Abstract
BXSB mice develop a lupus-like autoimmune syndrome. We have previously identified several intervals that were linked to disease and, in an attempt to characterise lupus susceptibility genes within these intervals, we have sought to isolate differentially expressed genes. Representational difference analysis was used to compare gene expression between BXSB and C57BL/10 mice using spleen and thymus as a source of mRNA. The majority of differentially expressed sequences identified were immunoglobulin and gp70-related sequences, overexpression of these being characteristic of the disease. Among other isolated sequences were a sialyltransferase gene, a mouse tumour virus superantigen gene (Mtv-3), and the virus-related sequence, hitchhiker. In BXSB the sialyltransferase gene not only overexpressed spliced transcripts, but also produced high levels of unspliced mRNA. Further analysis demonstrated that the copy number of the three linked sequences: sialyltransferase, Mtv-3 and hitchhiker, was amplified in BXSB and that the structural organisation of this locus varies in different mouse strains. This locus consists of three parts, Mtv-3–hitchhiker–sialyltransferase, hitchhiker–sialyltransferase, and sialyltransferase alone. Different combinations of the regions are present in different mouse strains. Linkage analysis demonstrated that this region at the distal end of chromosome 11 is weakly linked to phenotypic markers of disease.
INTRODUCTION
Systemic lupus erythematosus (SLE) is a chronic autoimmune disease characterised by the development of autoantibodies to a spectrum of nuclear antigens. Patients present with a wide range of symptoms, from a facial rash to severe glomerulonephritis, raising the possibility that SLE is in fact a syndrome of related disorders. Disease susceptibility is inherited as a complex trait involving both environmental and multiple genetic factors. There is limited information about the genes that predispose to the development of SLE. Studies in both human and mouse models of the disease have identified several genomic intervals as important in disease susceptibility (reviewed in 1–4), while recent data supports a role in disease pathology for reduced expression of DNASE 1 as a result of gene mutation (5). Due to the complexity of the disease, at both the genetic and phenotypic levels, in-bred mouse strains that spontaneously develop SLE provide important models for the study of this disease. We have focussed our studies on the BXSB mouse, a recombinant in-bred strain derived from an original cross between a C57BL/6 (B6) female and an SB/Le male, which spontaneously develops an autoimmune disease very similar to human SLE (6). Phenotypically the BXSB mouse is characterised by the production of autoantibodies to nuclear antigens and a severe diffuse proliferative glomerulonephritis (7). We have performed genome-wide linkage studies in crosses between the non-autoimmune strain C57BL/10 (B10) and BXSB mice and identified several intervals on chromosomes 1, 3, 4, 10, 13 and 19 that were linked to disease phenotypes (8,9). The challenge is to identify the disease susceptibility loci within these intervals.
Traditional methods to identify the individual genes include the candidate gene approach and positional cloning using congenic mice (10). In this way, we have successfully identified Fcgr2b as a strong candidate gene (11) and, currently, congenic mice carrying BXSB intervals on a B10 background are being developed. However, recent developments in the techniques of differential gene expression are proving to be complementary to linkage studies in the identification of susceptibility genes within disease intervals. The use of cDNA and oligonucleotide microarrays has been successfully used to identify Cd36 as an insulin-resistance gene causing defective fatty acid and glucose metabolism in hypertensive rats (12). Using serial analysis of gene expression, 79 differentially expressed genes have been identified in endothelial cells derived from blood vessels of normal compared with malignant colorectal tissues (13). While these two technologies are proving successful, they are expensive to perform. Other techniques, such as differential display PCR (14), representational difference analysis (RDA) (15) and suppression subtractive hybridisation (16) have also been widely applied to investigate the molecular events underlying a variety of diseases, and these techniques offer convenient and inexpensive methods for the study of differentially expressed genes. Here, we describe the use of cDNA RDA, a differential hybridisation amplification method, to isolate genes that are differentially expressed between BXSB and B10.
MATERIALS AND METHODS
Mouse strains
Mice were bred and maintained under identical conditions at the Imperial College School of Medicine from original stocks: BXSB from the Jackson Laboratory (Bar Harbor, ME) and B10 from Harlan Olac Ltd (Bicester, UK). For RDA analysis, all mice were 2–4-month-old males, before the onset of the autoimmune phenotype.
RNA isolation, cDNA synthesis and cDNA RDA
Total RNA was isolated from thymus and spleen using RNAzol B reagent (Biogenesis Ltd). Poly(A)+ RNA was purified further from total RNA using a mRNA isolation kit (Qiagen). Double-stranded cDNA was synthesised from mRNA using the cDNA synthesis kit (Gibco BRL) and RDA was performed as described (17). The driver cDNA was obtained from B10 thymus and spleen, and tester was from BXSB thymus and spleen.
Reverse transcriptase–PCR and genomic PCR
For reverse transcriptase–PCR (RT–PCR) analysis, ∼1 µg of poly(A)+ RNA was reverse-transcribed by the Superscript II first-strand cDNA synthesis kit (Gibco BRL). Conditions for PCR were 20–30 cycles of 1 min each at 94°C, 55–63°C (dependent on the primers) and 72°C with Taq DNA polymerase (Bioline). Primers are listed in Table 1. For semi-quantitative PCR, the control primers for the RT–PCR were designed from the ubiquitously expressed gene, growth factor receptor-bound protein 2 (Grb2) (18), while those for genomic PCR were from the Ifi202 promoter region (19).
Table 1. Primers used in PCR reactions.
Positions of the primers are illustrated in Figure 5. All numbering is taken from the GenBank entry for each sequence.
Genome walking and rapid amplification of cDNA ends (RACE) analysis
Genome walking was performed with a Genome Walker kit (Clontech) according to the manufacturer’s protocol. Amplification of the 3′ cDNA ends was performed using the SMART RACE cDNA amplification kit (Clontech).
Southern and northern blot analysis
Northern blot analysis was performed as described (20). Total RNA (10 µg) was size-fractionated on 1.2% agarose/2.2 M formaldehyde gels and transferred to nylon membranes. For Southern blot analysis genomic DNA was isolated from liver, digested with restriction enzymes, size-fractionated on 0.8% agarose gels containing ethidium bromide, and transferred onto nylon membranes (Stratagene). The sialyltransferase and hitchhiker probes were generated by PCR using primer pairs 7–8 and 14–15 (Table 1) designed from RDA sequences. Probe identity was confirmed by automated DNA sequence analysis. PCR products were radiolabelled with 25 µCi of [α-32P]dCTP using Megaprime (Amersham Pharmacia). The Southern filters were hybridised in Church and Gilbert hybridisation buffers (21) and washed as described (20). For copy number determination, autoradiographs of Southern blots were scanned with an Appligene Image analysis system (Appligene) and densitometry was performed with National Institutes of Health IMAGE 1.52 software.
Radiation hybrid mapping
The T31 radiation hybrid panel was purchased from Research Genetics and PCR was performed as recommended by the supplier.
Linkage analysis
Phenotype and genotype data were collected and χ2 and QTL analyses were performed as described previously (8,9).
RESULTS
RDA analysis of spleen and thymus cDNA
In order to identify potential disease susceptibility genes in BXSB we utilised RDA directed to identify sequences that were overexpressed in BXSB compared with B10. All mice analysed were 2–4-month-old males, and RNA was extracted from both thymus and spleen. After three rounds of subtraction, 624 individual clones were amplified by PCR. The products were digested with DpnII, electrophoresed through duplicate agarose gels, and blotted onto nylon membranes. Filters were probed with radiolabelled first-round cDNA from BXSB and B10. Fragments (260) exhibiting differential hybridisation were sequenced and compared with the GenBank database via a BLAST search to determine their identity. The majority of these sequenced fragments were immunoglobulin- and gp70-related sequences, which would be expected to be upregulated in active lupus. To confirm that the non-Ig fragments were differentially expressed, either the fragments were radiolabelled and used as probes against northern blots of RNA from BXSB and B10, and/or primers were designed from RDA fragments and used for semi-quantitative RT–PCR analysis. The fragments that exhibited strong differential expression were investigated for potential susceptibility to disease using linkage analysis in backcross mice as described previously (8,9).
Identification of three linked sequences: a murine mammary tumour virus (MMTV) superantigen, a sialyltransferase gene and a virus-related sequence
Several RDA fragments were identified that were derived from the same parent gene sequences: a MMTV superantigen (Mtv-3); a sialyltransferase gene (ST6GalNac1); and a sequence termed ‘hitchhiker’ (22). A 12 kb genomic sequence had previously been described that encoded hitchhiker and an Mtv-3 integration site (22). However, in the published study it was not described that the same fragment also contained some sequence from the sialyltransferase gene encoded on the opposite strand. Thus, these three sequences, all overexpressed in BXSB, were contained within a single genomic fragment. We tested this region for linkage to disease phenotypes by both QTL analysis with MAPMAKER and by χ2 analysis as described previously (8,9). There was weak linkage to nephritis (χ2 = 6.8, P = 0.009), splenomegaly (χ2 = 5.2, P = 0.02) and to the production of anti-ssDNA antibodies (LOD = 0.75, P = 0.06).
A BLAST search against the protein databases revealed that the predicted amino acid sequence from many parts of the hitchhiker genomic sequence, including the introns, had homology to the murine leukemia virus (MuLV) gag and pol genes; 31–45% identity over stretches of 56–187 amino acids. These sequences cannot be translated into functional protein, due to frame shifts and in-frame termination codons in the sequences, but suggest that hitchhiker may have had a viral origin.
The sialyltransferase and hitchhiker sequences were overexpressed in BXSB in comparison with B10, as demonstrated in northern blot analysis (data not shown) and semi-quantitative RT–PCR (Fig. 1). Mtv-3 expression was restricted to BXSB, which was consistent with the previously reported finding that Mtv-3 is absent from the B10 genome (23). In BXSB thymus and spleen, both the hitchhiker and sialyltransferase genes not only overexpressed the normal spliced transcripts, but also produced high levels of unspliced or incompletely spliced transcripts (Fig. 1). Genomic contamination of the RNA was excluded since in the absence of RT no amplicons were detectable. The presence of unspliced mRNA in the cytoplasm was supported by the observation that the majority of the RDA fragments isolated for both the sialyltransferase and hitchhiker genes contained intronic sequence.
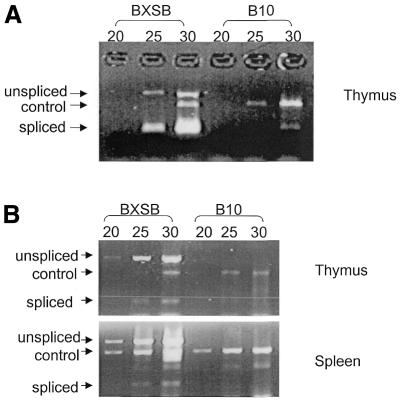
Figure 1.
Semi-quantitative RT–PCR analysis of the hitchhiker sequence and sialyltransferase gene. (A) RT–PCR using hitchhiker primers 12 and 13 together with control primers were performed at 20, 25 and 30 cycles, using an equal amount of BXSB and B10 thymus cDNA as template. (B) Sialyltransferase gene primers 7 and 8 were used in semi-quantitative RT–PCR analysis, with an equal amount of BXSB and B10 thymus cDNA (top) and spleen cDNA (bottom) as templates. Bands for unspliced and spliced transcripts together with the control products are indicated.
Characterisation of two forms of the sialyltransferase gene
The published 12 kb sequence for this region contained only part of the sialyltransferase gene sequence. Therefore, we performed primer walking and 3′ RACE to isolate and analyse the entire cDNA from B10 mouse spleen mRNA. Subsequently, a cDNA sequence for ST6GalNac1 was published (24) and its sequence contained a perfect match to a 4.5 kb region in the hitchhiker fragment of the 12 kb genomic sequence (22). The cDNA sequences we derived also matched the published sequence exactly, with the exception of an anomaly within the 3′ untranslated region (UTR). We identified two sequence populations for this region, one of which matched the published sequence, and a second, novel sequence. PCR analysis using primers 1 and 2 specific for the second sequence ruled out the possibility that this sequence might be an artifact and confirmed that it was present in both mRNA and genomic DNA from all mouse strains tested (Fig. 2A). In contrast, the hitchhiker region-derived variant 3′ end was absent from several mouse strains (Fig. 2B). Thus, the published sequence (24) represented a variant form of the sialyltransferase gene, in which a viral sequence (hitchhiker) has inserted into the 3′ UTR of the gene. Primer walking and 3′ RACE experiments were performed to identify the corresponding genomic sequence of the normal form of the gene and to determine the polyadenylation site. BALB/c genomic DNA and spleen cDNA were used as templates since the hitchhiker sequence is absent from this strain (see below). We sequenced 1343 bp of genomic sequence downstream of the gene and the polyadenylation site was identified at 819 bp 3′ to the termination codon (sequence deposited with GenBank, accession no. AF351128).
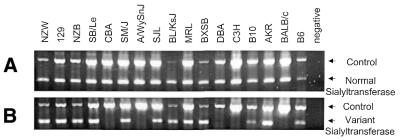
Figure 2.
PCR analysis of genomic DNA for the presence of the normal and variant sialyltransferase sequences. PCR was performed using primers 1 and 2, specific for the normal sialyltransferase gene (A), and using primers 17 and 5 specific for the variant sialyltransferase gene with the hitchhiker insertion (B), in additon to control primers.
Differential expression of normal and variant forms of the sialyltransferase gene
Competitive RT–PCR analyses were performed to investigate the expression patterns of these two forms of the sialyltransferase gene. Three primers were used, the reverse primer (primer 2) was specific for the common region of the 3′ end of the sialyltransferase gene, while the two forward primers, primers 1 and 3, were specific for the 3′ UTRs of the normal (384 bp amplicon) and variant (665 bp amplicon) forms, respectively. In spleen and thymus of BXSB, the variant form of the sialyltransferase gene expressed at a significantly higher level than the normal gene. In contrast, in B10 spleen the variant form expressed at a lower level than the normal gene and in B10 thymus both were expressed at a similar level. BALB/c only expressed the normal gene (Fig. 3A), consistent with the absence of hitchhiker sequence from the BALB/c genome (Fig. 2B).
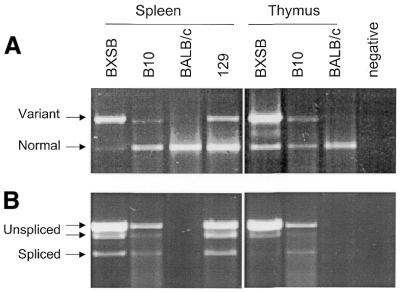
Figure 3.
RT–PCR analysis of the sialyltransferase gene. (A) RT–PCR analysis of variant and normal forms of the sialyltransferase gene. The RT–PCR reactions contain three primers: primer 2 which is common to both forms, primer 1 which is specific to normal form, and primer 3 which is specific to variant form. (B) RT–PCR analysis of splicing patterns of variant form of sialyltransferase gene. PCR using primers 6 and 16 which span two introns was performed. Spliced and unspliced bands are indicated.
In order to determine whether the unspliced sialyltransferase transcripts were derived from the normal or variant form, we performed an RT–PCR using primer 1 which specifically targets the 3′ UTR of the normal gene, within the last exon, and primer 6 which targets the third to last exon. Only spliced transcripts were produced in the spleen and thymus of BXSB and B10 (data not shown). In contrast, an RT–PCR using primer 16, specific to the 3′ UTR of the variant form of the gene and the common exon primer 6 gave two unspliced transcripts in addition to the normally spliced transcript, in both spleen and thymus of BXSB and B10 (Fig. 3B). Thus, the unspliced transcripts are derived from the variant gene only.
Genomic organisation of the Mtv-3, hitchhiker and sialyltransferase gene loci
As we have shown (Fig. 2A), all mouse strains contained the normal form of the sialyltransferase gene. Therefore, we looked for the presence of the hitchhiker sequence with and without insertion into sialyltransferase. Using Southern blot analysis (data not shown) we were unable to detect the presence of the hitchiker sequence in the absence of the surrounding sialyltransferase gene.
In order to characterise this region further, we studied the Mtv-3 insertion into the hitchhiker gene by PCR. The primer pair 17 and 5 crosses the entire Mtv-3 insertion site and only amplifies in the absence of Mtv-3, while primers 4 and 5 amplify the boundary between Mtv-3 and hitchhiker. In BXSB, both primer pairs produced amplicons of the expected size, while in B10, only primers 17 and 5 amplified the DNA (Figs 2B and 4), indicating that BXSB has both species, whereas B10 only has hitchhiker without Mtv-3 insertion. Thus, in BXSB this locus can be divided into three regions (Fig. 5): region 1, normal sialyltransferase; region 2, hitchhiker–(variant) sialyltransferase; and region 3, Mtv-3–hitchhiker–(variant) sialyltransferase. In B10 mice, region 3 is absent.
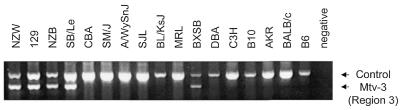
Figure 4.
PCR analysis for the presence of Mtv-3 (region 3). PCR was performed using primers 4 and 5, together with control primers, indicating the presence or absence of region 3, respectively.
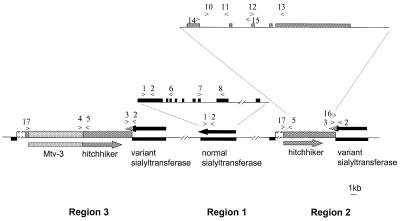
Figure 5.
Schematic representation of the genomic organisation of the locus containing the sialyltransferase gene. The sialyltransferase gene, hitchhiker sequence and Mtv-3 are represented by thick bars, and their transcripts by arrowed thick bars. Exon–intron organisation is illustrated for the sialyltransferase and hitchiker loci. Primers and their orientation are indicated by numbered arrowheads (Table 1). The differences between the 3′ UTRs of the normal and variant form of the sialyltransferase gene are indicated by shading of the arrowheads.
To study this variation in more detail, we examined a number of common mouse strains in the same way (Figs 2 and 4). We found that mouse strains can be classified into three categories dependent on which combination of regions 1–3 they contain. Thus, BXSB, NZW, 129, NZB and SB/Le have all three regions; B10, SM/J, SJL, C57BL/KsJ, MRL, B6 and AKR have both regions 1 and 2 while BALB/c, CBA, A/WySnJ, DBA and C3H have just region 1. These data resolve a controversy surrounding the 129 strain, since a previous study (22) identified both intact hitchhiker sequence (region 2) and hitchhiker with an Mtv-3 insertion (region 3). This was thought to be due to a contaminated BAC library. However, our data demonstrates that, in fact, both sequences are present in the 129 genome.
To determine whether regions 1–3 were linked together, we performed genomic mapping using a radiation hybrid panel (derived from 129 mice). Mtv-3 is known to be located on chromosome 11 (22), very close to the thymidine kinase 1 (Tk1) gene (78 cM) which was used as the dominant selectable marker in the construction of the radiation hybrid panel. Primers were used specific to each region, primers 4 and 5 for Mtv-3 (region 3), primers 5 and 17 for hitchhiker–sialyltransferase (region 2) and primers 1 and 2 for normal sialyltransferase (region 1). Normally, approximately 20–30 of the 100 hybrid cell lines tested would be expected to test positive in such a mapping analysis. In this analysis, region 1 was positive in 89 of the hybrid cell lines, region 2 was positive in 78 and region 3 was positive in 63. This is powerful evidence that all three are closely linked to each other and to Tk1. All of the radiation hybrid cell lines positive in regions 2 and 3 are also positive in region 1, while regions 2 and 3 only share 52 positive cell lines. Therefore, the likely genomic organisation would place region 1 between regions 2 and 3 in the telomeric region of chromosome 11, with a possible gene order of region 2–Tk1–region 1–region 3.
Amplification of the genome containing the loci Mtv-3, hitchhiker and ST6GalNac1
When we performed genomic PCR on sialyltransferase and the hitchhiker gene, a surprising result was observed. Although equal amounts of genomic DNA were used as templates, the PCR products in BXSB always gave a stronger signal than B10. We suspected that the genomic region containing these genes in BXSB was amplified. To test this hypothesis, we used PCR products (primers 14 and 15 for hitchhiker, and 7 and 8 for sialyltransferase gene) as probes to hybridise to BamHI-digested genomic DNA from BXSB and B10 (Fig. 6A and B). Both loci showed significant amplification in BXSB. Subsequent analyses have revealed that all mouse strains tested which contained Mtv-3 have genomic amplification of the hitchhiker and sialyltransferase genes. In addition, the AKR mouse showed genomic amplification of these genes in the absence of Mtv-3 insertion.
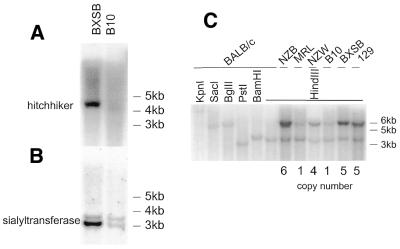
Figure 6.
Southern blot analysis of DNA from different mouse strains. (A) Genomic DNA from BXSB and B10 was digested with BamHI, transferred to nylon membrane and hybridised with a radiolabelled hitchhiker gene probe generated by PCR using primers 14 and 15. (B) The same Southern blot shown in (A) was rehybridised with a sialyltransferase gene probe generated from a PCR product amplified using primers 7 and 8. The upper bands represent the normal form of the gene while the lower bands are the variant form of the gene. (C) The same probe used in (B) was hybridised to a blot of KpnI, SacI, BglII, PstI, BamHI and HindIII digested BALB/c DNA, and HindIII digested NZB, MRL, NZW, B10, BXSB and 129 DNA. A single band was detected in BALB/c DNA regardless of the enzyme used, indicating the presence of a single copy of the sialyltransferase gene. The upper bands in other strains, representing the variant form of the gene, were analysed by densitometry to obtain copy numbers (indicated on the figure). The lower band was used as a single copy control.
To determine the copy number in this region, we performed a series of Southern hybridisations using as probe a 1 kb fragment from the sialyltransferase gene, the PCR product of primers 7 and 8. DNA from BALB/c mice always gave one restriction fragment, regardless of which enzyme was used (Fig. 6C). This implied the presence of a single copy of the normal sialyltransferase gene in BALB/c mice, supporting our previous data. However, using HindIII as enzyme, other DNA samples gave two bands, one at ∼3.5 kb, corresponding to the normal form of the gene (present in BALB/c), and an additional fragment from the variant form of the gene at 6 kb. Copy number was therefore derived by comparing the intensity of the two bands, assuming the 3.5 kb fragment represented a single copy of the normal form of the sialyltransferase gene. BXSB showed a 5-fold increase in copy number, while there was a 4-fold increase in NZW, 6-fold in NZB, with B10 and MRL showing no amplification.
DISCUSSION
The analysis of differentially expressed genes is a powerful technique to identify potential disease susceptibility loci. A study on synovial tissue from rheumatoid patients and lower intestinal mucosa from patients with Crohn’s disease, using microarrays of candidate cDNA sequences, verified the involvement of many genes in these diseases, but also revealed the novel participation of IL-3, amongst others, in both disease processes (25). In this study, we identified a number of genes that are overexpressed in lupus-prone BXSB spleen compared with non-autoimmune B10 spleen. The majority of sequences identified were Ig-related and retroviral proteins, both of which are known to be overexpressed in SLE. Due to the subtractive nature of the RDA analysis, these sequences overwhelmed the data and prevented the identification of many novel sequences; a serious drawback of this approach, which would not occur with the use of microarrays. However, a number of sequences were identified which were overexpressed in the BXSB spleen and thymus. Of these, the most interesting has proven to be an extended locus on chromosome 11, which varies between different mouse strains and is weakly linked to the lupus phenotype.
Interestingly, approximately the same interval has also been linked to the lupus phenotype in the MRL mouse (26), with the gene lying in the distal region of chromosome 11, lending support to a role for this region in disease. The function of hitchhiker transcripts is unknown; however, the sialyltransferase and Mtv superantigen are immunologically relevant genes. Sialyltransferases catalyse the transfer of sialic acid to the terminal positions of glycoproteins and glycolipids. Sialic acids are thought to play an important role in the regulation of the immune response (reviewed in 27). In most circumstances, the insertion of a retrovirus into immunological or apoptotic genes will have deleterious effects, which may result in disease (28,29). However, linkage to a disease phenotype in this study is weak and it is difficult to determine whether this region is playing a role in disease aetiology.
Combining data from DNA sequence analysis, Southern hybridisations and PCR, we have characterised the genomic structure of this locus in different mouse strains (Fig. 5). The evolution of these different organisations probably involved two separate insertion processes and several duplications. First, the sialyltransferase gene (region 1) underwent a duplication event, simultaneously with an insertion of the MuLV-related hitchhiker into the 3′ UTR of the sialyltransferase gene. We have found no example of a duplicated normal sialyltransferase gene and therefore presume these events took place together. However, it is possible that the insertion event promotes the duplication, as has apparently happened with the second insertion involving Mtv-3. The fact that the AKR strain has multiple copies of sialyltransferase–hitchhiker (region 2) supports a role for insertion in promoting amplification. The second insertion event involved the integration of the MMTV-related virus, Mtv-3, into the beginning of the hitchhiker sequences. The AKR strain provides support for the duplication taking place before insertion, rather than simultaneously. This second insertion event may have resulted in significant amplification in BXSB, 129, NZB, NZW and SB/Le mouse strains. Thus, this region of the genome has supported multiple insertion/amplification events, which may be mutually causative, and leads us to propose that this is an unstable region of the mouse genome.
For the sialyltransferase gene, the two forms of the gene are differentially expressed in different strains. Indeed, it was this variation that contributed to the identification of these sequences in the RDA analysis. However, the expression level, as determined by RT–PCR, was not proportional to the copy number of each gene within the mouse genome, since in 129 and BXSB mice there were similar numbers of copies of the variant form, but the expression levels of each form varied in the two mouse strains. Based on preliminary data that expression of the normal gene does not appear to vary, regardless of the genomic organisation, we suggest that the expression of the variant form is differentially regulated in the different strains; e.g. upregulated in BXSB and downregulated in B10 and 129 mice. However, this will need to be confirmed in future studies.
The reasons for the production of unspliced or incomplete transcripts of sialyltransferase and hitchhiker sequences are currently unknown. However, retroviruses have the ability to express cytoplasmic mRNAs that retain one or more introns. Because the hitchhiker sequences were retroviral in origin, the presence of high levels of unspliced RNAs from the hitchhiker and sialyltransferase genes might have some connection with the ancient virus sequence. A comparison between the constitutive transport element (CTE) of Mason-Pfizer monkey virus (MPMV), a sequence known to direct the export of viral RNAs to the cytoplasm (30,31), and the hitchhiker and sialyltransferase gene sequences revealed weak homologies (65% identity) between the CTE and two sequences in the complex: one of the hitchhiker introns (nucleotides 1951–2091; numbering as in ref. 22); and the 3′ UTR of the variant form of sialyltransferase gene (nucleotides 8535–8675). Significantly, no homology to the CTE was apparent in the normal form of sialyltransferase, from which no unspliced mRNAs were derived. If this were indeed the case it would be, to the best of our knowledge, the first demonstration of retrovirally-directed production of unspliced RNAs from cellular genes.
In summary, using cDNA RDA we have isolated several overexpressed genes in BXSB thymus and spleen and have subsequently identified a previously undescribed unstable genomic region, at the distal end of mouse chromosome 11, which has undergone repeated insertions and amplification. We have demonstrated that this region and its expression pattern varies in different mouse strains, and we have found limited linkage (maximum P = 0.009) of this unstable region to the lupus phenotype. It is possible that viral insertion into somatic genes and subsequent duplication/amplification of them, might play a role in the evolution of complex genomes and in the development of disease.
Acknowledgments
ACKNOWLEDGEMENT
This work was supported by the Arthritis Research Campaign UK.
DDBJ/EMBL/GenBank accession no. AF351128
REFERENCES
- 1.Wakeland E.K., Wandstrat,A.E., Liu,K. and Morel,L. (1999) Genetic dissection of systemic lupus erythematosus. Curr. Opin. Immunol., 11, 701–707. [DOI] [PubMed] [Google Scholar]
- 2.Vyse T.J. and Kotzin,B.L. (1998) Genetic susceptibility to systemic lupus erythematosus. Annu. Rev. Immunol., 16, 261–292. [DOI] [PubMed] [Google Scholar]
- 3.Tsao B.P. (2000) Lupus susceptibility genes on human chromosome 1. Int. Rev. Immunol., 19, 319–334. [DOI] [PubMed] [Google Scholar]
- 4.Kono D.H. and Theofilopoulos,A.N. (2000) Genetic studies in systemic autoimmunity and aging. Immunol. Res., 21, 111–122. [DOI] [PubMed] [Google Scholar]
- 5.Yasutomo K., Horiuchi,T., Kagami,S., Tsukamoto,H., Hashimura,C., Urushihara,M. and Kuroda,Y. (2001) Mutation of DNASE1 in people with systemic lupus erythematosus. Nature Genet., 28, 313–314. [DOI] [PubMed] [Google Scholar]
- 6.Murphy E.D. and Roths,J.B. (1979) A Y chromosome associated factor in strain BXSB producing accelerated autoimmunity and lymphoproliferation. Arthritis Rheum., 22, 1188–1194. [DOI] [PubMed] [Google Scholar]
- 7.Theofilopoulos A.N. and Dixon,F.J. (1985) Murine models of systemic lupus erythematosus. Adv. Immunol., 37, 269–389. [DOI] [PubMed] [Google Scholar]
- 8.Hogarth M.B., Slingsby,J.H., Allen,P.J., Thompson,E.M., Chandler,P., Davies,K.A., Simpson,E., Morley,B.J. and Walport,M.J. (1998) Multiple lupus susceptibility loci map to chromosome 1 in BXSB mice. J. Immunol., 161, 2753–2761. [PubMed] [Google Scholar]
- 9.Haywood M.E.K., Hogarth,M.B., Slingsby,J.H., Rose,S.J., Allen,P.J., Thompson,E.M., Maibaum,M.A., Chandler,P., Davies,K.A., Simpson,E., Walport,M.J. and Morley,B.J. (2000) Identification of intervals on chromosomes 1, 3 and 13 linked to the development of lupus in BXSB mice. Arthritis Rheum., 43, 349–355. [DOI] [PubMed] [Google Scholar]
- 10.Boehm T. (1998) Positional cloning and gene identification. Methods, 14, 152–158. [DOI] [PubMed] [Google Scholar]
- 11.Pritchard N.R., Cutler,A.J., Uribe,S., Chadban,S.J., Morley,B.J. and Smith,K.G. (2000) Autoimmune-prone mice share a promoter haplotype associated with reduced expression and function of the Fc receptor FcgammaRII. Curr. Biol., 24, 227–230. [DOI] [PubMed] [Google Scholar]
- 12.Aitman T.J., Glazier,A.M., Wallace,C.A., Cooper,L.D., Norsworthy,P.J., Wahid,F.N., Al-Majali,K.M., Trembling,P.M., Mann,C.J., Shoulders,C.C., Graf,D. et al. (1999) Identification of Cd36 (Fat) as an insulin-resistance gene causing defective fatty acid and glucose metabolism in hypertensive rats. Nature Genet., 21, 76–83. [DOI] [PubMed] [Google Scholar]
- 13.St Croix B., Rago,C., Velculescu,V., Traverso,G., Romans,K.E., Montgomery,E., Lal,A., Riggins,G.J., Lengauer,C., Vogelstein,B. and Kinzler,K.W. (2000) Genes expressed in human tumor endothelium. Science, 289, 1197–1202. [DOI] [PubMed] [Google Scholar]
- 14.Liang P. and Pardee,A.B. (1992) Differential display of eukaryotic messenger RNA by means of the polymerase chain reaction. Science, 257, 967–971. [DOI] [PubMed] [Google Scholar]
- 15.Lisitsyn N., Lisitsyn,N. and Wigler,M. (1993) Cloning the differences between two complex genomes. Science, 259, 946–951. [DOI] [PubMed] [Google Scholar]
- 16.Diatchenko L., Lau,Y.F., Campbell,A.P., Chenchik,A., Moqadam,F., Huang,B., Lukyanov,S., Lukyanov,K., Gurskaya,N., Sverdlov,E.D. and Siebert,P.D. (1996) Suppression subtractive hybridization: a method for generating differentially regulated or tissue-specific cDNA probes and libraries. Proc. Natl Acad. Sci. USA, 93, 6025–6030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Hubank M. and Schatz,D.G. (1994) Identifying differences in mRNA expression by representational difference analysis of cDNA. Nucleic Acids Res., 22, 5640–5648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Suen K.L., Bustelo,X.R., Pawson,T. and Barbacid,M. (1993) Molecular cloning of the mouse grb2 gene: differential interaction of the Grb2 adaptor protein with epidermal growth factor and nerve growth factor receptors. Mol. Cell. Biol., 13, 5500–5512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Choubey D., Snoddy,J., Chaturvedi,V., Toniato,E., Opdenakker,G., Thakur,A., Samanta,H., Engel,D.A. and Lengyel,P. (1989) Interferons as gene activators. Indications for repeated gene duplication during the evolution of a cluster of interferon-activatable genes on murine chromosome 1. J. Biol. Chem., 264, 17182–17189. [PubMed] [Google Scholar]
- 20.Mitchell T.J., Naughton,M., Norsworthy,P., Davies,K.A., Walport,M.J. and Morley,B.J. (1996) IFN-gamma up-regulates expression of the complement components C3 and C4 by stabilization of mRNA. J. Immunol., 156, 4429–4434. [PubMed] [Google Scholar]
- 21.Church G.M. and Gilbert,W. (1984) Genomic sequencing. Proc. Natl Acad. Sci. USA, 81, 1991–1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Stegalkina S.S., Guerrero,A., Walton,K.D., Liu,X., Robinson,G.W. and Hennighausen,L. (1999) Transcription originating in the long terminal repeats of the endogenous mouse mammary tumor virus Mtv-3 is activated in Stat5a-null mice and picks up hitchhiking exons. J. Virol., 73, 8669–8676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Fairchild S., Knight,A.M., Dyson,P.J. and Tomonari,K. (1991) Co-segregation of a gene encoding a deletion ligand for Tcrb-V3+ T cells with Mtv-3. Immunogenetics, 34, 227–230. [DOI] [PubMed] [Google Scholar]
- 24.Kurosawa N., Takashima,S., Kono,M., Ikehara,Y., Inoue,M., Tachida,Y., Narimatsu,H. and Tsuji,S. (2000) Molecular cloning and genomic analysis of mouse GalNAc alpha 2,6-sialyltransferase (ST6GalNAcI). J. Biochem. (Tokyo), 127, 845–854. [DOI] [PubMed] [Google Scholar]
- 25.Heller R.A., Schena,M., Chai,A., Shalon,D., Bedilion,T., Gilmore,J., Woolley,D.E. and Davis,R.W. (1997) Discovery and analysis of inflammatory disease-related genes using cDNA microarrays. Proc. Natl Acad. Sci. USA, 94, 2150–2155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gu L., Weinreb,A., Wang,X.P., Zack,D.J., Qiao,J.H., Weisbart,R. and Lusis,A.J. (1998) Genetic determinants of autoimmune disease and coronary vasculitis in the MRL-lpr/lpr mouse model of systemic lupus erythematosus. J. Immunol., 161, 6999–7006. [PubMed] [Google Scholar]
- 27.Pilatte Y., Bignon,J. and Lambre,C.R. (1993) Sialic acids as important molecules in the regulation of the immune system: pathophysiological implications of sialidases in immunity. Glycobiology, 3, 201–218. [DOI] [PubMed] [Google Scholar]
- 28.Watanabe-Fukunaga R., Brannan,C.I., Copeland,N.G., Jenkins,N.A. and Nagata,S. (1992) Lymphoproliferation disorder in mice explained by defects in Fas antigen that mediates apoptosis. Nature, 356, 314–317. [DOI] [PubMed] [Google Scholar]
- 29.Wu J., Zhou,T., He,J. and Mountz,J.D. (1993) Autoimmune disease in mice due to integration of an endogenous retrovirus in an apoptosis gene. J. Exp. Med., 178, 461–468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Cullen B.R. (1998) Retroviruses as model systems for the study of nuclear RNA export pathways. Virology, 249, 203–210. [DOI] [PubMed] [Google Scholar]
- 31.Kiss-Laszlo Z. and Hohn,T. (1996) Pararetro- and retrovirus RNA: splicing and the control of nuclear export. Trends Microbiol., 4, 480–485. [DOI] [PubMed] [Google Scholar]