Abstract
Quantitative assessment of brain myelination has gained attention for both research and diagnosis of neurological diseases. However, conventional pulse sequences cannot directly acquire the myelin‐proton signals due to its extremely short T2 and T2* values. To obtain the myelin‐proton signals, dedicated short T2 acquisition techniques, such as ultrashort echo time (UTE) imaging, have been introduced. However, it remains challenging to isolate the myelin‐proton signals from tissues with longer T2. In this article, we extended our previous two‐dimensional ultrashort echo time magnetic resonance fingerprinting (UTE‐MRF) with dual‐echo acquisition to three dimensional (3D). Given a relatively low proton density (PD) of myelin‐proton, we utilized Cramér–Rao Lower Bound to encode myelin‐proton with the maximal SNR efficiency for optimizing the MR fingerprinting design, in order to improve the sensitivity of the sequence to myelin‐proton. In addition, with a second echo of approximately 3 ms, myelin‐water component can be also captured. A myelin‐tissue (myelin‐proton and myelin‐water) fraction mapping can be thus calculated. The optimized 3D UTE‐MRF with dual‐echo acquisition is tested in simulations, physical phantom and in vivo studies of both healthy subjects and multiple sclerosis patients. The results suggest that the rapidly decayed myelin‐proton and myelin‐water signal can be depicted with UTE signals of our method at clinically relevant resolution (1.8 mm isotropic) in 15 min. With its good sensitivity to myelin loss in multiple sclerosis patients demonstrated, our method for the whole brain myelin‐tissue fraction mapping in clinical friendly scan time has the potential for routine clinical imaging.
Keywords: 3D, Cramér–Rao lower bound, magnetic resonance fingerprinting, myelin tissue fraction mapping, myelin‐proton, ultrashort echo time
In this article, we proposed a novel three‐dimensional ultrashort echo time magnetic resonance fingerprinting method (UTE‐MRF) with dual‐echo acquisition for simultaneously quantifying both myelin‐proton, myelin‐water and regular long T2 tissues. In particular, we proposed to utilize the Cramér–Rao Lower Bound to encode myelin‐proton to improve the SNR efficiency and signal sensitivity of myelin‐proton.

Abbreviations
- CC
Corpus Callosum
- CRLB
Cramér–Rao Lower Bound
- FA
Flip angle
- FLAIR
Fluid‐attenuated inversion recovery
- FIM
Fisher information matrix
- FOV
Field‐of‐view
- GM
Gray matter
- GRASE
Gradient and spin‐echo
- MPF
Myelin‐proton fraction
- MPRAGE
Magnetization prepared rapid acquisition gradient echo
- MRF
Magnetic resonance fingerprinting
- MRI
Magnetic resonance imaging
- MS
Multiple sclerosis
- MTF
Myelin‐tissue fraction
- MWF
Myelin‐water fraction
- MWI
Myelin water imaging
- NE
Normalized error
- NUFFT
nonuniform fast Fourier transform
- PD
Proton density
- RF
Radio frequency
- ROI
Region‐of‐interest
- SNR
Signal‐to‐noise ratio
- TI
Inversion time
- TR
Repetition time
- UTE
Ultrashort echo time
- UTE‐MRF
Ultrashort echo time MR fingerprinting
- WM
White matter
- ZTE
Zero echo time
1. INTRODUCTION
Myelin is a multilayered lipid–protein lamellar membranous structure enveloping axons in both central and peripheral nervous system of vertebrates (Laule et al., 2007), with the myelin‐water trapped between the layers of the myelin sheath. Thus, myelin‐tissue is composed of nonaqueous protons within myelin sheath and myelin‐water trapped between myelin sheath. For simplicity, we will refer the former to “myelin‐proton.” Myelin is vitally important for functions of healthy central nervous system. Demyelination results in high degree of the impairment of signal transmission, along with the loss of cognition, sensation and motor functioning, thus leading to the development of neuro‐degenerative disorders, such as multiple sclerosis (MS). Therefore, noninvasive imaging of myelin‐tissue can greatly aid in the in‐depth studies of the development of demyelinating diseases.
Many long‐term efforts have been put into developing various magnetic resonance imaging (MRI) methods for myelin‐tissue detection. In clinical practice, fluid‐attenuated inversion recovery (FLAIR) (Bachmann et al., 2006) and magnetization‐prepared rapid acquisition gradient echo (MPRAGE) (Nair et al., 2013) play an important role in the diagnosis of MS, but they are not specific to demyelination. The widely used myelin‐specific methods exploit the shorter relaxation time of myelin‐water molecules trapped inside the myelin sheath, referred to as myelin water imaging, such as acquiring multi‐echo T2 data using the multi‐echo spin‐echo sequence (MacKay et al., 1994) or the multi‐echo gradient and spin‐echo sequence (Does & Gore, 2000), and acquiring multi‐echo T2* data with multigradient‐echo sequences (Chen et al., 2021; Du et al., 2007). In addition, the direct visualization of the myelin water, referred to as ViSTa, was also achieved by exploiting the shorter T1 and T2 properties (Liao et al., 2021; Oh et al., 2013; Wu et al., 2018). These methods can detect the signal produced by myelin‐water protons within and around solid myelin‐proton, which is a potential markers for the myelin‐tissue presence, but nevertheless only an indirect indicator of myelin‐proton (Laule et al., 2007). Because the rapid signal decays of myelin‐proton are undetectable at the conventional echo time (TE) with the aforementioned sequences, its possibility of being directly imaged is hindered by the extremely short T2/T2* relaxation time of myelin protons.
To address this limitation, the ultrashort echo time (UTE) technique was developed to directly visualize the ultrashort T2/T2* components with TE less than 1 ms (Du, Sheth, et al., 2014; Muller et al., 2021; Waldman et al., 2003), which was attributed to myelin proton. Zero echo time (Weiger et al., 2013) was also applied to explore myelin‐proton imaging (Seifert et al., 2017; Weiger et al., 2020). Nevertheless, even in myelin‐tissue‐rich white matter (WM) regions, a large part of the UTE signal originates from long T2 tissues, still very challenging to directly image the myelin proton. The inversion recovery preparation was used for suppression of long T2 tissues signals by nulling the longitudinal magnetization using the specific inversion time (TI) (Fan et al., 2017; Horch et al., 2012; Ma et al., 2019). However, an accurate TI is difficult to achieve and even a slight offset from the accurate TI will lead to the severe signal contamination from long T2 tissues (Ma, Jang, et al., 2020; Ma, Searleman, et al., 2020). Recently, short repetition time (TR) adiabatic inversion recovery prepared UTE sequence (Ma, Jang, et al., 2020) and double‐echo sliding inversion recovery UTE sequence (Ma, Searleman, et al., 2020) were developed to efficiently suppress the long T2 tissues regardless of T1 variations in WM. These techniques improve the outcome of direct imaging of myelin‐proton within WM, but at the expense of scan time and spatial resolution in the whole brain imaging.
Magnetic resonance fingerprinting (MRF) is a fast multiparametric mapping technique, which can simultaneously generate multiple tissue parameters in a single scan (Ma et al., 2013). In MRF, each signal evolution or fingerprint was matched with precalculated dictionary to identify tissue parameters, including T1, T2, T2*, and proton density (PD) (Boyacioglu et al., 2021; Cao et al., 2017, 2019, 2022; Chen et al., 2019; Chen, Panda, et al., 2019; Jiang et al., 2015; Korzdorfer et al., 2019; Liao et al., 2017, 2018). With a multiple component analysis method, it is possible to estimate tissue fraction, such as WM, gray matter (GM), and myelin‐water fraction (MWF) (Chen et al., 2019; Cui et al., 2021; Liao et al., 2018; Nagtegaal et al., 2020). More recently, a number of research has been focused on optimizing the MRF acquisition and dictionary generation based on Cramér–Rao Lower Bound (CRLB) (Daeun et al., 2017; Lahiri et al., 2020; Lee et al., 2019; Zhao et al., 2019), while other methods were also developed (Jordan et al., 2021; Scope Crafts et al., 2022; Zou et al., 2021). However, challenges arise when applying MRF to quantify the properties of ultrashort T2/T2* tissues. First, the minimum TE in the conventional MRF sequence is several milliseconds, resulting in difficulty on the detection of ultrashort T2/T2* tissues. Second, due to the low PD of ultrashort T2/T2* tissues, the signal‐to‐noise ratio (SNR) is diminished, resulted in significantly lower accuracy of tissue properties estimation. Third, because myelin‐proton has an ultrashort T2/T2*, the relatively long readout duration of spiral trajectory (usually 5–10 ms) in conventional MRF is sensitive to the decay of myelin proton, leading to the visible image blurring (Qian & Boada, 2008).
To address the aforementioned problems, our previous work has proposed a 2D ultrashort echo time MR fingerprinting (UTE‐MRF) method (Li et al., 2019) with dual‐echo radial readout trajectories. This method combines the advantages of UTE and MRF; in addition, it is able to simultaneously quantify both long T2 and ultrashort T2/T2* tissues. This method eliminates the difficulties associated with the selection of an accurate TI for long T2 tissue suppression in WM. However, due to the radio frequency (RF) power and gradient amplitude limitations, this method was limited by using the half pulse slice excitation scheme, resulting in the difficulty on obtaining the whole brain volume coverage with an isotropic resolution in a clinical friendly scan time.
In this article, we extend the two‐dimensional (2D) UTE‐MRF to a three‐dimensional (3D) version with a hard pulse for volume excitation, which provides a higher SNR efficiency with faster acquisition. More importantly, an optimized sequence acquisition parameter pattern was designed by utilizing the CRLB to maximize SNR for myelin proton, and to improve the quantification precision of the ultrashort T2/T2* components. In order to obtain myelin‐proton fraction (MPF), a multicomponent analysis without a priori assumption about the exact number of tissue components or their relaxation properties in each voxel (Nagtegaal et al., 2020) called sparsity promoting iterative joint nonnegative least squares (SPIJN) algorithm was utilized for the separation of myelin‐proton from long T2 tissues in the ultrashort echo signal. In addition, with a second TE of approximately 3 ms, MWF could be also obtained. The potential utility of the proposed sequence was demonstrated in simulation, phantom, and in vivo experiments with 10 healthy subjects and 10 MS patients.
2. METHODS
2.1. Pulse sequence
The sequence diagram for 3D UTE‐MRF is shown in Figure 1a. A hard pulse with a duration of 0.2 ms was employed for volume excitation. Following the IR pulse, a train of center‐out dual‐echo acquisition (n = 564 TRs, or frames) was used in the MRF unit, with a typical TI of 10 ms (Jiang et al., 2017; Ma et al., 2013) for T1 quantification in MRF. In this article, we varied the FA and TE among different TRs while other parameters were remained the same following the previous 2D UTE‐MRF sequence (Li et al., 2019). The FA and ultrashort TE varying patterns in our sequence were optimized to generate a train of signals that is more sensitive to myelin‐proton. TE for the second echo was adding 2.86 ms to the echo 1. Section 2.2 describes how we optimized the aforementioned FA and ultrashort TE varying patterns. TR was fixed to 7 ms. In order to reduce T2* blurring, the radial trajectories were selected for minimizing the readout data collection window. With the ramp sampling, the readout duration was set to 1.28 ms (0.15 ms for ramp, and 1.13 ms for plateau), and the readout bandwidth was set to 890 Hz/pixel. A delay time of 2000 ms was set between each repetition, which is the waiting period after spoiler gradients for spin recovery to return to their initial state. An optimized 2D golden‐angle method (Zhang et al., 2019) was chosen to cover the 3D k‐space and to maximize the signal spatial–temporal incoherence, the trajectory intermixes in‐plane and through‐plane rotations along both frames and repetitions. Each of 564 frames has 160 repetitions to improve SNR with an acceptable scan time, from which we could get 564 under‐sampled images, each based on 160 radial spokes. Among repetitions, the radial trajectories were uniformly distributed over a unit sphere. For the reconstruction, a sliding window reconstruction algorithm (Cao et al., 2017) was utilized to further reduce the under‐sampling rate. To balance the tissue specificity and under‐sampling rate blurring, a window size of 60 was used in this study, and the reconstructed spatial resolution was estimated to be 1.8 mm isotropic from the corresponding point spread function of under‐sampling radial trajectories. Figure 1c depicts the k‐space coverage of the first frame under 160 repetitions. The total scan time was 15 min. Other parameters are listed in Figure 1b. It is noted that, since ultrashort T2* tissue signals fully decay at end of each TR and will not generate stimulated echo, we could only measure the T2* for ultrashort T2/T2* tissues. Thus, we obtained T2* values for ultrashort T2/T2* tissues while T2 value for long T2 tissues from our sequence.
FIGURE 1.

Sequence diagram of the Cramér–Rao lower bound‐derived three‐dimensional (3D) ultrashort echo time magnetic resonance fingerprinting (UTE‐MRF). (a) Sequence diagram with flip angle (FA) and echo time (TE, echo 1) pattern of two sequences, with the blue dashed part in right top panel the zoomed‐in part of FA and ultrashort TE. Each repetition has 564 TRs and delay time between each repetition is 2000 ms. (b) Sequence parameters are shown in the table. (c) The 3D k‐space coverage of frame 1 using a two‐dimensional golden angle method under repetition of 160 and 800, in other words each under‐sampled image was reconstructed based on 160 and 800 radial spokes. The radial trajectories are going from center to the edge and back to the center as blue lines show.
2.2. CRLB optimization
Given that the estimation precision depends on the sensitivity of the sequence to the parameters of interest and noise, the CRLB method (Daeun et al., 2017; Lahiri et al., 2020; Lee et al., 2019; Zhao et al., 2019) was adopted to optimize the acquisition parameter (FA and ultrashort TE) pattern of our 3D UTE‐MRF sequence to improve the detection of myelin‐proton. The bound was calculated using autograd differentiation (Lee et al., 2019).
For the unknown parameters, , where is the number of parameters; according to the assumption of the white Gaussian noise, the CRLB has a simplified form of the estimated parameter, , which can be formulated as , where is the Jacobian matrix of each UTE‐MRF signal with respect to the unknown parameters . Note that is measured at the corresponding TE after each RF pulse. Here, constrained by the computational burden, we mainly focus on the six parameters: T1 and T2 values of three representative MR tissues of myelin‐proton, WM, and GM. The target tissue parameters at 3 T were assumed to be T1/T2 = 300/1 ms for myelin‐proton (Du, Sheth, et al., 2014), 800/60 ms for WM, and 1300/80 ms for GM (Buonincontri et al., 2021; Cao et al., 2019; Deoni et al., 2005; Liao et al., 2018; Ma et al., 2017, 2018; Wansapura et al., 1999). For the sequence optimization, a weighting matrix was introduced as CRLBs are at very different scales for different parameters (Zhao et al., 2019). In this article, we set the entries of weighting matrix to be the inverse of the parameter squares according to (Lee et al., 2019), expressed as . We further added the smoothness constraints on FA and ultrashort TE variations to improve the quantification precision for highly under‐sampling experiment (Zhao et al., 2019). The final optimization problem can be mathematically expressed as:
| (1) |
The UTE‐MRF signal was simulated using the extended phase graphs (EPG) formalism (Weigel, 2015). We used the Python library of autograd, for signal and CRLB calculation (Lee et al., 2019). The optimization was initialized with different acquisition parameter patterns, shown in Table 1. Scheme A and B were taken from our previous UTE‐MRF (Li et al., 2019, 2020). The FA patterns of Scheme C, D, E were obtained from optimized scheme b, c, j (Jordan et al., 2021), while TEs were fixed to 0.05 ms for UTE considerations. Sequences of optimized b, c, j in (Jordan et al., 2021) were chosen because their TR numbers are larger than our TR number, 564. As TR was fixed in this study, the spiked‐TR‐pattern proposed in Jordan et al. (2021) was not included in our initialization.
TABLE 1.
The different schemes with optimization initialization of echo time (TE) and flip angle (FA) patterns
| TE (ms) | FA (°) | |
|---|---|---|
| Scheme A | (Li et al., 2019) | FA pattern from Li et al. (2019) |
| Scheme B | 0.05 (Li et al., 2020) | FA pattern from Li et al. (2020) |
| Scheme C | 0.05 | FA pattern of optimized b from Table S1 (Jordan et al., 2021) |
| Scheme D | 0.05 | FA pattern of optimized c from Table S1 (Jordan et al., 2021) |
| Scheme E | 0.05 | FA pattern of optimized j from Table S1 (Jordan et al., 2021) |
Sequences were optimized using a Python toolbox SciPy. After a number of runs of optimization algorithms with different initialization seeds (different FA and ultrashort TE pattern), the most promising sequence with the lowest cost function (Equation (1)) was selected for further numerical simulation, phantom, and in vivo experiments.
2.3. Numerical simulations
The simulations were designed to investigate on: (a) the ability of 3D UTE‐MRF sequence on detecting myelin‐proton; and (b) the efficiency of the CRLB on improving the accuracy of tissue quantification. Numerical simulations were implemented on a 3D digital phantom (with a size of for whole cubic region) with its transverse view shown in Figure 2a, including 15 tubes with five mixed tubes (tubes 1–5) of myelin‐proton, myelin‐water, and WM, and single‐component tubes with different T1, T2, and T2′ (tubes 6–15). Since we were interested in separating myelin‐proton from long T2 tissues in WM, the mixture of myelin‐proton and GM was not considered. Mixture tubes were used for tissue fraction estimation and single‐component tubes were utilized for T1/T2 mapping. For signal generation, we considered two different sets of acquisitions: (1) Original 3D UTE‐MRF proposed previously (Li et al., 2020); and (2) the CRLB‐derived 3D UTE‐MRF scheme. The Ideal MRF signals were generated from the dictionary. T2* effect was compensated by multiplying the ideal MRF signals with an additional exponential T2′ term due to the TE variation between each frame (Li et al., 2019). The T2′ values were chosen (as listed in Figure 2b) leading to T2* value of 0.5 ms for myelin‐proton (Du, Ma, et al., 2014), 19 ms for myelin‐water (Alonso‐Ortiz et al., 2018) and 54 ms for WM (Peters et al., 2007). For single‐component tubes, different T2′ were set to study the T2* effects on UTE‐MRF matching. To evaluate the feasibility of CRLB method for improving parameter mapping quality without increasing k‐space coverage rate, we also simulated the experiment under repetitions of 800, corresponding to five times increase in scan time, namely to reconstruct each under‐sampled image based on 800 radial spokes.
FIGURE 2.
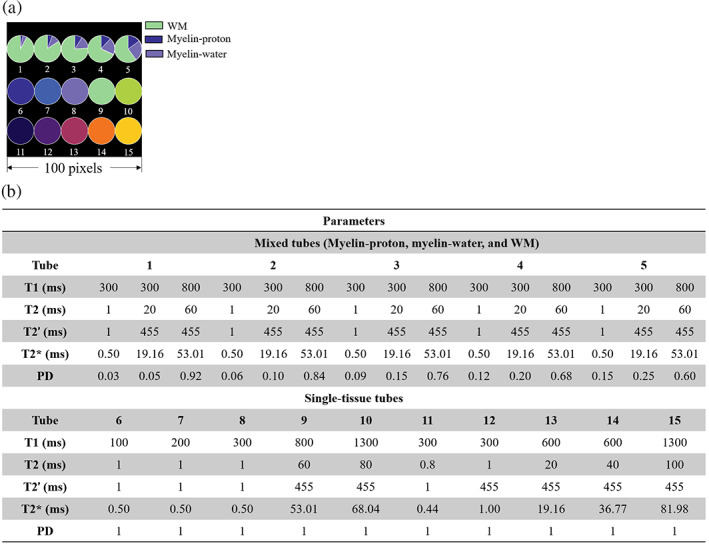
Transverse view of the digital phantom and simulation parameters used. (a) 15 tubes, with tube 1–5 for the mixture of myelin‐proton, myelin‐water, and WM and tube 6–15 of pure tissues for T1 and T2 matching. Different colors indicate different tissues. Note that the pie charts only indicate the portions of the different tissue and the digital phantom is uniform in each tube. Tissue parameters are listed in (b) with T1, T2, T2′, T2* and proton density (PD) in fraction.
2.4. Physical phantom and in vivo experiments
MR acquisition for both physical phantom and in vivo experiments were performed on a 3 T MAGNETOM Prisma scanner (Siemens Healthineers) using a 64‐channel head coil.
For physical phantom experiments, six ultrashort T2* phantoms were created by doping distilled water with manganese chloride () at different concentrations. The acquisitions were performed as follows:
The CRLB optimized 3D UTE‐MRF sequence with FA and ultrashort TE pattern plotted in Figure 1a, with FOV = , TI = 10 ms. The resolution is . Other parameters are listed in Figure 1b.
A B1 mapping sequence using product sequence of tfl_b1map, with TR = 6150 ms, TE = 1.9 ms, FA = 8°, FOV = , slice thickness = 5 mm, matrix = , and total scan time = 12 s.
An inversion recovery UTE sequence for a reference T1 map, with TIs of 5, 6, 7, 8, 10, 20 ms, TR = 3000 ms, TE = 0.05 ms, and total scan time = 2 h 29 min.
A UTE sequence for the reference T2* map, with TEs = 0.05, 0.1, 0.2, 0.5, 1, 2 ms, TR = 1500 ms, and total scan time = 47 min.
The in vivo studies were approved by the local institutional review board. Ten healthy volunteers (5 M/5F; age: ), and 10 MS patients (3 M /7F: age: ) were scanned with written informed consent. The acquisitions were performed as follows:
A product clinical axial FLAIR scan: TR = 9000 ms; TE = 89 ms; FA = 150°; FOV = ; matrix size = ; slice thickness = 1.4 mm; TI = 2500 ms; and scan time = 3 min.
A product MPRAGE scan: TR = 2300 ms; TE = 2.2 ms; FA = 9°; FOV = ; matrix = ; slice thickness = 0.9 mm; and scan time = 5 min.
The same CRLB optimized 3D UTE‐MRF scan used in the physical phantom study was implemented for the in vivo study.
The B1 mapping using product sequence of tfl_b1map: TR = 6150 ms; TE = 1.9 ms, FA = 8°; FOV = ; slice thickness = 5 mm; matrix = ; and scan time = 12 s.
The Subject 1 was scanned twice with 2 days apart for reproducibility study and a MS patient was scanned twice with 2 months apart for a longitudinal compare on lesion detection.
An MS patient was scanned with another sequence called mGRE for MWF, the protocol was as follows, mGRE, a high‐resolution 3D mGRE for simultaneous whole brain myelin‐water imaging: 32‐echo bi‐polar readout train with the first echo of 2.7 ms, echo spacing = 1.5 ms, TR = 62 ms, flip angle = 20°, readout bandwidth = 930 Hz/pixel, FOV = ; slice thickness = 2 mm; matrix = , and scan time = 16 min.
2.5. Data processing
The reconstruction for MRF was performed on a Linux workstation (2 × 2.80 GHz 20‐Core Intel Xeon CPU, 252GB RAM) using MATLAB R2018a (Mathworks), and it took approximately 2 h to reconstruct for each measurement. The dictionary was generated using EPG. T1 values varied from 1 to 3500 ms, in range of [1:0.1:10, 11:1:20, 22:2:800, 820:20:2000, 2100:100:3500] ms. The T2 values varied from 0.1 to 2000 ms, in range of [0.1:0.1:5, 10:5:150, 200:50:2000] ms. The B1 scaling factor varied from 0.75 to 1.25, in range of [0.75:0.05:1.25] for B1+ inhomogeneity correction (Ma et al., 2017). A two‐step dictionary generation method was utilized to model the partial recovery effect (Liao et al., 2017; Ma et al., 2018). The nonuniform fast Fourier transform (Fessler & Sutton, 2003) was utilized to transform the k‐space data into image space. Furthermore, T1 and T2 maps for the single‐component tubes for both simulations and physical phantoms were generated using a template matching algorithm (Ma et al., 2013).
For tissue fraction mapping, a multicomponent analysis was used for the tissue segmentation of the reconstructed data for both mixture tubes data in the numerical simulations and data from in vivo studies (Chen et al., 2019; Cui et al., 2021; Deshmane et al., 2017; Liao et al., 2018; Ma et al., 2013). An open toolbox called SPIJN (https://github.com/MNagtegaal/SPIJN) (Nagtegaal et al., 2020) was utilized to do the tissue‐fraction segmentation without making a prior assumption about the exact number of tissue components or their T1/T2 values in each voxel. For mGRE data, magnitude part of the complex data was filtered first to increase the SNR of T2* decay according to Wu et al. (2017). Then SPIJN algorithm was performed on the filtered mGRE magnitude data for MWF.
After SPIJN process, we calculated region of interest (ROI)‐based mean values for measures from UTE‐MRF on body, genu and splenium of corpus callosum (CC). The ROI‐based measurements were compared between the healthy and MS patient groups (Healthy subjects and MS patients) using t‐test implemented in SPSS (Version 20, SPSS Inc.).
3. RESULTS
3.1. Optimized pulse sequence
Since our focus is on the myelin‐proton imaging and for computation burden consideration, we optimized the sequence based on the CRLB of T1/T2 of myelin‐proton, WM, and GM. Compared with the original sequence, the normalized CRLB ( and ) values of the optimized sequences were decreased significantly for both T1 and T2 for all three tissues (Figure 3). In particular, the Scheme A provided the smallest overall CRLB (168.64), and it was selected for further simulations, phantom, and in vivo experiments.
FIGURE 3.
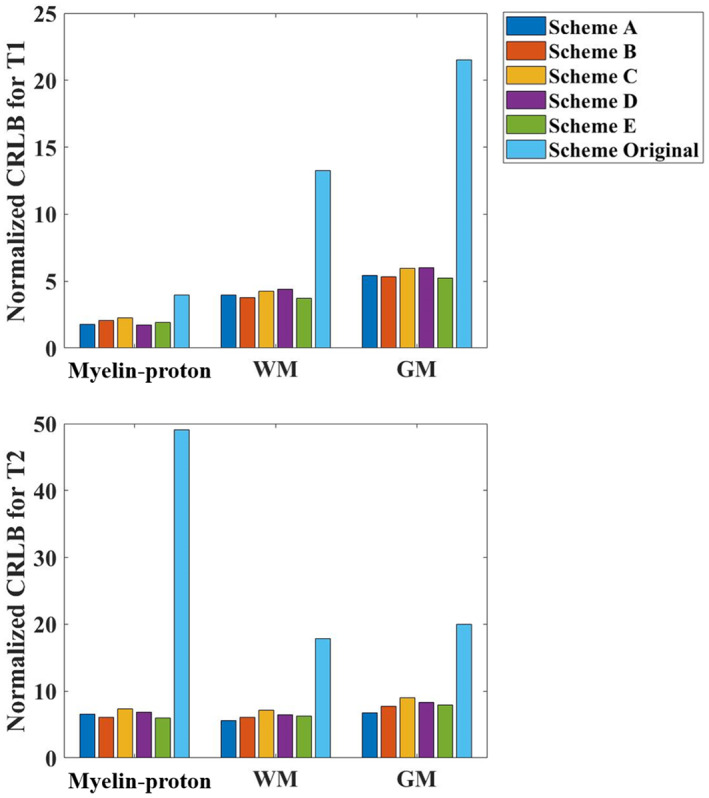
Normalized Cramér–Rao lower bound values for different optimized and original ultrashort echo time magnetic resonance fingerprinting schemes with respect to T1 and T2 of myelin‐proton, white matter and gray matter.
3.2. Numerical simulations
Simulation results of digital phantoms from CRLB‐optimized 3D UTE‐MRF sequence under 160 repetitions were shown in Figures 4 and 5. Figure 4 only shows the estimated fraction maps of mixed tubes 1–5, while Figure 5 shows the matched T1/T2 maps of single‐component tissue tubes 6–15.
FIGURE 4.

Comparison of simulation results of mixed tissue tubes 1–5 between Cramér–Rao lower bound‐optimized three‐dimensional ultrashort echo time magnetic resonance fingerprinting under 160 repetitions and the ground truth. (a) The sample labels with tubes 1–5 enclosed in red rectangular. (b,c) The estimated myelin‐proton fraction (MPF) from echo 1, and myelin‐water fraction (MWF) maps from echo 2. (d) The distribution of estimated T1 and T2 values for myelin‐proton and myelin‐water. The comparisons of mean MPF (e) and MWF (f) values for each tube with ground truth are plotted, with mean ± SD from region of interest covering each individual tube
FIGURE 5.

Comparison of simulation results of single‐tissue tubes 6–15 between Cramér–Rao lower bound‐optimized three‐dimensional ultrashort echo time magnetic resonance fingerprinting (UTE‐MRF) under 160 repetitions and the ground truth. (a) The sample labels. (b,d) The reconstructed maps and error maps of T1, T2 (for tubes 6–15). The comparisons of mean values for each tube with reference were plotted, with (c) for T1 map, (e) and (f) for T2 map. Note that T2 is shown at different display scales (d). The UTE‐MRF estimated T2 values of long T2 tissue tubes have good agreement with ground truth (e) while that of ultrashort T2* tissue tubes tend toward the nominal T2* values (f), with mean ± SD from region of interests covering each individual tube.
Figure 4a shows the sample label, with tubes 1–5 marked in red rectangular which are used for tissue fractional mapping. For the ultrashort echo signal, only two components were matched with their T1/T2 = 320/1.1 ms and 800/60 ms. These two were assumed to be myelin‐proton and WM. With a second echo signal, two components with T1/T2 = 300/12 ms and 800/60 ms were matched and assumed to be myelin‐water and WM. Figure 4b,c shows the estimated MPF from echo 1 and MWF maps from echo 2 of tubes 1–5, with the first row showing the ground truth, the second row showing reconstructed maps and the third row showing the error maps. Figure 4d shows the T1 and T2 values of the extracted myelin‐proton and myelin‐water components, with the red circle representing the ground truth and the blue filled dot the matched T1 and T2 values. Figure 4e,f is the comparisons between estimated mean MPF and mean MWF values of each tube with their corresponding ground truth.
Figure 5a shows the sample labels, with tubes 6–15 enclosed in red rectangular. Figure 5b presents the reconstructed T1 map and error map with the comparison between estimated mean T1 value of each tube and its corresponding ground truth (Figure 5c). Since the digital phantom has a large T2 range from 1 to 100 ms, the estimated T2 maps are displayed at two scales: [0, 120] ms for long T2 tissues, and [0, 2.5] ms for the ultrashort T2* tissues (Figure 5d). Compared with the original sequence where the ultrashort T2* values failed to be accurately estimated, with error even larger than the ground truth (Figure S1c), the CRLB‐optimized sequence showed significantly lower error (Figure S1f). Estimated T2 values for long T2 tissues displayed good agreement with the ground truth, while the estimated T2 values for ultrashort T2* tissues tended to approach to the nominal T2* values (Figure 5f, Figure S1c,f), due to the T2* effects which was also reported in previous paper (Li et al., 2019). Similar results under 800 repetitions are shown in Figure S2–S4. Overall, the accuracy of T1, T2 quantification, and fractional estimation improved with increasing repetitions. RMSE of T1 reduced from 35.88 (160 repetitions) to 13.35 (800 repetitions), RMSE of T2 from 3.17 (160 repetitions) to 1.12 (800 repetitions), and RMSE of MPF/MWF from 0.0109/0.0154 (160 repetitions) to 0.0059/0.0068 (800 repetitions).
3.3. Physical phantom and in vivo studies
3.3.1. Physical phantom studies
Figure 6 shows the phantom maps from CRLB‐optimized 3D UTE‐MRF and the reference methods. It can be observed that the T1 and T2 maps obtained from ultrashort TE signal of CRLB‐derived 3D UTE‐MRF method are consistent with the conventional reference maps. Figure 6e,f shows the ROI‐based quantitative comparisons conducted between the proposed method and reference maps. In addition, it can also be observed that the estimated T2 values from UTE‐MRF approach the reference T2* as demonstrated in Figure 6, which is consistent with our simulation results.
FIGURE 6.

Physical phantom results. Reconstructed T1 (b) and T2 (d) maps from Cramér–Rao lower bound‐optimized three‐dimensional ultrashort echo time magnetic resonance fingerprinting and reference T1 (a) and T2* (c), with the comparison of T1 (e) and T2/T2* (f) quantification of six phantoms
3.3.2. In vivo studies
Using data from MRF sequence, we could simultaneously obtain PD, T1, and T2 maps. A representative result from a healthy subject is shown in Figure 7. Figure 8 shows the three‐orthogonal tissue segmentation results of SPIJN algorithm for the subject from dual‐echo signals. There are four components matched by the SPIJN algorithm for echo 1 and echo 2. The estimated components from echo 1 are with T1/T2 = 252/0.8 ms, 738/55 ms, 1400/70 ms, and 3300/1800 ms and to be assumed to be related to be myelin‐proton, WM, GM, and cerebrospinal fluid (CSF), respectively. When TE is approximately 3 ms, the myelin‐proton signals were substantially decayed and the estimated components from echo 2 were with T1/T2 = 252/15 ms, 738/55 ms, 1400/70 ms, and 3300/1800 ms assumed as myelin‐water, WM, GM, and CSF, respectively.
FIGURE 7.

Reconstructed proton density (PD), T1, T2 maps from a typical volunteer
FIGURE 8.

Three‐orthogonal tissue segmentation results of sparsity promoting iterative joint nonnegative least squares algorithm from a representative healthy subject. WMF: white matter fraction, GMF: gray matter fraction, CSFF: cerebrospinal fluid (CSF) fraction. There are four components for each echo. Myelin‐proton from the ultrashort echo time (TE) is matched with T1/T2 = 252/0.8 ms with the myelin‐water from the second echo with T1/T2 = 252/15 ms. White matter, gray matter, and CSF were matched with T1/T2 = 738/55, 1400/70, and 3300/1800 ms from both echo signals. The T1 and T2 distribution is shown in the right.
In Figure S5, two repetitive scans of the same subject were compared to assess the reproducibility. The MPF/MWF maps are consistent to each other.
Figure 9 shows two scans of three representative imaging slices obtained from a MS patient using proposed method, and clinical T2‐weighted sequences. The left panel was from the first scan on 21st May, while the right one was results from the second scan on 12th September. The first column shows the images of T2w‐FLAIR and the second to fourth column in each panel are component fractional maps from UTE‐MRF with dual‐echo acquisition. MS lesions were defined with clinical radiological images by higher signal intensity on T2‐weighted images. The lesion areas identified on clinical T2w‐FLAIR images are accurately and clearly exhibited on MPF and MWF, which can be observed as reduction in MPF and MWF. MPF and MWF (white squares [b1, c1, f1, g1, b2, c2]) detected the lesions while WMF (white matter fraction) (red squares [d1, h1, d2]) did not show obvious changes, which demonstrates the possible superiority of MPF and MWF. When looking at the second row, the lesions which were not detected by T2w‐FLIAR (white arrow in [e1]) in the first scan was developed and detected by the second scan (white arrows in [e2]). However, this lesion could be detected as slightly decreased MPF (white circle in [f1], MPF of lesion vs MPF of peripheral area: 0.0501 vs. 0.1242) in the first scan but not showing any obvious changes on MWF (g1, MWF of lesion vs. MWF of peripheral area: 0.1239 vs. 0.1303). With lesions developed, in the second scan, both decreased MPF and MWF depicted the lesion. The different behaviors of MPF and MWF could indicate that MPF may provide new information in early detection of lesion over MWF. Last but not least, when comparing these two scans, MPF/MWF could detect the lesion in the first scan (white circle [f1] and white triangles [j1, k1]) while it could not be detected by the first scan of T2w‐FLAIR (e1, i1). Another MS case is shown in Figure S6. MPF map in Figure S6 also demonstrates that the peripheral areas of larger lesion areas (purple arrows) depicted on T2w‐FLAIR show more dominant demyelination than the smaller lesions (green arrows). This could indicate that MPF from UTE‐MRF shows superiority and has the potential of evaluating the severity of the lesion. However, MWF derived from mGRE seems to show the opposite trend, that is larger demyelination lesion depicted by the green arrow than the purple one Figure S6g.
FIGURE 9.

Two scans of the same multiple sclerosis patient, including T2w‐fluid‐attenuated inversion recovery, myelin‐proton fraction, myelin‐water fraction, WMF from ultrashort echo time magnetic resonance fingerprinting
Figure 10 shows the comparisons of MPF and MWF on three regions of CC between healthy and MS patient groups. Both MPF and MWF showed significant differences between the two groups.
FIGURE 10.

Group comparison on region of interest‐based MPF and MWF. Mean myelin‐proton fraction and myelin‐water fraction on three regions of Corpus Callosum are plotted, with group averages and SE bars shown. An asterisk indicates a significant difference between groups (**p < .01, ***p < .001) based on t‐test.
4. DISCUSSION
In this article, the 3D UTE‐MRF was proposed to simultaneously quantify both the ultrashort T2*, short T2, and long T2 tissues, and to map their fractions, which is summarized in Figure S7. The CRLB method was utilized to design the sequence acquisition parameter patterns for improving ultrashort T2*/T2 component parameter mapping. To extend the 2D UTE‐MRF to 3D, a hard pulse with duration of 0.2 ms was utilized for volume excitation. A multicomponent analysis was utilized to separate the ultrashort T2* tissues, myelin‐proton, from long T2 tissues in WM region. With this development, the whole brain mapping with an isotropic resolution of 1.8 mm could be accomplished in 15 min. The novel sequence was verified and tested by simulation and phantom, as well as in vivo human brain experiments.
Following the optimization, the CRLBs for both T1 and T2 of each tissue had been systematically reduced. In particular, the CRLB of T2 of myelin‐proton was much lower than the original one (from 49.07 [original scheme] to 6.53 [Scheme A]). According to this optimization, it was clearly shown that the CRLB method has the potential of improving the precision for myelin‐proton. This could be explained by the fact that using varied ultrashort TE, which is comparable to T2* of myelin‐proton, the ultrafast T2* decay of myelin‐proton is better encoded in the acquisition process, thus making the signal to be more reflective of the myelin‐proton. Simply put, despite the fact that different initializations result in different acquisition parameter patterns (local minima) shown in Figure S8, they still share qualitative pattern features of reaching their maximum and minimum values, which were similar to previous works (Lee et al., 2019; Zhao et al., 2019), and have very close cost function values, which imply that different initializations may have slight impact on the experiment designs.
MRF can simultaneously quantify multiple parameters by modeling tissue properties into the under‐sampled signal evolutions. In the proposed 3D UTE‐MRF method, the radial trajectory with a short readout duration was used to reduce the T2* blurring. However, it would cause a much higher under‐sampling rate than the spiral arm, resulting in inefficiency of data acquisition. Furthermore, by extending a 2D MRF method into 3D, dramatically increased under‐sampling aliasing degrades the quantification precision. Solutions had been focused on either using advanced reconstruction methods or increasing the k‐space coverage (Cao et al., 2017, 2019, 2022; Liao et al., 2017; Zhao et al., 2018). Here in our study, an optimized 2D golden‐angle method was chosen to cover the 3D k‐space to maximize the trajectory incoherence. Besides, 160 repetitions for each radial spoke with sliding window reconstruction algorithm was used for further reducing the under‐sampling rate, resulting in reduced full width at half maximum of the point spread function (Figure S9).
The simulation results indicate that the proposed CRLB‐optimized 3D UTE‐MRF has the potential to quantifying the ultrashort T2* tissues. Compared with the original 3D UTE‐MRF, the optimized one is shown with much better accuracy for T2 quantification, especially for ultrashort T2* tissue, which verified the improvement of utilizing CRLB method. It should be noted that since ultrashort T2* tissue signals decay totally at the end of each TR, it will not generate stimulated echo as long T2 tissues. Thus, the measured T2 of the ultrashort T2* tissues from the CRLB‐derived 3D UTE‐MRF approaches their nominal T2* values, which has also been reported on the previous study (Li et al., 2019). However, varying TE had little impact on long T2 tissues due to the much smaller TE compared with their T2 and T2*. According to our findings, mappings with 160 repetitions provide a good agreement with references, and 800 repetitions with four times longer scan time will further improve the accuracy. This enables us to “obtain the most important features” out of available scan time of 15 mins, which is an acceptable scan time, while still contains required features. Compared with the previous methods for myelin‐tissue‐related imaging, our method provides better scan efficiency, with higher resolution in comparable scan time (Table S1). It should be noted that in the Table S1, we denoted “direct imaging of myelin” as signals from myelin‐protons while other methods, either signals from myelin‐water or signal alteration related to myelin‐tissue, were denoted as “indirect imaging.”
In addition, we also took MT effects between myelin‐proton and WM into account. MT effect of MRF mainly came from the adiabatic IR pulse at the beginning. We used EPG‐X for the dictionary generation as stated in Hilbert et al. (2020) and Malik et al. (2018). The results show that MT effects are negligible when comparing between the reconstructed results of the simulations with/without MT effects considered, which is shown in Figure S10. This could be explained by the fact that the MT phenomenon only affects the first few frames after adiabatic IR pulse. Further, with ultrashort TE, we could obtain the signals from myelin‐proton, cancelling out the part of the MT effects. Our results are also consistent with the paper (Liu et al., 2017) which stated that the MT effects are more severe with larger TE.
For in vivo studies, in the multicomponent model analysis, we chose the SPIJN algorithm which assumes no prior number of tissue components and the properties of the tissues in each voxel. Here for in vivo data, the results of the ultrashort echo 1 were converged to four components with myelin‐proton (T1 = 252 ms, T2 = 0.8 ms), WM (T1 = 738 ms, T2 = 55 ms), GM (T1 = 1400 ms, T2 = 70 ms), and CSF (T1 = 3300 ms, T2 = 1800 ms). With a second TE of approximately 3 ms, myelin‐water could also be matched with T1 = 252 ms, T2 = 15 ms. The result from the repetition experiment also supported the reliability and reproducibility of the proposed sequence. For the MS patients, the MPF and MWF maps showed obvious reduction in MS lesions which had been depicted with clinical MRI sequences. More importantly, the normal‐appearing tissue around the lesions shown in clinical T2‐weighted is clearly demonstrated as the myelin loss on MPF maps (white circle on Figure 9f1, white triangle on Figure 9j1, k1), which developed and later detected by the clinical T2‐FLAIR sequence. These results may imply that directly imaging myelin‐proton has superiority in sensitivity to demyelination, leading to the possibility of earlier detection and monitoring disease progress using our propose sequence, which is consistent with previous studies (Sheth et al., 2016, 2017).
In the in vivo study, we did not extract the myelin‐water component from the ultrashort echo signal, which is partially consistent with the in vivo results from other 3 T studies (Cui et al., 2021; Nagtegaal et al., 2020). The result in Cui et al. (2021) suggested that with single IR pulses, myelin‐water component is not easy to be extracted. However, the MPF results were estimated with little bias. When the TE reached approximately 3 ms, the myelin‐proton signal with ultrashort T2* is substantially decayed, and predominated short T1/T2 component is mainly from myelin‐water component, which was matched with SPIJN algorithm. Thus, our proposed UTE‐MRF sequence with dual‐echo acquisition could capture both myelin‐proton and myelin‐water components. When comparing ROI‐based MPF/MWF in CC in the healthy and MS patient groups, we found significant differences, which demonstrate potential value of our methods in clinical use.
There are several limitations in our work. First, due to the limit of the highly under‐sampling rate of radial trajectories, it is of great challenge to improve the resolution. For further increasing the spatial resolution, we could optimize the sequence by the following potential approaches. (1) Higher gradient amplitude could be used to reach higher kmax at each radial spoke; (2) Reducing the full width at half maximum by optimizing the radial sampling k‐space to achieve higher incoherence at each MRF time point; (3) Increasing radial spokes; however, the scan time will also be increased; therefore, advanced reconstruction methods, such as subspace reconstruction for 3D highly accelerated MRF data (Cao et al., 2022; Hu et al., 2021; Zhao et al., 2018) should be utilized to reduce the scan time; (4) Increasing the sampling efficiency using radial cone trajectory (Johnson, 2017) or modified radial trajectories by deep learning (Peng et al., 2022), while maintaining the acquisition quality at central k‐space for ultrashort T2* components. However, the 3D whole brain MPF and MWF maps obtained from our proposed method can still provide the valuable data in clinical friendly scanning time, such as possibility of earlier detection of lesions, which might be implemented as an additional routine clinical scanning. Second, we did not extract myelin‐water component from the ultrashort echo signal. However, it was extracted from the echo 2. Further study should consider using the multi‐IR pulses for myelin‐water component extraction and for better MPF estimation. Third, water exchange between tissue components has not been considered in the simulation. In our views, water exchange could have impacts on the signals of intra‐axonal and extra‐axonal components in WM with the diffusion sequences (which is with TE of tens of milliseconds) and thus leading to the fractional estimation biases (Li et al., 2017). However, both of these two compartments are with T2 of tens of milliseconds (Gong et al., 2020) and are grouped into the long‐T2 tissue component group in our study which could have less effects on fractional mapping of myelin‐protons and myelin‐water. Furthermore, it is believed that with larger TE, the impacts become more prominent. In this article, with ultrashort TE of 0.05 ms, the impact of water exchange could be neglected. Future studies should be conducted with water exchange considered.
5. CONCLUSIONS
The CRLB‐optimized 3D UTE‐MRF method with dual‐echo radial acquisition was proposed to quantify and directly image myelin‐proton, which were validated in three different settings, including simulations, physical phantom, and in vivo experiments. With the second echo of approximately 3 ms, myelin‐water component could also be obtained with SPIJN algorithm. The studies of 10 healthy volunteers and 10 MS patients demonstrated the capability and sensitivity of the proposed method in detecting myelin loss, which may be helpful for detecting an early brain abnormality. The sequence with the whole brain MPF map obtained in clinical friendly scanning time may be implemented as an additional routine clinical scanning tool, in aiding more accurate diagnosis.
AUTHOR CONTRIBUTION
Zihan Zhou: Conceptualization, Software, Formal analysis, Investigation, Writing‐original draft, Writing‐review & editing, Visualization. Qing Li: Conceptualization, Software, Investigation, Writing‐review & editing. Congyu Liao: Conceptualization, Software, Writing‐review & editing. Xiaozhi Cao: Software, Writing‐review & editing. Hui Liang: Writing‐review. Quan Chen: Software. Run Pu: Software. Huihui Ye: Writing‐review & editing. Qiqi Tong: Writing‐review & editing, Funding acquisition. Hongjian He: Conceptualization, Writing‐review & editing, Supervision, Visualization, Project administration, Funding acquisition. Jianhui Zhong: Conceptualization, Writing‐review & editing, Visualization, Supervision, Project administration, Funding acquisition.
ACKNOWLEDGEMENT
The authors thank Dr. Liang hui for the MS patients recruitments.This article was supported by the National Key R&D Program of China (2020AAA0109502 and 2019YFB1311800), National Natural Science Foundation of China (81871428), Fundamental Research Funds for the Central Universities (2021FZZX002‐19), the Youth Innovation Team Project of the College of Biomedical Engineering & Instrument Science, and Major Scientific Project of Zhejiang Lab (No. 2020ND8AD01), Open Research Fund of the State Key Laboratory of Cognitive Neuroscience and Learning.
CONFLICT OF INTEREST
The authors declare there is no conflicts of interest.
Supporting information
Data S1: Supporting Information
Zhou, Z. , Li, Q. , Liao, C. , Cao, X. , Liang, H. , Chen, Q. , Pu, R. , Ye, H. , Tong, Q. , He, H. , & Zhong, J. (2023). Optimized three‐dimensional ultrashort echo time: Magnetic resonance fingerprinting for myelin tissue fraction mapping. Human Brain Mapping, 44(6), 2209–2223. 10.1002/hbm.26203
Zihan Zhou and Qing Li contributed equally to this study.
Funding information Fundamental Research Funds for the Central Universities, Grant/Award Number: 2021FZZX002‐19; Major Scientific Project of Zhejiang Laboratory, Grant/Award Number: 2020ND8AD01; National Natural Science Foundation of China, Grant/Award Number: 81871428; Open Research Fund of the State Key Laboratory of Cognitive Neuroscience and Learning; the National Key R&D Program of China, Grant/Award Numbers: 2019YFB1311800, 2020AAA0109502
DATA AVAILABILITY STATEMENT
All data, models, or code generated or used in this article are available from the corresponding author by request.
REFERENCES
- Alonso‐Ortiz, E. , Levesque, I. R. , & Pike, G. B. (2018). Multi‐gradient‐echo myelin water fraction imaging: Comparison to the multi‐echo‐spin‐echo technique. Magnetic Resonance in Medicine, 79, 1439–1446. [DOI] [PubMed] [Google Scholar]
- Bachmann, R. , Reilmann, R. , Schwindt, W. , Kugel, H. , Heindel, W. , & Kramer, S. (2006). FLAIR imaging for multiple sclerosis: A comparative MR study at 1.5 and 3.0 tesla. European Radiology, 16, 915–921. [DOI] [PubMed] [Google Scholar]
- Boyacioglu, R. , Wang, C. , Ma, D. , McGivney, D. F. , Yu, X. , & Griswold, M. A. (2021). 3D magnetic resonance fingerprinting with quadratic RF phase. Magnetic Resonance in Medicine, 85, 2084–2094. [DOI] [PubMed] [Google Scholar]
- Buonincontri, G. , Kurzawski, J. W. , Kaggie, J. D. , Matys, T. , Gallagher, F. A. , Cencini, M. , Donatelli, G. , Cecchi, P. , Cosottini, M. , Martini, N. , Frijia, F. , Montanaro, D. , Gomez, P. A. , Schulte, R. F. , Retico, A. , & Tosetti, M. (2021). Three dimensional MRF obtains highly repeatable and reproducible multi‐parametric estimations in the healthy human brain at 1.5T and 3T. NeuroImage, 226, 117573. [DOI] [PubMed] [Google Scholar]
- Cao, X. , Liao, C. , Iyer, S. S. , Wang, Z. , Zhou, Z. , Dai, E. , Liberman, G. , Dong, Z. , Gong, T. , He, H. , Zhong, J. , Bilgic, B. , & Setsompop, K. (2022). Optimized multi‐axis spiral projection MR fingerprinting with subspace reconstruction for rapid whole‐brain high‐isotropic‐resolution quantitative imaging. Magnetic Resonance in Medicine, 88(1), 133–150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cao, X. , Liao, C. , Wang, Z. , Chen, Y. , Ye, H. , He, H. , & Zhong, J. (2017). Robust sliding‐window reconstruction for accelerating the acquisition of MR fingerprinting. Magnetic Resonance in Medicine, 78, 1579–1588. [DOI] [PubMed] [Google Scholar]
- Cao, X. , Ye, H. , Liao, C. , Li, Q. , He, H. , & Zhong, J. (2019). Fast 3D brain MR fingerprinting based on multi‐axis spiral projection trajectory. Magnetic Resonance in Medicine., 82(1), 289–301. [DOI] [PubMed] [Google Scholar]
- Chen, Q. , She, H. , & Du, Y. P. (2021). Whole brain myelin water mapping in one minute using tensor dictionary learning with low‐rank plus sparse regularization. IEEE Transactions on Medical Imaging, 40, 1253–1266. [DOI] [PubMed] [Google Scholar]
- Chen, Y. , Chen, M. H. , Baluyot, K. R. , Potts, T. M. , Jimenez, J. , Lin, W. , & U.U.B.C.P. Consortium . (2019). MR fingerprinting enables quantitative measures of brain tissue relaxation times and myelin water fraction in the first five years of life. NeuroImage, 186, 782–793. [DOI] [PubMed] [Google Scholar]
- Chen, Y. , Panda, A. , Pahwa, S. , Hamilton, J. I. , Dastmalchian, S. , McGivney, D. F. , Ma, D. , Batesole, J. , Seiberlich, N. , Griswold, M. A. , Plecha, D. , & Gulani, V. (2019). Three‐dimensional MR fingerprinting for quantitative breast imaging. Radiology, 290, 33–40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cui, D. , Hui, E. S. , & Cao, P. (2021). A multi‐inversion‐recovery magnetic resonance fingerprinting for multi‐compartment water mapping. Magnetic Resonance Imaging, 81, 82–87. [DOI] [PubMed] [Google Scholar]
- Kim, D. , Wisnowski, J. L. , & Haldar, J. P. (2017). Improved efficiency for microstructure imaging using high‐dimensional MR correlation spectroscopic imaging. In 51st Asilomar conference on signals, systems, and computers (pp. 1264–1268). [Google Scholar]
- Deoni, S. C. , Peters, T. M. , & Rutt, B. K. (2005). High‐resolution T1 and T2 mapping of the brain in a clinically acceptable time with DESPOT1 and DESPOT2. Magnetic Resonance in Medicine, 53, 237–241. [DOI] [PubMed] [Google Scholar]
- Deshmane, A. , McGivney, D. , Badve, C. , Gulani, V. , & Griswold, M. A. (2017). Dictionary approach to partial volume estimation with MR fingerprinting: Validation and application to brain tumor segmentation [Conference presentation]. 25th Annual Meeting and Exhibition of the International Society for Magnetic Resonance in Medicine. Honolulu, HI, USA.
- Does, M. D. , & Gore, J. C. (2000). Rapid acquisition transverse relaxometric imaging. Journal of Magnetic Resonance, 147, 116–120. [DOI] [PubMed] [Google Scholar]
- Du, J. , Ma, G. , Li, S. , Carl, M. , Szeverenyi, N. M. , VandenBerg, S. , Corey‐Bloom, J. , & Bydder, G. M. (2014). Ultrashort echo time (UTE) magnetic resonance imaging of the short T2 components in white matter of the brain using a clinical 3T scanner. NeuroImage, 87, 32–41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Du, J. , Sheth, V. , He, Q. , Carl, M. , Chen, J. , Corey‐Bloom, J. , & Bydder, G. M. (2014). Measurement of T1 of the ultrashort T2* components in white matter of the brain at 3T. PLoS One, 9, e103296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Du, Y. , Chu, R. , Hwang, D. , Brown, M. S. , Kleinschmidt‐DeMasters, B. K. , Singel, D. , & Simon, J. H. (2007). Fast multislice mapping of the myelin water fraction using multicompartment analysis of T2* decay at 3T: A preliminary postmortem study. Magnetic Resonance in Medicine, 58, 865–870. [DOI] [PubMed] [Google Scholar]
- Fan, S. , Ma, Y. , Chang, E. Y. , Bydder, G. M. , & Du, J. (2017). Inversion recovery ultrashort echo time imaging of ultrashort T2 tissue components in ovine brain at 3 T: A sequential D2 O exchange study. NMR in Biomedicine, 30, e3767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fessler, J. A. , & Sutton, B. P. (2003). Nonuniform fast fourier transforms using min‐max interpolation. IEEE Transactions on Signal Processing, 51, 560–574. [Google Scholar]
- Gong, T. , Tong, Q. , He, H. , Sun, Y. , Zhong, J. , & Zhang, H. (2020). MTE‐NODDI: Multi‐TE NODDI for disentangling non‐T2‐weighted signal fractions from compartment‐specific T2 relaxation times. NeuroImage, 217, 116906. [DOI] [PubMed] [Google Scholar]
- Hilbert, T. , Xia, D. , Block, K. T. , Yu, Z. , Lattanzi, R. , Sodickson, D. K. , Kober, T. , & Cloos, M. A. (2020). Magnetization transfer in magnetic resonance fingerprinting. Magnetic Resonance in Medicine, 84, 128–141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horch, R. A. , Gochberg, D. F. , Nyman, J. S. , & Does, M. D. (2012). Clinically compatible MRI strategies for discriminating bound and pore water in cortical bone. Magnetic Resonance in Medicine, 68, 1774–1784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hu, Y. , Li, P. , Chen, H. , Zou, L. , & Wang, H. (2021). High‐quality MR fingerprinting reconstruction using structured low‐rank matrix completion and subspace projection. IEEE Transactions on Medical Imaging, 41(5), 1150–1164. [DOI] [PubMed] [Google Scholar]
- Jiang, Y. , Ma, D. , Jerecic, R. , Duerk, J. , Seiberlich, N. , Gulani, V. , & Griswold, M. A. (2017). MR fingerprinting using the quick echo splitting NMR imaging technique. Magnetic Resonance in Medicine, 77, 979–988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang, Y. , Ma, D. , Seiberlich, N. , Gulani, V. , & Griswold, M. A. (2015). MR fingerprinting using fast imaging with steady state precession (FISP) with spiral readout. Magnetic Resonance in Medicine, 74, 1621–1631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson, K. M. , (2017). Hybrid radial‐cones trajectory for accelerated MRI. Magnetic Resonance in Medicine, 77, 1068–1081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jordan, S. P. , Hu, S. , Rozada, I. , McGivney, D. F. , Boyacioglu, R. , Jacob, D. C. , Huang, S. , Beverland, M. , Katzgraber, H. G. , Troyer, M. , Griswold, M. A. , & Ma, D. (2021). Automated design of pulse sequences for magnetic resonance fingerprinting using physics‐inspired optimization. Proceedings of the National Academy of Sciences of the United States of America, 118, e2020516118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Korzdorfer, G. , Kirsch, R. , Liu, K. , Pfeuffer, J. , Hensel, B. , Jiang, Y. , Ma, D. , Gratz, M. , Bar, P. , Bogner, W. , Springer, E. , Lima Cardoso, P. , Umutlu, L. , Trattnig, S. , Griswold, M. , Gulani, V. , & Nittka, M. (2019). Reproducibility and repeatability of MR fingerprinting relaxometry in the human brain. Radiology, 292, 429–437. [DOI] [PubMed] [Google Scholar]
- Lahiri, A. , Fessler, J. A. , & Hernandez‐Garcia, L. (2020). Optimizing MRF‐ASL scan design for precise quantification of brain hemodynamics using neural network regression. Magnetic Resonance in Medicine, 83, 1979–1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laule, C. , Vavasour, I. M. , Kolind, S. H. , Li, D. K. B. , Traboulsee, T. L. , Moore, G. R. W. , & MacKay, A. L. (2007). Magnetic resonance imaging of myelin. Neurotherapeutics, 4, 460–484. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee, P. K. , Watkins, L. E. , Anderson, T. I. , Buonincontri, G. , & Hargreaves, B. A. (2019). Flexible and efficient optimization of quantitative sequences using automatic differentiation of Bloch simulations. Magnetic Resonance in Medicine, 82, 1438–1451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li, H. , Jiang, X. , Xie, J. , Gore, J. C. , & Xu, J. (2017). Impact of transcytolemmal water exchange on estimates of tissue microstructural properties derived from diffusion MRI. Magnetic Resonance in Medicine, 77, 2239–2249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li, Q. , Cao, X. , Ye, H. , Liao, C. , He, H. , & Zhong, J. (2019). Ultrashort echo time magnetic resonance fingerprinting (UTE‐MRF) for simultaneous quantification of long and ultrashort T2 tissues. Magnetic Resonance in Medicine, 82, 1359–1372. [DOI] [PubMed] [Google Scholar]
- Li, Q. , Cao, X. , Ye, H. , Zhou, Z. , He, H. , & Zhong, J. (2020). 3D UTE‐MRF for multiple parameteric maps with sub‐millimeter isotropic resolution using multi‐dimensional golden‐angle radial trajectory. International Society for Magnetic Resonance in Medicine. [Google Scholar]
- Liao, C. , Bilgic, B. , Manhard, M. K. , Zhao, B. , Cao, X. , Zhong, J. , Wald, L. L. , & Setsompop, K. (2017). 3D MR fingerprinting with accelerated stack‐of‐spirals and hybrid sliding‐window and GRAPPA reconstruction. NeuroImage, 162, 13–22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liao, C. , Cao, X. , Gong, T. , Wu, Z. , Zhou, Z. , He, H. , Zhong, J. , & Setsompop, K. (2021). High‐resolution myelin‐water fraction (MWF) and T1/T2/proton‐density mapping using 3D ViSTa‐MR fingerprinting with subspace reconstruction. International Society for Magnetic Resonance in Medicine. Virtual meeting. [Google Scholar]
- Liao, C. , Wang, K. , Cao, X. , Li, Y. , Wu, D. , Ye, H. , Ding, Q. , He, H. , & Zhong, J. (2018). Detection of lesions in mesial temporal lobe epilepsy by using MR fingerprinting. Radiology, 288, 804–812. [DOI] [PubMed] [Google Scholar]
- Liu, F. , Velikina, J. V. , Block, W. F. , Kijowski, R. , & Samsonov, A. A. (2017). Fast realistic MRI simulations based on generalized multi‐Pool exchange tissue model. IEEE Transactions on Medical Imaging, 36, 527–537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma, D. , Coppo, S. , Chen, Y. , McGivney, D. F. , Jiang, Y. , Pahwa, S. , Gulani, V. , & Griswold, M. A. (2017). Slice profile and B1 corrections in 2D magnetic resonance fingerprinting. Magnetic Resonance in Medicine, 78, 1781–1789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma, D. , Gulani, V. , Seiberlich, N. , Liu, K. , Sunshine, J. L. , Duerk, J. L. , & Griswold, M. A. (2013). Magnetic resonance fingerprinting. Nature, 495, 187–192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma, D. , Jiang, Y. , Chen, Y. , McGivney, D. , Mehta, B. , Gulani, V. , & Griswold, M. (2018). Fast 3D magnetic resonance fingerprinting for a whole‐brain coverage. Magnetic Resonance in Medicine, 79, 2190–2197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma, Y. , Jang, H. , Wei, Z. , Cai, Z. , Xue, Y. , Lee, R. R. , Chang, E. Y. , Bydder, G. M. , Corey‐Bloom, J. , & Du, J. (2020). Myelin imaging in human brain using a short repetition time adiabatic inversion recovery prepared ultrashort Echo time (STAIR‐UTE) MRI sequence in multiple sclerosis. Radiology, 297, 392–404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma, Y. , Searleman, A. C. , Jang, H. , Wong, J. , Chang, E. Y. , Corey‐Bloom, J. , Bydder, G. M. , & Du, J. (2020). Whole‐brain myelin imaging using 3D double‐echo sliding inversion recovery ultrashort echo time (DESIRE UTE) MRI. Radiology, 294, 362–374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma, Y. , Zhao, W. , Wan, L. , Guo, T. , Searleman, A. , Jang, H. , Chang, E. Y. , & Du, J. (2019). Whole knee joint T1 values measured in vivo at 3T by combined 3D ultrashort echo time cones actual flip angle and variable flip angle methods. Magnetic Resonance in Medicine, 81, 1634–1644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- MacKay, A. , Whittall, K. , Adler, J. , Li, D. , Paty, D. , & Graeb, D. (1994). In vivo visualization of myelin water in brain by magnetic resonance. Magnetic Resonance in Medicine, 31, 673–677. [DOI] [PubMed] [Google Scholar]
- Malik, S. J. , Teixeira, R. , & Hajnal, J. V. (2018). Extended phase graph formalism for systems with magnetization transfer and exchange. Magnetic Resonance in Medicine, 80, 767–779. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muller, M. , Egger, N. , Sommer, S. , Wilferth, T. , Meixner, C. R. , Laun, F. B. , Mennecke, A. , Schmidt, M. , Huhn, K. , Rothhammer, V. , Uder, M. , Dorfler, A. , & Nagel, A. M. (2021). Direct imaging of white matter ultrashort T2(*) components at 7 Tesla. Magnetic Resonance Imaging, 86, 107–117. [DOI] [PubMed] [Google Scholar]
- Nagtegaal, M. , Koken, P. , Amthor, T. , de Bresser, J. , Madler, B. , Vos, F. , & Doneva, M. (2020). Myelin water imaging from multi‐echo T2 MR relaxometry data using a joint sparsity constraint. NeuroImage, 219, 117014. [DOI] [PubMed] [Google Scholar]
- Nair, G. , Absinta, M. , & Reich, D. S. (2013). Optimized T1‐MPRAGE sequence for better visualization of spinal cord multiple sclerosis lesions at 3T. AJNR. American Journal of Neuroradiology, 34, 2215–2222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oh, S. H. , Bilello, M. , Schindler, M. , Markowitz, C. E. , Detre, J. A. , & Lee, J. (2013). Direct visualization of short transverse relaxation time component (ViSTa). NeuroImage, 83, 485–492. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peng, W. , Feng, L. , Zhao, G. , & Liu, F. (2022). Learning optimal k‐space acquisition and reconstruction using physics‐informed neural networks [Conference presentation]. IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), New Orleans, Louisiana, US, 20794–20803.
- Peters, A. M. , Brookes, M. J. , Hoogenraad, F. G. , Gowland, P. A. , Francis, S. T. , Morris, P. G. , & Bowtell, R. (2007). T2* measurements in human brain at 1.5, 3 and 7 T. Magnetic Resonance Imaging, 25, 748–753. [DOI] [PubMed] [Google Scholar]
- Qian, Y. , & Boada, F. E. (2008). Acquisition‐weighted stack of spirals for fast high‐resolution three‐dimensional ultra‐short echo time MR imaging. Magnetic Resonance in Medicine, 60, 135–145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scope Crafts, E. , Lu, H. , Ye, H. , Wald, L. L. , & Zhao, B. (2022). An efficient approach to optimal experimental design for magnetic resonance fingerprinting with B‐splines. Magnetic Resonance in Medicine, 88, 239–253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seifert, A. C. , Li, C. , Wilhelm, M. J. , Wehrli, S. L. , & Wehrli, F. W. (2017). Towards quantification of myelin by solid‐state MRI of the lipid matrix protons. NeuroImage, 163, 358–367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sheth, V. , Shao, H. , Chen, J. , Vandenberg, S. , Corey‐Bloom, J. , Bydder, G. M. , & Du, J. (2016). Magnetic resonance imaging of myelin using ultrashort Echo time (UTE) pulse sequences: Phantom, specimen, volunteer and multiple sclerosis patient studies. NeuroImage, 136, 37–44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sheth, V. R. , Fan, S. , He, Q. , Ma, Y. , Annese, J. , Switzer, R. , Corey‐Bloom, J. , Bydder, G. M. , & Du, J. (2017). Inversion recovery ultrashort echo time magnetic resonance imaging: A method for simultaneous direct detection of myelin and high signal demonstration of iron deposition in the brain ‐ A feasibility study. Magnetic Resonance Imaging, 38, 87–94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waldman, A. , Rees, J. H. , Brock, C. S. , Robson, M. D. , Gatehouse, P. D. , & Bydder, G. M. (2003). MRI of the brain with ultra‐short echo‐time pulse sequences. Neuroradiology, 45, 887–892. [DOI] [PubMed] [Google Scholar]
- Wansapura, J. P. , Holland, S. K. , Dunn, R. S. , & Ball, W. S. (1999). NMR relaxation times in the human brain at 3.0 Tesla. Journal of Magnetic Resonance Imaging, 9, 531–538. [DOI] [PubMed] [Google Scholar]
- Weigel, M. (2015). Extended phase graphs: Dephasing, RF pulses, and echoes ‐ Pure and simple. Journal of Magnetic Resonance Imaging, 41, 266–295. [DOI] [PubMed] [Google Scholar]
- Weiger, M. , Brunner, D. O. , Dietrich, B. E. , Muller, C. F. , & Pruessmann, K. P. (2013). ZTE imaging in humans. Magnetic Resonance in Medicine, 70, 328–332. [DOI] [PubMed] [Google Scholar]
- Weiger, M. , Froidevaux, R. , Baadsvik, E. L. , Brunner, D. O. , Rosler, M. B. , & Pruessmann, K. P. (2020). Advances in MRI of the myelin bilayer. NeuroImage, 217, 116888. [DOI] [PubMed] [Google Scholar]
- Wu, Z. , Bilgic, B. , He, H. , Tong, Q. , Sun, Y. , Du, Y. , Setsompop, K. , & Zhong, J. (2018). Wave‐CAIPI ViSTa: Highly accelerated whole‐brain direct myelin water imaging with zero‐padding reconstruction. Magnetic Resonance in Medicine, 80, 1061–1073. [DOI] [PubMed] [Google Scholar]
- Wu, Z. , He, H. , Sun, Y. , Du, Y. , & Zhong, J. (2017). High resolution myelin water imaging incorporating local tissue susceptibility analysis. Magnetic Resonance Imaging, 42, 107–113. [DOI] [PubMed] [Google Scholar]
- Zhang, J. , Feng, L. , Otazo, R. , & Kim, S. G. (2019). Rapid dynamic contrast‐enhanced MRI for small animals at 7T using 3D ultra‐short echo time and golden‐angle radial sparse parallel MRI. Magnetic Resonance in Medicine, 81, 140–152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao, B. , Haldar, J. P. , Congyu, L. , Dan, M. , Yun, J. , Griswold, M. A. , Setsompop, K. , & Wald, L. L. (2019). Optimal experiment design for magnetic resonance fingerprinting: Cramer‐Rao bound meets spin dynamics. IEEE Transactions on Medical Imaging, 38, 844–861. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao, B. , Setsompop, K. , Adalsteinsson, E. , Gagoski, B. , Ye, H. , Ma, D. , Jiang, Y. , Ellen Grant, P. , Griswold, M. A. , & Wald, L. L. (2018). Improved magnetic resonance fingerprinting reconstruction with low‐rank and subspace modeling. Magnetic Resonance in Medicine, 79, 933–942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zou, L. , Liang, D. , Ye, H. , Su, S. , Zhu, Y. , Liu, X. , Zheng, H. , & Wang, H. (2021). Quantitative MR relaxation using MR fingerprinting with fractional‐order signal evolution. Journal of Magnetic Resonance, 330, 107042. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data S1: Supporting Information
Data Availability Statement
All data, models, or code generated or used in this article are available from the corresponding author by request.


