Abstract
COVID-19 highlighted the need for use of real-world data (RWD) in critical care as a near real-time resource for clinical, research, and policy efforts. Analysis of RWD is gaining momentum and can generate important evidence for policy makers and regulators. Extracting high quality RWD from electronic health records (EHRs) requires sophisticated infrastructure and dedicated resources. We sought to customize freely available public tools, supporting all phases of data harmonization, from data quality assessments to de-identification procedures, and generation of robust, data science ready RWD from EHRs. These data are made available to clinicians and researchers through CURE ID, a free platform which facilitates access to case reports of challenging clinical cases and repurposed treatments hosted by the National Center for Advancing Translational Sciences/National Institutes of Health in partnership with the Food and Drug Administration. This commentary describes the partnership, rationale, process, use case, impact in critical care, and future directions for this collaborative effort.
Keywords: critical care, data harmonization, drug repurposing, electronic health record, real-world data, Observational Medical Outcomes Partnership (OMOP)
KEY POINTS
Question: The researchers pursued an answer to the automation of gathering, structuring, and harmonization of real-world data (RWD) from electronic health records for the purpose of generating real-world evidence.
Findings: The CURE Drug Repurposing Collaboratory and its various partners developed the Edge Tool, which is being successfully applied to use cases, including COVID-19, for structured and harmonized data.
Meanings: The implementation of the Edge Tool can result in the gathering of RWD and the harmonization of various data from different sources into the Observational Medical Outcomes Partnership Common Data Model with the purpose of updating the CURE ID platform.
Electronic health records (EHRs) provide the opportunity to leverage an abundance of real-world data (RWD) for many purposes including observational research, hypothesis generation, and external controls for nonrandomized clinical trials (1). In critical care, RWD may also support clinical and operational decision-making as well as enhance quality metrics (2–4). Currently, randomized controlled trials (RCTs) are considered the gold standard to assess the effectiveness of new drugs and therapeutics (5). However, in the face of diseases without adequate approved therapies, clinicians often use existing drugs approved for other indications (i.e., drug repurposing). While evidence of the performance of these therapies traditionally require RCTs, conducting RCTs are very expensive and time-intensive (6). Critical care providers treating emerging diseases (e.g., COVID-19) thus rely on repurposed drugs (e.g., remdesivir, tocilizumab) until enough evidence is gathered to obtain emergency use authorization (7). Treatments for other critical care conditions, such as sepsis and acute respiratory distress syndrome (ARDS), are challenging to study because of significant heterogeneity in pathology and lack of consensus around cohort definitions (8–11).
RWD and real-world evidence (RWE), or the potential benefits or risks of a medical product derived from analysis of RWD, increase availability of translational research data and evaluate effectiveness of repurposed therapeutics (12). The Food and Drug Administration (FDA) have provided guidance for utilizing RWD and RWE, which can be obtained through EHRs, insurance claims, product registries, among other resources (13). However, these data are often hard to obtain, and information pertinent to the problem of interest is difficult to extract. RWD has been plagued with data quality issues, including the nonstandardized nature of the data (14). Similarly, data lacks standardization when retrieved from different sources and requires harmonization to unify dissimilar data formats, definitions, units, etc. Therefore, publicly available tools designed to automate the extraction, harmonization, and standardization, as well as quality validation of critical care RWD, would increase its utility in developing RWE.
Other efforts and tools have been developed with a similar goal, although none make data completely public, accept nonstandardized data format, and are generalized to many diseases. For example, the National COVID Cohort Collaborative (15) require sites use a common data model (CDM) structure prior to contributing data. A platform that allows healthcare institutions, including those that have not transitioned to a CDM, to leverage harmonized RWD for COVID-19 and other diseases would be beneficial. The CURE Drug Repurposing Collaboratory (CDRC) is a broad coalition of stakeholders to provide public access to RWD through the CURE ID platform, thereby empowering clinical and translational research and decision efforts.
PARTNERS
CDRC is a public-private partnership between the Critical Path Institute, FDA, and National Center for Advancing Translational Sciences/National Institutes of Health. In 2021, FDA obtained funding from the U.S. Department of Health and Human Services Assistant Secretary for Planning and Evaluation’s Patient-Centered Outcomes Research Trust Fund to develop and deploy open access tools that automate extraction of RWD from EHRs and registries to populate the publicly available CURE ID platform with case reports for COVID-19, and in the future, other diseases.
CDRC brought together key partners (Fig. 1): the Society for Critical Care Medicine Discovery Critical Care Research Network and its Viral Infection and Respiratory Illness Universal Study COVID-19 Registry, Emory University, Johns Hopkins University, and the Infectious Diseases Data Observatory of the University of Oxford, Oxford, United Kingdom. These partners share expertise in automated data extraction and standardization from EHRs and registries and willingness to share data from their health systems and institutions. CDRC partners serve substantial roles in leading the development of the data flow and processes to transfer RWD from a variety of institutions to CURE ID.
Figure 1.
Partners in the CURE Drug Repurposing Collaboratory (CRDC). FDA = Food and Drug Administration, IDDO = Infectious Diseases Data Observatory, ISARIC DB = International Severe Acute Respiratory and emerging Infection Consortium Data Base, NCATS = National Center for Advancing Translational Sciences, NIH = National Institutes of Health, PCC = Partnership Coordinating Center, SCCM = Society for Critical Care Medicine, VIRUS = Viral Infection and Respiratory Illness Universal Study.
PIONEERING IN CRITICAL CARE
Delivery of evidence-based medicine in a critical care environment has unique challenges which have been amplified during the COVID-19 pandemic (16). Among other solutions, a source of rapid, robust, and reliable RWD can inform clinical practice in near real-time (2). Advanced analytics on intensive care treatments can positively impact outcomes for critically ill patients (17). The pandemic has also lowered the barriers among institutions and government agencies for pooling EHR data and making it broadly available. Curation of such data requires data harmonization and quality assessment solutions that work across multiple EHR vendors to lower the burden of manual data extraction, thereby making the process viable long-term (18). For instance, mechanical ventilation produces a wide range of measurements, settings, and event data (e.g., obstructions) often in a live stream integrated into the EHR. Merging data from multiple institutions is quite time consuming, however, which often limits the participation in large, observational studies to those institutions which have the resources to merge the datasets. Some challenges are relatively simple, such as aligning the labels for the variable (e.g., tidal volume might be recorded as “t_volume,” “tv,” or “tidal”) or ventilation mode (e.g., “volume-control,” “V/C,” etc.), or the measurements themselves may need to be harmonized if one hospital records positive end-expiratory pressure as a string with units included (e.g., “6 cm H2O”) and another records an integer (e.g., “6”). Other more complicated issues might include variations in data structure and formatting. Mapping of critically ill patients’ data to a CDM uniquely positions the critical care medicine community for large-scale collaboration for existing medical challenges and for future pandemic preparedness (19).
THE EDGE TOOL
The Observational Health Data Sciences and Informatics (OHDSI) collaborative developed the Observational Medical Outcomes Partnership (OMOP) CDM to facilitate scalable and reproducible observational research using EHRs. OMOP is a widely used model for longitudinal patient records (20). The OMOP CDM, which includes hundreds of affiliated projects on GitHub developed over the past decade, organizes EHR data into a standard set of tables containing standardized vocabularies (21), which allow for the organization and standardization of medical terms used across various clinical domains (Table 1).
TABLE 1.
Domains and Vocabulary Used to Extract Data From Electronic Health Records Using the Observational Medical Outcomes Partnership Common Data Model
| Vocabulary | Domain |
|---|---|
| Logical Observation Identifiers Names and Codes | Laboratory tests |
| RxNorm | Pharmacy |
| Systemized Nomenclature of Medicine-Clinical Terms | Clinical terms |
| International Classification of Diseases, 10th Revision | Mortality |
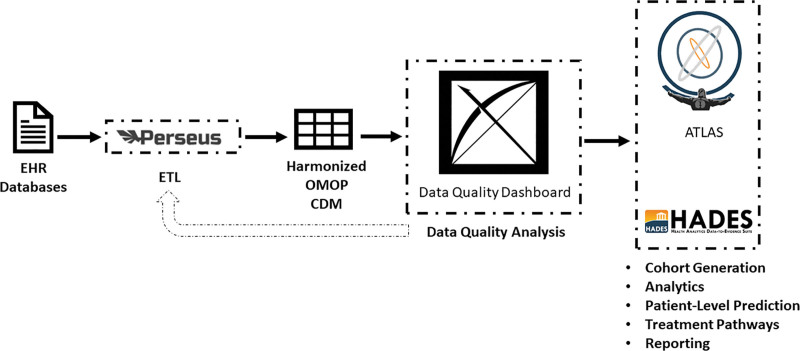
Through the Edge Tool, a group of public and free projects (Table 2) that enable the conversion of proprietary EHR data (i.e., data not stored in a CDM) to OMOP, relevant data can be extracted from its original EHR source, transformed to a format required by OMOP, and loaded into the CDM. This process is known as extract, transform, and load (ETL), and it involves extracting the data from its original source, transforming it by deduplicating, converting, cleaning, and combining it, and then loading it into a target database. This process allows data from different EHRs to be aggregated and analyzed together. Each project assists in the ETL process in tandem (Fig. 2) with the purpose of easing the ETL of existing EHR data, along with defining cohorts, treatment pathways, outcomes, and a data quality assessment that includes over 3,000 quality checks into a OMOP CDM that is interoperable. Through a user interface, The Edge Tool reduces the effort required to complete an ETL by generating coding and logic, which is normally written manually by an expert analyst or data scientist. The process of converting proprietary data models to the OMOP CDM had previously taken more than 2,000 dedicated hours from analysts and data scientists and has thus far reduced that number to 200 hours, with the hopes of eventually moving to as few as 50 hours of dedicated effort through continual improvement processes. The Edge Tool also includes a modern de-identification procedure that shifts the dates of occurrence by a static time period while maintaining the inter-temporal relationships within the patient, which preserves these temporal relationships in a Health Insurance Portability and Accountability Act (HIPAA)-compliant manner (22). This process meets the de-identification requirements under HIPAA, meaning that granularity can be retained in a de-identified or limited dataset, allowing researchers to calculate the time between events for specific analyses without requiring specific new variables be captured a priori (e.g., a researcher could calculate the time from admission to first dose of antibiotics, or the time between the first dose of dexamethasone and the first dose of tocilizumab).
TABLE 2.
Freely Available Data Management Tools Developed to Enable the Conversion of Proprietary Electronic Health Record Data to Observational Medical Outcomes Partnership
| Software Tools | Purpose |
|---|---|
| ATLAS | Facilitates the design and execution of analyses on standardized, patient-level, observational data in the OMOP CDM format |
| Health Analytics Data-To-Evidence Suite | Collection of open-source R packages to perform a complete observational study, including estimates, supporting statistics, figures, and tables |
| Data quality dashboard | Performs a harmonized data quality assessment to data standardized in the OMOP CDM |
| Perseus | Pre-processing, mapping, and conversion to OMOP CDM |
| More information about software and architecture is available at: www.ohdsi.org/software-tools/ | |
CDM = common data model, OMOP = Observational Medical Outcomes Partnership.
Figure 2.
Structure of tools within the Edge Tool software. CDM = common data model, EHR = electronic health record, ETL = extract, transform, and load, HADES = Health Analytics Data-To-Evidence Suite, OMOP = Observational Medical Outcomes Partnership.
One of the primary goals of the Edge Tool is lowering the barriers to implementing the OMOP CDM. The investigators worked closely with the Perseus team within OHDSI to enable efficiencies in the ETL process, specifically in refactoring three separate ETL tools (Table 2) in a graphic user interface to support mapping concepts from proprietary data models to OMOP. These tools are packaged together and implemented by healthcare institutions through existing OHDSI infrastructure with the purpose of generating hypotheses regarding the effectiveness of off-label drugs in a hospital encounter using RWD with higher quality to potentially inform future clinical trials designed to confirm said hypotheses. Once implemented, these tools can be re-run in the future with little resource effort, thus, significantly reducing barriers to sharing, as well as costs for collaboration.
COVID-19 AS USE CASE
COVID-19 was of high interest to CURE ID and provided a unique opportunity for automated data extraction in multiple critical aspects. First, as an emerging disease, there was ample opportunity for drug repurposing research (1). The high prevalence of the disease also allows many health systems to benefit from improved data collection and curation for research and quality purposes.
Second, cases were readily identifiable. Diagnosis of COVID-19 is made through a single laboratory test, which was widely available and captured in structured fields in the first year of the pandemic. Additionally, International Classification of Diseases, 10th Revision codes can be used to help phenotypically define cohorts describing COVID-19 cases.
Considerable early research efforts focused on radiological findings and natural history of COVID-19, requiring data from narrative reports (e.g., imaging interpretation, provider notes). CURE ID research interests, however, centered on treatment and mortality as outcome, thereby removing the requirement of abstracting from unstructured data, and instead focusing on structured data elements to answer the hypotheses related to these outcomes (e.g., laboratory results, medication administration) (23).
Lastly, COVID-19 has an acute disease course with data typically confined to a single hospital admission, reducing the challenges of linking visits between different facilities. However, resource limitations and improvisations, including construction of makeshift ICUs and “tent hospitals,” resulted in challenges since these efforts may not have used EHRs in the same way as the main wards.
RESULTS
CDRC supports 20 health systems of varying size and technical experience implementing new or expanding existing instances of the OMOP CDM. One system completed all phases of ETL and established data transfer processes, submitting more than 10,000 cases to CURE ID. Planned analyses include causal inference modeling of specific repurposed drugs. Sites interested in using the Edge Tool and researchers interested in accessing our RWD repository should contact our team (CDRC@c-path.org) or join the OHDSI community (www.ohdsi.org).
FUTURE DIRECTIONS AND LIMITATIONS
CDRC is working to expand the application of the Edge Tool to other diseases within the critical care space (e.g., sepsis, meningitis, ARDS), to other emerging diseases (e.g., mpox), and to diseases of high unmet medical need (e.g., osteomyelitis, sarcoma). Each disease poses unique challenges such as the need for incorporating natural language processing (NLP) to extract data from imaging (ARDS), microbiology (meningitis), pathology (sarcoma), or other narrative reports (mpox) or tracking patients across multiple encounters (osteomyelitis). Strategically developing solutions to the challenges presented by these and other diseases will allow the continued expansion of this technology.
The current limitations of the Edge Tool are the lack of implementation of an NLP tool for necessary data extraction, as well as the inability to connect patients to other encounters outside the healthcare site, confirm that a patient was administered a treatment, and resolve dates less than a year due to the time-shifted nature of the de-identification procedure used here.
Ultimately, tools that automate extraction of data from EHRs could be employed in clinical research, thus reducing the infrastructure needed for community centers and frontline workers to participate in clinical trials as a complement to traditional mechanisms. For repurposed drugs with well-established safety profiles, trials can be further decentralized through telemedicine, digital tools, and electronic patient-reported outcomes. Notably, expanding access of these tools to rural and underserved populations would improve the equity of study participation in minority populations, as well as diversity among study participants.
ACKNOWLEDGMENTS
We gratefully acknowledge the contributions and leadership of Heather Stone, MPH, and Leonard Sacks, MD.
Footnotes
This work was supported in part by the Intramural Research Program of the National Center for Advancing Translational Sciences, National Institutes of Health (1ZIATR000056-07). Critical Path Institute is supported by the Food and Drug Administration (FDA) of the U.S. Department of Health and Human Services (HHS) and is 54.2% funded by the FDA/HHS, totaling $13,239,950, and 45.8% funded by nongovernment source(s), totaling $11,196,634.
The views expressed are those of the author(s) and do not necessarily represent the official views of, nor an endorsement by, Food and Drug Administration/U.S. Department of Health and Human Services or the U.S. Government.
The authors have disclosed that they do not have any potential conflicts of interest.
Dr. Heavner led writing and review of this article, provided supervision, project administration and resources management for the project, validated findings, and assisted with conceptualization and funding acquisition. Dr. Anderson assisted with writing and coordinating review and editing. Ms. Dasher provided project administration and assisted with writing, review, and editing. Dr. Kashyap and Ms. Merson assisted with project administration and review and editing. Dr. Guerin and Mr. Weaver provided review and editing. Drs. Robinson, Schito, Kumar, and Nagy assisted with conceptualization, supervision, funding acquisition, writing, review, and editing.
REFERENCES
- 1.Concato J, Corrigan-Curay J: Real-World evidence — where are we now? N Engl J Med 2022; 386:1680–1682 [DOI] [PubMed] [Google Scholar]
- 2.Walkey AJ, Sheldrick RC, Kashyap R, et al. : Guiding principles for the conduct of observational critical care research for coronavirus disease 2019 pandemics and beyond: The Society of Critical Care Medicine Discovery Viral Infection and Respiratory Illness Universal Study Registry. Crit Care Med 2020; 48:e1038–e1044 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Peacock FW, Hauser GR, Toshev P, et al. : 59 Artificial intelligence occult sepsis detection in the emergency department: A large, multicenter real-world data study. Ann Emerg Med 2021; 78:S24 [Google Scholar]
- 4.Williams E, Szakmany T, Spernaes I, et al. : Discrete-event simulation modeling of critical care flow: New hospital, old challenges. Crit Care Explor 2020; 2:e0174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Derman BA, Belli AJ, Battiwalla M, et al. : Reality check: Real-world evidence to support therapeutic development in hematologic malignancies. Blood Rev 2022; 53100913. [DOI] [PubMed] [Google Scholar]
- 6.Corrigan-Curay J, Sacks L, Woodcock J: Real-World evidence and real-world data for evaluating drug safety and effectiveness. JAMA 2018; 320:867–868 [DOI] [PubMed] [Google Scholar]
- 7.Singh TU, Parida S, Lingaraju MC, et al. : Drug repurposing approach to fight COVID-19. Pharmacol Rep 2020; 72:1479–1508 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Sahoo BM, Ravi Kumar BVV, Sruti J, et al. : Drug repurposing strategy (DRS): Emerging approach to identify potential therapeutics for treatment of novel coronavirus infection. Front Mol Biosci 2021; 8:628144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ghosh CC, Thamm K, Berghelli AV, et al. : Drug repurposing screen identifies Foxo1-dependent angiopoietin-2 regulation in sepsis. Crit Care Med 2015; 43:e230–e240 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Fink MP, Warren HS: Strategies to improve drug development for sepsis. Nat Rev Drug Discov 2014; 13:741–758 [DOI] [PubMed] [Google Scholar]
- 11.Garrido-Mesa J, Adams K, Galvez J, et al. : Repurposing tetracyclines for acute respiratory distress syndrome (ARDS) and severe COVID-19: A critical discussion of recent publications. Expert Opin Investig Drugs 2022; 31:475–482 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Katkade VB, Sanders KN, Zou KH: Real world data: An opportunity to supplement existing evidence for the use of long-established medicines in health care decision making. J Multidiscip Healthc 2018; 11:295–304 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Dagenais S, Russo L, Madsen A, et al. : Use of real-world evidence to drive drug development strategy and inform clinical trial design. Clin Pharmacol Ther 2022; 111:77–89 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Cave A, Kurz X, Arlett P: Real-world data for regulatory decision making: Challenges and possible solutions for Europe. Clin Pharmacol Ther 2019; 106:36–39 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Haendel MA, Chute CG, Bennett TD, et al. ; N3C Consortium: The National COVID Cohort Collaborative (N3C): Rationale, design, infrastructure, and deployment. J Am Med Inform Assoc 2021; 28:427–443 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Walkey AJ, Kumar VK, Harhay MO, et al. : The viral infection and respiratory illness universal study (VIRUS). Crit Care Explor 2020; 2:e0113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Domecq JP, Lal A, Sheldrick CR, et al. ; Society of Critical Care Medicine Discovery Viral Infection and Respiratory Illness Universal Study (VIRUS): COVID-19 Registry Investigator Group: Outcomes of patients with coronavirus disease 2019 receiving organ support therapies: The international viral infection and respiratory illness universal study registry. Crit Care Med 2021; 49:437–448 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Khin NA, Grandinetti C, Dixey H, et al. : Tackling challenging data integrity topics in 2020: Update on good clinical practice perspectives from the US FDA and MHRA UK. Clin Pharmacol Ther 2022; 112:31–43 [DOI] [PubMed] [Google Scholar]
- 19.Paris N, Lamer A, Parrot A: Transformation and evaluation of the MIMIC database in the OMOP Common Data Model: Development and usability study. JMIR Med Inform 2021; 9:e30970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Lamer A, Abou-Arab O, Bourgeois A, et al. : Transforming anesthesia data into the observational medical outcomes partnership common data model: Development and usability study. J Med Internet Res 2021; 23:e29259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Seto T, Sung L, Posada J, et al. : Integrating flowsheet data in OMOP common data model for clinical research. arXiv Preprint posted online September 16, 2021. doi: 10.48550/arXiv.2109.08235 [Google Scholar]
- 22.Hripcsak G, Mirhaji P, Low AF, et al. : Preserving temporal relations in clinical data while maintaining privacy. J Am Med Inform Assoc 2016; 23:1040–1045 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Taksler GB, Dalton JE, Perzynski AT, et al. : Opportunities, pitfalls, and alternatives in adapting electronic health records for health services research. Med Decis Making 2021; 41:133–142 [DOI] [PMC free article] [PubMed] [Google Scholar]