Abstract
Background
Based on the constitution theroy, infants are classified into balanced constitution (BC) and unbalanced constitution. Yin-deficiency constitution (YINDC) is a common type of unbalanced constitutions in Chinese infants. An infant's gut microbiota directly affects the child's health and has long-term effects on the maturation of the immune and endocrine systems throughout life. However, the gut microbiota of infants with YINDC remains unknown. Herein, we aimed to evaluate the intestinal flora profiles and urinary metabolites in infant with YINDC, find biomarkers to identify YINDC, and promote our understanding of infant constitution classification.
Methods
Constitutional Medicine Questionnaires were used to assess the infants’ constitution types. 47 infants with 21 cases of YINDC and 26 cases of BC were included, and a cross-sectional sampling of stool and urine was conducted. Fecal microbiota was characterized using 16S rRNA sequencing, and urinary metabolomics was profiled using UPLC-Q-TOF/MS method. YINDC markers with high accuracy were identified using receiver operating characteristic (ROC) analysis.
Results
The diversity and composition of intestinal flora and urinary metabolites differed significantly between the YINDC and BC groups. A total of 13 obviously different genera and 55 altered metabolites were identified. Stool microbiome shifts were associated with urine metabolite changes. A combined marker comprising two genera may have a high potential to identify YINDC with an AUC of 0.845.
Conclusions
Infants with YINDC had a unique gut microbiota and metabolomic profile resulting in a constitutional microclassification. The altered gut microbiome in YINDC may account for the higher risk of cardiovascular diseases. Metabolomic analysis of urine showed that metabolic pathways, including histidine metabolism, proximal tubule bicarbonate reclamation, arginine biosynthesis, and steroid hormone biosynthesis, were altered in infants with YINDC. Additionally, the combined bacterial biomarker had the ability to identify YINDC. Identifying YINDC in infancy and intervening at an early stage is crucial for preventing cardiovascular diseases.
Keywords: Infant, Microbiota, Metabolomics, Biomarker, Traditional Chinese medicine, Yin deficiency constitution
1. Introduction
The TCM constitution (TCMC) is a comparatively consistent internal feature formed on the basis of inborn endowments and later characters, combined with one's lifetime morphological structure, physiological function and psychological state [1]. TCMC divides infants into nine constitutions: balanced constitution (BC), which indicates a comprehensive state of physical, mental and social well-being and less susceptibility toward illness, and eight unbalanced constitutions with their own unique biological characteristics and predisposition to different diseases [2,3]. For example, constipation is one of the most common diseases with a reported frequency of up to 30% [4], and 96% of cases are due to functional causes [5]. However, current diagnostic methods for functional constipation often fail to achieve cooperation among children or are harmful to their health. Furthermore, the evidence supporting the main therapeutic approaches is insufficient [6]. According to TCMC, constipation is a sign of yin-deficiency constitution (YINDC), which is a very common constitution in infants that is influenced by both congenital and acquired aspects. From the congenital aspect, the yin and yang bias of the parents' constitutions will affect the constitution of the offspring. Additionally, an improper diet during the mother's nurturing process will also cause YINDC in the offspring. Acquired aspects such as excessive consumption of fried foods, uncontrolled sexual life, over-intense emotions, specific natural and social environments, disease effects, and medical negligence can lead to YINDC [7]. YINDC is characterized by a deficiency of fluid (such as water) and the consequent presence of internal heat. Lack of fluid in the body will manifest as dry skin, lips, and eyes and constipation; internal heat presents as hot palms and soles, redness of the face and a preference for cold water. When the stomach yin is insufficient to fumigate the tongue, there is an uneven distribution of the tongue coating. At night, the deficient yin qi cannot restrain the yang qi, which forces sweating from the skin [8]. These YINDC manifestations can be treated with psychological adjustments, physical exercise, food therapy, herbal medicine, acupuncture, moxibustion, or massage [9].
Constitution can be identified by screening significantly different microbiota and metabolites and subsequently finding specific biomarkers. Increasing evidence shows that different constitutions have their own gut microbiota and metabolic characteristics [[10], [11], [12], [13]]; nevertheless, relevant researches have been performed mainly on adults. Recently, the gut microbiota of infants has drawn considerable attention because it directly affects the health of children at different developmental stages and has a long-term effects on the maturation of the immune and endocrine systems [[14], [15], [16]]. In particular, the first 2 years of life are a pivotal period of early childhood growing and developing when numerous microbiota gather and become stabilized, and the microbial community of the gastrointestinal tract serves as a key player in multiple pathways [17]. Accordingly, we focused on evaluating the correlation between 0 and 2 years old YINDC infants and intestinal microbiota in this study. Moreover, the metabolites generated by intestinal flora-host interactions are closely related to the phenotype and can directly reflect what happened in the body. Drawing blood samples from infants is inconvenient, easily infected, and does not indicate the total condition of the body [18], and stool samples are not as abundant as urine samples. Thus, we selected urine samples for metabolite detection. In this study, high-throughput 16S rRNA sequencing and untargeted metabolomics were used to gain insight into differential gut flora and metabolite characteristics in infants with YINDC aged 0–2 years old. This could help identify biomarkers and treat the corresponding diseases in infants with YINDC.
2. Materials and methods
2.1. Participants
0–2 years old infants were recruited via posters and social-media posts from Beijing and Guangdong, China. Constitutional Medicine Questionnaires were completed by the infants’ guardians, and only infants who met the criteria for conversion scores were screened for the next set of inclusion and exclusion criteria. The questionnaire comprised 43 items with 9 subscales, each measuring one constitution, namely, BC, items 1–3 and 43; qi–deficiency constitution, items 4–9; yang–deficiency constitution, items 10–13; YINDC, items 14–20; phlegm–dampness constitution, items 21–24; damp–heat constitution, items 25–29; blood stasis constitution, items 30–33; qi stagnation constitution, items 34–37; and inherited special constitution, items 38–42 (Supplemental file 1). Then, the conversion scores of the different constitutions corresponding to these items were calculated [19]. Individuals with BC score ≥60 and other conversions score <30 were considered to be BC. Individuals with YINDC were the ones with the highest YINDC score of ≥40 among all unbalanced constitutions.
YINDC and BC individuals were enrolled in fecal flora and urine metabolomics studies. The infants' guardians, generally their parents, were thoroughly informed of the trial procedure and signed informed consent forms. Ethical approval was acquired from the Ethics Committee of Beijing University of Chinese Medicine (No. 2020BZYLL122).
2.2. Inclusion and exclusion criteria
Participants were required to meet the following criteria: (1) infants aged 0–2 years old; (2) parents who had cared for the infants for a long time and had a detailed knowledge of the infant's basic condition; (3) infants with YINDC or BC diagnosed using the Constitutional Medicine Questionnaires.
Participants meeting one or more of the following criteria were excluded: (1) infants who received antibiotics and probiotics within 3 months prior to sample collection; (2) infants with a mental illness or severe disease that were unable to participate in and complete clinical studies in a standardized manner; (3) infants living far from the hospital where stool and urine samples were not readily available; (4) infants with incomplete clinical data.
2.3. Instrumentations
The following instrumentations were used: Qubit@ 2.0 Fluorometer (Thermo Scientific), Agilent Bioanalyzer 2100 system (Agilent), Bio-rad T100 thermal cycler (Bio-rad), NovaSeq6000 (Illumin), 1290 Infinity LC ultra-high pressure liquid chromatograph (Agilent Technologies), Triple TOF 6600+ Mass Spectrometer (AB SCIEX), Centrifuges (Eppendorf 5430R), 2.1 mm × 100 mm ACQUIY UPLC BEH 1.7 μm column (Waters, Ireland), Vortex (QiTe QT-1, Shanghai), MP Fastprep-24 Automated Homogenizer (MP Biomedicals), Ultrasonic Liquid Processors (Scientz JY92-II,Ningbo), Concentrator plus/Vacufuge (Eppendorf Concentrator Plus), −80 °C ultra-low temperature refrigerator (Thermo Scientific), Electronic balance (METLER TOLED AL104).
2.4. Chemicals
The following chemicals were used: Magnetic Soil And Stool DNA Kit (TIANGEN), Phusion®High-Fidelity PCR Master Mix (New England Biolabs), AxyPrepDNA Gel Extraction Kit (AXYGEN), TruSeq® DNA PCR-Free Sample Preparation Kit (TruSeq), Acetonirile (CAN, I592230123, Merck), NH3·H2O (221228, Sigma-Aldrich), Formic Acid (FA, 06450, Fluka), Ammonium acetate (70221, Sigma-Aldrich), MeOH (A456-4, Fisher Chemical).
2.5. Stool sample collection and sequencing
Freshly produced stool samples that did not contain urine were collected at home; >3 g (one-third the volume of the stool tube or two scoops) from the middle of the stool was scooped into a sterile stool cup. Then, the stool samples were transported with ice packs to a hospital within 1 h and stored at −80 °C after rapid freezing. Bacterial DNA was obtained by CTAB/SDS, and primers (341 F-806 R) were designed to amplify the 16S rRNA V3–V4 genes. Amplicons were mixed and purified, and amplicon libraries were generated. Lastly, the sequencing of the library yielded 250/300 bp paired-end reads [20].
2.6. Gut microbiota analysis
Paired-end reads were assembled and controlled for quality on 16S rRNA sequencing data. The sequences were performed with UPARSE. Sequences with ≥97% similarity were attributed to the same OTUs. Alpha diversity was used to evaluate the diversity of the microbial community within the sample and included the Chao 1, ACE, and Shannon. Beta diversity was applied to investigate the variation of biological groups between infants with YINDC and those with BC in species complexity. The linear discriminant analysis (LDA) of effect size (LEfSe) was able to discover taxa affecting the sample division. PICRUSt (version 1.1.1;) was used to predict the gene composition to analyze the functional differences in the KEGG database [21].
2.7. Urine sample collection and profiling
Middle part of urine were gathered in the morning in a sterile urine cup (>1.5 mL). Thereafter, the urine samples were transported to the hospital in ice bags in 1 h and centrifuged for 30 min (2000×g, 4 °C). The supernatant is then quickly frozen and stored at −80 °C. For analysis, the supernatant is thawed and mixed with pre-cooled methanol/acetonitrile solvent at a ratio of 1:4. The mixture was mixed by vortexing, sonicated at low temperature, held at −20 °C for 60 min, and centrifuged for 20 min (14000×g, 4 °C). After collecting, drying, redissolving the supernatant in acetonitrile/water solvent, it was again mediated by mixing for 1 min and centrifuged for 15 min. Finally, the supernatant was used for chromatographic analysis and subsequent Q-TOF mass analysis [22].
2.8. Urine metabolism analysis
ProteoWizard was used to convert the original data into mzXML format [23], and then XCMS was used for peak alignment, retention time correction and peak area extraction [24]. Metabolite structure identification was performed in a self-constructed laboratory database. 1 set of missing ion peak data >50% was removed. Data were preprocessed by Pareto scaling and subjected to multidimensional statistical analysis, represented as PCoA plots, and univariate analysis using Student 's t-test and fold change analysis, represented as volcano plots. The MetaboAnalyst was used for metabolic pathway analysis. The significance of the enrichment pathways was evaluated using Fisher's exact test.
2.9. Correlation analysis, ROC analysis, and statistical analysis
The correlation analysis was used to find the correlation for the differential flora and metabolites by Spearman. Heatmap and Network diagram analyses were performed using R and Cytoscape. The markers was measured by the area under the receiver operating characteristic (ROC) curve (AUC). For statistical analysis, SPSS and GraphPad Prism were used. P < 0.05 was considered a statistically significant difference.
3. Results
3.1. Basic information of infants
Among the 52 subjects recruited for this study, one subject without available samples, two whose parents were too busy to collect samples, and two whose samples failed sequencing or quality control were excluded. This left 47 infants, including 26 with BC and 21 with YINDC. Gut microbiota is well acknowledged to be affected by age, gender, delivery mode, and feeding pattern [[25], [26], [27], [28], [29]], so the YINDC and BC groups were compared using binary logistic regression analysis.No significant differences were observed (Table 1).
Table 1.
Influencing factors of the YINDC and BC groups.
| Influencing factors | YINDC | BC | P value |
|---|---|---|---|
| Age (months) | 12.48 ± 4.68 | 10.42 ± 6.67 | 0.104 |
| Gender | |||
| Boys | 16 (76.2%) | 14 (53.8%) | 0.113 |
| Girls | 5 (23.8%) | 12 (46.2%) | |
| Delivery Mode | |||
| Vaginal Delivery | 13 (61.9%) | 14 (53.8%) | 0.579 |
| Caesarean Delivery | 8 (38.1%) | 12 (46.2%) | |
| Feeding Pattern | |||
| Breast Feeding | 11 (52.4%) | 18 (69.2%) | 0.237 |
| Non-Breast Feeding | 10 (47.6%) | 8 (30.8%) | |
Note: Age values are shown as the mean ± standard deviation.
3.2. Alteration of gut microbiota diversity in infants with YINDC
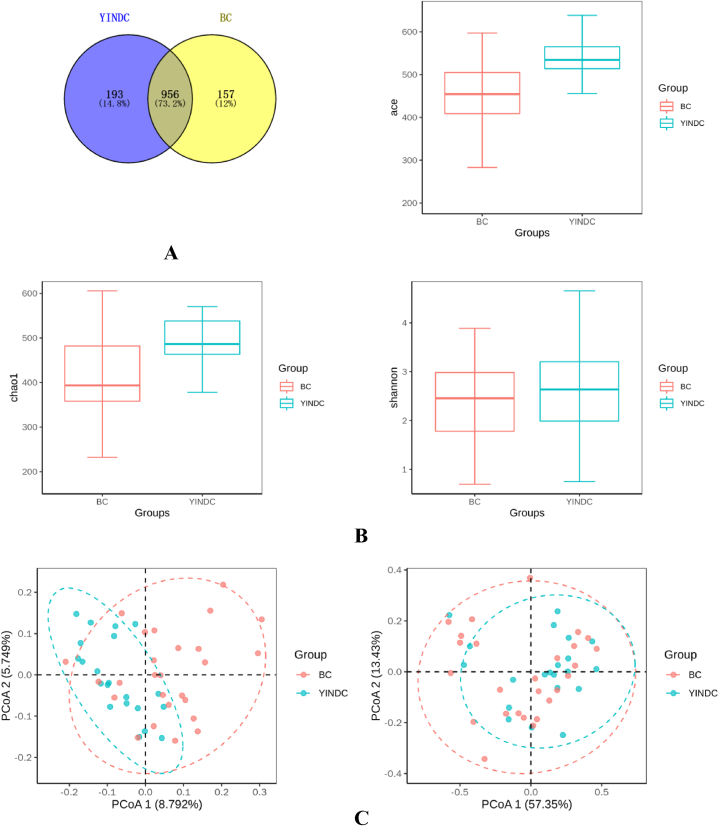
The stool samples yielded 110,207 and 124,236 reads of the YINDC and BC groups, respectively. They were classified into 193 and 157 unique OTUs in the YINDC and BC groups, respectively (Fig. 1A). Although the species evenness index Shannon was similar in two groups (p = 0.297), the species richness indices ACE and Chao 1 were higher in the YINDC group than in the BC group (all p < 0.01), which suggested that YINDC had a higher intestinal flora diversity than BC did (Fig. 1B). To investigate the clustering of microbial taxa in both groups, PCoA was constructed (Fig. 1C). Results revealed inconsistent YINDC and BC clusters based on unweighted (p = 0.00059) and weighted (p = 0.00068) unifrac distances, indicating that the microbial community structure significantly differed in both groups.
Fig. 1.
Intestinal microbial diversity analysis. (A) Venn diagram demonstrating the existence of different OTUs in the YINDC and BC groups. (B) Species alpha diversity differences between the two groups based on the ACE, Chao 1, and Shannon indices. (C) The overall microbial structure of the two groups as represented by PCoA of unweighted (left) and weighted (right) unifrac distances.
3.3. Alteration of gut microbiota composition in infants with YINDC
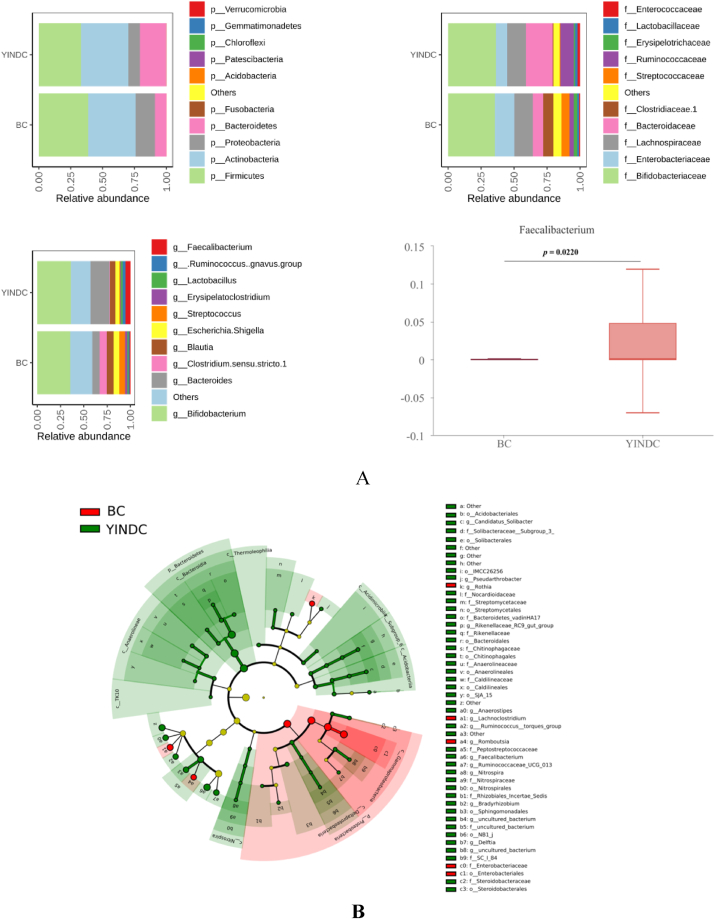
The relative proportions of dominant taxa at different levels were assessed, and specific taxa with statistically different proportions were identified in each group (Fig. 2A). At the phylum level, the YINDC and BC groups primarily comprised Actinobacteria, Firmicutes, Bacteroidetes, and Proteobacteria. Compared to the BC group, the relative abundance of Bacteroidetes (BC, 8.83%; YINDC, 20.54%) significantly increased (p = 0.0233), whereas the relative abundance of Proteobacteria (BC, 15.29%; YINDC, 9.04%) decreased (p = 0.0443) in the YINDC group. At the family level, Lachnospiraceae, Ruminococcaceae, and Bacteroidaceae were abundant in the YINDC and BC groups, consistent with the flora characteristic of infants on complementary foods [30]; At the genus level, Bifidobacterium and Bacteroides were abundant in both groups, and the proportion of Faecalibacterium (BC, 0.81%; YINDC, 5.27%) was higher in the YINDC group than in the BC group (p = 0.0220).
Fig. 2.
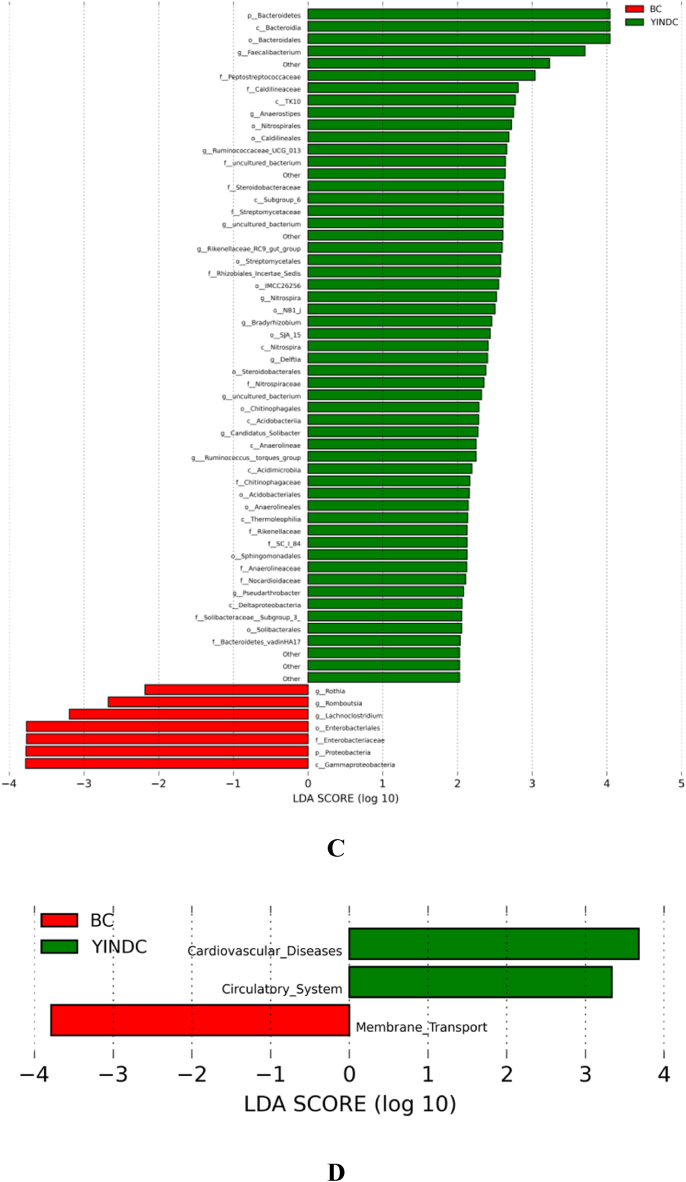
Alterations in gut microbiota associated with YINDC. (A) Relative abundance of the most prevalent bacterial phyla, families, genera and Faecalibacterium in the YINDC and BC groups. (B) Cladogram showing the distribution of intestinal microbial phylogeny. (C) Histograms of different taxa. (D) KEGG pathways of YINDC-associated intestinal flora.
To identify the significantly different intestinal flora between the YINDC and BC groups, LEfSe analysis was performed according to LDA score (log10) > 2.0, with p < 0.05. As expected, discriminative bacteria were identified in both groups. The YINDC group had a predominantly of phylum Bacteroidetes, and genera Ruminococcus__torques_group, Anaerostipes, Bradyrhizobium, Candidatus_Solibacter, Delftia, Faecalibacterium, Nitrospira, Pseudarthrobacter, Rikenellaceae_RC9_gut_group, Ruminococcaceae_UCG_013. Meanwhile, the microbiome in the BC group was characterized by the predominance of phylum Proteobacteria and genera Rothia, Romboutsia, and Lachnoclostridium (Fig. 2B and C).
3.4. Functional analysis of the microbiome in infants with YINDC
PICRUSt analyses were applied to evaluate the gut microbiota function involved in YINDC. Genes associated with cardiovascular disease (p = 0.003) and circulatory system (p = 0.003) were increased and genes associated with membrane transport (p = 0.029) were decreased in the YINDC group (Fig. 2D).
3.5. Altered urine metabolic profile in infants with YINDC
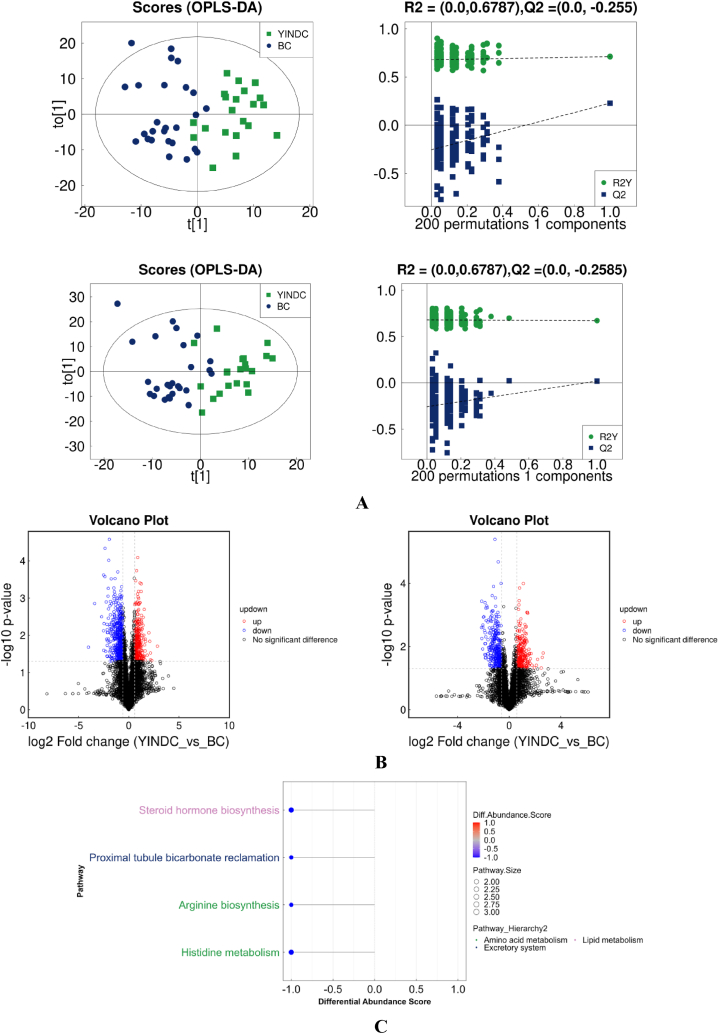
To explore the effect of YINDC on metabolism, urine samples were collected and assessed using untargeted metabonomics The samples of two groups were distinguished on different sides in two ion modes (Fig. 3A), suggesting that the metabolic profile of infants with YINDC was significantly altered.
Fig. 3.
Alterations in urine metabolites associated with YINDC. (A) OPLS-DA score plot in the positive- (above) and negative-ion (below) modes with VIP >1.0 and p < 0.05. (B) Volcano map of the significantly different metabolites in the positive (left) and negative (right) ion modes. Upregulated metabolites were detected by FC > 1.5 and p < 0.05, whereas downregulated metabolites were detected by FC < 0.67 and p < 0.05. (C) KEGG pathways of YINDC-associated metabolites.
3.6. Significantly different urine metabolites between the YINDC and BC groups
Univariate statistical analysis based on volcano plots was used to visualize the overall distribution difference of metabolites between the YINDC and BC groups. The upregulated and downregulated metabolites were shown in Fig. 3B. To further screen YINDC-related metabolites, OPLS-DA analysis was used. Finally, 55 metabolites were obtained and considered to be significantly different metabolites (Table 2). And the structural illustrations of the differential metabolites were shown in Supplemental file 2. Collectively, these results demonstrated that YINDC infants had a unique metabolic profile.
Table 2.
Differential metabolites between the YINDC and BC groups.
| Name | RT (s) | m/z | Adduction | VIP | p value |
|---|---|---|---|---|---|
| Mitoxantrone | 53.0615 | 443.17175 | [M − H]- | 10.08 | 0.0020 |
| d-Quinovose | 212.492 | 223.08057 | (M + CH3COO)- | 1.95 | 0.0025 |
| Amoxicillin | 53.131 | 729.18481 | [2M − H]- | 1.14 | 0.0030 |
| Arctigenin | 39.984 | 371.15127 | [M − H]- | 1.31 | 0.0032 |
| Cortisol 21-sulfate | 41.122 | 441.15545 | [M − H]- | 3.06 | 0.0050 |
| 1,5-anhydro-d-sorbitol | 214.764 | 163.05991 | [M − H]- | 1.84 | 0.0062 |
| Estrone glucuronide | 39.314 | 427.17627 | [M-H-H2O]- | 4.15 | 0.0063 |
| 3-dehydroepiandrosterone sulfate | 56.666 | 367.15666 | [M − H]- | 2.35 | 0.0081 |
| Alpha-d-Glucose | 433.422 | 179.05481 | (M − H)- | 1.20 | 0.0082 |
| [glu1] trh | 39.226 | 379.19233 | [M − H]- | 1.44 | 0.0096 |
| 2′-fucosyllactose | 431.907 | 487.16385 | [M − H]- | 2.03 | 0.0108 |
| Boldenone sulfate | 38.019 | 365.14084 | [M − H]- | 1.04 | 0.0113 |
| 1h-indazole-1-pentanoic acid,3-[[[1-(aminocarbonyl)-2,2-dimethylpropyl]amino]carbonyl]- | 57.347 | 373.16638 | [M − H]- | 1.82 | 0.0113 |
| Osajin | 86.01 | 403.17668 | [M − H]- | 1.53 | 0.0132 |
| Propionylglycine | 391.0055 | 130.0497 | [M − H]- | 1.11 | 0.0136 |
| Cortisone acetate | 79.548 | 401.19847 | (M − H)- | 1.93 | 0.0136 |
| 3,4-dihydroxyhydrocinnamic acid | 352.812 | 181.00802 | [M − H]- | 1.59 | 0.0142 |
| Acetic acid, 2-[4-[3-(4-acetyl-3-hydroxy-2-propylphenoxy)propoxy]phenoxy]- | 113.211 | 401.16014 | [M − H]- | 2.46 | 0.0169 |
| L-Galactono-1,4-lactone | 348.0265 | 215.0912 | (M + K–2H)- | 1.89 | 0.0170 |
| Alpha-ketoglutarate | 369.937 | 145.01292 | [M − H]- | 2.01 | 0.0170 |
| (2r,3s,4s,5r,6r)-2-[[(2s,3r,4r)-3,4-dihydroxy-4-(hydroxymethyl)oxolan-2-yl]oxymethyl]-6-[4-(4-hydroxyphenyl)butan-2-yloxy]oxane-3,4,5-triol | 114.254 | 459.16622 | [M − H]- | 4.39 | 0.0177 |
| pregnenolone sulfate | 70.595 | 395.18675 | (M − H)- | 3.64 | 0.0214 |
| Propylpyrazoletriol | 57.292 | 385.16626 | [M − H]- | 4.64 | 0.0216 |
| 3-(n-maleimidopropionyl)biocytin | 83.069 | 425.19754 | [M-H-C4H3NO2]- | 2.06 | 0.0240 |
| 4-pyridoxic acid | 52.365 | 182.04462 | [M − H]- | 9.70 | 0.0279 |
| Allantoin | 188.214 | 157.03556 | [M − H]- | 1.55 | 0.0286 |
| Cis,cis-muconic acid | 351.4185 | 141.01667 | [M − H]- | 11.68 | 0.0316 |
| Hydroxysebacic acid | 354.2715 | 217.10703 | [M − H]- | 3.34 | 0.0357 |
| (1-acetyloxy-3-hydroxy-6,8a-dimethyl-7-oxo-3-propan-2-yl-2,3a,4,8-tetrahydro-1h-azulen-4-yl) 4-hydroxybenzoate | 59.415 | 429.19212 | [M − H]- | 6.34 | 0.0394 |
| N-acetyl-l-glutamate | 385.099 | 188.05591 | [M − H]- | 1.85 | 0.0497 |
| 4-methyl-1h-pyrazole | 365.0525 | 83.05883 | [M+H]+ | 1.77 | 0.0011 |
| Erucamide | 40.5455 | 338.34094 | [M+H]+ | 6.62 | 0.0014 |
| 4-ketopimelic acid | 369.5635 | 129.06416 | [M + H–CH2O2]+ | 4.66 | 0.0030 |
| 5-methyl-2′-deoxycytidine | 205.012 | 126.06452 | [M + H–C5H8O3]+ | 2.31 | 0.0031 |
| Benzeneethanamine, 3,5-dimethoxy-.alpha.-methyl-4-propoxy- | 379.45 | 254.16003 | [M+H]+ | 2.36 | 0.0033 |
| Pro-Asn-Arg | 357.508 | 386.21833 | [M+H]+ | 1.44 | 0.0048 |
| 15-deoxy-goyazensolide | 139.501 | 284.33021 | [M+K]+ | 1.18 | 0.0049 |
| Ephedrine | 182.737 | 166.12144 | [M+H]+ | 3.68 | 0.0058 |
| Lumichrome | 72.1785 | 243.08667 | [M+H]+ | 4.38 | 0.0069 |
| (2r)-3-hydroxyisovaleroylcarnitine | 302.943 | 262.16418 | [M+H]+ | 2.91 | 0.0103 |
| 1-methylhistamine | 364.3375 | 126.08971 | [M+H]+ | 1.22 | 0.0121 |
| Didodecyl 3,3′-thiodipropionate oxide | 38.2345 | 531.40674 | [M+H]+ | 9.13 | 0.0148 |
| 1,5-pentanediamine | 264.4155 | 86.09527 | [M + H–NH3]+ | 4.33 | 0.0151 |
| N8-Acetylspermidine | 440.246 | 188.17435 | (M + H)+ | 1.65 | 0.0154 |
| Anserine | 301.8185 | 241.15358 | [M+H]+ | 6.24 | 0.0157 |
| 2-naphthalenol, 1-[(2,4-dimethylphenyl)azo]- | 40.715 | 277.1425 | [M+H]+ | 1.30 | 0.0171 |
| Asp-Asp-Arg | 431.97 | 203.06519 | [M+2H]2+ | 3.67 | 0.0209 |
| Adipoyl-l-carnitine | 390.888 | 290.15906 | [M+H]+ | 11.85 | 0.0263 |
| Gly-Arg | 374.997 | 232.15034 | [M+H]+ | 1.03 | 0.0303 |
| Asp-Asp-Lys | 373.823 | 189.07474 | [M+2H]2+ | 1.26 | 0.0311 |
| Vanylglycol | 50.5185 | 167.06886 | [M + H–H2O]+ | 1.49 | 0.0315 |
| Diethyl 2,4-dimethylpyrrole-3,5-dicarboxylate | 221.678 | 240.10067 | [M+H]+ | 1.13 | 0.0403 |
| Fusarenone-x | 131.117 | 377.1041 | [M+Na]+ | 1.18 | 0.0420 |
| N-acetylputrescine | 339.865 | 131.11631 | [M+H]+ | 2.84 | 0.0488 |
| 5-aminovaleric acid betaine | 387.571 | 160.13156 | [M+H]+ | 17.11 | 0.0497 |
3.7. Pathway analysis of the significantly different metabolites in infants with YINDC
To further investigate metabolic mechanisms of YINDC, the changed metabolites were mapped to physiological pathways. Differential abundance analysis scores showed that YINDC affected histidine metabolism, proximal tubule bicarbonate reclamation, arginine biosynthesis, and steroid hormone biosynthesis, with negative scores corresponding to the excretory system, amino acid metabolism, and lipid metabolism in the hierarchy 2 pathway (Fig. 3C).
3.8. Correlation between the significantly different flora and metabolites
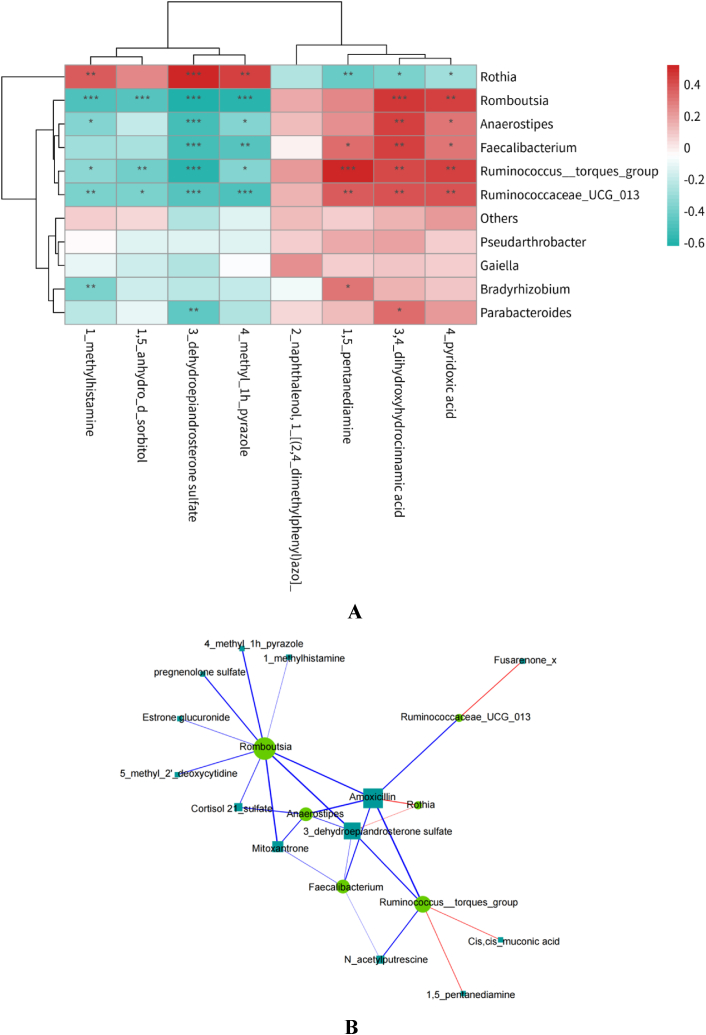
To determine if metabolic alterations related to intestinal flora, spearman analysis was performed. Several bacteria were very closely related to metabolites (Fig. 4A). We further narrowed the correlation coefficient to 0.5 < |r| < 1.26 pairs of significantly correlated bacteria and metabolites were screened, of which five pairs were positively correlated and 21 pairs were negatively correlated (Fig. 4B). Genera Romboutsia, Ruminococcus__torques_group, Anaerostipes, and Faecalibacterium were closely associated with the altered metabolites. Moreover, the correlation coefficient between Romboutsia and 3-dehydroepiandrosterone sulfate was >0.6, indicating that they may have a larger impact on YINDC.
Fig. 4.
Correlation analysis conducted by Spearman. (A) Heatmap of different microbiota and metabolites with |r| < 1. (B) Network diagram of significantly related microbiota and metabolites with 0.5 < |r| < 1. The red line and blue line indicate positive and negative correlation respectively. A darker color indicated a stronger correlation. (For interpretation of the references to color in this figure legend, the reader is referred to the Web version of this article.)
3.9. Microbial and metabolic markers of YINDC
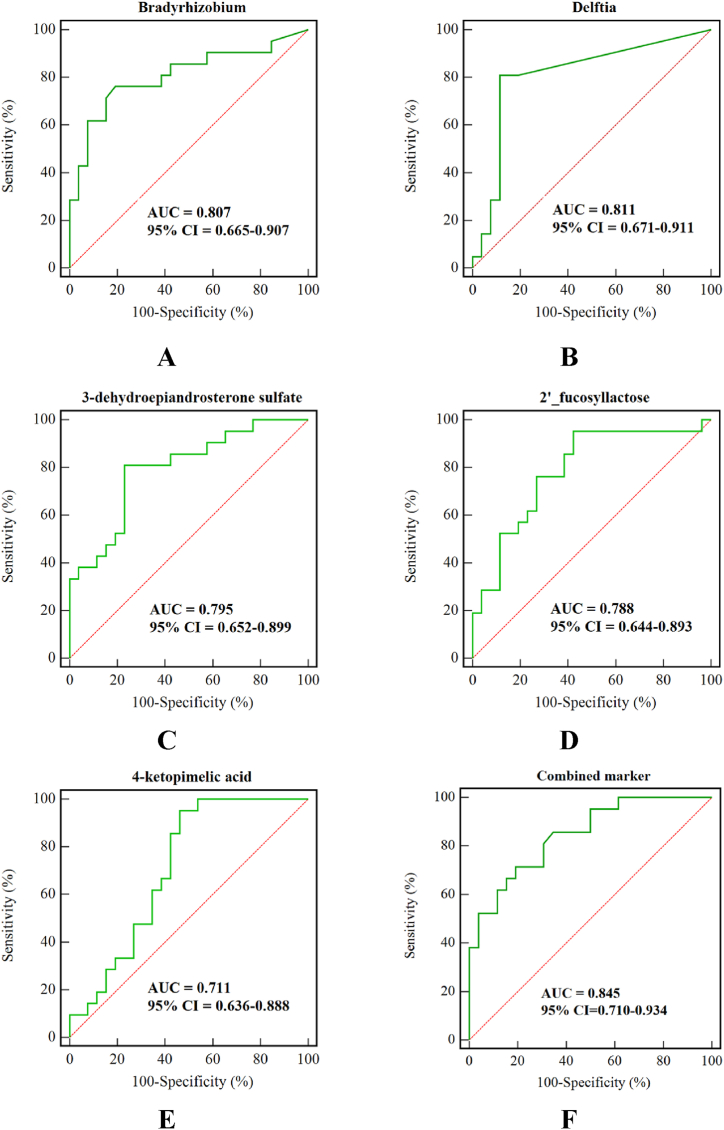
As an analytical method widely used to evaluate the predictive ability of models, ROC was used for the marker identification of YINDC. The top three metabolites were 3-dehydroepiandrosterone sulfate (AUC = 0.795, 95% CI = 0.652–0.899), 2′-fucosyllactose (AUC = 0.788, 95% CI = 0.644–0.893), and 4-pyridoxic acid (AUC = 0.780, 95% CI = 0.636–0.888). Genera with AUC >0.8 included Bradyrhizobium (AUC = 0.807, 95% CI = 0.665–0.907) and Delftia (AUC = 0.811, 95% CI = 0.671–0.911) (all p < 0.001) (Fig. 5A–E). Furthermore, the combination of two genus markers whose AUCs were >0.8 was found to have a higher diagnostic accuracy for YINDC than the markers alone (AUC = 0.845, 95% CI = 0.710–0.934, p < 0.001) (Fig. 5F). The combined marker had a sensitivity and specificity of 71.43% and 80.77%.
Fig. 5.
ROC curves for differential genera and metabolites of YINDC. (A) Genus Bradyrhizobium. (B) Genus Delftia. (C) Metabolite 3-dehydroepiandrosterone sulfate. (D) Metabolite 2′-fucosyllactose. (E) Metabolite 4-pyridoxic acid. (F) Combined marker composed of the two genus markers.
4. Discussion
Due to the fast development of modern detection methods and TCMC theory, more and more researchers are focusing on the microscopic study of YINDC. Four years ago, adults with YINDC were found to be associated with cardiovascular diseases (CVDs), infectious diseases, and certain cancers through salivary RNA [18]. In the current study, we found an increase in genes related to CVDs and the circulatory system in infants with YINDC via the intestinal microbiota. This suggests that YINDC may be associated with cardiovascular pathology, which remains a major contributor to global morbidity as well as mortality [31]. In a recently published meta-analysis, YINDC accounted for 14.8% of all patients with high normal blood pressure, which is a known risk factor for CVDs [32]. Therefore, identifying YINDC in infancy and intervening early are of critical importance in preventing CVDs. In this study, different gut microbiota of YINDC infants was identified, some of which were closely related to various CVDs.
The phylum Bacteroidetes, and its class Bacteroidia, and genus Bacteroides, which were abundant in the YINDC group, are considered to be the dominant flora in coronary heart disease. Bacteroidetes is the most numerous phylum of bacteria in the gut of the human body. It is engaged in polysaccharide fermentation, nitrogenous substances utilization, propionates production, and is often described as a beneficial bacterium in prior studies [33,34]. However, it is also participating in the process of protein hydrolysis with the release of toxic substances, thus promoting inflammation [35]. Shin et al. [36] found that Bacteroidetes was negatively related to HDL and acted as a counterpart in obesity. In this study, Bacteroidetes were abundant in YINDC where individuals are mostly thin, in agreement with the above study. Apart from the above bacteria, infants with YINDC had an increased abundance of Anaerostipes, Rikenellaceae_RC9_gut_group and Ruminococcus_torques_group and decreased abundance of Lachnoclostridium. Anaerostipes is reportedly associated with the development of CVDs [37]. Treatment with traditional Chinese medicine decreases its abundance in patients with chronic stable angina [38]. Some bacteria of this genus can reduce glucose levels and also promote propionate production through inositol or phytate, thereby reducing the risk of metabolic diseases [39,40]. Considerable changes in the Rikenellaceae_RC9_gut_group may be involved in acute myocardial ischemia through multiple pathways [41]. Ruminococcus torques is predictive of cardiac survival outcomes [42]. Lachnoclostridium counts are lower among those with higher maternal consumption of meat [43]. According to TCMC, YINDC is partially caused by excessive meat intake, which may be associated with reduced Lachnoclostridium level. Interestingly, the biomarker Bradyrhizobium, which can discriminate YINDC from BC, has been found to be negatively related to water intake and positively correlated with meat intake [44]. This could explained the preference for greasy, heavy-tasting food and the characteristics of water deficiency in individuals with YINDC.
In addition to being associated with CDVs, some genera are associated with the body shape of YINDC. For example, Ruminococcus_torques_group possibly modifies fat turnover. Low abundance of the group facilitates body fat control and enhances the contribution of resistant starch to abdominal obesity [45]. Conversely, an high abundance may exert adverse effects. Positive effect of probiotics on non-obese diabetic mice involved a reduction of this group [40]. Bacteria belonging to Ruminococcuss_torques_group probably consist of species that are involved in small intestinal permeability and immune responses during the first two years of life [46]. Specifically, it utilizes intestine mucin, whose increased mucin degradation allows luminal antigens to pass through the intestinal barrier, stimulating the immune system and causing intestinal diseases [47]. Infants with YINDC are usually underweight, possibly because Ruminococcus__torques_group increases intestinal permeability, thus nutrients cannot be completely digested and absorbed. A lower relative abundance of Anaerostipes is observed in obese patients and mice [48]. The consumption of lactic acid-producing bacteria increases the abundance of Anaerostipes, which is accompanied by weight loss [44]. The relative abundance of Lachnoclostridium is significantly higher in obese mice [49], which has been linked to the formation of visceral fat and is positively correlated with cholesterol levels [50,51]. Faecalibacterium is a butyrate producing bacterium with protective effects against obesity [52]. These changes in obesity-associated bacteria were also observed in the present study. Additionally, a high-fat diet reportedly results in the progression of primary Sjögren's syndrome [53], decreased Faecalibacterium is commonly observed among Sjogren's syndrome subjects [54]. The relationship between Faecalibacterium and dry symptoms requires further investigation.
“Was your child always excited” is an important personality item to determine whether an infant belongs to YINDC. If a child is always excited, he/she is more likely to behave as an individual with YINDC. The abundance of specific microbiota varies among people of different personalities. Participants with autism spectrum disorder have a decreased relative abundance of Ruminococcaceae UCG 013 [55], whereas excited YINDC individuals had an increased number of Ruminococcaceae UCG 013. Faecalibacterium, affiliated with the Ruminococcaceae family, is negatively correlated with the severity of depressive symptoms [56] and is more abundant in individuals with YINDC. Therefore, we speculated that the personality of YINDC infants may be correlated with the genera Ruminococcaceae UCG 013 and Faecalibacterium.
Several metabolites associated with CVDs were also identified in infants with YINDC, including upregulated metabolites, such as 4-pyridoxic acid and downregulated metabolites, such as 3-dehydroepiandrosterone sulfate. The concentration of 4-Pyridoxic acid correlates with cardiovascular risk, and a high 4-pyridoxic acid/pyridoxine ratio is considered to be relevant to the global cardiovascular risk [57]. Conversely, dehydroepiandrosterone sulfate levels is negatively associated with CVDs [58,59]. The major metabolic pathways of YINDC included histidine metabolism, arginine biosynthesis, steroid hormone biosynthesis, and proximal tubule bicarbonate reclamation. Consistent with the present study, Tabrez et al. [30] found histidine metabolism and arginine metabolism alteration by untargeted metabolomics in patients with CVDs. Histidine metabolism and arginine biosynthesis belong to amino acid metabolism. Current evidence confirms that a defect in branched chain amino acids (BCAAs) catabolism, and increased BCAAs levels are significantly correlated with CVDs risk [[60], [61], [62]]. Thus, BCAAs and their derivatives are supported as the potential biomarkers of CVDs [63]. A relative deficiency in steroid hormones, such as dehydroepiandrosterone, may be causally correlated with the development of a series of diseases associated with aging, including CVDs [64]. Cardiovascular risk is well known to increase in obese patients, but some overweight and obese individuals remain cardiometabolically healthy [65]. Innovatively, our study found an association between YINDC and CVDs, suggesting that thin individuals are also at risk of CVDs.
The microbial community structure significantly differed between the YINDC and BC groups, but there was partial overlap between the two groups of samples, probably due to the small sample size and within-sample variation. Spearman's correlation analysis showed that the level of critical metabolite 3-dehydroepiandrosterone sulfate was negatively related to the number of genera Ruminococcaceae UCG 013, Faecalibacterium, Ruminococcus__torques_group and positively correlated with the abundance of genus Rothia, which was in line with their separate trends. On the basis of the significantly different bacteria and metabolites, we found a combined marker comprising two bacterial markers Bradyrhizobium and Delftia to highly identify YINDC. These findings could help in the early diagnosis of YINDC so that early initiation of treatment could improve the imbalance state in infancy and prevent CVDs as early as possible.
5. Study limitations
This study had several limitations: (1) The age range of 0–2 years is considered a critical period for the functioning of the gut flora. Identification of structural changes in the gut flora during this time and narrowing of the age range requires further studies. (2) The sample size was limited and potential errors may exist. The use of 16S rRNA sequencing to analyze the gut microbiota has low phylogenetic power at the species level and poor discriminatory power for some genera; thus, our study findings need to be validated in macro-genomic studies with larger samples. (3) This study found that infants with YINDC and BC were not proportionally equal in terms of gender and feeding pattern, and the differences were not statistically significant. This may be influenced by the characteristics of different constitutions and requires studies related to the factors associated with constitution formation.
6. Conclusions
In conclusion, the present study revealed disordered gut microbiota and urinary metabolome in infants with YINDC, providing critical clues to its pathogenesis. Infants with YINDC may be at risk for CVDs, and this study provided several new targets for intervention in these complex diseases. The main metabolic pathways in infants with YINDC included histidine metabolism and arginine biosynthesis, etc. Additionally, the combined presence of genera Bradyrhizobium and Delftia had high diagnostic accuracy for YINDC. Multi-center studies with large sample sizes, such as bacterial transplantation experiments, in infants with YINDC are required to further study the key flora, metabolites, and their interactions identified in this study.
Author contribution statement
Haihong Zhao: Conceived and designed the experiments; Performed the experiments; Analyzed and interpreted the data; Wrote the paper.
Qiqi Ren; Huai-yu Wang: Analyzed and interpreted the data.
Yuhan Zong: Performed the experiments.
Weibo Zhao; Yaqi Wang; Miao Qu: Contributed reagents, materials, analysis tools or data.
Ji Wang: Conceived and designed the experiments.
Funding statement
National Key Research and Development Program of China (2020YFC2003100, 2020YFC2003103), the Project Supported by the Foundation of National Infrastructures for Translational Medicine (Shanghai) (TMSK-2021-414), Innovation Team and Talents Cultivation Program of National Administration of Traditional Chinese Medicine (ZYYCXTD-C-202001), Research Fund of Chinese Nutrition Society: Feihe Constitution Nutrition and Health (CNS-Feihe2018A01).
Data availability statement
Data will be made available on request.
Declaration of interest’s statement
The authors declare no conflict of interest.
Acknowledgments
We are grateful to all the participants in the study, including infants and their guardians. And we would like to thank Shanghai Applied Protein Technology Co. Ltd (Shanghai, China) for 16S rRNA sequencing and metabolomics service.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.heliyon.2023.e14684.
Appendix A. Supplementary data
The following are the Supplementary data to this article:
References
- 1.Wang Q. People's Health Publishing House; Beijing: 2009. Constitution of Chinese Medicine (2008) pp. 159–175. [Google Scholar]
- 2.Li L., Yao H., Wang J., et al. The role of Chinese medicine in health maintenance and disease prevention: application of constitution theory. Am. J. Chin. Med. 2019;47(3):495–506. doi: 10.1142/S0192415X19500253. [DOI] [PubMed] [Google Scholar]
- 3.Li Z. Beijing University of Chinese Medicine; Beijing: 2021. Compilation and Evaluation of Wang Qi's Nine Body Types in Constitutional Medicine Questionnaire (0-3 Years Version) [Google Scholar]
- 4.Van den Berg M.M., Benninga M.A., Di Lorenzo C. Epidemiology of childhood constipation: a systematic review. Am. J. Gastroenterol. 2006;101(10):2401–2409. doi: 10.1111/j.1572-0241.2006.00771.x. [DOI] [PubMed] [Google Scholar]
- 5.Loening-Baucke V. Prevalence, symptoms and outcome of constipation in infants and toddlers. J. Pediatr. 2005;146(3):359–363. doi: 10.1016/j.jpeds.2004.10.046. [DOI] [PubMed] [Google Scholar]
- 6.Tabbers M.M., DiLorenzo C., Berger M.Y., et al. Evaluation and treatment of functional constipation in infants and children: evidence-based recommendations from ESPGHAN and NASPGHAN. J. Pediatr. Gastroenterol. Nutr. 2014;58(2):258–274. doi: 10.1097/MPG.0000000000000266. [DOI] [PubMed] [Google Scholar]
- 7.Li Y., Wang Q. Discussion on the cause of yin-deficiency constitution. China J. Trad. Chin. Med. Pharm. 2012;27(12):3026–3028. [Google Scholar]
- 8.Wang Q. Classification and diagnosis basis of nine basic constitutions in Chinese medicine. J. Beijing Univer. Trad. Chin. Med. 2005;28(4):1–8. [Google Scholar]
- 9.Wang J., Li Y., Ni C., et al. Cognition research and constitutional classification in Chinese medicine. Am. J. Chin. Med. 2011;39(4):651–660. doi: 10.1142/S0192415X11009093. [DOI] [PubMed] [Google Scholar]
- 10.Jing Y., Han S., Chen J., et al. Gut microbiota and urine metabonomics alterations in constitution after Chinese medicine and lifestyle intervention. Am. J. Chin. Med. 2021;49(5):1165–1193. doi: 10.1142/S0192415X21500567. [DOI] [PubMed] [Google Scholar]
- 11.Ma K., Chen J., Kuang L., et al. Qi-deficiency related increases in disease susceptibility are potentially mediated by the intestinal microbiota. Evid. Bas. Comp. Alternat. Med. 2018;2018 doi: 10.1155/2018/1304397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Jing C., Xiao N., Zhang X., et al. Study on the composition and abundance of intestinal flora in phlegm-dampness constitution subjects based on 16S rDNA sequencing. J. Tradit. Chin. Med. 2019;60(23):5. [Google Scholar]
- 13.Liu Q., Luo B., Liang Jf, et al. Analysis of blood biochemical indexes and intestinal microflora diversity in women with constitution of yin deficiency and constitution of yin-yang harmony. Acta. Chin. Med. 2020;35(7):1514–1519. [Google Scholar]
- 14.Sarkar A., Yoo J.Y., Valeria Ozorio Dutra S., et al. The association between early-life gut microbiota and long-term health and diseases. J. Clin. Med. 2021;10(3):459. doi: 10.3390/jcm10030459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Li N., Liang S., Chen Q., et al. Distinct gut microbiota and metabolite profiles induced by delivery mode in healthy Chinese infants. J. Proteonomics. 2021;232 doi: 10.1016/j.jprot.2020.104071. [DOI] [PubMed] [Google Scholar]
- 16.Clarke G., Grenham S., Scully P., et al. The microbiome-gut-brain axis during early life regulates the hippocampal serotonergic system in a sex-dependent manner. Mol. Psychiatr. 2013;18(6):666–673. doi: 10.1038/mp.2012.77. [DOI] [PubMed] [Google Scholar]
- 17.Robertson R.C., Manges A.R., Finlay B.B., et al. The human microbiome and child growth - first 1000 Days and beyond. Trends Microbiol. 2019;27(2):131–147. doi: 10.1016/j.tim.2018.09.008. [DOI] [PubMed] [Google Scholar]
- 18.Chen Y., Wu Y., Yao H., et al. miRNA expression profile of saliva in subjects of yang deficiency constitution and yin deficiency constitution. Cell. Physiol. Biochem. 2018;49(5):2088–2098. doi: 10.1159/000493769. [DOI] [PubMed] [Google Scholar]
- 19.Zhao H., Zong Y., Li W., et al. Damp-heat constitution influences gut microbiota and urine metabolism of Chinese infants. Heliyon. 2022;9(2) doi: 10.1016/j.heliyon.2022.e12424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Xu Q., Su Z., Yang W., et al. Patchouli alcohol attenuates the cognitive deficits in a transgenic mouse model of Alzheimer's disease via modulating neuropathology and gut microbiota through suppressing C/EBPβ/AEP pathway. J. Neuroinflammation. 2023;20(1):19. doi: 10.1186/s12974-023-02704-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Langille M.G., Zaneveld J., Caporaso J.G., et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 2013;31(9):814–821. doi: 10.1038/nbt.2676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hu L., Che L., Wu C., et al. Metabolomic profiling reveals the difference on reproductive performance between high and low lactational weight loss sows. Metabolites. 2019;9(12):295. doi: 10.3390/metabo9120295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Chambers M.C., Maclean B., Burke R., et al. A cross-platform toolkit for mass spectrometry and proteomics. Nat. Biotechnol. 2012;30(10):918–920. doi: 10.1038/nbt.2377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Jia H., Shen X., Guan Y., et al. Predicting the pathological response to neoadjuvant chemoradiation using untargeted metabolomics in locally advanced rectal cancer. Radiother. Oncol. 2018;128(3):548–556. doi: 10.1016/j.radonc.2018.06.022. [DOI] [PubMed] [Google Scholar]
- 25.Palmer C., Bik E.M., DiGiulio D.B., et al. Development of the human infant intestinal microbiota. PLoS Biol. 2007;5(7):e177. doi: 10.1371/journal.pbio.0050177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Fallani M., Amarri S., Uusijarvi A., et al. Determinants of the human infant intestinal microbiota after the introduction of first complementary foods in infant samples from five European centres. Microbiology (Read.) 2011;157(Pt 5):1385–1392. doi: 10.1099/mic.0.042143-0. [DOI] [PubMed] [Google Scholar]
- 27.Kozyrskyj A.L., Kalu R., Koleva P.T., et al. Fetal programming of overweight through the microbiome: boys are disproportionately affected. J. Dev. Orig. Heal. Dis. 2016;7(1):25–34. doi: 10.1017/S2040174415001269. [DOI] [PubMed] [Google Scholar]
- 28.Biasucci G., Rubini M., Riboni S., et al. Mode of delivery affects the bacterial community in the newborn gut. Early Hum. Dev. 2010;86(Suppl 1):13–15. doi: 10.1016/j.earlhumdev.2010.01.004. [DOI] [PubMed] [Google Scholar]
- 29.Li N., Yan F., Wang N., et al. Distinct gut microbiota and metabolite profiles induced by different feeding methods in healthy Chinese infants. Front. Microbiol. 2020;11:714. doi: 10.3389/fmicb.2020.00714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Laursen M.F. Gut microbiota development: influence of diet from infancy to toddlerhood. Ann. Nutr. Metab. 2021:1–14. doi: 10.1159/000517912. [DOI] [PubMed] [Google Scholar]
- 31.Tabrez S., Shait Mohammed M.R., Jabir N.R., et al. Identification of novel cardiovascular disease associated metabolites using untargeted metabolomics. Biol. Chem. 2021;402(6):749–757. doi: 10.1515/hsz-2020-0331. [DOI] [PubMed] [Google Scholar]
- 32.Mu T.Y., Zhu Q.Y., Chen L.S., et al. Traditional Chinese Medicine constitution types of high-normal blood pressure: a meta-analysis. Heliyon. 2023;9(2) doi: 10.1016/j.heliyon.2023.e13438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Hayden H.S., Eng A., Pope C.E., et al. Fecal dysbiosis in infants with cystic fibrosis is associated with early linear growth failure. Nat. Med. 2020;26(2):215–221. doi: 10.1038/s41591-019-0714-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Sequeira R.P., McDonald J.A.K., Marchesi J.R., et al. Commensal Bacteroidetes protect against Klebsiella pneumoniae colonization and transmission through IL-36 signalling. Nat. Microbiol. 2020;5(2):304–313. doi: 10.1038/s41564-019-0640-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Jia W., Rajani C., Xu H., et al. Gut microbiota alterations are distinct for primary colorectal cancer and hepatocellular carcinoma. Protein Cell. 2021;12(5):374–393. doi: 10.1007/s13238-020-00748-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Shin J., Li T., Zhu L., et al. Obese individuals with and without phlegm-dampness constitution show different gut microbial composition associated with risk of metabolic disorders. Front. Cell. Infect. Microbiol. 2022;12 doi: 10.3389/fcimb.2022.859708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Van den Munckhof I.C.L., Kurilshikov A., Ter Horst R., et al. Role of gut microbiota in chronic low-grade inflammation as potential driver for atherosclerotic cardiovascular disease: a systematic review of human studies. Obes. Rev. 2018;19(12):1719–1734. doi: 10.1111/obr.12750. [DOI] [PubMed] [Google Scholar]
- 38.Zhao X., Chen Y., Li L., et al. Effect of DLT-SML on chronic stable angina through ameliorating inflammation, correcting dyslipidemia, and regulating gut microbiota. J. Cardiovasc. Pharmacol. 2021;77(4):458–469. doi: 10.1097/FJC.0000000000000970. [DOI] [PubMed] [Google Scholar]
- 39.Bui T.P.N., Mannerås-Holm L., Puschmann R., et al. Conversion of dietary inositol into propionate and acetate by commensal Anaerostipes associates with host health. Nat. Commun. 2021;12(1):4798. doi: 10.1038/s41467-021-25081-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Ioannou G.N. The role of cholesterol in the pathogenesis of NASH. Trends Endocrinol. Metabol. 2016;27(2):84–95. doi: 10.1016/j.tem.2015.11.008. [DOI] [PubMed] [Google Scholar]
- 41.Sun L., Jia H., Li J., et al. Cecal gut microbiota and metabolites might contribute to the severity of acute myocardial ischemia by impacting the intestinal permeability, oxidative stress, and energy metabolism. Front. Microbiol. 2019;10:1745. doi: 10.3389/fmicb.2019.01745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Tian R., Liu H., Feng S., et al. Gut microbiota dysbiosis in stable coronary artery disease combined with type 2 diabetes mellitus influences cardiovascular prognosis. Nutr. Metabol. Cardiovasc. Dis. 2021;31(5):1454–1466. doi: 10.1016/j.numecd.2021.01.007. [DOI] [PubMed] [Google Scholar]
- 43.Fan H.Y., Tung Y.T., Yang Y.S.H., et al. Maternal vegetable and fruit consumption during pregnancy and its effects on infant gut microbiome. Nutrients. 2021;13(5):1559. doi: 10.3390/nu13051559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Yuan C., Jin X., He Y., et al. Association of dietary patterns with gut microbiota in kidney stone and non-kidney stone individuals. Urolithiasis. 2022;50(4):389–399. doi: 10.1007/s00240-022-01325-2. [DOI] [PubMed] [Google Scholar]
- 45.Png C.W., Lindén S.K., Gilshenan K.S., et al. Mucolytic bacteria with increased prevalence in IBD mucosa augment in vitro utilization of mucin by other bacteria. Am. J. Gastroenterol. 2010;105(11):2420–2428. doi: 10.1038/ajg.2010.281. [DOI] [PubMed] [Google Scholar]
- 46.Kaczmarczyk M., Löber U., Adamek K., et al. The gut microbiota is associated with the small intestinal paracellular permeability and the development of the immune system in healthy children during the first two years of life. J. Transl. Med. 2021;19(1):177. doi: 10.1186/s12967-021-02839-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Ganesh B.P., Klopfleisch R., Loh G., et al. Commensal Akkermansia muciniphila exacerbates gut inflammation in Salmonella Typhimurium-infected gnotobiotic mice. PLoS One. 2013;8(9) doi: 10.1371/journal.pone.0074963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Zhou L., Xiao X., Zhang Q., et al. Improved glucose and lipid metabolism in the early life of female offspring by maternal dietary genistein is associated with alterations in the gut microbiota. Front. Endocrinol. 2018;9:516. doi: 10.3389/fendo.2018.00516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Jo J.K., Seo S.H., Park S.E., et al. Gut microbiome and metabolome profiles associated with high-fat diet in mice. Metabolites. 2021;11(8):482. doi: 10.3390/metabo11080482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Liao R., Xie X., Lv Y., et al. Ages of weaning influence the gut microbiota diversity and function in Chongming white goats. Appl. Microbiol. Biotechnol. 2021;105(9):3649–3658. doi: 10.1007/s00253-021-11301-2. [DOI] [PubMed] [Google Scholar]
- 51.Nogal A., Louca P., Zhang X., et al. Circulating levels of the short-chain fatty acid acetate mediate the effect of the gut microbiome on visceral fat. Front. Microbiol. 2021;12 doi: 10.3389/fmicb.2021.711359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Lopez-Siles M., Duncan S.H., Garcia-Gil L.J., et al. Faecalibacterium prausnitzii: from microbiology to diagnostics and prognostics. ISME J. 2017;11(4):841–852. doi: 10.1038/ismej.2016.176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Zhang M., Liang Y., Liu Y., et al. High-fat diet-induced intestinal dysbiosis is associated with the exacerbation of Sjogren's syndrome. Front. Microbiol. 2022;13 doi: 10.3389/fmicb.2022.916089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Moon J., Yoon C.H., Choi S.H., et al. Can gut microbiota affect dry eye syndrome? Int. J. Mol. Sci. 2020;21(22):8443. doi: 10.3390/ijms21228443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Chen Y.C., Lin H.Y., Chien Y., et al. Altered gut microbiota correlates with behavioral problems but not gastrointestinal symptoms in individuals with autism. Brain Behav. Immun. 2022;106:161–178. doi: 10.1016/j.bbi.2022.08.015. [DOI] [PubMed] [Google Scholar]
- 56.Jiang H., Ling Z., Zhang Y., et al. Altered fecal microbiota composition in patients with major depressive disorder. Brain Behav. Immun. 2015;48:186–194. doi: 10.1016/j.bbi.2015.03.016. [DOI] [PubMed] [Google Scholar]
- 57.Obeid R., Geisel J., Nix W.A. 4-Pyridoxic acid/pyridoxine ratio in patients with type 2 diabetes is related to global cardiovascular risk scores. Diagnostics. 2019;9(1):28. doi: 10.3390/diagnostics9010028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Barrett-Connor E., Khaw K.T., Yen S.S. A prospective study of dehydroepiandrosterone sulfate, mortality, and cardiovascular disease. N. Engl. J. Med. 1986;315(24):1519–1524. doi: 10.1056/NEJM198612113152405. [DOI] [PubMed] [Google Scholar]
- 59.Mitchell L.E., Sprecher D.L., Borecki I.B., et al. Evidence for an association between dehydroepiandrosterone sulfate and nonfatal, premature myocardial infarction in males. Circulation. 1994;89(1):89–93. doi: 10.1161/01.cir.89.1.89. [DOI] [PubMed] [Google Scholar]
- 60.Ruiz-Canela M., Toledo E., Clish C.B., et al. Plasma branched-chain amino acids and incident cardiovascular disease in the PREDIMED trial. Clin. Chem. 2016;62(4):582–592. doi: 10.1373/clinchem.2015.251710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Shah S.H., Bain J.R., Muehlbauer M.J., et al. Association of a peripheral blood metabolic profile with coronary artery disease and risk of subsequent cardiovascular events. Circ. Cardiovas. Genet. 2010;3(2):207–214. doi: 10.1161/CIRCGENETICS.109.852814. [DOI] [PubMed] [Google Scholar]
- 62.Gilstrap L.G., Wang T.J. Biomarkers and cardiovascular risk assessment for primary prevention: an update. Clin. Chem. 2012;58(1):72–82. doi: 10.1373/clinchem.2011.165712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Nie C., He T., Zhang W., et al. Branched chain amino acids: beyond nutrition metabolism. Int. J. Mol. Sci. 2018;19(4):954. doi: 10.3390/ijms19040954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Savineau J.P., Marthan R., Dumas de la Roque E. Role of DHEA in cardiovascular diseases. Biochem. Pharmacol. 2013;85(6):718–726. doi: 10.1016/j.bcp.2012.12.004. [DOI] [PubMed] [Google Scholar]
- 65.Mangge H., Zelzer S., Prüller F., et al. Branched-chain amino acids are associated with cardiometabolic risk profiles found already in lean, overweight and obese young. J. Nutr. Biochem. 2016;32:123–127. doi: 10.1016/j.jnutbio.2016.02.007. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Data will be made available on request.