Abstract
Viral co-infection has been found in animals; however, the mechanisms of co-infection are unclear. The abundance and diversity of viruses in water make fish highly susceptible to co-infection. Here, we reported a co-infection in fish, which resulted in reduced host lethality and illustrated the intracellular molecular mechanism of viral co-infection. The spring viremia of carp virus (SVCV) is a highly lethal virus that infects Cyprinidae, such as zebrafish. The mortality of SVCV infection was significantly reduced when co-infected with the grass carp reovirus (GCRV). The severity of tissue damage and viral proliferation of SVCV was also reduced in co-infection with GCRV. The transcriptome bioinformatics analysis demonstrated that the effect on the host transcripts in response to SVCV infection was significantly reduced in co-infection. After excluding the extracellular interactions of these two viruses, the intracellular mechanisms were studied. We found that the GCRV NS38 remarkably decreased SVCV infection and viral proliferation. The interaction between GCRV NS38 and SVCV nucleoprotein (N) and phosphoprotein (P) proteins was identified, and NS38 downregulated both N and P proteins. Further analysis demonstrated that the N protein was degraded by NS38 indispensable of the autophagy receptor, sequestosome 1 (p62). Meanwhile, K63-linked ubiquitination of the P protein was reduced by NS38, leading to ubiquitinated degradation of the P protein. These results reveal that the intracellular viral protein interactions are a crucial mechanism of co-infection and influence the host pathology and expand our understanding in intracellular viral interactions co-infection.
Keywords: Co-infection, Spring viremia of carp virus (SVCV), Grass carp reovirus (GCRV), Antagonize, Autophagy, Ubiquitination
Highlights
-
•
The GCRV markedly repressed the influence of SVCV infection on fish.
-
•
The transcriptome changes caused by SVCV were substantially defused by GCRV.
-
•
GCRV NS38 interacted with and degraded SVCV N and P proteins.
-
•
The N protein was degraded through autophagy by NS38.
-
•
The P protein was degraded by NS38 via decreasing K63-linked ubiquitination.
1. Introduction
Pathogen co-infection is receiving increasing attention. Co-infection refers to the concurrent infection of one cell or host by at least two pathogens. In humans, co-infection has been identified in several cases. Compared to single infection, the mechanisms and pathologies of co-infection are more complicated and intricated (Griffiths et al., 2011). Normally, bacteria and viruses are the two major pathogens that infect hosts in a random combination to form co-infection. Bacteria-bacteria co-infection are usually observed in pneumonia, and treatment with antibiotics is an effective tool (Hoshino et al., 2022). A virus can co-infect by either virus-bacteria or virus-virus infection and are difficult to treat. Clinical studies of the novel coronavirus disease 2019 (COVID-19) caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and the co-infection of the SARS-CoV-2 with other microorganisms showed how co-infection complicated the accurate diagnosis, treatment, and even enhanced the mortality rates in patients (McCullers, 2014; Hoque et al., 2021). In these patients, an association has been made between disease severity and the detection of co-infection with bacterial pathogens, such as Acinetobacter baumannii, Klebsiella pneumoniae, and Mycoplasma pneumoniae (Langford et al., 2020). Due to the susceptibility to infections of the respiratory tract, co-infection is understandable in respiratory diseases. In another recent study of COVID-19, SARS-CoV-2 infections have been found to be associated with 16 different viral family co-infection, such as tombusvirus, victorivirus, and partitivirus (Hoque et al., 2021). Virus-virus co-infection has been identified in some viral diseases. The mode of spread in the human immunodeficiency virus (HIV), hepatitis B virus (HBV), and hepatitis C virus (HCV) overlap and include sexual transmission and vertical transmission, which lead to the high probability of co-infection. It has been reported that the prevalence of triple co-infection of HIV, HBV, and HCV is 3.3%. In addition, up to 11% of individuals with HIV have a Mycobacterium tuberculosis co-infection (Zhang et al., 2014; Arora et al., 2021). Viral co-infection is found in other animal hosts. In an examination of 47 diseased chickens, 30 were detected to have infection with at least two viruses, including marek's disease virus (MDV), avian leukosis virus (ALV), and reticuloendotheliosis virus (REV) (Nishitha et al., 2021). In plants, viral co-infection has been widely reported. The environments of cropping in greenhouses and the prevalence of whitefly infestations lead to the co-infection of tomato yellow leaf curl virus (TYLCV) and tomato chlorosis virus (ToCV) in tomatoes (Gul-Seker and Elibuyuk, 2019). In addition, the different genotypes of viruses could also cause co-infection. The serotypes O and A of foot-and-mouth disease virus (FMDV) co-infect cattle and co-infection with porcine of circovirus 4 (PCV4) and PCV2 (Arzt et al., 2021; Wang et al., 2022). These co-infections are accompanied by aggravated infection status and increased mortality (Xu et al., 2022).
Generally, two main results in co-infection are particularly concerning, disease aggravation and remission. For aggravation, influenza A virus (IAV) co-infection remarkably enhances the infectivity of SARS-CoV-2, increasing SARS-CoV-2 viral proliferation and resulting in severe lung damage. Such disease aggravation is not observed with some other respiratory viruses (Bai et al., 2021). However, in the common cold, which is caused by the rhinovirus (RV), co-infection with IAV and RV have been shown to result in a negative correlation with the reduction of the severity and viral proliferation. During times of increased influenza cases, the prevalence of RV is significantly reduced (Nickbakhsh et al., 2019). In these cases, the explanation for the differences in co-infection is that one virus inhibits the host production of interferon (IFN), causing aggravation, or the cell surface receptors are competing for binding, leading to the remission of one virus. To date, though the phenomenon and results of viral co-infection have been reported, the mechanisms are still unclear.
Numerous viruses can infect fish and are contained in water. The mobility of water and fish increases the viral transmission and the possibility of viral co-infection (Kotob et al., 2016; Wolf et al., 2020). Synergistic or antagonistic viral co-infection has been reported in fish. For example, scale drop disease virus (SDDV) and lates calcarifer herpes virus (LCHV) co-infection downregulates barramundi's immune response. In contrast, experiments have demonstrated that infectious pancreatic necrosis virus (IPNV) inhibited infectious hematopoietic necrosis virus (IHNV) proliferation in vitro (Xu et al., 2020; Domingos et al., 2021). Therefore, in fish viral diseases, co-infection is more common; however, the cellular mechanisms of viral co-infection are still undiscovered.
Here, we reveal a unique antagonistic interaction in a fish viral co-infection. The severe symptoms and high viral titers caused by spring viremia of carp virus (SVCV) infections are significantly reduced when co-infected with the grass carp reovirus (GCRV). The GCRV NS38 protein degrades the SVCV nucleoprotein (N) by autophagy-lysosome manner and the SVCV phosphoprotein (P) by ubiquitin (Ub)-proteasome pathway. These intracellular degradations are pivotal to realize that one virus's proliferation suppresses the other virus in co-infection. These results suggest that viral protein regulations potentially cause complex pathologies in viral co-infection.
2. Materials and methods
2.1. Cells, viruses, and fish
Zebrafish liver (ZFL) cells (American Type Culture Collection, ATCC) were cultured at 28 °C in 5% CO2 in Ham's F12 nutrient mixture medium (Invitrogen, Grand Island, U.S.A) supplemented with 10% fetal bovine serum (FBS) (Gibco, Grand Island, U.S.A). Epithelioma papulosum cyprini (EPC) cells and ctenopharyngodon idellus kidney (CIK) cells were obtained from China Center for Type Culture Collection (CCTCC) and were maintained at 28 °C in 5% CO2 in medium 199 (Invitrogen) supplemented with 10% FBS. SVCV (strain OMG067), a negative ssRNA virus, was propagated in EPC cells until a cellular cytopathic effect (CPE) was observed. Then the harvested cell culture fluid containing SVCV was centrifuged at 4000 ×g for 20 min to remove the cell debris, and the supernatant was stored at −80 °C until used. GCRV (strain 873, group Ⅰ) was provided by Prof. Wuhan Xiao (Institute of Hydrobiology, Chinese Academy of Sciences). GCRV was propagated in CIK cells and harvested in a similar way to SVCV. The carassius auratus herpesvirus (CaHV) was bred in wild type crucian carp and obtained by isolation from infected tissues in our lab. The GCRV-II (strain 106, group Ⅱ) was obtained by isolation from infected grass carp in our lab.
Wild-type zebrafish (four months of age, both sexes, weighing 0.7 ± 0.2 g, body lengths 1–2 cm) were obtained from China Zebrafish Resource Center (CZRC) and kept in recirculating water at 28 °C. All zebrafish were maintained according to institutional and national ethical and animal welfare guidelines in the laboratory for at least two weeks prior to the experiments for acclimatization and evaluation of overall fish health. Only the healthy fish, as determined by the general appearance and the level of activity, were used for studies. For zebrafish survival experiments, wild-type zebrafish were intraperitoneally (i.p.) injected with 10 μL mixture of SVCV suspension [∼ 5 × 109 50% tissue culture infective dose (TCID50)/mL, 5 μL/individual] plus PBS (5 μL/individual) or GCRV (∼ 5 × 107 TCID50/mL, 5 μL/individual). As regards CaHV, zebrafish were i.p. injected with 10 μL mixture of CaHV suspension (5 μL/individual) plus PBS (5 μL/individual). The i.p. injection of PBS was used as mock infection. The survival rates of fish were counted daily.
2.2. Plasmid construction and reagents
The sequence of S9 (GenBank accession number: AF403395.1) that encodes NS38 was obtained from the NCBI (National Center for Biotechnology Information) website (http://www.ncbi.nlm.nih.gov/). Using the cDNA of the cell lysate from GCRV infected CIK cells as template, the open reading frame (ORF) of NS38 was amplified by polymerase chain reaction (PCR) and cloned into pCMV-Tag2C, pCMV-Myc, and pCMV-HA vectors (Clontech, Mountain View CA, U.S.A), respectively. The ORFs of calcium binding and coiled-coil domain 2 (NDP52, ZDB-GENE-050522-346), optineurin (OPTN, ZDB-GENE-030131-8103), sequestosome 1 (p62, ZDB-GENE-040426-2204), neighbour of breast cancer 1 gene (NBR1) autophagy cargo receptor a (NBR1a, ZDB-GENE-030131-9112), and toll interacting protein (Tollip, ZDB-GENE-030131-8820) were obtained from the ZFIN website (https://zfin.org/) and also subcloned into pCMV-Myc, pCMV-HA, and pCMV-Tag2C vectors, respectively. For subcellular localization, the NS38 was inserted into pEGFP-N3 (Clontech) or pCS2-mCherry vectors (BD Clontech, Mountain View CA, U.S.A). The ORFs of N protein (DQ097384.2), P protein (DQ097384.2), and G protein (KT321307.1) were also inserted into subcloned into pCMV-Myc, pCMV-HA, pCMV-Tag2C, pEGFP-N3, and pDsRed-N1 vectors (Clontech). Ub-K63O-HA and Ub-K48O-HA were expression plasmids for HA-tagged Lys-63 or Lys-48-only Ub mutants (all lysine residues except Lys-63/48 are mutated). All constructs were confirmed by DNA sequencing. The primers including the restriction enzyme cutting sites used for plasmid construction are listed in Supplementary Table S1. MG132 (Cat. No. M7449), 3-Methyladenine (3-MA, Cat. No. M9281), chloroquine (CQ, Cat. No. C6628) were obtained from Sigma-Aldrich (German). BafilomycinA1 (Baf-A1, Cat. No. S1413) was obtained from Selleck (Houston, Texas, U.S.A).
2.3. Transcriptomic analysis
Total RNA was extracted using the TRIzol reagent according to the manufacturer's protocol. RNA purity and quantification were evaluated using the NanoDrop 2000 spectrophotometer (Thermo Scientific, Waltham, U.S.A). RNA integrity was assessed using the Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, U.S.A). The transcriptome sequencing and analysis were conducted by OE Biotech Co., Ltd. (Shanghai). The raw sequencing data was submitted to the NGDC (National Genomics Data Center) (GSA accession number: CRA008409).
2.4. Histopathology
The heart, liver, and spleen tissues of zebrafish from mock group, SVCV-infected group, and SVCV&GCRV co-infected group at 72 h post infection (hpi) were dissected, and fixed in 10% phosphate-buffered formalin overnight. Then the samples were dehydrated in ascending grades of alcohol and embedded into paraffin. Sections at 5 μm thickness were taken and stained with hematoxylin and eosin (H&E). Histological changes were examined by optical microscopy at ×40 magnification and were analyzed using the Aperio ImageScope software (Leica, German).
2.5. Transient transfection and virus infection
Transient transfections were performed in EPC cells seeded in 6-well or 24-well plates by using FishTrans (MeiSenTe Biotechnology, Guangdong, China). For the antiviral assay using 24-well plates, EPC cells were transfected with 0.25 μg NS38-Myc or the empty vector. Antiviral assays were implemented as described (Li et al., 2019; Lu et al., 2021).
2.6. RNA interference (RNAi)
The short hairpin RNA (shRNA) of S9 and p62 were designed by BLOCK-iT™ RNAi Designer, and cloned into the pLKO.1-TRC cloning vector. The oligos targeted for S9 and p62 were listed in Supplementary Table S1. For RNAi of S9, EPC cells were seeded in 6-well plates overnight and transfected with 2 μg shS9-pLKO.1 or the negative control (sh-NC) for 6 h, and then GCRV (MOI = 0.01) was added. After 24 h, cells were harvested for RNAi effectiveness detection.
2.7. RNA extraction, reverse transcription, and quantitative PCR (qPCR)
For zebrafish tissue samples, at 72 h post-injection, all fish were anaesthetized with tricaine methanesulfonate (MS-222), dissected, and then livers were harvested and immediately frozen in liquid nitrogen and stored at −80 °C for further qPCR assays. Cell samples were harvested at 24 h after stimulation or infection. Total RNA was extracted by the TRIzol reagent (Invitrogen). First-strand cDNA was synthesized using PrimeScript reverse transcription reagent kit (TaKaRa, Kyoto, Japan) according to the manufacturer's instructions. qPCR was performed with Fast SYBR green PCR master mix (Bio-Rad, California, U.S.A) on the CFX96 real-time system (Bio-Rad). PCR conditions were as follows: 95 °C for 5 min and then 40 cycles of 95 °C for 20 s, 60 °C for 20 s, and 72 °C for 20 s. The β-actin gene was used as an internal control. The relative fold changes were calculated by comparison to the corresponding controls using the 2−ΔΔCt method.
2.8. Co-immunoprecipitation (Co-IP) assay
For Co-IP experiments, EPC cells seeded in 10 cm2 dishes overnight were transfected with a total of 10 μg of the plasmids indicated on the figures. At 24 h post-transfection, the medium was removed carefully, and the cell monolayer was washed twice with 10 mL ice-cold PBS. Then the cells were lysed in 1 mL of radioimmunoprecipitation (RIPA) lysis buffer [1% NP-40, 50 mmol/L Tris-HCl, pH 7.5, 150 mmol/L NaCl, 1 mmol/L EDTA, 1 mmol/L NaF, 1 mmol/L sodium orthovanadate (Na3VO4), 1 mmol/L phenylmethylsulfonyl fluoride (PMSF), 0.25% sodium deoxycholate] containing protease inhibitor cocktail (Sigma-Aldrich) at 4 °C for 1 h on a rocker platform. The cellular debris was removed by centrifugation at 12,000 ×g for 15 min at 4 °C. The supernatant was transferred to a fresh tube and incubated with 30 μL anti-Flag/HA/Myc affinity gel (Sigma-Aldrich) overnight at 4 °C with constant rotating incubation. These samples were further analyzed by immunoblotting (IB). Immunoprecipitated proteins were collected by centrifugation at 5000 ×g for 1 min at 4 °C, washed for three times with lysis buffer and resuspended in 100 μL 1× SDS sample buffer. The immunoprecipitates and whole cell lysates (WCLs) were analyzed by IB with the indicated antibodies (Abs).
2.9. Immunoblot analysis
Immunoprecipitates or whole cell lysates (WCLs) were analyzed as described (Lu et al., 2021). Antibodies (Abs) were diluted as follows: anti-β-actin (ABclonal, AC026) at 1:3000, anti-Flag (Sigma-Aldrich, F1804) at 1:3000, anti-HA (Covance, MMS-101R) at 1:3000, anti-Myc (Santa Cruz Biotechnology, sc-40) at 1:3000, anti-LC3 (Abcam, ab48394) at 1:1000, and HRP-conjugated anti-mouse/rabbit IgG (Thermo Scientific, 31430/31460) at 1:5000. The indicated Abs of SVCV proteins (includes N, and P, and G) at 1:1000 and were prepared by our lab (Li et al., 2019).
2.10. Immunofluorescence (IF)
EPC cells were plated onto glass coverslips in 6-well plates and infected with SVCV (MOI = 1) or co-infected with SVCV and GCRV (MOI = 0.001/0.01) for 24 h. Then the cells were washed with PBS and fixed in 4% paraformaldehyde (PFA) at room temperature for 1 h and permeabilized with 0.2% Triton X-100 in ice-cold PBS for 15 min. The samples were blocked for 1 h at room temperature in PBS containing 2% bovine serum albumin (BSA, Sigma-Aldrich). After additional PBS washing, the samples were incubated with anti-N or anti-P Abs in PBS containing 2% BSA for 2–4 h at room temperature. After three times washed by PBS, the samples were incubated with secondary Ab (Alexa Fluor 488 AffiniPure Donkey anti-Rabbit IgG (H + L) (34206ES60, 1:10,000) in PBS containing 2% BSA for 1 h at room temperature. After additional PBS washing, the cells were finally stained with 1 μg/ml 4′, 6-diamidino-2-phenylindole (DAPI; Beyotime Institute of Biotechnology, Shanghai, China) for 10 min in the dark at room temperature. Finally, the coverslips were washed and observed with a confocal microscope under a 10× immersion objective (SP8; Leica). Heart, liver, and spleen of zebrafish were isolated and fixed overnight at 4 °C in 4% PFA, then rinsed in PBS and dehydrated by permeation with 30% sucrose overnight at 4 °C. After being fixed in O.C.T. compound (SAKURA, U.S.A) and frozen at −80 °C, 10 μm tissue slices were sliced using a frozen sectioning machine for further treatment similar to the cells.
2.11. Transmission electron microscopy (TEM)
EPC cells were seeded in 6-well plates and transfected with indicated plasmids using FishTrans (MeiSenTe Biotechnology) for 24 h. For pretreatment, cells were washed with PBS, trypsinized, and transferred to 1.5 mL centrifuge tube. Cell precipitation was collected by centrifugation at 2000 ×g for 5 min. The cell pellets were resuspended with 2.5% glutaraldehyde in 0.075 mol/L phosphate buffer (pH 7.4) for 4 h at 4 °C for prefixation. Then the cells were washed for three times with solution containing 0.075 mol/L phosphate and 0.19 mol/L sucrose for 15 min each and post-fixed in 1% osmium tetroxide (OsO4) in 0.24 mol/L phosphate buffer (pH 7.4) for 2 h. After being washed for three times for 15 min each in 0.075 mol/L phosphate buffer and 0.19 mol/L sucrose buffer at 4 °C, the cells were dehydrated with a graded series of ethanol and acetone, and then gradually infiltrated with epoxy resin. Samples were sequentially polymerized at 37 °C overnight and then 60 °C for 48 h. Ultrathin sections (74 nm) were cut using microtome (UC7; Leica) and mounted on copper slot grids. Sections were doubly stained with 3% uranyl acetate-lead citrate for 10 min and observed under transmission electron microscope (HT7700; Hitachi, Tokyo, Japan).
2.12. In vitro ubiquitination assay
Transfected EPC cells were washed twice with 10 mL ice-cold PBS and then digested with 1 mL 0.25% trypsin-EDTA (1×) (Invitrogen) for 2–3 min until the cells were dislodged. 100 μL FBS was added to neutralize the trypsin and the cells were resuspended into 1.5 mL centrifuge tube, centrifuged at 2000 ×g for 5 min. The supernatant was discarded and the cell precipitations were resuspended using 1 mL PBS and centrifuged at 2000 ×g for 5 min. The collected cell precipitations were lysed using 100 μL PBS containing 1% SDS and denatured by heating for 10 min. The supernatants were diluted with lysis buffer until the concentration of SDS was decreased to 0.1%. The diluted supernatants were incubated with 20 μL anti-Myc/Flag affinity gel overnight at 4 °C with constant agitation. These samples were further analyzed by IB. Immunoprecipitated proteins were collected by centrifugation at 5000 ×g for 1 min at 4 °C, washed for three times with lysis buffer and resuspended in 100 μL 1× SDS sample buffer.
2.13. Fluorescent microscopy
Fluorescence confocal detection of EPC cells was generated as described (Lu et al., 2021).
2.14. Statistics analysis
For fish survival analysis, Kaplan-Meier survival curves were generated and analyzed by Log-rank test. For the bar graph, one representative experiment of at least three independent experiments is shown, and each was done in triplicate. qPCR data analyzed for significance were performed as mean ± standard error of the mean (SEM). The P values were calculated using the Student's t-test. A P value < 0.05 was considered statistically significant.
3. Results
3.1. GCRV significantly suppresses the infection and proliferation of SVCV in vivo
SVCV is a highly lethal virus that infects zebrafish. Only 10.3% of zebrafish survive in SVCV infection. SVCV infected fish co-infected with GCRV-Ⅱ (RNA virus) or CaHV (DNA virus), and the mortality was unaffected. Interestingly, co-infection with SVCV and GCRV-Ⅰ (RNA virus) significantly suppressed the lethality of SVCV, and nearly 90.9% of the fish survived (Fig. 1A). The lethality of just GCRV-Ⅰ infection was checked. The fish mortality in only GCRV infection was weaker than in SVCV, and 86.7% of the fish survived (Fig. 1B). The reduced mortality rate in GCRV means that the GCRV is a mild virus in zebrafish different from the severe infection from SVCV. Three zebrafish organs were chosen as representatives for analysis of SVCV-GCRV co-infection, the heart, liver, and spleen. In an H&E staining assay, severe tissue damage was observed in SVCV infection. However, there was remarkable less tissue damage in the co-infection group and it was almost indistinguishable from the un-infected group (Fig. 1C). In IF, the green fluorescence represented the SVCV N protein and was strongly detected in these tissues, while it was undetectable in the SVCV-GCRV co-infected groups (Fig. 1D). At the protein level, both SVCV N and P proteins were detected in the SVCV infected group, consistent with the results above. These viral proteins could not be detected in the co-infected group (Fig. 1E). Finally, the increase in viral transcripts that indicates proliferation of SVCV was suppressed in the SVCV-GCRV co-infected group (Fig. 1F). These data demonstrated that SVCV infections in fish were significantly attenuated by GCRV co-infection.
Fig. 1.
GCRV blocks the amplification of SVCV in zebrafish. A Survival rates of zebrafish infected with different viruses in combination with SVCV. Adult zebrafish were injected with SVCV [5 × 109 50% tissue culture infective dose (TCID50)/mL, 5 μL/individual] and GCRV/GCRV-II/CaHV (5 × 107 TCID50/mL, 5 μlLindividual). Mortality recorded over eight days (number of individuals = 15). B Survival rates of zebrafish decreased with combined injection of GCRV and SVCV. Adult zebrafish were injected with PBS only (10 μL/individual), SVCV (5 × 109 TCID50/mL, 5 μL/individual and 5 μL PBS to make up 10 μL/individual), GCRV (5 × 107 TCID50/mL, 5 μL/individual and 5 μL PBS to make up 10 μL/individual), or SVCV&GCRV [(5 × 109 TCID50/mL, 5 μL/individual) for SVCV and (5 × 107 TCID50/mL, 5 μL/individual) for GCRV]. Mortality recorded over eight days (number of individuals = 35). C Microscopy of hematoxylin and eosin (H&E)-stained heart, liver, and spleen sections from male and female zebrafish treated with SVCV (5 × 109 TCID50/mL, 5 μL/individual) or SVCV&GCRV [(5 × 109 TCID50/mL, 5 μL/individual) for SVCV and (5 × 107 TCID50/mL, 5 μL/individual) for GCRV] for 72 h. Histological changes were examined by optical light microscopy for histological changes at ×40 magnification. The location of the pathological area was indicated by white arrows. D Tissue immunofluorescence (IF) of liver, spleen, and heart tissue sections of SVCV infected zebrafish. The green fluorescence of tissue slices was observed with a confocal microscope under a 20× immersion objective (SP8; Leica) and indicates proliferation of SVCV. Scale bar, 10 μm. E Immunoblotting (IB) detection of virus abundance in the tissues of zebrafish in Null, SVCV, and SVCV&GCRV treatment groups. Scale bar, 20 μm. F Transcript levels of SVCV genes in zebrafish tissues of indicated treatment groups. The β-actin gene was used as an internal control and the relative expressions of SVCV genes were represented as fold induction relative to the expression level in control cells (set to 1). Data were expressed as mean ± standard error of the mean (SEM), n = 3. Statistical analysis was performed by the Student's t-test. Asterisks indicate significant differences from control (∗P < 0.05).
3.2. GCRV attenuates the dysregulation of the host immune response and cellular metabolism caused by SVCV
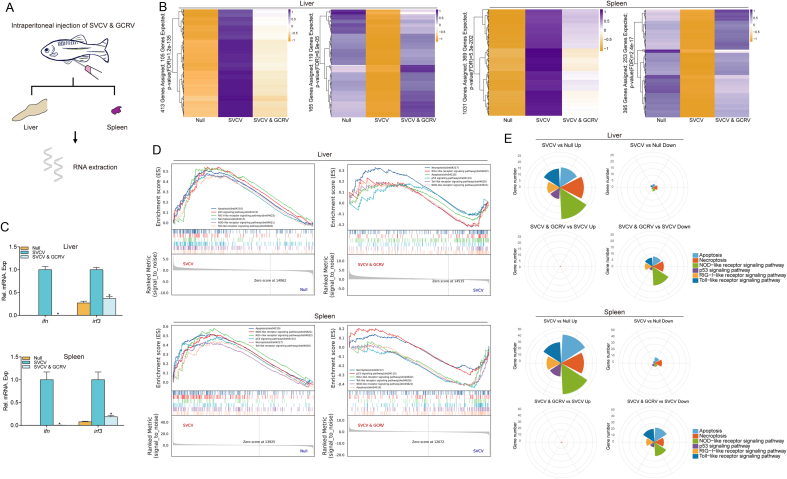
Co-infection of SVCV and GCRV reduced the lethality of SVCV, but the impact of the co-infection on the host immune system needs further studies. Total RNAs from zebrafish liver and spleen were extracted and subjected to transcriptome analysis (Fig. 2A). Short Time-series Expression Miner (STEM) software was used to cluster the genes with similar changes (Ernst and Bar-Joseph, 2006), and P-values were corrected by the False Discovery Rate method (FDR). Those with P-values less than 0.05 were selected as significant modules. There were four modules screened in gene set of liver tissue and five modules in gene set of spleen tissue (Supplementary Fig. S1). The SVCV infections led to the activation of biological process related to defense of virus and suppression of the metabolic process (Supplementary Fig. S2). In contrast, these same genes were expressed at normal levels in the co-infection group. Overall, the disruption of gene expression was caused by SVCV infection and was rescued by GCRV co-infection (Fig. 2B). This was also verified by qPCR assays on ifn and IFN regulatory 3 (irf3) (Fig. 2C). In addition, the UniGene sequences of the four groups were searched in the Kyoto Encyclopedia of Genes and Genomes (KEGG) databases. A fold change (FC) value greater than or equal to 1.5 was set as the threshold for expression pattern differences. Those genes were annotated as differentially expressed genes (DEGs). In the two comparison groups, SVCV versus Null and SVCV-GCRV versus SVCV, the KEGG-enriched top 20 signaling pathways that associated with immune response and cell cycle were selected for further analysis, including three typical pattern recognition receptor (PRR) signaling pathways: nucleotide-binding oligomerization domain (NOD)-like receptors (NLRs) signaling pathway, retinoic acid-inducible gene I (RIG-I)-like receptors (RLRs) signaling pathway, and Toll-like receptors (TLRs) signaling pathway. We also selected three representative signaling pathways for the cell cycle: necroptosis, apoptosis, and the p53 signaling pathway (Supplementary Fig. S3). Gene Set Enrichment Analysis (GSEA) was performed on those pathways. As shown in Fig. 2D, the plotting of the GSEA enrichment results was used to visualize these perturbed pathways. The results showed that these signaling pathways were activated overall in response to SVCV infection, while these signaling pathways are not activated in the co-infection group. The rose plots of DEGs of the above signaling pathways showed that most genes were upregulated after SVCV infection and downregulated in the SVCV-GCRV co-infected group (Fig. 2E). Collectively, these data demonstrated that GCRV co-infection inhibited the proliferation of SVCV in zebrafish and resulted in a significant reduction in the gene expression changes caused by SVCV on the immune system and metabolism.
Fig. 2.
The effect of SVCV on zebrafish gene transcription is eliminated by GCRV. A Schematic representation of zebrafish tissue dissection and RNA extraction for transcriptome sequencing. The liver and spleen tissues from male and female zebrafish injected with PBS (10 μL/individual), SVCV [5 × 109 50% tissue culture infective dose (TCID50)/mL, 5 μL/individual and 5 μL PBS to make up 10 μL/individual] and SVCV&GCRV (5 × 107 TCID50/mL, 5 μL/individual) for 72 h. Total RNAs were extracted and used for transcriptome sequencing and analysis. B Heat maps of representative gene modules of STEM analysis in different tissues. The P-values were corrected for the False Discovery Rate (FDR) method, and modules with P-values less than 0.05 were selected as significant. C Quantitative PCR (qPCR) validates changes in representative genes of immune system. The β-actin gene was used as an internal control, and the relative expressions of ifn and irf3 genes were represented as fold induction relative to the expression level in control cells (set to 1). Data were expressed as mean ± standard error of the mean (SEM), n = 3. Statistical analysis was performed by the Student's t-test. Asterisks indicate significant differences from control (∗P < 0.05). D GSEA enrichment plots in SVCV versus Null and SVCV&GCRV versus SVCV groups. E Rose plots showing the number of DEGs at SVCV versus Null and at SVCV&GCRV versus SVCV in selected pathways (FC > 1.5, adjusted P < 0.05).
3.3. Identification of the intracellular interactions between SVCV and GCRV
Since the interference of GCRV on SVCV proliferation has been characterized in vivo, the mechanisms were explored by the study in vitro. First, a crucial point needed to be clarified, whether the regulation happened extracellular or intracellular. Three groups of viral infections were tested: SVCV infected and then GCRV infected 8 h later, GCRV infected and then SVCV infected 8 h later, and SVCV and GCRV co-infected at the same time point. Interestingly, in the co-infected groups, the expression of SVCV viral proteins such as N and P were reduced compared to the non-co-infected SVCV group (Fig. 3A–C). These data suggested that the viral components regulated alterations seen in co-infection intracellularly. The effect of GCRV on SVCV replication was observed. SVCV viral genome transcription and protein expression were significantly repressed when co-infected with GCRV (Fig. 3D and E). In an IF assay, the green fluorescence represented SVCV N or P proteins and were observed in abundance, whereas these signals decreased as the GCRV titer increased (Fig. 3F). Compared with the severity of cells infected with SVCV alone, the co-infection with GCRV significantly reduced the severity of SVCV. There was less CPE exhibited in a dose-dependent manner with GCRV titers, and the viral titer in the co-infection group also remarkably declined (Fig. 3G and H). The data above demonstrates that GCRV negatively regulates SVCV proliferation intracellularly.
Fig. 3.
GCRV affects proliferation SVCV in fish cell lines. A–C SVCV is degraded by GCRV in all three different co-infection modes. EPC cells were seeded in 6-well plates overnight and infected with SVCV and GCRV in three modes: SVCV infected and then GCRV infected 8 h later, GCRV infected and then SVCV infected 8 h later, and SVCV and GCRV co-infected at the same time point. After 24 h of subsequent infection, cells were collected for immunoblotting (IB), using the indicated antibodies (Abs). D, E The viral gene transcription and protein expression of SVCV at different titers of GCRV co-infection with SVCV. EPC cells (for IB) were seeded in 6-well plates overnight and co-infected with SVCV (MOI = 1) and GCRV (MOI = 0.01) for 36 h. The total RNAs were extracted to examine the mRNA levels of SVCV genes. The β-actin was used as an internal control for normalization, and the relative expression was represented as fold induction relative to the expression level in control cells (set to 1). Data were expressed as mean ± standard error of the mean (SEM), n = 3. Statistical analysis was performed by the Student's t-test. Asterisks indicate significant differences from control (∗P < 0.05). The expressions of SVCV proteins were determined by IB with the anti-N protein, anti-P protein, and anti-β-actin Abs. F Immunofluorescence (IF) analysis of SVCV proteins at different titers of GCRV co-infection with SVCV. The green fluorescence of N and P proteins were observed under a 20× immersion objective (SP8; Leica). Scale bar, 50 μm. G, H Detection of viral amplification of SVCV at co-infection with different titers of GCRV. EPC cells were seeded in 24-well plates overnight and infected with the indicated concentrations of GCRV (MOI = 0.0001/0.001/0.01) and SVCV (MOI = 1) for 48 h. The cells were fixed with 4% PFA and stained with 1% crystal violet for visualizing CPE, and the viral titers of the supernatants were determined by 50% tissue culture infective dose (TCID50) assays.
3.4. The S9 segment of GCRV encoded viral protein inhibits SVCV proliferation
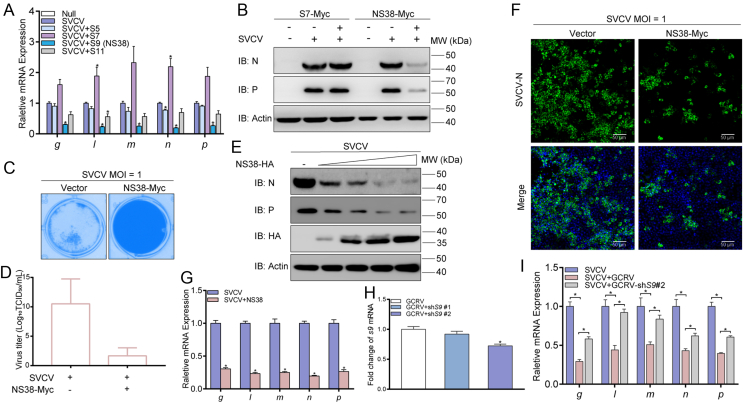
Since we identified the negative regulation pattern in GCRV-SVCV co-infection occurred intracellularly, we proposed the hypothesis that this regulation was mediated by viral components. Segments of the GCRV genome were cloned and overexpressed in cells infected with SVCV. The S9 segment, the viral segment that encodes the NS38 protein, had a significant inhibition on SVCV gene replication (Fig. 4A). At the protein level, NS38 remarkably decreased SVCV viral protein expressions. This decrease was not observed in the control S7 segment group (Fig. 4B). The proliferation of SVCV affected by NS38 was further explored. Compared with the control group, the CPE caused by SVCV was significantly impaired when NS38 was expressed, and the viral titer assay demonstrated that NS38 declined SVCV proliferation (Fig. 4C and D). Besides the decreased SVCV viral proteins correlating with NS38 in a dose-dependent manner, green fluorescence represented SVCV N proteins have almost disappeared when NS38 was overexpressed (Fig. 4E and F). As anticipated, all the SVCV viral genes were downregulated by NS38 overexpression (Fig. 4G). According to the nucleic acid sequence of S9, two shRNA were designed and one of which was effective. The effective shRNA was employed and successfully prevented the GCRV-associated reduction of SVCV replication (Fig. 4H and I). These data suggests that NS38 of GCRV plays a negative role on SVCV in co-infection.
Fig. 4.
The NS38, S9 segment-encoded protein of GCRV, inhibits SVCV proliferation. A, B Effect of overexpression of GCRV segments on the SVCV proliferation. EPC cells were seeded in 6-well plates and transfected with indicated plasmids (1 μg each) for 24 h, then cells were infected with SVCV (MOI = 1) for 24 h prior to being harvested for RNA extraction and quantitative PCR (qPCR). The β-actin was used as an internal control for normalization. The relative expressions of SVCV genes were represented as fold induction relative to the expression level in control cells (set to 1). Data were expressed as mean ± standard error of the mean (SEM), n = 3. Statistical analysis was performed by the Student's t-test. Asterisks indicate significant differences from control (∗P < 0.05). The expressions of SVCV proteins were determined by immunoblotting (IB) with the anti-N protein, anti-P protein, and anti-β-actin antibodies (Abs). C, D Detection of the viral titer of SVCV effected by overexpression of NS38. EPC cells were seeded in 24-well plates overnight and transfected with 2 μg NS38-Myc or vector. After 24 h, cells were infected with SVCV (MOI = 1) for 48 h. The cells were fixed with 4% PFA and stained with 1% crystal violet for visualizing cellular cytopathic effect (CPE) (C), and the viral titers of the supernatants were determined by 50% tissue culture infective dose (TCID50) assays (D). E NS38 degrades SVCV on a dose-dependent manner. EPC cells were seeded in 6-well plates and transfected with 0.5, 1, 1.5, or 2.0 μg NS38-Myc or empty vector for 24 h, then cells were infected with SVCV (MOI = 1) for 24 h prior to being harvested for IB analysis of WCLs with the indicated Abs. F–I NS38 inhibits proliferation SVCV. Immunofluorescence (IF) analysis was used to indicate amplification of SVCV, with green fluorescence representing the N protein (F). The mRNA levels of SVCV genes were detected upon overexpression and knockdown of S9 (G, I). H Transcript levels of S9 tested the efficiency of knockdown. Data were expressed as mean ± SEM, n = 3. Statistical analysis was performed by the Student's t-test. Asterisks indicate significant differences from control (∗P < 0.05).
3.5. GCRV NS38 interacts with the SVCV N and P protein
Since NS38 interferes intracellularly with SVCV proliferation, we speculated that NS38 subverted the viral components of SVCV replication. To verify this, the protein interaction between NS38 and the SVCV proteins was determined. First, in a subcellular localization assay, NS38 is located in the same cytoplasmic region as the P protein. In addition, the N protein aggregates in the cytoplasmic region partially overlapped with NS38 (Fig. 5A). The cytoplasmic localization of these proteins facilitated potential protein interactions. We found that NS38 was associated with the SVCV N and P proteins but not with the SVCV glycoprotein (G). These results proved that the interaction between the viral proteins of the two viruses occurred (Fig. 5B). These were confirmed by the results of the reverse IP assay (Fig. 5C). Based on the observation that NS38 interfered with SVCV infections and that the SVCV protein interacted with NS38, we presumed that SVCV viral proteins were downregulated by NS38. Meanwhile, in an IF experiment, the green fluorescence of the N or P proteins was significantly reduced when NS38 was overexpressed. This observation was confirmed by IB for the N or P proteins and fluorescence intensity statistics. The G protein expression was not changed in the NS38 overexpression group (Fig. 5D–F, Supplementary Fig. S4). We verified that SVCV N and P proteins decreased and were directly regulated by NS38 resulting in the suppression of SVCV proliferation. The N and P proteins were tagged with a Flag-tag and were co-expressed with NS38. The expression of N or P proteins was dramatically reduced in association with NS38 expression and the effect was confirmed to be a dose-dependent effect (Fig. 5G–I). These findings suggested that the SVCV N and P proteins interacted with and were reduced by GCRV NS38.
Fig. 5.
NS38 associates with and degrades N and P proteins. A NS38 localizes in the cytoplasm. EPC cells were plated onto coverslips in 6-well plates and transfected with indicated plasmids (1 μg each). After 24 h, the cells were fixed and subjected for confocal microscopy analysis. Scale bar, 5 μm. B, C NS38 associates with N and P proteins. EPC cells seeded in 10-cm2 dishes overnight were transfected with the indicated plasmids (5 μg each). After 24 h, cell lysates were immunoprecipitated (IP) with anti-Myc/Flag affinity gels. Then, the immunoprecipitates and cell lysates were analyzed by immunoblotting (IB) with indicated antibodies (Abs). D–F NS38 degrades N and P proteins and barely affects G protein expression. EPC cells were plated onto coverslips in 6-well plates and transfected with indicated plasmids (1 μg each). After 24 h, the cells were fixed and subjected for confocal microscopy analysis. Green or red signals represent overexpressed N, P, and G proteins (original magnification 10×; non-immersion objective). Scale bar, 300 μm. The fluorescence intensity (arbitrary unit, a.u.) was recorded by the LAS X software, and the data were expressed as mean ± SD, n = 5. Statistical analysis was performed by the Student's t-test. Asterisks indicate significant differences from control (∗P < 0.05). IB with indicated Abs was used to detect the expression level of N/P/G proteins. G–I NS38 degrades N and P proteins on a dose-dependent manner. EPC cells were seeded in 6-well plates and transfected with 1 μg N-Flag or P-Flag and 1 μg NS38-Myc (G) or 0.5, 1.0, or 1.5 μg NS38-Myc (H, I). At 24 h post-transfection, the cells were harvested for IB with the indicated Abs.
3.6. NS38 targets the SVCV N protein for autophagic degradation
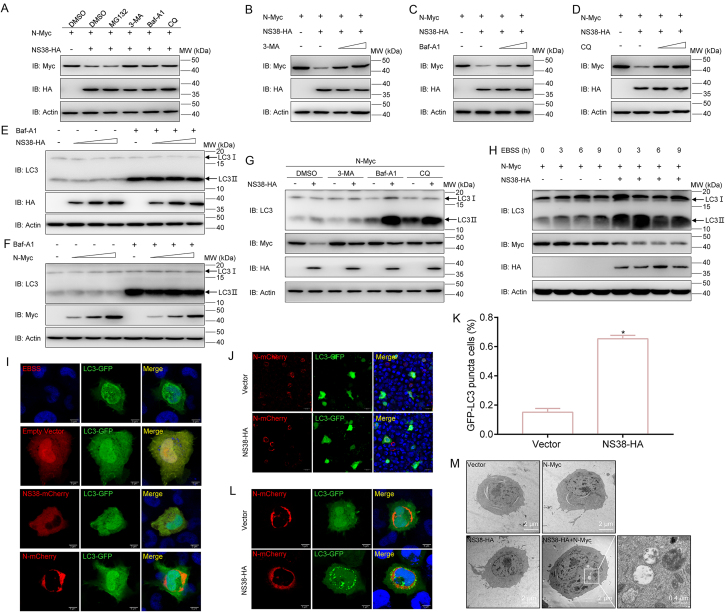
According to our previous results, the SVCV N protein interacted with and was reduced when NS38 was expressed; therefore, the regulatory mechanisms were verified. There are two classical protein degradation mechanisms: Ub-proteasome degradation and autophagy-lysosome degradation. These pathways can be inhibited by MG132, which inhibits the Ub-proteasome pathway; and 3-MA, Baf-A1, or CQ which inhibit the autophagy-lysosome pathway. When treated with different inhibitors, the reduction in N protein regulated by NS38 was not blocked in the MG132 group; however, the autophagic pathway inhibitors, 3-MA, Baf-A1, and CQ significantly inhibited the reduction of the N protein. These results suggested that the N protein was degraded by the autophagic-lysosome pathway (Fig. 6A). These data were confirmed by different inhibitor doses, elucidating that NS38 degraded the N protein via the autophagic-lysosome pathway (Fig. 6B–D). LC3-II is considered an autophagy indicator and was used to verify the process of N protein degradation by NS38. First, we explored if the N protein or NS38 induced cellular autophagy. Interestingly, when either NS38 or the N protein alone was overexpressed, the amount of LC3-II did not increase and suggested that alone NS38 or the N protein did not evoke cellular autophagy (Fig. 6E and F). The co-expression of the N protein and NS38 resulted in increased LC3-II expression indicating increased cellular autophagy (Fig. 6G). These results demonstrate that cellular autophagy is initiated only when both the N protein and NS38 are expressed. We also verified if the normal host autophagy response degraded the N protein. Starvation has been shown to induce cellular autophagy. To induce starvation, cells were treated with Earle's Balanced Salt Solution (EBSS) (Deng et al., 2021) for 9 h. The expression of the N protein was unaffected when the cells were starved, and the N protein was only degraded when NS38 was expressed, even during starvation (Fig. 6H). These results indicated that the degradation of the N protein is regulated by NS38. LC3 puncta formation is a sign of autophagosome flux. In a confocal microscopy analysis, during starvation, which we used as a positive control, increased cellular LC3 puncta accumulation was observed. However, LC3 accumulation was not observed when only NS38 or the N protein was expressed (Fig. 6I). When the N protein and NS38 were co-expressed, there was a strong green fluorescent protein (GFP) signal indicating the formation of LC3 puncta and the initiation of autophagy (Fig. 6J–L). We further verified the formation of autophagosomes through a TEM analysis. Autophagosome-like vesicles, which contained cytosolic contents, were observed in the co-overexpressed cells with the N protein and NS38. However, there were few autophagosome-like vesicles when the N protein or NS38 was individually expressed (Fig. 6M). These results illustrates that the SVCV N protein is degraded by NS38 in an autophagic manner, and this autophagy is only evoked when both the N protein and NS38 are expressed.
Fig. 6.
NS38 degrades N proteins in an autophagy-dependent manner. A Inhibitors of autophagy progress block NS38-mediated degradation of N protein. EPC cells were seeded in 6-well plates overnight and co-transfected the indicated plasmids (1 μg each). At 18 h post-transfection, the cells were treated with DMSO, MG132 (20 μmol/L), 3-MA (2 mmol/L), Baf-A1 (100 nmol/L), or CQ (100 μmol/L) for 6 h. The cell lysates were subjected to immunoblotting (IB) with the indicated antibodies (Abs). B–D The NS38-induced N protein degradation is rescued by 3-MA, Baf-A1, and CQ on a dose-dependent manner. EPC cells were seeded in 6-well plates overnight and co-transfected with the indicated plasmids. At 18 h post-transfection, the cells were treated with 3-MA (1 or 2 mmol/L), Baf-A1 (50 or 100 nmol/L) or CQ (50 or 100 μmol/L) for 6 h. Then, the cells were harvested for IB with the indicated Abs. E, F The presence of NS38 or N-only does not affect autophagy flux. EPC cells were seeded in 6-well plates and transfected with N-Myc or NS38-HA (0.5, 1, or 1.5 μg). At 18 h post-transfection, the cells were treated with DMSO or Baf-A1 (100 nmol/L) for 6 h. Then, the cells were harvested for IB with the indicated Abs. G, H Co-expression of NS38 and N protein specifically activates autophagy. EPC cells were seeded in 6-well plates overnight and co-transfected the indicated plasmids (1 μg each). At 18 h post-transfection, the cells were treated with DMSO, 3-MA (2 mmol/L), Baf-A1 (100 nmol/L), or CQ (100 μmol/L) for 6 h. The cell lysates were subjected to IB with the anti-LC3, anti-Myc, anti-HA and anti-β-actin Abs, respectively. EBSS was used to exclude the effect of non-specific cellular autophagy activation on N protein expression. I–L Co-expression of NS38 and N protein induces aggregation of LC3. EPC cells were plated onto coverslips in 6-well plates and co-transfected with 1 μg pCS2-mCherry, NS38-mCherry, or N-mCherry plus 1 μg LC3-GFP. After 24 h, the cells were fixed and observed by confocal microscopy. Red signals represent overexpressed NS38 or N protein, green signals represent overexpressed LC3 (original magnification 63×; oil immersion objective). Scale bar, 5 μm. K The percentage of cells in which LC3 aggregates (J) was quantified under a 20× immersion objective (SP8; Leica) and data were expressed as mean ± SD, n = 3. Statistical analysis was performed by the Student's t-test. Asterisks indicate significant differences from control (∗P < 0.05). M Autophagosome-like structures detection by transmission electron microscopy (TEM). EPC cells were seeded in 6-well plates overnight and transfected with indicated plasmids (1 μg each) for 24 h. The cells were then analyzed by TEM, enlarged section indicating autophagic vesicles. Scale bar, 2 or 0.4 μm.
3.7. Autophagy receptor p62 is crucial for GCRV NS38-mediated autophagic degradation of the SVCV N protein
Since the SVCV N protein is autophagic degraded by GCRV NS38, we explored the precise process. Autophagy receptors are a class of bridging proteins that play a pivotal role in cellular autophagy and act as a connecting protein to link the target substrates to the autophagosome (Deng et al., 2017). By screening several autophagy receptors, p62, a typical autophagy receptor that interacts and transfers substrates to the autophagosome, was selected (Tao et al., 2020). The p62 associated with NS38, indicating that p62 is crucial for the NS38 autophagic function (Fig. 7A and B). The role of p62 in NS38 degradation of the N protein was further explored. The N protein was not affected when only p62 was expressed; however, when p62 was expressed with NS38, there was increased N protein degradation (Fig. 7C). These results suggest that p62 promotes NS38 degrading of the N protein. However, knockdown of p62 by shRNA significantly reduced N protein degradation by NS38 (Fig. 7D). These data suggest p62 is the autophagy receptor involved in NS38 autophagic degradation of the N protein.
Fig. 7.
NS38 manipulates p62 to degrade N protein and affects biological function of N protein. A, B NS38 associates with autophagy receptor p62. EPC cells seeded in 10-cm2 dishes overnight were transfected with the indicated plasmids (5 μg each). After 24 h, cell lysates were immunoprecipitated (IP) with anti-Flag affinity gels or anti-HA affinity gels. Then, the immunoprecipitates and cell lysates were analyzed by immunoblotting (IB) with anti-Flag, anti-Myc and anti-HA antibodies (Abs), respectively. C, D The p62 is essential for NS38 degradation of N protein. Overexpression of p62 promotes the degradation of N protein by NS38 (C). EPC cells were seeded in 6-well plates overnight and co-transfected the indicated plasmids. After 24 h, the cell lysates were subjected to IB with indicated Abs. Knockdown of p62 eliminates the degradation of N protein by NS38 (D). EPC cells were seeded in 6-well plates overnight and transfected with 2 μg shp62#1-pLKO.1, shp62#2-pLKO.1 or the negative control (sh-NC) with other indicated plasmids for 24 h. The cell lysates were then subjected to IB with indicated Abs.
3.8. NS38 attenuates the K63-linked ubiquitination and promotes degradation of the SVCV P protein
As mentioned above, NS38 interacts and impairs both the SVCV N and P proteins. Besides the autophagic degradation of the N protein, we also studied the downregulation of the P protein by NS38. Treatment with MG132 significantly blocked the attenuation of the P protein. However, when treated with 3-MA, Baf-A1, or CQ, the P protein was not affected, distinguishing with the N protein (Fig. 8A). To verify that NS38 ubiquitinated the P protein, different doses of MG132 treatments were given, and the results were consistent with the blocking of P protein attenuation (Fig. 8B). We next explored the Ub levels of P protein degradation. The ubiquitination of the P protein declined in NS38-mediated P protein degradation (Fig. 8C). K48-linked polyubiquitin chain modification (Ub-K48) targets protein for proteasome degradation, while the K63-linked polyubiquitin chain modification (Ub-K63) improves the stability of the target protein. Therefore, identification of which Ub modification on the P protein was crucial. The wild-type Ub and Ub-K63 were significantly reduced when the P protein was degraded, and Ub-K48 was unchanged (Fig. 8D). These results suggested that NS38 decreased the K63-linked ubiquitination of P protein to negatively regulate the P protein. These data demonstrated that P protein was negatively regulated by NS38 through decreased Ub-K63.
Fig. 8.
NS38 degrades P protein via the ubiquitin-proteasome pathway and inhibits its biological function. A, B MG132 blocks NS38-mediated degradation of P protein. EPC cells were seeded in 6-well plates overnight and co-transfected the indicated plasmids (1 μg each). At 18 h post-transfection, the cells were treated with DMSO, MG132 (20 μmol/L or a dose gradient of 10 μmol/L, 20 μmol/L), 3-MA (2 mmol/L), Baf-A1 (100 nmol/L), or CQ (100 μmol/L) for 6 h. The cell lysates were subjected to immunoblotting (IB) with the indicated antibodies (Abs). C, D NS38 reduces the ubiquitination of P protein through inhibiting K63-linked endogenous ubiquitination of P protein. EPC cells were transfected with 5 μg P-Myc, 5 μg NS38-Flag or empty vector, and 1 μg Ub-HA, Ub-K48O-HA or Ub-K63O-HA. At 18 h post-transfection, the cells were treated with MG132 (20 μmol/L) for 6 h. At 24 h post-transfection, cell lysates were immunoprecipitated (IP) with anti-Myc affinity gel. Then the immunoprecipitates and WCLs were analyzed by IB with the indicated Abs.
3.9. NS38 influences the biological functions of N and P proteins
On the cellular level, we examined the effect of NS38 on N protein-mediated cell antiviral capacity abrogation. Under SVCV infection, overexpression of N protein accelerated the CPE formation compared with the normal infection, while such disruption mediated by N protein was significantly blocked when co-expressed with NS38. The viral titer assay also demonstrated that a higher viral titer was observed in N protein group, while it was suppressed in NS38 co-expressed group (Fig. 9A and B). It was suggested that the molecular function of SVCV N protein was dampened by NS38. Furthermore, N protein polymerization is considered necessary for viral proliferation, and it was also checked with NS38 existence. As shown in Fig. 9C, N protein with a Flag-tag interacted with a Myc-tagged N protein, and such association is the base of polymerization. However, the interaction of N proteins was blunted by NS38 overexpression. In addition, the biological function of the P protein when NS38 is expressed was identified. In our previous study, the P protein negatively regulated the host cell antiviral defense (Li et al., 2016), therefore, the immune evasion of P protein was also measured. Overexpression of the P protein caused the cells to be vulnerable for SVCV infections, while the cellular antiviral capacity was almost restored when co-expressed with NS38 (Fig. 9D and E). Dimerization of the P protein is necessary for viral replication (Bruhn et al., 2019). Therefore, the effect of NS38 on P protein dimerization was explored. In the control group, the P protein with a Myc-tag interacted with a Flag-tagged P protein, suggesting that there was dimerization. However, such interactions were remarkably reduced when NS38 was overexpressed (Fig. 9F). These data suggested that the cellular biologic functions of N protein and P protein were suppressed by NS38.
Fig. 9.
NS38 inhibits the biological functions of N and P proteins. A, B, D, E The effect of N/P on promoting SVCV amplification is attenuated by NS38. EPC cells were seeded in 24-well plates overnight and transfected with indicated plasmids (0.25 μg each) for 24 h, and were then infected with SVCV (MOI = 1). The viral titers of the supernatants were determined by 50% tissue culture infective dose (TCID50) assays. C, F N/P proteins dimerization is attenuated by NS38. EPC cells seeded in 10-cm2 dishes overnight were transfected with the indicated plasmids (5 μg each). After 24 h, cell lysates were immunoprecipitated (IP) with anti-Flag affinity gels. Then, the immunoprecipitates and cell lysates were analyzed by immunoblotting (IB) with indicated antibodies.
4. Discussion
Diseases caused by viral co-infection can be easily and accurately diagnosed in humans, for example, hepatitis B caused by HBV and acquired immune deficiency syndrome (AIDS) caused by HIV (Sulkowski, 2008). Most land animals are infected with a single virus at a time, whereas co-infection of multiple viruses is more common in fish. There are at least three main reasons for that co-infection in fish is common. Firstly, there is a larger variety and number of aquatic viruses than is currently known. Yuri et al. reported that more the 4500 distinct RNA viruses were identified in approximately 10 L of water from a harbor near Shanghai, China, which doubled the previously known number of viruses (Wolf et al., 2020). The fact that there are so many viruses present in water increases the chances of co-infection in fish. Secondly, water increases the survival time of viruses relative to land and facilitates the spread. Therefore, different viruses carried by fishes are more likely to form co-infection. Thirdly, many fish viruses are infectious but not lethal. For example, the SVCV can infect and be carried by gibel carp but not lead to death unless they were injected with an extremely high viral titer (Simkova et al., 2022). There are three types of GCRV, GCRV-I, GCRV-II, and GCRV-III, to our knowledge (Pei et al., 2014; Zhang and Gui, 2015). In recent years, only GCRV-II has been shown to be a high risk for grass carp, whereas infections with GCRV-I or GCRV-III cause asymptomatic infection (Zhang and Gui, 2018). The asymptomatic fish become a natural carrier for GCRV, leaving other viruses able to co-infect. The co-infection would lead to either disease aggravation, mitigation, or have no effect. In this study, we reported a disease mitigation case caused by co-infection of SVCV and GCRV. Our findings showed that the existence of intracellular viral proteins interfered with other viral processes, providing another perspective on viral co-infection.
Transcriptomic analysis of the viral-infected samples showed that the severity of viral infection is usually positively correlated with the high expression of immune-relevant genes in vivo (Gupta and Rao, 2011; Rosenberg et al., 2018). Transcriptomic analysis of nervous necrosis virus (NNV)-infected Epinephelus malabaricus showed that significant differentially expressed transcripts included immune-relevant pathways (Tso and Lu, 2018), which is consistent with our data. In our research, the GCRV-SVCV co-infection resulted in a marked alleviation of zebrafish viral-associated damage and a marked inhibition of SVCV proliferation in zebrafish tissues. We speculate that GCRV ameliorated the immune system disruption caused by SVCV. Consistent with this idea, upregulation of gene pathways associated with immune system activation in DEGs of the SVCV-infected group was observed. It showed an opposite trend in the co-infection group (Fig. 2D and E). Furthermore, STEM analysis of the infected tissues of zebrafish showed that gene expression of the GCRV-only infected group had a subtle trend change compared to the SVCV-infected group. In addition, a number of genes showed expression trends consistent with SVCV infections, and the expression of these genes may have been more severely altered during the viral co-infection. Genes typified by such changes being concentrated in Gene Ontology (GO) analysis (Supplementary Fig. S5). Although GCRV may exacerbate the perturbation of some genes, inhibition of SVCV replication by GCRV protects the host from more serious damage.
Viral co-infection has been reported in a few cases in land animals (Koh et al., 2019; Jones et al., 2020; Palinski et al., 2022). Besides the reported IAV and SARS-CoV-2 co-infection, co-infection of both adenovirus and SARS-CoV-2 have been detected in some cases of severe hepatitis among young children (Brodin and Arditi, 2022). Thus, the cause of hepatitis due to co-infection cannot be ruled out, although the exact reason has not been identified. Another type of viral co-infection can be caused by extracellular events. For instance, T-cell immunoglobulin and mucin domain 1 (TIM-1) is the same receptor for both the Zaire ebola virus (EBOV) and the lake victoria marburgvirus (MARV) (Kondratowicz et al., 2011). These findings suggest that the means for co-infection could be supplied by the cells. In our study, we could not exclude the possibility that an unknown receptor on the cellular surface for both SVCV and GCRV was also a reason for the viral interference because few studies about the receptor of these two viruses have reported.
Our data reveal that NS38 degrades SVCV N protein by autophagy-lysosome manner and SVCV P protein by Ub-proteasome pathway. This means that one viral protein of GCRV possesses multiple functions. The possible reason is that RNA viruses exhibit small-sized genomes that only encode a limited set of proteins compared to large DNA viruses, thus their proteins need to be multifunctional (Vidalain and Tangy, 2010). For instance, the N protein of coronavirus is also a multi-tasking protein, playing a critical role in virus replication and various cellular response including cell cycle, cell stress responses, immune system and so on (McBride et al., 2014).
It is widely accepted that K48-linked ubiquitination plays a central role in the proteasome-dependent proteolysis, and K63-linked ubiquitination has been found to be involved in numerous cellular events that rely on proteasome-independent processes. However, emerging in vitro and in vivo studies implied that K63-linked ubiquitination also serves as the proteasomal degradation signal (Kirkpatrick et al., 2006; Kim et al., 2007; Saeki et al., 2009). For instance, Hofmann et al. found that purified 26 S proteasomes could degrade synthetic K63-linked tetra-Ub chains (Hofmann and Pickart, 2001). In accord with these findings, our data revealed that MG132 treatment blocked NS38-mediated attenuation of P protein by decreasing K63-linked ubiquitination. Since NS38 does not have the typical characteristics of an E3 Ub ligase, we believe that it may affect the Ub levels of P proteins through using of Ub ligases that interact with P protein.
It should also be noted that besides NS38, there are several other viral proteins produced by GCRV that suppress SVCV viral transcription, such as S5 and S11. In addition to this, the protein encoded by the S7 even exhibited a facilitative effect on the transcriptional level of SVCV. This suggests that a viral protein may also be used by another virus to promote its replication. In contrast, the overall co-infection of GCRV with SVCV inhibited SVCV proliferation, suggesting that the mechanism behind this phenomenon can be more complicated. Our research focused on the interactions of NS38 and SVCV viral proteins; however, the roles of other GCRV viral proteins on the interference of SVCV proteins also need to be investigated. One possible mechanism of GCRV-SVCV co-infection interference is a similar receptor being targeted by both GCRV and SVCV. In our previous studies, the factors of the IFN signaling pathway: mitochondrial antiviral signaling protein (MAVS) and TANK-binding kinase 1 (TBK1), were inhibited by the SVCV N and P proteins, respectively (Li et al., 2016; Lu et al., 2016). Therefore, if GCRV viral proteins block the inhibition of IFN signaling by SVCV, the result is the suppression of SVCV proliferation. GCRV-II VP41, encoded by the S8 segment, could interact with TBK1 and IRF3 activation mediator (MITA) to evade the host immune response (Lu et al., 2017). We believe that such association between host IFN regulation and viral proteins is present in GCRV-I. However, another modulation mechanism might exist. Besides the apparent protein-protein interaction, other interactions are also crucial and cannot be ignored, such as interactions between proteins and nucleic acids. Whether GCRV viral proteins regulate SVCV transcription is still unclear. However, transient protein interactions might be happening, for example, phosphorylation. This interaction is an important but quick biologic process and is easily missed by regular experimental procedures such as Co-IP. Overall, previous reports described viral co-infection but offered little insight into the mechanisms. Our study demonstrates that intracellular interactions are crucial in viral co-infection interference; however, more studies are needed to elucidate all the possible co-infection interactions.
5. Conclusions
The complex viral environment of water enhances the co-infection of fish, while the mechanisms of animal co-infection are still unclear. Here, we found the GCRV infection significantly repressed the influence of SVCV infection in the fish, and the host tissue damage and transcriptome changes caused by the SVCV were substantially defused when co-infected. Further, the study suggested that GCRV NS38 interacted with and degraded SVCV N and P proteins. The N protein was degraded through autophagy by NS38, and p62 was pivotal in this degradation. The P protein was degraded by NS38 via decreasing K63-linked ubiquitination. These data bring to light the intracellular viral protein interactions in co-infection.
Data availability
All of the data supporting this study are available within the manuscript or upon request to the corresponding authors.
Ethics statement
The in vivo experiments of zebrafish described in this study were conducted at the Institute of Hydrobiology (IHB), Chinese Academy of Sciences (CAS) according to European Union guidelines for handling of laboratory animals (2010/63/EU). The whole study was in compliance with all ethical regulations and was approved by the Committee on the Ethics of Animal Experiments of the IHB, CAS (No. 2019-040).
Author contributions
Zhuo-Cong Li: conceptualization, methodology, software, data curation, writing-original draft preparation. Long-Feng Lu: conceptualization, methodology, software, data curation, writing-original draft preparation. Can Zhang: visualization, investigation. Xue-Li Wang: software, validation. Jing-Feng Tong: visualization, investigation. Ke-Jia Han: software, validation. Dan-Dan Chen: supervision. Xi-Yin Li: supervision. Zhou Li: supervision. Jian-Fang Gui: writing-reviewing and editing. Shun Li: conceptualization, methodology, software, writing-reviewing and editing.
Conflict of interest
The authors have no financial conflicts of interest.
Acknowledgments
We thank Fang Zhou (Analysis and Testing Center, IHB, CAS) for assistance with confocal microscopy analysis and Dr. Feng Xiong (China Zebrafish Resource Center, IHB, CAS) for assistance with qPCR analysis. We thank Xin Wang and Guang-Xin Wang (Analysis and Testing Center, IHB, CAS) for assistance with Histopathology. We thank Yuan Xiao and Zhen-Fei Xing (Analysis and Testing Center, IHB, CAS) for assistance with TEM observation. This work was supported by the Strategic Priority Research Program of the Chinese Academy of Sciences (XDA24030203), the National Key Research and Development Program of China (2018YFD0900504, 2018YFD0900204, and 2021YFD1200804), the National Natural Science Foundation of China (32073009, 31873036, 32173023, and 32002431), the Youth Innovation Promotion Association, the Key Program of Frontier Sciences of the Chinese Academy of Sciences (QYZDY-SSW-SMC025), the China Agriculture Research System of MOF and MARA (CARS-45-07), and the Autonomous Project of the State Key Laboratory of Freshwater Ecology and Biotechnology.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.virs.2022.12.003.
Appendix A. Supplementary data
The following are the Supplementary data to this article:
References
- Arora U., Garg P., Agarwal S., Nischal N., Shalimar Wig N. Complexities in the treatment of coinfection with hiv, hepatitis b, hepatitis c, and tuberculosis. Lancet Infect. Dis. 2021;21:e399–e406. doi: 10.1016/S1473-3099(20)30765-9. [DOI] [PubMed] [Google Scholar]
- Arzt J., Fish I.H., Bertram M.R., Smoliga G.R., Hartwig E.J., Pauszek S.J., Holinka-Patterson L., Diaz-San Segundo F.C., Sitt T., Rieder E., Stenfeldt C. Simultaneous and staggered foot-and-mouth disease virus coinfection of cattle. J. Virol. 2021;95 doi: 10.1128/JVI.01650-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bai L., Zhao Y., Dong J., Liang S., Guo M., Liu X., Wang X., Huang Z., Sun X., Zhang Z., Dong L., Liu Q., Zheng Y., Niu D., Xiang M., Song K., Ye J., Zheng W., Tang Z., Tang M., Zhou Y., Shen C., Dai M., Zhou L., Chen Y., Yan H., Lan K., Xu K. Coinfection with influenza a virus enhances sars-cov-2 infectivity. Cell Res. 2021;31:395–403. doi: 10.1038/s41422-021-00473-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brodin P., Arditi M. Severe acute hepatitis in children: investigate sars-cov-2 superantigens. Lancet Gastroenterol. Hepatol. 2022;7:594–595. doi: 10.1016/S2468-1253(22)00166-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bruhn J.F., Hotard A.L., Spiropoulou C.F., Lo M.K., Saphire E.O. A conserved basic patch and central kink in the nipah virus phosphoprotein multimerization domain are essential for polymerase function. Structure. 2019;27:660–668 e664. doi: 10.1016/j.str.2019.01.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deng Z., Li X., Blanca Ramirez M., Purtell K., Choi I., Lu J.H., Yu Q., Yue Z. Selective autophagy of akap11 activates camp/pka to fuel mitochondrial metabolism and tumor cell growth. Proc. Natl. Acad. Sci. U. S. A. 2021;118 doi: 10.1073/pnas.2020215118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deng Z.Q., Purtell K., Lachance V., Wold M.S., Chen S., Yue Z.Y. Autophagy receptors and neurodegenerative diseases. Trends Cell Biol. 2017;27:491–504. doi: 10.1016/j.tcb.2017.01.001. [DOI] [PubMed] [Google Scholar]
- Domingos J.A., Shen X., Terence C., Senapin S., Dong H.T., Tan M.R., Gibson-Kueh S., Jerry D.R. Scale drop disease virus (sddv) and lates calcarifer herpes virus (lchv) coinfection downregulate immune-relevant pathways and cause splenic and kidney necrosis in barramundi under commercial farming conditions. Front. Genet. 2021;12 doi: 10.3389/fgene.2021.666897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ernst J., Bar-Joseph Z. Stem: a tool for the analysis of short time series gene expression data. BMC Bioinf. 2006;7:191. doi: 10.1186/1471-2105-7-191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Griffiths E.C., Pedersen A.B., Fenton A., Petchey O.L. The nature and consequences of coinfection in humans. J. Infect. 2011;63:200–206. doi: 10.1016/j.jinf.2011.06.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gul-Seker M., Elibuyuk I.O. Occurrence of tomato yellow leaf curl virus and tomato chlorosis virus mixed infections in protected tomato plants, antalya, Turkey. Phytoparasitica. 2019;47:441–449. [Google Scholar]
- Gupta N., Rao P.V. Transcriptomic profile of host response in Japanese encephalitis virus infection. Virol. J. 2011;8:92. doi: 10.1186/1743-422X-8-92. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hofmann R.M., Pickart C.M. In vitro assembly and recognition of lys-63 polyubiquitin chains. J. Biol. Chem. 2001;276:27936–27943. doi: 10.1074/jbc.M103378200. [DOI] [PubMed] [Google Scholar]
- Hoque M.N., Rahman M.S., Ahmed R., Hossain M.S., Islam M.S., Islam T., Hossain M.A., Siddiki A.Z. Diversity and genomic determinants of the microbiomes associated with covid-19 and non-covid respiratory diseases. Gene Rep. 2021;23 doi: 10.1016/j.genrep.2021.101200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoshino T., Nishima D., Enseki M., Umehara N., Fukasawa C., Ishiwada N. Pediatric parapneumonic effusion/pleural empyema in Japan: a nationwide survey. Pediatr. Infect. Dis. J. 2022;41:20–23. doi: 10.1097/INF.0000000000003353. [DOI] [PubMed] [Google Scholar]
- Jones B.A., Mahapatra M., Chubwa C., Clarke B., Batten C., Hicks H., Henstock M., Keyyu J., Kock R., Parida S. Characterisation of peste des petits ruminants disease in pastoralist flocks in ngorongoro district of northern Tanzania and bluetongue virus co-infection. Viruses. 2020;12:389. doi: 10.3390/v12040389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim H.T., Kim K.P., Lledias F., Kisselev A.F., Scaglione K.M., Skowyra D., Gygi S.P., Goldberg A.L. Certain pairs of ubiquitin-conjugating enzymes (e2s) and ubiquitin-protein ligases (e3s) synthesize nondegradable forked ubiquitin chains containing all possible isopeptide linkages. J. Biol. Chem. 2007;282:17375–17386. doi: 10.1074/jbc.M609659200. [DOI] [PubMed] [Google Scholar]
- Kirkpatrick D.S., Hathaway N.A., Hanna J., Elsasser S., Rush J., Finley D., King R.W., Gygi S.P. Quantitative analysis of in vitro ubiquitinated cyclin b1 reveals complex chain topology. Nat. Cell Biol. 2006;8:700–710. doi: 10.1038/ncb1436. [DOI] [PubMed] [Google Scholar]
- Koh C., Da B.L., Glenn J.S. Hbv/hdv coinfection: a challenge for therapeutics. Clin. Liver Dis. 2019;23:557–572. doi: 10.1016/j.cld.2019.04.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kondratowicz A.S., Lennemann N.J., Sinn P.L., Davey R.A., Hunt C.L., Moller-Tank S., Meyerholz D.K., Rennert P., Mullins R.F., Brindley M., Sandersfeld L.M., Quinn K., Weller M., McCray P.B., Chiorini J., Maury W. T-cell immunoglobulin and mucin domain 1 (tim-1) is a receptor for zaire ebolavirus and lake victoria marburgvirus. Proc. Natl. Acad. Sci. U. S. A. 2011;108:8426–8431. doi: 10.1073/pnas.1019030108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kotob M.H., Menanteau-Ledouble S., Kumar G., Abdelzaher M., El-Matbouli M. The impact of co-infections on fish: a review. Vet. Res. 2016;47:98. doi: 10.1186/s13567-016-0383-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Langford B.J., So M., Raybardhan S., Leung V., Westwood D., MacFadden D.R., Soucy J.R., Daneman N. Bacterial co-infection and secondary infection in patients with covid-19: a living rapid review and meta-analysis. Clin. Microbiol. Infect. 2020;26:1622–1629. doi: 10.1016/j.cmi.2020.07.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li S., Lu L.F., Wang Z.X., Lu X.B., Chen D.D., Nie P., Zhang Y.A. The p protein of spring viremia of carp virus negatively regulates the fish interferon response by inhibiting the kinase activity of tank-binding kinase 1. J. Virol. 2016;90:10728–10737. doi: 10.1128/JVI.01381-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li S., Lu L.F., Liu S.B., Zhang C., Li Z.C., Zhou X.Y., Zhang Y.A. Spring viraemia of carp virus modulates p53 expression using two distinct mechanisms. PLoS Pathog. 2019;15 doi: 10.1371/journal.ppat.1007695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu L.F., Li S., Wang Z.X., Du S.Q., Chen D.D., Nie P., Zhang Y.A. Grass carp reovirus vp41 targets fish mita to abrogate the interferon response. J. Virol. 2017;91 doi: 10.1128/JVI.00390-17. 003900-e417. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu L.F., Li S., Lu X.B., LaPatra S.E., Zhang N., Zhang X.J., Chen D.D., Nie P., Zhang Y.A. Spring viremia of carp virus n protein suppresses fish ifnphi1 production by targeting the mitochondrial antiviral signaling protein. J. Immunol. 2016;196:3744–3753. doi: 10.4049/jimmunol.1502038. [DOI] [PubMed] [Google Scholar]
- Lu L.F., Zhang C., Li Z.C., Zhou X.Y., Jiang J.Y., Chen D.D., Zhang Y.A., Xiong F., Zhou F., Li S. A novel role of zebrafish tmem33 in negative regulation of interferon production by two distinct mechanisms. PLoS Pathog. 2021;17 doi: 10.1371/journal.ppat.1009317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McBride R., van Zyl M., Fielding B.C. The coronavirus nucleocapsid is a multifunctional protein. Viruses-Basel. 2014;6:2991–3018. doi: 10.3390/v6082991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McCullers J.A. The co-pathogenesis of influenza viruses with bacteria in the lung. Nat. Rev. Microbiol. 2014;12:252–262. doi: 10.1038/nrmicro3231. [DOI] [PubMed] [Google Scholar]
- Nickbakhsh S., Mair C., Matthews L., Reeve R., Johnson P.C.D., Thorburn F., von Wissmann B., Reynolds A., McMenamin J., Gunson R.N., Murcia P.R. Virus-virus interactions impact the population dynamics of influenza and the common cold. Proc. Natl. Acad. Sci. U. S. A. 2019;116:27142–27150. doi: 10.1073/pnas.1911083116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishitha Y., Priyanka E., Vamshi Krishna S., Kannaki T.R. Co-infection of marek's disease virus with different oncogenic immunosuppressive viruses in chicken flocks. Virusdisease. 2021;32:804–809. doi: 10.1007/s13337-021-00731-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Palinski R.M., Brito B., Jaya F.R., Sangula A., Gakuya F., Bertram M.R., Pauszek S.J., Hartwig E.J., Smoliga G.R., Obanda V., Omondi G.P., VanderWaal K., Arzt J. Viral population diversity during co-infection of foot-and-mouth disease virus serotypes sat1 and sat2 in african buffalo in Kenya. Viruses. 2022;14:897. doi: 10.3390/v14050897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pei C., Ke F., Chen Z.Y., Zhang Q.Y. Complete genome sequence and comparative analysis of grass carp reovirus strain 109 (gcrev-109) with other grass carp reovirus strains reveals no significant correlation with regional distribution. Arch. Virol. 2014;159:2435–2440. doi: 10.1007/s00705-014-2007-5. [DOI] [PubMed] [Google Scholar]
- Rosenberg B.R., Depla M., Freije C.A., Gaucher D., Mazouz S., Boisvert M., Bedard N., Bruneau J., Rice C.M., Shoukry N.H. Longitudinal transcriptomic characterization of the immune response to acute hepatitis c virus infection in patients with spontaneous viral clearance. PLoS Pathog. 2018;14 doi: 10.1371/journal.ppat.1007290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saeki Y., Kudo T., Sone T., Kikuchi Y., Yokosawa H., Toh-e A., Tanaka K. Lysine 63-linked polyubiquitin chain may serve as a targeting signal for the 26s proteasome. EMBO J. 2009;28:359–371. doi: 10.1038/emboj.2008.305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simkova A., Civanova K., Vetesnik L. Heterosis versus breakdown in fish hybrids revealed by one-parental species-associated viral infection. Aquaculture. 2022;546 [Google Scholar]
- Sulkowski M.S. Viral hepatitis and hiv coinfection. J. Hepatol. 2008;48:353–367. doi: 10.1016/j.jhep.2007.11.009. [DOI] [PubMed] [Google Scholar]
- Tao M.M., Liu T., You Q.D., Jiang Z.Y. P62 as a therapeutic target for tumor. Eur. J. Med. Chem. 2020;193 doi: 10.1016/j.ejmech.2020.112231. [DOI] [PubMed] [Google Scholar]
- Tso C.H., Lu M.W. Transcriptome profiling analysis of grouper during nervous necrosis virus persistent infection. Fish Shellfish Immunol. 2018;76:224–232. doi: 10.1016/j.fsi.2018.03.009. [DOI] [PubMed] [Google Scholar]
- Vidalain P.O., Tangy F. Virus-host protein interactions in rna viruses. Microb. Infect. 2010;12:1134–1143. doi: 10.1016/j.micinf.2010.09.001. [DOI] [PubMed] [Google Scholar]
- Wang D., Mai J., Yang Y., Xiao C.T., Wang N. Current knowledge on epidemiology and evolution of novel porcine circovirus 4. Vet. Res. 2022;53:38. doi: 10.1186/s13567-022-01053-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolf Y.I., Silas S., Wang Y., Wu S., Bocek M., Kazlauskas D., Krupovic M., Fire A., Dolja V.V., Koonin E.V. Doubling of the known set of rna viruses by metagenomic analysis of an aquatic virome. Nat. Microbiol. 2020;5:1262–1270. doi: 10.1038/s41564-020-0755-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu L.M., Liu M., Zhao J.Z., Ren G.M., Dong Y., Shao Y.Z., Lu T.Y., Zhang Q.Y. Infectious pancreatic necrosis virus inhibits infectious hematopoietic necrosis virus at the early stage of infection in a time dependent manner during co-infection in chinook salmon embryo cell lines. Fish Shellfish Immunol. 2020;102:361–367. doi: 10.1016/j.fsi.2020.05.010. [DOI] [PubMed] [Google Scholar]
- Xu T., Hou C.Y., Zhang Y.H., Li H.X., Chen X.M., Pan J.J., Chen H.Y. Simultaneous detection and genetic characterization of porcine circovirus 2 and 4 in henan province of China. Gene. 2022;808 doi: 10.1016/j.gene.2021.145991. [DOI] [PubMed] [Google Scholar]
- Zhang F., Zhu H., Wu Y., Dou Z., Zhang Y., Kleinman N., Bulterys M., Wu Z., Ma Y., Zhao D., Liu X., Fang H., Liu J., Cai W.P., Shang H. Hiv, hepatitis b virus, and hepatitis c virus co-infection in patients in the China national free antiretroviral treatment program, 2010-12: a retrospective observational cohort study. Lancet Infect. Dis. 2014;14:1065–1072. doi: 10.1016/S1473-3099(14)70946-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Q.Y., Gui J.F. Virus genomes and virus-host interactions in aquaculture animals. Sci. China Life Sci. 2015;58:156–169. doi: 10.1007/s11427-015-4802-y. [DOI] [PubMed] [Google Scholar]
- Zhang Q.Y., Gui J.F. Diversity, evolutionary contribution and ecological roles of aquatic viruses. Sci. China Life Sci. 2018;61:1486–1502. doi: 10.1007/s11427-018-9414-7. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All of the data supporting this study are available within the manuscript or upon request to the corresponding authors.