, Adrian L, Edwards EA, Konstantinidis KT, Müller JA, Fullerton H, Zinder SH, Spormann AM. Dehalococcoides mccartyi gen. nov., sp. nov., obligately organohalide-respiring anaerobic bacteria relevant to halogen cycling and bioremediation, belong to a novel bacterial class, Dehalococcoidia classis nov., order Dehalococcoidales ord. nov. and family Dehalococcoidaceae fam. nov., within the phylum Chloroflexi. Int. J. Syst. Evol. Microbiol. 2013;63(2):625–635. doi: 10.1099/ijs.0.034926-0. [DOI] [PubMed] [Google Scholar]
- 33.King CE, King GM. Thermomicrobium carboxidum sp. nov., and Thermorudis peleae gen. nov., sp. nov., carbon monoxide-oxidizing bacteria isolated from geothermally heated biofilms. Int. J. Syst. Evol. Microbiol. 2014;64(8):2586–2592. doi: 10.1099/ijs.0.060327-0. [DOI] [PubMed] [Google Scholar]
- 34.Wu D, Raymond J, Wu M, Chatterji S, Ren Q, Graham JE, Bryant DA, Robb F, Colman A, Tallon LJ, Badger JH, Madupu R, Ward NL, Eisen JA. Complete genome sequence of the aerobic CO-oxidizing thermophile Thermomicrobium roseum. PLoS One. 2009;4(1):e4207. doi: 10.1371/journal.pone.0004207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Hanada S. The Prokaryotes. Springer; 2014. The Phylum Chloroflexi, the family Chloroflexaceae, and the related phototrophic families Oscillochloridaceae and Roseiflexaceae; pp. 515–532. [Google Scholar]
- 36.Yabe S, Sakai Y, Abe K, Yokota A, Také A, Matsumoto A, Sugiharto A, Susilowati D, Hamada M, Nara K, Made Sudiana I, Otsuka S. Dictyobacter aurantiacus gen. nov., sp. nov., a member of the family Ktedonobacteraceae, isolated from soil, and emended description of the genus Thermosporothrix. Int. J. Syst. Evol. Microbiol. 2017;67(8):2615–2621. doi: 10.1099/ijsem.0.001985. [DOI] [PubMed] [Google Scholar]
- 37.Gupta RS. The Prokaryotes. Springer; 2014. The Phylum Aquificae; pp. 417–445. [Google Scholar]
- 38.Ming H, Zhao Z, Ji W, Ding C-L, Cheng L, Niu M, Li M, Yi B, Xia T, Nie G. Thermus thermamylovorans sp. Nov., isolated from a hot spring. Int. J. Syst. Evol. Microbiol. 2020;70(3):1729–1737. doi: 10.1099/ijsem.0.003965. [DOI] [PubMed] [Google Scholar]
- 39.Albuquerque L, da Costa MS. The Prokaryotes. Springer; 2014. The Family Thermaceae; pp. 955–987. [Google Scholar]
- 40.Issotta F, Moya-Beltrán A, Mena C, Covarrubias PC, Thyssen C, Bellenberg S, Sand W, Quatrini R, Vera M. Insights into the biology of acidophilic members of the Acidiferrobacteraceae family derived from comparative genomic analyses. Res. Microbiol. 2018;169(10):608–617. doi: 10.1016/j.resmic.2018.08.001. [DOI] [PubMed] [Google Scholar]
- 41.Lee KCY, Dunfield PF, Stott MB. The Prokaryotes. Springer; 2014. The Phylum Armatimonadetes; pp. 447–458. [Google Scholar]
- 42.Niederberger TD, Götz DK, McDonald IR, Ronimus RS, Morgan HW. Ignisphaera aggregans gen. nov., sp. nov., a novel hyperthermophilic crenarchaeote isolated from hot springs in Rotorua and Tokaanu, New Zealand. Int. J. Syst. Evol. Microbiol. 2006;56(5):965–971. doi: 10.1099/ijs.0.63899-0. [DOI] [PubMed] [Google Scholar]
- 43.St John E, Liu Y, Podar M, Stott MB, Meneghin J, Chen Z, Lagutin K, Mitchell K, Reysenbach A-L. A new symbiotic nanoarchaeote (Candidatus Nanoclepta minutus) and its host (Zestosphaera tikiterensis gen. nov., sp. nov.) from a New Zealand hot spring. Syst. Appl. Microbiol. 2019;42(1):94–106. doi: 10.1016/j.syapm.2018.08.005. [DOI] [PubMed] [Google Scholar]
- 44.Itoh T. The Prokaryotes. Springer; 2014. The Family Thermoproteaceae; pp. 389–401. [Google Scholar]
- 45.Belkova NL, Tazaki K, Zakharova JR, Parfenova VV. Activity of bacteria in water of hot springs from Southern and Central Kamchatskaya geothermal provinces, Kamchatka Peninsula, Russia. Microbiol. Res. 2007;162(2):99–107. doi: 10.1016/j.micres.2006.01.006. [DOI] [PubMed] [Google Scholar]
- 46.Sahm K, John P, Nacke H, Wemheuer B, Grote R, Daniel R, Antranikian G. High abundance of heterotrophic prokaryotes in hydrothermal springs of the Azores as revealed by a network of 16S rRNA gene-based methods. Extremophiles. 2013;17(4):649–662. doi: 10.1007/s00792-013-0548-2. [DOI] [PubMed] [Google Scholar]
- 47.Slobodkin AI, Slobodkina GB. Diversity of sulfur-disproportionating microorganisms. Microbiology. 2019;88(5):509–522. doi: 10.1134/s0026261719050138. [DOI] [Google Scholar]
- 48.Allioux M, Yvenou S, Slobodkina G, Slobodkin A, Shao Z, Jebbar M, Alain K. Genomic characterization and environmental distribution of a thermophilic anaerobe Dissulfurirhabdus thermomarina SH388T involved in disproportionation of sulfur compounds in shallow sea hydrothermal vents. Microorganisms. 2020;8(8):1132. doi: 10.3390/microorganisms8081132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Roy M, Gupta S, Patranabis S, Ghosh A. The oligomeric plasticity of Hsp20 of Sulfolobus acidocaldarius protects environment-induced protein aggregation and membrane destabilization. Biochim. Biophys. Acta Biomembr. 2018;1860(12):2549–2565. doi: 10.1016/j.bbamem.2018.09.005. [DOI] [PubMed] [Google Scholar]
- 50.Lulchev P, Klostermeier D. Reverse gyrase—recent advances and current mechanistic understanding of positive DNA supercoiling. Nucleic Acids Res. 2014;42(13):8200–8213. doi: 10.1093/nar/gku589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Charbonnier F, Forterre P, Erauso G, Prieur D. Purification of plasmids from thermophilic and hyperthermophilic archaebacteria. In: Robb FT, Place AR, DasSarma S, Schreier HJ, Fleischmann EM, editors. Archaea: A Laboratory Manual. Cold Spring Harbor Laboratory Press; 1995. pp. 87–90. [Google Scholar]
- 52.Koster J, Rahmann S. Snakemake—a scalable bioinformatics workflow engine. Bioinformatics. 2012;28(19):2520–2522. doi: 10.1093/bioinformatics/bts480. [DOI] [PubMed] [Google Scholar]
- 53.Shaiber A, Willis AD, Delmont TO, Roux S, Chen L-X, Schmid AC, Yousef M, Watson AR, Lolans K, Esen ÖC, Lee STM, Downey N, Morrison HG, Dewhirst FE, Mark Welch JL, Eren AM. Functional and genetic markers of niche partitioning among enigmatic members of the human oral microbiome. Genome Biol. 2020;21:1. doi: 10.1186/s13059-020-02195-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Eren AM, Kiefl E, Shaiber A, Veseli I, Miller SE, Schechter MS, Fink I, Pan JN, Yousef M, Fogarty EC, Trigodet F, Watson AR, Esen ÖC, Moore RM, Clayssen Q, Lee MD, Kivenson V, Graham ED, Merrill BD, Willis AD. Community-led, integrated, reproducible multi-omics with anvi’o. Nat. Microbiol. 2021;6(1):3–6. doi: 10.1038/s41564-020-00834-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Nurk S, Meleshko D, Korobeynikov A, Pevzner PA. metaSPAdes: A new versatile metagenomic assembler. Genome Res. 2017;27(5):824–834. doi: 10.1101/gr.213959.116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Alneberg J, Bjarnason BS, de Bruijn I, Schirmer M, Quick J, Ijaz UZ, Lahti L, Loman NJ, Andersson AF, Quince C. Binning metagenomic contigs by coverage and composition. Nat. Methods. 2014;11(11):1144–1146. doi: 10.1038/nmeth.3103. [DOI] [PubMed] [Google Scholar]
- 57.Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat. Methods. 2012;9(4):357–359. doi: 10.1038/nmeth.1923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R. The sequence alignment/map format and SAMtools. Bioinformatics. 2009;25(16):2078–2079. doi: 10.1093/bioinformatics/btp352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW. CheckM: Assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 2015;25(7):1043–1055. doi: 10.1101/gr.186072.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Parks DH, Rinke C, Chuvochina M, Chaumeil P-A, Woodcroft BJ, Evans PN, Hugenholtz P, Tyson GW. Recovery of nearly 8,000 metagenome-assembled genomes substantially expands the tree of life. Nat. Microbiol. 2017;2:1533–1542. doi: 10.1038/s41564-017-0012-7. [DOI] [PubMed] [Google Scholar]
- 61.Chaumeil P-A, Mussig AJ, Hugenholtz P, Parks DH. GTDB-Tk: A toolkit to classify genomes with the Genome Taxonomy Database. Bioinformatics. 2019 doi: 10.1093/bioinformatics/btz848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Parks DH, Chuvochina M, Chaumeil P-A, Rinke C, Mussig AJ, Hugenholtz P. A complete domain-to-species taxonomy for Bacteria and Archaea. Nat. Biotechnol. 2020;38(9):1079–1086. doi: 10.1038/s41587-020-0501-8. [DOI] [PubMed] [Google Scholar]
- 63.Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar X, Buchner A, Lai T, Steppi S, Jobb G, Förster W, Brettske I, Gerber S, Ginhart AW, Gross O, Grumann S, Hermann S, Jost R, König A, Liss T, Schleifer KH. ARB: A software environment for sequence data. Nucleic Acids Res. 2004;32(4):1363–1371. doi: 10.1093/nar/gkh293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Hedlund BP, Chuvochina M, Hugenholtz P, Konstantinidis KT, Murray AE, Palmer M, Parks DH, Probst AJ, Reysenbach AL, Rodriguez-R LM, Rossello-Mora R, Sutcliffe IC, Venter SN, Whitman WB. SeqCode: A nomenclatural code for prokaryotes described from sequence data. Nat. Microbiol. 2022;7:1702–1708. doi: 10.1038/s41564-022-01214-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Gu Z, Gu L, Eils R, Schlesner M, Brors B. circlize implements and enhances circular visualization in R. Bioinformatics. 2014;30(19):2811–2812. doi: 10.1093/bioinformatics/btu393. [DOI] [PubMed] [Google Scholar]
- 66.Gu Z, Eils R, Schlesner M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics. 2016;32(18):2847–2849. doi: 10.1093/bioinformatics/btw313. [DOI] [PubMed] [Google Scholar]
- 67.Seemann T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics. 2014;30(14):2068–2069. doi: 10.1093/bioinformatics/btu153. [DOI] [PubMed] [Google Scholar]
- 68.Karp PD, Midford PE, Billington R, Kothari A, Krummenacker M, Latendresse M, Ong WK, Subhraveti P, Caspi R, Fulcher C, Keseler IM, Paley SM. Pathway Tools version 23.0 update: Software for pathway/genome informatics and systems biology. Brief. Bioinform. 2021;22(1):109–126. doi: 10.1093/bib/bbz104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Caspi R, Billington R, Keseler IM, Kothari A, Krummenacker M, Midford PE, Ong WK, Paley S, Subhraveti P, Karp PD. The MetaCyc database of metabolic pathways and enzymes—a 2019 update. Nucleic Acids Res. 2020;48(D1):D445–D453. doi: 10.1093/nar/gkz862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Neukirchen S, Sousa FL. DiSCo: A sequence-based type-specific predictor of Dsr-dependent dissimilatory sulphur metabolism in microbial data. Microbial Genom. 2021;7:7. doi: 10.1099/mgen.0.000603. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The metagenome bins generated and analyzed during the current study are available in the European Nucleotide Archive (ENA) (https://www.ebi.ac.uk/ena/browser/home), under the Project PRJEB46766 (Table 1).