Summary
Elucidating a basic blueprint of osteoclast-osteoblast coordination in skeletal remodeling and understanding how this coordination breaks down with age and disease is essential for addressing the growing skeletal health problem in our aging population. The paucity of simple, activatable, biologically relevant models of osteoclast-osteoblast coordination has hindered our understanding of how skeletal remolding is regulated. Here, we describe an inducible ex vivo model of osteoclast-osteoblast progenitor coordination. Induction activates the release of osteoclastogenic factors from osteoprogenitors, which elicits the differentiation and fusion of neighboring preosteoclasts. In turn, multinucleated osteoclasts release soluble coupling factors, RANK+ extracellular vesicles and promote osteoprogenitor proliferation, recapitulating aspects of perturbed coordination in diseases underpinned by excessive osteoclast formation. We expect this model to expedite the investigation of cell-cell fusion, osteoclast-osteoblast progenitor coordination, and extracellular vesicle signaling during bone remodeling and offer a powerful tool for evaluating signaling cascades and novel therapeutic interventions in osteoclast-linked skeletal disease.
Subject area(s): Biological sciences, Cell biology, Stem cells research
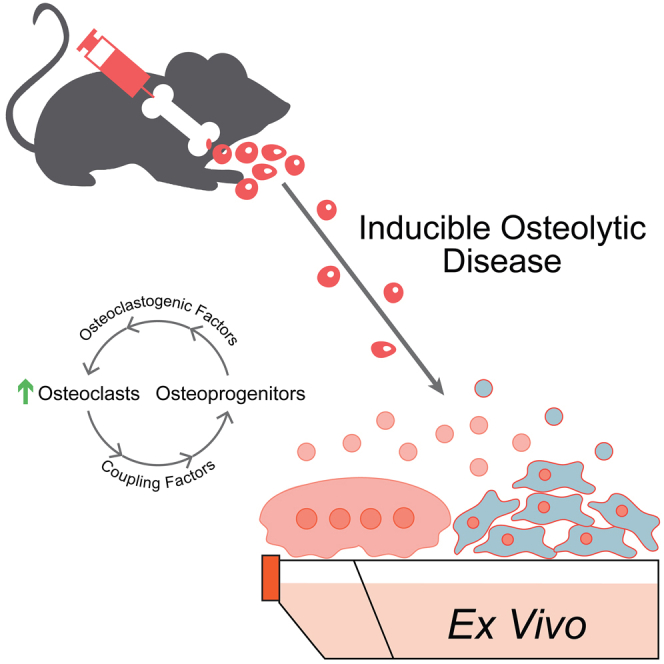
Graphical abstract
Highlights
-
•
Rapid, inducible model of osteoclast formation and function in osteolytic disease
-
•
Ex vivo model recapitulates osteoclast-osteoblast precursor coordination
-
•
A highly adaptable tool for characterizing perturbations in fibrous dysplasia
-
•
Excessive osteoclastogenesis promotes osteoprogenitor proliferation
Biological sciences; Cell biology; Stem cells research
Introduction
Metabolic bone diseases are a collection of disorders resulting in diminished skeletal integrity and changes in bone mass because of perturbations in the life-long process of bone remodeling.1 Throughout life, the skeleton relies on a basic multicellular unit (BMU) that forms a bone remodeling niche that proceeds along the skeletal matrix and continually remodels bone.2 This BMU is primarily composed of a bipartite remodeling system (i.e., osteoclasts and osteoblasts) and two regulatory cell types that come from the osteoblast lineage and work to orchestrate remodeling through signaling cascades (i.e., bone lining cells and osteocytes). The remodeling system is primarily led by osteoclasts, which erode the inorganic, mineral phase of old/damaged bone and degrade the organic, collagen-rich phase through the release of proteases. In turn, osteoblasts secrete osteoid and mineralize it to form new bone to replace what was removed. In this bipartite remodeling machine, osteoblast lineage cells release a variety of signals that regulate the formation of osteoclasts.3,4 In turn, recent evidence demonstrates that osteoclasts produce a variety of soluble coupling factors (e.g., leukemia inhibitor factor (LIF)) and package and release microRNA rich extracellular vesicles, both of which regulate the differentiation/proliferation of osteoblast precursors (osteoprogenitors).5,6,7,8,9 Together, the BMU continually erodes old or damaged bone and deposits new to renew the skeletal system throughout life.10 Here we offer a rapid model system for resolving how key remodeling members of the BMU – osteoprogenitors and preosteoclasts - coordinate their activities and malfunction with age and disease.
Osteoblast lineage cells activate the formation of multinucleated osteoclasts from mononucleated osteoclast precursors (preosteoclasts), of the myeloid lineage, through the release of osteoclastogenic cytokines (e.g., macrophage colony stimulating factor (M-CSF) and receptor activator of NF-κB ligand (RANKL)). First, M-CSF promotes the differentiation of monocytic lineage cells into preosteoclasts. Next, RANKL binding to the preosteoclast membrane receptor RANK is both necessary and sufficient to upregulate the transcription factors Nfatc1 and cFos and commit precursors to an osteoclast fate.11 Committed osteoclasts then migrate, adhere to one another and fuse plasma membranes (PMs) producing multinucleated, syncytial osteoclasts that typically reach 5–12 nuclei/cell in humans,12,13 although osteoclasts sizes can differ considerably between individuals and increase with age.14,15 Osteoclast number and size are tightly regulated during bone remodeling, and the number of fusion events forming a multinucleated osteoclast (i.e., its nuclear multiplicity) directly correlates with its ability to resorb bone (i.e., higher nuclear multiplicity = higher bone resorption).15,16,17,18 In addition to eliciting osteoclast formation, osteoblast lineage cells also negatively regulate the number and size of osteoclasts by releasing the RANKL decoy receptor osteoprotegerin (OPG).19 Thus, tight osteoblast-osteoclast coordination within the bone remodeling compartment manages the number, size, differentiation state and biological activities of this bipartite bone remodeling machine.
Perturbations in the number, size and nuclear multiplicity of osteoclasts underpin the development of diverse metabolic bone diseases that impact >13% of adults over age 50 world-wide (>20% of women).20 Each metabolic bone disease (e.g., osteoporosis, Paget’s disease, fibrous dysplasia (FD), osteopetrosis) presents with unique phenotypes, rises from distinct etiologies and progress with disparate severities, but all are underpinned by a breakdown in osteoclast formation/function.21,22,23,24 These perturbations of osteoclast formation/function either stem from or cause dysfunctional osteoclast-osteoblast coordination. To date, we still lack a detailed understanding of how osteoclasts and osteoblasts coordinate their functions in skeletal biology.
Elucidating a basic blueprint of osteoclast-osteoblast coordination in healthy skeletal remodeling and characterizing how the systems that coordinate this bipartite machine breakdown with age and disease is an essential first step in addressing the growing skeletal health problem in our aging population. Unfortunately, many common models of osteoclast or osteoblast function do not recapitulate the bipartite nature of the bone remodeling machine, but rather recapitulate the isolated function of a single cell type. Immortalized preosteoclast and osteoprogenitor cell lines (i.e., RAW 264.7 and MC3T3-E1 subclones, respectively) are convenient workhorses that suffer from well documented limitations in recapitulating skeletal biology.25,26,27 Primary osteoclast formation is modeled in vitro via enriching primary monocytes that are differentiated stepwise into preosteoclasts and then fusogenic osteoclasts via activation by recombinant M-CSF and RANKL.28 Unfortunately, these approaches suffer from the difficulty of obtaining fresh, primary monocytes, the inconsistent propensity of monocytes from different individuals to differentiate and fuse in vitro, the significant expense of recombinant M-CSF and RANKL and the inability to passage these cells.15,29 Obtaining primary osteoprogenitors with consistent, reliable behavior is even more challenging by comparison, requiring the investigator to obtain biopsies from donors or undergo challenging protocols for isolating, enriching and culturing primary cells from murine sources.30,31 In contrast, animal models do recapitulate the bipartite nature of the BMU and offer in vivo context for the underlying biology being evaluated but introduce significantly longer timelines for experiments; often require specialized equipment for sectioning, demineralizing and imaging cells on bone; introduce greater challenges for collecting high resolution image data; and generally introduce greater complexity and diminished resolution by the nature of osteoclasts and osteoblasts being minor components of the population of cell types that occupy bone.
Here we have developed an inducible explant culture model that recapitulates osteoclast-osteoblast progenitor signaling and coordination in the context of exacerbated osteoclastogenesis, a phenotype common to many skeletal diseases. Fibrous dysplasia (FD) is a metabolic bone disease caused by somatic, mosaic gain-of-function mutations in GNAS within osteoblast lineage cells, leading to aberrant cell proliferation, suppressed osteoprogenitor differentiation/commitment and the excessive release of osteoclastogenic factors, including RANKL and IL-6.32,33,34 We utilized an inducible murine model of FD to develop a tractable ex vivo culture model of osteoprogenitor-to-preosteoclast signaling and osteoclastogenesis. Whole bone marrow isolates from the hind limbs of this FD mouse produce a complex cell culture system enriched in osteoprogenitors and preosteoclasts. Doxycycline (Dox.) addition to these complex cultures activates GαsR201C expression and RANKL and IL-6 release from osteoprogenitors, which elicits the differentiation and fusion of neighboring preosteoclasts. In turn, resulting multinucleated osteoclasts release soluble coupling factors and RANK+ extracellular vesicles – recently recognized mediators of osteoclast-to-osteoprotenitor signaling – that correlate with changes in the proliferation and differentiation of osteoprogenitors. Moreover, these dynamic biological events all occur within 4 days of Dox. induction.
This relatively simple ex vivo model offers an exaggerated environment to study the complex osteoclast-osteoprogenitor coordination employed in the maintenance of skeletal integrity throughout life. The inducible, rapid formation of multinucleated osteoclasts and their quick impact on the function of surrounding osteoprogenitors greatly facilitates the analysis of the osteoprogenitor-osteoclast signaling in the context of exacerbated osteoclastogenesis. Moreover, based on the reproducibility of these ex vivo cultures and their adaptability to a variety of assays, we expect that this new model will expedite the investigation of osteoclast-osteoblast progenitor coordination in bone remodeling and offer a powerful tool for evaluating signaling cascades and novel therapeutic interventions in osteoclast-linked skeletal disease.
Results
Inducible GαsR201C in osteoprogenitors faithfully elicits osteoclast formation in murine marrow explants
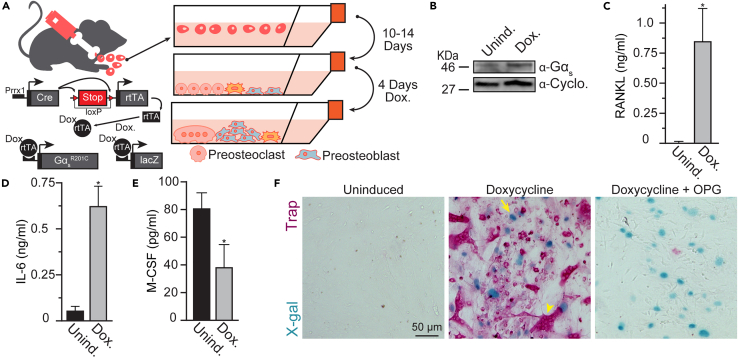
We isolated bone marrow from the tibia and femur of a previously described tet-on GαsR201C murine model crossed with a tet-on βgal reporter line to produce a quadruple transgenic model Prrx1-Cre/LSL-rtTA-IRES-GFP/TetO-GαsR201C/TetO-βgal. Expression of both GαsR201C and β-gal are inducible specifically in Prrx1, osteoprogenitors because of the conditional introduction of the reverse tetracycline transactivator (trTR) via Prrx1-Cre (Figure 1A).24,35 When introduced via Prrx1-Cre, rtTA can be specifically activated via Dox. addition in Prrx1 progeny (i.e., postnatal osteoprogenitors of the skeletal stem cell linage within the limbs) and induce the expression of both GαsR201C and β-gal via binding to their rtTA specific promotors (Figure 1A). Using these mice, the tibia and femur from freshly euthanized animals were dissected from both hindlimbs. The marrow space of the tibia and femur was opened by boring small holes in the epiphyses with hypodermic needles, and whole marrow was flushed out using serum free media and triturated until a homogeneous mixture of cells was obtained. These non-adherent isolates were maintained in complete explant media until the majority of cells began to adhere to the culture plastic (∼14 days), whereupon cultures were expanded and passaged for the downstream experiments described below (Figure 1A). Induction of marrow explant cultures via Dox. activates rtTA and elicits expression of GαsR201C and β-gal transgenes specifically in osteoprogenitors from the skeletal stem cell linage leading to an increase in the steady-state level of Gαs (Figure 1B). Dox. induction of GαsR201C led to the release of RANKL and IL-6 from osteoprogenitors and the rapid formation of multinucleated osteoclasts (Figure 1C, 1D, 1F). Moreover, marrow explants produced their own M-CSF before Dox. induction that likely facilitates the growth and survival of preosteoclasts in the cultures (Figure 1E). Intriguingly, M-CSF appears to be downregulated by Dox. induction, however, the mechanisms and physiological meaning of this downregulation remain to be clarified. β-gal expression was also observed via x-gal staining and provided a linage tracer demonstrating that rtTA activation and transgene expression occur only in osteoprogenitors and not in TRAP+ osteoclasts (Figure 1F). Moreover, the formation of TRAP+, multinucleated osteoclasts was dependent on the RANKL produced by Dox. induction of osteoprogenitors, as addition of OPG blocked both the production of TRAP and the observation of multinucleated osteoclasts in induced cultures but not the rtTA induction of β-gal (Figure 1F).
Figure 1.
Induction of GαsR201C in osteoprogenitors induces osteoclast formation in marrow cultures
(A) A cartoon schematic the genetic tools utilized and our approach for implementing them to model oteoprogenitor-osteoclast signaling/coordination in the formation of osteoclasts.
(B) A representative Western blot evaluating Gαs expression with (Dox.) or without (Unind.) 4 days induction.
(C) ELISA evaluation of soluble RANKL in the culture media of marrow explants with (Dox.) or without (Unind.) 4 days induction. (n = 7).
(D) ELISA evaluation of soluble IL-6 in the culture media of marrow explants with (Dox.) or without (Unind.) 4 days induction. (n = 4).
(E) ELISA evaluation of soluble M-CSF in the culture media of marrow explants with (Dox.) or without (Unind.) 4 days induction. (n = 5).
(F) Representative color brightfield images of marrow explants from GαsR201C following 4 days with (Dox.), without (Unind.) or with induction in the presence of OPG (Dox.+OPG). Arrow denotes x-gal staining of the nucleus of an induced osteoprogenitor, Arrowhead denotes a multinucleated, TRAP+ osteoclast. Error bars = ±SEM. Significance was assessed via paired t-test, where ∗p=<0.05, ∗∗p=<0.01, ∗∗∗p=<0.001.
Osteoprogenitor signaling elicits robust osteoclastogenesis in a Rankl-dependent manner
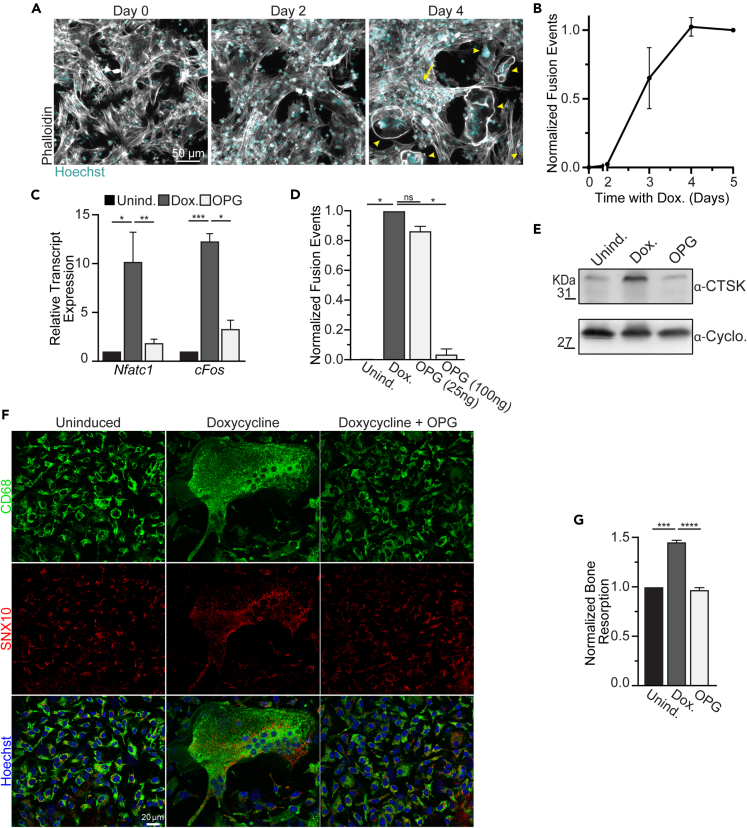
Tet-on GαsR201C marrow explant cultures offer a rapid, tractable model for analyzing osteoclast formation elicited by osteoprogenitors in an ex vivo environment, modeling the multicellular coordination between osteoclast and osteoblast progenitors in the BMU. Dox-induced osteoprogenitor-to-preosteoclast signaling within these cultures activated osteoclastogenesis in preosteoclasts that fuse and form syncytia with readily distinguishable actin borders in ∼4 days (Figure 2A). We observed negligible multinucleated osteoclast formation before 2 days of induction and a near linear increase in osteoclast fusion until 4 days of induction, when fusion plateaued (Figure 2B). RANKL production was responsible for the observed osteoclast formation from preosteoclasts, as Dox. induction activated the upregulation of the osteoclastogenic transcription factors Nfatc1 and cFos, both of which were blocked by the addition of OPG, the RANKL decoy receptor (Figure 2C). Upregulation of Nfatc1 and cFos coincided with the formation of TRAP+, multinucleated osteoclasts. Moreover, the expression of TRAP and extent of fusion between osteoclasts depended on RANKL production, as multinucleated cells were not observed in cultures without Dox. induction and OPG suppressed their formation in a concentration dependent manner (Figure 2D). Osteoclasts not only fused in response to Dox. induction, but the formation of multinucleated osteoclasts was associated with their production of the resorptive enzyme cathepsin k (Figure 2E). In addition to general cytoskeletal and chromogenic staining, we found that immunofluorescence imaging of the pan-macrophage lineage marker CD68 and the osteoclast enriched sorting nexin, Snx10, easily facilitated the detection of preosteoclasts and subsequent multinucleated osteoclasts among the cell types present in our complex cultures (Figure 2F). Finally, Dox. induction of marrow explant cultures not only elicited the formation of osteoclast syncytia, but also activated their resorptive activity in a RANKL dependent manner (Figure 2G).
Figure 2.
Dox. induction elicits osteoclast formation
(A) Representative images of ex vivo marrow cultures following Dox. induction (Gray = Phalloidin-Alexa488, Cyan = Hoechst).
(B) Quantification of the number of fusion events over time following addition of Dox. Each time course was normalized to the number of fusion events observed on day 5. (n = 3).
(C) qPCR evaluation of the osteoclastogenesis transcription factors Nfatc1 and cFos under uninduced (black), Dox. induced (gray) or Dox. induced +100 ng/mL OPG (white) conditions. Fold expression is relative to Actb RNA. (n = 3).
(D) Quantification of the number of fusion events in ex vivo marrow cultures treated as in (C). (n = 3).
(E) Representative Western blot evaluation of the steady-state level of the osteoclast resorption enzyme CTSK in ex vivo marrow cultures treated as in (C).
(F) Representative immunofluorescence images of the pan macrophage lineage marker CD68 (green), the osteoclast sorting nexin SNX10 and Hoechst in ex vivo marrow cultures treated as in (C).
(G) Quantification of bone resorption by ex vivo marrow cultures treated as in (C). In (A and D), Arrowheads = multinucleated osteoclasts, Arrows = proliferative preosteoblasts. Error bars = ±SEM. Statistical significance was assessed via paired t-test, where ∗p=<0.05, ∗∗p=<0.01, ∗∗∗p=<0.001.
Osteoclast formation promotes osteoprogenitor proliferation
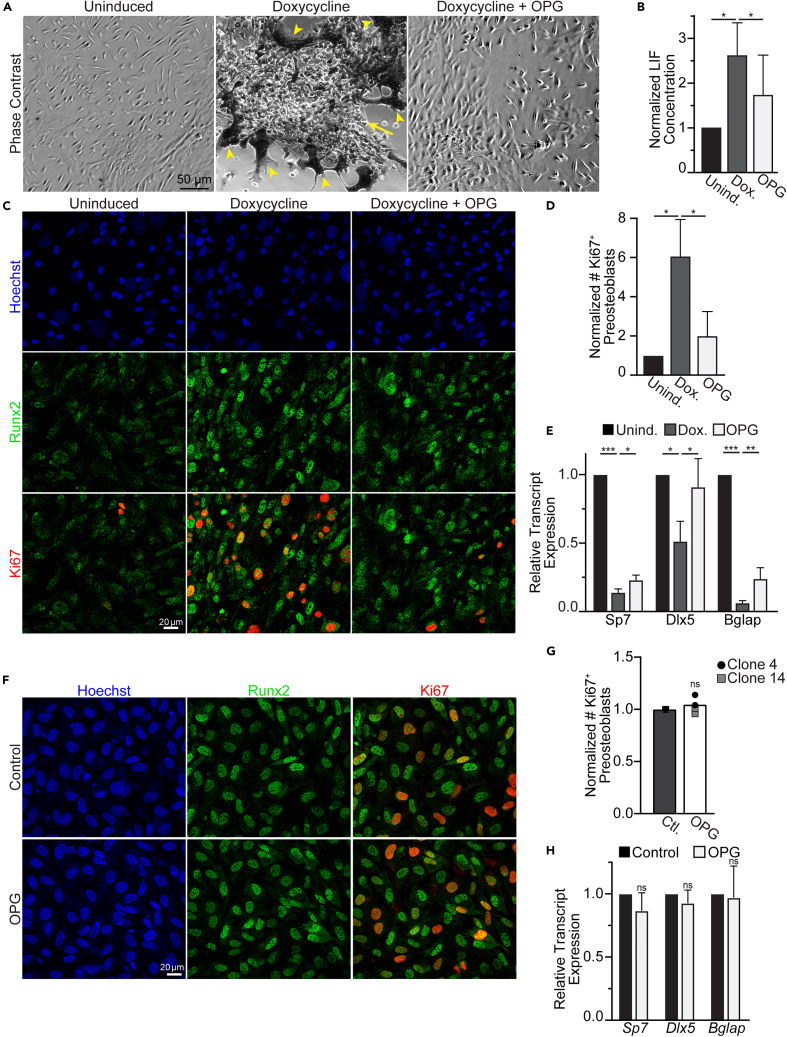
The formation of multinucleated osteoclasts was also accompanied by the development of fibrous, multicellular lesions, analogous to those observed in patients with gain-of-function mutations in Gαs (Figures 2A, 3A, arrows). The formation of multinucleated osteoclasts was also accompanied by an increase in the release of soluble LIF, a previously characterized osteoclast-to-osteoprogenitor coupling factor produced by osteoclasts (Figure 3B).8,9 Staining of these lesions demonstrated that the bulk of cells exhibited enriched, nuclear staining for the osteoprogenitor marker Runx2 (Figure 3C). Although preosteoblasts were readily observed in uninduced cultures, neither enrichment nor the clumping phenomenon was observed in the absence of Dox. induction (Figure 3A). Further underscoring the proliferative nature of these multicellular lesions following Dox. induction, the preosteoblasts demonstrated a dramatic nuclear enrichment of the proliferative marker Ki67 (Figures 3C and 3D). Moreover, in addition to the increased proliferation observed in preosteoblasts, we observed a marked decrease in the osteoblast differentiation markers Sp7, Dxl5 and Bglap36 in our cultures following Dox. induction, suggesting reduced osteogenic commitment accompanied this increase in preosteoblast proliferation (Figure 3E). As with the proliferative phenotype, both the production of LIF and the marked decrease in osteoblast differentiation observed were partially rescued by blocking the formation of osteoclasts via OPG, suggesting that osteoclasts contribute to the observed changes in LIF production and preosteoblast proliferation/differentiation (Figures 3B–3E).
Figure 3.
Osteoclast formation in ex vivo cultures elicits osteoprogenitor proliferation
(A) Representative phase contrast images of TRAP stained marrow explants following 4 days with (Dox.), without (Unind.) or with induction in the presence of OPG (Dox.+OPG). Arrow denotes lesion of cells, Arrowheads denote TRAP+, multinucleated osteoclasts, which appear dark in phase contrast.
(B) ELISA evaluation of soluble LIF in the culture media of marrow explants without (Unind.), with 4 days induction (Dox.) or 4 days induction and 100 ng/mL OPG (OPG). (n = 6 for OPG or n = 7 for Unind. and Dox.).
(C) Representative immunofluorescence images of the preosteoblast marker Runx2, the proliferation marker Ki67 and the nuclear stain Hoechst in cultures treated as in (A).
(D) Quantification of the fraction of Runx2+ preosteoblasts that exhibit staining for the proliferation marker Ki67. (n = 3).
(E) qPCR evaluation of the osteogenic markers Sp7, Dlx5 and Bglap under uninduced (black), Dox. induced (gray) or Dox. induced +100 ng/mL OPG (white) conditions. Fold expression is relative to 18s rRNA. (n = 5).
(F) Representative immunofluorescence images of the preosteoblast marker Runx2, the proliferation marker Ki67 and the nuclear stain Hoechst in MC3T3-E1 clone 14 preosteoblasts treated with (OPG) or without (control) 100 ng/mL OPG.
(G) Quantification of the fraction of Runx2+ preosteoblasts that exhibit staining for the proliferation marker Ki67 treated as in (F). (n = 3 for clone 4(circle), n = 3 for clone 14(square). Bar denotes summary average of both clones 4 and 14).
(H) qPCR evaluation of the osteogenic markers Sp7, Dlx5 and Bglap in clones 4 and 14 under control (black), 100 ng/mL OPG (white) conditions. Fold expression is relative to 18s rRNA. (n = 3) Error bars = ±SEM. Significance was assessed via paired t-test, where ∗p=<0.05, ∗∗p=<0.01, ∗∗∗p=<0.001.
In addition to releasing soluble RANKL, preosteoblasts express a transmembrane RANKL variant that has been recently implicated in osteoblast differentiation.37 To test whether changes in preosteoblast proliferation/differentiation elicited by OPG may be caused by some direct effect OPG has on preosteoblasts, we treated two preosteoblast cell lines (MC3T3-E1 clone 4 and clone 14) with OPG and looked for changes in Ki67 staining. In both clones, we observed no change in the expression of Runx2 or Ki67 staining, suggesting that the effect of OPG addition on the proliferation of preosteoblasts in our explant cultures is not direct (Figures 3F and 3G). Likewise, the addition of OPG to preosteoblasts alone had no impact on the steady-state levels of the osteoblast differentiation marks observed in our ex vivo cultures (Figures 3H and 3E). Together, these data suggest that osteoclasts themselves contribute to the changes in osteoprogenitor proliferation/differentiation observed in our explant model following Dox. induction, and that these observations are not a direct effect of OPG on preosteoblasts.
Osteoprogenitor proliferation correlates with the release of Rank+ extracellular vesicles (EVs)
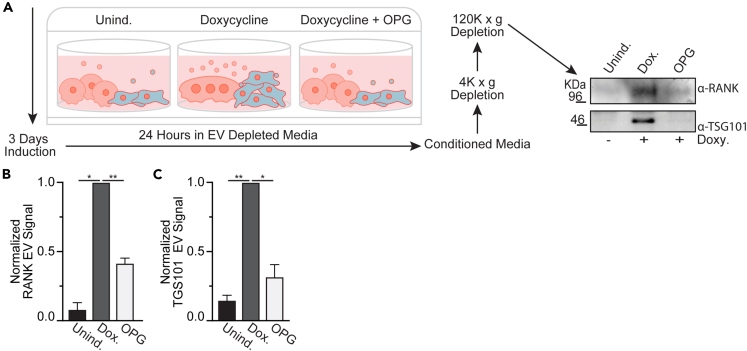
Recent reports have highlighted the role of EVs as mediators of osteoclast-to-osteoprogenitor signaling and regulation in coordinated bone remodeling.5,36,38,39 Previous characterizations of osteoclast EVs have identified both RANK and TGS101 as markers of osteoclast derived EVs that impact osteoclast and osteoblast differentiation.5,6,7,40 To assess whether our system models the osteoclast production of EVs that coordinate osteoclast-osteoblast activity, we collected conditioned media from our cultures and evaluated whether they contained EVs released by osteoclasts. We induced our cultures and waited until we observed multinucleated osteoclasts (3–4 days). Next, we placed our cultures in complete explant media made with EV depleted FBS, allowed cultures to condition the media for 24 h, collected the media and enriched an EV fraction using a simple centrifugation approach (Figure 4A). Biochemical assessment of these EV-enriched fractions demonstrated that Dox. induction led to an enrichment of the osteoclast transmembrane receptor RANK and the exosome marker TSG101 in the EV fraction (Figure 4A). The enrichment of RANK and TSG101 in the EV fraction produced by explant cultures was not observed without Dox. induction and was suppressed by OPG inhibition of osteoclast formation (Figures 4A–4C). Identical results were also obtained using commercially available EV isolation kits used according to manufactures instructions (QIAGEN ExoEasy). These findings indicated that osteoprogenitor-dependent osteoclast formation in our model is accompanied by the release of osteoclast-generated, RANK+ EVs and correlates with subsequent changes in osteoprogenitor proliferation/differentiation.
Figure 4.
Osteoclast formation in ex vivo marrow cultures is accompanied by EV release
(A) An illustrated schematic depicting an approach for enriching an EV fraction from uninduced, Doxy. induced and Doxy. induced +100 ng/mL OPG ex vivo marrow cultures (left). A representative Western blot evaluating EV enriched fractions for the osteoclast membrane receptor RANK and the exosome marker TSG101 (right).
(B) Quantification of RANK signal in representative Western blots of EV enriched fractions, as in (A). (n = 3).
(C) Quantification of TSG101 signal in representative Western blots of EV enriched fractions, as in (A). (n = 5) Error bars = ±SEM. Significance was assessed via paired t-test, where ∗p=<0.05, ∗∗p=<0.01, ∗∗∗p=<0.001.
In summary, we have established a simple, inducible, tractable tool for the study of osteoprogenitor induced osteoclast formation that can be implemented to study how osteoclasts and osteoblast lineage cells coordinate their functions in health and disease.
Discussion
Here, we describe a primary culture model for recapitulating the dynamic interactions and regulatory relationships that coordinate osteoclast and osteoblast progenitors in skeletal diseases underpinned by excessive osteoclastogenesis. Although the cellular make-up of the explant cultures described and the biological activities represented are complex, employing this model is relatively simple. The training required to master the dissections required is minimal, we observe high phenotypic reproducibility between cultures and for ≥4 passages within individual cultures and the cost in maintaining and implementing this model of osteoclast/osteoprogenitor biology is modest, as the only media additive required for activation is Dox.
Our primary aim was to develop an inducible model of osteoclast formation in response to osteogenic signaling rather than the supraphysiological addition of recombinant factors. Here, we have developed a culture model where osteoprogenitors are specifically induced to overexpress a gain-of-function mutation that elevates cAMP and promotes the release of endogenous osteoclastogenic factors (e.g., RANKL, IL-6, likely others). These factors activate osteoclastogenesis, promote osteoclast fusion, elicit osteoclast production of resorptive enzymes and lead to osteoclast-dependent demineralization in a short timeline (4 days following Dox. addition). Moreover, these cultures easily adapt to larger culture ware for biochemical sample collection (e.g., T-75 flasks, multi-well plates for screening assays, polymer bottom μ-slides for high-resolution imaging and mineralized biomimetics of bone) with no gross impact on their ability to faithfully produced osteoprogenitor-dependent multinucleated osteoclasts.
In addition to osteoclasts, we find that osteoblast lineage cells are well represented in the cultures. Skeletal stem cell progeny of the Prxx1 lineage conditionally express rtTR, facilitating inducible expression of the tet-on GαsR201C cassette responsible for RANKL production, as documented in the murine model from which our cultures are derived.24 Moreover, our addition of a tet-on β-gal cassette to this model allows for simple linage tracing and the detection of induced cells. GαsR201C is a common gain-of-function mutation in osteoprogenitors that leads to the excessive production of RANKL and the ectopic osteoclast formation that underpins the pathophysiology of FD. By taking advantage of this FD model, we have developed a tool that facilitates the ability to follow osteoprogenitor-to-osteoclast signaling with high spatial and temporal resolution. It is our hope that future work will further develop our model as a tool for evaluating the propensity of the osteoblast lineage cells in our cultures to secrete matrix and mineralize it. Although we have focused on cells of the osteoprogenitor and preosteoclast lineage, we cannot dismiss the possible presence or contribution of other cell types to the ability to passage, grow and faithfully induce cultures to elicit the osteoprogenitor-dependent formation of multinucleated osteoclasts. However, after the initial passage of the cell types that adhere to culture plastic, we observe that the vast majority of cells within our cultures exhibit morphology consistent with the CD68+ macrophages or Runx2+ osteoblast precursors we typically stain in our experiments, and that nearly all cells stain positively for either β-gal or TRAP activity following Dox. induction.
In addition to observing changes in osteoprogenitors and osteoclasts, our model offers an excellent opportunity to study the complex cell-cell signaling that coordinates the functions of these cells during bone remodeling. Well established intercellular signaling cascades tightly regulate the formation and activity of the multicellular unit (e.g., RANKL, TGF-β, BMPs), but growing evidence suggests that a complex array of cytokines and secondary messengers contribute to the formation of osteoblasts, osteoclasts and coordinate their activities. Moreover, new evidence demonstrates a role for both soluble factors and EVs in coordinating osteoclast-osteoblast function and suggests that these messengers act as a feedback mechanism for managing biological activity in bone remodeling.5,39,40 Recent work has also demonstrated dramatic changes in EV signaling in bone disease and injury, highlighting the clinical need for reliable systems with which to identify the signals released by osteoclasts/osteoblasts during bone remolding and to characterize how therapeutic interventions modulate the release and makeup of these critical secondary messengers.38,41,42 Here, we demonstrate that our ex vivo cultures are rich sources of both soluble factors like LIF and EV secondary messengers exchanged by osteoclast and osteoblast progenitors. We expect that these cultures will be an excellent tool for studying the contents, biological activity, signaling properties and downstream effects of these and other secondary messengers in the process of coordinated bone remodeling. For example, Dox. induction of our cultures is accompanied by a dramatic increase in the release of soluble IL-6, an inflammatory cytokine associated with the progression of FD.33,43 IL-6 has a paradoxical role in osteoclastogenesis and has been described to directly stimulate osteoclast formation, indirectly elicit osteoclast formation, directly suppress osteoclast formation and both directly stimulate and directly suppress the formation of osteoclasts depending on the concentration of RANKL available.44,45,46,47 The confusing information concerning the role of IL-6 in osteoclast formation recently culminated in a failed clinical trial assessing the efficacy of an α-IL-6 monoclonal antibody for the treatment of FD.48 Our current work limits our ability to comment on the mechanistic role of IL-6 in our system. However, we can say that our cultures recapitulate the production of excessive IL-6 previously associated skeletal disease underpinned by excessive osteoclast formation. We expect that our model represents an excellent opportunity for further resolving the role of IL-6 and other cytokines with complex roles in the coordination of osteoclast and osteoblast precursors.
Unfortunately, many current models of osteoclast/osteoblast function only replicate one constituent of the BMU that remodels bone or a complete remodeling compartment in vivo. These approaches either suffer from a lack of biological context or come with a level of complexity that makes the high-resolution evaluation of a relatively minor component of bone challenging. Here we describe a conditional, inducible, ex vivo approach that facilitates the rapid assessment of osteoprogenitor-preosteoclast coordination, the formation of osteoclasts and osteoclast-to-osteoprogenitor signaling in a fast, highly adaptive, cost-effective bone disease model (Figure 5).
Figure 5.
Ex vivo FD marrow explants as a model of osteoclast formation and osteoclast-osteoprogenitor coordination
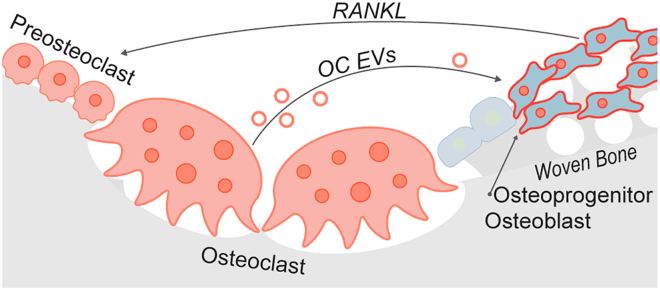
An illustration of the biological process modeled in our ex vivo marrow cultures. Doxycycline induction elicits osteoprogenitor expression of GαsR201C and release of RANKL. RANKL binding initiates osteoclastogenesis and the formation of multinucleated, bone resorbing osteoclasts. Osteoclasts release EVs that correlate with preosteoblast proliferation. Peach cells are of the monocyte>osteoclast lineage. Blue cells are of the skeletal stem cell>osteoblast lineage.
We expect our model to complement single cell, co-culture and in vivo models in an effort to better understand the signaling, coordination and behavior of osteoclasts/osteoprogenitors in diseases of exacerbated osteoclastogenesis. For example, there exist several, well-established osteoblast-osteoclast co-culture systems that our field has utilized for decades to lay the foundation of our understanding of osteoclast-osteoblast coordination.49,50,51 We believe that our model complements this gold standard in a number of ways (e.g., our system is inducible, facilitating synchronicity, discrete timelines, simple before and after conditions; our system is highly proliferative, passageable and scalable for large biochemical experiments (ideal for secondary messenger evaluation); our system allows an investigator to introduce genetic material, wait for marker expression (e.g., eGFP) and then induce osteoclastogenesis and coordination; and our system models osteoclast and osteoblast lineage cells at a variety of different states of differentiation, similar to an in vivo scenario). However, our model has limitations (e.g., it relies on an overexpressed gain of function mutation, it is much less characterized), and it will never replace classic co-culture systems. We ourselves utilize single cell, co-culture, in vivo and our ex vivo model simultaneously. We find that the ex vivo model has become an incredible, complementary tool for us, and we hope others find even better ways to implement it than we ever would by ourselves.
Although our system is built from genetic tools used to model FD, we would like to emphasize that its applications far surpass the study of this single disease. In this model, we have a multicellular composition where osteoclast formation, activity and signaling is directly inducible under the guidance of an osteoprogenitor conductor. In addition, the behavior of this osteoprogenitor conductor is subsequently influenced by the osteoclasts it elicits. Although we cannot exclude additional effects of GαsR201C induction in osteoprogenitors (e.g., basal proliferative, metabolic changes), we can say that OPG blockade of osteoclastogenesis significantly inhibits osteoprogenitor proliferation and partially rescues the expression of osteoprogenitor differentiation markers.
In addition to its obvious applications in studying skeletal diseases underpinned by excessive osteoclast formation, we believe our culture model will be useful for identifying biological pathways, molecular players and dynamic mechanisms underpinning the basic formation and coordination of osteoclasts and osteoprogenitors in healthy skeletal physiology. This acute model of excessive osteoclast formation can help us better resolve how osteoclast-osteoblast coordination is accomplished. As evidence of this model’s dual potential, we have utilized it in two recent manuscripts to identify a novel regulator of osteoclast fusion and assess the molecular mechanism of Denosumab treatment of FD.52,53 It is now our hope to disseminate this tool and partner with our colleagues to implement its use in answering new, vital questions concerning how bone remodeling is managed in health and disease.
Limitations of the study
As noted above, our ex vivo model of osteoclast-osteoblast progenitor coupling in exacerbated osteoclastogenesis exhibits a number of advantages that have led us to consider it a premier tool for the study of osteoclast/osteoprogenitor disease biology. However, we openly recognize that our model has a number of limitations that highlight the importance of implementing additional supportive models (e.g., individual cell lines, in vivo models, human equivalents, etc.). First, the osteoclastogenic induction our model utilizes relies on the overexpression of a gain of function mutation in Gαs, specifically in osteoprogenitors, that is known to elevate cAMP levels.24 Inducible expression of GαsR201C has the advantageous benefit of activating osteoclastogenic singling in a selective manner, however, we cannot exclude additional off target effects of elevated cAMP levels in the osteoprogenitors in our cultures. Second, based on our best assessments of immunofluorescent imaging with osteoprogenitor and osteoclast/osteoclast precursor markers and morphological assessment, we believe that the vast majority of cells in our cultures are of the osteoprogenitor or osteoclast linage; however, we cannot exclude the contribution of other minor cell types in our cultures. Finally, although our model recapitulates many biologically relevant aspects of the osteoclast-osteoblast progenitor signaling that underpins diseases of excessive osteoclastogenesis, our cultures lose additional signaling inputs from other cell types, the spatial organization of a native BMU, physical attributes of native substrates, and many other biologically relevant characteristics inherent to in vivo bone remodeling.
Our model is one useful tool in the expanding toolbox available to the skeletal biologist. When one addresses any of a number of simple home projects, do they not make use of a screwdriver, a hammer, a wrench when required? Each plays its part. How foolish would we be to suggest that the mystery of the osteoclast or of the osteoprogenitor or the majesty of their hidden conversations could be unraveled by a simple screwdriver? For that we will need the whole toolbox. For that we will need each other.
STAR★Methods
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| Rabbit polyclonal anti-snx10 antibody | Sigma | Cat#sab2107086 |
| Mouse monoclonal anti-tgs101 antibody | Novus | Cat#nb200-112 |
| Rat monoclonal anti-CD68 antibody | Abcam | Cat#ab53444 |
| Mouse monoclonal anti-runx2 antibody | Abcam | Cat#ab76956 |
| Rabbit monoclonal anti-ki67 antibody | Cell signaling technologies | Cat#ab76956 |
| Mouse monoclonal anti-rank antibody | Abcam | Cat#ab13918 |
| Rabbit polyclonal anti-ctsk antibody | Invitrogen | Cat#pa5-102483 |
| Rabbit polyclonal anti-gnas antibody | Millipore | Cat#371732 |
| Rabbit monoclonal anti-cyclophilin b antibody | Cell signaling technologies | Cat#43603s |
| Goat polyclonal anti-mouse secondary antibody | Cell signaling technologies | Cat#4408 |
| Goat polyclonal anti-mouse secondary antibody | Cell signaling technologies | Cat#4409 |
| Goat polyclonal anti-rabbit secondary antibody | Cell signaling technologies | Cat#4412 |
| Goat polyclonal anti-rabbit secondary antibody | Cell signaling technologies | Cat#4413 |
| Goat polyclonal anti-rat secondary antibody | Invitrogen | Cat#a-1106 |
| Goat polyclonal anti-mouse secondary antibody | Abcam | Cat#7069 |
| Goat polyclonal anti-rabbit secondary antibody | Abcam | Cat#7091 |
| Biological samples | ||
| Murine bone marrow isolates | Collins laboratory | N/A |
| Chemicals, peptides, and recombinant proteins | ||
| Doxycycline hyclate | Sigma | Cat#d9891 |
| cOmplete protease inhibitor | Sigma | Cat#118361700010 |
| Normocin | Invivogen | Cat#ant-nr-1 |
| ECF substrate | Cytiva | Cat# rpn5785 |
| Hoechst 33342 | Invitrogen | Cat#h3570 |
| Phalloidin-Alexa488 | Invitrogen | Cat#a30106 |
| Recombinant mouse osteoprotegerin (fc tag) | Cell Sciences | Cat#crm761b-fc |
| Critical commercial assays | ||
| Bone demineralization assay | CosmoBio | Cat#csr-bra-24kit |
| TRAP staining assay | CosmoBio | Cat#pmc-ak04f-cos |
| Murine m-csf elisa assay | Abcam | Cat#ab199084 |
| Murine rankl elisa assay | Abcam | Cat#ab100749 |
| Murine il-6 elisa assay | Abcam | Cat#ab222503 |
| Murine lif elisa assay | Abcam | Cat#ab238261 |
| Beta-galactosidase staining assay | Sigma | Cat#gals-1kt |
| PureLink RNA extraction kit | Invitrogen | Cat#12183018a |
| iQ SYBR Green Supermix | Biorad | Cat#1708882 |
| iScript RNA-to-cDNA kit | Biorad | Cat#1708841 |
| Experimental models: Cell lines | ||
| MC3T3-E1 clone 4 | ATCC | Cat#crl-2593 |
| MC3T3-E1 clone 14 | ATCC | Cat#crl-2594 |
| Experimental models: Organisms/strains | ||
| Prrx1-Cre/LSL-rtTA-IRES-GFP/TetO-GαsR201C/TetO-βgal mice | Collin’s Laboratory NICHD, NIH | N/A |
| Oligonucleotides | ||
| Nfactc1 primers | Sigma | PrimerPairID# M1_Nfatc1 |
| Fos primers | Sigma | PrimerPairID# M1_Fos |
| Sp7 primers | Sigma | PrimerPairID# M1_Sp7 |
| Dlx5 primers | Sigma | PrimerPairID# M1_Dxl5 |
| Bglap primers | Sigma | PrimerPairID# M1_Bglap |
| 18s rRNA primers | Sigma | PrimerPairID# M1_Rn18s |
| Actin primers | Sigma | PrimerPairID# M1_Actb |
Resource availability
Lead contact
Requests for further information, reagents and/or resources may be directed to our lead contact, Dr. Jarred M. Whitlock (Jarred.Whitlock@nih.gov).
Materials availability
The Prrx1-Cre/LSL-rtTA-IRES-GFP/TetO-GαsR201C/TetO-βga transgenic mouse model generated as a part of this study is available upon request.
Experimental model and subjectdetails
The immortalized osteoprogenitor lines MC3T3-E1 clones 4 and 14 were obtained from the AmericanType Culture Collection (Manassas, VA, USA) immediately before use in this study. The MC3T3-E1 lines were originally taken from calvaria isolated from 9 C57BL/6 mice 24 hours after birth. The sexes of the mice were not reported.54,55
For our ex vivo explant cultures, We isolated bone marrow from the tibia and femur of a previously described tet-on GαsR201C murine model crossed with a tet-on βgal reporter line to produce a quadruple transgenic model Prrx1-Cre/LSL-rtTA-IRES-GFP/TetO-GαsR201C/TetO-βgal. Expression of both GαsR201C and β-galactosidase are inducible specifically in Prrx1 progeny because of the conditional introduction of the reverse tetracycline transactivator (rtTA) via Prrx1-Cre.24,35 When introduced via Prrx1-Cre, rtTa can be specifically activated in postnatal cells of the skeletal stem cell linage (e.g., osteoprogenitors, osteoblasts) within the limbs. Addition of doxycycline induces binding of rtTA to the tet-on promotors of both TetO-GαsR201C and TetO-βgal and activates the expression of both GαsR201C and β-galactosidase.
Holes were drilled into the epyphises of each bone using a 22-gauge hypodermic needle, and the bone marrow was flushed into a culture dish using alpha MEM (Gibco). These bone marrow isolates were further dissociated through a fresh 22-gauge hypodermic needle to obtain a single cell suspension, and cultured in alpha MEM plus 20% FBS, 1x pen/strep (complete explant culture media) and 1x Normocin (InvivoGen) for 12-14 days in T-75 culture flasks. Cells that adhered to the flask were washed 3 times with PBS and passaged using 0.05% Trypsin and a cell scraper and cultured for up to 4 passages in complete explant culture media, less Normocin. For GαsR201C expression induction in the bone marrow stromal cell subset of the explants, cells were plated at ∼40% confluency in 6-well plates, 24-well plates, and angiogenesis μ-slides (Ibidy) for EV collection and staining, respectively. For induction, cells were treated with 1μM doxycycline (Sigma). During induction, media were refreshed daily. OPG was added at 100ng/ml.
All cells were incubated at 37°C in a humidified incubator supplemented with 5% CO2.
Methods details
Fluorescence microscopy imaging
In the immunofluorescence experiments we washed the cells with PBS and fixed with warm freshly prepared 4% formaldehyde in PBS (Sigma, F1268) at 370C. The cells were washed three times with PBS. To permeabilize the cells, we incubated them for 10 min in PBS with 0.1% Triton X100 in PBS and 5% FBS (IF Buffer) to suppress non-specific binding. Then the cells were incubated with primary antibodies overnight in IF Buffer. After 5 washes in IF Buffer, we placed the cells for 2h at room temperature in IF Buffer with secondary antibodies. Cells were then again washed 3 times with IF Buffer followed by 2 times with PBS. Images were captured on a Zeiss LSM 800 airyscan, confocal microscope using a C-Apochromat 63x/1.2 water immersion objective.
Bone resorption assay
Bone resorption was evaluated using bone resorption assay kits from Cosmo Bio USA according to the manufacturer’s instructions. In short, fluoresceinamine-labeled chondroitin sulfate was used to label 24-well, calcium phosphate-coated plates. Cultures were plated and differentiated as described above, using alpha MEM without phenol red. Media were collected at 4-5 days post doxycycline addition, and fluorescence intensity within the media was evaluated as recommended by the manufacture.
Fusion assays
Osteoclast fusion was evaluated by fluorescence microscopy.56 In detail, cells were fixed with 4% paraformaldehyde at the timepoints of interest described, permeabilized with 0.1% Triton X-100 and blocked with 5% FBS. Cells were then stained with phallodin-Alexa488 and Hoechst to label cells’ actin cytoskeleton and nuclei, respectively. 8 randomly selected fields of view were imaged using Alexa488, Hoechst and phase contrast compatible filter sets (BioTek) on a Lionheart FX microscope using a 10x/0.3 NA Plan Fluorite WD objective lense (BioTek) using Gen3.10 software (BioTek). Osteoclast fusion efficiency was evaluated as the number of fusion events between osteoclasts with obvious actin boarders and ≥3 nuclei in these images, as described previously.57 In detail, since regardless of the sequence of fusion events, the number of cell-to-cell fusion events required to generate syncytium with N nuclei is always equal to N-1, we calculated the fusion number index as Σ (Ni − 1) = Ntotal − Nsyn, where Ni = the number of nuclei in individual syncytia and Nsyn = the total number of syncytia. In contrast to traditional fusion index measurements, this approach gives equal consideration to fusion between two mononucleated cells, one mononucleated cell and one multinucleated cell and two multinucleated cells. In traditional fusion index calculations, fusion between two multinucleated cells does not change the percentage of nuclei in syncytia. If instead one counts the number of syncytia, a fusion event between two multinucleated is not just missed but decreases the number of syncytia. In contrast, the fusion number index is inclusive of all fusion events.
RNA extraction and transcript analysis
For real-time PCR, total RNA was collected from cell lysates using PureLink RNA kit following the manufacturer’s instructions (Invitrogen). cDNA was generated from total RNA via reverse transcription reaction using an iScript RNA-to-cDNA kit according to the manufacturer’s instructions (BioRad). cDNA was then amplified using the iQ SYBR Green Supermix (Biorad). All primers were predesigned KiCqStart SYBR Green primers with the highest rank score specific for the gene of interest or 18s control and were used according to the manufacturer’s instructions (Sigma). The catalogue numbers for all KiCqStart primers used is listed in the resource table above. Moreover, the sequences for each primer are listed in the table below.
| Gene | Fwd Primer Sequence | Rvs Primer Sequence |
|---|---|---|
| Nfatc1 | CCAAGTCAGTTTCTATGTCTG | ATAATTGGAACATTGGCAGG |
| Fos | GAAGGGAACGGAATAAGATG | CATCTTCAAGTTGATCTGTCTC |
| Sp7 | TGCTTGAGGAAGAAGCTC | CTTCTTTGTGCCTCCTTTC |
| Dlx5 | AGAGAAGGTTTCAGAAGACTC | AGATTTTCACCTGTGTTTGC |
| Bglap | ACCATGAGGACCATCTTTC | GGACATGAAGGCTTTGTC |
| 18s | CAGTTATGGTTCCTTTGGTC | TTATCTAGAGTCACCAAGCC |
All Real-time PCR reactions were performed and analyzed on a CFX96 real-time system (Biorad) for 40 cycles, using 18s ribosomal RNA as an internal control. Fold-change of gene expression was determined using the ΔΔCt method.58 3-4 independent experiments were performed (as defined in the figure legend), and each was analyzed in duplicate.
Enrichment of extracellular vesicle fractions
Ex vivo cultures were or were not induced under the conditions described for 4 days and then switched to complete explant culture media made with FBS depleted of EVs (depleted via ultracentrifugation at 150,000xg for >2 hours). Cultures were allowed to condition the media for 24 hours. Conditioned media were collected, and cells/large cell debris were depleted via centrifugation (15 minsat 4,000xg). Next, an EV fraction was enriched via centrifugation (150,000xg for 1.5 hours). EV enriched fractions were evaluated via Western Blot using α-TSG101 the α-RANK antibodies described above. Similar results were also obtained using the exoEasy Maxi Kit (Qiagen) following the manufacture’s instructions. TSG101 and RANK signals were quantified using densitometry via ImageJ.
Protein extraction, gel electrophoresis and Western blotting
Cells were collected via cell scraper in cold lysis buffer (PBS (Gibco) supplemented with 1% Tx-100), sonicated on ice via pulse sonication and rotated end over end at 4°C for 45 minutes, all in the presence of protease inhibitors (cOmplete, Sigma). Next, insoluble material was removed via centrifugation at 15,000 times gravity at 4°C for 15 minutes. The protein concentrations of lysates were assessed via BCA assay (Pierce) following the manufacture’s instructions. 7.5 micrograms of lysates denatured in loading buffer (Biorad), were separated via sodium dodecyl-sulfate polyacrylamide gel electrophoresis using 5-20% polyacrylamide, mini, precast gels (Biorad). Steady-state protein levels were evaluated via SDS-PAGE followed by immunoblotting. PVDF membranes were blocked in 5% milk prepared in PBS with 1% tween-20 (PBST) for 60 minutes. Primary antibodies were incubated with membranes in 1% milk prepared in PBST overnight at 4°C, and secondary antibodies were incubated with membranes for in 1% milk prepared in PBST for 1 hour at room temperature. The antibodies described were used at a 1:500 dilution, with the exception of a-Cyclophilin B which was used at a 1:2000 dilution. For Secondary antibodies, we used α-Rb (Abcam) or α-Ms (Abcam) conjugated to alkaline phosphatase at a dilution of 1:3000. EFC substrate was incubated with membranes for 5-10 minutes (cytiva, RPN5785) and the resulting bands were visualized via a GE Typhoon TLA 7000 imager.
Elisa assays
Conditioned media were collected from separate wells of a multiwell plate for each experimental condition described 4 days after induction. Media were quickly frozen and stored at -30°C. Elisa assays were utilized to evaluate the presence of soluble factors in the conditioned media following the manufacure’s instructions.
Quantification and statistical analysis
Statistical analyses were performed using Prism software (GraphPad Prism version 8.0.0). Differences between groups were analyzed by Student’s t test. The results of quantitative data presented here are all presented as mean ± the standard error of the mean (SEM). Statistical significance was evaluated as p=<0.05, where ∗p=<0.05, ∗∗p=<0.01, ∗∗∗p=<0.001.
Ethics approval
All animal studies were carried out according to NIH-Intramural Animal Care and Use Committee (ACUC) of the National Institute of Dental and Craniofacial Research approved protocols (ASP #19-897), in compliance with the Guide for the Care and Use of Laboratory Animals.
Acknowledgments
We thank Rebeca Galisteo and Tiffani Slaughter of M.T.C’s laboratory for providing the sacrificed mice from which explant cultures were isolated. The research in L.V.C. laboratory was supported by the Intramural Research Program of the Eunice Kennedy Shriver National Institute of Child Health and Human Development, National Institutes of Health. The research in A.M.B.’ and M.T.C.’ laboratories was supported by the Intramural Research Program of the National Institute of Dental and Craniofacial Research, National Institutes of Health. Work in MTC lab and LVC labs was also supported by the Office of Research on Women’s Health (ORWH) through the Bench to Bedside Program award #884515.
Author contributions
J.M.W., L.F.C.D., and L.V.C. designed the model described and the experiments to implement its use. J.M.W. performed the experiments described, analyzed the data and produced all figures. M.T.C and A.M.B. contributed intellectually to experimental approaches for assessing osteoclast-to-osteoprogenitor crosstalk. L.F.C.D. maintained the mouse model described and carried out genetic crosses to introduce the β-gal lineage tracer and ensure the fidelity of the model. J.M.W. and L.V.C. wrote the manuscript with assistance from L.F.C.D., M.T.C., and A.M.B.
Declaration of interests
The authors declare no competing interests.
Inclusion and diversity
We support inclusive, diverse, and equitable conduct of research.
Published: March 23, 2023
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.isci.2023.106470.
Supplemental information
Data and code availability
All data used to generate the figures in this manuscript, including uncropped, original Western blot images are available as Data S1 in the supplemental materials associated with this manuscript. Microscopy data will be shared by the lead contact upon request. No original code was written as part of this study. Any additional information required to reanalyze the data reported in this paper is available upon request from the lead contact listed above.
References
- 1.Feng X., McDonald J.M. Disorders of bone remodeling. Annu. Rev. Pathol. 2011;6:121–145. doi: 10.1146/annurev-pathol-011110-130203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Raggatt L.J., Partridge N.C. Cellular and molecular mechanisms of bone remodeling. J. Biol. Chem. 2010;285:25103–25108. doi: 10.1074/jbc.R109.041087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Kodama H., Nose M., Niida S., Yamasaki A. Essential role of macrophage colony-stimulating factor in the osteoclast differentiation supported by stromal cells. J. Exp. Med. 1991;173:1291–1294. doi: 10.1084/jem.173.5.1291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Nakashima T., Hayashi M., Fukunaga T., Kurata K., Oh-Hora M., Feng J.Q., Bonewald L.F., Kodama T., Wutz A., Wagner E.F., et al. Evidence for osteocyte regulation of bone homeostasis through RANKL expression. Nat. Med. 2011;17:1231–1234. doi: 10.1038/nm.2452. [DOI] [PubMed] [Google Scholar]
- 5.Li D., Liu J., Guo B., Liang C., Dang L., Lu C., He X., Cheung H.Y.S., Xu L., Lu C., et al. Osteoclast-derived exosomal miR-214-3p inhibits osteoblastic bone formation. Nat. Commun. 2016;7:10872. doi: 10.1038/ncomms10872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Sun W., Zhao C., Li Y., Wang L., Nie G., Peng J., Wang A., Zhang P., Tian W., Li Q., et al. Osteoclast-derived microRNA-containing exosomes selectively inhibit osteoblast activity. Cell Discov. 2016;2:16015. doi: 10.1038/celldisc.2016.15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Huynh N., VonMoss L., Smith D., Rahman I., Felemban M.F., Zuo J., Rody W.J., Jr., McHugh K.P., Holliday L.S. Characterization of regulatory extracellular vesicles from osteoclasts. J. Dent. Res. 2016;95:673–679. doi: 10.1177/0022034516633189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Weivoda M.M., Chew C.K., Monroe D.G., Farr J.N., Atkinson E.J., Geske J.R., Eckhardt B., Thicke B., Ruan M., Tweed A.J., et al. Identification of osteoclast-osteoblast coupling factors in humans reveals links between bone and energy metabolism. Nat. Commun. 2020;11:87. doi: 10.1038/s41467-019-14003-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Koide M., Kobayashi Y., Yamashita T., Uehara S., Nakamura M., Hiraoka B.Y., Ozaki Y., Iimura T., Yasuda H., Takahashi N., Udagawa N. Bone Formation is coupled to resorption via suppression of sclerostin expression by osteoclasts. J. Bone Miner. Res. 2017;32:2074–2086. doi: 10.1002/jbmr.3175. [DOI] [PubMed] [Google Scholar]
- 10.Manolagas S.C. Birth and death of bone cells: basic regulatory mechanisms and implications for the pathogenesis and treatment of osteoporosis. Endocr. Rev. 2000;21:115–137. doi: 10.1210/edrv.21.2.0395. [DOI] [PubMed] [Google Scholar]
- 11.Kim J.H., Kim N. Regulation of NFATc1 in osteoclast differentiation. J. Bone Metab. 2014;21:233–241. doi: 10.11005/jbm.2014.21.4.233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Roodman G.D., Windle J.J. Paget disease of bone. J. Clin. Invest. 2005;115:200–208. doi: 10.1172/JCI24281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Abdallah D., Jourdain M.L., Braux J., Guillaume C., Gangloff S.C., Jacquot J., Velard F. An optimized method to generate human active osteoclasts from peripheral blood monocytes. Front. Immunol. 2018;9:632. doi: 10.3389/fimmu.2018.00632. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Møller A.M.J., Delaissé J.M., Olesen J.B., Madsen J.S., Canto L.M., Bechmann T., Rogatto S.R., Søe K. Aging and menopause reprogram osteoclast precursors for aggressive bone resorption. Bone Res. 2020;8:27. doi: 10.1038/s41413-020-0102-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Møller A.M.J., Delaissé J.M., Olesen J.B., Canto L.M., Rogatto S.R., Madsen J.S., Søe K. Fusion potential of human osteoclasts in vitro reflects age, menopause, and in vivo bone resorption levels of their donors-A possible involvement of DC-STAMP. Int. J. Mol. Sci. 2020;21:6368. doi: 10.3390/ijms21176368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Makris G.P., Saffar J.L. Quantitative relationship between osteoclasts, osteoclast nuclei and the extent of the resorbing surface in hamster periodontal disease. Arch. Oral Biol. 1982;27:965–969. doi: 10.1016/0003-9969(82)90104-2. [DOI] [PubMed] [Google Scholar]
- 17.Piper K., Boyde A., Jones S.J. The relationship between the number of nuclei of an osteoclast and its resorptive capability in vitro. Anat. Embryol. 1992;186:291–299. doi: 10.1007/BF00185977. [DOI] [PubMed] [Google Scholar]
- 18.Boissy P., Saltel F., Bouniol C., Jurdic P., Machuca-Gayet I. Transcriptional activity of nuclei in multinucleated osteoclasts and its modulation by calcitonin. Endocrinology. 2002;143:1913–1921. doi: 10.1210/endo.143.5.8813. [DOI] [PubMed] [Google Scholar]
- 19.Boyce B.F., Xing L. The RANKL/RANK/OPG pathway. Curr. Osteoporos. Rep. 2007;5:98–104. doi: 10.1007/s11914-007-0024-y. [DOI] [PubMed] [Google Scholar]
- 20.Sarafrazi N., Wambogo E.A., Shepherd J.A. NCHS Data Brief; 2021. Osteoporosis or Low Bone Mass in Older Adults: United States, 2017-2018; pp. 1–8. [PubMed] [Google Scholar]
- 21.Stattin E.L., Henning P., Klar J., McDermott E., Stecksen-Blicks C., Sandström P.E., Kellgren T.G., Rydén P., Hallmans G., Lönnerholm T., et al. SNX10 gene mutation leading to osteopetrosis with dysfunctional osteoclasts. Sci. Rep. 2017;7:3012. doi: 10.1038/s41598-017-02533-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Eastell R., O'Neill T.W., Hofbauer L.C., Langdahl B., Reid I.R., Gold D.T., Cummings S.R. Postmenopausal osteoporosis. Nat. Rev. Dis. Primers. 2016;2:16069. doi: 10.1038/nrdp.2016.69. [DOI] [PubMed] [Google Scholar]
- 23.Kravets I. Paget's disease of bone: diagnosis and treatment. Am. J. Med. 2018;131:1298–1303. doi: 10.1016/j.amjmed.2018.04.028. [DOI] [PubMed] [Google Scholar]
- 24.Zhao X., Deng P., Iglesias-Bartolome R., Amornphimoltham P., Steffen D.J., Jin Y., Molinolo A.A., de Castro L.F., Ovejero D., Yuan Q., et al. Expression of an active Galphas mutant in skeletal stem cells is sufficient and necessary for fibrous dysplasia initiation and maintenance. Proc. Natl. Acad. Sci. USA. 2018;115:E428–E437. doi: 10.1073/pnas.1713710115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Hwang P.W., Horton J.A. Variable osteogenic performance of MC3T3-E1 subclones impacts their utility as models of osteoblast biology. Sci. Rep. 2019;9:8299. doi: 10.1038/s41598-019-44575-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Song C., Yang X., Lei Y., Zhang Z., Smith W., Yan J., Kong L. Evaluation of efficacy on RANKL induced osteoclast from RAW264.7 cells. J. Cell. Physiol. 2019;234:11969–11975. doi: 10.1002/jcp.27852. [DOI] [PubMed] [Google Scholar]
- 27.Ng A.Y., Tu C., Shen S., Xu D., Oursler M.J., Qu J., Yang S. Comparative characterization of osteoclasts derived from murine bone marrow macrophages and RAW 264.7 cells using quantitative proteomics. JBMR Plus. 2018;2:328–340. doi: 10.1002/jbm4.10058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Lacey D.L., Timms E., Tan H.L., Kelley M.J., Dunstan C.R., Burgess T., Elliott R., Colombero A., Elliott G., Scully S., et al. Osteoprotegerin ligand is a cytokine that regulates osteoclast differentiation and activation. Cell. 1998;93:165–176. doi: 10.1016/s0092-8674(00)81569-x. [DOI] [PubMed] [Google Scholar]
- 29.Henriksen K., Karsdal M.A., Taylor A., Tosh D., Coxon F.P. Generation of human osteoclasts from peripheral blood. Methods Mol. Biol. 2012;816:159–175. doi: 10.1007/978-1-61779-415-5_11. [DOI] [PubMed] [Google Scholar]
- 30.Nemeth K., Mayer B., Sworder B.J., Kuznetsov S.A., Mezey E. A practical guide to culturing mouse and human bone marrow stromal cells. Curr. Protoc. Immunol. 2013;102:22F.12.1–22F.12.13. doi: 10.1002/0471142735.im22f12s102. [DOI] [PubMed] [Google Scholar]
- 31.Gartland A., Buckley K.A., Dillon J.P., Curran J.M., Hunt J.A., Gallagher J.A. Isolation and culture of human osteoblasts. Methods Mol. Med. 2005;107:29–54. doi: 10.1385/1-59259-861-7:029. [DOI] [PubMed] [Google Scholar]
- 32.Piersanti S., Remoli C., Saggio I., Funari A., Michienzi S., Sacchetti B., Robey P.G., Riminucci M., Bianco P. Transfer, analysis, and reversion of the fibrous dysplasia cellular phenotype in human skeletal progenitors. J. Bone Miner. Res. 2010;25:1103–1116. doi: 10.1359/jbmr.091036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Yamamoto T., Ozono K., Kasayama S., Yoh K., Hiroshima K., Takagi M., Matsumoto S., Michigami T., Yamaoka K., Kishimoto T., Okada S. Increased IL-6-production by cells isolated from the fibrous bone dysplasia tissues in patients with McCune-Albright syndrome. J. Clin. Invest. 1996;98:30–35. doi: 10.1172/JCI118773. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.de Castro L.F., Burke A.B., Wang H.D., Tsai J., Florenzano P., Pan K.S., Bhattacharyya N., Boyce A.M., Gafni R.I., Molinolo A.A., et al. Activation of RANK/RANKL/OPG pathway is involved in the pathophysiology of fibrous dysplasia and associated with disease burden. J. Bone Miner. Res. 2019;34:290–294. doi: 10.1002/jbmr.3602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Vitale-Cross L., Amornphimoltham P., Fisher G., Molinolo A.A., Gutkind J.S. Conditional expression of K-ras in an epithelial compartment that includes the stem cells is sufficient to promote squamous cell carcinogenesis. Cancer Res. 2004;64:8804–8807. doi: 10.1158/0008-5472.CAN-04-2623. [DOI] [PubMed] [Google Scholar]
- 36.Amarasekara D.S., Kim S., Rho J. Regulation of osteoblast differentiation by cytokine networks. Int. J. Mol. Sci. 2021;22:2851. doi: 10.3390/ijms22062851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Ikebuchi Y., Aoki S., Honma M., Hayashi M., Sugamori Y., Khan M., Kariya Y., Kato G., Tabata Y., Penninger J.M., et al. Coupling of bone resorption and formation by RANKL reverse signalling. Nature. 2018;561:195–200. doi: 10.1038/s41586-018-0482-7. [DOI] [PubMed] [Google Scholar]
- 38.Herrmann M., Diederichs S., Melnik S., Riegger J., Trivanović D., Li S., Jenei-Lanzl Z., Brenner R.E., Huber-Lang M., Zaucke F., et al. Extracellular vesicles in musculoskeletal pathologies and regeneration. Front. Bioeng. Biotechnol. 2020;8:624096. doi: 10.3389/fbioe.2020.624096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Yuan F.L., Wu Q.Y., Miao Z.N., Xu M.H., Xu R.S., Jiang D.L., Ye J.X., Chen F.H., Zhao M.D., Wang H.J., Li X. Osteoclast-derived extracellular vesicles: novel regulators of osteoclastogenesis and osteoclast-osteoblasts communication in bone remodeling. Front. Physiol. 2018;9:628. doi: 10.3389/fphys.2018.00628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Liang M., Yin X., Zhang S., Ai H., Luo F., Xu J., Dou C., Dong S., Ma Q. Osteoclast-derived small extracellular vesicles induce osteogenic differentiation via inhibiting ARHGAP1. Mol. Ther. Nucleic Acids. 2021;23:1191–1203. doi: 10.1016/j.omtn.2021.01.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Shao J.L., Li H., Zhang X.R., Zhang X., Li Z.Z., Jiao G.L., Sun G.D. Identification of serum exosomal MicroRNA expression profiling in menopausal females with osteoporosis by high-throughput sequencing. Curr. Med. Sci. 2020;40:1161–1169. doi: 10.1007/s11596-020-2306-x. [DOI] [PubMed] [Google Scholar]
- 42.Huo C., Li Y., Qiao Z., Shang Z., Cao C., Hong Y., Xiao H. Comparative proteomics analysis of microvesicles in human serum for the evaluation of osteoporosis. Electrophoresis. 2019;40:1839–1847. doi: 10.1002/elps.201900130. [DOI] [PubMed] [Google Scholar]
- 43.Riminucci M., Kuznetsov S.A., Cherman N., Corsi A., Bianco P., Gehron Robey P. Osteoclastogenesis in fibrous dysplasia of bone: in situ and in vitro analysis of IL-6 expression. Bone. 2003;33:434–442. doi: 10.1016/s8756-3282(03)00064-4. [DOI] [PubMed] [Google Scholar]
- 44.Yoshitake F., Itoh S., Narita H., Ishihara K., Ebisu S. Interleukin-6 directly inhibits osteoclast differentiation by suppressing receptor activator of NF-kappaB signaling pathways. J. Biol. Chem. 2008;283:11535–11540. doi: 10.1074/jbc.M607999200. [DOI] [PubMed] [Google Scholar]
- 45.Feng W., Liu H., Luo T., Liu D., Du J., Sun J., Wang W., Han X., Yang K., Guo J., et al. Combination of IL-6 and sIL-6R differentially regulate varying levels of RANKL-induced osteoclastogenesis through NF-kappaB, ERK and JNK signaling pathways. Sci. Rep. 2017;7:41411. doi: 10.1038/srep41411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Udagawa N., Takahashi N., Katagiri T., Tamura T., Wada S., Findlay D.M., Martin T.J., Hirota H., Taga T., Kishimoto T., Suda T. Interleukin (IL)-6 induction of osteoclast differentiation depends on IL-6 receptors expressed on osteoblastic cells but not on osteoclast progenitors. J. Exp. Med. 1995;182:1461–1468. doi: 10.1084/jem.182.5.1461. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Tamura T., Udagawa N., Takahashi N., Miyaura C., Tanaka S., Yamada Y., Koishihara Y., Ohsugi Y., Kumaki K., Taga T., et al. Soluble interleukin-6 receptor triggers osteoclast formation by interleukin 6. Proc. Natl. Acad. Sci. USA. 1993;90:11924–11928. doi: 10.1073/pnas.90.24.11924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Chapurlat R., Gensburger D., Trolliet C., Rouanet S., Mehsen-Cetre N., Orcel P. Inhibition of IL-6 in the treatment of fibrous dysplasia of bone: the randomized double-blind placebo-controlled TOCIDYS trial. Bone. 2022;157:116343. doi: 10.1016/j.bone.2022.116343. [DOI] [PubMed] [Google Scholar]
- 49.Akatsu T., Tamura T., Takahashi N., Udagawa N., Tanaka S., Sasaki T., Yamaguchi A., Nagata N., Suda T. Preparation and characterization of a mouse osteoclast-like multinucleated cell population. J. Bone Miner. Res. 1992;7:1297–1306. doi: 10.1002/jbmr.5650071109. [DOI] [PubMed] [Google Scholar]
- 50.Udagawa N., Takahashi N., Akatsu T., Tanaka H., Sasaki T., Nishihara T., Koga T., Martin T.J., Suda T. Origin of osteoclasts: mature monocytes and macrophages are capable of differentiating into osteoclasts under a suitable microenvironment prepared by bone marrow-derived stromal cells. Proc. Natl. Acad. Sci. USA. 1990;87:7260–7264. doi: 10.1073/pnas.87.18.7260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Takahashi N., Akatsu T., Udagawa N., Sasaki T., Yamaguchi A., Moseley J.M., Martin T.J., Suda T. Osteoblastic cells are involved in osteoclast formation. Endocrinology. 1988;123:2600–2602. doi: 10.1210/endo-123-5-2600. [DOI] [PubMed] [Google Scholar]
- 52.Whitlock J.M., Leikina E., Melikov K., De Castro L.F., Mattijssen S., Maraia R.J., Collins M.T., Chernomordik L.V. Cell surface-bound La protein regulates the cell fusion stage of osteoclastogenesis. Nat. Commun. 2023;14:616. doi: 10.1038/s41467-023-36168-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.de Castro L.F., Whitlock J.M., Michel Z., Pan K., Taylor J., Szymczuk V., Paravastu S., Savoury B., Papadakis G.Z., Chernomordik L.V., et al. RANKL inhibition with denosumab improves fibrous dysplasia by decreasing lesional cell proliferation and increasing osteogenesis. medRxiv. 2022 doi: 10.1101/2022.10.24.22281375. Preprint at. [DOI] [Google Scholar]
- 54.Wang D., Christensen K., Chawla K., Xiao G., Krebsbach P.H., Franceschi R.T. Isolation and characterization of MC3T3-E1 preosteoblast subclones with distinct in vitro and in vivo differentiation/mineralization potential. J. Bone Miner. Res. 1999;14:893–903. doi: 10.1359/jbmr.1999.14.6.893. [DOI] [PubMed] [Google Scholar]
- 55.Kodama H.a., Amagai Y., Sudo H., Kasai S., Yamamoto S. Establishment of a clonal osteogenic cell line from newborn mouse calvaria. Jpn. J. Oral Biol. 1981;23:899–901. [Google Scholar]
- 56.Verma S.K., Leikina E., Melikov K., Gebert C., Kram V., Young M.F., Uygur B., Chernomordik L.V. Cell-surface phosphatidylserine regulates osteoclast precursor fusion. J. Biol. Chem. 2018;293:254–270. doi: 10.1074/jbc.M117.809681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Verma S.K., Chernomordik L.V., Melikov K. An improved metrics for osteoclast multinucleation. Sci. Rep. 2018;8:1768. doi: 10.1038/s41598-018-20031-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Livak K.J., Schmittgen T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25:402–408. doi: 10.1006/meth.2001.1262. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All data used to generate the figures in this manuscript, including uncropped, original Western blot images are available as Data S1 in the supplemental materials associated with this manuscript. Microscopy data will be shared by the lead contact upon request. No original code was written as part of this study. Any additional information required to reanalyze the data reported in this paper is available upon request from the lead contact listed above.