Abstract
Despite an abundance of gene expression surveys, comparatively little is known about Hox gene function in Chelicerata. Previous investigations of paralogs of labial (lab) and Deformed (Dfd) in a spider have shown that these play a role in tissue maintenance of the pedipalp segment (lab-1) and in patterning the first walking leg identity (Dfd-1), respectively. However, extrapolations of these data across chelicerates are hindered by the existence of duplicated Hox genes in arachnopulmonates (e.g., spiders and scorpions), which have resulted from an ancient whole genome duplication (WGD) event. Here, we investigated the function of the single-copy ortholog of lab in the harvestman Phalangium opilio, an exemplar of a lineage that was not subject to this WGD. Embryonic RNA interference against lab resulted in two classes of phenotypes: homeotic transformations of pedipalps to chelicerae, as well as reduction and fusion of the pedipalp and leg 1 segments. To test for combinatorial function, we performed a double knockdown of lab and Dfd, which resulted in a homeotic transformation of both pedipalps and the first walking legs into cheliceral identity, whereas the second walking leg is transformed into a pedipalpal identity. Taken together, these results elucidate a model for the Hox logic of head segments in Chelicerata. To substantiate the validity of this model, we performed expression surveys for lab and Dfd paralogs in scorpions and horseshoe crabs. We show that repetition of morphologically similar appendages is correlated with uniform expression levels of the Hox genes lab and Dfd, irrespective of the number of gene copies.
Keywords: Arthropoda, serial homology, Opiliones, tritocerebrum, Xiphosura, Hox1
Introduction
The study of body plan evolution has a rich history, punctuated by advances in comparative embryology, genomics, and functional approaches. Arthropod body plan evolution has served as a particularly fertile proving ground for numerous concepts in evolutionary developmental biology, such as tests of serial homology, cooption of regulatory networks, the evolution of cis-regulatory elements, and developmental systems drift (e.g. Gompel et al. 2005; Jager et al. 2006; Oliver et al. 2014; Auman and Chipman 2018; Setton and Sharma 2018). Among these central contributions is the mechanistic understanding of arthropod Hox genes, a family of conserved transcription factors that play a role in regionalizing the antero-posterior axis across Bilateria. Numerous expression surveys have highlighted conserved and labile aspects of Hox expression domains across the arthropod tree of life, and advances in functional toolkits for emerging model species have pinpointed both canonical and non-canonical roles of these genes in body plan patterning (e.g., Abzhanov and Kaufman 1999; Hughes and Kaufman 2002a; Brena et al. 2006; Mahfooz et al. 2007; Medved et al. 2015; Martin et al. 2016). However, the availability of functional datasets for Hox genes is highly asymmetrical, with much of what is known about anterior Hox genes stemming from pancrustacean (and particularly, insect) datasets. Whereas fundamentals of Hox gene dynamics are well understood in pancrustacean models like the fruit fly Drosophila melanogaster and the amphipod Parhyale hawaiensis, the functions of their homologs remain poorly understood in Chelicerata (e.g., sea spiders, horseshoe crabs, arachnids), the sister group to the remaining arthropods.
The bauplan of Chelicerata—and specifically, of arachnids—generally consists of two tagmata, the anterior prosoma (which bears the mouthparts and walking legs) and the posterior opisthosoma (which bears modified appendages or may lack appendages altogether in different groups) (Sharma, Schwager, et al. 2014). The mouthparts of chelicerates typically consist of a pair of chelicerae, innervated by the deutocerebrum (middle region of the tripartite brain; Damen et al. 1998; Telford and Thomas 1998); and a pair of pedipalps, innervated by the tritocerebrum (posterior region of the tripartite brain). Posterior to these are segments that bear four pairs of legs, which are variably modified across chelicerate diversity. Modifications of this general architecture are found in Pycnogonida (sea spiders), which lack an opisthosoma and may bear additional leg pairs (e.g., the ovigers; 10- and 12-legged species in some genera) (Arnaud and Bamber 1988; Ballesteros et al. 2021). Additionally, Xiphosura (the horseshoe crabs) exhibit anatomically identical pedipalps and walking legs with respect to the number of podomeres (leg segments) (Snodgrass 1952; Shultz 1989); only the last pair of walking legs (the pusher leg) exhibits a distinct morphology, owing to its larger size, leaf-like terminal ornamentation, and the presence of the flabellum (an exite; see discussion in [Bruce 2021]). Whereas the pedipalp (tritocerebral appendage) of Xiphosura is modified in adult males (comparable to the pedipalps of spiders or the third walking legs of Ricinulei), in females, this appendage is indistinguishable from the walking legs in the three immediately posterior segments, barring minor differences in size. As a result, Xiphosura is variably described in the literature as having pedipalps and four pairs of legs, or as having five pairs of legs (i.e., segmental homonomy) (Snodgrass 1952; Shultz 1989).
Functional datasets informing the patterning of chelicerate prosomal segments have long remained fragmentary. In the spider Parasteatoda tepidariorum, it was previously shown that one copy of the Hox gene labial (termed labial-1) is necessary for the maintenance of the pedipalp and L1 segment; maternal RNA interference (RNAi) against this gene resulted in the reduction or complete deletion of the tritocerebral segment (and occasionally, also the L1 segment; Pechmann et al. 2015). The implied function of segment maintenance is contrary to the broadly understood canonical role of Hox genes as drivers of homeosis. By contrast, RNAi against a copy of Deformed (termed Deformed-1; abbr. Dfd-1) in the same species resulted in a homeotic transformation of the first walking leg into a pedipalp identity, as inferred from both the morphology of the transformed appendage and the increased expression of lab-1 in the ectopic pedipalp (Pechmann et al. 2015). No functional outcomes were reported from RNAi against the paralogs of each gene (i.e., spider lab-2 and Dfd-2). The duplicates of these Hox genes are attributable to a shared genome duplication in the common ancestor of Arachnopulmonata, a group of six orders that includes spiders, scorpions, and pseudoscorpions (Sharma, Kaluziak, et al. 2014; Schwager et al. 2017; Ontano et al. 2021). Genomic datasets have shown that nearly all 20 Hox duplicates have been retained across arachnopulmonate lineages, paralleling the evolutionary history of the vertebrates (Leite et al. 2018; Gainett and Sharma 2020; Harper et al. 2021; Ontano et al. 2021).
However, as these duplicates are restricted to a subset of arachnid orders, it is not clear how well their dynamics reflect the ancestral condition of single-copy patterning genes, particularly given the possibility of subfunctionalization or neofunctionalization of new gene copies in arachnopulmonate exemplars (Sharma, Schwager, et al. 2014; Turetzek et al. 2015, 2017; Benton et al. 2016). In addition, their incidence and retention in models like P. tepidariorum raise the specter of functional redundancy and compensatory effects between paralogs in RNAi experiments. For this reason, functional datasets from chelicerate taxa that did not undergo genome duplication events are especially valuable for comparison. Accordingly, recent comparative studies have focused on the unprepossessing harvestman species Phalangium opilio, which exhibits an unduplicated genome and is amenable to gene silencing techniques (Sharma et al. 2012a, 2012b, 2013; Gainett et al. 2021). Functional datasets in the harvestman P. opilio have shown that the single-copy Dfd ortholog is required for patterning the identity of the first two pairs of walking legs; knockdown of Dfd resulted in the homeotic transformation of both L1 and L2 legs (first and second walking leg pairs) into pedipalps. Whereas RNAi against Sex combs reduced (Scr) had no effect by itself, double knockdown of Dfd and Scr resulted in homeotic transformations of L1–L3 legs toward pedipalp identity (Gainett et al. 2021). To date, no other functional data are available for the Hox genes of this species.
Here, we focused on discovering the functions of single-copy orthologs of anterior Hox genes in P. opilio, with the aim of understanding the Hox logic of the arachnid head segments. We show that knockdown of lab results in homeotic transformation of the pedipalp into cheliceral identity, whereas double knockdown of both lab and Dfd transforms both pedipalps and first walking legs into chelicerae. Having established a ground plan for the patterning of segmental identities in the harvestman, we explored the expression dynamics of lab and Dfd paralogs in a scorpion and a horseshoe crab, as points of comparison to spider and harvestman Hox genes. This survey substantiates a correlation between heterogeneity of Hox expression levels and the morphological disparity of prosomal appendages.
Results
Expression of Harvestman labial
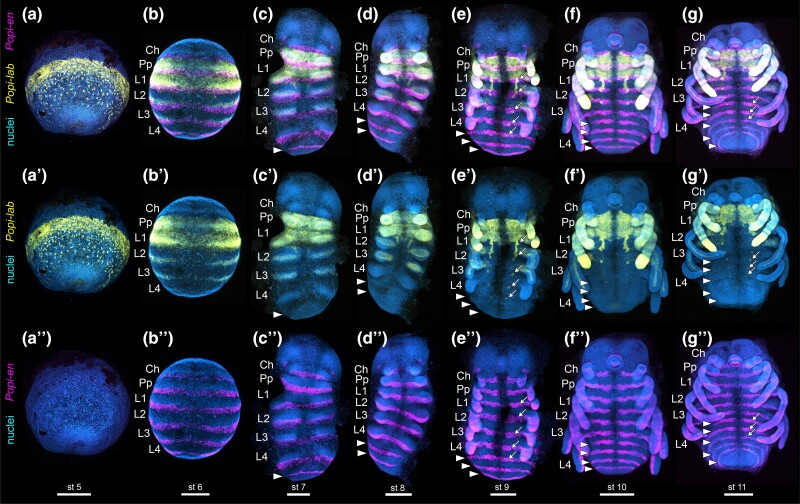
The single copy Popi-labial (Popi-lab) is expressed as a ring in the equator of the embryo prior to the formation of the germ band, and in dispersed cells towards the center of the encircled labial domain (fig. 1a–a″). In the stage where the antero-posterior axis forms and the anterior tagma (prosoma) becomes segmented, strong expression localizes to the entire tritocerebral and L1 segment, as marked by the segment polarity gene Popi-en (fig. 1b–b″). Popi-lab is also expressed in the posterior part of the L2 and L3 segments (fig. 1b). During the formation of limb buds, Popi-lab is strongly expressed in the ectoderm of the pedipalp and L1 limb buds, and in the mesoderm of L2 and L3 nascent limb buds (fig. 1c–d″). With the sequential addition of posterior body segments, additional paired dots of expression appear in sequence adjacent to the ventral midline (fig. 1e–g″, arrows). Two antero-posterior stripes of expression occur in the ventral ectoderm of the L2 segment (fig. 1c–g″). A sharp anterior boundary of expression in the tritocerebral segment, as well as strong expression in the tritocerebral and L1 appendages and in the ventral ectoderm of these segments, are maintained throughout the stages investigated (fig. 1c–g″).
Fig. 1.
Wild type expression of Popi-lab and Popi-en across embryonic stages. Anterior at top. (a) Wholemount, ventrolateral view of the germ disc. (b) Wholemount, ventral view. (c–g): Flatmount, ventral view. Nuclei in blue (Hoechst). Popi-en: magenta. Popi-lab: yellow. (a–g): Hoechst, Popi-lab and Popi-en multiplexed. (a′–g′): Hoechst and Popi-lab, same embryos as previous row. (a″–g″): Hoechst and Popi-en, same embryos as previous row. Arrowheads: Popi-en stripes marking the posterior of each segment. Arrows: dots of Popi-lab expression adjacent to the ventral midline. Ch, chelicera; Pp, pedipalp; L1–L4, L1–L4 legs; st, stage number. Scale bars: 200 µm.
Knockdown of lab Results in Homeotic Pedipalp-to-Chelicera Transformations and Defects in the Tritocerebral Segment
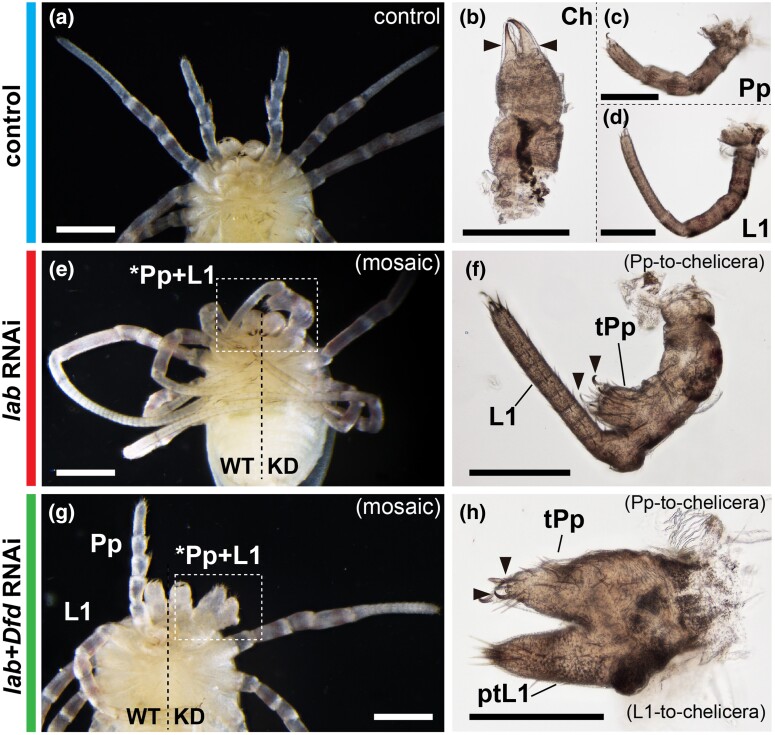
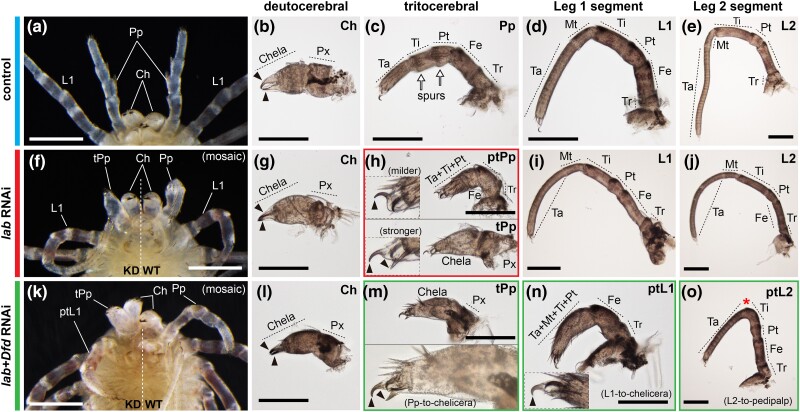
To investigate the function of Popi-lab, we conducted RNAi via embryonic injections of double-stranded RNA (dsRNA). Popi-lab RNAi hatchlings presented a spectrum of defects affecting the tritocerebral segment. The chelicera and leg identities were unaffected in hatchlings, whereas the pedipalps were transformed into cheliceral identity (n = 35/74) (fig. 2a–j; figs. S1 and S2, Supplementary material online), as evidenced by a proximal podomere and a distal podomere with two claws (chela) (fig. 2b, g, and l). Milder transformation consisted of partial truncation of the pedipalp podomeres and unaffected claw (fig. 2h, insets), whereas stronger homeosis consisted of a tritocerebral appendage with proximal podomere and an ectopic claw in the distal podomere (fig. 2h, insets). We note that the most distal part of the transformed appendages did not completely transform into a chela, as the claws retained an identity similar to the pedipalp and leg claws. Homeosis in the embryonic appendages was evidenced by the branching and reduced length of the tritocerebral limb (figs. S1a and b, Supplementary Material online).
Fig. 2.
Popi-lab knockdown results in homeotic transformations. Brightfield images of hatchlings (postembryos) of control (dH2O-injected) (a–e), Popi-lab RNAi (f–j), and Popi-lab + Popi-Dfd RNAi (k–o) experiments. (a), (f) and (k) are ventral views of the anterior prosoma. Other panels are dissected appendages in lateral view with proximal at right. (b, g, and l): Deutocerebral appendage (chelicera). (c, h, and m): Tritocerebral appendage. (d, i, and n): Appendage of the L1 segment. (e, j, and o): Appendage of the L2 segment. Note that because the first instar's cuticle is already secreted and visible inside the postembryo cuticle, two sets of claws are discernible in some panels. Black arrowhead, claw; red asterisk, missing metatarsus (Mt); white arrows, pedipalpal spurs; Ch, chelicera; Pp, pedipalp; tPp, transformed pedipalp; ptPp, partially transformed pedipalp; L1–L2, L1–L2 appendages; ptL1–ptL2, partially transformed L1–L2 appendages; Ta, tarsus; Mt, metatarsus; Ti, tibia; Pt, patella; Fe, femur; Tr, trochanter; Px, proximal cheliceral segment; WT, wild type side of individual; KD, knockdown side of individual. Scale bars: 200 µm.
In addition to homeosis, a subset of Popi-lab RNAi hatchlings (11%; n = 4/35) exhibited defects in the segment boundaries of the tritocerebral segment, as evidenced by the fusion of the tritocerebral appendage to either the chelicera or to L1 (fig. 3a–f; fig. S3, Supplementary Material online). Popi-lab RNAi embryos assayed for the segmentation marker Popi-engrailed (Popi-en) revealed a reduction in the tritocerebral and L1 segment relative to other prosomal segments, and juxtaposition of consecutive Popi-en expression stripes (fig. S3, Supplementary Material online). Taken together, these results suggest that P. opilio labial is necessary for both the specification of the tritocerebral segment and for the identity of the pedipalp.
Fig. 3.
Popi-lab knockdown results in fusion of the pedipalp and L1 leg. Brightfield images of hatchlings (postembryos) of control (dH2O-injected) (a–d), Popi-lab RNAi (e–f), and Popi-lab + Popi-Dfd RNAi (g–h) experiments. Phenotypes in (g–h) present both homeosis and fusion. (a, e, and g) are ventral views of the anterior prosoma. Other panels are dissected appendages in lateral view. Mosaic individuals in (e) and (g) are affected on the right side of the picture. Black arrowhead, claw; Ch, chelicera; Pp, pedipalp; tPp, transformed pedipalp; L1, L1 appendage; ptL1, partially transformed L1 leg; WT, wildtype side of individual; KD, knockdown side of individual. Scale bars: 200 µm.
Double Knockdown of lab + Dfd Results in Transformations of Pedipalps and L1 Legs into Chelicerae
While RNAi against lab results in the transformation to cheliceral identity in the pedipalp segment, we observed that the identity of L1 was not comparably affected, despite the presence and maintenance of lab transcript abundance in L1 territory in the wild type condition. We postulated that the absence of homeotic function in L1 may be attributable to functional redundancy with another anteriorly expressed Hox gene such as Dfd and proboscipedia (pb), though the latter gene is only expressed in subdomains of the prosomal appendages (Sharma et al. 2012a). To test this possibility, we conducted RNAi against proboscipedia (pb) and double RNAi against Popi-lab and Popi-Dfd. Preliminary efforts with RNAi against pb resulted in no observable phenotype (data not shown); this outcome, together with the restriction of pb expression to only some of the tissue from the pedipalp through the L4 segment (Sharma et al. 2012a), suggested that pb may not act as a canonical Hox gene in P. opilio.
Double RNAi against Popi-lab + Popi-Dfd resulted in homeotic transformations (n = 43/76) affecting tritocerebral, L1 and L2 segments (fig. 2k–o). Fusions of the tritocerebral appendage with chelicera or L1 leg (n = 9/76) were also observed, accompanied by homeosis in the pedipalp, L1 leg, or both appendages (fig. 3g and h). A subset of the hatchlings (n = 22/43) exhibit an additive phenotype, that is, the combination of both pedipalp-to-chelicera (as in single Popi-lab RNAi) and leg-to-pedipalp (as in single Popi-Dfd RNAi [Gainett et al. 2021]), as evidenced by the truncation of the tritocerebral appendage, and loss of the metatarsus in L1 leg. Notably, the effect in the legs in these cases is similar to the weak phenotypes of single Popi-Dfd RNAi (Gainett et al. 2021), with a partial transformation of L1 and L2 legs into pedipalps. A smaller subset of hatchlings exhibited either a single Popi-lab RNAi phenotype (1/43) or a single Popi-Dfd RNAi (3/43) phenotype (see supplementary fig. S2, Supplementary Material online).
A new synergistic phenotype was also observed (17/43), in which both pedipalp-to-chelicera and the L1 leg-to-chelicera homeosis occurred (fig. 2k–o; supplementary fig. S2, Supplementary Material online). Notably, we observed only partial L2 leg-to-pedipalp transformations, as evidenced by the lack of a metatarsus and reduced number of tarsomeres (fig. 2o).
These results suggest that both lab and Dfd repress cheliceral identity, and are consistent with a posterior prevalence model wherein lab is required for patterning pedipalpal identity, and Dfd for patterning leg identity.
Expression of lab and Dfd Duplicates in a Scorpion and a Horseshoe Crab
A notable complexity in chelicerate Hox cluster evolution is the occurrence of paralogs incurrent from whole genome duplications (WGDs), which parallels the history of the vertebrates (Wagner et al. 2003; Dehal and Boore 2005). Furthermore, the WGDs in Arachnopulmonata (spiders, scorpions, and four other arachnid orders) and in Xiphosura (horseshoe crabs) are inferred to have occurred independently (Schwager et al. 2017; Shingate, Ravi, Prasad, Tay, Garg, et al. 2020; Ontano et al. 2021). Hox gene copies are of interest in comparative studies because they may undergo subfunctionalization or neofunctionalization, which in turn may correlate with the diversification of body plans (Wagner et al. 2003; Sharma, Schwager, et al. 2014; Schwager et al. 2017). To test the validity of the Hox logic established herein, as well as obtain further insights into the fate of lab paralogs in Chelicerata, we investigated the expression patterns of lab paralogs in the scorpion Centruroides sculpturatus and in the horseshoe crab Limulus polyphemus.
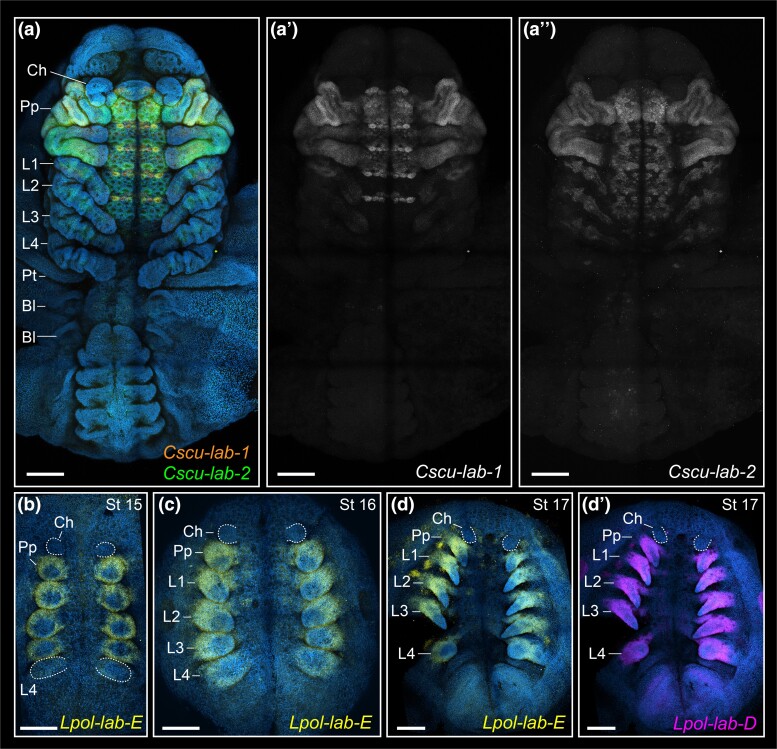
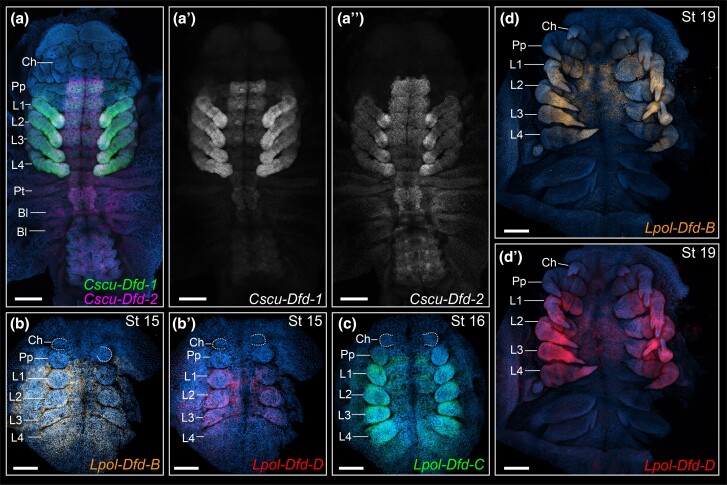
The two labial paralogs of C. sculpturatus (Cscu-lab-1, Cscu-lab-2) have a clear anterior boundary of expression in the tritocerebral segment, and strong expression in the tritocerebral appendage (fig. 4a–a″). The two paralogs have complex patterns of expression on the ventral midline and developing nervous system, with several overlapping domains (fig. 4a–a″). The two Dfd paralogs (Cscu-Dfd-1, Cscu-Dfd-2) have an anterior boundary of expression in the fourth head segment (L1) (fig. 5a–a″). Cscu-Dfd-1 is uniformly expressed in the L1–L4 legs and the ventral neuromeres (fig. 5a–a″). Cscu-Dfd-2 has largely overlapping expression with respect to its paralog; it differs from Cscu-Dfd-1 in that it is more strongly expressed in the tips of the legs, it is slightly more anteriorly expressed in the L1 neuromere, and its expression in the opisthosoma is ubiquitous (fig. 5a–a″). Therefore, the unique identity of the scorpion pedipalp with respect to the legs correlates with the strong expression of both labial paralogs on this appendage, whereas the identity of the four pairs of legs correlates with uniform expression of both Deformed paralogs in all legs.
Fig. 4.
HCR in situ hybridization of labial paralogs in the scorpion C. sculpturatus (a–a″) and the horseshoe crab L. polyphemus (b–d′). Maximum intensity projections of flat mounted embryos in ventral view. Hoechst counter staining in blue. Same letters indicate same embryo (multiplexed). (a) Merged projections of Cscu-lab-1 (orange), Cscu-lab-2 (green). Overlap of lab paralogs appears in yellow. Note that apparent labial expression in the posterior chelicera and labrum is in the tritocerebral tissue; overlapping of these regions is incurring by the maximum intensity projection. (a′) Single channel projection of Cscu-lab-1. (a″) Single channel projection of Cscu-lab-2. (b–d) Lpol-lab-E expression (yellow). (d″) Lpol-lab-D expression (magenta). Ch, chelicera; Pp, pedipalp; L1–L4, L1–L4 legs. Scale bars: 200 µm.
Fig. 5.
HCR in situ hybridization of Deformed paralogs in the scorpion C. sculpturatus (a–a″) and the horseshoe crab L. polyphemus (b–d′). Maximum intensity projections of flat mounted embryos in ventral view. Hoechst counter staining in blue. Same letters indicate same embryo (multiplexed). (a) Merged projections of Cscu-Dfd-1 (green), Cscu-Dfd-2 (magenta). (b) Lpol-Dfd-B expression (orange). (b′) Lpol-Dfd-D expression (red). (c) Lpol-Dfd-C expression (green). (d) Lpol-Dfd-B expression (orange). (d′) Lpol-Dfd-D expression (red). Ch, chelicera; Pp, pedipalp; L1–L4, L1–L4 legs. Scale bars: 200 µm.
Four labial paralogs (Lpol-lab-A, Lpol-lab-B, Lpol-lab-D, Lpol-lab-E) are present in the genome of L. polyphemus (five paralogs occur in Tachypleus gigas and Carcinoscorpius rotundicauda) (Kenny et al. 2015; Shingate, Ravi, Prasad, Tay, Garg, et al. 2020; Shingate, Ravi, Prasad, Tay, and Venkatesh 2020). Lpol-lab-E has an anterior boundary of expression in the tritocerebral segment, being expressed in the tritocerebral appendage and posterior prosomal appendages throughout embryonic stages analyzed (fig. 4b–d). Lpol-lab-D has a largely overlapping expression domain with respect to Lpol-lab-E, with a sharp anterior boundary in the tritocerebral segment (fig. 4d′), but slightly more distally expressed in the appendages than Lpol-lab-E. The short length of Lpol-lab-A CDS (469 bp) precluded reliable detection of transcripts via hybridization chain reaction (HCR) in situ hybridization. We did not analyze the putative homolog Lpol-lab-B, given its unusual sequence and ambiguous annotation (see supplementary Methods, Supplementary Material online).
Five Dfd paralogs occur in L. polyphemus (Lpol-Dfd-A–E). Lpol-Dfd-B, Lpol-Dfd-C, and Lpol-Dfd-D have a clear anterior boundary of expression in the L1 segment (fig. 5b–d′) and are expressed in the L1–L4 legs. The three paralogs have largely overlapping expression domains, with the exception that Lpol-Dfd-B expression extends to the lateral margin of the germ band and more posteriorly into the opisthosoma at the germband stage (fig. 5b). We were not able to detect expression of Lpol-Dfd-A and Lpol-Dfd-E. Therefore, the shared morphology of the tritocerebral appendage and the subsequent prosomal appendage pairs (legs) correlates with a uniform expression of labial paralogs across all post-cheliceral prosomal appendages, and largely overlapping lab and Dfd paralog expression across the L1–L4 legs.
Discussion
labial Function in Arthropoda
In the fruit fly D. melanogaster, lab is expressed in the tritocerebral segment (the intercalary segment in insects, which is reduced and lacks appendages) during embryogenesis and is necessary for normal head involution (Diederich et al. 1989; Merrill et al. 1989). However, there is no reported homeotic function during embryogenesis, which is also the case for a beetle and a milkweed bug (Angelini et al. 2005; Posnien and Bucher 2010; Schaeper et al. 2010). Homeosis in lab mutant flies is only detected in adults, as a shift from head-to-thorax identity of the bristles (Merril et al 1989). This homeosis contrasts with the archetypal transformation towards anterior identity upon disruption of most Hox genes.
By contrast, the tritocerebral segment of non-insect (and non-myriapod) arthropods (i.e., Chelicerata and the crustaceans) bears a pair of appendages that express lab (Telford and Thomas 1998; Abzhanov and Kaufman 1999; Jager et al. 2006; Sharma et al. 2012a; Serano et al. 2016). The only functional data point available for these groups, in the spider P. tepidariorum, demonstrated a role in tritocerebral (and in some cases, L1) tissue maintenance, similarly to insects, but no homeotic function in tritocerebral appendage identity upon lab-1 knockdown (Pechmann et al. 2015). In that work, the knockdown of Dfd-1 was also shown to result in the homeotic transformation of L1 to pedipalps, with the ectopic pedipalps strongly expressing lab-1 (Pechmann et al. 2015). This result is suggestive of the possibility that lab may play a role as a pedipalp determinant, but the evidence is indirect, as homeosis induces transformation of the entire pedipalp program and its associated gene expression patterns.
On the other hand, the spectrum of embryonic defects in P. opilio upon lab RNAi suggests a dual role for this gene, both in tritocerebral and L1 segment specification (segmental fusions in the phenotypic spectrum) and in canonical patterning of segmental identity (homeotic transformations of pedipalpal to cheliceral identity). These data reconcile the gap between the unexpected loss of the tritocerebral segment in spiders upon RNAi against lab-1 and the previously unknown identity of the Hox gene that functions as the pedipalpal selector. It is possible that the lab-1 segmentation phenotype in the spider reflects the severe end of the phenotypic spectrum, as a consequence of the mode and timing of delivery (maternal RNAi). Such a dosage-dependent mechanism could potentially be tested via embryonic RNAi against lab-1 in P. tepidariorum, with the prediction that lower concentrations of dsRNA injected at a later point in embryogenesis may elicit the homeotic phenotype.
More broadly, the significance of the P. opilio lab RNAi phenotype is that it represents the only known case of a homeotic function for lab aside from D. melanogaster. Given the phylogenetic relationship of these two species (a chelicerate and a hexapod), our results suggest that lab may play a role as a conserved tritocerebral selector across Arthropoda.
The Relationship Between Diminution of Hox Expression and Segmental Deletions
Severe loss-of-function phenotypes of lab homologs exhibit tritocerebral segment loss and/or fusion in two chelicerates (the spider P. tepidariorum [Pechmann et al. 2015] and the harvestman P. opilio). Increased cell death in lab loss-of-function phenotypes has been reported in the spider P. tepidariorum (overall in the prosoma) and in the beetle T. castaneum (particularly in the tritocerebral segment in early embryogenesis) (Schaeper et al. 2010; Pechmann et al. 2015), suggesting that labial may directly or indirectly regulate anterior segment specification by controlling apoptosis. Similarly, in D. melanogaster the Hox genes Dfd and Abdominal-B have been shown to promote cell death, which shapes segment boundaries (Lohmann et al. 2002). These observations substantiate the possibility that loss or reduction of a Hox gene may be a potential mechanism for some segmental deletions across Panarthropoda. This inference contrasts with the archetypal role of Hox genes as selector genes that confer segmental identity. Deletion of Hox genes from the Hox cluster has been linked to a segmental reduction in multiple panarthropod taxa, such as acariform mites (loss of segmentation posterior to the second opisthosomal segment, in tandem with loss of abdominal-A (abd-A) [Grbić et al. 2011; Barnett and Thomas 2013]) and tardigrades (loss of a large intermediate region of the antero-posterior axis, in tandem with multiple Hox genes [Smith et al. 2016]). Similarly, in sea spiders, the absence of a segmented opisthosoma has been tentatively linked to atypical patterns of sequence evolution in abd-A (Manuel et al. 2006), but expression patterns of posterior Hox genes are entirely missing in this group (Jager et al. 2006). The mechanism of Hox gene pseudogenization and loss was thought to be released selection in the wake of segmental deletions, resulting in accumulation of mutations and eventual loss of nonfunctional genes associated with the deleted regions—Hox gene loss was understood to reflect the consequence of segmental loss, and not the cause (Barnett and Thomas 2013; Smith et al. 2016).
Nevertheless, it has been recently reported that diminished Ubx and abd-A expression in the ant Camponotus floridanus results in posterior body truncations, which has been linked to a re-wiring of these genes to a lineage-specific function in early germ cell patterning (Rafiqi et al. 2020). In addition, several members of the homeobox gene family are classically known to be key to segmentation across Arthropoda, such as even-skipped and caudal (Hughes and Kaufman 2002b; Mito et al. 2007; Schönauer et al. 2016). Among these are the Hox3 paralog bicoid in cyclorrhaphan flies and fushi tarazu (ftz) in a subset of insects. Indeed, in Arthropoda, evidence for a canonical Hox function is entirely absent for Hox3 (Stauber et al. 1999) and greatly limited for ftz (Löhr et al. 2001). Our results support the possibility that diminution or loss of lab expression may underlie the independent origins of the appendage-free and highly reduced intercalary segment of hexapods and myriapods, as previously articulated by Pechmann et al. (2015). Given the absence of advanced functional toolkits for either the spider or the harvestman, tests of this hypothesis could capitalize upon the availability of genome editing tools in the amphipod crustacean P. hawaiensis (Martin et al. 2016; Serano et al. 2016), with the goal of understanding how fine-tuning of lab expression regulates the transition between a homeotic transformation and a segmental deletion. More broadly, gene silencing tools developed in the tardigrade model Hypsibius exemplaris could be leveraged to test whether a non-canonical role of lab as a segmentation gene is an ancestral feature of Panarthropoda (Tenlen et al. 2013; Smith et al. 2016).
Hox Logic and Evolution of the Chelicerate Prosoma
Leg identity in arachnids has been previously shown to require the expression of Dfd, as knockdown of Dfd results in leg-to-pedipalp homeotic transformation in the daddy-longlegs and in a spider (Pechmann et al. 2015; Gainett et al. 2021), with a redundant role of Scr in the case of L3 (Gainett et al. 2021). Similarly, it was previously shown that homothorax is required in the absence of Hox input for cheliceral identity (Sharma et al. 2015). The RNAi experiments with lab (and both lab and Dfd) presented herein thus enable the completion of a model for anterior prosomal fate specification in the harvestman (fig. 6), wherein cheliceral identity is specified by homothorax in the absence of Hox input; pedipalpal identity requires high levels of lab and low levels of Dfd; and leg identity requires Dfd expression, with a redundant role of Scr at least in the case of the L3 appendage. The genetic basis for L4 identity remains unknown, but is likely attributable to ftz, which is strongly expressed in this territory in the manner of a canonical Hox gene (Sharma et al. 2012a).
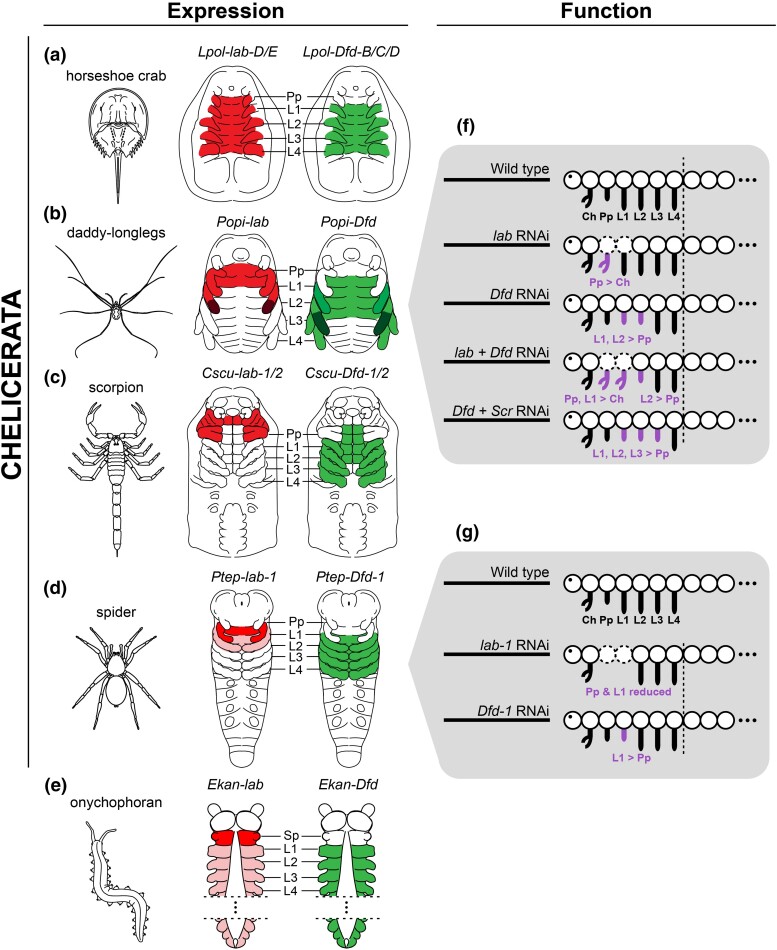
Fig. 6.
Summary of labial (red; left schematics) and Deformed (green; right schematics) expression and functional data in Chelicerata and Onychophora (outgroup). Darker shades reflect concentration of expression. (a) Horseshoe crab L. polyphemus. (b) Daddy-longlegs P. opilio, after Sharma et al. (2012a), and Gainett et al. (2021). (c) Scorpion C. sculpturatus. (d) Spider P. tepidariorum, after Schwager et al. ([2017]). (e) Onychophoran E. kanangrensis, after Eriksson et al (2010) and Janssen et al. (2014). (f) Summary of phenotypic outcomes of Hox RNAi experiments in P. opilio. Dotted body segments indicate segment reduction. Data for the single knockdown of Popi-Dfd and Popi-Scr knockdown after Gainett et al. (2021). (g) Summary of phenotypic outcomes of Ptep-lab-1 and Ptep-Dfd-1 RNAi experiments in P. tepidariorum, after Pechmann et al. (2015). Ch, chelicera; Pp, pedipalp; L1–L4, L1–L4 legs; Sp, slime papilla.
This refined understanding of anterior Hox function in this chelicerate led us to revisit patterns of morphological disparity across the prosomal segments of Chelicerata more broadly. In the harvestman, each prosomal segment exhibits a distinguishable identity; in the walking legs, this is observed as unique counts of tarsomeres (tarsal articles) associated with each walking leg pair. By contrast, the walking legs of spiders and scorpions are morphologically more similar, being mainly distinguished by allometric differences (e.g., walking legs become longer towards the posterior in scorpions; the first pair of walking legs is the longest in most spiders). Yet, like the harvestman, the pedipalps of both spiders and scorpions are clearly morphologically distinct. Horseshoe crabs exhibit another condition still, with no morphological distinction between the tritocerebral appendages (or “pedipalps”) and the posterior three leg pairs, except in adult males (where the tritocerebral appendage is modified into a clasper for reproduction). Only the posterior-most pusher leg has a distinct, unique morphology by comparison to anterior leg pairs in both sexes of horseshoe crabs (Snodgrass 1952).
Our gene expression surveys revealed an intriguing correlation between the presence of a morphologically distinct tritocerebral segment identity (e.g., pedipalp) and the concentration of lab expression in this segment (fig. 6a). The only arthropod surveyed to date that has strong, uniform lab expression on all post-deutocerebral appendages is the horseshoe crab L. polyphemus. This condition correlates with the morphological similarity of their tritocerebral appendage and post-tritocerebral prosomal appendages (i.e., these five appendage pairs have the same number of segments and the first four are indistinguishable in females) (fig. 6a). The only comparable case is the protonymphon larvae of sea spiders (Pycnogonida), where lab is uniformly expressed in the third and fourth head segments, which bear anatomically identical larval appendages at this developmental stage (Jager et al. 2006).
In contrast to the horseshoe crab and sea spider larvae, the lab homologs of all other arthropods with a morphologically distinct tritocerebral identity (second antenna, pedipalps, or intercalary segment) exhibit anteriorly restricted expression domains, with the highest concentration of expression levels in the tritocerebral (and in some cases, also the fourth head) segment (fig. 6b–d). This correlation further holds for the onychophoran Euperipatoides kanangrensis, in which lab is more strongly expressed in the tritocerebral segment and its appendage (and weakly in all posterior segments), correlating with the unique identity of the slime papillae in contrast to posterior lobopods (fig. 6e) (Eriksson et al. 2010; Janssen et al. 2014).
Furthermore, we discovered that Dfd homologs exhibit a similar trend: morphologically similar legs exhibit uniform expression of Deformed. Although Dfd homologs always bore an anterior boundary in the fourth head segment, expression patterns and transcript levels of Dfd copies are highly similar in the L1–L4 segments of spiders (Schwager et al. 2017) and scorpions (this study); and in L1–L4 of horseshoe crabs. This correlation holds upon inclusion of Hox surveys (albeit fragmentary datasets) from other chelicerate groups; uniform expression levels of Dfd were detected in the L1–L3 legs of the hexapod larva of an acariform mite (Telford and Thomas 1998). In contrast to these groups, expression levels of the single-copy ortholog of Dfd in the harvestman are markedly heterogeneous across L1–L4, with a unique expression pattern and level of intensity in each leg (Gainett et al. 2021). The general correlation we observe across these groups is that homonomous blocks of segments (i.e., segments bearing morphologically similar appendages) tend to exhibit similar combinations of Hox expression levels.
These patterns suggest that modifications of pedipalp and leg morphology across Chelicerata may be attributable to modulation of Hox levels, driven by variation in cis-regulation of lab and Dfd to achieve unique combinations of transcript levels in morphologically distinct appendages. The presence of Hox duplications in Arachnopulmonata, while intriguing from a macroevolutionary perspective, may not be the mechanism that underlies morphological disparity of chelicerate prosomal architecture. We found no evidence that prosomal Hox genes have achieved additional segmental identities in arachnopulmonates by spatial subdivision of paralogs’ expression domains (in contrast to opisthosomal Hox copies in spiders and scorpions; (Sharma et al. 2012a; Sharma, Schwager, et al. 2014). This inference is further underscored by the marked morphological disparity between walking legs across groups of acariform mites, which are comparably diverse to spiders, but are not part of the shared genome duplication at the root of Arachnopulmonata (Schwager et al. 2017; Ontano et al. 2021).
These hypotheses could be tested further by investigating arachnopulmonates with extreme modifications of specific appendage pairs, such as Amblypygi (whip spiders), Uropygi (vinegaroons), and Solifugae (camel spiders), groups that exhibit marked elongation of the first walking leg pair. Recently developed tools for the whip spider Phrynus marginemaculatus may prove useful in this regard (Gainett and Sharma 2020). With respect to the establishment of specific leg-bearing segment identities in the harvestman, advanced tools for misexpression of Hox genes are required in this system for a nuanced understanding of Hox transcript combinatorics. Ectopic expression of harvestman lab in posterior prosomal segments (L2–L4), for example, is central to testing the hypothesis that co-expression of lab and Dfd underlies L1 fate specification. We note that such an experiment, if successful in generating four pairs of morphologically identical appendages in P. opilio, would strongly accord with the interpretation that homonomous segmentation can evolve secondarily in lineages like Xiphosura—a scenario that is suggested by the phylogenetic distribution of Hox domains across Panarthropoda (fig. 6), as well as recent phylogenetic work that has recovered horseshoe crabs as derived arachnids (Ballesteros and Sharma 2019; Ballesteros et al. 2022; Ban et al. 2022)
Materials and Methods
Embryo Collection and Gene Identification
Adult P. opilio individuals were collected from Bascom Hill, Madison, WI, USA along the exterior walls of nearby buildings. Gravid females of the scorpion C. sculpturatus were hand-collected from sites in Arizona by citizen-scientist collaborators. Embryos of the horseshoe crab L. polyphemus were collected in Woods Hole, MA, USA in June 2022. Embryonic staging nomenclature for the harvestman and horseshoe crab follows Gainett et al. (2022) and Sekiguchi et al. (1982), respectively. Details of collection, maintenance and fixation of embryos are described in the supplementary Methods, Supplementary Material online.
The complete sequences of P. opilio Hox genes labial (lab), probocipedia (pb), and Deformed (Dfd) were previously isolated from the P. opilio genome (GCA_019434445.1; [Gainett et al. 2021]) (see supplementary table S1, Supplementary Material online). The engrailed (en) ortholog in P. opilio was identified by Sharma et al. (2012a; supplementary table S1, Supplementary Material online). The scorpion C. sculpturatus, lab paralogs (Cscu-lab-1, Cscu-lab-2) and Dfd paralogs (Cscu-Dfd-1, Cscu-Dfd-2) were identified from the reference genome (GCF_000671375.1; [Schwager et al. 2017]), matching the A/B nomenclature (here paralogs 1 and 2, respectively) of their original description (Schwager et al. 2017). The numerical nomenclature for the scorpion genes was used to differentiate paralogs of the arachnopulmonate WGD from the independently derived Hox paralogs of the horseshoe crab WGDs. Horseshoe crab Hox paralogs are referenced here using alphabetic nomenclature, following the convention established in the C. rotundicauda and T. gigas genome assemblies (Shingate, Ravi, Prasad, Tay, Garg, et al. 2020; Shingate, Ravi, Prasad, Tay, and Venkatesh 2020). The horseshoe crab L. polyphemus lab paralogs (Lpol-lab-A, Lpol-lab-B, Lpol-lab-D, Lpol-lab-E) and Dfd paralogs (Lpol-Dfd-A, Lpol-Dfd-B, Lpol-Dfd-C, Lpol-Dfd-D, Lpol-Dfd-E) were retrieved from the reference genome annotation (GCF_000517525.1; Kenny et al. 2015; Battelle et al. 2016). Accession numbers (see supplementary table S2, Supplementary Material online) and phylogenetic analysis of lab (see supplementary fig. S4, Supplementary Material online) and Dfd (see supplementary fig. S5, Supplementary Material online) homologs in the horseshoe crab species are described in the Supplementary material online.
RNAi via dsRNA Embryonic Injections
Gene cloning and dsRNA synthesis are detailed in the supplementary Methods, Supplementary Material online (see supplementary table S3, Supplementary Material online). For the single knockdown of Popi-lab, two clutches of embryos were injected (n = 194). Three clutches were injected in the double knockdown of Popi-lab + Popi-Dfd (n = 217). An additional three clutches were injected with Popi-lab dsRNA, and an additional two were injected with Popi-lab + Popi-Dfd dsRNA, which were then fixed and exclusively used in both colorimetric and fluorescent in situ hybridization. Two clutches (n = 157) were injected with deionized water as a negative control. Injection mixes were prepared with Rhodamine dextran (1:20) for visualization.
Colorimetric and Fluorescent in Situ Hybridization
Colorimetric in situ hybridization gene expression assays of P. opilio used sense (control) and antisense RNA probes labeled with DIG RNA labeling mix (Roche, Basel, Switzerland) and followed published protocols (Sharma et al. 2012a). Images were obtained on a Nikon SMZ25 fluorescent stereomicroscope with a DS-Fi2 digital color camera and driven by Nikon Elements. Fluorescent in situ hybridization followed a modified version of the Molecular Instruments (Los Angeles, CA, USA) HCR v.3 protocol (Choi et al. 2018; Bruce et al. 2021). HCR probes for Hox paralogs were designed to avoid regions of high similarity to circumvent non-specific binding. Probe sequences were designed by Molecular Instruments or in an open-source probe design program (Kuehn et al. 2022). Details of probe design and probe sequences are available in the supplementary Methods, Supplementary Material online (see supplementary tables S4–S12, Supplementary Material online). Imaging was performed on a Zeiss 710 and Zeiss 780 confocal microscope at the Newcomb Imaging Center, UW-Madison, USA. Z-stacks were projected with maximal intensity mode, and linearly adjusted for brightness and contrast in FIJI (v. 2.9.0/1.53t). Figures were assembled in Adobe Illustrator 2022.
Supplementary Material
Acknowledgments
We are indebted to Sarah Swanson and the Newcomb Imaging Center (Department of Botany) at the University of Wisconsin-Madison for the imaging infrastructure. Comments from Emily V. W. Setton improved the first draft of the manuscript. We thank two anonymous reviewers and the associate editor for comments on the first draft. This work was supported by the National Science Foundation NSF IOS-2016141 grant to P.P.S.
Contributor Information
Guilherme Gainett, Department of Integrative Biology, University of Wisconsin-Madison, Madison, WI.
Benjamin C Klementz, Department of Integrative Biology, University of Wisconsin-Madison, Madison, WI.
Pola O Blaszczyk, Department of Integrative Biology, University of Wisconsin-Madison, Madison, WI.
Heather S Bruce, Marine Biological Laboratory, Woods Hole, MA.
Nipam H Patel, Marine Biological Laboratory, Woods Hole, MA; Organismal Biology & Anatomy, University of Chicago, Chicago, IL.
Prashant P Sharma, Department of Integrative Biology, University of Wisconsin-Madison, Madison, WI.
Supplementary Material
Supplementary data are available at Molecular Biology and Evolution online.
Data Availability
The data underlying this article are available in the article and in its online Supplementary material.
References
- Abzhanov A, Kaufman TC. 1999. Homeotic genes and the arthropod head: expression patterns of the labial, proboscipedia, and Deformed genes in crustaceans and insects. Proc Natl Acad Sci U S A. 96:10224–10229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Angelini DR, Liu PZ, Hughes CL, Kaufman TC. 2005. Hox gene function and interaction in the milkweed bug Oncopeltus fasciatus (Hemiptera). Dev Biol. 287:440–455. [DOI] [PubMed] [Google Scholar]
- Arnaud F, Bamber RN. 1988. The Biology of Pycnogonida. Adv Mar Biol 24:1–96. [Google Scholar]
- Auman T, Chipman AD. 2018. Growth zone segmentation in the milkweed bug Oncopeltus fasciatus sheds light on the evolution of insect segmentation. BMC Evol Biol. 18:178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ballesteros JA, Santibáñez-López CE, Baker CM, Benavides LR, Cunha TJ, Gainett G, Ontano AZ, Setton EVW, Arango CP, Gavish-Regev E, et al. 2022. Comprehensive species sampling and sophisticated algorithmic approaches refute the monophyly of Arachnida. Mol Biol Evol. 39:msac021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ballesteros JA, Setton EVW, López CES, Arango CP, Brenneis G, Brix S, Corbett KF, Sánchez EC, Dandouch M, Dilly GF, et al. 2021. Phylogenomic resolution of sea spider diversification through integration of multiple data classes. Mol Biol Evol. 38:686–701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ballesteros JA, Sharma PP. 2019. A critical appraisal of the placement of Xiphosura (Chelicerata) with account of known sources of phylogenetic error. Syst Biol. 33:440–422. [DOI] [PubMed] [Google Scholar]
- Ban X, Shao Z, Wu L, Sun J, Xue X. 2022. Highly diversified mitochondrial genomes provide new evidence for interordinal relationships in the Arachnida. Cladistics 38:452–464. [DOI] [PubMed] [Google Scholar]
- Barnett AA, Thomas RH. 2013. Posterior Hox gene reduction in an arthropod: Ultrabithorax and Abdominal-B are expressed in a single segment in the mite Archegozetes longisetosus. EvoDevo 4:23–23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Battelle B-A, Ryan JF, Kempler KE, Saraf SR, Marten CE, Warren WC, Minx PJ, Montague MJ, Green PJ, Schmidt SA, et al. 2016. Opsin repertoire and expression patterns in horseshoe crabs: evidence from the genome of Limulus polyphemus (Arthropoda: Chelicerata). Genome Biol Evol. 8:1571–1589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benton MA, Pechmann M, Frey N, Stappert D, Conrads KH, Chen Y-T, Stamataki E, Pavlopoulos A, Roth S. 2016. Toll genes have an ancestral role in axis elongation. Curr Biol. 26:1609–1615. [DOI] [PubMed] [Google Scholar]
- Brena C, Chipman AD, Minelli A, Akam M. 2006. Expression of trunk Hox genes in the centipede Strigamia maritima: sense and anti-sense transcripts. Evol Dev. 8:252–265. [DOI] [PubMed] [Google Scholar]
- Bruce HS. 2021. How to align arthropod legs. bioRxiv. 2021.01.20.427514.dx.doi.org/10.17504/protocols.io.bunznvf6
- Bruce HS, Jerz G, Kelly SR, McCarthy J, Pomerantz A, Senevirathne G, Sherrard A, Sun DA, Wolff C, Patel NH. 2021. Hybridization chain reaction (HCR) in situ protocol V.1. Protocols.io. dx.doi.org/10.17504/protocols.io.bunznvf6
- Choi HMT, Schwarzkopf M, Fornace ME, Acharya A, Artavanis G, Stegmaier J, Cunha A, Pierce NA. 2018. Third-generation in situ hybridization chain reaction: multiplexed, quantitative, sensitive, versatile, robust. Development 145:dev165753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Damen WGM, Hausdorf M, Seyfarth E-A, Tautz D. 1998. A conserved mode of head segmentation in arthropods revealed by the expression pattern of Hox genes in a spider. Proc Natl Acad Sci U S A. 95:10665–10670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dehal P, Boore JL. 2005. Two rounds of whole genome duplication in the ancestral vertebrate. PLoS Biol. 3:e314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diederich RJ, Merrill VK, Pultz MA, Kaufman TC. 1989. Isolation, structure, and expression of labial, a homeotic gene of the Antennapedia Complex involved in Drosophila head development. Genes Dev. 3:399–414. [DOI] [PubMed] [Google Scholar]
- Eriksson BJ, Tait NN, Budd GE, Janssen R, Akam M. 2010. Head patterning and Hox gene expression in an onychophoran and its implications for the arthropod head problem. Dev Genes Evol. 220:117–122. [DOI] [PubMed] [Google Scholar]
- Gainett G, Crawford AR, Klementz BC, So C, Baker CM, Setton EVW, Sharma PP. 2022. Eggs to long-legs: embryonic staging of the harvestman Phalangium opilio (Opiliones), an emerging model arachnid. Front Zool. 19:11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gainett G, González VL, Ballesteros JA, Setton EVW, Baker CM, Gargiulo LB, Santibáñez-López CE, Coddington JA, Sharma PP. 2021. The genome of a daddy-long-legs (Opiliones) illuminates the evolution of arachnid appendages. Proc R Soc B. 288:20211168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gainett G, Sharma PP. 2020. Genomic resources and toolkits for developmental study of whip spiders (Amblypygi) provide insights into arachnid genome evolution and antenniform leg patterning. EvoDevo 11:18–18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gompel N, Prud’homme B, Wittkopp PJ, Kassner VA, Carroll SB. 2005. Chance caught on the wing: cis-regulatory evolution and the origin of pigment patterns in Drosophila. Nature 433:481–487. [DOI] [PubMed] [Google Scholar]
- Grbić M, Leeuwen TV, Clark RM, Rombauts S, Rouzé P, Grbić V, Osborne EJ, Dermauw W, Ngoc PCT, Ortego F, et al. 2011. The genome of Tetranychus urticae reveals herbivorous pest adaptations. Nature 479:487–492. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harper A, Gonzalez LB, Schönauer A, Janssen R, Seiter M, Holzem M, Arif S, McGregor AP, Sumner-Rooney L. 2021. Widespread retention of ohnologs in key developmental gene families following whole genome duplication in arachnopulmonates. G3 (Bethesda) 11:jkab299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hughes CL, Kaufman TC. 2002a. Hox genes and the evolution of the arthropod body plan. Evol Dev. 4:459–499. [DOI] [PubMed] [Google Scholar]
- Hughes CL, Kaufman TC. 2002b. Exploring myriapod segmentation: the expression patterns of even-skipped, engrailed, and wingless in a centipede. Dev Biol. 247:47–61. [DOI] [PubMed] [Google Scholar]
- Jager M, Murienne J, Clabaut C, Deutsch J, Guyader HL, Manuel M. 2006. Homology of arthropod anterior appendages revealed by Hox gene expression in a sea spider. Nature 441:506–508. [DOI] [PubMed] [Google Scholar]
- Janssen R, Eriksson BJ, Tait NN, Budd GE. 2014. Onychophoran Hox genes and the evolution of arthropod Hox gene expression. Front Zool. 11:22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kenny NJ, Chan KW, Nong W, Qu Z, Maeso I, Yip HY, Chan TF, Kwan HS, Holland PWH, Chu KH, et al. 2015. Ancestral whole-genome duplication in the marine chelicerate horseshoe crabs. Heredity 116:190–199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuehn E, Clausen DS, Null RW, Metzger BM, Willis AD, Özpolat BD. 2022. Segment number threshold determines juvenile onset of germline cluster expansion in Platynereis dumerilii. J Exp Zool Part B Mol Dev Evol. 338:225–240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leite DJ, Baudouin-Gonzalez L, Iwasaki-Yokozawa S, Lozano-Fernandez J, Turetzek N, Akiyama-Oda Y, Prpic N-M, Pisani D, Oda H, Sharma PP, McGregor AP. 2018. Homeobox gene duplication and divergence in arachnids. Mol Biol Evol 35:2240–2253. 10.1093/molbev/msy125 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lohmann I, McGinnis N, Bodmer M, McGinnis W. 2002. The Drosophila Hox gene Deformed sculpts head morphology via direct regulation of the apoptosis activator reaper. Cell 110:457–466. [DOI] [PubMed] [Google Scholar]
- Löhr U, Yussa M, Pick L. 2001. Drosophila fushi tarazu a gene on the border of homeotic function. Curr Biol. 11:1403–1412. [DOI] [PubMed] [Google Scholar]
- Mahfooz N, Turchyn N, Mihajlovic M, Hrycaj S, Popadić A. 2007. Ubx regulates differential enlargement and diversification of insect hind legs. PLoS ONE 2:e866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manuel M, Jager M, Murienne J, Clabaut C, Guyader HL. 2006. Hox genes in sea spiders (Pycnogonida) and the homology of arthropod head segments. Dev Genes Evol. 216:481–491. [DOI] [PubMed] [Google Scholar]
- Martin A, Serano JM, Jarvis E, Bruce HS, Wang J, Ray S, Barker CA, O’Connell LC, Patel NH. 2016. CRISPR/Cas9 mutagenesis reveals versatile roles of Hox genes in crustacean limb specification and evolution. Curr Biol. 26:14–26. [DOI] [PubMed] [Google Scholar]
- Medved V, Marden JH, Fescemyer HW, Der JP, Liu J, Mahfooz N, Popadić A. 2015. Origin and diversification of wings: insights from a neopteran insect. Proc Natl Acad Sci U S A. 112:15946–15951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Merrill VKL, Diederich RJ, Turner FR, Kaufman TC. 1989. A genetic and developmental analysis of mutations in labial, a gene necessary for proper head formation in Drosophila melanogaster. Dev Biol. 135:376–391. [DOI] [PubMed] [Google Scholar]
- Mito T, Kobayashi C, Sarashina I, Zhang H, Shinahara W, Miyawaki K, Shinmyo Y, Ohuchi H, Noji S. 2007. even-skipped has gap-like, pair-rule-like, and segmental functions in the cricket Gryllus bimaculatus, a basal, intermediate germ insect (Orthoptera). Dev Biol. 303:202–213. [DOI] [PubMed] [Google Scholar]
- Oliver JC, Beaulieu JM, Gall LF, Piel WH, Monteiro A. 2014. Nymphalid eyespot serial homologues originate as a few individualized modules. Proc R Soc B Biol Sci. 281:20133262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ontano AZ, Gainett G, Aharon S, Ballesteros JA, Benavides LR, Corbett KF, Gavish-Regev E, Harvey MS, Monsma S, Santibáñez-López CE, et al. 2021. Taxonomic sampling and rare genomic changes overcome long-branch attraction in the phylogenetic placement of pseudoscorpions. Mol Biol Evol. 38:2446–2467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pechmann M, Schwager EE, Turetzek N, Prpic N-M. 2015. Regressive evolution of the arthropod tritocerebral segment linked to functional divergence of the Hox gene labial. Proc R Soc B Biol Sci. 282:20151162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Posnien N, Bucher G. 2010. Formation of the insect head involves lateral contribution of the intercalary segment, which depends on Tc-labial function. Dev Biol. 338:107–116. [DOI] [PubMed] [Google Scholar]
- Rafiqi AM, Rajakumar A, Abouheif E. 2020. Origin and elaboration of a major evolutionary transition in individuality. Nature 585:1–30. [DOI] [PubMed] [Google Scholar]
- Schaeper ND, Pechmann M, Damen WGM, Prpic N-M, Wimmer EA. 2010. Evolutionary plasticity of collier function in head development of diverse arthropods. Dev Biol. 344:363–376. [DOI] [PubMed] [Google Scholar]
- Schönauer A, Paese CLB, Hilbrant M, Leite DJ, Schwager EE, Feitosa NM, Eibner C, Damen WGM, McGregor AP. 2016. The Wnt and Delta-Notch signalling pathways interact to direct pair-rule gene expression via caudal during segment addition in the spider Parasteatoda tepidariorum. Development 143:2455–2463. [DOI] [PubMed] [Google Scholar]
- Schwager EE, Sharma PP, Clarke T, Leite DJ, Wierschin T, Pechmann M, Akiyama-Oda Y, Esposito L, Bechsgaard J, Bilde T, et al. 2017. The house spider genome reveals an ancient whole-genome duplication during arachnid evolution. BMC Biol. 15:62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sekiguchi K, Yamamichi Y, Costlow JD. 1982. Horseshoe crab developmental studies I. Normal embryonic development of Limulus polyphemus compared with Tachypleus tridentatus. Prog Clin Biol Res. 81:53–73. [PubMed] [Google Scholar]
- Serano JM, Martin A, Liubicich DM, Jarvis E, Bruce HS, La K, Browne WE, Grimwood J, Patel NH. 2016. Comprehensive analysis of Hox gene expression in the amphipod crustacean Parhyale hawaiensis. Dev Biol. 409:297–309. [DOI] [PubMed] [Google Scholar]
- Setton EVW, Sharma PP. 2018. Cooption of an appendage-patterning gene cassette in the head segmentation of arachnids. Proc Natl Acad Sci U S A. 115:E3491–E3500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharma PP, Kaluziak ST, Pérez-Porro AR, González VL, Hormiga G, Wheeler WC, Giribet G. 2014. Phylogenomic interrogation of Arachnida reveals systemic conflicts in phylogenetic signal. Mol Biol Evol. 31:2963–2984. [DOI] [PubMed] [Google Scholar]
- Sharma PP, Schwager EE, Extavour CG, Giribet G. 2012a. Hox gene expression in the harvestman Phalangium opilio reveals divergent patterning of the chelicerate opisthosoma. Evol Dev. 14:450–463. [DOI] [PubMed] [Google Scholar]
- Sharma PP, Schwager EE, Extavour CG, Giribet G. 2012b. Evolution of the chelicera: a dachshund domain is retained in the deutocerebral appendage of Opiliones (Arthropoda, Chelicerata). Evol Dev. 14:522–533. [DOI] [PubMed] [Google Scholar]
- Sharma PP, Schwager EE, Extavour CG, Wheeler WC. 2014. Hox gene duplications correlate with posterior heteronomy in scorpions. Proc Biol Sci. 281:20140661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharma PP, Schwager EE, Giribet G, Jockusch EL, Extavour CG. 2013. Distal-less and dachshund pattern both plesiomorphic and apomorphic structures in chelicerates: RNA interference in the harvestman Phalangium opilio (Opiliones). Evol Dev. 15:228–242. [DOI] [PubMed] [Google Scholar]
- Sharma PP, Tarazona OA, Lopez DH, Schwager EE, Cohn MJ, Wheeler WC, Extavour CG. 2015. A conserved genetic mechanism specifies deutocerebral appendage identity in insects and arachnids. Proc R Soc B Biol Sci. 282:20150698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shingate P, Ravi V, Prasad A, Tay B-H, Garg KM, Chattopadhyay B, Yap L-M, Rheindt FE, Venkatesh B. 2020. Chromosome-level assembly of the horseshoe crab genome provides insights into its genome evolution. Nat Commun. 11:2322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shingate P, Ravi V, Prasad A, Tay B-H, Venkatesh B. 2020. Chromosome-level genome assembly of the coastal horseshoe crab (Tachypleus gigas). Mol Ecol Resour. 20:1748–1760. [DOI] [PubMed] [Google Scholar]
- Shultz JW. 1989. Morphology of locomotor appendages in Arachnida: evolutionary trends and phylogenetic implications. Zool J Linn Soc 97:1–55. [Google Scholar]
- Smith FW, Boothby TC, Giovannini I, Rebecchi L, Jockusch EL, Goldstein B. 2016. The compact body plan of tardigrades evolved by the loss of a large body region. Curr Biol. 26:224–229. [DOI] [PubMed] [Google Scholar]
- Snodgrass ERL. 1952. Textbook of Arthropod Anatomy: Cornell University Press. 20–40. http://www.jstor.org/stable/10.7591/j.ctvn1tb6g.6. [Google Scholar]
- Stauber M, Jäckle H, Schmidt-Ott U. 1999. The anterior determinant bicoid of Drosophila is a derived Hox class 3 gene. Proc Natl Acad Sci U S A. 96:3786–3789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Telford MJ, Thomas RH. 1998. Expression of homeobox genes shows chelicerate arthropods retain their deutocerebral segment. Proc Natl Acad Sci U S A. 95:10671–10675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tenlen JR, McCaskill S, Goldstein B. 2013. RNA Interference can be used to disrupt gene function in tardigrades. Dev Genes Evol. 223:171–181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Turetzek N, Khadjeh S, Schomburg C, Prpic N-M. 2017. Rapid diversification of homothorax expression patterns after gene duplication in spiders. BMC Evol Biol. 17:1–12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Turetzek N, Pechmann M, Schomburg C, Schneider J, Prpic N-M. 2015. Neofunctionalization of a duplicate dachshund gene underlies the evolution of a novel leg segment in arachnids. Mol Biol Evol. 33:109–121. [DOI] [PubMed] [Google Scholar]
- Wagner GP, Amemiya C, Ruddle F. 2003. Hox cluster duplications and the opportunity for evolutionary novelties. Proc Natl Acad Sci U S A. 100:14603–14606. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The data underlying this article are available in the article and in its online Supplementary material.