Abstract
Single-cell technologies, particularly single-cell RNA sequencing (scRNA-seq) methods, together with associated computational tools and the growing availability of public data resources, are transforming drug discovery and development. New opportunities are emerging in target identification owing to improved disease understanding through cell subtyping, and highly multiplexed functional genomics screens incorporating scRNA-seq are enhancing target credentialling and prioritization. ScRNA-seq is also aiding the selection of relevant preclinical disease models and providing new insights into drug mechanisms of action. In clinical development, scRNA-seq can inform decision-making via improved biomarker identification for patient stratification and more precise monitoring of drug response and disease progression. Here, we illustrate how scRNA-seq methods are being applied in key steps in drug discovery and development, and discuss ongoing challenges for their implementation in the pharmaceutical industry.
Subject terms: Computational biology and bioinformatics, Gene expression profiling, Target identification
There have been significant recent advances in the development of single-cell technologies, providing remarkable opportunities for drug discovery and development. Here, Ferran and colleagues discuss how single-cell technologies, primarily single-cell RNA sequencing methods, are being applied in the drug discovery pipeline, from target identification to clinical decision-making. Ongoing challenges and potential future directions are discussed.
Introduction
Drug discovery is generally an inefficient process characterized by rising costs1,2, long timelines3 and high rates of attrition4. These inefficiencies are partly rooted in our limited understanding of human biology, in particular, disease-related mechanisms, actionable therapeutic targets and disease response heterogeneity5,6. The lack of sufficiently representative preclinical models, and the limitations of necessarily reductionist disease models, compound the challenges of understanding human systems.
Before single-cell (SC) approaches, cell and tissue characteristics could only be assessed in bulk and from relatively large amounts of starting material. Amplification-based techniques, such as microarrays, bulk RNA sequencing (RNA-seq) and quantitative PCR with reverse transcription (qRT–PCR)7, measured mRNA transcripts in pools of cells and could not distinguish relevant signals from heterogeneous subpopulations or rare cell types. Techniques capable of SC resolution, such as fluorescence-activated cell sorting (FACS), immunohistochemistry and cytometry by time of flight (CyTOF), were limited by the relatively small scale of testable targets and the need for a priori biological insights to enable experimental design8–10.
SC technologies that have been developed in the past decade (reviewed in refs. 11–13) have made significant inroads towards resolving some of these limitations, while at the same time being complementary to bulk applications that are still commonly used. Among the growing range of technologies, single-cell RNA sequencing (scRNA-seq; Box 1) has advanced substantially14,15 since the demonstration of whole-transcriptome profiling from a single cell in 2009 (ref. 16), and has reached the point where it is being applied in the pharmaceutical industry to investigate key questions in drug discovery and development (Fig. 1). Consequently, scRNA-seq is the focus of this article. SC technologies that extend beyond mRNA to DNA, epigenetic, proteomic and other features17 are also highlighted.
Fig. 1. How single-cell sequencing can inform decisions across the drug discovery and development pipeline.
Single-cell technologies are being applied to answer key questions at various stages in the drug discovery and development pipeline. These applications are anticipated to increase the probability of success in the clinic by improving the quality of both the drug candidates emerging from discovery programmes and the clinical development plans for those drug candidates in stratified disease populations.
The rapid and simultaneous development of scalable plate-based and microfluidic-based methods capable of profiling large numbers of single cells has enhanced the utility of SC techniques for industrial-scale applications. Novel computational techniques and other methods (Fig. 2; Supplementary Table 1; Boxes 2 and 3) have also played a key part in leveraging SC data, supported by a growing user community that has helped to improve public data access and generate best practices. The combination of SC profiling platforms and sophisticated computational methods is driving step-change improvements in our knowledge of disease biology and pharmacology. For example, the availability of SC sequencing data for animal model systems is improving our understanding of translatability to humans18. ScRNA-seq has enabled identification of molecular pathways that allow prediction of survival19, response to therapy20, likelihood of resistance21,22 and candidacy for alternative intervention23. Further capabilities provided by SC technologies include the identification of novel cell types24 and subtypes25, the refinement of cell differentiation trajectories and the dissection of heterogeneously manifested human traits26 or constituent cell types that compose multicellular organs or tumours27.
Fig. 2. Computational methods used in single-cell data analysis for drug discovery and development.
Representation of the computational tools and/or methods (see Supplementary Table 1 for further details and URLs for the various tools), currently used by pharmaceutical companies for data handling and to probe biological insights through cell-type annotation to reveal genotype and/or phenotype and functional assignment. B cell receptor; CNV, copy number variation; eQTL, expression quantitative trait loci; scATAC-seq, single-cell sequencing assay for transposase-accessible chromatin; scDNA-seq, single-cell DNA sequencing; scRNA-seq, single-cell RNA sequencing; SNV, single-nucleotide variant; ST, spatial transcriptomics; TCR, T cell receptor.
In this Review, we illustrate how SC technologies, primarily scRNA-seq methods, are being applied in the various steps of the drug discovery pipeline, from target identification to clinical decision-making. Ongoing challenges related to study design and data accessibility are also highlighted, as well as potential future directions for the use of SC techniques in drug discovery and development.
Box 1 Fundamentals of single-cell RNA sequencing.
A typical single-cell RNA sequencing (scRNA-seq) workflow has three key phases: library generation, pre-processing and post-processing. The library generation process includes the isolation of individual cells or nuclei, mRNA capture and sequencing (see figure). Once sequences are obtained, the subsequent steps are computational. Pre-processing includes the initial analyses to count and clean the data. In post-processing, dimensionality is reduced, gene signatures and cell types are identified, and visualizations may be generated. Data integration and batch correction are optional steps, and ultimately may support the inference analyses. All or a subset of these steps are often performed iteratively to optimize outcomes. Key phases in the typical scRNA-seq workflow are described in more detail below and illustrated in Supplementary Fig. 1.
Library generation and sequencing
Library generation transforms cells or nuclei into sequencer-ready samples. Sample preparation is a crucial step, which often requires tissue dissociation with mechanical or enzymatic stress, depending on sample type. This unavoidably releases RNA into the suspension, contributing to high background or noise if not removed during data processing. Fresh samples are ideal for high-quality scRNA-seq, and single-nucleus RNA sequencing is usually preferable when samples must be frozen.
Samples are then separated into reaction chambers for lysis and RNA capture, most commonly using 10X Chromium technology, which combines an aqueous flow of cells, barcoded primers carried in beads, lysis buffer and reverse transcription enzymes with oil to create microdroplet reaction chambers. Plate-based technologies perform this step in microwells, and automated microfluidic devices use other forms of microchamber. The common feature is that individual cells must be trapped within a space that is not continuous with spaces containing any other individual cells.
Next, the RNA transcripts of each cell are tagged with a barcoded unique molecular identifier (UMI), to help distinguish between cell transcripts and extraneous PCR amplicons generated during processing. A cDNA library is created by reverse transcription and amplified; depending on the tagging strategy, multiple amplification steps may be needed, and adapter sequences that bind to the flow cell may be ligated as well. The cDNA is then processed, similarly to bulk RNA-seq techniques, by fragmentation to create a homogeneously sized pool of molecules and the addition of index sequences useful for the identification of read origin (for example, to allow multiplexing). Like any sequencing protocol, this workflow contains several purification and quantification steps to ensure high quality. Multiple samples, with different indices, are finally loaded onto a flow cell and sequenced.
Sequence data pre-processing
Reads from plate-based technologies (for example, SMART-seq2 (ref. 201)) can be analysed by traditional bulk genome or transcriptome alignment and quantification pipelines. Droplet-based platforms require specific tools to handle highly cell-multiplexed data to correctly assign UMI counts to cell barcodes. For all methods, an RNA capture rate of between 10% and 20% is common and must be accounted for during analysis202.
The Cell Ranger pipeline from 10X Genomics is widely used to process 10X data. It is based on the STAR method for RNA-seq alignment and offers additional features such as cell counting and quality control summary reporting203. Academic efforts strengthened by the open-source community provide more recent solutions such as STARsolo204, Alevin205 and Kallisto-BUStools206,207.
For all platforms, the next steps are to determine counts for each gene in each cell to generate a cell-by-gene matrix. For processing in droplet platforms, pre-emptive filtering to distinguish cells from empty droplets may first be applied208,209. Further filtering of ambient RNA210,211 and/or methods for removing doublets are also used212–214, and together help to clean the data and reduce data volume. The matrix is then normalized to take into account discrepancies in RNA capture for each cell215–217 and finally, highly variable genes in a sample may be flagged for downstream analysis.
Sequence data post-processing
Downstream of matrix generation and normalization, typical scRNA-seq workflows include unsupervised clustering218 to group together cells with similar expression profiles, and dimensionality reduction, via methods such as t-distributed stochastic neighbour embedding (t-SNE)219 or uniform manifold approximation and projection (UMAP)220 that enable visualization of cell clustering in a 2D or 3D space. Marker genes associated with each cluster are detected via differential expression analysis. Cell-type annotation methods, integrative analysis to correct batch effects, trajectory mapping to trace cell differentiation and cell communication analysis can provide additional insights. Downstream analyses may need to be iteratively performed to optimize the analyses.
Box 2 Other single-cell technologies.
Single-cell CRISPR screening technologies: pooled CRISPR screening is an efficient and scalable approach to drug-target discovery but is restricted to low-content readouts and can only identify genes yielding distinct phenotypes. To overcome this, single-cell (SC) CRISPR screening technologies such as Perturb-seq84,86,221,222 were developed, coupling pooled CRISPR screening with single-cell RNA sequencing (scRNA-seq) or SC multi-omics. Several computational frameworks (MIMOSCA84, scMAGeCK223, MUSIC224, Mixscape222) and a screening platform85 allow decoding of the effect of individual perturbations on gene expression, their interactions or their cell-state dependence and prioritization of the cell types most sensitive to CRISPR-mediated perturbations at a SC level.
Single-cell DNA sequencing technologies: these have been mainly used to infer cell lineage of cancers and to track cells with treatment-resistant mutations. To overcome technical limitations such as non-uniform coverage depth in scRNA-seq, several computational methods225–230 have been developed for the identification of single-nucleotide variants (SNVs); short insertions and deletions (indels) and copy number variation (CNV). CNV detection methods for other technologies (for example, array-CGH, single-nucleotide polymorphism (SNP) arrays and whole-genome sequencing (WGS) or whole-exome sequencing (WES)) were also extended and applied to scDNA-seq data231. However, scWGS is still very expensive. Therefore, computational methods such as CopyKat232 and InferCNV233 have been developed to characterize copy number and intratumoural heterogeneity using scRNA-seq data instead. These methods are also used to infer aneuploidy in cells from scRNA-seq cancer data sets to better delineate host from cancer cells. In addition, scRNA-seq-based point mutation detection approaches234,235 allow linkage of genotype to phenotype and make it possible to detect functional mutations that drive cell-type-specific gene expression. Best practices for mapping of single-cell expression quantitative trait loci (sc-eQTL) have been assessed236.
SC T cell receptor and B cell receptor sequencing technologies: scTCR-seq and scBCR-seq help to investigate the dynamics of T cell or B cell clones in tissues or peripheral blood by determining T cell or B cell clonotypes at a SC level. Cells from the adaptive immune system originating from a common ancestor and therefore sharing the same TCR or BCR are called clonotypes. Alternatively, TCR and BCR repertoire reconstruction and clonality inference can be made based on scRNA-seq by using computational methods237–241. The clonotype dynamics can be examined by using computational tools such as scRepertoire242 and CellaRepertorium243. Coupling scTCR-seq or scBCR-seq with scRNA-seq can reveal the relationship between clonotype and phenotype (or transcriptional states) in T or B cell populations244. Detailed characterization of T and B cells provided by SC technologies has helped in understanding of disease (for example, cancer microenvironment, multiple sclerosis antigens, etc.) and in improving of engineered T cell therapies such as chimeric antigen receptor (CAR) T cells.
SC epigenetics: various SC technologies capture epigenetic characteristics at near-nucleotide resolution. SC open chromatin structure (that is, transposase-accessible) can be revealed by single-cell sequencing assay for transposase-accessible chromatin (scATAC-seq)245, chromatin histone modifications by scCUT&Tag246 or scChIP–seq247, and DNA methylation patterns by scBS-seq248. Understanding promoters and enhancers that are activated in a certain cell type or state can help in identifying tissues, cell types and/or biological conditions in which a target is more abundantly expressed and the transcriptional programmes that lead to expression of the target. Moreover, these techniques help to identify causal non-coding variants associated with a disease discovered by genome-wide association studies (GWAS) and map them to a specific cell type.
SC proteomics methods: emerging SC proteomics methods decode the variation of the proteome across individual cells249. SC proteomics (sc-proteomics — see reviews250,251) methods typically focus on either absolute quantification of a small number of proteins or on highly multiplexed protein measurements. A method has been recently proposed for counting single proteins in single cells, based on nanopore single-molecule peptide reads, that is sensitive to single-amino acid substitutions within individual peptides252. This method opens the opportunity to develop single-molecule protein fingerprinting in the future.
SC multi-omics technologies: technologies such as ECCITE-seq97, scNMT-seq161 and DOGMA-seq253 now allow for the simultaneous measurement of different readouts (for example, RNA expression, surface protein expression, clonotypes, DNA methylation and/or chromatin accessibility) from the same single cells.
Emerging SC technologies and methods: methods for SC microRNA254 and SC long non-coding RNA (see review255) have expanded RNA transcriptomic profiling. SC metabolomics (sc-metabolomics) techniques were proposed for cataloguing the chemical contents of a single cell or even a single organelle256. scRibo-seq, for SC ribosomal profiling, opens the possibility to explore translation at SC level. Integrated with a machine learning approach, this method achieves single codon resolution257. Methods such as scSPRITE258 and Higashi259 allow detection of high-order 3D genome structures in single cells (scHi-C).
Spatially resolved omics approaches: SC technologies lose spatial information during the tissue dissociation step. Spatially resolved omics approaches have been recently developed260–262 to recover the spatial context. Excellent reviews on spatial transcriptomics and associated computational methods are available263–266.
Box 3 Computational methods used to infer insights from single-cell RNA sequencing data sets.
Single-cell (SC) sequence data pre-processing is required before insights can be generated from a SC data set. Once a gene expression matrix has been generated several methods exist to provide answers to relevant research questions. This box highlights pre-processing methods, focusing on areas of active development and concern.
Methods for addressing sparsity in scRNA-seq data sets: single-cell RNA sequencing (scRNA-seq) data sets are sparse in that many counts in the gene expression matrix are zero, that is, not a single RNA molecule is detected for those genes. The source of this higher prevalence of zeros in comparison with bulk samples is diverse. Biological sources of sparsity in a data set are mainly driven by absent gene expression in the various cell types captured in a sample. In addition, gene expression is a stochastic process, which also contributes to a higher frequency of zero read counts. Technical sources of sparsity are inefficiencies in mRNA capture and/or sampling effect owing to limited sequencing depth. How to deal with these challenges is under discussion and ranges from using appropriate statistical models, for example, zero-inflated Poisson models, to use of imputation techniques. This topic is nicely reviewed in ref. 267.
Batch-effect correction and data integration methods: SC data from large-scale or multiple studies are frequently generated by multiple institutions and/or in different experimental conditions. Two recent papers268,269 comprehensively evaluate the performance of batch correction, that is, removing the variability in the data due to technical or other less relevant variables, and data integration methods, that is, methods that combine several data sets in an embedded space or provide a common expression matrix. These tools help to facilitate integrative analyses of SC data from various sources. However, the application of batch correction methods to SC data from heterogeneous diseases (for example, tumours) may risk obscuring true biological signals. Proper experimental planning is important and directly empowers these tools270.
Single-cell multi-omics analysis: joint analysis of SC multi-omics data enhances the ability to more deeply characterize cell types and states and their association with disease progression and drug effect251. Weighted nearest neighbour (WNN) analysis in Seurat183, CiteFuse252, MOFA+253 and totalVI183,253–256 have been developed to improve the ability to resolve cell states and fates by integration of multimodal SC data. When generated from different cells, such multimodal measurements are projected into a common latent space by computational methods such as LIGER255,257, and canonical correlation analysis (CCA) in Seurat256 to jointly model variation across sample groups and data modalities.
Cell-type annotation: for scRNA-seq data, cell-type annotation can be performed based on unsupervised cell clustering and marker genes identified per cluster. This approach is very labour intensive. To facilitate cell-type annotation, automated cell-type annotation tools have been developed including Seurat label transfer271, Garnett272, scmap273, SingleR274 Cell-ID275 and more recently CellTypist189.
Once a properly integrated, normalized and annotated data set is available, insights can be derived from these data sets using a wide variety of methods.
Trajectory inference or pseudo-time analysis: cells experience dynamic processes such as differentiation, response to treatment and disease evolution. A heterogeneous sample of cells represents a snapshot of cells in various phases of these processes. Trajectory inference (TI) is used to determine the pattern of such a dynamic process. Widely used TI computational tools include Monocle276, PAGA277, Slingshot278, STEMNET279 and Scorpius280. Most TI methods require prior understanding of the anticipated topology and careful design considerations169. These methods are different from RNA velocity281, which exploits the presence of unspliced mRNA to derive an estimate of the rate of change of gene expression. Many methods have expanded upon this technique: Velocyto281 and scVelo282. CellRank283 combines TI and RNA velocity techniques.
Pathway analysis tools: these provide cell-type specific functional annotation and new biological insights into disease and response to treatment. GSVA284 and single sample gene set enrichment analysis (ssGSEA)285 were designed for bulk RNA-seq but can be applied to scRNA-seq data for this purpose. Tools such as Pagoda2 (ref. 286) and Vision287 were developed for the characterization of cell-type specific transcriptional heterogeneity. They allow interactive analysis of large SC data sets and identification of intercellular relationships in disease or in response to treatment.
Cell–cell communication analysis: disease can be caused by disruptions in cell–cell communications288, and a growing collection of computational tools support drawing inferences about these disruptions183,289–294, generating new hypotheses and potentially enhancing disease understanding295.
Cell-type deconvolution methods: most clinical transcriptomics data are currently generated with either bulk RNA-seq or microarray. Cell-type deconvolution methods296–301 enable the estimation of cell-type composition based on gene signatures derived from scRNA-seq data and are especially useful in the drug development pipeline.
Methods of mapping disease-associated variants to scRNA-seq data sets: methods are emerging that integrate genetic cues from genome-wide association studies (GWAS) with SC phenotypic data sets such as transcriptomics. SC Linker combines GWAS summary statistics with SC transcriptomics to quantify the heritability of a gene expression signature derived from scRNA-seq data sets (capturing either a cell type or a biological process)81. Another method called scDRS looks for enrichment of polygenic GWAS-derived signatures in SC gene expression profiles182.
Applications in drug discovery and development
SC technologies can be applied throughout drug discovery and development (Fig. 1). Improved disease understanding gained through subtyping based on altered cell compositions and cell states can guide the identification of novel cellular and molecular targets. Target credentialling and validation can benefit from the use of SC sequencing in the identification of relevant preclinical models for a given disease subtype. Highly multiplexed functional genomics screens that merge CRISPR and SC sequencing (scCRISPR screening; Box 2) can enhance target credentialling throughput and augment the perturbation readouts with mechanistic information to improve target prioritization. SC sequencing technologies can provide insights on cell-type-specific compound actions, off-target effects and heterogeneous responses to inform drug candidate selection. In clinical development, these technologies can contribute by helping to identify biomarkers for patient stratification, elucidating drug mechanisms of action or resistance, or monitoring drug responses and disease progression. Opportunities to characterize and improve engineered biologics and cell therapies using SC technologies are also emerging (Box 4).
Below, we discuss representative published studies that demonstrate how SC technologies, particularly scRNA-seq approaches, can be applied in key steps in drug discovery and development, with a focus on those that are widely used in the pharmaceutical industry.
Box 4 Single-cell analysis for biologics and cell therapies.
Monoclonal antibodies
Single-cell sequencing technologies can accelerate and improve therapeutic antibody identification and optimization. Charting the full antibody repertoire of an immunized animal, subsequently tracking its evolution during clonal selection, expansion and affinity maturation, and comparison with derived hybridoma cell lines at cellular resolution is enabled by high-throughput single-cell B cell receptor sequencing (scBCR-seq)302 (Box 2). These efforts can assist therapeutic antibody identification by expanding the available BCR repertoire, and may also improve the generation of diverse and large phage displays or the mining for therapeutic antibodies based on sequence similarity303. Moreover, technologies such as LIBRA-seq combine scBCR readouts with antigen specificity and thereby directly expedite lead discovery304. Finally, direct usage of the human B cell reservoir of convalescent donors as an antibody pool opens new avenues for the development of therapeutic monoclonal antibodies. This approach has been used to engineer neutralizing monoclonal antibodies for coronavirus disease 2019 (COVID-19)305.
CAR-T cell therapies
Chimeric antigen receptor (CAR)-T cell therapies have shown strong efficacy in the treatment of some B cell-originating haematological malignancies. Unfortunately, the toxicity induced by these treatments can be life-threatening, and efficacy is restricted to a subset of patients. Single-cell RNA sequencing (scRNA-seq) has been used as a complementary tool to investigate cellular heterogeneity and cell composition dynamics in the pre-treated patient peripheral blood mononuclear cell (PBMC) samples and post-CAR-T infusion time points306.
B cell maturation antigen (BCMA) CAR-T cells have demonstrated promising effects in patients with relapsed or refractory multiple myeloma. ScRNA-seq has been used to analyse the dynamics of BCMA CAR-T cells in a clinically successful case of relapsed or refractory primary plasma cell leukaemia (pPCL)307. At the peak phase, CAR-T cells were found to shift from a highly proliferative state to a highly cytotoxic state, finally changing to a memory-like state at remission phase.
Many SC studies focus on understanding factors that drive favourable outcomes in CAR-T cell therapies. In large B cell lymphoma (LBCL), complete response is associated with the increase in memory CD8+ T cells308. Multi-omic SC interrogation of T cells showed that interferon signalling controlled by IRF7 reduces persistence of CAR-T cells after treatment309. In parallel, efforts to better understand and control toxicity of these therapies are undertaken. In normal brain tissue, a small population of mural cells — which surround the endothelium and are crucial for blood–brain barrier integrity — were shown to express CD19 and are therefore potentially targeted by CD19 CAR-T cells310. These findings can explain the CAR-T cell-induced neurotoxicity, due to increased vascular permeability in the brain. Investigation of expression patterns of CD19 using human SC reference atlases such as the Human Cell Landscape (HCL), revealed potentially on-target off-tumour toxic effects of CD19 CAR-T cell treatment311.
Improvements in CAR-T cell therapy are also being explored using genome-wide genetic perturbation techniques. CRISPR perturbation studies revealed that knocking out TLE4 and IKZF2 (encoding Helios) in CAR-T cells boosted their antitumour efficacy312. In a different approach, OverCITE-seq, which overexpresses open reading frames (ORFs) in T cells in a high-throughput fashion, was developed and combined with SC transcriptomics and epitope profiling313. Applying this to CAR-T cells, the gene LTBR was discovered to increase resistance to exhaustion and to augment overall effector function f these cells.
Disease understanding
As most complex diseases involve multiple cell types, SC resolution can significantly advance disease understanding. ScRNA-seq captures differences in cell-type composition and changes in cellular phenotype that are characteristic of a pathological state. Moreover, the unbiased view of scRNA-seq can detect the presence of rare cell types that drive pathobiology.
SC technologies are providing detailed knowledge of underlying disease mechanisms, enabling the investigation of novel therapeutic approaches. Although an exhaustive review is outside the scope of this article, illustrative examples for cancer, neurodegenerative diseases, inflammatory and autoimmune diseases, as well as infectious diseases are presented.
Cancer
SC molecular phenotyping has been extensively used to understand cancer development. Notable examples include the application of SC technologies to identify the cell of origin or cells associated with prostate carcinogenesis, heterogeneous papillary renal cell carcinoma (pRCC) and Barrett’s oesophagus leading to oesophageal adenocarcinoma28–30.
ScRNA-seq has revealed extensive cellular and transcriptional cell-state diversity in cancer and enabled tracking of cancer cell heterogeneity. This has been combined with immunophenotyping techniques to provide a view of stromal–immune niches (ecosystems or ecotypes) with unique cellular composition characterizing different types of tumour. Certain ecotypes are sometimes associated with tumour initiation or progression, sensitivity or resistance to therapeutic agents or clinical outcome as demonstrated by the application of this approach to capture the heterogenicity of diffuse large B cell lymphoma, breast cancer, oesophageal squamous cell carcinoma tumours and papillary thyroid carcinoma31–34.
SC technologies such as Perturb-seq hold promise in the mapping of genotype to phenotype changes — not only for oncology but also in other diseases — by assessing the impact of rare and common human disease genetic variants. This has been applied to assess the phenotypic consequences of somatic coding variants in the oncogene KRAS and the tumour suppressor gene TP53 in an unbiased and high-throughput fashion35.
As the extensive transcriptional cell-state diversity found in cancer is often observed independently of genetic heterogeneity, many studies have investigated the epigenetic coding of malignant cell states. Understanding epigenetic mechanisms is vital as they may enable adaptation to challenging microenvironments and may contribute to therapeutic resistance. Multi-omics SC profiling (Box 2) has provided insights into intratumoural heterogeneity in glioma and identified epigenetic mechanisms that underlie gliomagenesis36,37.
Longitudinal studies provide insights into the biological mechanisms associated with tumour progression and fitness of polyclonal tumours. Most studies have been carried out using mouse models or patient-derived xenografts (PDXs). Examples of this approach include a longitudinal SC analysis of samples from a myeloma mouse model that led to the identification of the GCN2 stress response as a potential therapeutic target38, and multi-year time-series SC whole-genome sequencing (scWGS; Box 2) of breast epithelium and primary triple-negative breast cancer (TNBC) PDX, which revealed how clonal fitness dynamics was induced by TP53 mutations and cisplatin chemotherapy39.
SC studies have also improved understanding of metastasis. A Cas9-based, SC lineage tracer has been applied to study the rates, routes and drivers of metastasis in a lung cancer xenograft mouse model, revealing that metastatic capacity was heterogeneous, arising from pre-existing and heritable differences in gene expression, and uncovering a previously unknown suppressive role for KRT17 (ref. 40). This study demonstrated the power of tracing cancer progression at subclonal resolution and vast scale. Further, SC immune mapping of melanoma sentinel lymph nodes (SLNs) identified immunological changes that compromise anti-melanoma immunity and contribute to a high relapse rate41. The progressive immune dysfunction found to be associated with micro-metastasis in patients with stage I–III cutaneous melanoma may motivate new hypotheses for neoadjuvant therapy with potential to reinvigorate endogenous antitumour immunity42. A similar suppressed immune environment was observed in acral melanoma compared with that of cutaneous melanoma from non-acral skin43. Expression of multiple, therapeutically tractable immune checkpoints was observed, offering new options for clinical translation that may have been missed without SC approaches. Metastasis studies based on SC analysis of circulating tumour cells (CTCs) have also been carried out44,45. The spatial heterogeneity and the immune-evasion mechanism of CTCs in hepatocellular carcinoma (HCC) have been dissected using scRNA-seq44, identifying chemokine CCL5 as an important mediator of CTC immune evasion, and highlighting a potential anti-metastatic therapeutic strategy in HCC. Further, it was recently shown that the spread of breast cancer cells occurs predominantly during sleep. ScRNA-seq analysis of blood CTCs, which increase during rest in both patients and mouse models, revealed a marked upregulation of mitotic genes, exclusively during the resting phase, thus enabling metastasis proficiency45.
A step change in our understanding of cancer is anticipated from initiatives such as the Human Tumour Atlas Network (HTAN)46 established by the National Cancer Institute, the primary focus of which is to elucidate the evolution of cancer from its pre-malignant forms to the state of metastasis at SC and spatial resolution. HTAN will generate SC, multiparametric, longitudinal atlases and integrate them with clinical outcomes. This initiative has already resulted in studies that capture in detail tumour initiation and progression as demonstrated by the creation of a SC tumour atlas covering the transition of polyps to malignant adenocarcinoma in colorectal cancer (CRC)47.
Neurodegenerative diseases
Parkinson disease is caused by the degeneration of dopaminergic neurons in the substantia nigra48, but not all dopamine-producing neurons degenerate. SC genomic profiling of human dopamine neurons found that although there are ten transcriptionally defined dopaminergic subpopulations in the human substantia nigra, only one population selectively degenerates in Parkinson disease, and the transcriptional signature of this population is highly enriched for the expression of genes associated with Parkinson disease risk49. The vulnerability of this population of dopaminergic neurons may provide insights for potential therapeutic interventions.
A different approach was used to study somatic DNA changes in single Alzheimer disease neurons. By comparing more than 300 individual neurons from the hippocampus and the prefrontal cortex of patients with Alzheimer disease with matched controls using scWGS, genomic alterations implicating nucleotide oxidation in the impairment of neural function were identified50. This work provided a different perspective on disease evolution, suggesting that the known pathogenic mechanisms in Alzheimer disease may lead to genomic damage in neurons that can progressively impair their function.
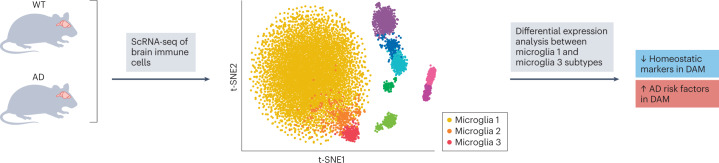
The role of immune cells in neurodegenerative diseases is posited in many recent studies. ScRNA-seq studies of brain tissues from both healthy mice and Alzheimer disease mouse models highlight disease-associated microglia, suggesting that a cell-state-targeting strategy may benefit patients with Alzheimer disease51 (Fig. 3). In addition, SC transcriptome and T cell receptor (TCR) profiling (Box 2) has revealed T cell compartments that are activated and expanded in Parkinson disease52.
Fig. 3. Single-cell RNA sequencing in disease understanding.
Single-cell RNA sequencing (scRNA-seq) reveals a novel microglia type in an Alzheimer disease (AD) mouse model. Unbiased clustering of single immune cells (CD45+) sorted from wild-type (WT) and AD mouse brains classified the cells into ten subpopulations, according to the expression patterns of the 500 most variable genes. The analysis thus allowed for de novo identification of rare subpopulations and revealed three microglia types: 1 (yellow), 2 (orange) and 3 (red). As the distinct microglia states of the orange and red clusters are found only in the AD model mice, they are called ‘disease-associated microglia’ (DAM). Microglia 1 cluster corresponds to homeostatic monocyte states found in both WT and AD. Differential expression analysis between DAM (microglia 3) and homeostatic microglia (microglia 1) from the AD mouse brain shows that DAMs are characterized by a significant downregulation of homeostatic markers and upregulation of several known AD risk factors. Microglia 2 is an intermediate Trem2-independent state between microglia 1 and microglia 3. t-Distributed stochastic neighbour embedding (t-SNE) map adapted with permission from ref. 51, Elsevier.
Novel SC technologies have been developed to study the brain. Examples include Patch-seq53,54 — a robust platform that combines scRNA-seq with patch clamp recording — and VINE-seq55, which is based on single-nucleus RNA sequencing (snRNA-seq). These approaches have been used to identify cell types in the neocortex that were selectively depleted in Alzheimer disease and to chart vascular and perivascular cell types at SC resolution in the human Alzheimer disease brain, respectively55,56.
Inflammatory and autoimmune diseases
ScRNA-seq was used to characterize a particular regulatory T cell present in spondyloarthritis57 and helped the discovery of cytotoxic T cells in the synovium in psoriatic arthritis. Clonal expansion of these synovial immune cells was demonstrated via complementary TCR-seq58. Differentiation of peripheral blood mononuclear cell (PBMC) samples of patients with anti-citrullinated peptide antibody-positive (ACPA+) and negative (ACPA−) rheumatoid arthritis at the SC level mapped immune correlates to each of these two different rheumatoid arthritis subtypes59, while profiling of the immune compartment of skin biopsies revealed that common dermatological inflammatory diseases each have distinct T cell resident memory, innate lymphoid cell and CD8+ T cell gene signatures59,60.
In multiple sclerosis, comparing PBMC samples at SC resolution from sets of twins discordant in multiple sclerosis revealed an inflammatory shift in a monocyte cluster, together with a subset of naive helper T cells that are IL-2-hyper-responsive in the multiple sclerosis cohort61. SC techniques have also helped to explain epidemiological evidence implicating Epstein–Barr virus (EBV) as a necessary aetiological factor in multiple sclerosis62. Using single-cell B cell receptor sequencing (scBCR-seq; Box 2) of both cerebrospinal fluid and blood from patients with multiple sclerosis revealed expansion of B cell clones in multiple sclerosis that bind a similar antigen in glia (GlialCAM) and EBV (EBNA1)63.
Further studies in rheumatoid arthritis, modelling expression quantitative trait loci (eQTLs) at SC resolution in memory T cells found several autoimmune variants enriched in cell-state-dependent eQTLs64, identifying risk variants for rheumatoid arthritis enriched near the ORMDL3 and CTLA4 genes. It is important to note that eQTLs depend on the functional cell state, thus their identification is complicated in studies that aggregate cells.
Technological advancements building on SC protocols can further enhance disease understanding. For example, tetramer-associated T cell antigen receptor sequencing (TetTCR-SeqHD) helped to unravel the role of cytotoxic T cells in type 1 diabetes by combining TCR-seq readouts with cognate antigen specificity, gene expression and surface marker presence65.
Infectious diseases
A prominent example of the use of SC approaches to advance understanding of infectious diseases is in the recent study of coronavirus disease 2019 (COVID-19) to identify immune correlates of disease severity in human tissue. Comparing bronchoalveolar lavages of patients with COVID-19 of different disease severity found local immune profiles associated with disease status66. Analyses of SC transcriptome, surface proteome and T and B lymphocyte antigen receptors of PBMC samples from patients with COVID-19 found a monocytic role in platelet aggregation, circulating follicular helper T cells in mild disease and clonal expansion of cytotoxic CD8+ T cells and an increased ratio of CD8+ effector T cells to effector memory T cells in the more severe cases67. These findings indicate cellular components that might be targeted therapeutically. Similarly, scRNA-seq of circulating immune cells and readouts of metabolites in plasma of patients with COVID-19 revealed an intricate interplay between immunophenotypes and metabolic reprogramming. Emerging rare, but metabolically dominant, T cell subpopulations were found, along with a bifurcation of monocytes into two metabolically distinct subsets that correlated with disease severity68. Further, combining SC transcriptomics and SC proteomics (Box 2) with mechanistic studies found that generation of the C3a complement protein fragment by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection drives differentiation of a CD16-expressing T cell population associated with severe COVID-19 disease outcomes69.
SC analysis of lung tissue samples collected post-mortem from patients with COVID-19 identified molecular fingerprints of hyperinflammation, alveolar epithelial cell exhaustion, vascular changes and fibrosis70. Data suggested FOXO3A suppression as a potential mechanism underlying the fibroblast-to-myofibroblast transition associated with COVID-19 pulmonary fibrosis, providing insights into potential symptomatic treatments for SARS-CoV-2. A complementary study compiling lethal COVID-19 multi-tissue SC data sets from scRNA-seq and snRNA-seq analyses identified potential disease-relevant mechanisms, such as defective alveolar type 2 differentiation, expansion of fibroblasts and putative TP63+ intrapulmonary basal-like progenitor cells in the lungs of dead patients71. A review of the SC immunology of SARS-CoV-2 infection has provided interactive and downloadable curated SC data sets72.
Other notable applications of SC technologies in infectious diseases include the study of bacterial heterogeneous clonal evolution during infection and the characterization of granulomas in tuberculosis.
Parallel sequential fluorescence in situ hybridization (Par-seqFISH) was developed to capture gene expression profiles of individual prokaryotic cells while preserving spatial context73. This technology showed heterogeneity in growing Pseudomonas aeruginosa populations and demonstrated that individual multicellular biofilms can contain coexisting but separated subpopulations with distinct physiological activities73.
Coupling sophisticated SC analyses with detailed in vivo measurements of Mycobacterium tuberculosis-associated granulomas was used to define the cellular and transcriptional properties of a successful host immune response during tuberculosis74. Lack of clearance of granulomas and persistence of M. tuberculosis was characterized by type 2 immunity and a wound-healing involvement, whereas granulomas that drove bacterial control were dominated by the presence of pro-inflammatory type 1, type 17 and cytotoxic T cells74.
Target discovery
The precision and granularity that SC technologies bring to disease understanding can not only accelerate the discovery of new drug targets, but also potentially reduce attrition by providing insights into issues that affect the likelihood that drug candidates modulating these targets will progress successfully. Below, we discuss examples that illustrate the general impact of SC technologies in target discovery, while being mindful that the terms associated with target progression, such as identification, validation, credentialling and qualification have different but overlapping meanings.
Target identification
Oncology is at the forefront of the application of SC approaches to target identification. A clear example of the use of SC analysis in the discovery of novel cell-type-specific targets is the identification of S100A4 as a novel immunotherapy target in glioblastoma, following an integrated analysis of >200,000 glioma, immune and other stromal cells from human glioma samples at the SC level. Deleting this target in non-cancer cells reprogrammed the immune landscape and significantly improved survival75. Developing strategies to directly target cancer cells remains a primary focus, and SC technologies can also provide significant benefits here. As an example, SC genomics has recently provided a map charting potential new tumour antigens76. These are ideal targets for cell-depleting therapeutic monoclonal antibodies, as has been demonstrated for haematological cancers (for example, rituximab or alemtuzumab).
SC techniques have been applied in target identification in other therapeutic areas besides oncology. Of particular interest are studies in diseases with a fibrotic component, as there are few therapeutic options currently available. For example, scRNA-seq in mice comparing healthy and ischaemic hearts identified CKAP4 as a potential target for preventing fibroblast activation and thereby reducing the risk of cardiac fibrosis77. In cardiac samples from patients with ischaemic heart disease, expression of CKAP4 positively correlated with genes known to be induced in activated cardiac fibroblasts. In human chronic kidney disease, the creation of a multi-model SC atlas facilitated the discovery of myofibroblast-specific naked cuticle homologue 2 (NKD2) as a candidate therapeutic target in kidney fibrosis78. In addition, in a mouse model of kidney fibrosis, the transcription factor RUNX1 was identified as a potential target to block myofibroblast differentiation, after further analysis of sparse single-cell sequencing assay for transposase-accessible chromatin (scATAC-seq; Box 2) data79.
Human genetic data are a key resource for target identification4. Integrating information on cell-type-specific expression with disease-associated genetic variants from genome-wide association studies (GWAS) — so-called sc-eQTL — can identify the cell types and effector genes that have a causal role in disease, providing insight into potential therapeutic approaches80. Other strategies that combine GWAS summary statistics with SC transcriptomics quantify the heritability of a gene expression signature derived from scRNA-seq data sets (capturing either a cell type or a biological process)81. Via a method called SC Linker (Box 3), novel relationships between GABAergic neurons in major depressive disorder, disease progression programmes in M cells in ulcerative colitis and a disease-specific complement cascade process in multiple sclerosis have been identified81.
Computational frameworks integrating complementary molecular information have been used extensively to prioritize potential drug targets. For example, GuiltyTargets annotates on protein–protein interaction networks with differentially expressed genes linked to a disease, learns an embedded representation and uses this to predict new targets82. The incorporation of SC data sets into these computational approaches enables the prediction of cell-specific targets. For example, a network-based approach based on SC data sets has been used to prioritize drug targets in arthritis83.
Target credentialling and validation
In target credentialling and validation, confidence in a gene target is established by acquiring and combining evidence from various sources (disease biology, target biology and tractability, genetic studies, etc.). The translational validity of study models may also be examined to better understand potential gaps between the models and the disease biology or therapeutic aim. ScRNA-seq data can inform each of these facets.
Routes to improving confidence in a target include validating functional linkages between the target and the disease biology. Gene targets, gene signatures and cell states affected by individual perturbations and their genetic interactions may all be assessed at once through a scCRISPR screen, allowing target categorization and prioritization. Traditionally, significant resources are involved in target credentialling, and so compromises are often made between the number of targets examined and the complexity and number of readouts. ScCRISPR screening alone or after a genome-wide pooled screen (Box 2) can mitigate this trade-off by allowing tens to hundreds of perturbations to be pooled and profiled at once84–86.
An application of this scCRISPR screening approach first involved the identification of regulators of T cell stimulation and immunosuppression using a genome-wide pooled CRISPR screen, with candidate hits followed up with functional assays and Perturb-seq to reveal affected gene programmes, leading to at least four potential antitumour targets87. More recently, the platform has been expanded to allow paired CRISPR activation (CRISPRa) and CRISPR interference (CRISPRi) screening and pooled scRNA-seq profiling, advancing the range and depth of target validation. Perturb-seq could also be performed in vivo88, allowing investigation of gene functions in multiple cell types in a physiological context.
Targets may be further credentialled and validated for their impact on disease-relevant mechanisms by using functional genomics or pharmacology studies in vitro or in vivo. Currently, readouts of these studies are usually low-dimensional, focusing on only dozens of predefined proteins or specific disease-related phenotypes89–91. However, coupling these studies with unbiased omics readouts can provide more granularity, allow exploration of drug mode of action (MoA) (see also next section) and even reveal any unexpected toxicity profiles. Transcriptomic readouts are often the most cost-effective and relatively straightforward to interpret, and SC transcriptomics has the additional advantage of high resolution, especially for complex models. For example, dual specificity phosphatase 6 (DUSP6) has been proposed as a potential target for inflammatory bowel disease (IBD)92 and the roles of Dusp6, which had remained unclear previously from a study using bulk RNA sequencing93, have been dissected in mice in a cell-type-specific manner using scRNA-seq94.
De-orphaning studies are typically needed if the target of the drug candidate is unknown. These studies are particularly interesting for drug combinations or bispecific treatments, because biological mechanisms that are different from those of the individual drugs may be involved. For example, scRNA-seq profiling of CD45+-enriched cells from livers of mice treated with an anti-CTLA4 immune checkpoint inhibitor (ICI), and/or the IDO1 inhibitor epacadostat showed that the combination promotes CD8+ T cell proliferation and activation, and the enrichment of an interferon-γ (IFNγ) gene signature95. Similarly, flow cytometry and CyTOF were applied to demonstrate that anti-CD47–PDL1 bispecific treatment reduced binding on red blood cells and enhanced selectivity to the tumour microenvironment (TME), compared with anti-CD47 and anti-PDL1 monotherapies or combination therapies96. ScRNA-seq enabled further exploration of the mechanism, including myeloid population reprogramming, activation of the innate immune system and T cell differentiation, which cannot be directly measured using traditional methods.
ScRNA-seq can be conveniently combined with scATAC-seq for chromatin information, DNA-barcoded antibody staining for surface and/or intracellular protein expression (such as CITE-seq/ECCITE-seq97 and INs-seq98) and is therefore useful when target modulation results in pre- and/or post-transcriptional changes (Box 2). For instance, to study ICI resistance (ICR), Perturb-seq was extended and coupled with antibody staining and TCR profiling99. This work targeted 248 genes of the ICR signature identified in a previous study22 and revealed novel ICR mechanisms including downregulation of CD58 along with known resistance mechanisms.
Preclinical studies
Selecting the appropriate models for target credentialling maximizes clinical translatability. In vitro models include cell lines, primary cells and patient-derived organoids (PDOs), the latter incorporating some elements of higher-order tissue organizational complexity. In vivo models include syngeneic models, in which murine cancer cells are isografted into genotypically similar mice, PDX in immunodeficient mice, and genetically engineered mouse models (GEMMs), which recapitulate genetic alterations crucial to human carcinogenesis. Before the advent of SC omics technologies, the relative translatability of derived research models could be assessed using bulk and/or antibody-targeted SC methods (for example, flow cytometry) capable of demonstrating that characteristics of patients or donors were, in fact, recapitulated by the research models100. SC sequencing methods expand the granularity with which model or patient fidelity can be examined by shifting assessments from wholesale pools or averages to measurements of cell-type composition, intra-tissue heterogeneity and detection of rare cell phenotypes.
It has long been suggested that therapeutic strategies that account for the cellular pathogenic diversity present in complex diseases such as cancer are more likely to be successful in patients. ScRNA-seq profiling of the Cancer Cell Line Encyclopedia (CCLE) revealed patterns of heterogeneity shared between tumour lineages and specific cell model lines, suggesting that derivative cell models are promising tools for the discovery of therapeutic strategies that are not compromised by cellular heterogeneity101.
Although cell lines are easy to manipulate and have limited associated costs, more complex biological model systems better recapitulate the cell–cell interplay and emergent functions of human physiology. Using scRNA-seq to expand and quantify the extent of this recapitulation helps to guide efforts towards the most translatable systems for preclinical development, and recent areas of focus include mouse102 and human organoids103. Human liver organoids have been shown to be highly predictive for drug-induced liver injury (DILI)104, and human PDOs derived from pancreatic duct adenocarcinoma malignant ductal cells have been assessed as a good model for the human counterpart105.
Taking model complexity a step further, SC sequencing studies of hepatoblastoma and lung adenocarcinoma have demonstrated that tumour state and heterogeneity are preserved in PDX models despite differences in TME106 and that they can help to identify heterogeneity in drug responses and likely associations with anti-drug resistance107.
Characterization of well-established GEMMs at SC resolution108 and compendiums of mouse SC transcriptomic data have facilitated the identification of genes with similar murine and human expression profiles109, ligand–receptor interactions across all cell types in a microenvironment of syngeneic mouse models110, and similarities across murine–human cell populations or subpopulations in lung cancer18 (Supplementary Fig. 2). Similarly, recent SC studies revealed mechanisms underlying chemotherapy-induced ototoxicity after comparing healthy and cisplatin-exposed mice111, as well as mechanisms of ICI-induced liver injury following comparisons of treated versus untreated mice95.
A growing number of public SC data sets, representing models of interest, healthy and diseased human donors, are enabling researchers to better assess translatability18,109,112 (Table 1).
Table 1.
Examples of publicly available single-cell data sets and their applications in different phases of drug discovery
| Resource | Utility | Data repository and associated URL |
|---|---|---|
| Application: target expression in healthy human tissue; cell-type annotation of new data sets | ||
| Human Cell Atlas186 | Community-generated multi-omic open SC data processed by a uniform pipeline | Query and/or download data via project portal https://data.humancellatlas.org/ |
| Human Cell Landscape173 | Reference SC atlas for human healthy tissue |
Raw sequence data on CNGB — the CNGB Nucleotide Sequence Archive accession number is CNP0000325. Expression matrix on GEO — the GEO accession number is GSE134355 Binary expression data on Figshare — and https://figshare.com/articles/dataset/HCL_DGE_Data/7235471 Online viewer via Cellxgene — https://cellxgene.cziscience.com/d/human_cell_landscape-3.cxg/ |
| Tabula Sapiens (https://tabula-sapiens-portal.ds.czbiohub.org/)174 | Reference SC atlas for human healthy tissue |
Raw sequence data on AWS S3 — https://registry.opendata.aws/tabula-sapiens/ Binary formatted expression matrix on FigShare — https://figshare.com/projects/Tabula_Sapiens/100973 Online viewer via Cellxgene — https://cellxgene.cziscience.com/collections/e5f58829-1a66-40b5-a624-9046778e74f5 |
| Application: target expression in healthy model organisms; cell-type annotation of new data sets | ||
| Tabula Muris (https://tabula-muris.ds.czbiohub.org/)109 | Reference SC atlas for murine healthy tissue |
Raw sequence data on GEO (GSE109774) Binary expression data on FigShare — https://figshare.com/projects/Tabula_Muris_Transcriptomic_characterization_of_20_organs_and_tissues_from_Mus_musculus_at_single_cell_resolution/27733 |
| Non-human primate SC atlas187 | Reference SC atlas for non-human primate Macaca fascicularis |
Raw sequence data are deposited to the CNGB Nucleotide Sequence Archive (CNP0001469) Count matrix data are available from https://db.cngb.org/nhpca/download Explorable database accessible at https://db.cngb.org/nhpca/ |
| Application: cell-type annotation of new data sets | ||
| Azimuth human PBMC188 | CITE-seq profiling of PBMCs from multiple human donors |
Raw sequencing data are available in the dbGaP under the accession number dbGaP: phs002315.v1.p1 CITE-Seq and ECITE-seq gene expression and ADT matrices are available on GEO: GSE164378 Data set can be explored online at https://atlas.fredhutch.org/nygc/multimodal-pbmc/ Azimuth provides query mapping facilities https://atlas.fredhutch.org/nygc/multimodal-pbmc/ |
| Cross-tissue immune cell atlas189 | scRNA-seq of immune cells across different tissues in healthy humans |
Raw SC sequencing data are available in the ArrayExpress database: E-MTAB-11536 Processed data can be downloaded and interactively explored at https://www.tissueimmunecellatlas.org |
| Application: disease understanding; target identification and validation | ||
| Pan-cancer blueprint190 | scRNA-seq profiling of human cancer biopsies for several cancer types (CRC, breast cancer, ovarian and lung cancer) | Raw sequencing reads of the SC RNA experiments have been deposited in the ArrayExpress database at EMBL-EBI (www.ebi.ac.uk/biostudies/arrayexpress) with accession number E-MTAB-8107, E-MTAB-6149 and E-MTAB-6653. Online SCope viewer is also available at http://blueprint.lambrechtslab.org/ |
| Spatially resolved breast cancer atlas32 | scRNA-seq profiling of human primary breast cancer biopsy samples covering common subtypes |
Raw sequence data are deposited with the European Genome-phenome Archive (EGAS00001005173) Expression matrices are available through the GEO (GSE176078) All processed scRNA-seq data are available for in-browser exploration at https://singlecell.broadinstitute.org/single_cell/study/SCP1039/a-single-cell-and-spatially-resolved-atlas-of-human-breast-cancers All spatially resolved transcriptomics data from this study are available from the Zenodo data repository (10.5281/zenodo.4739739) |
| Pan-cancer SC atlas of tumour infiltrating lymphocytes191 | SC atlas of cytotoxic T lymphocytes from the immune TME of a pan-cancer cohort of 316 patients covering 21 types of cancer |
Sequencing data are available at Genome Sequence Archive (PRJCA001702). Processed gene expression data are deposited in GEO (GSE156728) Online data browser is also available at: http://cancer-pku.cn:3838/PanC_T |
| Tumour Immune SC Hub (TISCH)192 | Repository of uniformly processed human and murine scRNA-seq data covering several cancer types | Data can be explored at http://tisch.comp-genomics.org. Individual data sets can be downloaded as expression matrices |
| Human Tumour Atlas Network (HTAN)46 | An NIH-funded initiative to capture tumour initiation and progression in spatial and SC tumour atlases | Data and publications can be explored and downloaded from the portal: https://humantumoratlas.org/ |
| Application: disease understanding; target identification and validation; cell-type annotation of new data sets | ||
| Tumour immune cell atlas193 | Integrated immune TME atlas covering several types of cancer | A binary version of the expression count matrix and metadata can be downloaded from https://zenodo.org/record/4263972#.YQfScVMzYTs (h5ad or SeuratObject is available) |
| Accelerating Medicines Partnership Rheumatoid Arthritis and Systemic Lupus Erythematosus (AMP RA/SLE) phase I project194 | SC atlas of immune cell phenotypes in rheumatoid arthritis and lupus nephritis |
The scRNA-seq data, bulk RNA-seq data, mass cytometry data, flow cytometry data, and the clinical and histological data for this study are available at ImmPort (https://www.immport.org/shared/study/SDY998, study accession code SDY998) The raw scRNA-seq data are deposited in dbGaP (phs001457.v1.p1) Data can be explored at https://immunogenomics.io/ or https://portals.broadinstitute.org/single_cell/study/amp-phase-1 |
| Application: target identification and validation | ||
| SOMA Data Portal195–198 |
Reference SC chromatin accessibility (sci-ATAC-seq) for Drosophila melanogaster embryonic tissue and murine healthy tissue. SC transcriptome (sci-RNA-seq) for Caenorhabditis elegans larval tissue and murine embryo |
Data can be queried and downloaded from the project’s data portal at: https://atlas.gs.washington.edu/hub/ |
| Application: target validation (interpretation of non-coding variants in GWAS) | ||
| SC chromatin accessibility data set199 | Reference SC chromatin accessibility (via sci-ATAC-seq) from 70 primary tissue samples collected from 25 distinct anatomical sites in four human donors. | Data are available from GEO under GSE165659 |
ATAC, assay for transposase-accessible chromatin; CRC, colorectal cancer; GEO, Gene Expression Omnibus; GWAS, genome-wise association studies; PBMC, peripheral blood mononuclear cell; SC, single-cell; scATAC-seq, single-cell sequencing assay for transposase-accessible chromatin; sci-RNA-seq, single-cell combinatorial indexing RNA sequencing; scRNA-seq, single-cell RNA sequencing; TME, tumour microenvironment.
Drug screening and MoA analysis
High-throughput screening (HTS) in drug discovery is traditionally performed using coarse (cell viability or proliferation) or highly specific (marker expression) readouts. If a more unbiased phenotypic assessment is chosen, using bulk assessments such as RNA-seq assumes that all cells in the assay behave similarly. In comparison with bulk RNA-seq, SC transcriptomics offers more detailed views of the responding cell types, and the corresponding cell-type-specific changes (pathway, off-target effects, dose–response profiles), allowing for separation of confounding factors such as cell cycles. Therefore, HTS approaches have recently been combined with scRNA-seq readouts. Standard HTS tests a much larger number of compounds but typically at a single dose and under very limited biological conditions, whereas the novel HTS approaches that use SC gene expression readouts test several doses and conditions at the same time and are well adapted for drug MoA studies (Fig. 4).
Fig. 4. Single-cell high-throughput screening.
a, Standard high-throughput screening (HTS) tests a much larger number of compounds than HTS using single cells, but typically at a single dose and a single biological condition. The most active compounds obtained by standard HTS must be further studied (for example, dose–response analysis) but finally provide hits that are the starting point for drug discovery of active and safe drugs. b, HTS using single-cell approaches allows for testing of several doses and conditions at the same time and it is mainly used for drug mode of action (MoA) studies. In the uniform manifold approximation and projection (UMAP) embeddings shown, each cell is coloured either by the type of perturbation or the perturbation dose. k, thousand; M, million; t-SNE, t-distributed stochastic neighbour embedding. Elements of part b adapted from: ref. 200, CC BY 4.0; ref. 115. © The Authors, some rights reserved; exclusive licensee AAAS.
To mitigate the costs of scRNA-seq as a readout for chemical perturbation studies and to increase its throughput, multiplexing techniques have been developed. Hundreds of compounds can now be simultaneously profiled, considering multiple doses, time points and cell types, leading to a comprehensive understanding of compound function at scale and SC resolution. Using pre-existing genetic diversity and barcode-labelled antibodies or lipids, samples originating from different experimental conditions (time points, compounds, dose) can be pooled together; techniques that are collectively called hashing. For example, MIX-seq increases throughput using single-nucleotide polymorphism (SNP)-based demultiplexing of scRNA-seq readouts of cell lines and has been used to identify treatment-induced transcriptional changes for 13 drugs on up to 99 cell lines113. Another application of this approach relied on transient transfection of cells with short oligo barcodes114. The technology was validated by first multiplexing cell samples from various species (human or mouse) and, in a subsequent experiment, by multiplexing different time exposures of a human chronic myelogenous leukaemia cell line to a drug perturbation (imatinib, a BCR–ABL-targeting drug). Multiplexing the response of this cell line to 45 drugs (mostly kinase inhibitors) revealed drug-induced differential gene expression. A recent extension of single-cell combinatorial indexing sequencing (sci-RNA-seq), called sci-Plex, introduces a precursory step for sample multiplexing by single-stranded DNA (ssDNA) oligo uptake in single nuclei. This technique has been applied to screen exposure of 188 compounds in three cancer cell lines and profiled up to 650,000 cells115. Common and dose-dependent pathways associated with HDAC inhibitors, interfering with epigenetic cellular mechanisms, across these three diverse cancer cell lines were discovered. A metabolic consequence to depletion of cellular acetyl-CoA reserves in HDAC-inhibited cells was found, providing insight into the MoA of histone deacetylase (HDAC) inhibitors.
The field of deep learning has embraced the rich and high-dimensional data sets generated by SC multiplexed perturbation experiments (see review116). These methods enable the prediction of the cellular changes induced by a drug117 or exploration of the prohibitively large combinatorial space when combining chemical perturbations (for example, compositional perturbation autoencoder (CPA)118). The latter can identify potential combination treatments from the large multiplex SC data sets generated by techniques such as sci-Plex.
SC approaches using human samples can also help to explore the MoA of drugs or vaccines. As an example, elucidating the nature of the induced immunological memory after SARS-CoV-2 vaccination from real-world evidence has complemented the preclinical and clinical studies of these vaccines. SC technologies were used to compare the immunological changes induced by natural infection, vaccine-based antigen exposure or a combination of the two. The immunological B cell response to BNT162b2 vaccination was charted using scRNA-seq and scBCR-seq (Box 2), and the effectiveness of this mRNA vaccine against emerging variants of concern was analysed119. On the basis of SC data, it was discovered that the antibody response resulting from hybrid exposure (previously infected people vaccinated with the BNT162b2 mRNA vaccine) has an increased potency for neutralization120. These findings were later proved to be clinically relevant in a much larger cohort of patients121. Regarding therapies, the RECOVERY trial established dexamethasone as an effective treatment for hospitalized patients with COVID-19 receiving oxygen or mechanical ventilation122. Subsequent SC studies unravelled the immunological components that underlie the effectiveness of dexamethasone. A prominent role for neutrophils in response to this potent corticosteroid in patients with severe COVID-19 was discovered123. These insights may thus help the development of more targeted treatment options for severe COVID-19.
Finally, SC expression profiling has also been applied to study the biological mechanisms of drug resistance at cellular resolution. Analysing SC data from pre- and multiple post-treatment time points from a lung adenocarcinoma cell line demonstrated the mechanism of acquired resistance to epidermal growth factor receptor (EGFR) tyrosine kinase inhibitors such as erlotinib in non-small-cell lung carcinoma and the existence of intracellular heterogeneity in treatment sensitivity, highlighting the importance of unbiased SC readouts124.
Biomarkers and patient stratification
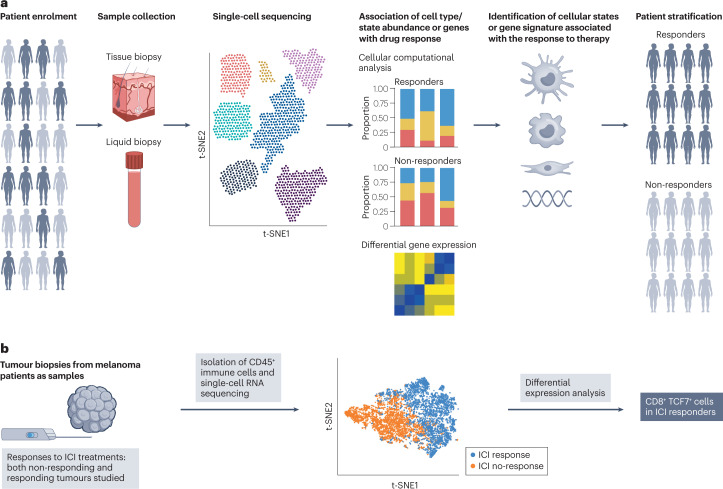
In some settings, patients can be stratified into refined populations on the basis of disease prognosis or therapeutically relevant markers that predict drug response. These prognostic or predictive biomarkers are often used as eligibility criteria in clinical trials to identify patients who are more likely to have disease progression or respond to a drug, respectively (Fig. 5a).
Fig. 5. Biomarker discovery and patient stratification.
a, Single-cell RNA sequencing or single-cell multi-omics technologies enable the identification of a predictive biomarker from a cohort of patients enrolled in an early-phase clinical study. Such a predictive biomarker can be used to identify patients who can benefit from a given treatment as a biomarker enrichment strategy. b, Single-cell analysis of immune cells from samples from patients with metastatic melanoma treated with immune checkpoint inhibitor (ICI) therapies uncovers a TCF7+ memory-like state in the cytotoxic T cell population associated with a positive outcome. t-SNE, t-distributed stochastic neighbour embedding. Elements of part b reprinted with permission from ref. 19, Elsevier.
Bulk transcriptomic signatures have been typically used to determine prognostic biomarkers in cancer, as in the case of the four consensus molecular subtypes (CMS1–4) defined by an international consortium for CRC125. However, the CMS classification has not yet proved convincingly useful in the clinic126. Bulk sequencing inherently lacks the resolution to capture crucial cell populations of CRC tumours and their complex microenvironment; and the underlying epithelial cell diversity remains unclear in the CMSs. Recently, scRNA-seq has helped to define more precise prognostic biomarkers in CRC127,128. Analysis of the transcriptomes of single cells from tumour and adjacent normal samples led to the definition of two epithelial cell groups with different intrinsic CMSs (named iCMS2 and iCMS3). Combining them with microsatellite instability and fibrosis status, a new classification called IMF has been proposed128. IMF includes five subtype classes, having distinct signalling pathways, mutational profiles and transcriptional programmes. Although promising, the value of this new classification is yet to be proved in the clinic.
ICI therapy has been successful in achieving durable responses in a subset of patients in a wide range of malignancies. However, there are still many unanswered questions around why not all patients respond to ICI therapy, and identification of predictive biomarkers for the response of ICI remains a key goal. Through these efforts, several predictive biomarkers, including tumour mutation burden (TMB), have been discovered129,130. Unfortunately, these predictive biomarkers fail to explain response to ICI for all patients. Recent SC sequencing studies have demonstrated the ability to identify new predictive biomarkers for the response or resistance to ICI. A study of CD8+ T cellular states at baseline19 revealed that responders to checkpoint inhibitors are enriched in the TCF7+CD8+ T cell state, which is also present in other indications responsive to checkpoint blockade (Fig. 5b). Beyond the conventional CD8+ T cell mediated mechanisms associated with ICI response, SC sequencing is also highlighting other cell types that shape response, such as TREM2hi macrophages, γδ T cells, CXCL9+ tumour-associated macrophages, T cell exclusion signatures and lung cancer activation module (LCAMhi) characterized by PDCD1+CXCL13+ activated T cells, IgG+ plasma cells and SPP1+ macrophages131–136. Promisingly, some of these cell types and states have been recurrent in multiple independent studies across tumour types137 and have outperformed currently used predictors such as TMB, tumour infiltrating lymphocyte (TIL) levels and PDL1 expression. In addition to scRNA-seq, there are examples of SC spatial analysis being applied to identification of potential predictive biomarkers of response. The proximity of exhausted CD8+ T cells to PDL1+ cells has been reported to predict the clinical response of combined PARP and PD1 inhibition in ovarian cancer138, while the proximity of antigen-presenting cells to stem-like CD8 T cells in intratumoural tertiary lymphoid structures has been reported to predict ICI efficacy139,140.
ScRNA-seq has also been applied to characterize chemotherapy resistance processes in cancer, as exemplified by a study in high-grade serous ovarian cancer (HGSOC). SC analysis of tissue samples collected before and after chemotherapy showed that stress-associated cancer cell populations pre-exist and are subclonally enriched during chemotherapy. The stress-associated gene signature also predicted poor prognosis in HGSOC141. In addition, scRNA-seq may be applied to predict future relapse, as seen in MLL-rearranged acute lymphoblastic leukaemia (ALL) by quantifying the proportion of cells that are identified as resistant or sensitive to treatment142. In this study, the relapse prediction outperformed the current risk stratification scheme143.
Outside oncology, SC studies are, for the first time, providing an opportunity to stratify disease into actionable subtypes. In IBD, scRNA-seq identified a cellular module called GIMATS in inflamed tissues from patients with Crohn’s disease144, consisting of IgG plasma cells, inflammatory mononuclear phagocytes, activated T cells and stromal cells. A high GIMATS score in patients was associated with failure to achieve durable remission after antitumour necrosis factor (TNF) therapy. In addition, profiling patients with ulcerative colitis and healthy individuals identified immune and stromal cells (including inflammation-associated fibroblasts) associated with resistance to anti-TNF treatment145. Furthermore, scRNA-seq analysis of PBMCs from patients with acute Kawasaki disease revealed the decreased abundance of CD16+ monocytes and downregulation of pro-inflammatory cytokines such as TNF and IL-1β in response to high-dose intravenous immunoglobulin (IVIG) therapy146. There have now also been several studies that have applied scRNA-seq approaches to diseased tissues and reported on biomarkers predictive of drug response or resistance124,131,147; however, there is still a gap in terms of understanding how well these findings translate into the clinic.
Although these SC studies are limited in terms of patient numbers, conditions and samples, methods such as cell-type deconvolution allow them to be used to complement existing bulk RNA-seq studies that typically have more mature response and outcome data22.
Monitoring of drug response and disease progression
Clinical monitoring of both disease progression and response to therapy with SC sequencing approaches is starting to influence clinical decision-making. The field of oncology has taken the lead in this area. The concept of minimal residual disease (MRD) as a metric to indicate remaining cancer cells during or after completing therapy has been a central tenet in measuring drug response. For example, patients with acute myeloid leukaemia (AML) often harbour multiple subclones, each with complex molecular abnormalities148. Clinical practice today defines complete remission as <5% blasts detected by morphological evaluation in the bone marrow without an assessment of subclonal molecular abnormalities or their evolution during therapy. Evidence is mounting that MRD assessments below this 5% threshold are a relapse risk factor and could therefore guide treatment decisions149. MRD assessment with SC mutational profiling (in contrast to more traditional MRD methods) allows for subclonal assessment at lower detection limits and for analysis of subclonal evolution throughout treatment150. SC mutational profiling improved sensitivity and specificity of MRD detection and was also able to identify relapse-causing resistant clones.
The relapse risk associated with MRD is partially explained by the presence of persister cells that are induced in response to treatment. This type of drug resistance is often driven by non-genetic adaptive mechanisms, although these are poorly understood. To study the rare and transiently resistant persister cells, a high-complexity lentiviral barcode library called Watermelon was developed to simultaneously trace the clonal lineage, proliferation status and transcriptional profile of individual cells during drug treatment151 (Supplementary Fig. 3). This approach identified rare cancerous persister lineages that are preferentially poised to proliferate under drug pressure and found that upregulation of antioxidant gene programmes and a metabolic shift to fatty acid oxidation are associated with persister proliferative capacity. Obstructing oxidative stress or rewiring of the metabolic programme of these cells alters their proportion. In human tumours, programmes associated with cycling persisters are induced in response to multiple targeted therapies. Persister cell states should thus be targeted to delay or even prevent cancer recurrence. In addition, the PERSIST-SEQ consortium (https://persist-seq.org/) was initiated to create a SC atlas of persister cells to improve the understanding of therapeutic resistance in cancer. Similarly, initiatives like HTAN46 could potentially contribute to consistent mapping of persister cell states among the set of clinical transitions of adult and paediatric malignancies when exploring therapeutic resistance. A study in TNBC showed that treatment-resistant clones originated from pre-existing cancer cells. By combining bulk whole-exome sequencing (WES) with SC transcriptomics, it was demonstrated that some of these adaptive changes were not induced by somatic mutations but were characterized by transcriptional reprogramming of these cells152.
As discussed previously, ICI therapy is a promising new therapeutic modality for some cancer patients, and understanding which subpopulation benefits from this treatment option is important. In addition, monitoring of pharmacodynamic changes and closely following response to ICI treatment from a molecular level are required for better patient selection and overall treatment outcome improvement. Mechanisms by which PD1/PDL1 blockade either revives pre-existing TILs or recruits novel T cells have been examined recently with the application of paired scRNA-seq and scTCR-seq on site-matched tumours from patients with basal or squamous cell carcinoma before and after anti-PD1 therapy153. Analysis of TCR clones and their transcriptional phenotypes revealed that drug response is driven by the expansion of novel T cell clones not previously observed in the same tumour, probably derived from a distinct repertoire of T cell clones that recently migrated into the tumour. Another SC study154 showed that CXCL13+CD8+ T cells were expanded in response to PDL1 treatment and identified a circulating T cell subtype that shared higher levels of TCR clones with tumour CXCL13+CD8+ T cells. The number of T cell clonotypes induced during early treatment provides a good proxy for future treatment success. This metric was used to identify SC changes induced by successful ICI treatment during a window of opportunity study155. These findings have also been recently confirmed in a multiple tumour type study155,156, thereby not only providing insight into the PD1/PDL1 blockade MoA, but also suggesting that liquid biopsies that sample TCR repertoire and identify clonal changes upon treatment may provide an actionable pharmacodynamic response.
Current challenges
Several challenges remain for industry to harness the transformational capabilities of scRNA-seq technologies, which will require changes to infrastructure and ways of working. Moreover, as the generation of scRNA-seq data in the public domain has outpaced that of internal efforts from any single pharmaceutical company, effective integration of all relevant scRNA-seq data is particularly challenging. In addition, owing in part to sample requirements and cost of scRNA-seq data generation, it is not likely to quickly replace bulk molecular profiling of early discovery or clinical samples, and so effective integration of scRNA-seq and bulk molecular profiling data is also needed.
Study design and implementation
Standardized design and implementation of SC experiments is still in its infancy. Although SC resolution has the potential to improve understanding of cell states and subsets of rare populations, discerning a cell type precisely and consistently across different experiments for rare cell populations is difficult, especially when fine distinctions guide cell-type identification. A uniform analysis pipeline, together with consistent methodology and vocabulary, are prerequisites to addressing this. Multi-omics approaches, by providing orthogonal indicators including cell surface and intracellular proteins or epigenetic markers, can further refine cell-state delineation but also imply new analysis challenges157–161.
SC sequencing throughput is primarily limited by the cost, but also by sample processing and computation capacity. For scRNA-seq, tissue samples need to be dissociated and processed immediately after collection to preserve high RNA quality145,162. SC library preparation poses a challenge to clinical sites where personnel may not necessarily be trained to handle sample preparation and specialized equipment. Sample quality and consistency are also hard to control, especially in large-scale multi-site clinical studies. Technology development of single-nucleus sequencing on cryopreserved or even formalin-fixed paraffin embedded (FFPE) samples provides a potential solution to this issue, allowing clinical sites to bank biopsies for later processing163–165. This technology also makes it possible to take advantage of banked samples from previous studies. However, care should be taken when selecting technologies as each has its own limitations166,167.
An online calculator (https://satijalab.org/howmanycells/) can help to determine the number of cells to be interrogated in a sample given prior assumptions on the diversity and relative composition of cells in the biology under investigation. Guidance in deciding which protocol to use or how deeply to sequence the collected cells has been provided168. In addition, design considerations for setting up longitudinal SC experiments have been reported169.
Design of SC experiments presents unique opportunities and challenges compared with bulk transcriptomics assays. On one hand, the availability of many SC samples within the experiment allows application of machine learning approaches that may be inappropriate for the typically powered bulk experiment. However, the results may have limited generalizability, owing to the low number of biological samples used to generate the SC data. On the other hand, compared with bulk RNA-seq, scRNA-seq is more expensive, and samples are more difficult to access and process. Bulk techniques have been optimized to deal with poor-quality RNA, frozen samples and even FFPE samples, whereas SC technology is only recently expanding beyond the use of fresh tissue. Enabling technologies, such as cryopreservation170 or snRNA-seq165, are still undergoing considerable optimization. A balance in complexity and budget can be achieved by combining bulk and scRNA-seq in a single experiment. SC samples can be used to computationally deconvolute cell-type abundance from bulk samples collected using an experimental set-up that favours fewer SC and more bulk sequenced samples. In addition, leveraging publicly available SC data sets can mitigate budget constraints.
Data accessibility
The current organization of public SC data generally falls short of the FAIR principles for data stewardship in several aspects171, in particular with respect to data accessibility. Ongoing cataloguing efforts (for example, the BROAD Single Cell Portal — https://singlecell.broadinstitute.org/single_cell, spreadsheet of data set metadata172) and international collaborations to generate healthy reference databases (for example, Human Cell Landscape (HCL)173, Tabula Sapiens174 — https://tabula-sapiens-portal.ds.czbiohub.org/) provide an initial entry point for discovery of data sets. However, none of these initiatives is comprehensive, resulting in the need to manually search the publication databases (for example, PubMed) and omics repositories (for example, GEO). Without uniform metadata across these databases, the search strategy must also be varied between various resources to ensure completeness.
Within a given organization, some data are likely to be accessible only to a subset of analysts. Tracking designations flagging permissible data use in the metadata versus in an external system each present different barriers related to internal risk management and compliance, as well as to scientists and analysts seeking to use those data or to build on previously completed analyses. For public data sets, similar issues exist — data access might be restricted behind security portals, as in the case of dbGaP and EGA, because of privacy laws, contractual considerations or the sensitivity of human data. This is especially true for raw reads from full transcript protocols such as Smart-Seq2 and is equally likely to be applicable to internally generated data.
Data interoperability and reusability
Most SC transcriptomics data sets of published work are made available publicly. Unfortunately, there is considerable variability in the format and layout of data. Digital formats for expression or count matrices (scRNA-seq) and experimental metadata are not standardized175. In addition, lack of comprehensive sample metadata is a common problem. Therefore, the interoperability of these data sets is limited.
Moreover, the non-uniformity of data processing, including the quality control (QC), cell-type annotation and the lack of a well-defined cell-type nomenclature (that is, either ‘flat’ or ‘shallow’ nomenclatures are used, with different levels of detail across studies), necessitates reprocessing of the data sets to interrogate them for new research questions.
Currently, the pharmaceutical industry either resorts to in-house curation efforts to augment their internal library of SC data sets with uniformly processed public entries and/or engages with external vendors for this service (see Box 5 for an example from a company and Box 6 for general use of SC public data sets by industry). The maturity, range and type of services provided by vendors varies greatly, from project-based and ad hoc curation of a small set of data sets, to platforms that house an industrialized pipeline, SC web viewers and exploratory research environments. The extent of the curation is also highly variable: some vendors start from raw sequence reads, whereas others reuse published gene expression matrices and cell-type annotations. Another big challenge to overcome is technical variations in SC data introduced by multiple factors such as laboratories and conditions. It is crucial to properly handle technical variations in the data integration and curation step (see Box 3 for computational tools for batch-effect correction and data integration). However, these approaches are expensive and time-consuming. To avoid duplication of work across companies and academic institutions, the community could benefit from collaboratively adopting and developing common standards. The academic sector has clearly paved the way by showing the value generated by creating repositories of uniformly processed and/or integrated data sets (Table 1).
Direct exploration of published data sets is being facilitated by both online viewers hosted by some researchers and general purpose scRNA-seq platforms that provide more elaborate exploratory analysis capabilities. Researcher-hosted viewers are useful to quickly check the expression of a gene but do not support maximal reuse of published data sets. Even the most advanced viewers, such as Cellxgene176 limit the scope of interrogation to selected use cases. These viewers are not a durable resource and often rely on temporary web hosting and are therefore more appropriate for accessing the data immediately after publication. By contrast, general purpose platforms such as Cumulus/Pegasus, which runs on Terra.Bio177, provide a cloud infrastructure tailored to run scRNA-seq bioinformatics pipelines and a notebook system for exploratory analysis. The EMBL-EBI Single Cell Expression Atlas (SCEA)178 has built a uniform pipeline for transcript quantification, quality control and cell-type annotation, and it runs on the browser-based Galaxy platform179. A final example, the HCA Data Coordination Platform (DCP), is a public, cloud-based platform on which scientists can share, organize and interrogate SC data.
Box 5 Harmonizing metadata across single-cell data sets.
Single-cell (SC) sequencing performs unbiased profiling of individual cells and enables evaluation of rare cellular populations, often missed using bulk sequencing. However, the diversity and multiplicity of the SC data sets pose a challenge, further exacerbated when working with large data sets typically generated by complex organizations such as the Human Cell Atlas (HCA) consortium. Merging public domain SC data sets with those generated within the private sector adds another complication. As the number and scale of SC data sets increase, there is an unmet technological need to develop suitable database platforms to evaluate key biological hypotheses across this multiplicity of data sets. In addition to the absence of a common processing workflow mapping raw sequences to gene expression matrices in a uniform way, the lack of standardized metadata collection is a primary challenge.
To address this challenge, the REVEAL:SingleCell platform, built by a pharma company on top of SciDB, provides unified scientific data management and computational tools to load, store, retrieve and query multiple SC data sets314. Its data model accommodates FAIR access to heterogeneous, multi-attribute data as well as metadata such as ontologies and reference data sets. Multiple users can load, read and write data in a secure, transactionally safe manner. REVEAL:SingleCell provides purpose-built data schema, interfaces and task-focused functionality, using a controlled vocabulary. R and Python APIs provide direct, ad hoc access and analysis, as well as extensibility via the integration of additional library packages. A FLASK REST API implements a web interface. A Shiny GUI supports data visualization and exploration by non-programmers.
The platform was applied to coronavirus disease 2019 (COVID-19) research; integrating a collection of 32 disease-related data sets available at that time (from 2.2 million cells in all), including public data from HCA Census of Immune Cells data set and COVID-19 Cell Atlas314. As the data sets were generated by different groups and metadata standardization was completely lacking, the company harmonized metadata for cell-type annotations, a crucial factor when performing cross-data set analysis. Harmonizing of cell-type annotations (T cell, B cell, etc.) is highly desirable because they are typically captured as free text and under variable names (Cell type, CellType, etc.). To solve the lack of metadata standardization, a workflow that identified and captured the cell-type information for each data set in a predefined variable name (Celltype.select) was created and mapped back to unique Ontobee cell ontology CL identifiers (https://ontobee.org/ontology/CL). This step harmonized the cell-type annotations from a free text format to controlled Ontobee CL identifiers. On the other hand, raw expression data from the multiple SC studies were normalized into a common format. These expression counts, along with the harmonized metadata, were then loaded into SciDB, which allows profiling queries across data sets with user-defined thresholds of gene expression values and metadata features to select cells of interest. For example, using this platform it was found that more than 40% of gallbladder cells co-express ACE2 and TMPRSS2 and can thus be infected by the virus. The workflow is generalizable for other metadata features such as tissues and diseases.
Box 6 Public single-cell data in drug discovery and development.
The vast array of publicly available single-cell (SC) data is crucial for the industrial use of SC technologies. Table 1 shows selected key public SC data resources of interest to pharmaceutical companies. Some of these resources originate from academic initiatives to assemble pre-existing data sets into harmonized resources and atlases. The original data sets and these secondary resources can be used to complement internal research programmes in several ways.
Access to a uniform pipeline is a first step that many companies take to ensure compatibility between internally generated and public data. Unfortunately, reprocessing of public data at each company still results in considerable duplication of effort. As with bulk RNA-seq projects (ARCHS4 (ref. 315), recount2 (ref. 316) or UCSC Toil317), academic initiatives are also leading in the creation of uniform catalogues or integrated SC data sets. Sometimes this is because of an immediate need (for example, Conquer318 created a benchmark of SC data sets to assess differential expression methods), but most initiatives were driven by the added value generated. An example is the EMBL-EBI Single Cell Expression Atlas (SCEA)178, which, in addition to a uniform pipeline, also provides the original author cell-type labels as well as cell ontology-matched labels.
SC atlases, such as those produced by the Human Tumour Atlas Network46 initiative, can be used as a reference for cell-type annotation of internal research data sets (see Box 3 for relevant methods). Multimodal technologies that enrich SC transcriptomics with matched cell surface protein (for example, CITE-Seq or REAP-Seq) and/or open chromatin data, are also yielding public data sets. For instance, many CITE-Seq data sets have been generated, are publicly available and can be used to predict protein expression from internally generated single-cell RNA sequencing (scRNA-seq) experiments319.
Benchmarking of the many available computational methods in the SC field also benefits strongly from the availability of public data. Benchmarking is necessary to assess method performance and guide the development of best practices320. Synthetically generated data sets can help to assess methods, but creating such synthetic data sets is difficult321. Publicly available data sets can be used instead either to define the starting data for generative methods322 or to benchmark the generated data sets, for example, in Splatter323. Public data sets can also be used directly in other benchmarking exercises, for example, benchmarking trajectory inference methods that rely on a synthetic and public repository of data sets324.
Bulk transcriptomics assays to provide an unbiased view on the effect of a drug are now an integral part of internal research programmes in industry. The tools to deconvolute the cellular composition of bulk RNA-seq samples need prior knowledge of cell types present in the sample and their associated gene expression profiles or marker genes. Public scRNA-seq from matching tissues is an excellent source of this information. In addition, as recently illustrated using EcoTyper in diffuse large B cell lymphoma31, SC data can be used to reanalyse bulk RNA-seq from previous studies to further define cell states or classes linked to outcome. As there is a huge amount of public and internal bulk RNA-seq data available, re-analysis of public data with SC data sets focusing on specific clinical questions is of interest.
Similarly, integration of SC analysis with other types of internal or public bulk assay (for example, epigenomics, proteomics, metabolomics) would also be of value. In fact, this is an emerging frontier in research, with tools such as flux analysis and others being explored. However, although relevant for research, these approaches are not yet adopted by industry.
Public data can also serve as independent cohorts to verify internal findings, and integrative methods (for example, Harmony325) allow the generation of SC atlases by combining cellular spaces from several experiments, increasing the generalizability of exploratory research. This approach has been successfully applied to uncover biomarkers and improve disease understanding in lung fibrosis, when internal scRNA-seq data were combined with two public data sets with a similar experimental set-up (that is, control versus disease)326.
Finally, public data studies can serve as pilot experiments when performing power calculations (that is, to define the number of samples required to demonstrate predetermined effect size) and can be helpful for getting basic information related to experimental design (for example, to decide experimental protocols)168,327,328.
Conclusions and future perspectives
Most complex diseases for which treatment remains elusive have a multicellular aetiology, and a SC perspective could be crucial in advancing our understanding and ability to select the most therapeutically impactful cellular or molecular targets. SC protocols combined with sophisticated multiplex strategies have increased the scale and resolution at which assays can be performed. In addition, SC profiling of commonly used preclinical models enables researchers to select the model that best recapitulates essential human pathobiology. Interrogating human samples at cellular resolution can help to advance personalized medicine, by expediting the discovery of new biomarkers to help stratify patients on the basis of prognosis or prediction of treatment effect. A longitudinal SC view on diseased tissues during treatment can also provide physicians with a more direct and mechanistic view on response to treatment.
Having established the more mature scRNA-seq-based methods for routine use in industry, effort is increasingly focused on adopting other methods such as SC proteomics and spatial omics technologies, as industrial SC capabilities are expanded. As the core technologies become standardized, the requisite skills become more widely available and the costs fall, the rate of SC data generation is likely to continue to accelerate180,181.
As the technical challenges involved in SC data generation, curation and access are addressed, new opportunities are emerging. For example, upstream of target discovery, the focus is already shifting from the discovery of novel cell types and cellular marker genes towards hypothesis generation rooted in deeper understanding of cellular mechanisms. The integration of additional data types supports this shift as omics and other multiparametric data enhance the granularity of insight into the cellular environment. For example, mapping genetic cues on disease provided by GWAS on SC profiles from scRNA-seq experiments can help to elucidate cellular phenotypes linked to complex diseases81,182.
With the increasing maturity of spatial profiling technologies, we are beginning to better understand human tissue organization and microenvironment niches. Spatial profiling enables cell types to be accurately counted and localized within the broader tissue architecture. In addition, it facilitates the mapping of intricate auto- and paracrine interactions between cell types within a tissue. However, the resolution of the most unbiased and comprehensive approaches (for example, 10X Visium) remains supracellular. We expect that such approaches will evolve to provide SC resolution, and thus complement and extend the pipeline of methods applicable to intercellular interaction discovery from scRNA-seq (for example, CellPhoneDB183). Moreover, advances in spatial profiling are lining up with the recent progress made in digital pathology. Combined with automated feature extraction and molecular classification of digitized pathology images via deep learning techniques184, orthogonal informational cues assayed via sequencing or multiplex imaging technologies will enable researchers to develop a deeper knowledge of the complex biology involved in some diseases.
Given the enormous technical, computational and scientific complexities involved in SC data generation and translating those data into benefits to patients, collaboration has a key role. This is clearly demonstrated by the Accelerating Medicines Partnership and LifeTime initiatives, and the rapid growth of SC research around SARS-CoV-2 (ref. 185). LifeTime established a special task force to study COVID-19 and to identify SC-based biomarkers and novel modalities. In this case, HCA and LifeTime created a common framework for sharing knowledge, data, tools and other resources. As the scale and complexity of SC data and our understanding of human biology continue to deepen, collaborative efforts between academia and industry will be increasingly vital to realize the transformational potential of SC technologies.
Supplementary information
Acknowledgements
The authors thank I. Papatheodorou (Research Group Leader, EMBL-EBI), B. Kidd (Director, Bristol Myers Squibb (BMS)), R. Loos (Director, BMS) and M. Hall (Senior Scientific Officer, EMBL-EBI) for constructive criticism and proofreading of the original article before this revision.
Glossary
- Barcode
A short DNA sequence ‘tag’ to identify reads that originate from the same cell.
- Biomarkers
Readouts used to classify biological states, often in the context of patient stratification.
- Cell-type deconvolution
Estimation of the proportion of particular cell types in a bulk RNA sequencing sample, based on cell markers or a labelled single-cell expression matrix.
- CRISPR screening
A pooled or arrayed screen of cells harbouring CRISPR-mediated gene edits.
- Doublets
Sets of two (or more) cells mistakenly considered as single cells, owing to being captured and processed in the same droplet and thus with the same barcode in data.
- Hashing
A labelling technique that attaches barcoded antibodies to cell surface proteins, allowing multiplexing of samples for single-cell sequencing, and subsequent disambiguation of sample of origin during analysis.
- Metadata
A set of data that describe and give information about other data (Oxford dictionary). For example, patient or sample characteristics in an RNA sequencing experiment.
- Seurat
A popular R package for the quality control, analysis and exploration of single-cell RNA sequencing data.
- Target credentialling
Also called target qualification. Exploration of target quality more expansively than a straightforward target validation. May include contextually informed enquiries into biological characteristics such as network, pathway or interactome mapping, regulatory landscape or other investigations intended to either help rank target quality or inform on-target biology.
- t-Distributed stochastic neighbour embedding
(t-SNE). A popular dimensionality reduction technique for the visualization of single-cell experiments.
- Trajectory inference
Inference from single-cell data of the order of cells along a dynamic biological process (for example, developmental trajectory). Relies on the fact that a heterogeneous sample provides a snapshot view on a mixture of cells in different phases along the developmental or dynamic biological process. Also called ‘pseudo-time analysis’.
- Uniform manifold approximation and projection
(UMAP). A popular dimensionality reduction technique for the visualization of single-cell experiments, with some advantages in preservation of global data structure and performance compared with t-distributed stochastic neighbour embedding.
- Unique molecular identifier
(UMI). Reads with the same UMI are from the same mRNA molecule. UMIs help in the assessment of sequencing accuracy and precision.
- Unsupervised clustering
Analysis grouping of similar samples together that does not require labelling or prior knowledge.
Competing interests
N.K. is an employee and shareholder of BMS. M.M. is an employee and shareholder of GSK. B.V.d.S. is an employee and shareholder of UCB Pharma. M.K. is an employee and shareholder of GSK. J.H. is an employee of Boehringer Ingelheim Pharmaceuticals, Inc. B.N. is an employee of Eisai, Inc. J.S.L. is an employee and shareholder of Sanofi. Y.W. was previously a shareholder of BMS. J.P. was previously an employee and shareholder of Sanofi. J.W. is an employee of Pfizer. E.F. is a shareholder of Sanofi and Board Director of Pulmobiotics. A.L. is a GSK shareholder, has consulted for Astex Therapeutics, LifeArc and Syncona and has received research funding from Novo Nordisk and AstraZeneca. X.C. is a former employee and shareholder of AbbVie. E.M.-G., W.B. and J.M. declare no competing interests.
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
These authors contributed equally: Bram Van de Sande, Joon Sang Lee, Euphemia Mutasa-Gottgens.
Supplementary information
The online version contains supplementary material available at 10.1038/s41573-023-00688-4.
References
- 1.DiMasi JA, Grabowski HG, Hansen RW. Innovation in the pharmaceutical industry: new estimates of R&D costs. J. Health Econ. 2016;47:20–33. doi: 10.1016/j.jhealeco.2016.01.012. [DOI] [PubMed] [Google Scholar]
- 2.Wouters OJ, McKee M, Luyten J. Estimated research and development investment needed to bring a new medicine to market, 2009–2018. JAMA. 2020;323:844–853. doi: 10.1001/jama.2020.1166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Paul SM, et al. How to improve R&D productivity: the pharmaceutical industry’s grand challenge. Nat. Rev. Drug Discov. 2010;9:203–214. doi: 10.1038/nrd3078. [DOI] [PubMed] [Google Scholar]
- 4.Nelson MR, et al. The support of human genetic evidence for approved drug indications. Nat. Genet. 2015;47:856–860. doi: 10.1038/ng.3314. [DOI] [PubMed] [Google Scholar]
- 5.1000 Genomes Project Consortium. An integrated map of genetic variation from 1,092 human genomes. Nature. 2012;491:56–65. doi: 10.1038/nature11632. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Sernoskie SC, Jee A, Uetrecht JP. The emerging role of the innate immune response in idiosyncratic drug reactions. Pharmacol. Rev. 2021;73:861–896. doi: 10.1124/pharmrev.120.000090. [DOI] [PubMed] [Google Scholar]
- 7.Heid CA, Stevens J, Livak KJ, Williams PM. Real time quantitative PCR. Genome Res. 1996;6:986–994. doi: 10.1101/gr.6.10.986. [DOI] [PubMed] [Google Scholar]
- 8.Cheung RK, Utz PJ. CyTOF — the next generation of cell detection. Nat. Rev. Rheumatol. 2011;7:502–503. doi: 10.1038/nrrheum.2011.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Bendall SC, et al. Single-cell mass cytometry of differential immune and drug responses across a human hematopoietic continuum. Science. 2011;332:687–696. doi: 10.1126/science.1198704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Nassar AF, Ogura H, Wisnewski AV. Impact of recent innovations in the use of mass cytometry in support of drug development. Drug. Discov. Today. 2015;20:1169–1175. doi: 10.1016/j.drudis.2015.06.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Wen L, Tang F. Recent advances in single-cell sequencing technologies. Precis. Clin. Med. 2022;5:pbac002. doi: 10.1093/pcmedi/pbac002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Jovic D, et al. Single‐cell RNA sequencing technologies and applications: a brief overview. Clin. Transl. Med. 2022;12:e694. doi: 10.1002/ctm2.694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kashima Y, et al. Single-cell sequencing techniques from individual to multiomics analyses. Exp. Mol. Med. 2020;52:1419–1427. doi: 10.1038/s12276-020-00499-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Svensson V, Vento-Tormo R, Teichmann SA. Exponential scaling of single-cell RNA-seq in the past decade. Nat. Protoc. 2018;13:599–604. doi: 10.1038/nprot.2017.149. [DOI] [PubMed] [Google Scholar]
- 15.Aldridge S, Teichmann SA. Single cell transcriptomics comes of age. Nat. Commun. 2020;11:4307. doi: 10.1038/s41467-020-18158-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Tang F, et al. mRNA-Seq whole-transcriptome analysis of a single cell. Nat. Methods. 2009;6:377–382. doi: 10.1038/nmeth.1315. [DOI] [PubMed] [Google Scholar]
- 17.Navin NE, Rozenblatt-Rosen O, Zhang NR. New frontiers in single-cell genomics. Genome Res. 2021;31:ix–x. doi: 10.1101/gr.276129.121. [DOI] [Google Scholar]
- 18.Zilionis R, et al. Single-cell transcriptomics of human and mouse lung cancers reveals conserved myeloid populations across individuals and species. Immunity. 2019;50:1317–1334.e10. doi: 10.1016/j.immuni.2019.03.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Sade-Feldman M, et al. Defining T cell states associated with response to checkpoint immunotherapy in melanoma. Cell. 2018;175:998–1013.e20. doi: 10.1016/j.cell.2018.10.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Jang JS, et al. Molecular signatures of multiple myeloma progression through single cell RNA-Seq. Blood Cancer J. 2019;9:2. doi: 10.1038/s41408-018-0160-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Tanaka N, et al. Single-cell RNA-seq analysis reveals the platinum resistance gene COX7B and the surrogate marker CD63. Cancer Med. 2018;7:6193–6204. doi: 10.1002/cam4.1828. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Jerby-Arnon L, et al. A cancer cell program promotes T cell exclusion and resistance to checkpoint blockade. Cell. 2018;175:984–997.e24. doi: 10.1016/j.cell.2018.09.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Cohen YC, et al. Identification of resistance pathways and therapeutic targets in relapsed multiple myeloma patients through single-cell sequencing. Nat. Med. 2021;27:491–503. doi: 10.1038/s41591-021-01232-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Villani A-C, et al. Single-cell RNA-seq reveals new types of human blood dendritic cells, monocytes, and progenitors. Science. 2017;356:eaah4573. doi: 10.1126/science.aah4573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Park J-E, et al. A cell atlas of human thymic development defines T cell repertoire formation. Science. 2020;367:eaay3224. doi: 10.1126/science.aay3224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.GTEx Consortium. Landscape of X chromosome inactivation across human tissues. Nature. 2017;550:244–248. doi: 10.1038/nature24265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Ramachandran P, et al. Resolving the fibrotic niche of human liver cirrhosis at single-cell level. Nature. 2019;575:512–518. doi: 10.1038/s41586-019-1631-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Song H, et al. Single-cell analysis of human primary prostate cancer reveals the heterogeneity of tumor-associated epithelial cell states. Nat. Commun. 2022;13:141. doi: 10.1038/s41467-021-27322-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Wang Q, et al. Single-cell chromatin accessibility landscape in kidney identifies additional cell-of-origin in heterogenous papillary renal cell carcinoma. Nat. Commun. 2022;13:31. doi: 10.1038/s41467-021-27660-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Nowicki-Osuch K, et al. Molecular phenotyping reveals the identity of Barrett’s esophagus and its malignant transition. Science. 2021;373:760–767. doi: 10.1126/science.abd1449. [DOI] [PubMed] [Google Scholar]
- 31.Steen CB, et al. The landscape of tumor cell states and ecosystems in diffuse large B cell lymphoma. Cancer Cell. 2021;39:1422–1437.e10. doi: 10.1016/j.ccell.2021.08.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Wu SZ, et al. A single-cell and spatially resolved atlas of human breast cancers. Nat. Genet. 2021;53:1334–1347. doi: 10.1038/s41588-021-00911-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Zhang X, et al. Dissecting esophageal squamous-cell carcinoma ecosystem by single-cell transcriptomic analysis. Nat. Commun. 2021;12:5291. doi: 10.1038/s41467-021-25539-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Pu W, et al. Single-cell transcriptomic analysis of the tumor ecosystems underlying initiation and progression of papillary thyroid carcinoma. Nat. Commun. 2021;12:6058. doi: 10.1038/s41467-021-26343-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Ursu O, et al. Massively parallel phenotyping of coding variants in cancer with Perturb-seq. Nat. Biotechnol. 2022;40:896–905. doi: 10.1038/s41587-021-01160-7. [DOI] [PubMed] [Google Scholar]
- 36.Chaligne R, et al. Epigenetic encoding, heritability and plasticity of glioma transcriptional cell states. Nat. Genet. 2021;53:1469–1479. doi: 10.1038/s41588-021-00927-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Johnson KC, et al. Single-cell multimodal glioma analyses identify epigenetic regulators of cellular plasticity and environmental stress response. Nat. Genet. 2021;53:1456–1468. doi: 10.1038/s41588-021-00926-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Croucher DC, et al. Longitudinal single-cell analysis of a myeloma mouse model identifies subclonal molecular programs associated with progression. Nat. Commun. 2021;12:6322. doi: 10.1038/s41467-021-26598-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Salehi S, et al. Clonal fitness inferred from time-series modelling of single-cell cancer genomes. Nature. 2021;595:585–590. doi: 10.1038/s41586-021-03648-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Quinn JJ, et al. Single-cell lineages reveal the rates, routes, and drivers of metastasis in cancer xenografts. Science. 2021;371:eabc1944. doi: 10.1126/science.abc1944. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Yaddanapudi K, et al. Single-cell immune mapping of melanoma sentinel lymph nodes reveals an actionable immunotolerant microenvironment. Clin. Cancer Res. 2022;28:2069–2081. doi: 10.1158/1078-0432.CCR-21-0664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Lund AW. Standing watch: immune activation and failure in melanoma sentinel lymph nodes. Clin. Cancer Res. 2022;28:1996–1998. doi: 10.1158/1078-0432.CCR-22-0214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Li J, et al. Single-cell characterization of the cellular landscape of acral melanoma identifies novel targets for immunotherapy. Clin. Cancer Res. 2022;28:2131–2146. doi: 10.1158/1078-0432.CCR-21-3145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Sun Y-F, et al. Dissecting spatial heterogeneity and the immune-evasion mechanism of CTCs by single-cell RNA-seq in hepatocellular carcinoma. Nat. Commun. 2021;12:4091. doi: 10.1038/s41467-021-24386-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Diamantopoulou Z, et al. The metastatic spread of breast cancer accelerates during sleep. Nature. 2022;607:156–162. doi: 10.1038/s41586-022-04875-y. [DOI] [PubMed] [Google Scholar]
- 46.Rozenblatt-Rosen O, et al. The Human Tumor Atlas Network: charting tumor transitions across space and time at single-cell resolution. Cell. 2020;181:236–249. doi: 10.1016/j.cell.2020.03.053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Becker WR, et al. Single-cell analyses define a continuum of cell state and composition changes in the malignant transformation of polyps to colorectal cancer. Nat. Genet. 2022;54:985–995. doi: 10.1038/s41588-022-01088-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Arenas E. Parkinson’s disease in the single-cell era. Nat. Neurosci. 2022;25:536–538. doi: 10.1038/s41593-022-01069-7. [DOI] [PubMed] [Google Scholar]
- 49.Kamath T, et al. Single-cell genomic profiling of human dopamine neurons identifies a population that selectively degenerates in Parkinson’s disease. Nat. Neurosci. 2022;25:588–595. doi: 10.1038/s41593-022-01061-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Miller MB, et al. Somatic genomic changes in single Alzheimer’s disease neurons. Nature. 2022;604:714–722. doi: 10.1038/s41586-022-04640-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Keren-Shaul H, et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell. 2017;169:1276–1290.e17. doi: 10.1016/j.cell.2017.05.018. [DOI] [PubMed] [Google Scholar]
- 52.Wang P, et al. Single-cell transcriptome and TCR profiling reveal activated and expanded T cell populations in Parkinson’s disease. Cell Discov. 2021;7:52. doi: 10.1038/s41421-021-00280-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Cadwell CR, et al. Electrophysiological, transcriptomic and morphologic profiling of single neurons using Patch-seq. Nat. Biotechnol. 2016;34:199–203. doi: 10.1038/nbt.3445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Fuzik J, et al. Integration of electrophysiological recordings with single-cell RNA-seq data identifies neuronal subtypes. Nat. Biotechnol. 2016;34:175–183. doi: 10.1038/nbt.3443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Yang AC, et al. A human brain vascular atlas reveals diverse mediators of Alzheimer’s risk. Nature. 2022;603:885–892. doi: 10.1038/s41586-021-04369-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Berg J, et al. Human neocortical expansion involves glutamatergic neuron diversification. Nature. 2021;598:151–158. doi: 10.1038/s41586-021-03813-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Simone D, et al. Single cell analysis of spondyloarthritis regulatory T cells identifies distinct synovial gene expression patterns and clonal fates. Commun. Biol. 2021;4:1395. doi: 10.1038/s42003-021-02931-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Penkava F, et al. Single-cell sequencing reveals clonal expansions of pro-inflammatory synovial CD8 T cells expressing tissue-homing receptors in psoriatic arthritis. Nat. Commun. 2020;11:4767. doi: 10.1038/s41467-020-18513-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Wu X, et al. Single-cell sequencing of immune cells from anticitrullinated peptide antibody positive and negative rheumatoid arthritis. Nat. Commun. 2021;12:4977. doi: 10.1038/s41467-021-25246-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Liu Y, et al. Classification of human chronic inflammatory skin disease based on single-cell immune profiling. Sci. Immunol. 2022;7:eabl9165. doi: 10.1126/sciimmunol.abl9165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Ingelfinger F, et al. Twin study reveals non-heritable immune perturbations in multiple sclerosis. Nature. 2022;603:152–158. doi: 10.1038/s41586-022-04419-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Bjornevik K, et al. Longitudinal analysis reveals high prevalence of Epstein-Barr virus associated with multiple sclerosis. Science. 2022;375:296–301. doi: 10.1126/science.abj8222. [DOI] [PubMed] [Google Scholar]
- 63.Lanz TV, et al. Clonally expanded B cells in multiple sclerosis bind EBV EBNA1 and GlialCAM. Nature. 2022;603:321–327. doi: 10.1038/s41586-022-04432-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Nathan A, et al. Single-cell eQTL models reveal dynamic T cell state dependence of disease loci. Nature. 2022;606:120–128. doi: 10.1038/s41586-022-04713-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Ma K-Y, et al. High-throughput and high-dimensional single-cell analysis of antigen-specific CD8+ T cells. Nat. Immunol. 2021;22:1590–1598. doi: 10.1038/s41590-021-01073-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Wauters E, et al. Discriminating mild from critical COVID-19 by innate and adaptive immune single-cell profiling of bronchoalveolar lavages. Cell Res. 2021;31:272–290. doi: 10.1038/s41422-020-00455-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Stephenson E, et al. Single-cell multi-omics analysis of the immune response in COVID-19. Nat. Med. 2021;27:904–916. doi: 10.1038/s41591-021-01329-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Lee JW, et al. Integrated analysis of plasma and single immune cells uncovers metabolic changes in individuals with COVID-19. Nat. Biotechnol. 2022;40:110–120. doi: 10.1038/s41587-021-01020-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Georg P, et al. Complement activation induces excessive T cell cytotoxicity in severe COVID-19. Cell. 2022;185:493–512.e25. doi: 10.1016/j.cell.2021.12.040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Wang S, et al. A single-cell transcriptomic landscape of the lungs of patients with COVID-19. Nat. Cell Biol. 2021;23:1314–1328. doi: 10.1038/s41556-021-00796-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Delorey TM, et al. COVID-19 tissue atlases reveal SARS-CoV-2 pathology and cellular targets. Nature. 2021;595:107–113. doi: 10.1038/s41586-021-03570-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Tian Y, et al. Single-cell immunology of SARS-CoV-2 infection. Nat. Biotechnol. 2022;40:30–41. doi: 10.1038/s41587-021-01131-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Dar D, Dar N, Cai L, Newman DK. Spatial transcriptomics of planktonic and sessile bacterial populations at single-cell resolution. Science. 2021;373:eabi4882. doi: 10.1126/science.abi4882. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Gideon HP, et al. Multimodal profiling of lung granulomas in macaques reveals cellular correlates of tuberculosis control. Immunity. 2022;55:827–846.e10. doi: 10.1016/j.immuni.2022.04.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Abdelfattah N, et al. Single-cell analysis of human glioma and immune cells identifies S100A4 as an immunotherapy target. Nat. Commun. 2022;13:767. doi: 10.1038/s41467-022-28372-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Lareau CA, Parker KR, Satpathy AT. Charting the tumor antigen maps drawn by single-cell genomics. Cancer Cell. 2021;39:1553–1557. doi: 10.1016/j.ccell.2021.11.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Gladka MM, et al. Single-cell sequencing of the healthy and diseased heart reveals cytoskeleton-associated protein 4 as a new modulator of fibroblasts activation. Circulation. 2018;138:166–180. doi: 10.1161/CIRCULATIONAHA.117.030742. [DOI] [PubMed] [Google Scholar]
- 78.Kuppe C, et al. Decoding myofibroblast origins in human kidney fibrosis. Nature. 2021;589:281–286. doi: 10.1038/s41586-020-2941-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Li Z, et al. Chromatin-accessibility estimation from single-cell ATAC-seq data with scOpen. Nat. Commun. 2021;12:6386. doi: 10.1038/s41467-021-26530-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Cano-Gamez E, Trynka G. From GWAS to function: using functional genomics to identify the mechanisms underlying complex diseases. Front. Genet. 2020;11:424. doi: 10.3389/fgene.2020.00424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Jagadeesh KA, et al. Identifying disease-critical cell types and cellular processes by integrating single-cell RNA-sequencing and human genetics. Nat. Genet. 2022;54:1479–1492. doi: 10.1038/s41588-022-01187-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Muslu O, Hoyt CT, Lacerda M, Hofmann-Apitius M, Frohlich H. Guiltytargets: prioritization of novel therapeutic targets with network representation learning. IEEE/ACM Trans. Comput. Biol. Bioinform. 2022;19:491–500. doi: 10.1109/TCBB.2020.3003830. [DOI] [PubMed] [Google Scholar]
- 83.Gawel DR, et al. A validated single-cell-based strategy to identify diagnostic and therapeutic targets in complex diseases. Genome Med. 2019;11:47. doi: 10.1186/s13073-019-0657-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Dixit A, et al. Perturb-seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell. 2016;167:1853–1866.e17. doi: 10.1016/j.cell.2016.11.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Adamson B, et al. A multiplexed single-cell CRISPR screening platform enables systematic dissection of the unfolded protein response. Cell. 2016;167:1867–1882.e21. doi: 10.1016/j.cell.2016.11.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Datlinger P, et al. Pooled CRISPR screening with single-cell transcriptome readout. Nat. Methods. 2017;14:297–301. doi: 10.1038/nmeth.4177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Shifrut E, et al. Genome-wide CRISPR screens in primary human T cells reveal key regulators of immune function. Cell. 2018;175:1958–1971.e15. doi: 10.1016/j.cell.2018.10.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Jin X, et al. In vivo Perturb-Seq reveals neuronal and glial abnormalities associated with autism risk genes. Science. 2020;370:eaaz6063. doi: 10.1126/science.aaz6063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Lazo JS, et al. Credentialing and pharmacologically targeting PTP4A3 phosphatase as a molecular target for ovarian cancer. Biomolecules. 2021;11:969. doi: 10.3390/biom11070969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Wang W, et al. MAPK4 promotes triple negative breast cancer growth and reduces tumor sensitivity to PI3K blockade. Nat. Commun. 2022;13:245. doi: 10.1038/s41467-021-27921-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Wang P-X, et al. Targeting CASP8 and FADD-like apoptosis regulator ameliorates nonalcoholic steatohepatitis in mice and nonhuman primates. Nat. Med. 2017;23:439–449. doi: 10.1038/nm.4290. [DOI] [PubMed] [Google Scholar]
- 92.Bertin S, et al. Dual-specificity phosphatase 6 regulates CD4+ T-cell functions and restrains spontaneous colitis in IL-10-deficient mice. Mucosal Immunol. 2015;8:505–515. doi: 10.1038/mi.2014.84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Ruan J-W, et al. Dual-specificity phosphatase 6 deficiency regulates gut microbiome and transcriptome response against diet-induced obesity in mice. Nat. Microbiol. 2016;2:16220. doi: 10.1038/nmicrobiol.2016.220. [DOI] [PubMed] [Google Scholar]
- 94.Chang C-S, et al. Single-cell RNA sequencing uncovers the individual alteration of intestinal mucosal immunocytes in Dusp6 knockout mice. iScience. 2022;25:103738. doi: 10.1016/j.isci.2022.103738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Llewellyn HP, et al. T cells and monocyte-derived myeloid cells mediate immunotherapy-related hepatitis in a mouse model. J. Hepatol. 2021;75:1083–1095. doi: 10.1016/j.jhep.2021.06.037. [DOI] [PubMed] [Google Scholar]
- 96.Chen S-H, et al. Dual checkpoint blockade of CD47 and PD-L1 using an affinity-tuned bispecific antibody maximizes antitumor immunity. J. Immunother. Cancer. 2021;9:e003464. doi: 10.1136/jitc-2021-003464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Mimitou EP, et al. Multiplexed detection of proteins, transcriptomes, clonotypes and CRISPR perturbations in single cells. Nat. Methods. 2019;16:409–412. doi: 10.1038/s41592-019-0392-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Katzenelenbogen Y, et al. Coupled scRNA-Seq and intracellular protein activity reveal an immunosuppressive role of TREM2 in cancer. Cell. 2020;182:872–885.e19. doi: 10.1016/j.cell.2020.06.032. [DOI] [PubMed] [Google Scholar]
- 99.Frangieh CJ, et al. Multimodal pooled Perturb-CITE-seq screens in patient models define mechanisms of cancer immune evasion. Nat. Genet. 2021;53:332–341. doi: 10.1038/s41588-021-00779-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Schütte M, et al. Molecular dissection of colorectal cancer in pre-clinical models identifies biomarkers predicting sensitivity to EGFR inhibitors. Nat. Commun. 2017;8:14262. doi: 10.1038/ncomms14262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Kinker GS, et al. Pan-cancer single-cell RNA-seq identifies recurring programs of cellular heterogeneity. Nat. Genet. 2020;52:1208–1218. doi: 10.1038/s41588-020-00726-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Mead BE, et al. Screening for modulators of the cellular composition of gut epithelia via organoid models of intestinal stem cell differentiation. Nat. Biomed. Eng. 2022;6:476–494. doi: 10.1038/s41551-022-00863-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Bock C, et al. The organoid cell atlas. Nat. Biotechnol. 2021;39:13–17. doi: 10.1038/s41587-020-00762-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Shinozawa T, et al. High-fidelity drug-induced liver injury screen using human pluripotent stem cell-derived organoids. Gastroenterology. 2021;160:831–846.e10. doi: 10.1053/j.gastro.2020.10.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Krieger TG, et al. Single-cell analysis of patient-derived PDAC organoids reveals cell state heterogeneity and a conserved developmental hierarchy. Nat. Commun. 2021;12:5826. doi: 10.1038/s41467-021-26059-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Bondoc A, et al. Identification of distinct tumor cell populations and key genetic mechanisms through single cell sequencing in hepatoblastoma. Commun. Biol. 2021;4:1049. doi: 10.1038/s42003-021-02562-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Kim K-T, et al. Single-cell mRNA sequencing identifies subclonal heterogeneity in anti-cancer drug responses of lung adenocarcinoma cells. Genome Biol. 2015;16:127. doi: 10.1186/s13059-015-0692-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Hosein AN, et al. Cellular heterogeneity during mouse pancreatic ductal adenocarcinoma progression at single-cell resolution. JCI Insight. 2019;5:129212. doi: 10.1172/jci.insight.129212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Tabula Muris Consortium. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature. 2018;562:367–372. doi: 10.1038/s41586-018-0590-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Kumar MP, et al. Analysis of single-cell RNA-Seq identifies cell-cell communication associated with tumor characteristics. Cell Rep. 2018;25:1458–1468.e4. doi: 10.1016/j.celrep.2018.10.047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Taukulis IA, et al. Single-cell RNA-Seq of cisplatin-treated adult stria vascularis identifies cell type-specific regulatory networks and novel therapeutic gene targets. Front. Mol. Neurosci. 2021;14:718241. doi: 10.3389/fnmol.2021.718241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Yofe I, Dahan R, Amit I. Single-cell genomic approaches for developing the next generation of immunotherapies. Nat. Med. 2020;26:171–177. doi: 10.1038/s41591-019-0736-4. [DOI] [PubMed] [Google Scholar]
- 113.McFarland JM, et al. Multiplexed single-cell transcriptional response profiling to define cancer vulnerabilities and therapeutic mechanism of action. Nat. Commun. 2020;11:4296. doi: 10.1038/s41467-020-17440-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Shin D, Lee W, Lee JH, Bang D. Multiplexed single-cell RNA-seq via transient barcoding for simultaneous expression profiling of various drug perturbations. Sci. Adv. 2019;5:eaav2249. doi: 10.1126/sciadv.aav2249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Srivatsan SR, et al. Massively multiplex chemical transcriptomics at single-cell resolution. Science. 2020;367:45–51. doi: 10.1126/science.aax6234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Ji Y, Lotfollahi M, Wolf FA, Theis FJ. Machine learning for perturbational single-cell omics. Cell Syst. 2021;12:522–537. doi: 10.1016/j.cels.2021.05.016. [DOI] [PubMed] [Google Scholar]
- 117.Lotfollahi MJ, Wolf FA, Theis F. scGen predicts single-cell perturbation responses. Nat. Methods. 2019;16:715–721. doi: 10.1038/s41592-019-0494-8. [DOI] [PubMed] [Google Scholar]
- 118.Lotfollahi M, et al. Learning interpretable cellular responses to complex perturbations in high-throughput screens. bioRxiv. 2021 doi: 10.1101/2021.04.14.439903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Brewer RC, et al. BNT162b2 vaccine induces divergent B cell responses to SARS-CoV-2 S1 and S2. Nat. Immunol. 2022;23:33–39. doi: 10.1038/s41590-021-01088-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Andreano E, et al. Hybrid immunity improves B cells and antibodies against SARS-CoV-2 variants. Nature. 2021;600:530–535. doi: 10.1038/s41586-021-04117-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Hall V, et al. Protection against SARS-CoV-2 after Covid-19 vaccination and previous infection. N. Engl. J. Med. 2022;386:1207–1220. doi: 10.1056/NEJMoa2118691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.RECOVERY Collaborative Group. Dexamethasone in hospitalized patients with Covid-19. N. Engl. J. Med. 2021;384:693–704. doi: 10.1056/NEJMoa2021436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Sinha S, et al. Dexamethasone modulates immature neutrophils and interferon programming in severe COVID-19. Nat. Med. 2022;28:201–211. doi: 10.1038/s41591-021-01576-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Aissa AF, et al. Single-cell transcriptional changes associated with drug tolerance and response to combination therapies in cancer. Nat. Commun. 2021;12:1628. doi: 10.1038/s41467-021-21884-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Guinney J, et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 2015;21:1350–1356. doi: 10.1038/nm.3967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Mehrvarz Sarshekeh A, et al. Consensus molecular subtype (CMS) as a novel integral biomarker in colorectal cancer: a phase II trial of bintrafusp alfa in CMS4 metastatic CRC. JCO. 2020;38:4084–4084. doi: 10.1200/JCO.2020.38.15_suppl.4084. [DOI] [Google Scholar]
- 127.Khaliq AM, et al. Refining colorectal cancer classification and clinical stratification through a single-cell atlas. Genome Biol. 2022;23:113. doi: 10.1186/s13059-022-02677-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Joanito I, et al. Single-cell and bulk transcriptome sequencing identifies two epithelial tumor cell states and refines the consensus molecular classification of colorectal cancer. Nat. Genet. 2022;54:963–975. doi: 10.1038/s41588-022-01100-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Litchfield K, et al. Meta-analysis of tumor- and T cell-intrinsic mechanisms of sensitization to checkpoint inhibition. Cell. 2021;184:596–614.e14. doi: 10.1016/j.cell.2021.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Li H, van der Merwe PA, Sivakumar S. Biomarkers of response to PD-1 pathway blockade. Br. J. Cancer. 2022;126:1663–1675. doi: 10.1038/s41416-022-01743-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Leader AM, et al. Single-cell analysis of human non-small cell lung cancer lesions refines tumor classification and patient stratification. Cancer Cell. 2021;39:1594–1609.e12. doi: 10.1016/j.ccell.2021.10.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Xiong D, Wang Y, You M. A gene expression signature of TREM2hi macrophages and γδ T cells predicts immunotherapy response. Nat. Commun. 2020;11:5084. doi: 10.1038/s41467-020-18546-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Kieffer Y, et al. Single-cell analysis reveals fibroblast clusters linked to immunotherapy resistance in cancer. Cancer Discov. 2020;10:1330–1351. doi: 10.1158/2159-8290.CD-19-1384. [DOI] [PubMed] [Google Scholar]
- 134.Dominguez CX, et al. Single-cell RNA sequencing reveals stromal evolution into LRRC15+ myofibroblasts as a determinant of patient response to cancer immunotherapy. Cancer Discov. 2020;10:232–253. doi: 10.1158/2159-8290.CD-19-0644. [DOI] [PubMed] [Google Scholar]
- 135.Guo X, et al. Global characterization of T cells in non-small-cell lung cancer by single-cell sequencing. Nat. Med. 2018;24:978–985. doi: 10.1038/s41591-018-0045-3. [DOI] [PubMed] [Google Scholar]
- 136.Zheng C, et al. Landscape of infiltrating T cells in liver cancer revealed by single-cell sequencing. Cell. 2017;169:1342–1356.e16. doi: 10.1016/j.cell.2017.05.035. [DOI] [PubMed] [Google Scholar]
- 137.Pittet MJ, Michielin O, Migliorini D. Clinical relevance of tumour-associated macrophages. Nat. Rev. Clin. Oncol. 2022;19:402–421. doi: 10.1038/s41571-022-00620-6. [DOI] [PubMed] [Google Scholar]
- 138.Färkkilä A, et al. Immunogenomic profiling determines responses to combined PARP and PD-1 inhibition in ovarian cancer. Nat. Commun. 2020;11:1459. doi: 10.1038/s41467-020-15315-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Jansen CS, et al. An intra-tumoral niche maintains and differentiates stem-like CD8 T cells. Nature. 2019;576:465–470. doi: 10.1038/s41586-019-1836-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Vanhersecke L, et al. Mature tertiary lymphoid structures predict immune checkpoint inhibitor efficacy in solid tumors independently of PD-L1 expression. Nat. Cancer. 2021;2:794–802. doi: 10.1038/s43018-021-00232-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Zhang K, et al. Longitudinal single-cell RNA-seq analysis reveals stress-promoted chemoresistance in metastatic ovarian cancer. Sci. Adv. 2022;8:eabm1831. doi: 10.1126/sciadv.abm1831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Candelli T, et al. Identification and characterization of relapse-initiating cells in MLL-rearranged infant ALL by single-cell transcriptomics. Leukemia. 2022;36:58–67. doi: 10.1038/s41375-021-01341-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.Pieters R, et al. A treatment protocol for infants younger than 1 year with acute lymphoblastic leukaemia (Interfant-99): an observational study and a multicentre randomised trial. Lancet. 2007;370:240–250. doi: 10.1016/S0140-6736(07)61126-X. [DOI] [PubMed] [Google Scholar]
- 144.Martin JC, et al. Single-cell analysis of Crohn’s disease lesions identifies a pathogenic cellular module associated with resistance to anti-TNF therapy. Cell. 2019;178:1493–1508.e20. doi: 10.1016/j.cell.2019.08.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Smillie CS, et al. Intra- and inter-cellular rewiring of the human colon during ulcerative colitis. Cell. 2019;178:714–730.e22. doi: 10.1016/j.cell.2019.06.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Wang Z, et al. Single-cell RNA sequencing of peripheral blood mononuclear cells from acute Kawasaki disease patients. Nat. Commun. 2021;12:5444. doi: 10.1038/s41467-021-25771-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Zhang Y, et al. Single-cell analyses of renal cell cancers reveal insights into tumor microenvironment, cell of origin, and therapy response. Proc. Natl Acad. Sci. USA. 2021;118:e2103240118. doi: 10.1073/pnas.2103240118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Tyner JW, et al. Functional genomic landscape of acute myeloid leukaemia. Nature. 2018;562:526–531. doi: 10.1038/s41586-018-0623-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Schuurhuis GJ, et al. Minimal/measurable residual disease in AML: a consensus document from the European LeukemiaNet MRD Working Party. Blood. 2018;131:1275–1291. doi: 10.1182/blood-2017-09-801498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Ediriwickrema A, et al. Single-cell mutational profiling enhances the clinical evaluation of AML MRD. Blood Adv. 2020;4:943–952. doi: 10.1182/bloodadvances.2019001181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Oren Y, et al. Cycling cancer persister cells arise from lineages with distinct programs. Nature. 2021;596:576–582. doi: 10.1038/s41586-021-03796-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152.Kim C, et al. Chemoresistance evolution in triple-negative breast cancer delineated by single-cell sequencing. Cell. 2018;173:879–893.e13. doi: 10.1016/j.cell.2018.03.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Yost KE, et al. Clonal replacement of tumor-specific T cells following PD-1 blockade. Nat. Med. 2019;25:1251–1259. doi: 10.1038/s41591-019-0522-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154.Zhang Y, et al. Single-cell analyses reveal key immune cell subsets associated with response to PD-L1 blockade in triple-negative breast cancer. Cancer Cell. 2021;39:1578–1593.e8. doi: 10.1016/j.ccell.2021.09.010. [DOI] [PubMed] [Google Scholar]
- 155.Bassez A, et al. A single-cell map of intratumoral changes during anti-PD1 treatment of patients with breast cancer. Nat. Med. 2021;27:820–832. doi: 10.1038/s41591-021-01323-8. [DOI] [PubMed] [Google Scholar]
- 156.Wu TD, et al. Peripheral T cell expansion predicts tumour infiltration and clinical response. Nature. 2020;579:274–278. doi: 10.1038/s41586-020-2056-8. [DOI] [PubMed] [Google Scholar]
- 157.Stoeckius M, et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods. 2017;14:865–868. doi: 10.1038/nmeth.4380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Peterson VM, et al. Multiplexed quantification of proteins and transcripts in single cells. Nat. Biotechnol. 2017;35:936–939. doi: 10.1038/nbt.3973. [DOI] [PubMed] [Google Scholar]
- 159.Chen S, Lake BB, Zhang K. High-throughput sequencing of the transcriptome and chromatin accessibility in the same cell. Nat. Biotechnol. 2019;37:1452–1457. doi: 10.1038/s41587-019-0290-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Ma S, et al. Chromatin potential identified by shared single-cell profiling of RNA and chromatin. Cell. 2020;183:1103–1116.e20. doi: 10.1016/j.cell.2020.09.056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161.Clark SJ, et al. scNMT-seq enables joint profiling of chromatin accessibility DNA methylation and transcription in single cells. Nat. Commun. 2018;9:781. doi: 10.1038/s41467-018-03149-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Ren X, et al. COVID-19 immune features revealed by a large-scale single-cell transcriptome atlas. Cell. 2021;184:1895–1913.e19. doi: 10.1016/j.cell.2021.01.053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163.Mathys H, et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature. 2019;570:332–337. doi: 10.1038/s41586-019-1195-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164.Melms JC, et al. A molecular single-cell lung atlas of lethal COVID-19. Nature. 2021;595:114–119. doi: 10.1038/s41586-021-03569-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 165.Ding J, et al. Systematic comparison of single-cell and single-nucleus RNA-sequencing methods. Nat. Biotechnol. 2020;38:737–746. doi: 10.1038/s41587-020-0465-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 166.Thrupp N, et al. Single-nucleus RNA-Seq is not suitable for detection of microglial activation genes in humans. Cell Rep. 2020;32:108189. doi: 10.1016/j.celrep.2020.108189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 167.Der E, et al. Tubular cell and keratinocyte single-cell transcriptomics applied to lupus nephritis reveal type I IFN and fibrosis relevant pathways. Nat. Immunol. 2019;20:915–927. doi: 10.1038/s41590-019-0386-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168.Haque A, Engel J, Teichmann SA, Lönnberg T. A practical guide to single-cell RNA-sequencing for biomedical research and clinical applications. Genome Med. 2017;9:75. doi: 10.1186/s13073-017-0467-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 169.Ding J, Sharon N, Bar-Joseph Z. Temporal modelling using single-cell transcriptomics. Nat. Rev. Genet. 2022;23:355–368. doi: 10.1038/s41576-021-00444-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170.Guillaumet-Adkins A, et al. Single-cell transcriptome conservation in cryopreserved cells and tissues. Genome Biol. 2017;18:45. doi: 10.1186/s13059-017-1171-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 171.Wilkinson MD, et al. The FAIR guiding principles for scientific data management and stewardship. Sci. Data. 2016;3:160018. doi: 10.1038/sdata.2016.18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 172.Svensson V, da Veiga Beltrame E, Pachter L. A curated database reveals trends in single-cell transcriptomics. Database. 2020;2020:baaa073. doi: 10.1093/database/baaa073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 173.Han X, et al. Construction of a human cell landscape at single-cell level. Nature. 2020;581:303–309. doi: 10.1038/s41586-020-2157-4. [DOI] [PubMed] [Google Scholar]
- 174.Tabula Sapiens Consortium. The Tabula Sapiens: a multiple-organ, single-cell transcriptomic atlas of humans. Science. 2022;376:eabl4896. doi: 10.1126/science.abl4896. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 175.Füllgrabe A, et al. Guidelines for reporting single-cell RNA-seq experiments. Nat. Biotechnol. 2020;38:1384–1386. doi: 10.1038/s41587-020-00744-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 176.Meghill C, et al. Cellxgene: a performant, scalable exploration platform for high dimensional sparse matrices. bioRxiv. 2021 doi: 10.1101/2021.04.05.438318v1. [DOI] [Google Scholar]
- 177.Li B, et al. Cumulus provides cloud-based data analysis for large-scale single-cell and single-nucleus RNA-seq. Nat. Methods. 2020;17:793–798. doi: 10.1038/s41592-020-0905-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 178.Papatheodorou I, et al. Expression Atlas update: from tissues to single cells. Nucleic Acids Res. 2020;48:D77–D83. doi: 10.1093/nar/gkz947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 179.Moreno P, et al. User-friendly, scalable tools and workflows for single-cell RNA-seq analysis. Nat. Methods. 2021;18:327–328. doi: 10.1038/s41592-021-01102-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 180.Lähnemann D, et al. Eleven grand challenges in single-cell data science. Genome Biol. 2020;21:31. doi: 10.1186/s13059-020-1926-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 181.Angerer P, et al. Single cells make big data: new challenges and opportunities in transcriptomics. Curr. Opin. Syst. Biol. 2017;4:85–91. doi: 10.1016/j.coisb.2017.07.004. [DOI] [Google Scholar]
- 182.Zhang MJ, et al. Polygenic enrichment distinguishes disease associations of individual cells in single-cell RNA-seq data. Nat. Genet. 2022;54:1572–1580. doi: 10.1038/s41588-022-01167-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 183.Efremova M, Vento-Tormo M, Teichmann SA, Vento-Tormo R. CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 2020;15:1484–1506. doi: 10.1038/s41596-020-0292-x. [DOI] [PubMed] [Google Scholar]
- 184.Fu Y, et al. Pan-cancer computational histopathology reveals mutations, tumor composition and prognosis. Nat. Cancer. 2020;1:800–810. doi: 10.1038/s43018-020-0085-8. [DOI] [PubMed] [Google Scholar]
- 185.Warnat-Herresthal S, et al. Swarm Learning as a privacy-preserving machine learning approach for disease classification. bioRxiv. 2020 doi: 10.1101/2020.06.25.171009. [DOI] [Google Scholar]
- 186.Regev A, et al. The human cell atlas. eLife. 2017;6:e27041. doi: 10.7554/eLife.27041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 187.Han L, et al. Cell transcriptomic atlas of the non-human primate Macaca fascicularis. Nature. 2022;604:723–731. doi: 10.1038/s41586-022-04587-3. [DOI] [PubMed] [Google Scholar]
- 188.Hao Y, et al. Integrated analysis of multimodal single-cell data. Cell. 2021;184:3573–3587.e29. doi: 10.1016/j.cell.2021.04.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 189.Domínguez Conde C, et al. Cross-tissue immune cell analysis reveals tissue-specific features in humans. Science. 2022;376:eabl5197. doi: 10.1126/science.abl5197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 190.Qian J, et al. A pan-cancer blueprint of the heterogeneous tumor microenvironment revealed by single-cell profiling. Cell Res. 2020;30:745–762. doi: 10.1038/s41422-020-0355-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 191.Zheng L, et al. Pan-cancer single-cell landscape of tumor-infiltrating T cells. Science. 2021;374:abe6474. doi: 10.1126/science.abe6474. [DOI] [PubMed] [Google Scholar]
- 192.Sun D, et al. TISCH: a comprehensive web resource enabling interactive single-cell transcriptome visualization of tumor microenvironment. Nucleic Acids Res. 2021;49:D1420–D1430. doi: 10.1093/nar/gkaa1020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 193.Nieto P, et al. A single-cell tumor immune atlas for precision oncology. Genome Res. 2021;31:1913–1926. doi: 10.1101/gr.273300.120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 194.Zhang F, et al. Defining inflammatory cell states in rheumatoid arthritis joint synovial tissues by integrating single-cell transcriptomics and mass cytometry. Nat. Immunol. 2019;20:928–942. doi: 10.1038/s41590-019-0378-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 195.Cao J, et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature. 2019;566:496–502. doi: 10.1038/s41586-019-0969-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 196.Cusanovich DA, et al. A single-cell atlas of in vivo mammalian chromatin accessibility. Cell. 2018;174:1309–1324.e18. doi: 10.1016/j.cell.2018.06.052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 197.Cao J, et al. Comprehensive single-cell transcriptional profiling of a multicellular organism. Science. 2017;357:661–667. doi: 10.1126/science.aam8940. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 198.Cusanovich DA, et al. The cis-regulatory dynamics of embryonic development at single-cell resolution. Nature. 2018;555:538–542. doi: 10.1038/nature25981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 199.Zhang K, et al. A single-cell atlas of chromatin accessibility in the human genome. Cell. 2021;184:5985–6001.e19. doi: 10.1016/j.cell.2021.10.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 200.Cheng J, Liao J, Shao X, Lu X, Fan X. Multiplexing methods for simultaneous large‐scale transcriptomic profiling of samples at single‐cell resolution. Adv. Sci. 2021;8:2101229. doi: 10.1002/advs.202101229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 201.Picelli S, et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods. 2013;10:1096–1098. doi: 10.1038/nmeth.2639. [DOI] [PubMed] [Google Scholar]
- 202.Hwang B, Lee JH, Bang D. Single-cell RNA sequencing technologies and bioinformatics pipelines. Exp. Mol. Med. 2018;50:1–14. doi: 10.1038/s12276-018-0071-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 203.Dobin A, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21. doi: 10.1093/bioinformatics/bts635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 204.Kaminow B, Yunusov D, Dobin A. STARsolo: accurate, fast and versatile mapping/quantification of single-cell and single-nucleus RNA-seq data. bioRxiv. 2021 doi: 10.1101/2021.05.05.442755. [DOI] [Google Scholar]
- 205.Srivastava A, Malik L, Smith T, Sudbery I, Patro R. Alevin efficiently estimates accurate gene abundances from dscRNA-seq data. Genome Biol. 2019;20:65. doi: 10.1186/s13059-019-1670-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 206.Bray NL, Pimentel H, Melsted P, Pachter L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 2016;34:525–527. doi: 10.1038/nbt.3519. [DOI] [PubMed] [Google Scholar]
- 207.Melsted P, Ntranos V, Pachter L. The barcode, UMI, set format and BUStools. Bioinformatics. 2019;35:4472–4473. doi: 10.1093/bioinformatics/btz279. [DOI] [PubMed] [Google Scholar]
- 208.Lun ATL, et al. EmptyDrops: distinguishing cells from empty droplets in droplet-based single-cell RNA sequencing data. Genome Biol. 2019;20:63. doi: 10.1186/s13059-019-1662-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 209.Muskovic W, Powell JE. DropletQC: improved identification of empty droplets and damaged cells in single-cell RNA-seq data. Genome Biol. 2021;22:329. doi: 10.1186/s13059-021-02547-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 210.Yang S, et al. Decontamination of ambient RNA in single-cell RNA-seq with DecontX. Genome Biol. 2020;21:57. doi: 10.1186/s13059-020-1950-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 211.Young MD, Behjati S. SoupX removes ambient RNA contamination from droplet-based single-cell RNA sequencing data. GigaScience. 2020;9:giaa151. doi: 10.1093/gigascience/giaa151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 212.Wolock SL, Lopez R, Klein AM. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 2019;8:281–291.e9. doi: 10.1016/j.cels.2018.11.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 213.McGinnis CS, Murrow LM, Gartner ZJ. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 2019;8:329–337.e4. doi: 10.1016/j.cels.2019.03.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 214.DePasquale EAK, et al. DoubletDecon: deconvoluting doublets from single-cell RNA-sequencing data. Cell Rep. 2019;29:1718–1727.e8. doi: 10.1016/j.celrep.2019.09.082. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 215.Lun ATL, McCarthy DJ, Marioni JC. A step-by-step workflow for low-level analysis of single-cell RNA-seq data with Bioconductor. F1000Res. 2016;5:2122. doi: 10.12688/f1000research.9501.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 216.Hafemeister C, Satija R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 2019;20:296. doi: 10.1186/s13059-019-1874-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 217.Bacher R, et al. SCnorm: robust normalization of single-cell RNA-seq data. Nat. Methods. 2017;14:584–586. doi: 10.1038/nmeth.4263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 218.Duò A, Robinson MD, Soneson C. A systematic performance evaluation of clustering methods for single-cell RNA-seq data. F1000Res. 2020;7:1141. doi: 10.12688/f1000research.15666.3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 219.Kobak D, Berens P. The art of using t-SNE for single-cell transcriptomics. Nat. Commun. 2019;10:5416. doi: 10.1038/s41467-019-13056-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 220.Becht E, et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 2019;37:38–44. doi: 10.1038/nbt.4314. [DOI] [PubMed] [Google Scholar]
- 221.Jaitin DA, et al. Dissecting immune circuits by linking CRISPR-pooled screens with single-cell RNA-Seq. Cell. 2016;167:1883–1896.e15. doi: 10.1016/j.cell.2016.11.039. [DOI] [PubMed] [Google Scholar]
- 222.Papalexi E, et al. Characterizing the molecular regulation of inhibitory immune checkpoints with multimodal single-cell screens. Nat. Genet. 2021;53:322–331. doi: 10.1038/s41588-021-00778-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 223.Yang L, et al. scMAGeCK links genotypes with multiple phenotypes in single-cell CRISPR screens. Genome Biol. 2020;21:19. doi: 10.1186/s13059-020-1928-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 224.Duan B, et al. Model-based understanding of single-cell CRISPR screening. Nat. Commun. 2019;10:2233. doi: 10.1038/s41467-019-10216-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 225.Wang R, Lin D-Y, Jiang Y. SCOPE: a normalization and copy-number estimation method for single-cell DNA sequencing. Cell Syst. 2020;10:445–452.e6. doi: 10.1016/j.cels.2020.03.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 226.Zaccaria S, Raphael BJ. Characterizing allele- and haplotype-specific copy numbers in single cells with CHISEL. Nat. Biotechnol. 2021;39:207–214. doi: 10.1038/s41587-020-0661-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 227.Zafar H, Wang Y, Nakhleh L, Navin N, Chen K. Monovar: single-nucleotide variant detection in single cells. Nat. Methods. 2016;13:505–507. doi: 10.1038/nmeth.3835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 228.Dong X, et al. Accurate identification of single-nucleotide variants in whole-genome-amplified single cells. Nat. Methods. 2017;14:491–493. doi: 10.1038/nmeth.4227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 229.Luquette LJ, Bohrson CL, Sherman MA, Park PJ. Identification of somatic mutations in single cell DNA-seq using a spatial model of allelic imbalance. Nat. Commun. 2019;10:3908. doi: 10.1038/s41467-019-11857-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 230.Singer J, Kuipers J, Jahn K, Beerenwinkel N. Single-cell mutation identification via phylogenetic inference. Nat. Commun. 2018;9:5144. doi: 10.1038/s41467-018-07627-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 231.Mallory XF, Edrisi M, Navin N, Nakhleh L. Methods for copy number aberration detection from single-cell DNA-sequencing data. Genome Biol. 2020;21:208. doi: 10.1186/s13059-020-02119-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 232.Gao R, et al. Delineating copy number and clonal substructure in human tumors from single-cell transcriptomes. Nat. Biotechnol. 2021;39:599–608. doi: 10.1038/s41587-020-00795-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 233.Patel AP, et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science. 2014;344:1396–1401. doi: 10.1126/science.1254257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 234.Petti AA, et al. A general approach for detecting expressed mutations in AML cells using single cell RNA-sequencing. Nat. Commun. 2019;10:3660. doi: 10.1038/s41467-019-11591-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 235.Vu TN, et al. Cell-level somatic mutation detection from single-cell RNA sequencing. Bioinformatics. 2019;35:4679–4687. doi: 10.1093/bioinformatics/btz288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 236.Cuomo ASE, et al. Optimizing expression quantitative trait locus mapping workflows for single-cell studies. Genome Biol. 2021;22:188. doi: 10.1186/s13059-021-02407-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 237.Stubbington MJT, et al. T cell fate and clonality inference from single-cell transcriptomes. Nat. Methods. 2016;13:329–332. doi: 10.1038/nmeth.3800. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 238.Lindeman I, et al. BraCeR: B-cell-receptor reconstruction and clonality inference from single-cell RNA-seq. Nat. Methods. 2018;15:563–565. doi: 10.1038/s41592-018-0082-3. [DOI] [PubMed] [Google Scholar]
- 239.Song L, et al. TRUST4: immune repertoire reconstruction from bulk and single-cell RNA-seq data. Nat. Methods. 2021;18:627–630. doi: 10.1038/s41592-021-01142-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 240.Upadhyay AA, et al. BALDR: a computational pipeline for paired heavy and light chain immunoglobulin reconstruction in single-cell RNA-seq data. Genome Med. 2018;10:20. doi: 10.1186/s13073-018-0528-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 241.Rizzetto S, et al. B-cell receptor reconstruction from single-cell RNA-seq with VDJPuzzle. Bioinformatics. 2018;34:2846–2847. doi: 10.1093/bioinformatics/bty203. [DOI] [PubMed] [Google Scholar]
- 242.Borcherding N, Bormann NL, Kraus G. scRepertoire: an R-based toolkit for single-cell immune receptor analysis. F1000Res. 2020;9:47. doi: 10.12688/f1000research.22139.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 243.McDavid, A., Gu, Y. & VonKaenel, E. CellaRepertorium: data structures, clustering and testing for single cell immune receptor repertoires (scRNAseq RepSeq/AIRR-seq). https://rdrr.io/bioc/CellaRepertorium (2021).
- 244.Zhang Z, Xiong D, Wang X, Liu H, Wang T. Mapping the functional landscape of T cell receptor repertoires by single-T cell transcriptomics. Nat. Methods. 2021;18:92–99. doi: 10.1038/s41592-020-01020-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 245.Buenrostro JD, et al. Single-cell chromatin accessibility reveals principles of regulatory variation. Nature. 2015;523:486–490. doi: 10.1038/nature14590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 246.Wu SJ, et al. Single-cell CUT&Tag analysis of chromatin modifications in differentiation and tumor progression. Nat. Biotechnol. 2021;39:819–824. doi: 10.1038/s41587-021-00865-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 247.Grosselin K, et al. High-throughput single-cell ChIP-seq identifies heterogeneity of chromatin states in breast cancer. Nat. Genet. 2019;51:1060–1066. doi: 10.1038/s41588-019-0424-9. [DOI] [PubMed] [Google Scholar]
- 248.Clark SJ, et al. Genome-wide base-resolution mapping of DNA methylation in single cells using single-cell bisulfite sequencing (scBS-seq) Nat. Protoc. 2017;12:534–547. doi: 10.1038/nprot.2016.187. [DOI] [PubMed] [Google Scholar]
- 249.Slavov N. Learning from natural variation across the proteomes of single cells. PLoS Biol. 2022;20:e3001512. doi: 10.1371/journal.pbio.3001512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 250.Vistain LF, Tay S. Single-cell proteomics. Trends Biochem. Sci. 2021;46:661–672. doi: 10.1016/j.tibs.2021.01.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 251.Perkel JM. Single-cell proteomics takes centre stage. Nature. 2021;597:580–582. doi: 10.1038/d41586-021-02530-6. [DOI] [PubMed] [Google Scholar]
- 252.Brinkerhoff H, Kang ASW, Liu J, Aksimentiev A, Dekker C. Multiple rereads of single proteins at single–amino acid resolution using nanopores. Science. 2021;374:1509–1513. doi: 10.1126/science.abl4381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 253.Mimitou EP, et al. Scalable, multimodal profiling of chromatin accessibility, gene expression and protein levels in single cells. Nat. Biotechnol. 2021;10:1246–1258. doi: 10.1038/s41587-021-00927-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 254.Hücker SM, et al. Single-cell microRNA sequencing method comparison and application to cell lines and circulating lung tumor cells. Nat. Commun. 2021;12:4316. doi: 10.1038/s41467-021-24611-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 255.Gawronski KAB, Kim J. Single cell transcriptomics of noncoding RNAs and their cell-specificity: Single cell transcriptomics of noncoding RNAs. WIREs RNA. 2017;8:e1433. doi: 10.1002/wrna.1433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 256.Seydel C. Single-cell metabolomics hits its stride. Nat. Methods. 2021;18:1452–1456. doi: 10.1038/s41592-021-01333-x. [DOI] [PubMed] [Google Scholar]
- 257.VanInsberghe M, van den Berg J, Andersson-Rolf A, Clevers H, van Oudenaarden A. Single-cell Ribo-seq reveals cell cycle-dependent translational pausing. Nature. 2021;597:561–565. doi: 10.1038/s41586-021-03887-4. [DOI] [PubMed] [Google Scholar]
- 258.Arrastia MV, et al. Single-cell measurement of higher-order 3D genome organization with scSPRITE. Nat. Biotechnol. 2022;40:64–73. doi: 10.1038/s41587-021-00998-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 259.Zhang R, Zhou T, Ma J. Multiscale and integrative single-cell Hi-C analysis with Higashi. Nat. Biotechnol. 2022;40:254–261. doi: 10.1038/s41587-021-01034-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 260.Chen KH, Boettiger AN, Moffitt JR, Wang S, Zhuang X. Spatially resolved, highly multiplexed RNA profiling in single cells. Science. 2015;348:aaa6090. doi: 10.1126/science.aaa6090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 261.Rodriques SG, et al. Slide-seq: a scalable technology for measuring genome-wide expression at high spatial resolution. Science. 2019;363:1463–1467. doi: 10.1126/science.aaw1219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 262.Vickovic S, et al. High-definition spatial transcriptomics for in situ tissue profiling. Nat. Methods. 2019;16:987–990. doi: 10.1038/s41592-019-0548-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 263.Liu B, Li Y, Zhang L. Analysis and visualization of spatial transcriptomic data. Front. Genet. 2022;12:785290. doi: 10.3389/fgene.2021.785290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 264.Hu J, et al. Statistical and machine learning methods for spatially resolved transcriptomics with histology. Comput. Struct. Biotechnol. J. 2021;19:3829–3841. doi: 10.1016/j.csbj.2021.06.052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 265.Zeng Z, Li Y, Li Y, Luo Y. Statistical and machine learning methods for spatially resolved transcriptomics data analysis. Genome Biol. 2022;23:83. doi: 10.1186/s13059-022-02653-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 266.Palla G, Fischer DS, Regev A, Theis FJ. Spatial components of molecular tissue biology. Nat. Biotechnol. 2022;40:308–318. doi: 10.1038/s41587-021-01182-1. [DOI] [PubMed] [Google Scholar]
- 267.Jiang R, Sun T, Song D, Li JJ. Statistics or biology: the zero-inflation controversy about scRNA-seq data. Genome Biol. 2022;23:31. doi: 10.1186/s13059-022-02601-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 268.Luecken MD, et al. Benchmarking atlas-level data integration in single-cell genomics. Nat. Methods. 2022;19:41–50. doi: 10.1038/s41592-021-01336-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 269.Tran HTN, et al. A benchmark of batch-effect correction methods for single-cell RNA sequencing data. Genome Biol. 2020;21:12. doi: 10.1186/s13059-019-1850-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 270.Song F, Chan GMA, Wei Y. Flexible experimental designs for valid single-cell RNA-sequencing experiments allowing batch effects correction. Nat. Commun. 2020;11:3274. doi: 10.1038/s41467-020-16905-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 271.Stuart T, et al. Comprehensive integration of single-cell data. Cell. 2019;177:1888–1902.e21. doi: 10.1016/j.cell.2019.05.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 272.Pliner HA, Shendure J, Trapnell C. Supervised classification enables rapid annotation of cell atlases. Nat. Methods. 2019;16:983–986. doi: 10.1038/s41592-019-0535-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 273.Kiselev VY, Yiu A, Hemberg M. scmap: projection of single-cell RNA-seq data across data sets. Nat. Methods. 2018;15:359–362. doi: 10.1038/nmeth.4644. [DOI] [PubMed] [Google Scholar]
- 274.Aran D, et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 2019;20:163–172. doi: 10.1038/s41590-018-0276-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 275.Cortal A, Martignetti L, Six E, Rausell A. Gene signature extraction and cell identity recognition at the single-cell level with Cell-ID. Nat. Biotechnol. 2021;39:1095–1102. doi: 10.1038/s41587-021-00896-6. [DOI] [PubMed] [Google Scholar]
- 276.Qiu X, et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods. 2017;14:979–982. doi: 10.1038/nmeth.4402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 277.Wolf FA, et al. PAGA: graph abstraction reconciles clustering with trajectory inference through a topology preserving map of single cells. Genome Biol. 2019;20:59. doi: 10.1186/s13059-019-1663-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 278.Street K, et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics. 2018;19:477. doi: 10.1186/s12864-018-4772-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 279.Velten L, et al. Human haematopoietic stem cell lineage commitment is a continuous process. Nat. Cell Biol. 2017;19:271–281. doi: 10.1038/ncb3493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 280.Schlitzer A, et al. Identification of cDC1- and cDC2-committed DC progenitors reveals early lineage priming at the common DC progenitor stage in the bone marrow. Nat. Immunol. 2015;16:718–728. doi: 10.1038/ni.3200. [DOI] [PubMed] [Google Scholar]
- 281.La Manno G, et al. RNA velocity of single cells. Nature. 2018;560:494–498. doi: 10.1038/s41586-018-0414-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 282.Bergen V, Lange M, Peidli S, Wolf FA, Theis FJ. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 2020;38:1408–1414. doi: 10.1038/s41587-020-0591-3. [DOI] [PubMed] [Google Scholar]
- 283.Lange M, et al. CellRank for directed single-cell fate mapping. Nat. Methods. 2022;19:159–170. doi: 10.1038/s41592-021-01346-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 284.Hänzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinforma. 2013;14:7. doi: 10.1186/1471-2105-14-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 285.Subramanian A, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA. 2005;102:15545–15550. doi: 10.1073/pnas.0506580102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 286.Fan J, et al. Characterizing transcriptional heterogeneity through pathway and gene set overdispersion analysis. Nat. Methods. 2016;13:241–244. doi: 10.1038/nmeth.3734. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 287.DeTomaso D, et al. Functional interpretation of single cell similarity maps. Nat. Commun. 2019;10:4376. doi: 10.1038/s41467-019-12235-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 288.Wei C-J, Xu X, Lo CW. Connexins and cell signaling in development and disease. Annu. Rev. Cell Dev. Biol. 2004;20:811–838. doi: 10.1146/annurev.cellbio.19.111301.144309. [DOI] [PubMed] [Google Scholar]
- 289.Noël F, et al. Dissection of intercellular communication using the transcriptome-based framework ICELLNET. Nat. Commun. 2021;12:1089. doi: 10.1038/s41467-021-21244-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 290.Browaeys R, Saelens W, Saeys Y. NicheNet: modeling intercellular communication by linking ligands to target genes. Nat. Methods. 2020;17:159–162. doi: 10.1038/s41592-019-0667-5. [DOI] [PubMed] [Google Scholar]
- 291.Jin S, et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 2021;12:1088. doi: 10.1038/s41467-021-21246-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 292.Cabello-Aguilar S, et al. SingleCellSignalR: inference of intercellular networks from single-cell transcriptomics. Nucleic Acids Res. 2020;48:e55–e55. doi: 10.1093/nar/gkaa183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 293.Wang S, Karikomi M, MacLean AL, Nie Q. Cell lineage and communication network inference via optimization for single-cell transcriptomics. Nucleic Acids Res. 2019;47:e66. doi: 10.1093/nar/gkz204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 294.Dimitrov D, et al. Comparison of methods and resources for cell-cell communication inference from single-cell RNA-Seq data. Nat. Commun. 2022;13:3224. doi: 10.1038/s41467-022-30755-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 295.Zhang Q, et al. Landscape and dynamics of single immune cells in hepatocellular carcinoma. Cell. 2019;179:829–845.e20. doi: 10.1016/j.cell.2019.10.003. [DOI] [PubMed] [Google Scholar]
- 296.Wang X, Park J, Susztak K, Zhang NR, Li M. Bulk tissue cell type deconvolution with multi-subject single-cell expression reference. Nat. Commun. 2019;10:380. doi: 10.1038/s41467-018-08023-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 297.Erdmann-Pham DD, Fischer J, Hong J, Song YS. Likelihood-based deconvolution of bulk gene expression data using single-cell references. Genome Res. 2021;31:1794–1806. doi: 10.1101/gr.272344.120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 298.Wang J, Roeder K, Devlin B. Bayesian estimation of cell type-specific gene expression with prior derived from single-cell data. Genome Res. 2021;31:1807–1818. doi: 10.1101/gr.268722.120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 299.Sokolowski DJ, et al. Single-cell mapper (scMappR): using scRNA-seq to infer the cell-type specificities of differentially expressed genes. NAR Genom. Bioinform. 2021;3:lqab011. doi: 10.1093/nargab/lqab011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 300.Newman AM, et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 2019;37:773–782. doi: 10.1038/s41587-019-0114-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 301.Luca BA, et al. Atlas of clinically distinct cell states and ecosystems across human solid tumors. Cell. 2021;184:5482–5496.e28. doi: 10.1016/j.cell.2021.09.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 302.Goldstein LD, et al. Massively parallel single-cell B-cell receptor sequencing enables rapid discovery of diverse antigen-reactive antibodies. Commun. Biol. 2019;2:304. doi: 10.1038/s42003-019-0551-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 303.Marks C, Deane CM. How repertoire data are changing antibody science. J. Biol. Chem. 2020;295:9823–9837. doi: 10.1074/jbc.REV120.010181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 304.Setliff I, et al. High-throughput mapping of B cell receptor sequences to antigen specificity. Cell. 2019;179:1636–1646.e15. doi: 10.1016/j.cell.2019.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 305.Peng L, et al. Monospecific and bispecific monoclonal SARS-CoV-2 neutralizing antibodies that maintain potency against B.1.617. Nat. Commun. 2022;13:1638. doi: 10.1038/s41467-022-29288-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 306.Castellanos-Rueda R, Di Roberto RB, Schlatter FS, Reddy ST. Leveraging single-cell sequencing for chimeric antigen receptor T cell therapies. Trends Biotechnol. 2021;39:1308–1320. doi: 10.1016/j.tibtech.2021.03.005. [DOI] [PubMed] [Google Scholar]
- 307.Li X, et al. Single-cell transcriptomic analysis reveals BCMA CAR-T cell dynamics in a patient with refractory primary plasma cell leukemia. Mol. Ther. 2021;29:645–657. doi: 10.1016/j.ymthe.2020.11.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 308.Deng Q, et al. Characteristics of anti-CD19 CAR T cell infusion products associated with efficacy and toxicity in patients with large B cell lymphomas. Nat. Med. 2020;26:1878–1887. doi: 10.1038/s41591-020-1061-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 309.Chen GM, et al. Integrative bulk and single-cell profiling of premanufacture T-cell populations reveals factors mediating long-term persistence of CAR T-cell therapy. Cancer Discov. 2021;11:2186–2199. doi: 10.1158/2159-8290.CD-20-1677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 310.Parker KR, et al. Single-cell analyses identify brain mural cells expressing CD19 as potential off-tumor targets for CAR-T immunotherapies. Cell. 2020;183:126–142.e17. doi: 10.1016/j.cell.2020.08.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 311.Jing Y, et al. Expression of chimeric antigen receptor therapy targets detected by single-cell sequencing of normal cells may contribute to off-tumor toxicity. Cancer Cell. 2021;39:1558–1559. doi: 10.1016/j.ccell.2021.09.016. [DOI] [PubMed] [Google Scholar]
- 312.Wang D, et al. CRISPR screening of CAR T cells and cancer stem cells reveals critical dependencies for cell-based therapies. Cancer Discov. 2021;11:1192–1211. doi: 10.1158/2159-8290.CD-20-1243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 313.Legut M, et al. A genome-scale screen for synthetic drivers of T cell proliferation. Nature. 2022;603:728–735. doi: 10.1038/s41586-022-04494-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 314.Kumar N, et al. Rapid single cell evaluation of human disease and disorder targets using REVEAL: SingleCellTM. BMC Genomics. 2021;22:5. doi: 10.1186/s12864-020-07300-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 315.Lachmann A, et al. Massive mining of publicly available RNA-seq data from human and mouse. Nat. Commun. 2018;9:1366. doi: 10.1038/s41467-018-03751-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 316.Collado-Torres L, et al. Reproducible RNA-seq analysis using recount2. Nat. Biotechnol. 2017;35:319–321. doi: 10.1038/nbt.3838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 317.Vivian J, et al. Toil enables reproducible, open source, big biomedical data analyses. Nat. Biotechnol. 2017;35:314–316. doi: 10.1038/nbt.3772. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 318.Soneson C, Robinson MD. Bias, robustness and scalability in single-cell differential expression analysis. Nat. Methods. 2018;15:255–261. doi: 10.1038/nmeth.4612. [DOI] [PubMed] [Google Scholar]
- 319.Stuart T, Satija R. Integrative single-cell analysis. Nat. Rev. Genet. 2019;20:257–272. doi: 10.1038/s41576-019-0093-7. [DOI] [PubMed] [Google Scholar]
- 320.Luecken MD, Theis FJ. Current best practices in single-cell RNA-seq analysis: a tutorial. Mol. Syst. Biol. 2019;15:e8746. doi: 10.15252/msb.20188746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 321.Cannoodt R, Saelens W, Deconinck L, Saeys Y. Spearheading future omics analyses using dyngen, a multi-modal simulator of single cells. Nat. Commun. 2021;12:3942. doi: 10.1038/s41467-021-24152-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 322.Treppner M, et al. Synthetic single cell RNA sequencing data from small pilot studies using deep generative models. Sci. Rep. 2021;11:9403. doi: 10.1038/s41598-021-88875-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 323.Zappia L, Phipson B, Oshlack A. Splatter: simulation of single-cell RNA sequencing data. Genome Biol. 2017;18:174. doi: 10.1186/s13059-017-1305-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 324.Saelens W, Cannoodt R, Todorov H, Saeys Y. A comparison of single-cell trajectory inference methods. Nat. Biotechnol. 2019;37:547–554. doi: 10.1038/s41587-019-0071-9. [DOI] [PubMed] [Google Scholar]
- 325.Korsunsky I, et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods. 2019;16:1289–1296. doi: 10.1038/s41592-019-0619-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 326.Mayr CH, et al. Integrative analysis of cell state changes in lung fibrosis with peripheral protein biomarkers. EMBO Mol. Med. 2021;13:e12871. doi: 10.15252/emmm.202012871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 327.Nguyen QH, Pervolarakis N, Nee K, Kessenbrock K. Experimental considerations for single-cell RNA sequencing approaches. Front. Cell Dev. Biol. 2018;6:108. doi: 10.3389/fcell.2018.00108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 328.Dal Molin A, Di Camillo B. How to design a single-cell RNA-sequencing experiment: pitfalls, challenges and perspectives. Brief. Bioinform. 2019;20:1384–1394. doi: 10.1093/bib/bby007. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The current organization of public SC data generally falls short of the FAIR principles for data stewardship in several aspects171, in particular with respect to data accessibility. Ongoing cataloguing efforts (for example, the BROAD Single Cell Portal — https://singlecell.broadinstitute.org/single_cell, spreadsheet of data set metadata172) and international collaborations to generate healthy reference databases (for example, Human Cell Landscape (HCL)173, Tabula Sapiens174 — https://tabula-sapiens-portal.ds.czbiohub.org/) provide an initial entry point for discovery of data sets. However, none of these initiatives is comprehensive, resulting in the need to manually search the publication databases (for example, PubMed) and omics repositories (for example, GEO). Without uniform metadata across these databases, the search strategy must also be varied between various resources to ensure completeness.
Within a given organization, some data are likely to be accessible only to a subset of analysts. Tracking designations flagging permissible data use in the metadata versus in an external system each present different barriers related to internal risk management and compliance, as well as to scientists and analysts seeking to use those data or to build on previously completed analyses. For public data sets, similar issues exist — data access might be restricted behind security portals, as in the case of dbGaP and EGA, because of privacy laws, contractual considerations or the sensitivity of human data. This is especially true for raw reads from full transcript protocols such as Smart-Seq2 and is equally likely to be applicable to internally generated data.