Abstract
Since its emergence at the end of 2019, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has caused the infection of more than 600 million people worldwide and has significant damage to global medical, economic, and political structures. Currently, a highly mutated variant of concern, SARS-CoV-2 Omicron, has evolved into many different subvariants mainly including BA.1, BA.2, BA.3, BA.4/5, and the recently emerging BA.2.75.2, BA.2.76, BA.4.6, BA.4.7, BA.5.9, BF.7, BQ.1, BQ.1.1, XBB, XBB.1, etc. Mutations in the N-terminal domain (NTD) of the spike protein, such as A67V, G142D, and N212I, alter the antigenic structure of Omicron, while mutations in the spike receptor binding domain (RBD), such as R346K, Q493R, and N501Y, increase the affinity for angiotensin-converting enzyme 2 (ACE2). Both types of mutations greatly increase the capacity of Omicron to evade immunity from neutralizing antibodies, produced by natural infection and/or vaccination. In this review, we systematically assess the immune evasion capacity of SARS-CoV-2, with an emphasis on the neutralizing antibodies generated by different vaccination regimes. Understanding the host antibody response and the evasion strategies employed by SARS-CoV-2 variants will improve our capacity to combat newly emerging Omicron variants.
Keywords: SARS-CoV-2, Omicron, Immune evasion, Mutations, Booster, Bivalent mRNA vaccine
Graphical Abstract

1. Introduction
1.1. Zoonotic coronaviruses and human pathogenic coronavirus
Zoonotic coronaviruses (CoV) are a large group of pathogens that often infect humans or animals to cause severe diseases, which potentially threaten public health and social development [1], [2]. In 1965, Tyrrell and Bynoe isolated the first human coronavirus, named B814, from the nasal wash of a cold patient using a human embryo tracheal culture [3]. Electron microscopy revealed that there are spinous processes shaped like coronas on their envelopes, hence the name “coronavirus” was proposed [4]. Coronavirus can be classified into 4 different genera by the International Committee of Taxonomy of Viruses (ICTV), namely Alpha-, Beta-, Gamma-, and Delta-coronaviruses [5]. Alpha- and Beta-coronaviruses infect mammals only, whereas Gamma- and Delta-coronaviruses infect birds and occasionally also infect mammals [5]. Currently, 7 coronaviruses are known to be capable of infecting humans [6]. SARS-CoV-2 is the 7th coronavirus with a confirmed ability to infect humans and can cause mild to severe disease depending on pathogenicity, infectivity, and subvariants [5], [6], [7], [8], [9], [10]. Of these 7 coronaviruses, HCoV-229E and HCoV-NL63 are in the Alpha-coronavirus, while SARS-CoV, SARS-CoV-2, HCoV-HKU1, HCoV-OC43, and MERS-CoV are all in the Beta-coronaviruses [5], [11]. The genera of Beta-coronavirus can further be divided into subgroups based on their genomic structures and phylogenetic relationships [5]. HCoV-OC43 and HCoV-HKU1 belong to subgroup A, SARS-CoV, SARS-CoV-2 to subgroup B, and MERS-CoV to subgroup C [5], [11] ( Fig. 1). HCoV-229E, HCoV-NL63, HCoV-HKU1, and HCoV-OC43 cause humans to develop mild upper respiratory symptoms and are commonly referred to as “common cold” CoVs [5], [6], [7], [8], [9], [10]. The human lower respiratory tract can be infected by SARS-CoV and MERS-CoV, leading to severe respiratory disease [5], [6], [7], [8], [9], [10].
Fig. 1.
Classification of the Coronaviridae. The Coronaviridae family has been divided into Orthocoronavirnae and Torovirinae subfamilies. The subfamily Orthocoronavirnae is divided into 4 genera, Alpha-, Beta-, Gamma-, and Delta-coronavirus. Viruses in the Alpha-genus are represented in blue and the Beta-genus are represented in orange. Viruses in the Beta-genus are divided into 4 subgroups A, B, C, and D, with subgroup A, represented in light green, subgroup B in orange, subgroup C in lavender, and subgroup D in light yellow. SARS-CoV-2 belongs to subgroup B in the genus Beta-coronavirus. The Variants of concerns of SARS-CoV-2 are represented in yellow, and Omicron subvariants are represented in purple.
1.2. Genome structure, hosts, symptoms, and transmission routes of SARS-CoV-2
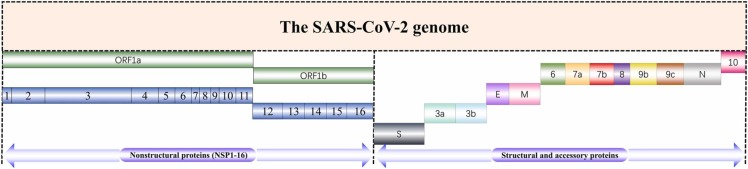
SARS-CoV-2 is a single-stranded RNA virus with about 30 kb long genome [12]. Its genome encodes 4 structural proteins called membrane (M), envelope (E), nucleocapsid (N), and spike (S) plus 16 nonstructural proteins and 9 accessory proteins [12] ( Fig. 2). SARS-CoV isolated from the 2003 pandemic was found to spread from bats to an intermediate host, the civet, which subsequently infected humans [6]. MERS-CoV, which caused Middle East Respiratory Syndrome (MERS) in 2012, was transmitted from bats to dromedary camels before infecting humans [6]. Analogous to SARS-CoV and MERS-CoV, bats are regarded to be natural hosts for SARS-CoV-2 [13]. However, the intermediate host of SARS-CoV-2 remains undetermined [13]. When infected by SARS-CoV-2, the most common COVID-19 symptoms in humans are fever, cough, myalgia, and tiredness [6], [12], [14], [15], [16]. More than half of the patients developed dyspnoea [6], [12], [17], [18]. In addition to respiratory symptoms, some patients may also develop gastrointestinal symptoms including diarrhea, vomiting, and abdominal pain [14]. How SARS-CoV-2 transmits among individuals is a major concern. It can spread through respiratory droplets, aerosols, contact transmission, and fecal-oral routes [19], [20], [21]. There is even evidence that SARS-CoV-2 can be transmitted from mothers to their offspring in a vertical manner during pregnancy and childbirth [22], [23], [24].
Fig. 2.
Overview of the SARS-CoV-2 genome encoding nonstructural proteins, structural proteins, and accessory proteins. Nonstructural proteins include ORF1a (NSP 1–11) and ORF1b (NSP 12–16). Structural proteins include membrane (M), envelope (E), nucleocapsid (N), and spike (S). Accessory proteins include ORF3a, ORF3b, ORF6, ORF7a, ORF7b, ORF8, ORF9b, ORF9c, and ORF10.
1.3. Variants of concerns and the dominating Omicron variant
Globally, as reported to The World Health Organization (WHO) on February 1st, 2023, there have been 753,651,712 confirmed cases of COVID-19, including 6,813,845 deaths. As of January 30th, 2023, a total of 13168935724 vaccine doses have been administered (https://covid19.who.int/). SARS-CoV-2 has evolved and mutated into newer variants over time and based on the risk level of the strains, WHO classifies the strains as variants under monitoring (VUM), variants of interest (VOI), variants of concern (VOC), variants of high consequence (VOHC). The detected SARS-CoV-2 variants were classified as VUM, VOI, and VOC, and none have yet been classified as VOHC. The VOCs have been the main type of variant causing several waves of the COVID-19 pandemic worldwide [25]. The SARS-CoV-2 VOCs are Alpha (B.1.1.7), Beta (B.1.351), Gamma (P.1), Delta (B.1.617.2), and Omicron (BA.1, etc.) [26]. This article focuses on Omicron, the major dominating and circulating variant worldwide at the moment. Omicron was initially detected in Botswana, Africa, on November 11th, 2021, and it was classified as a VOC by the WHO on November 26th, 2021 [27]. It rapidly caused the 4th wave of the SARS-CoV-2 epidemic in southern African countries [27]. Omicron swept the world in just a few weeks, causing a dramatic rise in infection numbers [27]. Over time, Omicron has evolved into several different subvariants, mainly including BA.1, BA.1.1, BA.2, BA.2.12.1, BA.2.13, BA.2.38, BA.2.75, BA.3, and BA.4/5 [28]. Currently, the most widespread and influential is BA.4/5, but new Omicron subvariants have recently appeared, including BA.2.75.2, BA.4.6, BA.4.7, BA.5.9, BF.7, BQ.1, BQ.1.1, BN.1, XBB, and CH.1.1 [29], [30], [31]. Over time, these new subvariants have contributed to the rise in the total number of COVID-19 infections, and BQ.1 is taking the lead now [32], [33]. The Omicron continues to evolve, and successively produces subvariants that are not only more transmissive but also more antibody-evasive [34].
1.4. The SARS-CoV-2 Omicron and neutralizing antibodies
1.4.1. Mutations on the Omicron S protein
Entry of coronavirus into host cells is mediated by the S protein [35] ( Fig. 3). A furin cleavage site divides the S protein into S1 and S2 subunits, where the S1 subunit is responsible for binding to the host cell receptor, and the S2 subunit is responsible for fusing the viral membrane to the cell membrane [1], [36]. The S protein serves as a target for neutralizing antibodies, drug development, and vaccine design [35], [37], [38], [39]. More than 30 mutations out of the over 50 mutations found in the Omicron variant are located on the S protein including about 20 mutations in the receptor binding domain (RBD) [39], [40]. BA.1 shares RBD mutations K417N, T478K, and N501Y with previous VOCs and contains many unique mutations including G339D, S371L, S373P, S375F, N440K, G446S, S477N, E484A, Q493R, Q496S, Q498R, Y505H [41] (Fig. 3). BA.1.1, in addition, possesses the mutation R346K [42]. BA.2 and BA.1 have multiple mutations in common, but BA.2 has some unique mutations in the RBD including S371F, T376A, D405N, and R408S [27], [42] (Fig. 3). BA.2.12.1 adds the mutation L452Q in the RBD [43]. BA.2.75 has the following additional mutations: G339H, G446S, and N460K in the RBD compared to BA.2, and BA.2.75.2 has mutations: R346T, F486S in the RBD in addition to the BA.2.75 mutations [29], [31], [44] (Fig. 3). BA.3 has fewer mutations compared to BA.2: T376A and R408S in the RBD are missing but has acquired one additional mutation: G446S [40]. BA.4/5 have acquired two missense mutations in the RBD: L452R and F486V, but the mutation Q493R is absent when compared to BA.2 [45]. BA.4.6 has two additional mutations (R346T, N658S) in the RBD than BA.4/5 [29], [31], [46], [47]. BA.4.7 has one more mutation (R346S) in the RBD compared to BA.4/5 [47] (Fig. 3). BA.5.9 contains an additional R346I mutation in the RBD [47] (Fig. 3). BF.7 has, likewise, the R346T mutation in the RBD [31]. BQ.1 has also acquired additional mutations compared to BA.4/5: K444T and N460K in the RBD [31]. BQ.1.1 harbors further one additional mutation (R346T) compared to BQ.1 [29], [31], [48] (Fig. 3). In contrast to the previously emerged SARS-CoV-2 variant, the large number of mutations in the S protein of the Omicron variant results in extensive antibody escape and creates a challenge to the effectiveness of existing vaccines and antibody therapies [49], [50] (Fig. 3). Firstly, A67V, Δ69–70, G142D, Δ143–145, N212I, Δ211, and ins214EPE on N-terminal Domain (NTD) lead to conformational changes and altered antigen structures in the N2, N3, N4, and N5 rings, thereby giving Omicron powerful immune evasion capabilities [28], [37], [39], [51], [52]. Secondly, although several point mutations reduce the binding of RBD to angiotensin-converting enzyme 2 (ACE2), the presence of compensatory mutations can restore the higher affinity of RBD to ACE2 [49] ( Fig. 4). For example, the R346K mutation affects the interaction network in the BA.1.1 RBD/hACE2 interface primarily through long-range modifications resulting in a higher affinity between the BA.1.1 RBD and hACE2 than BA.1 [53] (Fig. 4). T478K and Q498R establish hydrogen bonds or salt bridges with Q24, and D38 respectively, which increases the affinity between the RBD and ACE2 [51] ( Fig. 5). The Q493R mutation introduces a new salt bridge that replaces the hydrogen bond formed by the SARS-CoV-2 Wuhan-Hu-1 strain RBD, thereby reshaping the electrostatic interaction with ACE2 [49], [54] (Fig. 5). Mutations K417N, E484A, and G496S reduced RBD binding to ACE2, but mutation N501Y significantly enhanced RBD binding to ACE2 [51], [53], [55], [56], [57], [58] (Fig. 4). The N501Y mutation increases the binding of ACE2 to the RBD by 6-fold compared to the SARS-CoV-2 original Wuhan-Hu-1 strain RBD due to increased complementarity of shape between side chains [54], [59], [60] (Fig. 4).
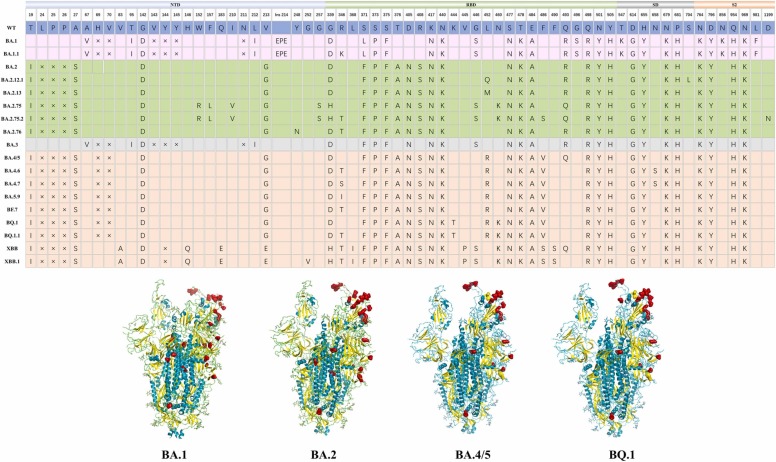
Fig. 3.
Comprehensive comparison of mutations between diverse Omicron subvariants. Mutations in the spike protein in Omicron subvariants wild type (WT), BA.1, BA.1.1, BA.2, BA.2.12.1, BA.2.13, BA.2.75, BA.2.75.2, BA.2.76, BA.3, BA.4/5, BA.4.6, BA.4.7, BF.7, BA.5.9, BQ.1, BQ.1.1, XBB, and XBB.1. All identical and different mutations on the receptor binding domain (RBD) between different Omicron subvariants are shown. “× ” represents a deletion. The spatial positions of mutations in BA.1, BA.2, BA.4/5, and BQ.1 are highlighted in structure models. The helix, loop, and sheet of S protein are colored in teal, grey, and yellow, respectively. Mutations are shown as red spheres (PDB 7EB4).
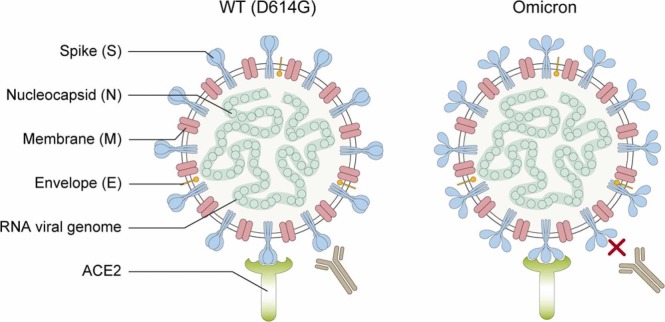
Fig. 4.
The mechanisms of viral attachment to host cells. The S protein enters the human body by binding to the angiotensin-converting enzyme 2 (ACE2) in human cells. In the left picture, the neutralizing antibody binds the S protein so the S protein cannot bind ACE2, thus preventing SARS-CoV-2 from entering the cell. In the right picture, the neutralizing antibody is unable to bind effectively to the S protein, thus allowing the S protein to bind ACE2 and SARS-CoV-2 can enter the cell. The increase of mutations in the Omicron S protein causes a conformational change in the S protein, which allows the S protein to bind the RBD better and prevents most neutralizing antibodies from being effective against Omicron.
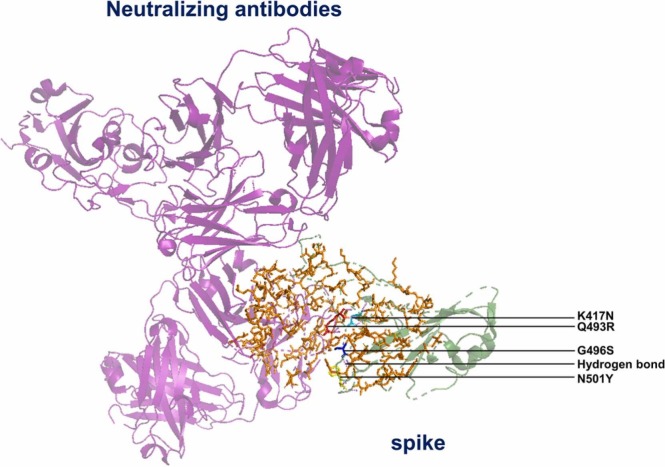
Fig. 5.
Interaction of S protein with neutralizing antibodies. Mutations K417N, Q493R, G496S, and N501Y are indicated by cyan, red, blue, and yellow, respectively. Hydrogen bonds are indicated in violet. The interaction residues between mutations K417N, Q493R, G496S, and N501Y on the S protein of Omicron and neutralizing antibodies are colored orange (PDB 7ZFD).
1.5. Omicron variants evade the humoral immune response induced by SARS-CoV-2 natural infection or vaccination
1.5.1. BA.1
1.5.1.1. Neutralization against BA.1 by sera from people with previous SARS-CoV-2 infection but not vaccinated
The neutralizing antibody titers of convalescent-phase sera against BA.1 are significantly reduced (1.7–33.8 times) compared to the D614G. In subjects previously infected with early pandemic virus (n = 32), Alpha (n = 18), Beta (n = 14), Gamma (n = 16), or Delta (n = 19), the relative sera dilution required to neutralize 50% of the live virus (focus reduction neutralization test (FRNT50)) against BA.1 were 28, 39, 28, 25, and 27, respectively, with decreases of 16.9-, 33.8-, 11.8-, 3.1-, and 1.7-fold for D614G, respectively [61] ( Fig. 6 ). Sera specimens from unvaccinated people infected with BA.1 (n = 11) or BA.2 (n = 8) demonstrated neutralizing antibody titers against BA.1 at 500 and 241, a 12.8- and 4.7-fold increase, respectively, in comparison to D614G [44] ( Table 1 ). These results show that BA.1 has the potential to escape the neutralization effect of antibodies raised against the previous SARS-CoV-2 VOCs, whereas antibodies raised against the BA.1 or BA.2 infection possess a much higher neutralization potential.
Fig. 6.
Neutralization ability of sera against Omicron by different vaccination and infection routines. Comparison of the neutralizing effect of respective sera from different groups. The number of antibodies represents the level of serum neutralization. A: SARS-CoV-2 infected but unvaccinated individuals, B: Vaccinated individuals without a history of SARS-CoV-2 infection, C: Individuals with SARS-CoV-2 breakthrough infection after vaccination, D: Individuals infected with SARS-CoV-2 before vaccination against Omicron.
Table 1.
Neutralizing capacity of Omicron and its subvariants in unvaccinated people previously infected with SARS-CoV-2.
| Classification | Variants of infection | Samples size | Experimental methods | Neutralizing titer | Fold change to D614G | Fold change to BA.1 | Refs. |
|---|---|---|---|---|---|---|---|
| BA.1 | Early pandemic virus | 32 | FRNT | BA.1: 28 | 16.9↓ | — | [61] |
| Alpha | 18 | FRNT | BA.1: 39 | 33.8↓ | — | ||
| Beta | 14 | FRNT | BA.1: 28 | 11.8↓ | — | ||
| Gamma | 16 | FRNT | BA.1: 25 | 3.1↓ | — | ||
| Delta | 19 | FRNT | BA.1: 27 | 1.7↓ | — | ||
| BA.1 | 11 | Pseudovirus neutralization | BA.1: 500 | 12.8↑ | — | [44] | |
| BA.2 | 8 | Pseudovirus neutralization | BA.1: 241 | 4.7↑ | — | ||
| BA.2 and its subvariants | BA.1 | 14 | Pseudovirus neutralization | BA.2: 142 | 1.0 | — | [77] |
| BA.2.75: 177 | 1.1↑ | — | |||||
| BA.1 | 11 | Pseudovirus neutralization | BA.2: 104 | 2.6↑ | 4.8↓ | [44] | |
| BA.2.75: 52 | 1.3↑ | 9.6↓ | |||||
| BA.2 | 8 | Pseudovirus neutralization | BA.2: 746 | 14.6↑ | 3.0↑ | ||
| BA.2.75: 155 | 3.0↑ | 1.5↓ | |||||
| BA.4/5 | BA.1 | 24 | FRNT | BA.4: 36 | — | 7.6↓ | [64] |
| BA.5: 37 | — | 7.4↓ | |||||
| BA.1 | 11 | Pseudovirus neutralization | BA.5: 18 | 2.1↓ | 27.7↓ | [44] | |
| BA.2 | 8 | Pseudovirus neutralization | BA.5: 103 | 2.0↑ | 2.3↓ |
Notes: Live-virus focus reduction neutralization test (FRNT)
1.5.1.2. Neutralization of BA.1 by sera from vaccine-only populations
Compared with D614G, serum neutralization of BA.1 is greatly reduced (8.5–23 times) in people previously not infected with SARS-CoV-2 but have been vaccinated. After two doses of the BNT162b2 vaccination (n = 24), the half maximal inhibitory concentration (IC50) by pseudovirus neutralization assay for BA.1 was 29, approximately 23 times lower than for D614G [62] (Fig. 6). Similarly, the average titer of BA.1 was less than 848, a decrease of more than 8.5-fold when compared to D614G in patients (n = 12) who received two shots of the mRNA-1273 vaccine [63] ( Table 2). Similarly, to the results from sera of infected but unvaccinated, sera from vaccinated-only correspondingly show a reduced neutralization capacity towards BA.1 compared to D614G.
Table 2.
Neutralizing capacity of Omicron and its subvariants in vaccinated populations.
| Classification | Vaccine name and schedule | Sample size | Experimental methods | Neutralizing titer | Fold change to D614G | Fold change to BA.1 | Refs. |
|---|---|---|---|---|---|---|---|
| BA.1 | Without infection→BNT162b2 × 2 | 24 | Pseudovirus neutralization | BA.1: 29 | 23.0↓ | — | [62] |
| Without infection→mRNA-1273 × 2 | 12 | Pseudovirus neutralization | BA.1: < 848 | > 8.5↓ | — | [63] | |
| BNT162b2/Ad26. COV2. S× 1/2→BA.1 | 15 | FRNT | BA.1: 507 | 4.0↓ | — | [64] | |
| CoronaVac× 3→BA.1 | 50 | Pseudovirus neutralization | BA.1: 852 | 1.8↓ | — | [30], [65] | |
| CoronaVac× 3→BA.2 | 39 | Pseudovirus neutralization | BA.1: 282 | 4.4↓ | — | [30], [65] | |
| CoronaVac× 3→BA.5 | 36 | Pseudovirus neutralization | BA.1: 209 | 5.4↓ | — | [30] | |
| BNT162b2 × 2→BA.1/BA.2 | 19 | Pseudovirus neutralization | BA.1: 5080 | 2.6↓ | — | [44] | |
| BNT162b2 × 3→BA.1/BA.2 | 9 | Pseudovirus neutralization | BA.1: 8364 | 4.2↓ | — | ||
| AZD1222 × 3 | 41 | FRNT | BA.1: 57 | 12.6↓ | — | [61] | |
| BNT162b2 × 3 | 20 | FRNT | BA.1: 649 | 14.2↓ | — | ||
| BBIBP-CorV× 3 | 10 | Pseudovirus neutralization | BA.1: 84 | 6.8↓ | — | [66] | |
| CoronaVac× 3 | 40 | Pseudovirus neutralization | BA.1: 122 | 5.3↓ | — | [30], [65] | |
| mRNA-1273 × 3 | 20 | Pseudovirus neutralization | BA.1: 850 | 2.9↓ | — | [67] | |
| BBIBP-CorV× 2+ZF2001 | 10 | Pseudovirus neutralization | BA.1: 172 | 9.4↓ | — | [66] | |
| CoronaVac× 2+ZF2001 | 38 | Pseudovirus neutralization | BA.1: 246 | 3.4↓ | — | [40] | |
| BNT162b2 × 2+mRNA-1273 | 20 | Pseudovirus neutralization | BA.1: 2852 | 6.5↓ | — | [44] | |
| CoronaVac× 2+BNT162b2 (without infection) | 75 | FRNT | — | 6.7↓ | — | [68] | |
| CoronaVac× 2+BNT162b2 (previous infection) | 26 | FRNT | — | 8.1↓ | — | ||
| BA.2 and its subvariants | Without infection→BNT162b2 × 2 | 24 | Pseudovirus neutralization | BA.2: 24 | 27.0↓ | 1.2↓ | [62] |
| Without Omicron infection→mRNA-1273/BNT162b2 × 1/2 | 22 | Pseudovirus neutralization | BA.2: 939 | 5.5↓ | 1.1↑ | [34] | |
| BA.2.12.1: 791 | 6.5↓ | 1.0 | |||||
| BNT162b2/mRNA-1273 × 1/2/3→BA.1 | 13 | Pseudovirus neutralization | BA.2: 3126 | 3.3↓ | — | [78] | |
| BA.2.12.1: 1622 | 6.3↓ | ||||||
| BA.2.75: 1587 | 6.5↓ | ||||||
| BNT162b2/mRNA-1273 × 3→BA.2 | 19 | Pseudovirus neutralization | BA.2: 800 | 1.2↓ | 1.6↑ | [69] | |
| BA.2.12.1: 552 | 1.8↓ | 1.1↑ | |||||
| BNT162b2 × 3/BNT162b2 × 1+mRNA-1273 × 2→BA.2 | 10 | FRNT | BA.2: 801 | 2.5↓ | — | [79] | |
| CoronaVac× 3→BA.1 | 50 | Pseudovirus neutralization | BA.2: 452 | 3.4↓ | 1.8↓ | [30], [65] | |
| BA.2.12.1: 244 | 6.3↓ | 3.4↓ | |||||
| BA.2.75: 197 | 7.9↓ | 4.3↓ | |||||
| BA.2.75.2: 34 | 45.7↓ | 25.0↓ | |||||
| CoronaVac× 3→BA.2 | 39 | Pseudovirus neutralization | BA.2: 696 | 1.7↓ | 2.4↑ | [30], [65] | |
| BA.2.12.1: 290 | 4.2↓ | 1.0 | |||||
| BA.2.75: 217 | 5.7↓ | 1.2↓ | |||||
| BA.2.75.2: 35 | 35.5↓ | 8.0↓ | |||||
| CoronaVac× 3→BA.5 | 36 | Pseudovirus neutralization | BA.2: 729 | 1.5↓ | 3.4↑ | [30] | |
| BA.2.75: 145 | 7.8↓ | 1.4↓ | |||||
| BA.2.75.2: 49 | 23.1↓ | 4.2↓ | |||||
| BNT162b2 × 2→BA.1/BA.2 | 19 | Pseudovirus neutralization | BA.2: 3372 | 4.0↓ | 1.5↓ | [44] | |
| BA.2.75: 2976 | 4.5↓ | 1.7↓ | |||||
| BNT162b2 × 3→BA.1/BA.2 | 9 | Pseudovirus neutralization | BA.2: 6470 | 5.4↓ | 1.2↓ | ||
| BA.2.75: 4393 | 8.0↓ | 1.9↓ | |||||
| BNT162b2/mRNA-1273 × 1/2/3/4→BA.1 | 13 | Pseudovirus neutralization | BA.2: 2142 | 3.3↓ | — | [80] | |
| mRNA vaccine× 2/3→BA.2 | 11 | Pseudovirus neutralization | BA.2: 1715 | 2.5↓ | — | ||
| mRNA vaccine× 3→BA.4/5 | 8 | Pseudovirus neutralization | BA.2: 5435 | 1.9↓ | — | ||
| BNT162b2 × 3→BA.1/2 (after 3 months) | 18 | Pseudovirus neutralization | BA.2.75.2: 600 | — | 18.0↓ | [29] | |
| BNT162b2 ×3→BA.1/2 (after 8 months) | 13 | Pseudovirus neutralization | BA.2.75.2: 200 | 40.0↓ | 15.0↓ | ||
| BNT162b2 ×3 | 30 | Pseudovirus neutralization | BA.2: 1725 | 3.7↓ | — | [81] | |
| BA.2.12.1: 1052 | 6.1↓ | ||||||
| BA.2.75: 1203 | 5.3↓ | ||||||
| mRNA-1273 ×3 | 20 | Pseudovirus neutralization | BA.2: 475 | 5.1↓ | 1.8↓ | [67] | |
| BA.2.75: 583 | 4.2↓ | 1.5↓ | |||||
| mRNA-1273 ×3/BNT162b2 ×3 | 15 | Pseudovirus neutralization | BA.2: 759 | 3.4↓ | — | [31] | |
| BA.2.75: 589 | 4.4↓ | ||||||
| BA.2.75.2: 54 | 48.4↓ | ||||||
| CoronaVac×3 | 40 | Pseudovirus neutralization | BA.2: 130 | 5.0↓ | 1.0 | [30], [65] | |
| BA.2.12.1: 105 | 6.2↓ | 1.1↓ | |||||
| BA.2.75: 90 | 7.2↓ | 1.3↓ | |||||
| BA.2.75.2: 22 | 29.6↓ | 5.5↓ | |||||
| BBIBP-CorV×3 | 20 | Pseudovirus neutralization | BA.2: <56 | >4.0↓ | >1.1↓ | [42] | |
| BBIBP-CorV×2+ZF2001 | 18 | Pseudovirus neutralization | BA.2: <42 | >12.7↓ | >2.5↓ | ||
| BNT162b2 ×2+mRNA-1273 | 20 | Pseudovirus neutralization | BA.2: 2288 | 8.2↓ | 1.2↓ | [44] | |
| BA.2.75: 1779 | 10.5↓ | 1.6↓ | |||||
| CoronaVac×2+ZF2001 | 38 | Pseudovirus neutralization | BA.2: 234 | 3.6↓ | 1.0 | [40] | |
| BA.2.12.1: 187 | 4.5↓ | 1.3↓ | |||||
| BA.4/5 and its subvariants | Without BA.1 infection→BNT162b2 ×1/2 | 18 | FRNT | BA.4: 211 | 19.5↓ | 1.0 | [64] |
| BA.5: 197 | 20.9↓ | 1.0 | |||||
| CoronaVac×3→BA.1 | 50 | Pseudovirus neutralization | BA.4/5: 107 | 14.4↓ | 7.9↓ | [30], [65] | |
| BQ.1: 32 | 48.6↓ | 26.6↓ | |||||
| BQ.1.1: 27 | 57.6↓ | 31.5↓ | |||||
| CoronaVac×3→BA.2 | 39 | Pseudovirus neutralization | BA.4/5: 175 | 7.1↓ | 1.6↓ | [30], [65] | |
| BQ.1: 45 | 27.6↓ | 6.2↓ | |||||
| BQ.1.1: 40 | 31.1↓ | 7.0↓ | |||||
| CoronaVac×3→BA.5 | 36 | Pseudovirus neutralization | BA.4/5: 508 | 2.2↓ | 2.4↑ | [30] | |
| BQ.1: 129 | 8.8↓ | 1.6↓ | |||||
| BQ.1.1: 77 | 14.7↓ | 2.7↓ | |||||
| CoronaVac×3→BA.1 | 50 | Pseudovirus neutralization | BA.4/5: 107 | 14.4↓ | 7.9↓ | [47] | |
| BA.5.9: 71 | 21.7↓ | 12.0↓ | |||||
| BA.4.6: 68 | 22.7↓ | 12.5↓ | |||||
| BA.4.7: 65 | 23.7↓ | 13.1↓ | |||||
| CoronaVac×3→BA.2 | 39 | Pseudovirus neutralization | BA.4/5: 175 | 7.1↓ | 1.6↓ | ||
| BA.5.9: 103 | 12.0↓ | 2.7↓ | |||||
| BA.4.6: 99 | 12.5↓ | 2.8↓ | |||||
| BA.4.7: 94 | 13.2↓ | 3.0↓ | |||||
| CoronaVac×3→BA.5 | 8 | Pseudovirus neutralization | BA.4/5: 481 | 2.9↓ | 2.9↑ | ||
| BA.5.9: 196 | 7.1↓ | 1.2↑ | |||||
| BA.4.6: 188 | 7.4↓ | 1.1↑ | |||||
| BA.4.7: 185 | 7.5↓ | 1.1↑ | |||||
| BNT162b2/mRNA-1273 ×3→BA.1 | 14 | Pseudovirus neutralization | BA.4/5: 263 | 5.4↓ | 4.2↓ | [69] | |
| BNT162b2/mRNA-1273 ×3→BA.2 | 19 | Pseudovirus neutralization | BA.4/5: 386 | 2.6↓ | 1.2↓ | [69] | |
| BNT162b2 ×3/BNT162b2 ×1+mRNA-1273 ×2→BA.2 | 10 | FRNT | BA.4.6: 204 | 9.8↓ | — | [79] | |
| BA.5: 155 | 12.9↓ | ||||||
| BNT162b2/mRNA-1273 ×1/2/3/4→BA.1 | 13 | Pseudovirus neutralization | BA.4/5: 647 | 11.2↓ | — | [80] | |
| BA.4.6: 686 | 10.6↓ | ||||||
| BA.4.7: 655 | 11.1↓ | ||||||
| BA.5.9: 718 | 10.1↓ | ||||||
| BF.7: 555 | 13.1↓ | ||||||
| mRNA vaccine×2/3→BA.2 | 11 | Pseudovirus neutralization | BA.4/5: 884 | 5.0↓ | — | [80] | |
| BA.4.6: 690 | 6.4↓ | ||||||
| BA.4.7: 815 | 5.4↓ | ||||||
| BA.5.9: 687 | 6.4↓ | ||||||
| BF.7: 699 | 6.3↓ | ||||||
| mRNA vaccine×3→BA.4/5 | 8 | Pseudovirus neutralization | BA.4/5: 3395 | 3.1↓ | — | [80] | |
| BA.4.6: 3140 | 3.3↓ | ||||||
| BA.4.7: 3316 | 3.2↓ | ||||||
| BA.5.9: 2959 | 3.5↓ | ||||||
| BF.7: 2953 | 3.5↓ | ||||||
| BNT162b2 ×3→BA.1/2 (after 3 months) | 18 | Pseudovirus neutralization | BA.5: 1500 | — | 7.0↓ | [29] | |
| BA.4.6: 1800 | 7.0↓ | ||||||
| BQ.1.1: 700 | 18.0↓ | ||||||
| BNT162b2 × 3→BA.1/2 (after 8 months) | 13 | Pseudovirus neutralization | BA.5: 400 | 20.0↓ | 7.5↓ | ||
| BA.4.6: 400 | 20.0↓ | 7.5↓ | |||||
| BQ.1.1: 200 | 40.0↓ | 15.0↓ | |||||
| BNT162b2 × 2→BA.1/BA.2 | 19 | Pseudovirus neutralization | BA.5: 878 | 15.5↓ | 5.7↓ | [44] | |
| BNT162b2 × 3→BA.1/BA.2 | 9 | Pseudovirus neutralization | BA.5: 1857 | 18.9↓ | 4.5↓ | ||
| CoronaVac× 3 | 40 | Pseudovirus neutralization | BA.4/5: 72 | 9.0↓ | 1.6↓ | [47] | |
| BA.5.9: 48 | 13.5↓ | 2.5↓ | |||||
| BA.4.6: 46 | 14.1↓ | 2.6↓ | |||||
| BA.4.7: 43 | 15.1↓ | 2.8↓ | |||||
| BNT162b2 × 3 | 27 | Pseudovirus neutralization | BA.4/5: 275 | 21.0↓ | 3.2↓ | [43] | |
| BNT162b2 × 3 | 21 | FRNT | BA.4.6: 51 | 16.5↓ | — | [79] | |
| BA.5: 47 | 17.7↓ | ||||||
| mRNA-1273 × 3 | 16 | Pseudovirus neutralization | BA.4/5: 449 | 35.6↓ | — | [46] | |
| BA.4.6: 225 | 71.1↓ | ||||||
| mRNA-1273/BNT162b2 × 3 | 15 | Pseudovirus neutralization | BA.4/5: 300 | 8.7↓ | — | [31] | |
| BA.4.6: 246 | 10.6↓ | ||||||
| BF.7: 238 | 10.9↓ | ||||||
| BQ.1: 140 | 18.6↓ | ||||||
| BQ.1.1: 114 | 22.9↓ | ||||||
| CoronaVac× 3 | 40 | Pseudovirus neutralization | BA.4/5: 72 | 9.0↓ | 1.6↓ | [30], [65] | |
| BQ.1: 27 | 24.1↓ | 4.5↓ | |||||
| BQ.1.1: 24 | 27.1↓ | 5.0↓ | |||||
| BNT162b2 × 2+mRNA-1273 | 20 | Pseudovirus neutralization | BA.5: 1057 | 17.7↓ | 2.6↓ | [44] | |
| CoronaVac× 2+ZF2001 | 38 | Pseudovirus neutralization | BA.4/5: 103 | 8.3↓ | 2.3↓ | [40] |
Notes: Live-virus focus reduction neutralization test (FRNT)
1.5.1.3. Neutralization of BA.1 by sera from people with Omicron breakthrough infection after vaccination
Compared to D614G, serum neutralization of BA.1 was slightly reduced (1.8–5.4 times) in people who developed Omicron breakthrough infection after vaccination. Samples (n = 15) were collected from South Africans infected with BA.1 after one or two doses of BNT162b2/Ad26. COV2. S vaccine showed that the FRNT50 was 507, a 4.0-fold decline relative to D614G [64] (Fig. 6). According to Yunlong Cao et al., the IC50 for BA.1 was 852, 282, and 209, a 1.8-, 4.4-, and 5.4-fold decrease when compared to D614G in serum samples infected with BA.1 (n = 50), BA.2 (n = 39), and BA.5 (n = 36) after three doses of CoronaVac vaccine [30], [65] (Table 2). Sera from people affected by BA.1 or BA.2 after two or three doses of the BNT162b2 vaccine was demonstrated to possess neutralizing antibody titers for BA.1 at 5080 and 8364, which are a decreased of 2.6- and 4.2- fold, when being compared to the D614G [44] (Fig. 6). Following a breakthrough infection of Omicron, the neutralization capacity of sera towards the BA.1 is still diminished compared to D614G, although the reduction seems less compared to sera from vaccinated-only or previous SARS-CoV-2 infected. This implies a higher immune evasion capacity of BA.1 compared to D614G.
1.5.1.4. Neutralization of BA.1 by sera from people who have received the first booster
1.5.1.4.1. A homologous first booster vaccine
The first dose of homologous booster increases the neutralizing activity against BA.1 to some extent but is still lower (2.9–14.2 times) than D614G. The three doses of AZD1222 or the BNT162b2 produced neutralizing antibodies against BA.1 with mean titers of 57 and 649, respectively, about 12.6- and 14.2-fold lower than those against the D614G [61] (Table 2). An infectious dose of 50% of exposed individuals (ID50) of 84 (6.8 times reduced compared to D614G) for BA.1 was found in sera specimens from healthy adults inoculated with three doses of the BBIBP-CorV vaccine [66] (Fig. 6). Vaccination with the three shots of the CoronaVac elicited neutralizing antibody with a mean titer of 122 against BA.1, a 5.3-fold decrease relative to D614G [30], [65]. Sera samples from adults inoculated with three shots of the mRNA-1273 vaccine were measured to possess a neutralizing antibody titer of 850 against BA.1, a 2.9-fold reduction compared to the D614G [67] (Table 2).
1.5.1.4.2. A heterologous first booster vaccine
The first dose of heterologous booster increases the neutralizing activity against BA.1 to some extent but is still lower (3.4–9.4 times) than D614G. Serum specimens from healthy adults who had received two shots of the BBIBP-CorV vaccine and one shot of the ZF2001 booster achieved an ID50 against BA.1 of 172, a 9.4-fold decrease relative to D614G [66]. The IC50 against BA.1 was 246, a 3.4-fold decrease relative to D614G in samples from a population inoculated with two CoronaVac shots and one ZF2001 shot [40] (Table 2). The IC50 against BA.1 was 2852, a 6.5-fold decrease relative to D614G in specimens from people inoculated with 2 BNT162b2 shots and 1 mRNA-1273 shot [44]. In subjects vaccinated with two shots of the CoronaVac vaccine and a BNT162b2 booster, the half-maximum plaque reduction neutralizing assays (PRNT50) against BA.1 decreased 6.7-fold relative to D614G in the group without a history of SARS-CoV-2 infection, and in the group with infection history, the PRNT50 against BA.1 decreased 8.1-fold relative to D614G [68] (Table 2).
The booster vaccine similar to the initial vaccination regime shows a reduced neutralization capacity when comparing BA.1 with D614G. However, the neutralization capacity against BA.1 still increases after the first booster dose, indicating a better immune response and lower immune evasion of BA.1 regardless of whether the booster vaccine is homologous or heterologous compared to the initial vaccine.
1.5.2. BA.2
1.5.2.1. Neutralization against BA.2 by sera from people with previous SARS-CoV-2 infection but not vaccinated
The neutralization levels against BA.1 and BA.2 range from 3.0 to 4.8-fold in the sera of convalescent individuals. The neutralizing antibody titers for BA.2 and BA.2.75 in serum samples from unvaccinated but infected BA.1 patients were 104 and 52, respectively, 4.8- and 9.6-fold lower than those for BA.1 [44]. Compared with BA.1, BA.2 had a neutralizing antibody titer of 746, a 3.0-fold increase, and BA.2.75 had a neutralizing antibody titer of 155, a 1.5-fold reduction in serum samples from unvaccinated but infected BA.2 [44] (Table 1). The neutralization capacity against BA.2 and BA.2.75 is very similar to BA.1 in infected but unvaccinated.
1.5.2.2. Neutralization against BA.2 by sera from vaccine-only populations
Neutralizing antibody titers against BA.1 and BA.2 by sera from vaccine-only populations (1.1–1.2 times difference) were not significantly different. Both were lower compared to D614G among vaccinated people with no history of Omicron infection. Vaccination with the two shots of the BNT162b2 elicited neutralizing antibody with a mean titer of 24 against BA.2, a 1.2-fold decrease when in contrast to BA.1 [62] (Fig. 6). Sera from people, who had received the mRNA vaccine but had no history of Omicron infection were found to have an ID50 for BA.2 of 939 and BA.2.12.1 of 791, which are not very far from the ID50 for BA.1 [34] (Table 2). As observed for BA.1 when comparing infected but unvaccinated and vaccinated-only, the neutralization capacity against BA.2 is similar in the two groups. Additionally, the neutralization capacity against BA.2 is comparable to that against BA.1.
1.5.2.3. Neutralization against BA.2 by sera from people with Omicron breakthrough infection after vaccination
Among the subvariants of BA.2, the neutralizing titers against BA.2.12.1 and BA.2.75 differed slightly, and BA.2.75.2 (4.2–25.0 times reduced in comparison with BA.1) having the lowest neutralizing antibody titers among vaccinated people with Omicron infection. Sera from recovering patients with BA.2 infection after three doses of the homologous or heterologous mRNA vaccine showed a 1.6- and 1.1-fold rise in neutralization titer against BA.2 (IC50: 800), and BA.2.12.1 (IC50: 552) compared to BA.1 [69] (Table 2). According to Yunlong Cao et al., the IC50 of BA.2, BA.2.12.1, BA.2.75, and BA.2.75.2 were 452, 244, 197, and 34, respectively, representing 1.8-, 3.4-, 4.3-, and 25.0-fold declines from BA.1, respectively [30], [65]. The IC50 for BA.2, BA.2.12.1, BA.2.75, and BA.2.75.2 were 696 (2.4 times higher than BA.1), 290 (similar to BA.1), 217 (1.2 times lower compared to BA,1), 35 (8.0 times lower compared to BA.1), respectively, according to the data of plasma samples from individuals, who received three shots of the CoronaVac vaccine and with BA.2 infection [30], [65]. The IC50 for BA.2, BA.2.75, and BA.2.75.2 were 729 (3.4 times higher than BA.1), 145 (1.4 times lower compared to BA.1), and 49 (4.2 times lower in comparison with BA.1), respectively, in plasma specimens from people with BA.5 infection after three shots of the CoronaVac vaccine [30], [65]. Sera from a population with breakthrough infection of BA.1 or BA.2 after two or three doses of the BNT162b2 vaccine were found to have neutralizing antibody titers for BA.2 of 3372 and 6470, respectively, a 1.5- and 1.2-fold decrease in contrast to BA.1 [44]. The titers for BA.2.75 were 2976 and 4393, a 1.7- and 1.9-fold decrease in comparison to BA.1, respectively [44] (Fig. 6). 3 months after the vaccination an effective dose of 50% (ED50) was 600 against BA.2.75.2, an 18-fold decrease when compared with BA.1, and the ED50 of BA.2.75.2 dropped sharply to 200, a 15.0-fold decline relative to BA.1, 8 months after vaccination in sera samples from healthcare workers (HCWs), who have received three injections of the BNT162b2 vaccine and experienced either BA.1 or BA.2 breakthrough infection [29] (Table 2). Following a breakthrough infection of Omicron, there is little difference in the neutralizing capacity of sera for BA.1 and BA.2. This demonstrates that the difference in immune evasion ability between BA.1 and BA.2 is minimal, and both are higher than D614G.
1.5.2.4. Neutralization of BA.2 by sera from people who have received the first booster
1.5.2.4.1. A homologous booster
There were minor differences in the neutralizing antibody titers between BA.1 and BA.2 (1.1–1.8 times lower than BA.1) of sera from patients inoculated with one dose of a homologous booster vaccine. Serum samples from adults inoculated with three shots of the mRNA-1273 vaccine were found to have an ID50 of 475 for BA.2 (1.8 times lower than BA.1), and 583 for BA.2.75 (1.5 times lower in comparison with BA.1) [67]. The IC50 against BA.2 was 130, approximately equal to that of BA.1, and those against BA.2.12.1, BA.2.75, and BA.2.75.2 were 105, 90, and 22, respectively, which is a decrease of 1.1, 1.3, and 5.5 compared to BA.1 in the plasma specimens from people receiving three doses of the CoronaVac vaccine [30], [65] (Table 2). In subjects inoculated with three doses of the BBIBP-CorV vaccine, the ID50 for BA.2 was measured to less than 56 whereas the ID50 for BA.1 was 1.1 times higher than this [42] (Fig. 6).
1.5.2.4.2. A heterologous booster
There were minor differences in the neutralizing antibody titers between BA.1 and BA.2 (1.2–2.5 times reduced compared to BA.1) of sera from patients inoculated with one dose of booster heterologous vaccine. Serum samples from people receiving two injections of the BBIBP-CorV vaccine and a ZF2001 heterologous booster were found to have an ID50 against BA.2 less than 42, a 2.5-fold reduction when compared to BA.1 [42]. Serum samples from patients receiving two injections of the BNT162b2 vaccine and one injection of the mRNA-1273 vaccine were found the neutralizing antibody titers for BA.2 and BA.2.75 to be 2288 (1.2 times lower in comparison with BA.1) and 1779 (1.6 times lower than BA.1), respectively [44]. Vaccination with the two shots of the CoronaVac and one dose of ZF2001 elicited an IC50 of 234 against BA.2, slightly less than that of BA.1, and an IC50 of 187 against BA.2.12.1, 1.3 times lower than BA.1 [40] (Table 2). These data show that there is no major difference in the immune evasion ability of BA.1 and BA.2 from neutralizing antibodies elicited by the first homologous or heterologous booster vaccine.
1.5.3. BA.4/5
1.5.3.1. Neutralization of BA.4/5 by sera from people with previous SARS-CoV-2 infection but not vaccinated
Compared with BA.1, serum neutralization of BA.4/5 is reduced (2.3–27.7 times compared to BA.1) in people not previously infected with SARS-CoV-2 but vaccinated. The neutralizing antibody titers for BA.5 in sera from unvaccinated individuals infected with either BA.1 or BA.2 were 18 and 103, respectively, a 27.7- and 2.3-fold decline in comparison to BA.1 [44]. The neutralizing antibody potencies of BA.4 and BA.5 in sera samples from unvaccinated but Omicron-infected South Africans were 36 and 37 respectively, a reduction of approximately 7.6 and 7.4 times compared to BA.1 [64] (Table 1). These data show that BA.4/5 has a higher immune evasion capacity than BA.1 from neutralizing antibodies elicited by serum in convalescent patients.
1.5.3.2. Neutralization of BA.4/5 by sera from vaccine-only individuals
The neutralizing antibody potency against BA.4 and BA.5 were 211 and 197, respectively, which were approximately equal to those of BA.1 in serum samples from a population, who received one or two doses of the BNT162b2 vaccine [64] (Table 2). The immune evasion capacity of BA.4/5 was higher than that of BA.1 from neutralizing antibodies elicited by serum from vaccination-only individuals.
1.5.3.3. Neutralization of BA.4/5 by sera from people with Omicron breakthrough infection after vaccination
Sera from patients recovered from BA.5 breakthrough infection showed high neutralization of BA.5-derived variants such as BQ.1 (1.6–26.6 times reduced compared to BA.1) and BQ.1.1 (2.7–31.5 times reduced compared to BA.1) [30] (Table 2). According to Yunlong Cao et al., the IC50 in plasma from a population injected with three injections of the CoronaVac vaccine but with BA.1 breakthrough infection against BA.4/5, BQ.1, and BQ.1.1 were 107, 32, and 27, respectively, a 7.9-, 26.6-, and 31.5-fold decline when compared to BA.1 [30], [65]. In individuals vaccinated with three doses of CoronaVac and infected with BA.2, the IC50 for BA.4/5, BQ.1, and BQ.1.1 were 175, 45, and 40, respectively, a reduction of 1.6, 6.2, and 7.0 times compared to BA.1 [30], [65]. Whereas in individuals vaccinated with three doses of CoronaVac and infected with BA.5, the IC50 for BA.4/5, BQ.1, and BQ.1.1 were 508 (2.4 times higher), 129 (1.6 times lower), and 77 (2.7 times lower), when compared to those of BA.1 [30] (Table 2). Similarly, plasma samples from people infected with Omicron after three doses of the CoronaVac vaccine and BA.1 breakthrough infection were demonstrated to have IC50 values for BA.4/5, BA.5.9, BA.4.6, and BA.4.7 of 107, 71, 68, and 65, with respective 7.9-, 12.0-, 12.5-, 13.1-fold decrease when compared to those of BA.1 [47]. In the group with BA.2 breakthrough infection, the IC50 values were 175, 103, 99, and 94, respectively, with 1.6-, 2.7-, 2.8-, and 3.0-fold lower than those of BA.1 [47]. Lastly, in the group with BA.5 breakthrough infection, the IC50 for BA.4/5, BA.5.9, BA.4.6, and BA.4.7 were 481 (2.9 times increased than BA.1), 196 (1.2 times higher compared to BA.1), 188 (1.1 times increased in comparison with BA.1), and 185 (1.1 times higher in comparison with BA.1), respectively [47] (Fig. 6). Sera specimens from patients affected by BA.1 or BA.2 after three injections of mRNA COVID-19 vaccine were found to have a neutralizing antibody potency against BA.4/5 of 263 and 386, respectively, a 4.2- and 1.2- fold decrease when comparing to BA.1 [69] (Fig. 6). In sera from HCWs with BA.1 or BA.2 infection after three shots of the BNT162b2 vaccine, 3 months after vaccination, the ED50 for BQ.1.1 was 700, an 18.0-fold decrease relative to BA.1, 8 months after vaccination, the ED50 for BQ.1.1 was 200, a 15.0-fold decline compared to BA.1 [29]. According to CheeWah Tan et al., the neutralizing antibody potency was measured to be 878 for BA.5, a 5.7-fold decrease relative to BA.1 in sera from people with breakthrough BA.1 or BA.2 infection after two doses of BNT162b2 vaccine [44] (Table 2). The above data suggest that following a breakthrough infection of Omicron, the neutralizing capacity of serum for BA.4/5 is reduced compared to BA.1.
1.5.3.4. Neutralization of BA.4/5 by sera from people who have received the first booster
1.5.3.4.1. A homologous booster
BA.4/5 (1.6–3.2 times reduced) had a slightly lower neutralizing power than BA.1 in serum specimens from individuals inoculated with the first dose of the homologous booster. In serum from individuals vaccinated with the first dose of homologous booster vaccination (three shots of the CoronaVac) elicited neutralizing antibodies with a mean titer of 72, 48, 46, and 43 against BA.4/5, BA.5.9, BA.4.6, BA.4.7 with 1.6-, 2.5-, 2.6-, and 2.8-fold reduction when compared to those of BA.1 [47] (Fig. 6). Sera from people injected with three doses of the BNT162b2 vaccine were found to have a neutralizing antibody titer against BA.4/5 of 275, a 3.2-fold decrease when comparing with BA.1 [43]. Plasma from people given three shots of the CoronaVac vaccine was demonstrated to have an IC50 against BA.4/5, BQ.1, and BQ.1.1 of 72, 27, and 24, respectively, a 1.6-, 4.5-, and 5.0-fold decline compared to BA.1 [30], [65] (Table 2).
1.5.3.4.2. A heterologous booster
The neutralizing capacity of BA.4/5 (2.3–2.6 times reduced) was slightly higher than that of BA.1 in serum specimens from individuals inoculated with the first dose of the heterologous booster. In plasma samples from people inoculated with two shots of the CoronaVac vaccine and one ZF2001 booster shot, the IC50 was 103 against BA.4/5, a 2.3-fold decrease relative to BA.1 [40] (Table 2). In subjects vaccinated with two doses of the BNT162b2 vaccine and one shot of the mRNA-1273 vaccine, the neutralizing antibody titer against BA.5 was 1057, a 2.6-fold decrease compared to BA.1 [44] (Fig. 6). These data show that BA.4/5 has a higher immune evasion capacity than BA.1 from neutralizing antibodies elicited by people who received their first dose of a homologous or heterologous booster vaccine. Furthermore, the emerging subvariants BA.4.6, BA.4.7, BA.5.9, BF.7, BQ.1, and BQ.1.1 derived from BA.4/5 also possess substantial immune evasion capabilities from a homologous or heterologous vaccine-elicited neutralizing antibodies (Table 2).
1.6. Neutralization of newly emerging Omicron subvariants by sera from people receiving the second monovalent or bivalent booster
As Omicron continues to evolve and more and more subvariants emerge, for example, BA.2.75.2 have evolved from BA.2.75, BA.2.75 have evolved from BA.2, and BA.4.6, BF.7, BQ.1, and BQ.1.1 subvariants are descended from BA.4/5 [31], [46], [48], [70]. These newly emerged subvariants have stronger immune evasion ability than BA.1, BA. 2, and BA.4/5 [31]. According to Delphine Planas et al., in HCWs vaccinated with three doses of BNT162b2, the neutralizing activity for BA.2.75.2, BA.4.6, and BQ.1.1 was undetectable or hardly detectable 4 months after vaccination [29].
The second booster (including monovalent and bivalent vaccine) was developed to combat the newly emerged Omicron variant [71]. Compared to three doses of vaccine, the neutralizing effect against Omicron of four doses of the vaccine was significantly improved [71]. The 4th dose of the bivalent vaccine has a higher neutralization capacity than the 4th dose of the monovalent vaccine [71]. Sera from participants, who received the 4th shot of the bivalent vaccine and participants, who received the 4th shot of the monovalent vaccine were separately evaluated and found to have a 50% fluorescence focal reduction neutralization titer (FFRNT50) after the 4th dose of monovalent the BNT162b2 vaccine of 2605, 236, 232, 99, 58, and 41 for wild type (WT), BA.4/5, BA.4.6, BA.2.75.2, BQ.1.1, and XBB.1, respectively, an increase of 3.0, 2.9, 2.3, 2.1, 1.8, and 1.5 times relative to the FFRNT50 measured with only three doses of the BNT162b2 vaccine [71]. Furthermore, the FFRNT50 after the 4th dose of bivalent the BNT162b2 vaccine were 3328, 856, 905, 196, 252, and 84 for WT, BA.4/5, BA.4.6, BA.2.75.2, BQ.1.1, and XBB.1, respectively, an increase of 5.8, 13.0, 11.1, 6.7, 8.7, and 4.8 times when compared with the FFRNT50 measured with three shots of the BNT162b2 vaccine [71]. The FRNT50 against BA.1, BA.5, BA.2.75.2, and BQ.1.1 in groups inoculated with a monovalent booster was 60, 50, 23, and 19, respectively, a 14.3-, 17.1-, 37.3-, and 45.1-fold decrease compared with those of D614G [72]. In the group vaccinated with two monovalent boosters, the FRNT50 for BA.1, BA.5, BA.2.75.2, and BQ.1.1 was 408, 250, 98, and 73, respectively, a reduction of 5.8-, 9.4-, 24.0-, and 32.2-fold compared with corresponding values of D614G, while in the group vaccinated with the bivalent booster, the FRNT50 for BA.1, BA.5, BA.2.75.2, and BQ.1.1 was 618, 576, 201, and 112, respectively, a decline of 4.0, 4.3, 12.3, and 22.2 times compared with titers against D614G [72]. Neutralizing antibody titers against BA.5 were increased from 184 to 2829 (15-fold higher) after the monovalent mRNA booster and were increased from 212 to 3693 (17-fold higher) after the bivalent mRNA booster [73]. Hence, these findings prove that the bivalent mRNA vaccine can enhance humoral immunity against the newly emerged Omicron subvariants such as BA.5, BQ.1.1.
Those who have been vaccinated or infected with SARS-CoV-2 at a previous time are still likely to be affected by Omicron as their sera have low neutralizing potency against Omicron [26], [63]. Two shots of the vaccine may not be adequate to protect the vaccine from Omicron infection, therefore boosters are recommended [26], [42], [63], [74]. A bivalent mRNA vaccine, as a booster vaccine encoding two Omicron S proteins (WT and BA.4/5), can induce protective immunity against both previous and current SARS-CoV-2 Omicron variants [75]. Simultaneously, another study also showed that a bivalent vaccine produces a greater breadth and strength of neutralizing antibodies than a monovalent vaccine [75]. Clinical trials have shown the safety and effectiveness of bivalent mRNA vaccine as demonstrated by Spyros Chalkias et al. [75], [76]. Furthermore, the second and even annual booster will be a powerful tool in fighting against new Omicron subvariants such as BA.4.6, BQ.1, and BQ.1.1 or other emerging variants during the global vaccination campaign.
2. Summary
BA.4/5 has the highest immune evasion capacity, followed by BA.2.75 and BA.2.12, and finally BA.1 and BA.2, and they all have a higher capacity for immune evasion than D614G. The recently emerged Omicron subvariants BA.4.6, BF.7, BQ.1, and BQ.1.1 have a greatly enhanced immune evasion capacity. Neutralizing antibodies induced by the current available COVID-19 vaccine continue to wane over time after vaccination. As new SARS-CoV-2 Omicron variants emerge, possibly not exclusively, BA.1, BA.1.1, BA.2, BA. 2.75, BA.2.75.2, BA.3, BA.4/5, BA.4.6, BA.5.9, BF.7, BQ.1, etc. Infection by such Omicron subvariants with a high capacity for immune invasion might make the scenario more complicated when vaccine options are relatively limited in the general population. Even though vaccine-induced neutralizing antibodies are observed to have a reduced ability against almost all Omicron variants, scheduled or annual boosters, especially bivalent vaccines, are still encouraged to strengthen the immune system in the future to protect the human population from severe diseases and death.
Credit authorship contribution statement
H.Z. conceived ideas and supervised the work. LD.W. wrote the initial draft and made figures. H.Z., M.M., and PF.W. reviewed and edited the manuscript. All authors contributed substantially to the discussion of the content, reviewed, edited, and approved the manuscript before submission.
Acknowledgments
We thank the support from Chengdu University of TCM (No. 030040018).
Conflict of interest
The authors declare that they have no conflict of interest.
Biographies

Lidong Wang, she received her Bachelor's degree from Qilu College of Medicine in 2021. She is currently a master's student at Chengdu University of Traditional Chinese Medicine. Her main research interest involves microbiology, immunology, and virology.

Dr. Michelle Møhlenberg, Ph.D. She received her Ph.D from Aarhus University in Denmark, where she investigated the effect of genetic variants within the human IFN system upon viral liver and lung infections. She has published several articles, three of which focused on SARS-CoV-2 (EMBO reports, Journal of Interferon and Cytokine Research, Clinical Microbiology Reviews). She is currently working as a Postdoc employed at both Aarhus University, Denmark, within the field of Human Genetics and Immunology with the aim of characterizing the physiological and cellular response towards viral infections, among others: SARS-CoV-2.

Prof. Pengfei Wang, Ph.D. He is a professor from the School of Life Sciences, Fudan University, Shanghai, China. He was working as a research scientist in Columbia University and a postdoctoral fellow in The Rockefeller University. He earned his Ph.D in Genetics at Fudan University and B.S in Biology at Beijing Normal University in China. His study has been focused on HIV and coronaviruses and has published more than 40 peer-reviewed journal articles (Nature, Cell Host & Microbe). Google Scholar h-index 31 (>6000 citations).

Dr. Hao Zhou, Ph.D. He is a professor from Chengdu University of Traditional Chinese Medicine (Sichuan, China). He received his Ph.D degree in molecular biology from Aarhus University (Aarhus, Denmark) and then worked as a postdoctoral fellow at the University of Massachusetts Medical School (Worcester, USA) and Grossman School of Medicine, New York University (New York, USA), studying infectious diseases (SARS-CoV-2), innate immunity and preventive and therapeutic tools (vaccines and antibodies) against SARS-CoV-2. He has published several peer-reviewed papers in high-profile journals (Nature Reviews Immunology, Clinical Microbiology Reviews, Cell Reports, mBio).
References
- 1.V'Kovski P., Kratzel A., Steiner S., Stalder H., Thiel V. Coronavirus biology and replication: implications for SARS-CoV-2. Nat. Rev. Microbiol. 2021;19(3):155–170. doi: 10.1038/s41579-020-00468-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ye Z.W., Yuan S., Yuen K.S., Fung S.Y., Chan C.P., Jin D.Y. Zoonotic origins of human coronaviruses. Int J. Biol. Sci. 2020;16(10):1686–1697. doi: 10.7150/ijbs.45472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Tyrrell D.A., Bynoe M.L. Cultivation of a novel type of common-cold virus in organ cultures. Br. Med J. 1965;1(5448):1467–1470. doi: 10.1136/bmj.1.5448.1467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Tyrrell D.A., Almeida J.D., Cunningham C.H., Dowdle W.R., Hofstad M.S., McIntosh K., Tajima M., Zakstelskaya L.Y., Easterday B.C., Kapikian A., Bingham R.W. Coronaviridae. Intervirology. 1975;5(1–2):76–82. doi: 10.1159/000149883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Cui J., Li F., Shi Z.L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 2019;17(3):181–192. doi: 10.1038/s41579-018-0118-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Tang D., Comish P., Kang R. The hallmarks of COVID-19 disease. PLoS Pathog. 2020;16(5) doi: 10.1371/journal.ppat.1008536. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Schneider W.M., Luna J.M., Hoffmann H.H., Sánchez-Rivera F.J., Leal A.A., Ashbrook A.W., Le Pen J., Ricardo-Lax I., Michailidis E., Peace A., Stenzel A.F., Lowe S.W., MacDonald M.R., Rice C.M., Poirier J.T. Vol. 184. 2021. Genome-Scale Identification of SARS-CoV-2 and Pan-coronavirus Host Factor Networks; pp. 120–132. (Cell). e14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hasoksuz M., Kilic S., Sarac F. Coronaviruses and SARS-COV-2. Turk. J. Med. Sci. 2020;50:549–556. doi: 10.3906/sag-2004-127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Zumla A., Chan J.F., Azhar E.I., Hui D.S., Yuen K.Y. Coronaviruses - drug discovery and therapeutic options. Nat. Rev. Drug Disco. 2016;15(5):327–347. doi: 10.1038/nrd.2015.37. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Chen B., Tian E.K., He B., Tian L., Han R., Wang S., Xiang Q., Zhang S., El Arnaout T., Cheng W. Overview of lethal human coronaviruses. Signal Transduct. Target Ther. 2020;5(1):89. doi: 10.1038/s41392-020-0190-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Chan J.F., Lau S.K., To K.K., Cheng V.C., Woo P.C., Yuen K.Y. Middle East respiratory syndrome coronavirus: another zoonotic betacoronavirus causing SARS-like disease. Clin. Microbiol Rev. 2015;28(2):465–522. doi: 10.1128/CMR.00102-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Malone B., Urakova N., Snijder E.J., Campbell E.A. Structures and functions of coronavirus replication-transcription complexes and their relevance for SARS-CoV-2 drug design. Nat. Rev. Mol. Cell Biol. 2022;23(1):21–39. doi: 10.1038/s41580-021-00432-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.de Wit E., van Doremalen N., Falzarano D., Munster V.J. SARS and MERS: recent insights into emerging coronaviruses. Nat. Rev. Microbiol. 2016;14(8):523–534. doi: 10.1038/nrmicro.2016.81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Pan L., Mu M., Yang P., Sun Y., Wang R., Yan J., Li P., Hu B., Wang J., Hu C., Jin Y., Niu X., Ping R., Du Y., Li T., Xu G., Hu Q., Tu L. Clinical characteristics of COVID-19 patients with digestive symptoms in Hubei, China: a descriptive, cross-sectional, multicenter study. Am. J. Gastroenterol. 2020;115(5):766–773. doi: 10.14309/ajg.0000000000000620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Menni C., Valdes A.M., Polidori L., Antonelli M., Penamakuri S., Nogal A., Louca P., May A., Figueiredo J.C., Hu C., Molteni E., Canas L., Österdahl M.F., Modat M., Sudre C.H., Fox B., Hammers A., Wolf J., Capdevila J., Chan A.T., David S.P., Steves C.J., Ourselin S., Spector T.D. Symptom prevalence, duration, and risk of hospital admission in individuals infected with SARS-CoV-2 during periods of omicron and delta variant dominance: a prospective observational study from the ZOE COVID Study. Lancet. 2022;399(10335):1618–1624. doi: 10.1016/S0140-6736(22)00327-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Safiabadi Tali S.H., LeBlanc J.J., Sadiq Z., Oyewunmi O.D., Camargo C., Nikpour B., Armanfard N., Sagan S.M., Jahanshahi-Anbuhi S. Tools and techniques for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)/COVID-19 Detection. Clin. Microbiol Rev. 2021;34(3) doi: 10.1128/CMR.00228-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Huang C., Wang Y., Li X., Ren L., Zhao J., Hu Y., Zhang L., Fan G., Xu J., Gu X., Cheng Z., Yu T., Xia J., Wei Y., Wu W., Xie X., Yin W., Li H., Liu M., Xiao Y., Gao H., Guo L., Xie J., Wang G., Jiang R., Gao Z., Jin Q., Wang J., Cao B. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 2020;395(10223):497–506. doi: 10.1016/S0140-6736(20)30183-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Su Y., Yuan D., Chen D.G., Ng R.H., Wang K., Choi J., Li S., Hong S., Zhang R., Xie J., Kornilov S.A., Scherler K., Pavlovitch-Bedzyk A.J., Dong S., Lausted C., Lee I., Fallen S., Dai C.L., Baloni P., Smith B., Duvvuri V.R., Anderson K.G., Li J., Yang F., Duncombe C.J., McCulloch D.J., Rostomily C., Troisch P., Zhou J., Mackay S., DeGottardi Q., May D.H., Taniguchi R., Gittelman R.M., Klinger M., Snyder T.M., Roper R., Wojciechowska G., Murray K., Edmark R., Evans S., Jones L., Zhou Y., Rowen L., Liu R., Chour W., Algren H.A., Berrington W.R., Wallick J.A., Cochran R.A., Micikas M.E., Wrin T., Petropoulos C.J., Cole H.R., Fischer T.D., Wei W., Hoon D.S.B., Price N.D., Subramanian N., Hill J.A., Hadlock J., Magis A.T., Ribas A., Lanier L.L., Boyd S.D., Bluestone J.A., Chu H., Hood L., Gottardo R., Greenberg P.D., Davis M.M., Goldman J.D., Heath J.R. Multiple early factors anticipate post-acute COVID-19 sequelae. Cell. 2022;185(5):881–895. doi: 10.1016/j.cell.2022.01.014. e20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Xiao F., Tang M., Zheng X., Liu Y., Li X., Shan H. Evidence for gastrointestinal infection of SARS-CoV-2. Gastroenterology. 2020;158(6):1831–1833. doi: 10.1053/j.gastro.2020.02.055. e3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Klompas M., Baker M.A., Rhee C. Airborne transmission of SARS-CoV-2 theoretical considerations and available evidence. Jama-J. Am. Med. Assoc. 2020;324(5):441–442. doi: 10.1001/jama.2020.12458. [DOI] [PubMed] [Google Scholar]
- 21.Wiersinga W.J., Rhodes A., Cheng A.C., Peacock S.J., Prescott H.C. Pathophysiology, transmission, diagnosis, and treatment of coronavirus disease 2019 (COVID-19): a review. Jama. 2020;324(8):782–793. doi: 10.1001/jama.2020.12839. [DOI] [PubMed] [Google Scholar]
- 22.Allotey J., Chatterjee S., Kew T., Gaetano A., Stallings E., Fernández-García S., Yap M., Sheikh J., Lawson H., Coomar D., Dixit A., Zhou D., Balaji R., Littmoden M., King Y., Debenham L., Llavall A.C., Ansari K., Sandhu G., Banjoko A., Walker K., O'Donoghue K., van Wely M., van Leeuwen E., Kostova E., Kunst H., Khalil A., Brizuela V., Broutet N., Kara E., Kim C.R., Thorson A., Oladapo O.T., Zamora J., Bonet M., Mofenson L., Thangaratinam S. SARS-CoV-2 positivity in offspring and timing of mother-to-child transmission: living systematic review and meta-analysis. Bmj. 2022;376 doi: 10.1136/bmj-2021-067696. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Pirkle C.M. Transmission of SARS-CoV-2 from mother to baby is rare. Bmj. 2022;376:o593. doi: 10.1136/bmj.o593. [DOI] [PubMed] [Google Scholar]
- 24.Dong L., Tian J., He S., Zhu C., Wang J., Liu C., Yang J. Possible vertical transmission of SARS-CoV-2 from an infected mother to her newborn. Jama. 2020;323(18):1846–1848. doi: 10.1001/jama.2020.4621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Boehm E., Kronig I., Neher R.A., Eckerle I., Vetter P., Kaiser L. Novel SARS-CoV-2 variants: the pandemics within the pandemic. Clin. Microbiol Infect. 2021;27(8):1109–1117. doi: 10.1016/j.cmi.2021.05.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zhou H., Møhlenberg M., Thakor J.C., Tuli H.S., Wang P., Assaraf Y.G., Dhama K., Jiang S. Sensitivity to vaccines, therapeutic antibodies, and viral entry inhibitors and advances to counter the SARS-CoV-2 omicron variant. Clin. Microbiol Rev. 2022;35(3) doi: 10.1128/cmr.00014-22. e0001422. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Viana R., Moyo S., Amoako D.G., Tegally H., Scheepers C., Althaus C.L., Anyaneji U.J., Bester P.A., Boni M.F., Chand M., Choga W.T., Colquhoun R., Davids M., Deforche K., Doolabh D., du Plessis L., Engelbrecht S., Everatt J., Giandhari J., Giovanetti M., Hardie D., Hill V., Hsiao N.Y., Iranzadeh A., Ismail A., Joseph C., Joseph R., Koopile L., Kosakovsky Pond S.L., Kraemer M.U.G., Kuate-Lere L., Laguda-Akingba O., Lesetedi-Mafoko O., Lessells R.J., Lockman S., Lucaci A.G., Maharaj A., Mahlangu B., Maponga T., Mahlakwane K., Makatini Z., Marais G., Maruapula D., Masupu K., Matshaba M., Mayaphi S., Mbhele N., Mbulawa M.B., Mendes A., Mlisana K., Mnguni A., Mohale T., Moir M., Moruisi K., Mosepele M., Motsatsi G., Motswaledi M.S., Mphoyakgosi T., Msomi N., Mwangi P.N., Naidoo Y., Ntuli N., Nyaga M., Olubayo L., Pillay S., Radibe B., Ramphal Y., Ramphal U., San J.E., Scott L., Shapiro R., Singh L., Smith-Lawrence P., Stevens W., Strydom A., Subramoney K., Tebeila N., Tshiabuila D., Tsui J., van Wyk S., Weaver S., Wibmer C.K., Wilkinson E., Wolter N., Zarebski A.E., Zuze B., Goedhals D., Preiser W., Treurnicht F., Venter M., Williamson C., Pybus O.G., Bhiman J., Glass A., Martin D.P., Rambaut A., Gaseitsiwe S., von Gottberg A., de Oliveira T. Rapid epidemic expansion of the SARS-CoV-2 Omicron variant in southern Africa. Nature. 2022;603(7902):679–686. doi: 10.1038/s41586-022-04411-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Iketani S., Liu L., Guo Y., Liu L., Chan J.F., Huang Y., Wang M., Luo Y., Yu J., Chu H., Chik K.K., Yuen T.T., Yin M.T., Sobieszczyk M.E., Huang Y., Yuen K.Y., Wang H.H., Sheng Z., Ho D.D. Antibody evasion properties of SARS-CoV-2 Omicron sublineages. Nature. 2022;604(7906):553–556. doi: 10.1038/s41586-022-04594-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Planas D., Bruel T., Staropoli I., Guivel-Benhassine F., Porrot F., Maes P., Grzelak L., Prot M., Mougari S., Planchais C., Puech J., Saliba M., Sahraoui R., Fémy F., Morel N., Dufloo J., Sanjuán R., Mouquet H., André E., Hocqueloux L., Simon-Loriere E., Veyer D., Prazuck T., Péré H., Schwartz O. Resistance of Omicron subvariants BA.2.75.2, BA.4.6, and BQ.1.1 to neutralizing antibodies. Nat. Commun. 2023;14(1):824. doi: 10.1038/s41467-023-36561-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Cao Y., Jian F., Wang J., Yu Y., Song W., Yisimayi A., Wang J., An R., Chen X., Zhang N., Wang Y., Wang P., Zhao L., Sun H., Yu L., Yang S., Niu X., Xiao T., Gu Q., Shao F., Hao X., Xu Y., Jin R., Shen Z., Wang Y., Xie X.S. Imprinted SARS-CoV-2 humoral immunity induces convergent Omicron RBD evolution. Nature. 2023;614(7948):521–529. doi: 10.1038/s41586-022-05644-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Qu P., Evans J.P., Faraone J.N., Zheng Y.M., Carlin C., Anghelina M., Stevens P., Fernandez S., Jones D., Lozanski G., Panchal A., Saif L.J., Oltz E.M., Xu K., Gumina R.J., Liu S.L. Enhanced neutralization resistance of SARS-CoV-2 Omicron subvariants BQ.1, BQ.1.1, BA.4.6, BF.7, and BA.2.75.2. Cell Host Microbe. 2023;31(1):9–17.e3. doi: 10.1016/j.chom.2022.11.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Iacobucci G. Covid-19: Hospital admissions rise in England as some trusts reinstate mask requirements. Bmj. 2022;379:o2440. doi: 10.1136/bmj.o2440. [DOI] [PubMed] [Google Scholar]
- 33.Mahase E. Covid-19: Cases rise in England and Wales as hospital admissions remain steady. Bmj. 2022;378:o2307. doi: 10.1136/bmj.o2307. [DOI] [PubMed] [Google Scholar]
- 34.Wang Q., Guo Y., Iketani S., Nair M.S., Li Z., Mohri H., Wang M., Yu J., Bowen A.D., Chang J.Y., Shah J.G., Nguyen N., Chen Z., Meyers K., Yin M.T., Sobieszczyk M.E., Sheng Z., Huang Y., Liu L., Ho D.D. Antibody evasion by SARS-CoV-2 Omicron subvariants BA.2.12.1, BA.4 and BA.5. Nature. 2022;608(7923):603–608. doi: 10.1038/s41586-022-05053-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Hoffmann M., Kleine-Weber H., Schroeder S., Krüger N., Herrler T., Erichsen S., Schiergens T.S., Herrler G., Wu N.H., Nitsche A., Müller M.A., Drosten C., Pöhlmann S. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell. 2020;181(2):271–280.e8. doi: 10.1016/j.cell.2020.02.052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Wrapp D., Wang N., Corbett K.S., Goldsmith J.A., Hsieh C.L., Abiona O., Graham B.S., McLellan J.S. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science. 2020;367(6483):1260–1263. doi: 10.1126/science.abb2507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Cerutti G., Guo Y., Zhou T., Gorman J., Lee M., Rapp M., Reddem E.R., Yu J., Bahna F., Bimela J., Huang Y., Katsamba P.S., Liu L., Nair M.S., Rawi R., Olia A.S., Wang P., Zhang B., Chuang G.Y., Ho D.D., Sheng Z., Kwong P.D., Shapiro L. Potent SARS-CoV-2 neutralizing antibodies directed against spike N-terminal domain target a single supersite. Cell Host Microbe. 2021;29(5):819–833. doi: 10.1016/j.chom.2021.03.005. e7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Dejnirattisai W., Zhou D., Ginn H.M., Duyvesteyn H.M.E., Supasa P., Case J.B., Zhao Y., Walter T.S., Mentzer A.J., Liu C., Wang B., Paesen G.C., Slon-Campos J., López-Camacho C., Kafai N.M., Bailey A.L., Chen R.E., Ying B., Thompson C., Bolton J., Fyfe A., Gupta S., Tan T.K., Gilbert-Jaramillo J., James W., Knight M., Carroll M.W., Skelly D., Dold C., Peng Y., Levin R., Dong T., Pollard A.J., Knight J.C., Klenerman P., Temperton N., Hall D.R., Williams M.A., Paterson N.G., Bertram F.K.R., Siebert C.A., Clare D.K., Howe A., Radecke J., Song Y., Townsend A.R., Huang K.A., Fry E.E., Mongkolsapaya J., Diamond M.S., Ren J., Stuart D.I., Screaton G.R. The antigenic anatomy of SARS-CoV-2 receptor binding domain. Cell. 2021;184(8):2183–2200. doi: 10.1016/j.cell.2021.02.032. e22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.McCallum M., De Marco A., Lempp F.A., Tortorici M.A., Pinto D., Walls A.C., Beltramello M., Chen A., Liu Z., Zatta F., Zepeda S., di Iulio J., Bowen J.E., Montiel-Ruiz M., Zhou J., Rosen L.E., Bianchi S., Guarino B., Fregni C.S., Abdelnabi R., Foo S.C., Rothlauf P.W., Bloyet L.M., Benigni F., Cameroni E., Neyts J., Riva A., Snell G., Telenti A., Whelan S.P.J., Virgin H.W., Corti D., Pizzuto M.S., Veesler D. N-terminal domain antigenic mapping reveals a site of vulnerability for SARS-CoV-2. Cell. 2021;184(9):2332–2347. doi: 10.1016/j.cell.2021.03.028. e16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Cao Y., Yisimayi A., Jian F., Song W., Xiao T., Wang L., Du S., Wang J., Li Q., Chen X., Yu Y., Wang P., Zhang Z., Liu P., An R., Hao X., Wang Y., Wang J., Feng R., Sun H., Zhao L., Zhang W., Zhao D., Zheng J., Yu L., Li C., Zhang N., Wang R., Niu X., Yang S., Song X., Chai Y., Hu Y., Shi Y., Zheng L., Li Z., Gu Q., Shao F., Huang W., Jin R., Shen Z., Wang Y., Wang X., Xiao J., Xie X.S. BA.2.12.1, BA.4 and BA.5 escape antibodies elicited by Omicron infection. Nature. 2022;608(7923):593–602. doi: 10.1038/s41586-022-04980-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Zhou T., Wang L., Misasi J., Pegu A., Zhang Y., Harris D.R., Olia A.S., Talana C.A., Yang E.S., Chen M., Choe M., Shi W., Teng I.T., Creanga A., Jenkins C., Leung K., Liu T., Stancofski E.D., Stephens T., Zhang B., Tsybovsky Y., Graham B.S., Mascola J.R., Sullivan N.J., Kwong P.D. Structural basis for potent antibody neutralization of SARS-CoV-2 variants including B.1.1.529. Science. 2022;376(6591) doi: 10.1126/science.abn8897. eabn8897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ai J., Wang X., He X., Zhao X., Zhang Y., Jiang Y., Li M., Cui Y., Chen Y., Qiao R., Li L., Yang L., Li Y., Hu Z., Zhang W., Wang P. Antibody evasion of SARS-CoV-2 Omicron BA.1, BA.1.1, BA.2, and BA.3 sub-lineages. Cell Host Microbe. 2022;30(8):1077–1083. doi: 10.1016/j.chom.2022.05.001. e4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Hachmann N.P., Miller J., Collier A.Y., Ventura J.D., Yu J., Rowe M., Bondzie E.A., Powers O., Surve N., Hall K., Barouch D.H. Neutralization escape by SARS-CoV-2 Omicron Subvariants BA.2.12.1, BA.4, and BA.5. New Engl. J. Med. 2022;387(1):86–88. doi: 10.1056/NEJMc2206576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Tan C.W., Lim B.L., Young B.E., Yeoh A.Y., Yung C.F., Yap W.C., Althaus T., Chia W.N., Zhu F., Lye D.C., Wang L.F. Comparative neutralisation profile of SARS-CoV-2 omicron subvariants BA.2.75 and BA.5. Lancet Microbe. 2022;3(12) doi: 10.1016/S2666-5247(22)00220-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Tuekprakhon A., Nutalai R., Dijokaite-Guraliuc A., Zhou D., Ginn H.M., Selvaraj M., Liu C., Mentzer A.J., Supasa P., Duyvesteyn H.M.E., Das R., Skelly D., Ritter T.G., Amini A., Bibi S., Adele S., Johnson S.A., Constantinides B., Webster H., Temperton N., Klenerman P., Barnes E., Dunachie S.J., Crook D., Pollard A.J., Lambe T., Goulder P., Paterson N.G., Williams M.A., Hall D.R., Fry E.E., Huo J., Mongkolsapaya J., Ren J., Stuart D.I., Screaton G.R. Antibody escape of SARS-CoV-2 Omicron BA.4 and BA.5 from vaccine and BA.1 serum. Cell. 2022;185(14):2422–2433. doi: 10.1016/j.cell.2022.06.005. e13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Hachmann N.P., Miller J., Collier A.Y., Barouch D.H. Neutralization escape by SARS-CoV-2 Omicron subvariant BA.4.6. N. Engl. J. Med. 2022;387(20):1904–1906. doi: 10.1056/NEJMc2212117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Jian F., Yu Y., Song W., Yisimayi A., Yu L., Gao Y., Zhang N., Wang Y., Shao F., Hao X., Xu Y., Jin R., Wang Y., Xie X.S., Cao Y. Further humoral immunity evasion of emerging SARS-CoV-2 BA.4 and BA.5 subvariants. Lancet Infect. Dis. 2022;22(11):1535–1537. doi: 10.1016/S1473-3099(22)00642-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Arora P., Kempf A., Nehlmeier I., Schulz S.R., Jäck H.M., Pöhlmann S., Hoffmann M. Omicron sublineage BQ.1.1 resistance to monoclonal antibodies. Lancet Infect. Dis. 2023;23(1):22–23. doi: 10.1016/S1473-3099(22)00733-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Mannar D., Saville J.W., Zhu X., Srivastava S.S., Berezuk A.M., Tuttle K.S., Marquez A.C., Sekirov I., Subramaniam S. SARS-CoV-2 Omicron variant: Antibody evasion and cryo-EM structure of spike protein-ACE2 complex. Science. 2022;375(6582):760–764. doi: 10.1126/science.abn7760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Focosi D., McConnell S., Casadevall A., Cappello E., Valdiserra G., Tuccori M. Monoclonal antibody therapies against SARS-CoV-2. Lancet Infect. Dis. 2022;22(11):e311–e326. doi: 10.1016/S1473-3099(22)00311-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Cui Z., Liu P., Wang N., Wang L., Fan K., Zhu Q., Wang K., Chen R., Feng R., Jia Z., Yang M., Xu G., Zhu B., Fu W., Chu T., Feng L., Wang Y., Pei X., Yang P., Xie X.S., Cao L., Cao Y., Wang X. Structural and functional characterizations of infectivity and immune evasion of SARS-CoV-2 Omicron. Cell. 2022;185(5):860–871. doi: 10.1016/j.cell.2022.01.019. e13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Cameroni E., Bowen J.E., Rosen L.E., Saliba C., Zepeda S.K., Culap K., Pinto D., VanBlargan L.A., De Marco A., di Iulio J., Zatta F., Kaiser H., Noack J., Farhat N., Czudnochowski N., Havenar-Daughton C., Sprouse K.R., Dillen J.R., Powell A.E., Chen A., Maher C., Yin L., Sun D., Soriaga L., Bassi J., Silacci-Fregni C., Gustafsson C., Franko N.M., Logue J., Iqbal N.T., Mazzitelli I., Geffner J., Grifantini R., Chu H., Gori A., Riva A., Giannini O., Ceschi A., Ferrari P., Cippà P.E., Franzetti-Pellanda A., Garzoni C., Halfmann P.J., Kawaoka Y., Hebner C., Purcell L.A., Piccoli L., Pizzuto M.S., Walls A.C., Diamond M.S., Telenti A., Virgin H.W., Lanzavecchia A., Snell G., Veesler D., Corti D. Broadly neutralizing antibodies overcome SARS-CoV-2 Omicron antigenic shift. Nature. 2022;602(7898):664–670. doi: 10.1038/s41586-021-04386-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Li L., Liao H., Meng Y., Li W., Han P., Liu K., Wang Q., Li D., Zhang Y., Wang L., Fan Z., Zhang Y., Wang Q., Zhao X., Sun Y., Huang N., Qi J., Gao G.F. Structural basis of human ACE2 higher binding affinity to currently circulating Omicron SARS-CoV-2 sub-variants BA.2 and BA.1.1. Cell. 2022;185(16):2952–2960. doi: 10.1016/j.cell.2022.06.023. e10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.McCallum M., Czudnochowski N., Rosen L.E., Zepeda S.K., Bowen J.E., Walls A.C., Hauser K., Joshi A., Stewart C., Dillen J.R., Powell A.E., Croll T.I., Nix J., Virgin H.W., Corti D., Snell G., Veesler D. Structural basis of SARS-CoV-2 Omicron immune evasion and receptor engagement. Science. 2022;375(6583):864–868. doi: 10.1126/science.abn8652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.McCallum M., Walls A.C., Sprouse K.R., Bowen J.E., Rosen L.E., Dang H.V., De Marco A., Franko N., Tilles S.W., Logue J., Miranda M.C., Ahlrichs M., Carter L., Snell G., Pizzuto M.S., Chu H.Y., Van Voorhis W.C., Corti D., Veesler D. Molecular basis of immune evasion by the Delta and Kappa SARS-CoV-2 variants. Science. 2021;374(6575):1621–1626. doi: 10.1126/science.abl8506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Collier D.A., De Marco A., Ferreira I., Meng B., Datir R.P., Walls A.C., Kemp S.A., Bassi J., Pinto D., Silacci-Fregni C., Bianchi S., Tortorici M.A., Bowen J., Culap K., Jaconi S., Cameroni E., Snell G., Pizzuto M.S., Pellanda A.F., Garzoni C., Riva A., Elmer A., Kingston N., Graves B., McCoy L.E., Smith K.G.C., Bradley J.R., Temperton N., Ceron-Gutierrez L., Barcenas-Morales G., Harvey W., Virgin H.W., Lanzavecchia A., Piccoli L., Doffinger R., Wills M., Veesler D., Corti D., Gupta R.K. Sensitivity of SARS-CoV-2 B.1.1.7 to mRNA vaccine-elicited antibodies. Nature. 2021;593(7857):136–141. doi: 10.1038/s41586-021-03412-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Yuan M., Huang D., Lee C.D., Wu N.C., Jackson A.M., Zhu X., Liu H., Peng L., van Gils M.J., Sanders R.W., Burton D.R., Reincke S.M., Prüss H., Kreye J., Nemazee D., Ward A.B., Wilson I.A. Structural and functional ramifications of antigenic drift in recent SARS-CoV-2 variants. Science. 2021;373(6556):818–823. doi: 10.1126/science.abh1139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Thomson E.C., Rosen L.E., Shepherd J.G., Spreafico R., da Silva Filipe A., Wojcechowskyj J.A., Davis C., Piccoli L., Pascall D.J., Dillen J., Lytras S., Czudnochowski N., Shah R., Meury M., Jesudason N., De Marco A., Li K., Bassi J., O'Toole A., Pinto D., Colquhoun R.M., Culap K., Jackson B., Zatta F., Rambaut A., Jaconi S., Sreenu V.B., Nix J., Zhang I., Jarrett R.F., Glass W.G., Beltramello M., Nomikou K., Pizzuto M., Tong L., Cameroni E., Croll T.I., Johnson N., Di Iulio J., Wickenhagen A., Ceschi A., Harbison A.M., Mair D., Ferrari P., Smollett K., Sallusto F., Carmichael S., Garzoni C., Nichols J., Galli M., Hughes J., Riva A., Ho A., Schiuma M., Semple M.G., Openshaw P.J.M., Fadda E., Baillie J.K., Chodera J.D., Rihn S.J., Lycett S.J., Virgin H.W., Telenti A., Corti D., Robertson D.L., Snell G. Circulating SARS-CoV-2 spike N439K variants maintain fitness while evading antibody-mediated immunity. Cell. 2021;184(5):1171–1187. doi: 10.1016/j.cell.2021.01.037. e20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Zhu X., Mannar D., Srivastava S.S., Berezuk A.M., Demers J.P., Saville J.W., Leopold K., Li W., Dimitrov D.S., Tuttle K.S., Zhou S., Chittori S., Subramaniam S. Cryo-electron microscopy structures of the N501Y SARS-CoV-2 spike protein in complex with ACE2 and 2 potent neutralizing antibodies. PLoS Biol. 2021;19(4) doi: 10.1371/journal.pbio.3001237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Starr T.N., Greaney A.J., Hilton S.K., Ellis D., Crawford K.H.D., Dingens A.S., Navarro M.J., Bowen J.E., Tortorici M.A., Walls A.C., King N.P., Veesler D., Bloom J.D. Deep mutational scanning of SARS-CoV-2 receptor binding domain reveals constraints on folding and ACE2 Binding. Cell. 2020;182(5):1295–1310. doi: 10.1016/j.cell.2020.08.012. e20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Dejnirattisai W., Huo J., Zhou D., Zahradník J., Supasa P., Liu C., Duyvesteyn H.M.E., Ginn H.M., Mentzer A.J., Tuekprakhon A., Nutalai R., Wang B., Dijokaite A., Khan S., Avinoam O., Bahar M., Skelly D., Adele S., Johnson S.A., Amini A., Ritter T.G., Mason C., Dold C., Pan D., Assadi S., Bellass A., Omo-Dare N., Koeckerling D., Flaxman A., Jenkin D., Aley P.K., Voysey M., Costa Clemens S.A., Naveca F.G., Nascimento V., Nascimento F., Fernandes da Costa C., Resende P.C., Pauvolid-Correa A., Siqueira M.M., Baillie V., Serafin N., Kwatra G., Da Silva K., Madhi S.A., Nunes M.C., Malik T., Openshaw P.J.M., Baillie J.K., Semple M.G., Townsend A.R., Huang K.A., Tan T.K., Carroll M.W., Klenerman P., Barnes E., Dunachie S.J., Constantinides B., Webster H., Crook D., Pollard A.J., Lambe T., Paterson N.G., Williams M.A., Hall D.R., Fry E.E., Mongkolsapaya J., Ren J., Schreiber G., Stuart D.I., Screaton G.R. SARS-CoV-2 Omicron-B.1.1.529 leads to widespread escape from neutralizing antibody responses. Cell. 2022;185(3):467–484. doi: 10.1016/j.cell.2021.12.046. e15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Yu J., Collier A.Y., Rowe M., Mardas F., Ventura J.D., Wan H., Miller J., Powers O., Chung B., Siamatu M., Hachmann N.P., Surve N., Nampanya F., Chandrashekar A., Barouch D.H. Neutralization of the SARS-CoV-2 Omicron BA.1 and BA.2 Variants. N. Engl. J. Med. 2022;386(16):1579–1580. doi: 10.1056/NEJMc2201849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Liu L., Iketani S., Guo Y., Chan J.F., Wang M., Liu L., Luo Y., Chu H., Huang Y., Nair M.S., Yu J., Chik K.K., Yuen T.T., Yoon C., To K.K., Chen H., Yin M.T., Sobieszczyk M.E., Huang Y., Wang H.H., Sheng Z., Yuen K.Y., Ho D.D. Striking antibody evasion manifested by the Omicron variant of SARS-CoV-2. Nature. 2022;602(7898):676–681. doi: 10.1038/s41586-021-04388-0. [DOI] [PubMed] [Google Scholar]
- 64.Khan K., Karim F., Ganga Y., Bernstein M., Jule Z., Reedoy K., Cele S., Lustig G., Amoako D., Wolter N., Samsunder N., Sivro A., San J.E., Giandhari J., Tegally H., Pillay S., Naidoo Y., Mazibuko M., Miya Y., Ngcobo N., Manickchund N., Magula N., Karim Q.A., von Gottberg A., Abdool Karim S.S., Hanekom W., Gosnell B.I., Lessells R.J., de Oliveira T., Moosa M.S., Sigal A. Omicron BA.4/BA.5 escape neutralizing immunity elicited by BA.1 infection. Nat. Commun. 2022;13(1):4686. doi: 10.1038/s41467-022-32396-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Cao Y., Song W., Wang L., Liu P., Yue C., Jian F., Yu Y., Yisimayi A., Wang P., Wang Y., Zhu Q., Deng J., Fu W., Yu L., Zhang N., Wang J., Xiao T., An R., Wang J., Liu L., Yang S., Niu X., Gu Q., Shao F., Hao X., Meng B., Gupta R.K., Jin R., Wang Y., Xie X.S., Wang X. Characterization of the enhanced infectivity and antibody evasion of Omicron BA.2.75. Cell Host Microbe. 2022;30(11):1527–1539.e5. doi: 10.1016/j.chom.2022.09.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Wang X., Zhao X., Song J., Wu J., Zhu Y., Li M., Cui Y., Chen Y., Yang L., Liu J., Zhu H., Jiang S., Wang P. Homologous or heterologous booster of inactivated vaccine reduces SARS-CoV-2 Omicron variant escape from neutralizing antibodies. Emerg. Microbes Infect. 2022;11(1):477–481. doi: 10.1080/22221751.2022.2030200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Shen X., Chalkias S., Feng J., Chen X., Zhou H., Marshall J.C., Girard B., Tomassini J.E., Aunins A., Das R., Montefiori D.C. Neutralization of SARS-CoV-2 Omicron BA.2.75 after mRNA-1273 Vaccination. N. Engl. J. Med. 2022;387(13):1234–1236. doi: 10.1056/NEJMc2210648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Pérez-Then E., Lucas C., Monteiro V.S., Miric M., Brache V., Cochon L., Vogels C.B.F., Malik A.A., De la Cruz E., Jorge A., De Los Santos M., Leon P., Breban M.I., Billig K., Yildirim I., Pearson C., Downing R., Gagnon E., Muyombwe A., Razeq J., Campbell M., Ko A.I., Omer S.B., Grubaugh N.D., Vermund S.H., Iwasaki A. Neutralizing antibodies against the SARS-CoV-2 Delta and Omicron variants following heterologous CoronaVac plus BNT162b2 booster vaccination. Nat. Med. 2022;28(3):481–485. doi: 10.1038/s41591-022-01705-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Muik A., Lui B.G., Bacher M., Wallisch A.K., Toker A., Finlayson A., Krüger K., Ozhelvaci O., Grikscheit K., Hoehl S., Ciesek S., Türeci Ö., Sahin U. Omicron BA.2 breakthrough infection enhances cross-neutralization of BA.2.12.1 and BA.4/BA.5. Sci. Immunol. 2022;7(77) doi: 10.1126/sciimmunol.ade2283. eade2283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Sheward D.J., Kim C., Fischbach J., Sato K., Muschiol S., Ehling R.A., Björkström N.K., Hedestam G.B.Karlsson, Reddy S.T., Albert J., Peacock T.P., Murrell B. Omicron sublineage BA.2.75.2 exhibits extensive escape from neutralising antibodies. Lancet Infect. Dis. 2022;22(11):1538–1540. doi: 10.1016/S1473-3099(22)00663-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Zou J., Kurhade C., Patel S., Kitchin N., Tompkins K., Cutler M., Cooper D., Yang Q., Cai H., Muik A., Zhang Y., Lee D.Y., Şahin U., Anderson A.S., Gruber W.C., Xie X., Swanson K.A., Shi P.Y. Neutralization of BA.4-BA.5, BA.4.6, BA.2.75.2, BQ.1.1, and XBB.1 with Bivalent Vaccine. N. Engl. J. Med. 2023;388(9):854–857. doi: 10.1056/NEJMc2214916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Davis-Gardner M.E., Lai L., Wali B., Samaha H., Solis D., Lee M., Porter-Morrison A., Hentenaar I.T., Yamamoto F., Godbole S., Liu Y., Douek D.C., Lee F.E., Rouphael N., Moreno A., Pinsky B.A., Suthar M.S. Neutralization against BA.2.75.2, BQ.1.1, and XBB from mRNA Bivalent Booster. N. Engl. J. Med. 2023;388(2):183–185. doi: 10.1056/NEJMc2214293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Collier A.Y., Miller J., Hachmann N.P., McMahan K., Liu J., Bondzie E.A., Gallup L., Rowe M., Schonberg E., Thai S., Barrett J., Borducchi E.N., Bouffard E., Jacob-Dolan C., Mazurek C.R., Mutoni A., Powers O., Sciacca M., Surve N., VanWyk H., Wu C., Barouch D.H. Immunogenicity of BA.5 Bivalent mRNA Vaccine Boosters. New Engl. J. Med. 2023;388(6):565–567. doi: 10.1056/NEJMc2213948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Bowen J.E., Addetia A., Dang H.V., Stewart C., Brown J.T., Sharkey W.K., Sprouse K.R., Walls A.C., Mazzitelli I.G., Logue J.K., Franko N.M., Czudnochowski N., Powell A.E., Dellota E., Jr., Ahmed K., Ansari A.S., Cameroni E., Gori A., Bandera A., Posavad C.M., Dan J.M., Zhang Z., Weiskopf D., Sette A., Crotty S., Iqbal N.T., Corti D., Geffner J., Snell G., Grifantini R., Chu H.Y., Veesler D. Omicron spike function and neutralizing activity elicited by a comprehensive panel of vaccines. Science. 2022;377(6608):890–894. doi: 10.1126/science.abq0203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Scheaffer S.M., Lee D., Whitener B., Ying B., Wu K., Liang C.Y., Jani H., Martin P., Amato N.J., Avena L.E., Berrueta D.M., Schmidt S.D., O'Dell S., Nasir A., Chuang G.Y., Stewart-Jones G., Koup R.A., Doria-Rose N.A., Carfi A., Elbashir S.M., Thackray L.B., Edwards D.K., Diamond M.S. Bivalent SARS-CoV-2 mRNA vaccines increase breadth of neutralization and protect against the BA.5 Omicron variant in mice. Nat. Med. 2023;29(1):247–257. doi: 10.1038/s41591-022-02092-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Chalkias S., Eder F., Essink B., Khetan S., Nestorova B., Feng J., Chen X., Chang Y., Zhou H., Montefiori D., Edwards D.K., Girard B., Pajon R., Dutko F.J., Leav B., Walsh S.R., Baden L.R., Miller J.M., Das R. Safety, immunogenicity and antibody persistence of a bivalent Beta-containing booster vaccine against COVID-19: a phase 2/3 trial. Nat. Med. 2022;28(11):2388–2397. doi: 10.1038/s41591-022-02031-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Qu P., Evans J.P., Zheng Y.M., Carlin C., Saif L.J., Oltz E.M., Xu K., Gumina R.J., Liu S.L. Evasion of neutralizing antibody responses by the SARS-CoV-2 BA.2.75 variant. Cell Host Microbe. 2022;30(11):1518–1526. doi: 10.1016/j.chom.2022.09.015. e4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Wang Q., Iketani S., Li Z., Guo Y., Yeh A.Y., Liu M., Yu J., Sheng Z., Huang Y., Liu L., Ho D.D. Antigenic characterization of the SARS-CoV-2 Omicron subvariant BA.2.75. Cell Host Microbe. 2022;30(11):1512–1517. doi: 10.1016/j.chom.2022.09.002. e4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Takashita E., Yamayoshi S., Halfmann P., Wilson N., Ries H., Richardson A., Bobholz M., Vuyk W., Maddox R., Baker D.A., Friedrich T.C., O'Connor D.H., Uraki R., Ito M., Sakai-Tagawa Y., Adachi E., Saito M., Koga M., Tsutsumi T., Iwatsuki-Horimoto K., Kiso M., Yotsuyanagi H., Watanabe S., Hasegawa H., Imai M., Kawaoka Y. In vitro efficacy of antiviral agents against omicron subvariant BA.4.6. New Engl. J. Med. 2022;387(22):2094–2097. doi: 10.1056/NEJMc2211845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Wang Q., Li Z., Ho J., Guo Y., Yeh A.Y., Mohri H., Liu M., Wang M., Yu J., Shah J.G., Chang J.Y., Herbas F., Yin M.T., Sobieszczyk M.E., Sheng Z., Liu L., Ho D.D. Resistance of SARS-CoV-2 omicron subvariant BA.4.6 to antibody neutralisation. Lancet Infect. Dis. 2022;22(12):1666–1668. doi: 10.1016/S1473-3099(22)00694-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Gruell H., Vanshylla K., Tober-Lau P., Hillus D., Sander L.E., Kurth F., Klein F. Neutralisation sensitivity of the SARS-CoV-2 omicron BA.2.75 sublineage. Lancet Infect. Dis. 2022;22(10):1422–1423. doi: 10.1016/S1473-3099(22)00580-1. [DOI] [PMC free article] [PubMed] [Google Scholar]