Abstract
There have been an increasing number of patients with degenerative disc diseases due to the aging population. In light of this, studies on the pathogenesis of intervertebral disc degeneration have become a hot topic, and gene knockout mice have become a valuable tool in this field of research. With the development of science and technology, constitutive gene knockout mice can be constructed using homologous recombination, zinc finger nuclease, transcription activator‐like effector nuclease technology and clustered regularly interspaced short palindromic repeats/Cas9 (CRISPR/Cas9) system, and conditional gene knockout mice can be constructed using the Cre/LoxP system. The gene‐edited mice using these techniques have been widely used in the studies on disc degeneration. This paper reviews the development process and principles of these technologies, functions of the edited genes in disc degeneration, advantages, and disadvantages of different methods and possible targets of the specific Cre recombinase in intervertebral discs. Recommendations for the choice of suitable gene‐edited model mice are presented. At the same time, possible technological improvements in the future are also discussed.
Keywords: construction, gene edition technology, gene targeting, guideline, intervertebral disc degeneration, knockout, mouse model, specific Cre recombinase
In this review, we described the development process and principles of the gene knockout technologies, functions of the edited genes in disc degeneration, advantages and disadvantages of different methods and possible targets of the specific Cre recombinase in intervertebral discs. Recommendations for the choice of suitable gene‐edited model mice for the study of disc degeneration are presented. At the same time, possible technological improvements in the future are also discussed.

1. INTRODUCTION
Low back pain (LBP) is a common musculoskeletal symptom that can take a huge toll on people's health, quality of life and economics. About 85.3% of people experience LBP at some time point in their lives. 1 Intervertebral disc degeneration (IVDD) is one of the important causes of LBP. 2 Studies have found that the cause of disc degeneration is multifactorial. Among them, genetics, biological, chemical, biomechanical, aging, and so forth, all take part in the pathogenesis. 3 However, the exact mechanism of IVDD is still not clarified.
From the anatomical point of view, intervertebral disc (IVD) consists of three main parts: the nucleus pulposus (NP), which is located in the middle of the IVD structure and serves to offset and transfer the axial pressure load of the spine. The annulus fibrosus (AF) that surrounds the NP and prevents it from herniating during spinal contortion. Two hyaline cartilaginous endplates (CEPs) which connect the upper and lower vertebrae and play an important role in the exchange of nutrients and metabolic waste to the IVD. 4 , 5 , 6 The molecular composition and property of different parts of the IVD are also different. NP cells can synthesize extracellular matrix (ECM), which is rich in type II collagen, proteoglycan, and elastin. AF is composed of alternating type I collagen fibers, and the ECM in the CEP is rich in proteoglycan and collagen fibers. 7 Different gene expressions in different tissues or cells provides a reason for making animal models by means of gene edition. With the development of genetic engineering technology in recent years, it has become a reliable method to use gene edition model animals to study the pathogenesis and verify the efficacy of treatment for IVDD.
Gene edition technology in animals is a product of genetic engineering and embryo engineering. Over the past few decades, gene‐edited animals have become increasingly common in studies of life science. At present, the gene‐editing technologies we use mainly include gene knockout (KO) technology, gene knock‐in (KI) technology, RNA interference (RNAi) technology, transgenic technology, and animal humanization technology. 8 The gene knockout mice are the most commonly used in the study of IVDD. According to the principle and scope of gene knockout, it can be divided into constitutive gene knockout (KO) and conditional gene knockout (cKO). Many practical scientific technologies, such as the classical homologous recombination (HR) technology, the newly‐developing zinc finger nuclease (ZFN) and transcription activator‐like effector nuclease (TALEN) technology, the current mainstream CRISPR/Cas9 technology, and the highly specific Cre/LoxP system, have made great contributions to the production of gene knockout mice to study disc degeneration.
Here, we summarize the technical principles, the advantages, and disadvantages of these gene knockout mice, as well as their applications in IVD research.
2. CONSTITUTIVE KO MICE
Constitutive KO mice refers to a mouse model that does not express the gene in all tissues or cells of the body by knocking out several important exons or functional regions or even all exons of the target gene through gene knockout technology.
2.1. Technical principles and characteristics
Nowadays, the most commonly used four techniques to completely knockout the target gene in the mice are the classic HR technique, the ZFN technique, the TALEN technique and the CRISPR/Cas9 system.
2.1.1. Classic HR technique
In 1989, Capecchi, 9 , 10 an American scientist, realized the site‐specific integration of exogenous genes of embryonic stem (ES) cells for the first time, and successfully obtained gene knockout mice. Although decades have passed, the traditional gene recombination technology is still the most common and classical method to explore gene function and construct animal models of human diseases. However, this method has a long cycle and is difficult to construct the target carrier, which costs a lot of manpower and material resources and has a low success rate 11 , 12 (Table 1).
TABLE 1.
Comparison of different gene knockout techniques
| Techniques | The authors and timing of the earlier paper that reported the technique | Advantages | Disadvantages |
|---|---|---|---|
| Constitutive KO | Completely knock out the target gene in the embryonic stage of mice. | Poor imitation of human gene mutation, potential for functional redundancy, embryo death | |
| HR | Capecchi (1989) 10 | Be widely used and the most classic | Long cycle, difficult to construct the target carrier, cost a lot, and low success rate |
| ZFN | Bibikova et al. (2002) 17 | Efficient and high specificity | High off target rate, high cytotoxicity, and high cost. |
| TALEN |
Moscou et al. (2009) 23 Boch et al. (2009) 24 |
TALEN is simpler to design and more specific than ZFN technology. | High cytotoxicity and high cost |
| CRISPR/Cas9 | Qi et al. (2013) 38 | Precise targeting, high efficiency, low cytotoxicity, low cost, and simplicity | Its specificity is not high and its safety is controversial. |
| Conditional KO | Conditional gene knockout achieves specificity of time and sites. | Cre may be expressed in tissues other than target site, causing unpredictable phenotypes, and impact on experiments. | |
| Cre/LoxP | Gu et al. (1994) 81 | Cre/LoxP realizes the specificity of gene knockout site and time. |
Flox mice are complicated and expensive to produce. |
Abbreviations: CRISPR/Cas9, clustered regularly interspaced short palindromic repeats/Cas9; HR, homologous recombination; KO, knockout; TALEN, transcription activator‐like effector nuclease; ZFN, zinc finger nuclease.
The principle of DNA HR is usually used to replace the target gene fragment with the designed homologous fragment, so as to achieve the goal of gene knockout. Using HR principle to make gene knockout mice in the study of disc degeneration is not complicated, which can be done in the following steps: (a) Construction of gene vector. 13 (b) The vector is introduced into homologous ES cells and recombined with the corresponding part of ES cell genome. 14 (c) The hit cells are selected by positive and negative selection (PNS) or polymerase chain reaction (PCR). (d) Reconstruction is constructed by blastocyst injection. (e) Mating chimeric mice to obtain homozygous gene knockout mice 15 (Figure 1).
FIGURE 1.
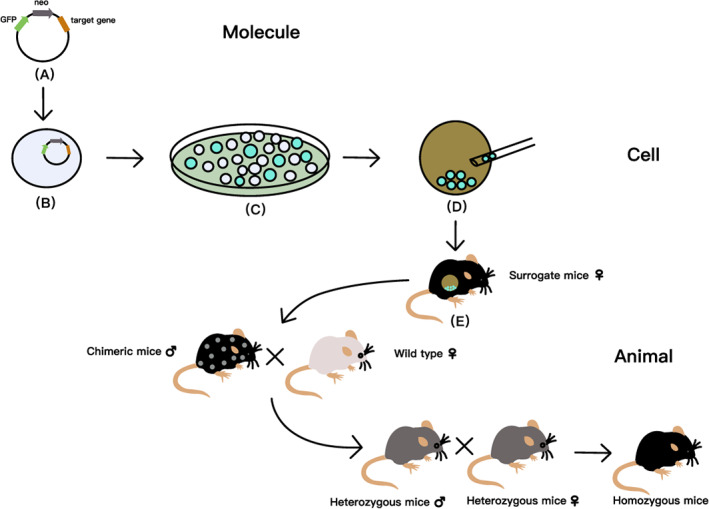
Steps of making knockout mice by homologous recombination (HR) principle. (A) Construction of gene vector. The target gene is recombined into a vector with marker genes such as neo or tk, so that it loses its physiological function and becomes a recombinant vector. (B) The vector is introduced into homologous embryonic stem (ES) cells in a certain way, and the exogenous DNA is recombined with the corresponding part of ES cell genome. (C) The hit cells are selected by positive and negative selection (PNS) or polymerase chain reaction (PCR). (D) Reconstruction is completed by blastocyst injection. (E) Chimeric animals are obtained after embryo transplantation, and then mating chimeric mice to obtain homozygous gene knockout mice. GFP, green fluorescent protein
2.1.2. Zinc finger nuclease
The ZFN technology has many functions, such as the ability to knock out, integrate, or modify target genes. 16 , 17 The structure of ZFN consists of two parts: DNA‐binding domain and the DNA‐cutting domain. 18 The DNA‐binding domain contains 3–6 zinc finger (ZF) repeats, each of which recognizes multiple base pairs. 19 We assemble ZF structures as modular assemblies. When ZF protein structure perfectly recognize the specific DNA sequence and dimerized FokI that contained in two DNA‐cutting domains at the same location of the targeted site‐specific double‐stranded DNA (dsDNA), DNA double‐strand break (DSB) is realized, thus completing gene knockout 20 , 21 (Figure 2A,C). ZFN is more efficient than HR and has high specificity, but it cannot replace traditional technology due to its high off target rate, high cytotoxicity and high cost 19 , 20 , 22 (Table 1).
FIGURE 2.
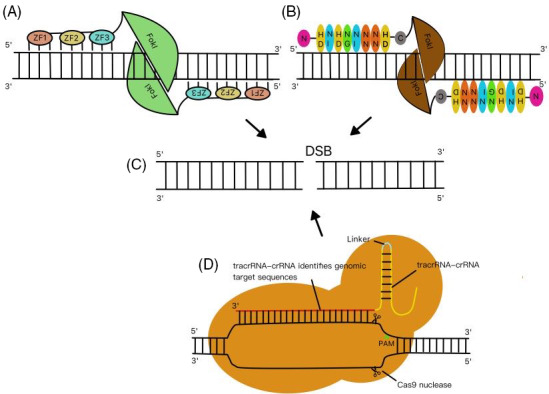
Principles of gene knockout by zinc finger nuclease (ZFN), transcription activator‐like effector nuclease (TALEN), and CRISPR/Cas9 technologies. (A) Zinc finger (ZF) structures are assembled as modular assemblies. When ZF DNA‐binding domain can be perfectly paired with the target DNA sequence and FokI is dimerized, targeted site‐specific DNA is cut off. (B) The most common repeat‐variable di‐residues (RVDs), which can recognize four types of bases such as A, G, C, and T, respectively, are NI, NN, HD, and NG. The DNA specific recognition domain on TALE combines with the DNA sequence on the target site, and then FokI cutting domain is dimerized to cut off the specific sites of the target DNA sequence. (C) DNA double‐strand break (DSB) is implemented by ZFN, TALEN, and CRISPR/Cas9. (D) Cas9, crRNA, and tracrRNA jointly establish the Cas9 protein‐RNA mechanism. Cas9 targets virus or plasmid DNA by recognizing protospacer adjacent motif (PAM), and then cuts the DNA at the target site.
2.1.3. Transcription activator‐like effector nuclease
Transcription activator‐like effector nuclease technology is characterized by very high specificity. 23 , 24 A typical TALEN unit consists of three main parts: N‐terminal domains, central domains containing typical tandem transcription activator‐like effector (TALE) repeats, and C‐terminal domains with FokI endonuclease function. The DNA specific recognition unit of TALE is 12 ~ 13 repeat‐variable di‐residues (RVDs), such as NI, NG, NN, and HD. 25 These RVDs recognize four types of bases: A and NI, T and NG, G and NN, C and HD. Their unique affinity makes the target location very specific. 26 , 27 Its principle is to combine the DNA specific recognition domain on TALE with the target DNA sequence, and then dimerize the FokI cutting domain. Under the action of FokI nuclease, target DNA sequence are cut off to achieve gene knockout 28 , 29 (Figure 2B,C). TALEN is simpler to design and more specific than ZFN technology, but it is not good enough to replace traditional technology at present due to its high cytotoxicity and high cost 28 , 30 (Table 1).
2.1.4. CRISPR/Cas9
Clustered regularly interspaced short palindromic repeats (CRISPR) was observed in Escherichia coli in 1987 by Yoshizumi Ishino and his colleagues. 31 Later, CRISPR/Cas9 technology developed from it has gradually been used in the field of genetic engineering research. 32 The principles of type II CRISPR/Cas9 technology are comprehensible: Mature CRISPR RNA (crRNA) and trans‐activating CRISPR RNA (tracrRNA) are two key RNA fragments in CRISPR/Cas9 system. There are partial complementary sequences in the two RNA fragments, which together constitute the double stranded RNA (dsRNA) structure. 33 crRNA, tracrRNA and Cas9 jointly establish the Cas9 protein‐RNA mechanism. Cas9 targets virus or plasmid DNA by recognizing protospacer adjacent motif (PAM) which is made up of NGG bases (N is any base), and then cuts the DNA at the target site to make DSB, thus causing the loss of gene fragments and realizing gene knockout 34 , 35 (Figure 2C,D). Organ cultures made by the CRISPR/Cas9 system in vitro have been proven to be suitable for studying the biological characteristics of IVD and the system is the main method for generating the disease models in vivo. 36 In a mouse model of IVD injury, injection of adenovirus expressing CRISPR/Cas9 lost β‐catenin function and alleviated IVDD. 37 In recent years, CRISPR/Cas9 has gradually become the mainstream of knockout technology due to its advantages of precise targeting, high efficiency, low cytotoxicity, low cost and simplicity, but its specificity is not high and its safety is controversial 32 , 34 , 38 (Table 1).
2.2. Current situation of applications
The target gene in all tissues or cells of KO mice was knocked out, and specific organ or site was studied according to the experimental protocol.
2.2.1. Genotype of the KO model
KO mice can be homozygous, or heterozygous. Sometimes double knockout mice can be constructed by knocking out two target genes simultaneously.
A homozygote is an individual with two identical alleles on a homologous chromosome. When researchers use certain knockout techniques to knock out two identical alleles on homologous chromosome of the mice, they can generate homozygous KO mice. Sometimes, in order to avoid other adverse factors, such as early death of mice, researchers use gene edition technology to knock out one of the two alleles on a homologous chromosome, creating heterozygous KO mice. Heterozygous KO mice with nonlethal genes can be a useful tool for the generation of same‐week‐old homozygous KO mice as well as wild‐type mice because of trait segregation in offspring produced by mating between heterozygous mice. In addition, double gene knockout (dKO) mice are increasingly used in medical research, and IVD study is no exception. By using the same gene knockout technology or the combination of different gene knockout technologies, the alleles of the two target genes were knocked out respectively according to the experimental requirements, so as to construct the dKO mice required by the researchers. According to this method, researchers can even knock out three or more target genes to study disc degeneration.
2.2.2. Scientific rationale and reasoning for choosing this model
Homozygous KO mice are used in most of the studies because the target gene in these mice is completely knocked out and the disc phenotype changes most dramatically. When the target gene is completely knocked out in mice without serious consequences, using homozygous KO mice is an excellent choice for animal models. However, when the target gene is knocked out and the mice have serious defects or even embryo death, which may interfere with the experimental results or affect the experimental process., heterozygous KO mice might be considered. Although using heterozygous mouse model may have some disadvantages such as insignificant disease phenotype, which makes it difficult to fully explore the relationship between target genes and diseases, the presence of heterozygous KO mice also provides a glimmer of hope for the research that has failed to advance with homozygous KO mice as animal models. For the dKO mice, it is more and more commonly used in the study of IVDD. When we want to investigate the relationship of two different genes in the progression of disc degeneration, dKO mice may be a good choice. However, experiments using dKO mice are extremely demanding for the screening of the target genes, because the expression of the two genes in the IVD and their possible association must be fully understood before the experiment. At the same time, dKO mice also have some disadvantages, such as a long production cycle and more consumption of manpower and capital that need to be considered in the process of animal experiments.
2.2.3. Examples in the studies of the IVD
Knocking out the genes with different functions results in various characteristic effects.
Some genes have protective effect for the IVD, and when these genes are knocked out, disc degeneration will develop or become aggravated. For example, in lumbar spine specimens from apolipoprotein E (Apoe) KO mice, Zhang et al. 39 found collagen levels (collagen I, II) increased in AF cells but decreased in NP cells. Increased expression of matrix metalloproteinase‐3 (Mmp3), 9 and 13 in disc cells and increased apoptosis of NP cells were also observed. Analogously, our previous study used insulin‐like growth factor 1 receptor +/− (Igf1r+/−) mice to observe disc degeneration. Type II collagen and proteoglycan significantly reduced and Mmp13 significantly increased in the lumbar discs of Igf1r+/− mice. 40 Thus, reduced Igf1r expression accelerates IVDD. As another example, Lin et al. 41 found that age‐related IVD degeneration (ARD) was accelerated in lumbar IVDs of Tenomodulin (Tnmd) KO mice, mainly manifested as macrophage infiltration in outer AF (OAF) and collagen fibril diameter reduction. However, when Tnmd and Chondromodulin I (Chm1) were knocked out at the same time, the progression of ARD was faster than that of Tnmd KO mice. When Tnmd gene is mutated in IVDs, Chm1 may play a compensatory role. 42 Many other genes have similar functions 43 , 44 , 45 , 46 , 47 , 48 , 49 , 50 , 51 , 52 , 53 , 54 , 55 , 56 , 57 , 58 , 59 , 60 , 61 , 62 , 63 , 64 , 65 (Table 2).
TABLE 2.
IVD phenotypes and function of genes that have been knocked out in KO mice.
| Model | Jax mice ID | IVD phenotypes or key information | Target gene | NCBI gene ID | Effect of each model on IVDD | References |
|---|---|---|---|---|---|---|
| Adamts5 KO mice | / | Loss of GAG content in IVDs reduced, proteolysis of aggrecan in IGD decreased | A disintegrin‐like and metallopeptidase (reprolysin type) with thrombospondin type 1 motif, 5 (aggrecanase‐2) | 23794 | Decrease | 68 |
| Ager KO mice | / | Accumulation of AGE, destruction of RAGE‐dependent collagen in AF | Advanced glycosylation end product‐specific receptor | 11596 | Decrease | 67 |
| Apoe KO mice | 002052 | Collagen increased in AF but decreased in NP cells. Expression of Mmp3, 9, and 13 in ECM and apoptosis of NP cells increased. | Apolipoprotein E | 11 816 | Increase | 39 |
| Bach1 KO mice | / |
The expression of HO‐1 increased in NP and the proportion of apoptosis decreased after the needle puncture. There was no significant difference between Bach KO mice and WT mice after a period of time. No significant effect on ARD |
BTB and CNC homology 1, basic leucine zipper transcription factor 1 | 12013 | Decrease/none | 76 |
| Bgn KO mice | / | IVDD occurred as the mice grew older. | Biglycan | 12111 | Increase | 43 |
| Bmi1 KO mice | 024584 | Fewer Col2a1 positive regions and more Col10 positive regions were observed. | Bmi1 polycomb ring finger oncogene | 12151 | Increase | 44 |
| Casp3 KO mice | / | Delay IRD, but accelerate ARD. | Caspase 3 | 12367 | Increase/decrease | 73 |
| Caveolin KO mice | 004585 | Cells are decreased in NP but collagen is increased in the ECM. | Caveolin 1, caveolae protein | 12389 | Increase | 45 |
| Cdkn1b KO mice | 003122 | NP becomes irregularly smaller and has an ambiguous boundary with AF. | cyclin‐Dependent kinase inhibitor 1B | 12576 | Increase | 46 |
| Cdkn2a KO mice | 029676 | IVD height, GAG and water content almost maintained in IVDs, Il1b, Il6, Tnfa, Col1a1 and Col10 expression decreased | cyclin dependent kinase inhibitor 2A | 12578 | Decrease | 69 |
| Chsy3 KO mice | / | The anabolism of NP cells decreased and catabolism increased. | Chondroitin sulfate synthase 3 | 78923 | Increase | 47 |
|
Col2a1 KO mice (heterozygous) |
/ | GAG decreased in AF and EP, EP thickened and ossified prematurely | Collagen, type II, alpha 1 | 12824 | Increase | 48 |
| Col9a1 KO mice | / | EP degeneration, new bone formation and cartilage disorganization | Collagen, type IX, alpha 1 | 12839 | Increase | 49 |
| Dmd/Utrn dKO mice | / | In IVDs, especially in the NP, PGs are disorganized or even lost. | Dystrophin, muscular dystrophy utrophin |
13405 22288 |
Increase | 50 |
| ERa KO mice | 004744 | The damage to the CEP is aggravated. | Estrogen receptor 1 (alpha) | 13982 | Increase | 51 |
| Flnb KO mice | 027233 | AF cells lost typical characteristics, Col10 expression, IVD tissue ossification, increased apoptotic cells | Filamin, beta | 286940 | Increase | 52 |
| Gdf5 KO mice | / | Abnormal proteoglycans and collagen | Growth differentiation factor 5 | 14563 | Increase | 53 |
| Gpr35 KO mice | / | The mice retained more proteoglycans and collagen, as well as alleviated IVDD. | G protein‐coupled receptor 35 | 64095 | Decrease | 70 |
| Igf1r KO mice (heterozygous) | / | Type II collagen and proteoglycan decreased and Mmp13 significantly increased | Insulin‐like growth factor I receptor | 16001 | Increase | 40 |
| Il1a/b dKO mice | / | The proportion of NP cells decreased, and AF showed obvious degeneration. |
Interleukin 1 alpha interleukin 1 beta |
16175 16176 |
Increase | 54 |
| Il1rn KO mice | / | Expression of Adamts4, Mmp3 and Mmp7 increased, proteoglycan decreased, and collagen structure lost | Interleukin 1 receptor antagonist | 16181 | Increase | 55 |
| Lepr KO (db/db) mice | 000697 | Vertebrae bone mass decreased, Mmp3 expression and apoptotic cells increased in T2DM mice | Leptin receptor | 16847 | Increase | 56 |
| Mir141 KO mice | / | Reducing the apoptosis of NP cells through Sirt1/NF‐κB signaling pathway | microRNA 141 | 387159 | Decrease | 71 |
| Mir146 KO mice | 034342 | Low expression of GAG, high expression of Mmp13 and Adamts5 | microRNA 146 | 387164 | Increase | 57 |
| Mstn(Gdf8) KO mice | / | Muscle mass and bone mass increased in the mice, while the discs degenerated. | Myostatin | 17700 | Increase | 58 |
| Nrf2 KO mice | 017009 | Decreased the expression of autophagy gene and inhibited the Keap1/Nrf2/p62 feedback loop | Nuclear factor, erythroid derived 2, like 2 | 18024 | Increase | 59 |
| Opg KO mice | 010672 | CEP calcification, IVD blood vessels formation, high expression of cytokines in GP and EP | Osteoprotegerin | 18383 | Increase | 60 |
| Panx3 KO mice | / |
AF injured by needle puncture retained the structure, while NP without injury showed degeneration. No significant effect on ARD |
Pannexin 3 | 208098 | Increase/none | 77 |
| Pgrn KO mice | 013175 | Activation of β‐catenin signaling and NF‐κB signaling, abnormal bone formation, and IVD matrix degeneration | Granulin | 14824 | Increase | 61 |
| Phlpp1 KO mice | / | Akt phosphorylation, cell proliferation, wound healing after the needle puncture | PH domain and leucine rich repeat protein phosphatase 1 | 98432 | Decrease | 66 |
| Smad3 KO mice | 003451 | CEP height declined, Tgfb, aggrecan and Col2a1 expression decreased, Col10α1 expression increased | SMAD family member 3 | 17127 | Increase | 62 |
| Sost KO mice | / | Inhibition of sclerostin enhanced IVD structure in mice. | Sclerostin | 74499 | Decrease | 72 |
| Sparc KO mice | 003728 | Accelerated IVDD and increased IVD stiffness | Secreted acidic cysteine rich glycoprotein | 20692 | Increase | 63 |
| Sulf1 KO mice | / | Type II collagen is reduced in the NP and type I collagen is denatured in the AF. | Sulfatase 1 | 240725 | Increase | 64 |
| Tnfa KO mice | 003008 | The expression of Ngf in injured discs was not significantly different from that in WT. | Tumor necrosis factor | 21926 | None | 74 |
| Tnmd KO mice | / | Accelerate ARD, Chondrocytes thickened in NP, macrophages infiltrated in OAF | Tenomodulin | 64103 | Increase | 41, 42 |
| Tnmd/Chm1 dKO mice | / | Chondromodulin | 16840 | |||
| TonEBP KO mice | / | EP and annular herniations, cytoskeletal rearrangements, changes in proinflammatory gene expression and matrix remodeling | Nuclear factor of activated T cells 5 | 54446 | Increase | 65 |
| Trpa1 KO mice | 006401 | GAG decreased in AF and GP. | Transient receptor potential cation channel, subfamily A, member 1 | 277328 | Increase | 75 |
| Trpv1 KO mice | 003770 | No significant change was observed in Trpv1 KO mice. | Transient receptor potential cation channel, subfamily V, member 1 | 193034 | None |
Abbreviations: Adamts, a disintegrin and metalloproteinase with thrombospondin motifs; AGE, advanced glycation end‐product; Akt, protein kinase B; ARD, age‐related IVD degeneration; CEP, cartilaginous endplates; Col1, type I collagen; Col10, type X collagen; Col2a1, collagen type II alpha 1 chain; ECM, extracellular matrix; EP, endplates; GAG, glycosaminoglycan; GP, growth plate; HO‐1, heme oxygenase 1; IGD, interglobular domain; IRD, injury‐related IVD degeneration; IVD, intervertebral disc; IVDD, intervertebral disc degeneration; Mmp, matrix metalloproteinase; Ngf, nerve growth factor; NP, nucleus pulposus; OAF, outer annulus fibrosus; PGs, proteoglycans; RAGE, the receptor for AGE; WT, wide type.
In contrast, some genes have deleterious effect for the IVD, and when these genes are knocked out, disc degeneration will be prevented or alleviated. For instance, Zhang et al. 66 found protein kinase B (Akt) phosphorylation and cell proliferation increased but apoptosis decreased in the caudal IVDs of PH domain leucine‐rich repeat protein phosphatase (Phlpp1) KO mice. After the IVD was injured by needle puncture, disc degeneration was less severe in the Phlpp1 KO mice than in the WT mice and knocking out Phlpp1 had a protective effect on the IVDs of the mice. Hoy et al. 67 used the receptor for AGEs (Ager) KO mice to study the effects of advanced glycation end products (AGEs) and RAGE on the quantity and quality of collagen in the AF of the caudal IVDs. They found that accumulation of AGE would lead to the disruption of RAGE‐dependent AF collagen. Therefore, Ager KO may have protective effect on collagen in AF. In the study of disc degeneration, a disintegrin‐like and metallopeptidase with thrombospondin type 1 motif, 5 (Adamts5) KO mice used by Ngo et al., 68 cyclin dependent kinase inhibitor 2A (Cdkn2a) KO mice used by Che et al., 69 G protein‐coupled receptor 35 (Gpr35) KO mice used by Chen et al., 70 microRNA 141 (Mir141) KO mice used by Ji et al. 71 and sclerostin (Sost) KO mice used by Kroon et al. 72 were also found that the target gene they studied could exacerbate disc degeneration (Table 2).
Some genes may have opposite effects on different types of disc degeneration (Table 2). For example, Ohnishi et al. 73 used Caspase‐3 (Casp3) KO C57BL/6 mice to analyze the different roles of Caspase‐3 in injury‐related IVD degeneration (IRD) and ARD. The critical factor of IRD is cell apoptosis, while the main cause of ARD is cell senescence, so their pathological mechanisms are different. In this study, researchers found that the histological degenerative score of the lumbar IVDs in Casp3 KO mice was lower than that in WT mice, and the disc degeneration in Casp3 KO mice was delayed as compared with WT mice by acupuncturing the IVD of both types of mice. Later, by comparing young and old Casp3 KO mice and WT mice, they found that the expression of p16INK4A (a marker of cellular senescence) and the ECM degradation enzyme was upregulated in the lumbar IVDs of the old Casp3 KO mice, but the number of apoptotic cells was significantly reduced. Therefore, Casp3 knockout could delay IRD, but accelerate ARD.
Some other genes have been studied in disc degeneration (Table 2). For instance, Yokozeki et al. 74 studied whether tumor necrosis factor‐α (Tnfa) could affect the expression of nerve growth factor (Ngf) in the mouse coccygeal discs injury model. They compared the Tnfa KO (C57B/6J background) mice with WT mice, and found that Ngf expression was similar without significant changes. Kameda et al. 75 used Transient receptor potential ankyrin 1 (Trpa1) KO C57BL/6 mice and Transient receptor potential vanilloid 1 (Trpv1) KO mice to study the caudal motor segments of Trpa1 and Trpv1 deficient mice. They found that glycosaminoglycan (GAG) decreased in AF and growth plate (GP) in mature Trpa1 KO mice, while no significant change in Trpv1 KO mice. Thus, it was concluded that Trpa1 could promote GAG production in mature IVDs. In addition, BTB and CNC homology 1, basic leucine zipper transcription factor 1 (Bach1) KO mice and pannexin 3 (Panx3) KO mice were used for ARD study, but the difference between the knockout group and the control group was not significant. 76 , 77
2.2.4. Authors' scientific perspectives on the strengths and limitations of the mouse model
With the development of genome sequencing technology, 99% of mouse genes have been found to have human homologue. 78 Therefore, it has become a very important method to study human diseases by KO mice, and models made by this technology have been widely used in the study of IVDD.
Constitutive knockout technology is mostly carried out in the embryonic stage of mice and it can completely knock out the target gene, which makes the construction of disc degeneration model more accurate and brings great convenience to the follow‐up research. 27 However, this technology also has some disadvantages. First, this technology is a poor imitation of human normal gene variation, which will lead to the complete loss of gene function. Second, there may be many members in the gene family as the target gene with the same function. When the target gene is knocked out, the phenotype of disc degeneration is not easily identified, possibly due to compensation by other genes which have the same function. Finally, when certain essential genes are knocked out, they can be fatal to the embryo, making them impossible to study 78 (Table 1).
3. CONDITIONAL KO MICE
cKO mice is a mouse model in which the target gene is deleted in a specific stage of the mouse life cycle or in a specific tissue, but is normally expressed in other tissues or cells, thus making the scope and time of the modification of the mouse genome in a controllable state.
3.1. Name of the Cre model
Each part of the IVD has different composition. For example, NP cells can synthesize ECM, which is rich in type II collagen, proteoglycan, and elastin; AF is composed of alternating type I collagen fibers; And the ECM in the CEP is rich in proteoglycan and collagen fibers. 7 When Cre is expressed in specific cells or tissue, the target gene at that site is knocked out. Therefore, the location of Cre recombinase activation in the IVD is critical. 79 According to the different activated sites of Cre recombinase, researchers have constructed a variety of Cre mouse models for IVD research, which are mainly divided into the following categories.
Col2 Cre mice, Col1 Cre mice, Shh Cre mice, Shh CreER mice, Acan CreER mice, Krt19 CreER mice, Noto Cre mice, Ella Cre mice, and Lepr Cre mice are used to study NP. Col2 CreER mice and Scx Cre mice are used to study inner AF (IAF). Col1 CreER mice, Lepr Cre mice, and Scx Cre mice are used to study OAF. Col2 Cre mice, Col2 CreER mice, Col1 Cre mice, Shh Cre mice, Ella Cre mice, Acan CreER mice, and Osx Cre mice are used to study CEP. Col2 Cre mice, Col2 CreER mice, Shh Cre mice, Osx Cre mice, and Acan CreER are used to study GP. At present, the expression sites of Col1 Cre and Ella Cre have not been confirmed, but there have been phenotypic changes in IVDs in related animal experiments (Figure 3).
FIGURE 3.
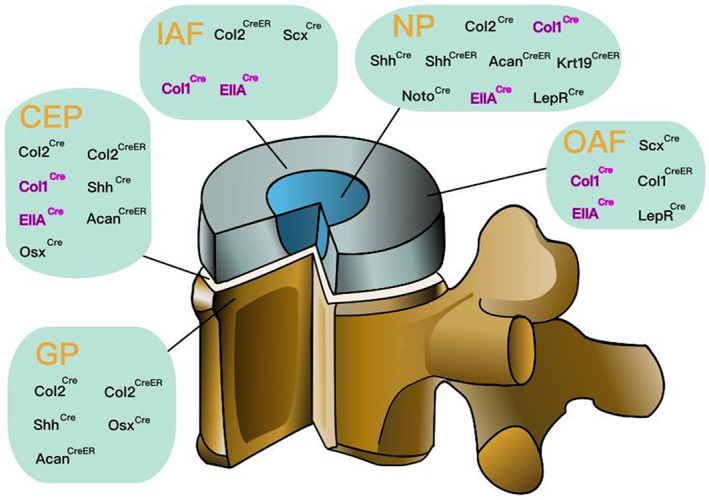
The location of Cre recombinase activation in the intervertebral disc (IVD). Some Cre mice target specific sites of the IVDs. Cre can be expressed in specific cells or tissue and the target gene at that site is knocked out. Therefore, the location of Cre recombinase activation in the IVD is critical. Cre recombinase activated in nucleus pulposus (NP): Col2 Cre , Col1 Cre , Shh Cre , Shh CreER , Acan CreER , Krt19 CreER , Noto Cre , Ella Cre , and Lepr Cre . Cre recombinase activated in inner annulus fibrosus (IAF): Col2 CreER and Scx Cre . Cre recombinase activated in outer annulus fibrosus (OAF): Col1 CreER , Lepr Cre , and Scx Cre . Cre recombinase activated in cartilaginous endplate (CEP): Col2 Cre , Col2 CreER , Col1 Cre , Shh Cre , Ella Cre , Acan CreER , and Osx Cre . Cre recombinase activated in growth plate (GP): Col2 Cre , Col2 CreER , Shh Cre , Osx Cre , and Acan CreER . Among them, the expression sites of Col1 Cre and Ella Cre , marked in rose red, have not been confirmed, but there have been phenotypic changes in IVDs in related experiments. Acan, aggrecan; Col1, type I collagen; Col2, type Il collagen; ER, estrogen receptor; Krt19, cytokeratin 19; Lepr, leptin receptor; Noto, notochord; Osx, osterix; Scx, Scleraxis; Shh, sonic hedgehog
3.2. Description of the gene construct
Conditional gene knockout is a special gene knockout method, which is mainly realized by chromosomal site‐specific recombinase system, such as Cre‐LoxP, FLP‐FRT and Dre‐Rox. 80 Cre‐LoxP and FLP‐FRT have the same basic principle. According to the literature, FLP‐FRT and Dre‐Rox methods have hardly been applied to the study of IVDD, and Cre/LoxP recombination system is most commonly used.
Since 1994, Gu et al. 81 have successfully developed tissue‐specific gene knockout mice using Cre/LoxP recombination system. Subsequently, the technology has been widely used. Cre/LoxP recombination system is composed of Cre recombinase and LoxP site. Cre recombinase cleaves DNA fragments between two LoxP sites by recognizing two LoxP sites in the same direction. In general, Flox mice were produced by inserting two LoxP in the same direction at both ends of one or more important exons of the target gene which was selected and has significant difference between healthy and degenerated disc through gene sequencing or other biotechnology. These Flox mice have no other changes in the target gene, and their phenotype remains the same as that of wild‐type mice. When Flox mice were crossed with mice that expressed a tissue‐specific Cre recombinase, the progeny contained both the Cre gene and the LoxP gene. 82 Therefore, the progeny can knock out the target gene in a specific tissue or cell, while the target gene is normally expressed in other tissues or cells, realizing the specificity of the knockout tissue 83 , 84 (Figure 4A).
FIGURE 4.
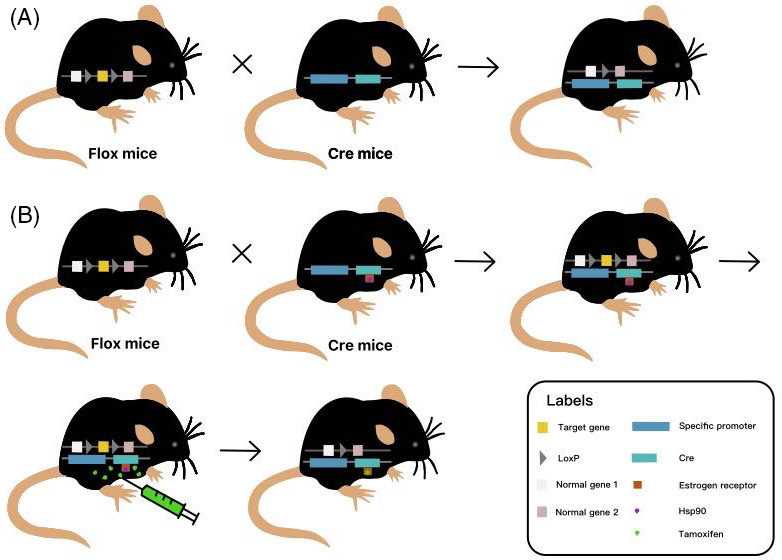
Basic principle of conditional gene knockout using Cre/LoxP system. (A) When Flox mice inserted by two LoxP loci in the same direction at both ends of one or more important exons of the target gene are crossed with mice that express a tissue‐specific Cre recombinase, the progeny which contains both the Cre gene and the LoxP gene can knock out the target gene in a specific tissue or cells. (B) tamoxifen (TAM) is most commonly used as an inducer that binds to the fused mutant estrogen receptor (ER) on the Cre recombinase, replacing heat shock protein 90 (Hsp90), and induces the activation of the Cre/(ER)T system to knockout target gene. Appropriate timing of tamoxifen injection is selected according to the conditions of mice to avoid embryo or premature death.
Another important application of the Cre/LoxP system is the ability to achieve gene knockout through induction of hormones, tetracycline, adenovirus, and interferon. 85 In this way, the target gene between LoxP can be knocked out in a certain period of time and in certain tissues by controlling Cre expression promoter activity or the inducibility of Cre recombinase expressed, so as to control the time of inducer administration or the time of Cre gene localization and expression system transfer to animals. Thus, the time of gene knockout can be controlled. 86 For example, in studies of disc degeneration, tamoxifen (TAM) is most commonly used as an inducer that binds to the fused mutant estrogen receptor (ER) on the Cre recombinase, replacing Hsp90, and induces the activation of the Cre/(ER)T system 87 (Figure 4B). In addition, tetO‐Cre recombinase system induced by tetracycline or doxycycline (Dox) has also been applied in the studies of disc degeneration, but this system is rarely used due to its complexity and the large workload of breeding and genotype identification.
It can be seen that target gene is knocked out at specific tissues or times according to the tissues or times in which Cre is expressed. Therefore, Cre recombinase plays an important role in maintaining tissue specificity and controllability of the time when the gene is knocked out. Realizing the controllability of the time is a great leap forward in the development of conditional knockout technology which can avoid premature death in mice to a large extent. Meanwhile, this feature has a huge role in simulating the specific onset period. For example, juvenile (JIS) or adolescent idiopathic scoliosis (AIS) tends to occur in young people between the ages of 10 and 16. It would not be appropriate to study this disease by knocking out the related gene from the embryonic stage of mice.
3.3. Expression profile of the gene
The site and time of Cre recombinase activation determine the specificity of conditional gene knockout. It is essential to fully understand the components of each tissue and the expression of related genes in the IVD to establish a Cre model.
3.3.1. Type lI collagen
Type lI collagen (Col2) is an important cartilage marker, widely distributed in IVD, joints, rib tips, tendon, ligament, nose, larynx, the rings of the trachea, jaw and other tissues. It is mainly expressed in hyaline matrix‐rich chondrocytes and cell‐rich fibrous chondrocytes. 88 , 89 Therefore, Col2 plays a very important role in the body. Typically, Col2 Cre consists of a 3 kb Col2a1 promoter, a 3.02 kb Col2a1 intron 1, and a Col2a1 exon 1. Col2 Cre mice were hybridized with Rosa26 (R26)‐mTmG mice, then obtaining IVDs of offspring Col2 Cre;R26‐mTmG mice at 2 weeks and 2 months of age. Green fluorescent protein (GFP) expression was detected in NP, AF, CEP, and GP of lumbar spine. 79 , 90 , 91 And the researchers found similar GFP expression levels in the discs of 2‐week‐old and 2‐month‐old mice. Thus, Col2 Cre recombinase is widely activated in IVD (Table 3). It may be due to the presence of ischemia and hypoxia in the IVD or the different distribution of ER in different parts of the IVD, the Col2 CreER recombinase activated sites induced by tamoxifen are different from the Col2 Cre recombinase activated sites. Col2 CreER consists of a 1 kb Col2a1 promoter, a 650 bp Col2a1 intron 1 and an ER. GFP positive cells were detected in the inner AF (IAF) along with EPs and GPs of the lumbar spine of Col2 Cre(ER)T2;R26‐mTmG mice when tamoxifen was injected at 2 weeks and 2 months of age, but not in the NP (Table 3). The researchers also found that the GFP positive rate in IAF and GP of 2‐week‐old mice was significantly higher than that of 2‐month‐old mice, and EP contained only a few GFP‐positive cells at all stages. 79 Wei et al. 92 injected tamoxifen intraperitoneally into Col2 CreER/tdTomato mice at P7, and punctured the coccygeal (Co) discs of offspring mice to induce IVDD. It was found that the IAF labeled with red fluorescent protein (RFP) gradually infringed into the NP.
TABLE 3.
Specific Cre mice and their use in IVD studies
| Cre recombinase | Jax mice ID | Activated sites in the disc | Research methods and findings | References | Activated in other sites | References |
|---|---|---|---|---|---|---|
| Acan CreER |
019148 |
NP, GP, Part of the CEP and AF | Agc1 Cre(ER)T2;R26‐mTmG mice were injected with tamoxifen (once a day for 5 days) at 2 weeks and 2 months of age, respectively, to observe the expression of GFP in IVD. All chondrocytes in NP and GP of Agc1 Cre(ER)T2;R26‐mTmG mice were GFP positive. The number of GFP positive cells in CEP of 2‐month‐old mice was more than that of 2‐week‐old mice, while the number of GFP positive cells in AF was opposite. |
Cartilage, aorta, heart, and so forth |
||
| Col1 Cre |
/ |
May be in AF, NP and EP |
— |
Bone, liver (hepatic stellate cell), and so forth. |
||
| Col1 CreER |
/ |
OAF | Col1a2 Cre(ER)T;R26mTmG mice were injected with tamoxifen at the age of 3 weeks, and the expression of tdTomato and GFP in IVD was observed 100 days later. |
Bone |
||
| Col2 Cre |
003554 |
NP, AF, CEP, GP |
The expression of GFP in the IVD of Col2 Cre;R26‐mTmG mice was observed at 2 weeks and 2 months of age, respectively. The researchers found similar levels of GFP expression in the IVDs of mice at 2 weeks and 2 months of age. |
Cartilage, meniscus, endosteum, ligament, bone, synovium, kidney (embryonic renal epithelium), and so forth. |
||
| Col2 CreER |
/ |
IAF along with EPs and GPs | Col2 Cre(ER)T2;R26‐mTmG mice were injected with tamoxifen (once a day for 5 days) at 2 weeks and 2 months of age, respectively, to observe the expression of GFP in IVD. GFP positive rate in IAF and GP of 2‐week‐old mice was significantly higher than that of 2‐month‐old mice, and EP contained only a few GFP‐positive cells at all stages. |
Cartilage, epiphyseal bone, and so forth. |
||
| Ella Cre | 003724 | May be in AF, NP and EP |
Cre recombinase widely activated in early embryos of the mice. |
A wide range of tissues |
||
| Krt19 CreER |
026925 |
Most NP cells |
Different doses of tamoxifen were injected into Krt19 Cre(ER)T; R26mTmG mice of different ages to observe the expression of GFP in IVD. The GFP positive expression rate increased gradually in the NP of neonatal mice, even reaching 92% at P8. As the mice grew older, the positive rate of GFP in the NP cells gradually decreased to 72.64%. |
Epithelial and peridermal cells of kidney, gallbladder, liver, and so forth. |
||
| Lepr Cre |
008320 |
OAF, Part of the NP |
Expression of tdTomato was observed in the IVD of 1 month old Lepr Cre;R26R‐tdTomato mice. The number of NP cells of the lumbar IVDs specifically targeted by Lepr Cre nearly 75% gradually decreased from the P0, and no longer decreased after reaching the plateau at 1 month. OAF can be labeled by Lepr Cre from the P14 to adulthood. |
Skeletal cell, neurons, and so forth. |
||
| Noto Cre |
/ |
Notochord (NP) |
Mating Noto Cre mice with R26‐LacZ to observe the expression of β ‐galactosidase in subsets of notochordal cells and mature NP cells. β‐galactosidase was expressed at E8.0, and solid rod‐like notochord appeared at E10.5. NP was segmented at E15.5 and β‐galactosidase was limited to NP at P100. |
/ |
||
| Osx Cre |
/ |
Part of the CEP and GP |
The expression of GFP in the IVD of Osx Cre;R26‐mTmG mice was observed at 2 weeks and 2 months of age, respectively. GFP expression levels were similar in IVDs of 2‐week‐old and 2‐month‐old mice. |
Bone, cartilage, bone marrow, glomerular cell, gastric, intestinal epithelium, and so forth. |
||
| Scx Cre |
/ |
AF, May also in other tissues of IVD |
Scx Cre‐H mice crossed with R26R mice to generate offspring with the genotype Scx Cre‐H;R26R , and high Cre expression activity was detected in the annulus fibrosus of the Scx Cre‐H;R26R mice. |
Ligaments and tendons, joints, and cartilage around the chondro‐tendinous/ligamentous junction, and so forth. |
||
| Shh Cre |
005622 |
NP, Part of the CEP and GP |
The expression of GFP in the IVD of Shh Cre;R26‐mTmG mice was observed at 2 weeks and 2 months of age, respectively. NP cells from 2‐week and 2‐month‐old mice were labeled with 100% GFP, whereas only some GFP‐positive cells were found in CEP and GP. |
Bone, dental epithelium, ventral midbrain progenitors, pulmonary vasculature, cartilage, neural stem cell, genital tubercle, and so forth. |
||
| Shh CreER |
005623 |
Few NP cells |
Shh Cre(ER)T2;R26‐mTmG mice were injected with tamoxifen (once a day for 5 days) at 2 weeks and 2 months of age, respectively, to observe the expression of GFP in IVD. Shh Cre(ER)T2;R26‐mTmG mice had 1.6% GFP‐positive cells at 2 weeks of age and only 0.5% at 2 months of age. |
Forebrain, taste bud progenitors, and so forth. |
Abbreviations: Acan, aggrecan; CEP, cartilaginous endplates; Col1, type I collagen; Col2, type Il collagen; EP, endplates; ER, estrogen receptor; GP, growth plate; IAF, inner annulus fibrosus; IVD, intervertebral disc; IVDD, intervertebral disc degeneration; Krt19, cytokeratin 19; Lepr, leptin receptor; Noto, notochord; NP, nucleus pulposus; OAF, outer annulus fibrosus; Osx, osterix; Scx, Scleraxis; Shh, sonic hedgehog.
3.3.2. Type I collagen
Type I collagen (Col1), mostly expressed in fibroblasts and cell‐rich fibrous chondrocytes, accounts for 90% of human body and is the most common protein type. 88 , 93 It is mainly distributed in skeleton, tendons and skin, and considered to be an important fibrous collagen. 94 , 95 , 96 , 97 Meanwhile, type I collagen is the main component of AF, so it is also of great value in the study of IVDs. However, this may also be a disadvantage of type I collagen, because it is widely distributed and does not have high specificity, so Col1 Cre mice is rarely used in the study of IVDD. At the same time, tamoxifen can induce activation of Col2 CreER recombinase in IAF, but not in OAF. Most studies on the composition and function of the AF have treated it as a whole, ignoring the differences between its interior and exterior parts. Bedore et al. 98 injected 3‐week‐old Col1a2 Cre(ER)T;R26mTmG mice with tamoxifen. The cells expressing Cre were labeled with GFP, and the other cells were labeled with tdTomato. The expression of Cre was found in the OAF of the lumbar spine. However, NP and IAF are not expressed. Therefore, crossing Col1 CreER mice with Flox mice to study IVDD can knock out the target gene located on the OAF (Table 3).
3.3.3. Sonic hedgehog
The notochord secretes a kind of protein called Sonic hedgehog (Shh), which induces the differentiation of motor neurons, ventral interneurons and floor plate cells. 99 , 100 , 101 , 102 , 103 , 104 , 105 , 106 , 107 In IVDs, the Shh promoter driven Cre recombinase was only expressed in NP cells. 108 , 109 , 110 Thus, the Shh Cre mice has high specificity and becomes an available tool for studying IVDs. Peck et al. 111 isolated notochord‐derived cells (NDCs) from mouse embryos using Shh Cre ;ROSA:YFP mice and analyzed them to study the embryogenesis of NP. Choi et al. 112 found that it specifically targeted NP cells in elderly mice and mouse embryos by fate‐mapping studies. Subsequently, Zheng et al. 79 produced Shh Cre;R26‐mTmG mice through hybridization. The NP cells were completely labeled with GFP, while some positive cells were also found in CEP and GP. Therefore, Cre recombinase can be activated in NP and part of the CEP and GP in IVD (Table 3). The Shh CreER mice, which requires tamoxifen induction, has been shown to specifically target NP cells in early embryo by Choi et al 113 Soon afterwards Zheng et al. 79 injected tamoxifen into Shh Cre(ER)T2; R26‐mTmG mice at the age of 2 weeks or 2 months, and found that only a few NP cells were GFP positive. As a consequence, Cre recombinase can be activated in a few NP cells (Table 3).
3.3.4. Notochord
Notochord is an important rod‐like structure during embryogenesis of the chordate and has been considered as a precursor of the spine for a long time. 114 , 115 It degenerates with age, but the remains of notochord can be observed in normal adult NP tissue. 116 Currently, notochord is thought to have two main functions: (1) It provides support for embryonic structures prior to the formation of the chordate NP; (2) Secrete signal molecules such as Shh to promote body development. 117 , 118 At the same time, it is thought to be the sole source of NP cells in mice. 112 , 119 , 120 Therefore, the notochord (Noto) gene plays a crucial role in the proper development of vertebrates. McCann et al. 120 mated Noto Cre mice with R26‐LacZ, and found that β‐galactosidase was expressed in subsets of notochordal cells at E8.0, and solid rod‐like notochord appeared at E10.5. Subsequently, there was an obviously segmented NP which contains residual notochord at E15.5. It was observed that β‐galactosidase was expressed throughout the thoracic spine to caudal tip of the tail at postnatal time points and was limited to NP in section at P100, whereas it was not present in other IVD tissues. Obviously, the activation of Noto Cre recombinase has a high specificity in the study of NP degeneration (Table 3).
3.3.5. Ella
Under the control of the adenovirus Ella promoter that induces Cre expression only in the zygote and early embryo, and even can transmit genetic alterations to the offspring germ cells. Therefore, the level of expression of Ella has low specificity and it is seen as an important component of a tool for simultaneously knocking out target gene in a wide range of tissues. 121
3.3.6. Leptin receptor
The leptin receptor (Lepr) gene belongs to the obesity gene family, which encodes leptin receptor that recognizes and transports leptin. This gene is expressed in a variety of tissues, such as the hypothalamus, IVDs, cartilage, and bone. 122 , 123 , 124 , 125 , 126 What's more, it is an important marker of stem cells derived from the periosteum, mesenchymal stem cells and notochord derived cells, and has the characteristics of stem cells. It has been suggested that Lepr‐expressing cells can differentiate into chondrocytes, fibroblasts, osteoblasts, and adipose cells. And it can also be found in NP and OAF of IVDs. Therefore, the majority of Lepr‐expressing cells may appear at the very early stage of cell differentiation. In the field of IVDs research, Gao et al. 127 found that NP cells can be divided into different subgroups. By studying the Lepr Cre;R26R‐tdTomato mice which is generated by hybridization, they found that the number of NP cells of the lumbar IVDs specifically targeted by Lepr Cre nearly 75% gradually decreased from the P0, and no longer decreased after reaching the plateau at 1 month. OAF can be labeled by Lepr Cre from the P14 to adulthood. Therefore, the target gene can be knocked out by activating Lepr Cre recombinase on NP and OAF, and its relationship with disc degeneration can be explored (Table 3).
3.3.7. Osterix
Osterix (Osx), also known as SP7, is an active marker of osteoblasts. It is widely expressed in osteoblasts, most osteocytes, prehypertrophic chondrocytes of adult mouse long bones and adipocytes, perivascular cells, stromal cells in the bone marrow. 128 Recently, Osx expression has also been studied in IVDs. By crossing Osx Cre mice with R26‐mTmG mice, Zheng et al. observed IVD tissue sections of Osx Cre;R26‐mTmG offspring mice at 2 weeks and 2 months of age, and found that about 30% and 50% of cells in CEP were GFP positive, respectively. And some prehypertrophic and hypertrophic chondrocytes in GP were also GFP positive (Table 3). So it can be used as a tool to target IVDs.
3.3.8. Scleraxis
Scleraxis (Scx) plays an early essential role in mesoderm formation and is also an important marker of ligamentocytes and tenocytes. 129 , 130 The sites of Scx expression vary at different stages of mouse embryonic growth and development. Scx was detected in the body wall and lateral plate mesoderm between E9.5 and E10.5. It was widely expressed in various tissues such as long bones and kidney by E12.5 but only expressed in IVDs, tendons, and ligaments by E14.5. Sugimoto et al. 131 crossed Scx Cre‐H mice with R26R mice to generate offspring with the genotype Scx Cre‐H;R26R , and detected high Cre expression activity in the AF of the IVD, ligaments, and tendons (Table 3). It is concluded that Scx Cre recombinase may have some potential application in the study of IVD diseases.
3.3.9. Aggrecan
Aggrecan (Agc, Acan) is a large chondroitin sulfate proteoglycan in the ECM, which is widely distributed in IVD, cartilage, aorta and heart. 132 , 133 It is present in hyaline cartilage as aggregates that create osmotic swelling pressure gradients, drawing water into tissue. 134 Acan CreER mice produced based on endogenous aggrecan genes are important tamoxifen‐induced mice. Zheng et al. 79 generated Agc1 Cre(ER)T2;R26‐mTmG mice by hybridization and injected with tamoxifen at 2 weeks and 2 months of age. All chondrocytes in GP and NP cells were GFP positive, while nearly 50% of the cells in CEP of newborn and adult mice were GFP positive. As the mice grew older, the positive expression of GFP in AF cells decreased significantly. Therefore, Cre recombinase can be widely activated in IVD under tamoxifen induction (Table 3).
3.3.10. Cytokeratin 19
Cytokeratin 19 (Ck19, Krt 19, or K19) is a keratin family gene, which plays an important role in stress reactions, signal transduction pathway, and apoptosis. It is highly expressed in epithelial and peridermal cells of kidney, gallbladder and other tissues, and considered as a marker of cell proliferation. 135 , 136 , 137 , 138 , 139 In recent years, Krt19 has been found to be expressed in NP tissues (Table 3). Mohanty et al. 140 hybridized R26mTmG mice with Krt19 Cre(ER)T mice to generate Krt19 Cre(ER)T;R26mTmG mice. With the accumulation of tamoxifen, the GFP positive expression rate increased gradually in the NP of neonatal mice lumbar IVDs, even reaching 92% at P8. As the mice grew older, the positive expression rate of GFP in the NP cells gradually decreased to 72.64%. It's very important to note that Krt19 Cre(ER)T was not activated in the AF, EP or GP, indicating the high specificity of the Krt19 CreER mice in the NP for IVDs.
3.4. Examples of IVD specific Cre‐Flox model and the findings
In recent years, specific Cre‐Flox model has been widely used in the studies of IVD, which not only improves the specificity of the research, but also increases the accuracy. However, researchers should also pay attention to the defects brought by them, so as to provide the ideological basis for developing more perfect experimental models in the future.
3.4.1. Col2 Cre mice
It is known that Bmal1 plays an important role in bone and cartilage metabolism. When Bmal1 is knocked out, the differentiation of osteoclasts is promoted but the differentiation of chondrocytes and osteoblasts is inhibited, following diseases such as osteoporosis, IVDD and osteoarthritis. 141 Wang et al. 142 crossed Col2a1 Cre mice with Bmal1 flox/flox mice to generate cartilage/IVD specific Bmal1 knockout mice (Col2a1 Cre Bmal1 flox/flox , Bmal1 cKO). They found degenerative changes of NP and AF of lumbar IVDs with degradation of their ECM at 6 months of age. Further studies demonstrated that the RhoA/ROCK pathway may be involved in the disc degeneration in Bmal1 cKO mice. Due to ischemia and hypoxia in cartilage tissue, pathological changes related to inhibition of Hif1a‐Vegf signaling pathway may also occur in cartilage/IVD specific Bmal1 cKO mice. At the same time, the altered genes associated with endochondral ossification, such as Ptch1, Gli1 and Col10a1, may further promote the degeneration of CEP and GP. 141 Therefore, deletion of Bmal1 gene may cause disc degeneration. Meanwhile, Tessier et al. 143 found in the histological sections of the discs of the Col2 Cre Arpc2 flox/flox mice, the characteristic changes such as total absence of NP, irregular AF and no type II collagen in the CEP. However, the survival rate of the mice was extremely low. Similar disc degenerative changes were found in Adgrg6 cKO mice, Foxo cKO mice, Has2 cKO mice, Nf1 cKO mice, and Tsc1 cKO mice by crossbreeding Col2 Cre mice with flox mice carrying these target genes 144 , 145 , 146 , 147 , 148 (Table 4).
TABLE 4.
IVD phenotypes and function of genes that have been knocked out in cKO mice
| Model | IVD phenotypes or key information | Target gene | NCBI gene ID | Effect of each model on IVDD | References |
|---|---|---|---|---|---|
| Adgrg6 cKO mice (Col2 Cre Adgrg6 flox/flox ) | STAT3 signaling was upregulated and IVD stiffness increased, while the adult mice developed EP‐oriented herniations. | Adhesion G protein‐coupled receptor G6 | 215798 | Increase | 147 |
| Adgrg6 cKO mice (ATC; Adgrg6 flox/flox ) | STAT3 activation increased, EP‐oriented disc degeneration occurred | 147 | |||
| Arpc2 cKO mice (Col2 Cre Arpc2 flox/flox ) |
NP completely lost, AF disorganized, no type II collagen deposition in GP. Low survival rate |
Actin related protein 2/3 complex, subunit 2 | 76709 | Increase | 143 |
| Arpc2 cKO mice (Acan Cre(ER)T2 Arpc2 flox/flox ) | The boundary between NP and AF was gradually lost and protein deposition in ECM of the AF was observed. | ||||
| Bmal1 cKO mice (Col2a1 Cre Bmal1 flox/flox ) | NP, AF, and ECM all had degenerative changes at 6 months of age. | Aryl hydrocarbon receptor nuclear translocator‐like | 11865 | Increase | 142 |
| Ccn2 cKO mice (Noto Cre Ccn2 flox/flox ) | The level of type I collagen was increased, but type II collagen and Aggrecan were decreased in the NP of neonatal mice. | Cellular communication network factor 2 | 14219 | Increase | 165 |
| Ctnnb1 cKO mice (Shh Cre(ER)T2 Ctnnb1 flox/flox ) | Compressive stiffness of lumbar IVD reduced, wnt signal transduction decreased, Ccnd1 and aggrecan decreased, Osx and Adamts5 increased in NP | Catenin (cadherin associated protein), beta 1 | 12387 | Increase | 164 |
| Fam20B cKO mice (Col1 Cre Fam20B flox/flox ) |
The level of Col1 was decreased but Col2a1 and Sox9 were increased in OAF, even causing severe spinal malformations and IVD defects. There are some changes at the molecular level in the MAPK pathway. |
FAM20B, glycosaminoglycan xylosylkinase | 215015 | Increase | 153 |
| Foxo cKO mice (Col2a1 Cre Foxo flox/flox ) |
The boundary between NP and AF broken, the disorganized AF lamellae in IAF, and EP calcified GP tissue was abnormal and vertebral diameter increased. |
Forkhead box O1 forkhead box O3 forkhead box O4 |
56458 56484 54601 |
Increase | 148 |
| Foxo cKO mice (Acan CreER Foxo flox/flox ) |
Foxo expression decreased more significantly in NP than AF, but aggrecan expression was higher in NP than AF. GP, vertebral body and subchondral bone were normal. |
||||
| Has cKO mice (Col2 Cre Has2 flox/flox ) | Uneven surface of NP, low abundance of ECM and other disc defects | Hyaluronan synthase 1 | 15116 | Increase | 144 |
| Hif1a cKO mice (Shh Cre Hif1a flox/flox ) | The number of NP cells, the content of type II collagen and proteoglycan were all decreased. | Hypoxia inducible factor 1, alpha subunit | 15251 | Increase | 158 |
| Ift80 cKO mice (Col2 Cre(ER)T Ift80 flox/flox ) | Chondrocyte structure destroyed in GP, cell size and number reduced in EP, and cell morphology changed in IAF and NP | Intraflagellar transport 80 | 68259 | Increase | 152 |
| Ift80 cKO mice (Col1 Cre(ER)T Ift80 flox/flox ) |
The collagen fiber structure was disorganized and the number of apoptosis increased in OAF. At the same time, vertebral bone mass decreased significantly. |
||||
| Nf1 cKO mice (Col2a1 Cre Nf1 flox/flox ) | NP cells significantly reduced, AF no longer evolved and IVD structure destroyed | Neurofibromin 1 | 18015 | Increase | 146 |
| p16INK4a cKO mice (Acan Cre(ER)T2 p16INK4a flox/flox ) |
The number of apoptotic cells decreased, but not significantly. The function of aggrecan and collagen composition were changed. |
Cyclin dependent kinase inhibitor 2A | 12578 | Decrease | 181 |
| Runx2 cKO mice (Agc1 CreER Runx2 flox/flox ) | A large number of vacuolar notochordal cells in NP decreased, the proteoglycan matrix decreased, GP cartilage tissue expansion | Runt related transcription factor 2 | 12393 | Increase | 179 |
| Sirt1 cKO mice (Ella Cre Sirt1 floxed/+ ) | NP area invaded by fibrocartilage, fibers ruptured in AF, and chondrocyte‐like cells lost | Sirtuin 1 | 93759 | Increase | 168 |
| Sox9 cKO mice (Col2a1 Cre(ER)T Sox9 flox/flox ) |
Sox9 gene was deleted in NP cells and the expression of CTGF was decreased. Proteoglycan and retained water were lost in NP tissues. |
SRY (sex determining region Y)‐box 9 | 20682 | Increase | 151 |
| Tgfbr2 cKO mice (Acan tm1(Cre/ERT2)Crm Tgfbr2 flox/flox ) | Severe loss of proteoglycan in IVD and the collagen become thinner in IAF | Transforming growth factor, beta receptor II | 21813 | Increase | 180 |
|
Tsc1 cKO mice (Col2a1 Cre Tsc1 flox/flox ) |
Disc height was significantly reduced and the classical structure of the disc was absent in mice. | TSC complex subunit 1 | 64930 | Increase | 145 |
Abbreviations: Adamts5, a disintegrin and metalloproteinase with thrombospondin motifs 5; Ccnd1, Cyclin D1; CTGF, connective tissue growth factor; ECM, extracellular matrix; EP, endplates; GP, growth plate; IAF, inner annulus fibrosus; IVD, intervertebral disc; NP, nucleus pulposus; OAF, outer annulus fibrosus; Osx, osterix.
However, conditional gene knockout does not mean absolute specificity. In addition to the sites mentioned above, type II collagen is also expressed in other organs. Col2 Cre mice have been used in the studies of polycystic kidney disease. 149 As we all know, vitamin D needs to be hydroxylated a second time in the kidneys for it to function. The direct and indirect relationship between vitamin D and disc degeneration has already been confirmed. 150 Therefore, the target gene specifically knocked out in Col2 Cre mice for IVD research can also affect the function of the kidney. Defective kidney function may influence disc degeneration through ways other than the knockout gene. In such situations, different site specific Cre mice should be considered.
3.4.2. Col2 CreER mice
It was found that Col2 CreER recombinase cannot be activated in NP. But Interestingly, it has been suggested that the target gene can be knocked out in NP tissues by using Col2a1 Cre(ER)T mice. Oh et al. 151 injected tamoxifen intraperitoneally at P20 of Col2 CreER Sox9 flox/flox mice to knock out the Sox9 gene. Subsequently, Sox9 deletion and decreased expression of connective tissue growth factor (CTGF) were found in NP cells, and proteoglycan and retained water are lost in NP tissues of the mice lumbar discs. This special case is worthy of further study and discussion. Studies in which Col2 CreER mice were crossed with Flox mice that carry the target gene and generated Ift80 cKO mice after tamoxifen injection, showed disc defects and degeneration 152 (Table 4).
3.4.3. Col1 Cre mice
Saiyin et al. 153 generated Col1 Cre Fam20B flox/flox mice by hybridization and found that OAF in such mice showed significant defects. For example, OAF cells showed chondrocyte phenotype without fibroblast‐like morphology, and the expression of type I collagen was low, while the expression of type II collagen and Sox9 was high. At the same time, other phenotypic changes were observed in the IVDs. Such as the decline of EP height and NP area, as well as some decrease of heparin sulfate and chondroitin sulfate in AF (Table 4). Therefore, we believe that Col1 Cre recombinase may be activated at AF, EP, and NP in IVDs, but the recombination efficiency is the highest at OAF (Table 3).
Col1 Cre recombinase has high bone specificity, while wntless (Wls) is considered to be a susceptibility locus for osteoporosis. Lu et al. 154 constructed conditional Wls null mice of which the number of tartrate‐resistant acid phosphatase (TRAP) staining positive cells at the junction of cartilage and bone was significantly increased by crossing Col1 Cre mice with the flox mice. Its bone metabolism was obviously disturbed, which made it more susceptible to osteoporosis. At the same time, the influence of osteoporosis on disc degeneration has been supported by some studies. 155 , 156 Therefore, when Col1 Cre mice were used in the study of disc degeneration, we should pay attention to whether the target gene that has been knocked out will cause the phenotype of osteoporosis and exclude the possible impact of osteoporosis on disc degeneration.
3.4.4. Col1 CreER mice
Li et al. 152 also found significant changes in the structure of collagen fibers in OAF of lumbar spine and tail by using Col1 Cre(ER)T Ift80 flox/flox mice, such as uneven surface or disconnected fiber alignment, and an obvious reduction in bone mass of the vertebrae. The number of apoptotic cells in OAF significantly increased compared with WT mice (Table 4). At the same time, Rix et al. 157 found that partial deletion of Ift80 resulted in extremely high embryo lethality, even up to 98%. Not to mention what would happen if the gene were completely knocked out. Therefore, conditional gene knockout with tissue specificity and time controllability has become an essential method most of the time.
3.4.5. Shh Cre mice
In the study of disc degeneration, Meng et al. 158 crossed Shh Cre mice with Hif1a flox/flox mice to generate NP‐specific Hif1a cKO mice. The number of NP cells and the content of collagen II and proteoglycan significantly decreased, disc degeneration aggravated (Table 4). However, the mechanism of Hif1a action on IVDs is very complex, and some studies revealed that Hif1a could also aggravate disc degeneration in the case of severe hypoxia. 159 Therefore, under different oxygen tension microenvironments, Hif1a may have different effects on IVDs.
Shh Cre mice have also been used in the studies of nervous system disease. Ren et al. 104 used Shh Cre mice to establish mouse models of which Lepr was deleted in neural stem cells and found abnormal differentiation of hypothalamic neurons in the early postnatal period. This change also makes it possible for neuropeptide Y neurons to become abnormal. However, neuropeptide Y in hypothalamus plays an important role in metabolic regulation and is thought to prevent apoptosis of NP cells 160 , 161 . Therefore, when using Shh Cre mice to study disc degeneration, we should think about whether deletion of the gene would cause the neurological changes to affect the IVDs, thus interfering with our results. Nevertheless, Lep and Lepr not only affect disc degeneration directly, but also indirectly through hypothalamus and sympathetic nervous system. 162 If we study disc degeneration using Lep or Lepr cKO mice which are constructed by Shh Cre mice, undoubtedly, the results will be affected by all these aspects.
3.4.6. Shh CreER mice
In the application of this type of mice, Maier et al. 163 constructed Shh Cre(ER)T2;R26R Foxa1 ‐/‐ Foxa2 C/C offspring mice by hybridization, and found NP compression, IVD defects and an increase in cell death in the tail after injecting the mice with tamoxifen. However, when Foxa1 or Foxa2 was deleted, the phenotype of the mice did not change significantly, so the gene function may be redundant. In this special dKO case, the researchers conditionally knocked out the Foxa2 gene, which is prone to embryonic death, while the Foxa1 gene was knocked out by the traditional method. This flexible way fully elucidates the relationship between Foxa1 and Foxa2. When we take advantage of gene knockout technology to explore the relationship between phenotype and two or more genes in mice, the genetic selection is demanding. With the same result, Ctnnb1 cKO mice used by Holguin et al. also showed severe disc degeneration 164 (Table 4).
3.4.7. Noto Cre mice
Bedore et al. 165 constructed Noto Cre Ccn2 flox/flox mice to study disc degeneration. They found increased levels of type I collagen and decreased levels of type II collagen and Aggrecan in the NP of neonatal mice lumbar spine (Table 4). Thus, the deletion of Ccn2 significantly accelerated the process of ARD and demonstrated the high specificity of Noto Cre expression in NP tissues. Meanwhile, since the NP is the most active and abundant part of the IVDs for physiological regulation, Noto Cre mice is of great significance for the study of disc degeneration.
3.4.8. Ella Cre mice
Ella Cre is a kind of Cre line which is expressed in germline. 166 Studies have shown that the Ella Cre recombinase widely activated in early embryos of the mice, so it may be less specific than Noto Cre . 167 Because the size of Sirt1 −/− mouse pups is smaller than WT and the survival time after birth is shorter than 1 month, Xia et al. 168 crossed Ella Cre mice with Sirt1 floxed/+ mice to generate heterozygous mice whose genotype is Sirt +/− . Severe defects such as invasion of the NP area by fibrocartilage, ruptured fibers in AF, and loss of chondrocyte‐like cells were found after puncturing the coccygeal discs in these mice (Table 4). Therefore, the deletion of Sirt1 can aggravate IVDs injury and Ella Cre may be expressed in various parts of the IVDs (Table 3). Consequently, the simultaneous use of conditional knockout technology and heterozygous mice may significantly reduce the probability of mice death to a great degree.
Extensive activation of Ella Cre recombinases determines the wide applicability of Ella Cre mice. For instance, Billon et al. 169 utilized Ella Cre mice to study cerebellar ataxia, and the target gene retinoic acid receptor‐related receptor α (Rora) plays an important role in maintaining immune function and metabolic regulation in the body. The expression of pro‐inflammatory markers, such as Tnfa and Ifnγ, was significantly reduced in Rora cKO mice. However, other studies have found that these substances are closely related to disc degeneration. 170 , 171 Moreover, Ella Cre mice have also been used in the fields of tooth development, heart defects, and renal fibrosis. 172 , 173 , 174 This interlocking and complex effect of different organs is likely to affect the accuracy of the studies on disc degeneration.
3.4.9. Lepr Cre mice
Lepr Cre mice have not been widely used in studies of disc degeneration, because Lepr expression in many systems is not highly specific in the disc. When the target gene is knocked out, it may be disturbed in terms of nervous system, endocrine system, immune function, and energy metabolism, resulting in multiple organ abnormalities. The pathophysiological and pathological changes in these organs may also have impacts on disc degeneration, which makes it difficult to explore the relationship between target genes and disc degeneration. However, Lepr Cre mice can be useful if we investigate the effects of deletion of target gene on sympathetic regulation of NP cells.
3.4.10. Osx Cre mice
Osx Cre mice that target CEP and GP cells can be established and hybridize with Flox mice which have the LoxP locus to generate cKO mice for disc degeneration studies by knocking out the degeneration related target gene. 79 , 175 However, the Osx Cre mice are not highly specific and may have defects during the physical growth of the mice. Huang et al. 176 found that the body weight of Osx Cre mice are less than WT, and there are many fracture callus spots on fibula and ribs and delayed skeleton mineralization at P6. These phenotypes may be due to the impairment of endogenous gene function during the production of such mice, and may also have negative effects on the discs. Therefore, when we study the IVD phenotype caused by target genes, we should also consider the possible influence of skeletal defects in Osx Cre mice.
3.4.11. Scx Cre mice
Currently, Scx Cre mice, as an important tool, are mainly used for muscle and tendon studies, but are rarely used in the studies of disc degeneration. Researches shown that changes in the morphology and biological properties of paraspinal muscles and ligaments have also become important factors causing disc degeneration. 177 , 178 The effect of muscle and ligament tissue changes on disc degeneration should also be considered if researchers intend to cross Scx Cre mice with Flox mice to generate gene knockout offspring mice to study disc degeneration.
3.4.12. Acan CreER mice and ATC mice
In studies related to disc degeneration, Tessier et al. 143 crossed Acan Cre(ER)T2 mice with Arpc2 floxflox mice to generate Acan CreERT2 Arpc2 floxflox mice when Col2 Cre Arpc2 flox/flox mice they used had very low survival rate. The Arpc2 cKO mice were generated when tamoxifen was injected into mice at the age of 4 months. Thus, we need to select suitable Cre mice to cross with Flox mice when using conditional knockout technology to knock out target genes, in order to avoid mice death or other adverse effects. As Arpc2 cKO mice grow, the boundary between the lumbar NP and AF gradually disappears and protein deposition in ECM of the AF appears. At the same time, articular cartilage defects also can be seen. Therefore, the deletion of Arpc2 has a serious impact on the physiology and anatomical structure of mice skeleton, and Arpc2 is also associated with multi‐system diseases such as blood system and gastrointestinal system, hence the study of this gene has profound significance. Studies in which Agc CreER mice were crossed with Flox mice that carry the target gene and generated Foxo cKO mice, Runx2 cKO mice and Tgfbr2 cKO mice after tamoxifen injection, showed disc defects and degeneration. 148 , 179 , 180 Meanwhile, p16INK4a cKO mice showed the opposite effect on the discs. 181 In a word, Aggrecan is an important component of IVDs, and the construction of Acan CreER mice brings great convenience to the study of disc degeneration (Table 4).
So far, Acan CreER mice have been widely used in the studies of skeleton, but rarely in other fields. Maurer et al. 133 also confirmed that there was no significant association between cardiovascular risk factors and disc degeneration. Therefore, Acan CreER mice are good candidates to be used in the study of disc degeneration.
In the meantime, Liu et al. 147 also took advantage of a type of mouse which belongs to aggrecan enhancer‐driven, tetracycline‐inducible transgenic Cre (ATC) strain and its genotype is ATC;Adgrg6 flox/flox . They found that deletion of Adgrg6 can increase Stat3 activation, followed by EP‐oriented disc degeneration in mice lumbar spine. As a consequence, using ATC mice for research is also a feasible method. Meanwhile, this kind of mice has also been used to study craniofacial development. 182
3.4.13. Krt19 CreER mice
Krt19 CreER mice were mainly used in the studies of hepatobiliary diseases previously, while liver and gallbladder may not have much to do with disc degeneration. However, when Krt19 CreER mice were used in the study of disc degeneration, the effects on the kidney through tissue‐renin‐angiotensin system, vitamin D/VDR levels, and other possible factors, should be noticed. 183 , 184
3.4.14. Other potential Cre mice
Several genes have been shown to express in IVDs, but tissue/IVD‐specific Cre mice have not been constructed for the study of disc degeneration. According to the literature, keratan sulfate is expressed in the outer layers of the AF. Gli1, Ptch1 and chondroitin sulfate are expressed in almost all cells of IVDs. 185 Therefore, it is possible to make specific Cre mice according to the site of expression of these genes and activate Cre recombinases in this site for the study of disc degeneration.
Some genes expressed in IVDs have been made into Cre mice based on the tissue they express, but have never been used in the study of disc degeneration. For instance, Brachyury Cre mice have been used to study limb bud positioning, digit patterning and outgrowth. 186 As Brachyury is expressed in NP cells, Brachyury Cre mice could be utilized to study degeneration of the NP. 185 Similarly, Col10 Cre mice play an important role in the studies of endochondral ossification. 187 , 188 , 189 As Col10 is a cartilage osteogenic gene highly expressed in CEP, Col10 Cre mice also should be good candidates for the study disc degeneration. 190 Sox9 Cre mice have been applied in a variety of studies including skin, reproduction, kidney and skeleton. Meanwhile, Sox9 is expressed in almost all cells of IVDs. 185 If Sox9 Cre mice are used to study disc degeneration, the effects of other organs must be considered. In general, all of these Cre mice have the potential to be applied in the study of disc degeneration.
The mechanism of disc degeneration is very complicated. In addition to the causes within the IVDs, many other factors such as nervous system, endocrine regulation, energy metabolism and gut‐disc axis regulation may have influence on disc degeneration. Hence, other organs/tissues specific Cre mice may also be useful tools for the study of disc degeneration. For example, Hcrt‐IRES cre mice and Crh Cre mice target in hypothalamic neurons, Cdh16 Cre mice target in kidney, Plin1 CreER mice target in cartilage, and Adipoq CreER mice target in adipose tissue. Using these Cre mice to explore the relationship between target genes and disc degeneration has been suggested in further studies.
3.5. Authors' scientific perspectives on the strengths and limitations of the mouse model
Cre/LoxP system has been widely used in the study of disc degeneration. Different from the constitutive knockout technique, this system utilizes the expression of tissue‐specific promoters to ensure the specificity of the gene knockout site; Furthermore, based on the Cre/LoxP system, an estrogen‐induced Cre/(ER)T system is developed to achieve the time specificity of target gene knockout in the IVDs. It avoids the consequences of early embryo or newborn mouse death. The targeted build of this Cre/LoxP system is generally more flexible than the targeted build of classical methods. 82
The mouse models produced by this technique also have limitations. The production of Flox mice is usually complex and costly. Cre may be expressed in tissues other than IVD, causing unpredictable phenotypes and varying degrees of impact on experiments 191 (Table 1).
4. OTHER IMPORTANT TECHNOLOGIES
As with gene knockout technology, gene knockdown and knock‐in technology are very important gene edition technologies which are also widely used in scientific research.
Gene knockdown technology usually uses small double‐stranded RNA to efficiently and specifically degrade intracellular homologous mRNA, thereby inhibiting the expression of target genes, resulting in the phenotype of gene deletion. 83 , 192 Unlike knockout technology, this kind of technology does not permanently silence the expression of target genes, but inhibits gene expression by degrading mRNA of target genes which have homologous sequences, and achieves spatiotemporal and reversibility of gene expression regulation. At the same time, it can quickly and easily assess gene function in mammalian cells. Sometimes the small double‐stranded RNA is not inhibitory. Therefore, how to design an effective small double‐stranded RNA sequence that can be stably expressed has become a very concerned issue. 83
Similar to gene knockout technology, gene knock‐in technology is based on the principle of HR. The exogenous gene is transferred into the cell and recombined with the homologous sequence in the cell genome to make the cell phenotype change. The emergence of new technologies such as CRISPR/Cas9 has also made the implementation of gene knock‐in more common. However, the efficiency of gene knock‐in using CRISPR/Cas9 system is extremely low when compared with gene knockout, which may be an important reason that gene knock‐in technology is not seen in the study of disc degeneration. 193
5. DECISION CONSIDERATIONS
There are many kinds of mice to study disc degeneration. It is necessary to select a scientific and reasonable mouse model when you conduct experiments. First, you can select genes of interest with significant expression differences between healthy and degenerative samples by literature review, bioinformatics technology, or gene sequencing. The genes of interest were searched by NCBI gene ID and fully understood. Then you can search for mouse models for the gene of interest by going to The Jackson Laboratory's website or another mouse ordering sites. The type of mouse model should be fully considered before making a choice. For example, if you want to select constitutive KO mice for research, you should consider whether the mice will be lethal after knocking out the gene of interest. If it is a nonlethal gene, you can use homozygous KO for experiment. Instead, heterozygous KO mice or cKO mice could be selected for study. Sometimes cKO mice are not readily available, and you need to buy specific Cre mice and Flox mice to generate gene knockout progeny. However, sometimes site‐specific Cre mice cannot meet the needs of the disease background. For example, some diseases occur in young or old age, or just site‐specific Cre mice will still lead to early death of mice. At this point, you need to consider using CreER mice and Flox mice to breed offspring, and tamoxifen induction at specific stages to study disc degeneration. To this end, we can provide a clear guideline to decision considerations for the use of knockout mice by referring to the following flow chart (Figure 5).
FIGURE 5.
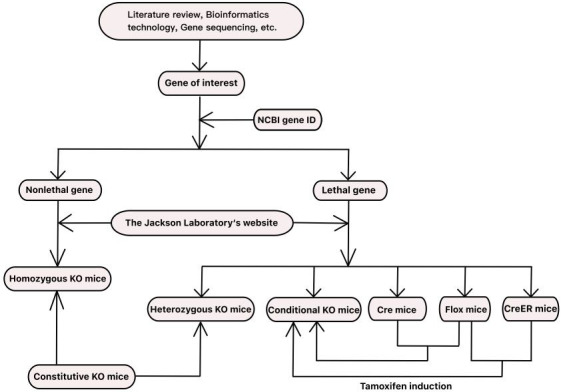
A clear guideline to decision considerations for the use of knockout mice. First, you can select genes of interest by literature review, bioinformatics technology, or gene sequencing. The genes of interest were searched by NCBI gene ID and fully understood. Then you can search mouse models by going to The Jackson Laboratory's website. If it is a nonlethal gene, you can use homozygous KO for experiment. Instead, heterozygous KO mice or cKO mice could be selected for study. Sometimes cKO mice are not readily available, and you need to buy specific Cre mice and Flox mice to generate gene knockout progeny. However, sometimes site‐specific Cre mice cannot meet the needs of the disease background. At this point, you need to consider using CreER mice and Flox mice to breed offspring, and tamoxifen induction at specific stages to study disc degeneration.
6. CONCLUSION AND FUTURE DIRECTION
With the development of science and technology, KO mice and cKO mice have been widely used and helped us to have a deeper understanding of the pathogenesis of disc degeneration. Although conditional knockout technology has become an inevitable trend, at present time constitutive knockout technology cannot be completely replaced. Conditional knockout technology has higher specificity, but it is not absolutely site‐specific. In this review, we described in detail the commonly used Cre mice and their characteristics in the study of disc degeneration. We also presented recommendations for the choice of suitable gene‐edited model mice. More and more accurate approaches will constantly be created on the base of shortcomings of previous technologies. We believe that the following problems are worth further discussing. First, several genes with high expression in IVDs have been engineered into Cre mice, but these Cre mice have not been used in studies of disc degeneration, and their usability needs to be further evaluated in the future. Second, we found that the studies about disc degeneration were carried out using mice as knockout animals. In the future, we can try in animals whose genotypes and phenotypes are more similar to human beings for verification. Finally, due to a variety of factors affecting disc degeneration, such as nervous system, gut‐disc axis regulation, endocrine disorders and immune regulation, the current knockout technology cannot rule out the impact of non‐target knockout site on target site research, so the technology needs to be further developed.
AUTHOR CONTRIBUTIONS
All authors have contributed to the drafting of the manuscript and its critical revision, and have approved the final version.
CONFLICT OF INTEREST
The authors declare no conflicts of interest.
ACKNOWLEDGMENTS
This work was supported by the National Natural Science Foundation of China (No. 82172474).
Lu, Z.‐Y. , Chen, P.‐B. , Xu, Q.‐Y. , Li, B. , Jiang, S.‐D. , Jiang, L.‐S. , & Zheng, X.‐F. (2023). Constitutive and conditional gene knockout mice for the study of intervertebral disc degeneration: Current status, decision considerations, and future possibilities. JOR Spine, 6(1), e1242. 10.1002/jsp2.1242
Ze‐Yu Lu and Peng‐Bo Chen contributed equally to this work and shared first authorship.
Funding information National Natural Science Foundation of China, Grant/Award Number: 82172474
Contributor Information
Lei‐Sheng Jiang, Email: jiangleisheng@xinhuamed.com.cn.
Xin‐Feng Zheng, Email: zhengxinfeng@xinhuamed.com.cn.
REFERENCES
- 1. Sato EM, Magalhães MO, Jenkins BC, et al. Low back pain in elderly from Belém‐Pa, Brazil: prevalence and association with functional disability. Healthcare (Basel, Switzerland). 2021;9(12):1658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Wang P, Zhang S, Liu W, et al. Selenium attenuates TBHP‐induced apoptosis of nucleus pulposus cells by suppressing mitochondrial fission through activating nuclear factor erythroid 2‐related factor 2. Oxid Med Cell Longev. 2022;2022:7531788. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Xin J, Wang Y, Zheng Z, Wang S, Na S, Zhang S. Treatment of intervertebral disc degeneration. Orthop Surg. 2022;14(7):1271‐1280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Xi Y, Ma J, Chen YJC. PTEN promotes intervertebral disc degeneration by regulating nucleus pulposus cell behaviors. Cell Biol Int. 2020;44(2):583‐592. [DOI] [PubMed] [Google Scholar]
- 5. Sampara P, Banala R, Vemuri S, Av G, Gpv SJG. Understanding the molecular biology of intervertebral disc degeneration and potential gene therapy strategies for regeneration: a review. Gene Ther. 2018;25(2):67‐82. [DOI] [PubMed] [Google Scholar]
- 6. Chen F, Liu H, Wang X, et al. Melatonin activates autophagy via the NF‐κB signaling pathway to prevent extracellular matrix degeneration in intervertebral disc. Osteoarthr Cartil. 2020;28(8):1121‐1132. [DOI] [PubMed] [Google Scholar]
- 7. Zhang G, Liu M, Chen H, et al. NF‐κB signalling pathways in nucleus pulposus cell function and intervertebral disc degeneration. Cell Prolif. 2021;54(7):e13057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Szymańska H. Genetically engineered mice: mouse models for cancer research. Postepy Hig Med Dosw (Online). 2007;61:639‐645. [PubMed] [Google Scholar]
- 9. Capecchi MR. The new mouse genetics: altering the genome by gene targeting. Trends Genetics. 1989;5(3):70‐76. [DOI] [PubMed] [Google Scholar]
- 10. Capecchi MR. Altering the genome by homologous recombination. Science (New York, NY). 1989;244(4910):1288‐1292. [DOI] [PubMed] [Google Scholar]
- 11. Vega MJH. Prospects for homologous recombination in human gene therapy. Human Genet. 1991;87(3):245‐253. [DOI] [PubMed] [Google Scholar]
- 12. Wijshake T, Baker D, van de Sluis B. Endonucleases: new tools to edit the mouse genome. Biochim Biophys Acta. 2014;1842(10):1942‐1950. [DOI] [PubMed] [Google Scholar]
- 13. Thomas K, Capecchi M. Site‐directed mutagenesis by gene targeting in mouse embryo‐derived stem cells. Cell. 1987;51(3):503‐512. [DOI] [PubMed] [Google Scholar]
- 14. Doetschman T, Gregg R, Maeda N, et al. Targetted correction of a mutant HPRT gene in mouse embryonic stem cells. Nature. 1987;330(6148):576‐578. [DOI] [PubMed] [Google Scholar]
- 15. Holmes A. Targeted gene mutation approaches to the study of anxiety‐like behavior in mice. Neurosci Biobehav Rev. 2001;25(3):261‐273. [DOI] [PubMed] [Google Scholar]
- 16. Kim YG, Cha J, Chandrasegaran S. Hybrid restriction enzymes: zinc finger fusions to Fok I cleavage domain. Proc Natl Acad Sci U S A. 1996;93(3):1156‐1160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Bibikova M, Golic M, Golic KG, Carroll D. Targeted chromosomal cleavage and mutagenesis in drosophila using zinc‐finger nucleases. Genetics. 2002;161(3):1169‐1175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Shankar S, Sreekumar A, Prasad D, Das AV, Pillai MR. Genome editing of oncogenes with ZFNs and TALENs: caveats in nuclease design. Cancer Cell Int. 2018;18:169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Semaan M, Ivanusic D, Denner J. Cytotoxic effects during Knock out of multiple porcine endogenous retrovirus (PERV) sequences in the pig genome by zinc finger nucleases (ZFN). PLoS One. 2015;10(4):e0122059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Cui X. Molecular aspects of zinc finger nucleases (ZFNs)‐mediated gene editing in rat embryos. Meth Mol Biol. 2019;1874:295‐312. [DOI] [PubMed] [Google Scholar]
- 21. Rémy S, Tesson L, Ménoret S, Usal C, Scharenberg AM, Anegon I. Zinc‐finger nucleases: a powerful tool for genetic engineering of animals. Transgenic Res. 2010;19(3):363‐371. [DOI] [PubMed] [Google Scholar]
- 22. Wu J, Kandavelou K, Chandrasegaran S. Custom‐designed zinc finger nucleases: what is next? Cell Mol Life Sci. 2007;64(22):2933‐2944. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Moscou MJ, Bogdanove AJ. A simple cipher governs DNA recognition by TAL effectors. Science (New York, NY). 2009;326(5959):1501. [DOI] [PubMed] [Google Scholar]
- 24. Boch J, Scholze H, Schornack S, et al. Breaking the code of DNA binding specificity of TAL‐type III effectors. Science (New York, NY). 2009;326(5959):1509‐1512. [DOI] [PubMed] [Google Scholar]
- 25. Lee JM, Kim U, Yang H, et al. TALEN‐mediated generation of Nkx3.1 knockout rat model. Prostate. 2021;81(3):182‐193. [DOI] [PubMed] [Google Scholar]
- 26. Mussolino C, Cathomen T. TALE nucleases: tailored genome engineering made easy. Curr Opin Biotechnol. 2012;23(5):644‐650. [DOI] [PubMed] [Google Scholar]
- 27. Sung YH, Baek IJ, Seong JK, Kim JS, Lee HW. Mouse genetics: catalogue and scissors. BMB Rep. 2012;45(12):686‐692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Mahata B, Biswas K. Generation of stable knockout mammalian cells by TALEN‐mediated locus‐specific gene editing. Meth Mol Biol. 2017;1498:107‐120. [DOI] [PubMed] [Google Scholar]
- 29. Bhardwaj A, Nain V. TALENs‐an indispensable tool in the era of CRISPR: a mini review. J Genet Eng Biotechnol. 2021;19(1):125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Li HL, Nakano T, Hotta A. Genetic correction using engineered nucleases for gene therapy applications. Dev Growth Differ. 2014;56(1):63‐77. [DOI] [PubMed] [Google Scholar]
- 31. Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J Bacteriol. 1987;169(12):5429‐5433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Wu W, Yang Y, Lei H. Progress in the application of CRISPR: from gene to base editing. Med Res Rev. 2019;39(2):665‐683. [DOI] [PubMed] [Google Scholar]
- 33. Mali P, Yang L, Esvelt KM, et al. RNA‐guided human genome engineering via Cas9. Science (New York, NY). 2013;339(6121):823‐826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Sarkar E, Khan A. Erratic journey of CRISPR/Cas9 in oncology from bench‐work to successful‐clinical therapy. Cancer Treat Res Commun. 2021;27:100289. [DOI] [PubMed] [Google Scholar]
- 35. Shojaei Baghini S, Gardanova ZR, Zekiy AO, Shomali N, Tosan F, Jarahian M. Optimizing sgRNA to improve CRISPR/Cas9 knockout efficiency: special focus on human and animal cell. Front Bioeng Biotechnol. 2021;9:775309. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 36. Krupkova O, Cambria E, Besse L, Besse A, Bowles R, Wuertz‐Kozak K. The potential of CRISPR/Cas9 genome editing for the study and treatment of intervertebral disc pathologies. JOR Spine. 2018;1(1):e1003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Fan Y, Zhao L, Lai Y, Lu K, Huang J. CRISPR‐Cas9‐mediated loss of function of β‐catenin attenuates intervertebral disc degeneration. Mol Ther Nucl Acids. 2022;28:387‐396. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Qi LS, Larson MH, Gilbert LA, et al. Repurposing CRISPR as an RNA‐guided platform for sequence‐specific control of gene expression. Cell. 2013;152(5):1173‐1183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Zhang D, Jin L, Reames DL, Shen FH, Shimer AL, Li X. Intervertebral disc degeneration and ectopic bone formation in apolipoprotein E knockout mice. J Orthop Res. 2013;31(2):210‐217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Li B, Zheng XF, Ni BB, et al. Reduced expression of insulin‐like growth factor 1 receptor leads to accelerated intervertebral disc degeneration in mice. Int J Immunopathol Pharmacol. 2013;26(2):337‐347. [DOI] [PubMed] [Google Scholar]
- 41. Lin D, Alberton P, Delgado Caceres M, et al. Loss of tenomodulin expression is a risk factor for age‐related intervertebral disc degeneration. Aging Cell. 2020;19(3):e13091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Pauli D, von Treuheim T, Torre OM, Ferreri ED, et al. Tenomodulin and chondromodulin‐1 are both required to maintain biomechanical function and prevent intervertebral disc degeneration. Cartilage. 2021;13:604‐614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Marfia G, Campanella R, Navone SE, et al. Potential use of human adipose mesenchymal stromal cells for intervertebral disc regeneration: a preliminary study on biglycan‐deficient murine model of chronic disc degeneration. Arthritis Res Ther. 2014;16(5):457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Zhang Q, Li J, Li Y, et al. Bmi deficiency causes oxidative stress and intervertebral disc degeneration which can be alleviated by antioxidant treatment. J Cell Mol Med. 2020;24(16):8950‐8961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Bach FC, Zhang Y, Miranda‐Bedate A, et al. Increased caveolin‐1 in intervertebral disc degeneration facilitates repair. Arthritis Res Ther. 2016;18:59. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Liu X, Wang D, Zhang Z, et al. Research on the function and related mechanism of P27 gene in the intervertebral disc degeneration of mice. Exp Ther Med. 2017;14(2):1141‐1145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Wei L, Cao P, Xu C, et al. Chondroitin synthase‐3 regulates nucleus pulposus degeneration through Actin‐induced YAP signaling. FASEB J. 2020;34(12):16581‐16600. [DOI] [PubMed] [Google Scholar]
- 48. Sahlman J, Inkinen R, Hirvonen T, et al. Premature vertebral endplate ossification and mild disc degeneration in mice after inactivation of one allele belonging to the Col2a1 gene for type II collagen. Spine. 2001;26(23):2558‐2565. [DOI] [PubMed] [Google Scholar]
- 49. Boyd LM, Richardson WJ, Allen KD, et al. Early‐onset degeneration of the intervertebral disc and vertebral end plate in mice deficient in type IX collagen. Arthritis Rheum. 2008;58(1):164‐171. [DOI] [PubMed] [Google Scholar]
- 50. Isaac C, Wright A, Usas A, et al. Dystrophin and utrophin "double knockout" dystrophic mice exhibit a spectrum of degenerative musculoskeletal abnormalities. J Orthop Res. 2013;31(3):343‐349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Kato M, Takaishi H, Yoda M, et al. GRIP1 enhances estrogen receptor alpha‐dependent extracellular matrix gene expression in chondrogenic cells. Osteoarthr Cartil. 2010;18(7):934‐941. [DOI] [PubMed] [Google Scholar]
- 52. Zieba J, Forlenza KN, Khatra JS, et al. TGFβ and BMP dependent cell fate changes due to loss of Filamin B produces disc degeneration and progressive vertebral fusions. PLoS Genet. 2016;12(3):e1005936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Feng C, Liu H, Yang Y, Huang B, Zhou Y. Growth and differentiation factor‐5 contributes to the structural and functional maintenance of the intervertebral disc. Cell Physiol Biochem. 2015;35(1):1‐16. [DOI] [PubMed] [Google Scholar]
- 54. Gorth DJ, Shapiro IM, Risbud MV. A new understanding of the role of IL‐1 in age‐related intervertebral disc degeneration in a murine model. J Bone Miner Res. 2019;34(8):1531‐1542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Phillips KL, Jordan‐Mahy N, Nicklin MJ, Le Maitre CL. Interleukin‐1 receptor antagonist deficient mice provide insights into pathogenesis of human intervertebral disc degeneration. Ann Rheum Dis. 2013;72(11):1860‐1867. [DOI] [PubMed] [Google Scholar]
- 56. Li X, Liu X, Wang Y, et al. Intervertebral disc degeneration in mice with type II diabetes induced by leptin receptor deficiency. BMC Musculoskelet Disord. 2020;21(1):77. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Gu SX, Li X, Hamilton JL, et al. MicroRNA‐146a reduces IL‐1 dependent inflammatory responses in the intervertebral disc. Gene. 2015;555(2):80‐87. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Hamrick MW, Pennington C, Byron CD. Bone architecture and disc degeneration in the lumbar spine of mice lacking GDF‐8 (myostatin). J Orthop Res. 2003;21(6):1025‐1032. [DOI] [PubMed] [Google Scholar]
- 59. Tang Z, Hu B, Zang F, Wang J, Zhang X, Chen H. Nrf2 drives oxidative stress‐induced autophagy in nucleus pulposus cells via a Keap1/Nrf2/p62 feedback loop to protect intervertebral disc from degeneration. Cell Death Dis. 2019;10(7):510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Li XF, Xue CC, Zhao YJ, et al. Deletion of Opg leads to increased neovascularization and expression of inflammatory cytokines in the lumbar intervertebral disc of mice. Spine. 2017;42(1):E8‐E14. [DOI] [PubMed] [Google Scholar]
- 61. Zhao YP, Tian QY, Liu B, et al. Progranulin knockout accelerates intervertebral disc degeneration in aging mice. Sci Rep. 2015;5:9102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Li CG, Liang QQ, Zhou Q, et al. A continuous observation of the degenerative process in the intervertebral disc of Smad3 gene knock‐out mice. Spine. 2009;34(13):1363‐1369. [DOI] [PubMed] [Google Scholar]
- 63. Whittal MC, Molladavoodi S, Zwambag DP, Millecamps M, Stone LS, Gregory DE. Mechanical consequence of induced intervertebral disc degeneration in the SPARC‐null mouse. J Biomech Eng. 2021;143(2):024501. [DOI] [PubMed] [Google Scholar]
- 64. Otsuki S, Alvarez‐Garcia O, Lotz MK, Neo M. Role of heparan sulfate 6‐0 endosulfatases in intervertebral disc homeostasis. Histol Histopathol. 2019;34(9):1051‐1060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. Tessier S, Tran VA, Ottone OK, et al. TonEBP‐deficiency accelerates intervertebral disc degeneration underscored by matrix remodeling, cytoskeletal rearrangements, and changes in proinflammatory gene expression. Matrix Biol. 2020;87:94‐111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Zhang C, Smith MP, Zhou GK, et al. Phlpp1 is associated with human intervertebral disc degeneration and its deficiency promotes healing after needle puncture injury in mice. Cell Death Dis. 2019;10(10):754. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Hoy RC, D'Erminio DN, Krishnamoorthy D, et al. Advanced glycation end products cause RAGE‐dependent annulus fibrosus collagen disruption and loss identified using in situ second harmonic generation imaging in mice intervertebral disk in vivo and in organ culture models. JOR Spine. 2020;3(4):e1126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68. Ngo K, Pohl P, Wang D, et al. ADAMTS5 deficiency protects mice from chronic tobacco smoking‐induced intervertebral disc degeneration. Spine. 2017;42(20):1521‐1528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Che H, Li J, Li Y, et al. p16 deficiency attenuates intervertebral disc degeneration by adjusting oxidative stress and nucleus pulposus cell cycle. Elife. 2020;9:e52570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70. Chen Z, Jiao Y, Zhang Y, et al. G‐protein coupled receptor 35 induces intervertebral disc degeneration by mediating the influx of calcium ions and upregulating reactive oxygen species. Oxid Med Cell Longev. 2022;2022:5469220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Ji ML, Jiang H, Zhang XJ, et al. Preclinical development of a microRNA‐based therapy for intervertebral disc degeneration. Nat Commun. 2018;9(1):5051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Kroon T, Bhadouria N, Niziolek P, et al. Suppression of Sost/Sclerostin and Dickkopf‐1 augment intervertebral disc structure in mice. J Bone Miner Res. 2022;37(6):1156‐1169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Ohnishi T, Yamada K, Iwasaki K, et al. Caspase‐3 knockout inhibits intervertebral disc degeneration related to injury but accelerates degeneration related to aging. Sci Rep. 2019;9(1):19324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Yokozeki Y, Uchida K, Kawakubo A, et al. TGF‐β regulates nerve growth factor expression in a mouse intervertebral disc injury model. BMC Musculoskelet Disord. 2021;22(1):634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75. Kameda T, Zvick J, Vuk M, et al. Expression and activity of TRPA1 and TRPV1 in the intervertebral disc: association with inflammation and matrix remodeling. Int J Mol Sci. 2019;20(7):1767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76. Ohta R, Tanaka N, Nakanishi K, et al. Heme oxygenase‐1 modulates degeneration of the intervertebral disc after puncture in Bach 1 deficient mice. Eu Spine J. 2012;21(9):1748‐1757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. Serjeant M, Moon PM, Quinonez D, Penuela S, Beier F, Séguin CA. The role of Panx3 in age‐associated and injury‐induced intervertebral disc degeneration. Int J Mol Sci. 2021;22(3):1080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78. Bhutta MF. Mouse models of otitis media: strengths and limitations. Otolaryngol Head Neck Surg. 2012;147(4):611‐614. [DOI] [PubMed] [Google Scholar]
- 79. Zheng Y, Fu X, Liu Q, et al. Characterization of Cre recombinase mouse lines enabling cell type‐specific targeting of postnatal intervertebral discs. J Cell Physiol. 2019;234(9):1‐10. doi: 10.1002/jcp.28166 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80. Weng W, Liu X, Lui K, Zhou B. Harnessing orthogonal recombinases to decipher cell fate with enhanced precision. Trends Cell Biol. 2022;32(4):324‐337. [DOI] [PubMed] [Google Scholar]
- 81. Gu H, Marth JD, Orban PC, Mossmann H, Rajewsky K. Deletion of a DNA polymerase beta gene segment in T cells using cell type‐specific gene targeting. Science (New York, NY). 1994;265(5168):103‐106. [DOI] [PubMed] [Google Scholar]
- 82. Kos C. Cre/loxP system for generating tissue‐specific knockout mouse models. Nutr Rev. 2004;62(6):243‐246. [DOI] [PubMed] [Google Scholar]
- 83. Miao X. Recent advances in the development of new transgenic animal technology. Cell Mol Life Sci. 2013;70(5):815‐828. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84. Bouabe H, Okkenhaug K. Gene targeting in mice: a review. Meth Mol Biol. 2013;1064:315‐336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85. Garcia E, Mills A. Getting around lethality with inducible Cre‐mediated excision. Semin Cell Dev Biol. 2002;13(2):151‐158. [DOI] [PubMed] [Google Scholar]
- 86. Kühn R, Torres RM. Cre/loxP recombination system and gene targeting. Meth Mol Biol. 2002;180:175‐204. [DOI] [PubMed] [Google Scholar]
- 87. Wamhoff B, Sinha S, Owens G. Conditional mouse models to study developmental and pathophysiological gene function in muscle. Handb Exp Pharmacol. 2007;178:441‐468. [DOI] [PubMed] [Google Scholar]
- 88. Alizadeh Sardroud H, Wanlin T, Chen X, Eames BF. Cartilage tissue engineering approaches need to assess fibrocartilage when hydrogel constructs are mechanically loaded. Front Bioeng Biotechnol. 2022;9:787538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89. Nagao M, Cheong CW, Olsen BR. Col2‐Cre and tamoxifen‐inducible Col2‐CreER target different cell populations in the knee joint. Osteoarthr Cartil. 2016;24(1):188‐191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90. Wei Y, Tower RJ, Tian Z, et al. Spatial distribution of type II collagen gene expression in the mouse intervertebral disc. JOR Spine. 2019;2(4):e1070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91. Li X, Yang S, Qin L, Yang S. Type II collagen‐positive embryonic progenitors are the major contributors to spine and intervertebral disc development and repair. Stem Cells Transl Med. 2021;10(10):1419‐1432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92. Wei Y, Tian Z, Tower RJ, et al. The inner annulus fibrosus encroaches on the nucleus pulposus in the injured mouse tail intervertebral disc. Am J Phys Med Rehabil. 2021;100(5):450‐457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93. Naomi R, Ridzuan PM, Bahari H. Current insights into collagen type I. Polymers. 2021;13(16):2642. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94. Cutroneo KR, White SL, Chiu JF, Ehrlich HP. Tissue fibrosis and carcinogenesis: divergent or successive pathways dictate multiple molecular therapeutic targets for oligo decoy therapies. J Cell Biochem. 2006;97(6):1161‐1174. [DOI] [PubMed] [Google Scholar]
- 95. Collette NM, Genetos DC, Economides AN, et al. Targeted deletion of Sost distal enhancer increases bone formation and bone mass. Proc Natl Acad Sci U S A. 2012;109(35):14092‐14097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96. Wang R, Ding Q, De Assuncao TM, et al. Hepatic stellate cell selective disruption of dynamin‐2 GTPase increases murine fibrogenesis through up‐regulation of sphingosine‐1 phosphate‐induced cell migration. Am J Pathol. 2017;187(1):134‐145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97. Bao Q, Li A, Chen S, et al. Disruption of bone morphogenetic protein type IA receptor in osteoblasts impairs bone quality and bone strength in mice. Cell Tissue Res. 2018;374(2):263‐273. [DOI] [PubMed] [Google Scholar]
- 98. Bedore J, Quesnel K, Quinonez D, Séguin CA, Leask A. Targeting the annulus fibrosus of the intervertebral disc: Col1a2‐Cre(ER)T mice show specific activity of Cre recombinase in the outer annulus fibrosus. J Cell Commun Signal. 2016;10(2):137‐142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99. Zhu XJ, Fang Y, Xiong Y, et al. Disruption of Wnt production in Shh lineage causes bone malformation in mice, mimicking human Malik‐Percin‐type syndactyly. FEBS Lett. 2018;592(3):356‐368. [DOI] [PubMed] [Google Scholar]
- 100. Xiong Y, Fang Y, Qian Y, et al. Wnt production in dental epithelium is crucial for tooth differentiation. J Dent Res. 2019;98(5):580‐588. [DOI] [PubMed] [Google Scholar]
- 101. Tang M, Luo SX, Tang V, Huang EJ. Temporal and spatial requirements of smoothened in ventral midbrain neuronal development. Neural Dev. 2013;8:8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102. Jiang M, Ku WY, Fu J, Offermanns S, Hsu W, Que J. Gpr177 regulates pulmonary vasculature development. Development. 2013;140(17):3589‐3594. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103. Gerhardt B, Leesman L, Burra K, et al. Notum attenuates Wnt/β‐catenin signaling to promote tracheal cartilage patterning. Dev Biol. 2018;436(1):14‐27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104. Ren Z, Liu Y, Hong W, et al. Conditional knockout of leptin receptor in neural stem cells leads to obesity in mice and affects neuronal differentiation in the hypothalamus early after birth. Mol Brain. 2020;13(1):109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105. Sreenivasan R, Gordon CT, Benko S, et al. Altered SOX9 genital tubercle enhancer region in hypospadias. J Steroid Biochem Mol Biol. 2017;170:28‐38. [DOI] [PubMed] [Google Scholar]
- 106. Wada Y, Yamauchi K, Murakami F, Tanabe Y. Temporally‐ and spatially regulated generation of distinct descendants by sonic hedgehog‐expressing progenitors in the forebrain. Dev Neurobiol. 2012;72(8):1099‐1113. [DOI] [PubMed] [Google Scholar]
- 107. Thirumangalathu S, Harlow DE, Driskell AL, Krimm RF, Barlow LA. Fate mapping of mammalian embryonic taste bud progenitors. Development. 2009;136(9):1519‐1528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108. Martí E, Bumcrot DA, Takada R, McMahon AP. Requirement of 19K form of sonic hedgehog for induction of distinct ventral cell types in CNS explants. Nature. 1995;375(6529):322‐325. [DOI] [PubMed] [Google Scholar]
- 109. Roelink H, Porter JA, Chiang C, et al. Floor plate and motor neuron induction by different concentrations of the amino‐terminal cleavage product of sonic hedgehog autoproteolysis. Cell. 1995;81(3):445‐455. [DOI] [PubMed] [Google Scholar]
- 110. Ericson J, Morton S, Kawakami A, Roelink H, Jessell TM. Two critical periods of sonic hedgehog signaling required for the specification of motor neuron identity. Cell. 1996;87(4):661‐673. [DOI] [PubMed] [Google Scholar]
- 111. Peck SH, McKee KK, Tobias JW, Malhotra NR, Harfe BD, Smith LJ. Whole transcriptome analysis of notochord‐derived cells during embryonic formation of the nucleus pulposus. Sci Rep. 2017;7(1):10504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112. Choi KS, Cohn MJ, Harfe BD. Identification of nucleus pulposus precursor cells and notochordal remnants in the mouse: implications for disk degeneration and chordoma formation. Develop Dyn. 2008;237(12):3953‐3958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113. Choi KS, Harfe BD. Hedgehog signaling is required for formation of the notochord sheath and patterning of nuclei pulposi within the intervertebral discs. Proc Natl Acad Sci U S A. 2011;108(23):9484‐9489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114. Di Gregorio A. The notochord gene regulatory network in chordate evolution: conservation and divergence from ciona to vertebrates. Curr Top Dev Biol. 2020;139:325‐374. [DOI] [PubMed] [Google Scholar]
- 115. Sayyid S, Wong P, Read W, et al. The clincoradiologic spectrum of notochordal derived masses. Clin Imaging. 2019;56:124‐134. [DOI] [PubMed] [Google Scholar]
- 116. Ramesh T, Nagula S, Tardieu G, et al. Update on the notochord including its embryology, molecular development, and pathology: a primer for the clinician. Cureus. 2017;9(4):e1137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117. Travill AA. The anatomical basis of clinical practice: an anatomy learning programme. Med Educ. 1977;11(6):377‐379. [DOI] [PubMed] [Google Scholar]
- 118. Corallo D, Trapani V, Bonaldo P. The notochord: structure and functions. Cell Mol Life Sci. 2015;72(16):2989‐3008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119. Lawson L, Harfe BD. Notochord to nucleus pulposus transition. Curr Osteoporos Rep. 2015;13(5):336‐341. [DOI] [PubMed] [Google Scholar]
- 120. McCann MR, Tamplin OJ, Rossant J, Séguin CA. Tracing notochord‐derived cells using a Noto‐cre mouse: implications for intervertebral disc development. Dis Model Mech. 2012;5(1):73‐82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121. Le Y, Sauer B. Conditional gene knockout using Cre recombinase. Mol Biotechnol. 2001;17(3):269‐275. [DOI] [PubMed] [Google Scholar]
- 122. Trayhurn P. Biology of leptin—its implications and consequences for the treatment of obesity. Int J Obes Relat Metab Disord. 2001;1:26‐28. [DOI] [PubMed] [Google Scholar]
- 123. de Vrind V, Van't Sant L, Rozeboom A, Luijendijk‐Berg M, Omrani A, Adan R. Leptin receptor expressing neurons in the substantia Nigra regulate locomotion, and in the ventral tegmental area motivation and feeding. Front Endocrinol. 2021;12:680494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124. Yen Y, Chien M, Wu P, Hung S. PP2A in LepR+ mesenchymal stem cells contributes to embryonic and postnatal endochondral ossification through Runx2 dephosphorylation. Commun Biol. 2021;4(1):658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125. Mo C, Guo J, Qin J, et al. Single‐cell transcriptomics of LepR‐positive skeletal cells reveals heterogeneous stress‐dependent stem and progenitor pools. EMBO J. 2022;41(4):e108415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126. Sadagurski M, Leshan RL, Patterson C, et al. IRS2 signaling in LepR‐b neurons suppresses FoxO1 to control energy balance independently of leptin action. Cell Metab. 2012;15(5):703‐712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127. Gao B, Yin J, Xu X, et al. Leptin receptor‐expressing cells represent a distinct subpopulation of notochord‐derived cells and are essential for disc homoeostasis. J Orthopaedic Transl. 2020;21:91‐99. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128. Chen J, Shi Y, Regan J, Karuppaiah K, Ornitz DM, Long F. Osx‐Cre targets multiple cell types besides osteoblast lineage in postnatal mice. PLoS One. 2014;9(1):e85161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129. Yoshimoto Y, Takimoto A, Watanabe H, Hiraki Y, Kondoh G, Shukunami C. Scleraxis is required for maturation of tissue domains for proper integration of the musculoskeletal system. Sci Rep. 2017;7:45010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130. Sun M, Luo EY, Adams SM, et al. Collagen XI regulates the acquisition of collagen fibril structure, organization and functional properties in tendon. Matrix Biol. 2020;94:77‐94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131. Sugimoto Y, Takimoto A, Hiraki Y, Shukunami C. Generation and characterization of ScxCre transgenic mice. Genesis. 2013;51(4):275‐283. [DOI] [PubMed] [Google Scholar]
- 132. Watanabe H. Aggrecan and versican: two brothers close or apart. Am J Physiol Cell Physiol. 2022;322(5):C967‐C976. [DOI] [PubMed] [Google Scholar]
- 133. Maurer E, Klinger C, Lorbeer R, et al. Association between cardiovascular risk factors and degenerative disc disease of the thoracolumbar spine in the general population: results from the KORA MRI study. Acta Radiol. 2022;63(6):750‐759. [DOI] [PubMed] [Google Scholar]
- 134. Cong L, Pang H, Xuan D, Tu GJ. Association between the expression of aggrecan and the distribution of aggrecan gene variable number of tandem repeats with symptomatic lumbar disc herniation in Chinese Han of northern China. Spine. 2010;35(14):1371‐1376. [DOI] [PubMed] [Google Scholar]
- 135. Mehrpouya M, Pourhashem Z, Yardehnavi N, Oladnabi M. Evaluation of cytokeratin 19 as a prognostic tumoral and metastatic marker with focus on improved detection methods. J Cell Physiol. 2019;234(12):21425‐21435. [DOI] [PubMed] [Google Scholar]
- 136. Stroescu C, Herlea V, Dragnea A, Popescu I. The diagnostic value of cytokeratins and carcinoembryonic antigen immunostaining in differentiating hepatocellular carcinomas from intrahepatic cholangiocarcinomas. J Gastrointestin Liver Dis. 2006;15(1):9‐14. [PubMed] [Google Scholar]
- 137. Olszewska E, Sudhoff H. Comparative cytokeratin distribution patterns in cholesteatoma epithelium. Compar Study. 2007;22(1):37‐42. [DOI] [PubMed] [Google Scholar]
- 138. Alix‐Panabières C, Vendrell J, Slijper M, et al. Full‐length cytokeratin‐19 is released by human tumor cells: a potential role in metastatic progression of breast cancer. Breast Cancer Res. 2009;11(3):R39. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139. Leelawat K, Narong S, Udomchaiprasertkul W, et al. Prognostic relevance of circulating CK19 mRNA in advanced malignant biliary tract diseases. World J Gastroenterol. 2012;18(2):175‐181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140. Mohanty S, Pinelli R, Dahia C. Characterization of Krt19 allele for targeting the nucleus pulposus cells in the postnatal mouse intervertebral disc. J Cell Physiol. 2020;235(1):128‐140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141. Chen G, Tang Q, Yu S, et al. The biological function of BMAL1 in skeleton development and disorders. Life Sci. 2020;253:117636. [DOI] [PubMed] [Google Scholar]
- 142. Wang D, Peng P, Dudek M, et al. Restoring the dampened expression of the core clock molecule BMAL1 protects against compression‐induced intervertebral disc degeneration. Bone Res. 2022;10(1):20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143. Tessier S, Doolittle AC, Sao K, et al. Arp2/3 inactivation causes intervertebral disc and cartilage degeneration with dysregulated TonEBP‐mediated osmoadaptation. JCI Insight. 2020;5(4):e131382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144. Roughley PJ, Lamplugh L, Lee ER, Matsumoto K, Yamaguchi Y. The role of hyaluronan produced by Has2 gene expression in development of the spine. Spine. 2011;36(14):E914‐E920. [DOI] [PubMed] [Google Scholar]
- 145. Yang C, Chen Y, Li Z, et al. Chondrocyte‐specific knockout of TSC‐1 leads to congenital spinal deformity in mice. Biomed Res Int. 2017;2017:8215805‐8215807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146. Wang W, Nyman JS, Ono K, Stevenson DA, Yang X, Elefteriou F. Mice lacking Nf1 in osteochondroprogenitor cells display skeletal dysplasia similar to patients with neurofibromatosis type I. Hum Mol Genet. 2011;20(20):3910‐3924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147. Liu Z, Easson GWD, Zhao J, et al. Dysregulation of STAT3 signaling is associated with endplate‐oriented herniations of the intervertebral disc in Adgrg6 mutant mice. PLoS Genet. 2019;15(10):e1008096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148. Alvarez‐Garcia O, Matsuzaki T, Olmer M, et al. FOXO are required for intervertebral disk homeostasis during aging and their deficiency promotes disk degeneration. Aging Cell. 2018;17(5):e12800. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149. Kolpakova‐Hart E, Nicolae C, Zhou J, Olsen B. Col2‐Cre recombinase is co‐expressed with endogenous type II collagen in embryonic renal epithelium and drives development of polycystic kidney disease following inactivation of ciliary genes. Matrix Biol. 2008;27(6):505‐512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150. Colombini A, De Luca P, Cangelosi D, et al. High‐throughput gene and Protein analysis revealed the response of disc cells to vitamin D, depending on the VDR FokI variants. Int J Mol Sci. 2021;22(17):9603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151. Oh CD, Yasuda H, Zhao W, et al. SOX9 directly regulates CTGF/CCN2 transcription in growth plate chondrocytes and in nucleus Pulposus cells of intervertebral disc. Sci Rep. 2016;6:29916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152. Li X, Yang S, Han L, Mao K, Yang S. Ciliary IFT80 is essential for intervertebral disc development and maintenance. FASEB J. 2020;34(5):6741‐6756. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153. Saiyin W, Li L, Zhang H, Lu Y, Qin C. Inactivation of FAM20B causes cell fate changes in annulus fibrosus of mouse intervertebral disc and disc defects via the alterations of TGF‐β and MAPK signaling pathways. Biochim Biophys Acta Mol Basis Dis. 2019;1865(12):165555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154. Lu C, Wan Y, Cao J, et al. Wnt‐mediated reciprocal regulation between cartilage and bone development during endochondral ossification. Bone. 2013;53(2):566‐574. [DOI] [PubMed] [Google Scholar]
- 155. Sun Q, Tian FM, Liu F, et al. Denosumab alleviates intervertebral disc degeneration adjacent to lumbar fusion by inhibiting endplate osteochondral remodeling and vertebral osteoporosis in ovariectomized rats. Arthritis Res Ther. 2021;23(1):152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156. Xiao ZF, He JB, Su GY, et al. Osteoporosis of the vertebra and osteochondral remodeling of the endplate causes intervertebral disc degeneration in ovariectomized mice. Arthritis Res Ther. 2018;20(1):207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157. Rix S, Calmont A, Scambler PJ, Beales PL. An Ift80 mouse model of short rib polydactyly syndromes shows defects in hedgehog signalling without loss or malformation of cilia. Hum Mol Genet. 2011;20(7):1306‐1314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158. Meng X, Zhuang L, Wang J, et al. Hypoxia‐inducible factor (HIF)‐1alpha knockout accelerates intervertebral disc degeneration in mice. Int J Clin Exp Pathol. 2018;11(2):548‐557. [PMC free article] [PubMed] [Google Scholar]
- 159. Li Y, Liu S, Pan D, et al. The potential role and trend of HIF‐1α in intervertebral disc degeneration: friend or foe? (review). Mol Med Rep. 2021;23(4):239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160. Rabal P, Coveñas R. Regulation of homeostasis by neuropeptide Y: involvement in food intake. Curr Med Chem. 2021;29(23):4026‐4049. [DOI] [PubMed] [Google Scholar]
- 161. Sun K, Zhu J, Sun J, et al. Neuropeptide Y prevents nucleus pulposus cells from cell apoptosis and IL‐1β‐induced extracellular matrix degradation. Cell Cycle. 2021;20(10):960‐977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162. Reid I, Baldock P, Cornish J. Effects of leptin on the skeleton. Endocr Rev. 2018;39(6):938‐959. [DOI] [PubMed] [Google Scholar]
- 163. Maier JA, Lo Y, Harfe BD. Foxa1 and Foxa2 are required for formation of the intervertebral discs. PLoS One. 2013;8(1):e55528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164. Holguin N, Silva MJ. In‐vivo nucleus pulposus‐specific regulation of adult murine intervertebral disc degeneration via Wnt/Beta‐catenin signaling. Sci Rep. 2018;8(1):11191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 165. Bedore J, Sha W, McCann MR, Liu S, Leask A, Séguin CA. Impaired intervertebral disc development and premature disc degeneration in mice with notochord‐specific deletion of CCN2. Arthritis Rheum. 2013;65(10):2634‐2644. [DOI] [PubMed] [Google Scholar]
- 166. Abdeen S, Del Mare S, Hussain S, et al. Conditional inactivation of the mouse Wwox tumor suppressor gene recapitulates the null phenotype. J Cell Physiol. 2013;228(7):1377‐1382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 167. Lakso M, Pichel JG, Gorman JR, et al. Efficient in vivo manipulation of mouse genomic sequences at the zygote stage. Proc Natl Acad Sci U S A. 1996;93(12):5860‐5865. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168. Xia X, Guo J, Lu F, Jiang J. SIRT1 plays a protective role in intervertebral disc degeneration in a puncture‐induced rodent model. Spine. 2015;40(9):E515‐E524. [DOI] [PubMed] [Google Scholar]
- 169. Billon C, Sitaula S, Burris T. Metabolic characterization of a novel RORα knockout mouse model without ataxia. Front Endocrinol. 2017;8:141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170. Hernandez P, Jacobsen T, Chahine N. Actomyosin contractility confers mechanoprotection against TNFα‐induced disruption of the intervertebral disc. Sci Adv. 2020;6(34):eaba2368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 171. Ye F, Lyu F, Wang H, Zheng Z. The involvement of immune system in intervertebral disc herniation and degeneration. JOR Spine. 2022;5(1):e1196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 172. Yu M, Liu Y, Wang Y, et al. Epithelial Wnt10a is essential for tooth root furcation morphogenesis. J Dent Res. 2020;99(3):311‐319. [DOI] [PubMed] [Google Scholar]
- 173. Lin X, Zhao W, Jia J, et al. Ectopic expression of Cripto‐1 in transgenic mouse embryos causes hemorrhages, fatal cardiac defects and embryonic lethality. Sci Rep. 2016;6:34501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 174. Xue H, Xiao Z, Zhang J, et al. Disruption of the Dapper3 gene aggravates ureteral obstruction‐mediated renal fibrosis by amplifying Wnt/β‐catenin signaling. J Biol Chem. 2013;288(21):15006‐15014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 175. Davis JL, Pokhrel NK, Cox L, Rohatgi N, Faccio R, Veis DJ. Conditional loss of IKKα in Osterix + cells has no effect on bone but leads to age‐related loss of peripheral fat. Sci Rep. 2022;12(1):4915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 176. Huang W, Olsen BR. Skeletal defects in Osterix‐Cre transgenic mice. Transgenic Res. 2015;24(1):167‐172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 177. Miki T, Naoki F, Takashima H, Takebayashi T. Associations between paraspinal muscle morphology, disc degeneration, and clinical features in patients with lumbar spinal stenosis. Prog Rehabil Med. 2020;5:20200015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 178. Ellingson AM, Shaw MN, Giambini H, An KN. Comparative role of disc degeneration and ligament failure on functional mechanics of the lumbar spine. Comput Methods Biomech Biomed Engin. 2016;19(9):1009‐1018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 179. Liao L, Jiang H, Fan Y, et al. Runx2 is required for postnatal intervertebral disc tissue growth and development. J Cell Physiol. 2019;234(5):6679‐6687. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 180. Alkhatib B, Liu C, Serra R. Tgfbr2 is required in Acan‐expressing cells for maintenance of the intervertebral and sternocostal joints. JOR Spine. 2018;1(2):e1025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 181. Novais EJ, Diekman BO, Shapiro IM, Risbud MV. p16(Ink4a) deletion in cells of the intervertebral disc affects their matrix homeostasis and senescence associated secretory phenotype without altering onset of senescence. Matrix Biol. 2019;82:54‐70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 182. Qi S, Wang Y, Wei X, et al. Expression of Cre recombinase in chondrocytes causes abnormal craniofacial and skeletal development. Transgenic Res. 2022;31(3):399‐411. [DOI] [PubMed] [Google Scholar]
- 183. Li Z, Wystrach L, Bernstein A, et al. The tissue‐renin‐angiotensin‐system of the human intervertebral disc. Eur Cell Mater. 2020;40:115‐132. [DOI] [PubMed] [Google Scholar]
- 184. Sun K, Sun X, Sun J, et al. Tissue renin‐angiotensin system (tRAS) induce intervertebral disc degeneration by activating oxidative stress and inflammatory reaction. Oxid Med Cell Longev. 2021;2021:3225439‐3225425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 185. Dahia CL, Mahoney E, Wylie C. Shh signaling from the nucleus pulposus is required for the postnatal growth and differentiation of the mouse intervertebral disc. PLoS One. 2012;7(4):e35944. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 186. Verheyden JM, Lewandoski M, Deng C, Harfe BD, Sun X. Conditional inactivation of Fgfr1 in mouse defines its role in limb bud establishment, outgrowth and digit patterning. Development. 2005;132(19):4235‐4245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 187. Park J, Gebhardt M, Golovchenko S, et al. Dual pathways to endochondral osteoblasts: a novel chondrocyte‐derived osteoprogenitor cell identified in hypertrophic cartilage. Biol Open. 2015;4(5):608‐621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 188. Rashid H, Chen H, Javed A. Runx2 is required for hypertrophic chondrocyte mediated degradation of cartilage matrix during endochondral ossification. Matrix Biol Plus. 2021;12:100088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 189. Jing Y, Zhou X, Han X, et al. Chondrocytes directly transform into bone cells in mandibular condyle growth. J Dent Res. 2015;94(12):1668‐1675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 190. Shafaei H, Bagernezhad H, Bagernajad H. Importance of floating chondrons in cartilage tissue engineering. World J Plast Surg. 2017;6(1):62‐67. [PMC free article] [PubMed] [Google Scholar]
- 191. Miura K, Ogura A, Kobatake K, Honda H, Kaminuma O. Progress of genome editing technology and developmental biology useful for radiation research. J Radiat Res. 2021;62:i53‐i63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 192. Kim D, Rossi J. RNAi mechanisms and applications. Biotechniques. 2008;44(5):613‐616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 193. Banan M. Recent advances in CRISPR/Cas9‐mediated knock‐ins in mammalian cells. J Biotechnol. 2020;308:1‐9. [DOI] [PubMed] [Google Scholar]


