Abstract
Over the past decade, substantial progress has been made in the discovery of alleles contributing to the risk of coronary artery disease (CAD). In addition to providing causal insights into disease, these endeavours have yielded and enabled refinement of polygenic risk scores (PRS). These scores predict incident CAD in multiple cohorts and indicate the clinical response to some preventive therapies in post hoc analyses of clinical trials. These observations and the widespread ability to calculate PRS from direct-to-consumer and healthcare-associated biobanks have raised many questions about responsible clinical adoption. In this Review, we describe technical and downstream considerations for the derivation and validation of PRS and current evidence for their efficacy and safety. We discuss the implementation of these scores in clinical medicine for uses including CAD risk prediction and screening algorithms, prioritization of patient subgroups that are likely to derive benefit from treatment, and efficient prospective clinical trial designs.
Introduction
Naturally occurring genetic variation has long been hypothesized to be an important risk factor for coronary artery disease (CAD) following the observation that severe, early-onset disease often clustered in families1-3. As the germline DNA sequence is established at conception, genetic risk could even be considered the “first risk factor”4,5, establishing a lifelong baseline risk trajectory that can be altered by clinical, environmental, lifestyle and stochastic factors6.
Polygenic disease risk represents the aggregate contribution of numerous common genetic variants (minor allele frequency [MAF] >0.01) individually conferring small-to-moderate effects. Genome-wide association studies (GWAS) allow investigators to compare single-nucleotide polymorphism (SNP) prevalence systematically and individually between individuals with CAD and those without CAD to generate SNP-level association statistics. These summary statistics represent the core of polygenic risk scores (PRS) — single, normally distributed quantitative factors that combine the genetic risk attributable to many individual genetic variants — to ultimately quantify polygenic risk for individuals in external datasets.
Early studies focused on the analysis of exceptional families with extreme phenotypes to search for rare genotypes (MAF <0.005) that conferred large, deleterious effects typically through linkage analyses and positional cloning. Such mutations are typically inherited in classical Mendelian patterns and enable powerful causal inference as they typically lead to extreme phenotypes through protein-coding disruptive variation. For example, familial hypercholesterolaemia (FH) affects approximately 1 in 300 individuals and is characterized by severely elevated low-density lipoprotein cholesterol (LDL-C) concentrations and resultant severe atherosclerotic cardiovascular disease7,8. FH inheritance follows an incomplete autosomal dominant pattern, whereby each copy of the inherited allele is additively correlated with FH phenotype severity. Single disruptive mutations in the LDL receptor (LDLR) gene account for approximately 90% of FH mutations identified, with most others being apolipoprotein B (APOB) gene mutations in the LDLR-binding domain, and proprotein convertase subtilisin/kexin type 9 (PCSK9) gene gain-of-function mutations9. According to a Scientific Expert Panel convened by the Journal of the American College of Cardiology, FH genetic testing is recommended for all individuals for whom FH is suspected to prioritize early and substantial LDL-C lowering often with specialty medicines to address excess CAD risk10,11.
The first cardiovascular disease GWAS for CAD was published in 2007 14-16. Since then, the advent of low-cost, high-throughput genotyping arrays, advances in the statistical imputation of non-genotyped alleles, incorporation of increasingly diverse genetic ancestries and progressively larger discovery datasets have enabled the identification of thousands of disease-associated variants. GWAS have also revealed the genetic architecture of risk factors for cardiovascular disease including smoking17,18, renal dysfunction19,20, blood pressure21,22 and blood lipids23-25. The genetic associations are observed not only for the risk factor involved, but also appear as association signals in the GWAS for CAD. As the number of disease alleles identified increases and corresponding disease risks are refined, calculating PRS for clinical risk prediction is now feasible.
In this Review, we describe technical and downstream considerations for the derivation and validation of PRS and current evidence for their efficacy and safety. We then discuss the implementation of these scores in clinical medicine for CAD risk prediction and screening algorithms, prioritization of patient subgroups that are likely to derive the greatest benefit from treatment, and efficient design of prospective clinical trials.
Principles and methods for PRS
Liability threshold model for disease
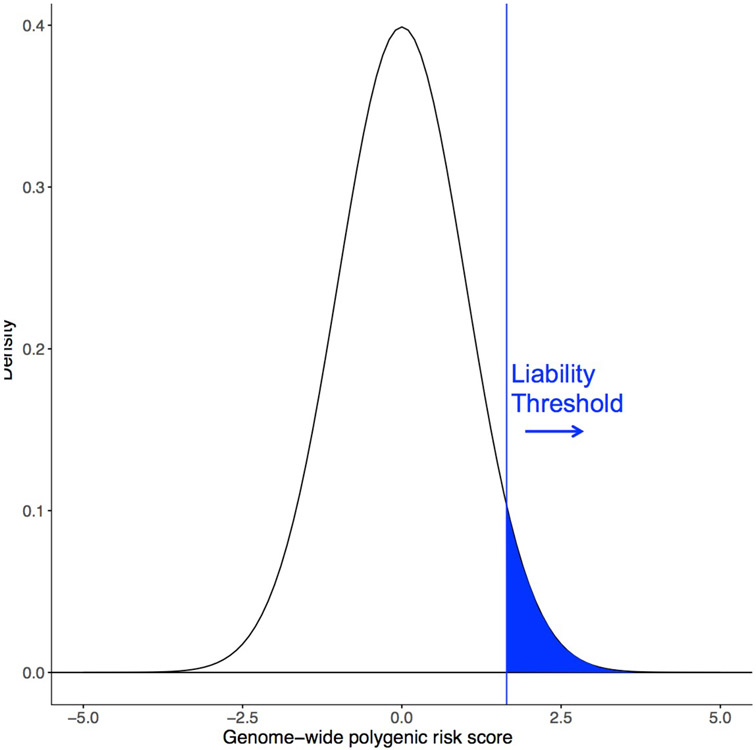
The findings of GWAS — with ever larger samples and increasing numbers of common alleles linked with cardiovascular disease — largely conform to the “common variant–common disease” hypothesis26. This hypothesis asserts that, for a fairly common condition, the genetic risk will be due largely to multiple genetic variants commonly observed in the population. When the genetic risk associated with multiple, previously identified genetic variants is combined into a single instrument, this is referred to as a PRS. A PRS captures the susceptibility to disease conferred by an individual’s genetic predisposition. Disease risk distribution for a binary outcome, such as CAD, is consistent with the ‘liability threshold’ model for disease (FIG. 1). This model proposes that a threshold exists within an underlying continuous distribution of disease risk that, once surpassed, results in disease manifestation. For the remainder of this Review, we will focus on how PRS influence risk for developing a binary disease outcome.
Fig. 1 ∣. The liability threshold model.
Risk for a given disease or binary outcome reaches a specific, unknown threshold (blue line) that, once surpassed, results in disease manifestation. Most polygenic risk score analyses model genetic risk based on the liability threshold model.
PRS are trained principally using genome-wide summary statistics from analysis of external, typically publicly available cardiovascular GWAS. To account for multiple testing and minimize the false discovery rate, GWAS include an accepted ‘genome-wide significance threshold’ of P < 5 × 10−8, which is calculated by dividing a conventional statistical significance threshold (P <0.05) by the approximate number of independent segments in the human genome (1,000,000)12. Independent replication of initial associations minimizes the risk of false discoveries that can be secondary to differences in genetic ancestry and biases associated with case ascertainment13. The resultant GWAS summary statistics define the genomic position, risk allele, corresponding effect estimate, associated standard error of the effect estimate and the P value association with the disease of interest for each allele tested across the genome in the discovery population. Each allele is inherited in classical inheritance patterns and effects are typically modelled in additive fashion. Given differences in genotyping array content, participant ancestries, and imputation panels across populations, performing appropriate quality control and harmonization of allele frequencies, genomic strand, and risk alleles from these summary statistics are critical for PRS construction. A study published in 2020 indicated that incorporating data from a transancestry meta-analysis improves the transferability of a single PRS across ancestry groups when compared with a PRS calculated from single-ancestry GWAS27. Once a set of harmonized genetic variants are identified, an investigator has multiple methodological choices for creating a PRS.
Unweighted restricted PRS
In the simplest PRS construction, an unweighted PRS represents the sum of the total number of genome-wide significant risk alleles from discovery studies present within an individual’s genome that have been previously demonstrated to be associated with disease. In this method, all risk alleles are assumed to have equal effects on the outcome of interest, and the resultant score simply reflects the number of risk alleles carried by an individual in the form of an integer. However, empirical data indicate that nearly all GWAS for cardiovascular disease have demonstrated varied effect estimates for risk alleles across the human genome. Although unweighted PRS are the simplest to construct, this method underperforms relative to weighted PRS methods28.
Weighted restricted PRS
Contemporary PRS typically utilize both the number of risk alleles and their corresponding effect size (or weights, denoted β) with the cardiovascular condition of interest. Statistically, a weighted PRS is represented by the following equation:
Where the score (φ) for each study participant (j), is the sum of the N risk alleles (x) at each variant (i), multiplied by the respective, per-allele effect size (β; calculated from GWAS summary statistics). A restricted PRS only considers genome-wide significant variants. The resultant score (φ) represents the weighted PRS for each individual that, at a population level, are normally distributed. In addition, variants that are selected for PRS should be independent, or their correlation should be accounted for through appropriate modelling.
Beyond genome-wide significant, independent variants
Early PRS analyses focused on variants that fell below the genome-wide significance threshold (P < 5 × 10−8)29. However, inclusion of additional variants that fall above this P value threshold is now known to improve PRS prediction30. The absolute P value cut-off varies on the basis of the genetic architecture of the disease of interest, the sample size of the GWAS utilized for PRS construction and the number of variants available (generally a function of the genotyping/imputation panel used for index analysis). Ultimately, there is not a single best threshold for the P value when constructing a PRS; rather, multiple thresholds are used as training parameters to derive the optimal P value cut-off.
Accounting for the correlation between associated genetic variants that are typically in close physical proximity represents another opportunity for training. Such correlation is referred to as linkage disequilibrium (LD), denoting the nonrandom association between to two alleles in the general population. Two main strategies are used to avoid excessively weighting genetic variants that are correlated to one another. The first option is to arbitrarily (or randomly) select one of the two correlated SNPs (pruning) or select the variant that is most strongly associated with the outcome (clumping). In this scenario, a final set of largely independent variants are generated, although artificial thresholds of correlation (R2) are selected in a manner similar to that for P values. The resultant P value threshold and pruning/clumping matrix of training parameters are employed together. The second option is to include both correlated genetic variants in the PRS and appropriately adjust or ‘shrink’ the effect estimates to account for the degree of correlation, level of statistical evidence, and estimates of the proportion of causal variants. Advances in computational methods have created multiple software tools31-33 for implementation of β value adjustment/shrinkage, which allow for larger numbers of genetic variants to be included in PRS construction34. Publications by Mak et al.33 and Patee & Pan35 highlight using a penalized regression model, and incorporating the LASSO (least absolute shrinkage and selection operator) or elastic net penalties, to model LD appropriately. Initial results indicate that these methods might improve accuracy of PRS in certain circumstances compared with simpler methods.
Extensions of current Bayesian PRS frameworks include external information to improve prediction. Whereas GWAS summary statistics are often overwhelmingly derived from individuals of European ancestry, LD information in training for a target non-European population could improve PRS performance in the non-European groups36. Moreover, including multiple haplotypes in multiancestry GWAS discovery efforts to ascertain transancestry effect estimates could improve portability27.
Genetic variants that exhibit associations across multiple traits or are linked to genomic functional consequences in experimental data, are more likely to be causal variants. Prioritizing pleiotropy37 or functional annotations38-40 in reweighting effects improves prediction. Restricting SNPs based on cell-type-specific regulatory annotations might also improve transancestry portability41.
Testing and validation of PRS
Expanded PRS require the consideration of diverse training parameters, therefore, sufficiently large testing and validation datasets are necessary. Typically, a series of PRS with a various P values and R2 cut-offs from an initial set of GWAS summary statistics are tested in an independent dataset without any sample overlap. This process minimizes the risk of overfitting and allows for an agnostic assessment of performance; a series of outcome and performance statistics can then be compared. In the setting of cardiovascular diseases with binary outcomes, such as CAD, investigators examine the odds ratio or hazard ratio per standard deviation change in PRS, the proportion of variance explained (pseudo-R2), the area under the receiver operating characteristic curve or C-statistic and the association P value. After a priori metrics are assessed, an optimally performing PRS is selected and used for downstream clinical analysis.
PRS reporting standards
To improve PRS research and translation, PRS reporting standards (PRS-RS) have been proposed42,43. PRS-RS metrics encompass background, study population and data, risk model development and application, risk model evaluation, limitations, clinical implications and data transparency and availability. The Polygenic Score Catalog is a repository of both score files (that is, risk alleles and weights as described above) and metadata corresponding to PRS-RS metrics. Consistency in reporting facilitates independent evaluation and improves the accuracy of downstream clinical implementation.
Clinical utility of PRS
As PRS research matures, investigators continue to explore the potential clinical utility of these scores. In the following sections, we will discuss how PRS could potentially be leveraged to enhance disease risk prediction and screening algorithms, identify patients who would derive the greatest benefit from medical therapies, and enrich clinical trial populations for individuals at greatest risk of developing disease end points.
Population-based screening
Unlike other biomarkers, germline genetics remain static and can be assayed at any time point. Cardiovascular PRS carry prognostic significance from early in life through to advanced age44-47. However, evidence suggests that, for CAD, genetic relative risk diminishes with age compared with risk conferred by nongenetic factors48. With continually expanding healthcare-associated biobanks and direct-to-consumer genetic testing, population-based implementation of PRS is conceptually feasible. Several studies have shown that a high PRS for CAD is associated with risk of CAD6, and that PRS for CAD approximates the risk discrimination observed for individual traditional clinical risk factors5. Clinical risk calculators used in practice were trained and perform optimally in middle-aged adults49. Among these individuals, a PRS for CAD modestly improves risk discrimination5,39,50-53. Whereas the prevalence of a high PRS for CAD is increased among individuals presenting with early-onset (age ≤55 years) myocardial infarction54, metrics of risk discrimination for incident CAD events among asymptomatic younger adults have not been well described. Expanding the availability of genetic data could provide complementary prognostic information at the population-level.
A single genotyping procedure with a fixed cost can generate PRS across multiple diseases beyond CAD. As low-coverage, whole-genome sequencing (which can be used to calculate PRS55 and genotype monogenic factors) becomes less expensive, PRS construction will become increasingly cost-effective for preventative health, particularly when performed early in life. Affordable, deep whole-genome sequencing will also allow for both common variant and rarer (Mendelian) alleles to be identified and potentially used to refine PRS.
Risk-enhancing factors
The 2018 AHA/ACC guidelines on the management of blood cholesterol11 and the 2019 ACC/AHA guidelines on the primary prevention of cardiovascular disease56 advanced the concept of ‘risk-enhancing factors’ to support the prescription of statins. Risk-enhancing factors, including chronic inflammatory conditions and premature menopause, are associated with increased risk of CAD in the population, independent of conventional risk factors. Among adults with an estimated 5.0–19.9% 10-year risk of atherosclerotic cardiovascular disease, an accumulation of risk-enhancing factors support a recommendation for statin administration11,56. Among subgroups clinically described to-date, the relative clinical benefit of statins is intimately tied to proportional LDL-C reduction57,58. Therefore, the anticipated clinical benefit of statins, or the ‘number needed to treat’ to prevent an event, is closely linked to predicted absolute risk. This notion underlies the concept of using risk-enhancing factors to support statin prescription.
PRS for CAD could have particular utility in the allocation of statins complementary to conventional clinical risk scores46,59-61. In epidemiological and clinical cohorts, a high PRS for CAD is associated with incident events to a similar degree as most other risk-enhancing factors5,50,60. Whereas family history of CAD is already considered a risk-enhancing factor, a high PRS for CAD is additively associated with CAD independent of a self-reported family history of CAD62. Furthermore, in a study using data from the Malmö Diet and Cancer Study50, the incremental gain in C-statistic from a PRS was higher than for any of the 11 traditional risk factors. Minimal correlation was observed between the PRS and 10-year risk assessed via the pooled cohort equations clinical risk estimator50. In 2020, Ripatti et al. demonstrated that PRS generated for LDL-C and triglyceride-rich lipoproteins were both associated with risk of CAD63. The highest 5% of PRS demonstrated an average increased CAD risk of 1.36 and 1.31 for LDL-C and triglyceride-rich lipoproteins, respectively63. When the investigators adjusted for the PRS in the association model, the associations between lipid PRS and CAD were only slightly attenuated. Notably, this these PRS for CAD were constructed using the CARDIoGRAMplusC4D GWAS64 summary statistics; whether greater attenuation would be observed when using data from a larger CAD GWAS preprint65 is unknown.
Post hoc subgroup analyses of completed primary prevention statin trials indicate that individuals with high PRS for CAD (utilizing 27-variant and 67-variant scores) derive greater clinical benefit from statins for a given amount of LDL-C reduction than those without a high PRS for CAD46,59. Therefore, a high PRS for CAD reduces the number needed to treat to prevent one event by having both prognostic and relative effects on statin response unique to other risk-enhancing factors. In geographically distinct and ethnically diverse US-based healthcare-system biobanks, 37% of middle-aged adults have a current indication for statin therapy for primary prevention of CAD60. This figure could increase by 4.1% if a PRS for CAD is also considered60. In a clinical trial, disclosure of a patient’s genetic risk in addition to a conventional 10-year risk score led to a reduction in LDL-C levels61.
Early surveillance and lifestyle modification
Current clinical risk scores are not well suited to identifying individuals who will develop premature CAD events, largely because age is the most important factor in 10-year risk prediction66. Lifetime, or 30-year, risk prediction remains challenging for several reasons —follow-up is limited in most cohorts; geographic and ethnic diversity is limited in samples with long-term follow-up; approaches to phenotyping vary over time; acquisition of clinical risk factors is time-dependent and multi-morbidity involves multiple competing risks; use of medicines modulating clinical risk factors varies during follow-up; and cohorts with prolonged follow-up are inherently noncontemporary.
Genotypes contributing to CAD risk are obtainable very early in life, even before the onset of risk factors, to provide a baseline lifetime risk trajectory4. Such information could be used to estimate absolute long-term risk, which can be refined by risk factors as they accrue and are modulated67. Theoretically, such early information could provide a strong impetus for primordial prevention — modification of risk factors even before they enter the abnormal range6.
Retrospective analyses in epidemiological data sets indicate that cardiometabolic PRS and lifestyle practices are additive with respect to cardiovascular risk29,39,68,69. These studies imply that, whereas lifestyle modification is beneficial regardless of genetic risk, absolute clinical benefit might be greater among those at heightened genetic risk because of overall greater lifetime risk. Motivation for lifestyle modification after prospective disclosure of a cardiometabolic PRS is variable70-73. Lifestyle modification programs are generally effective at preventing diseases with trivial risks in a cost-effective manner, yet incentives are not well-aligned across parties in most multi-payer health systems. Tailored investment to better promote lifestyle modification for asymptomatic younger adults, albeit those at increased genetic risk, might be an efficient and necessary approach.
Increased genetic risk of CAD might warrant selective use of cardiac imaging in asymptomatic individuals to refine risks and improve timing of preventive therapy. For example, coronary artery calcium (CAC) score is a robust predictor of intermediate-term cardiovascular disease risk74. In primary prevention guidelines, the presence of CAC supports the use of statins11. Age is a key determinant of CAC score75, posing challenges for identifying younger adults suitable for CAC scoring. Indeed, the presence of CAC in young adulthood is a strong predictor of intermediate-term and long-term cardiovascular disease risk as well as all-cause mortality76-78. Both sibling and parental history of premature CAD are independent predictors of premature CAC79. In small studies, a high PRS for CAD is independently predictive of the presence of CAC in younger adults46. Such observations suggest that a high PRS for CAD could justify early use of cardiac CT to identify the presence of incident CAC.
Such concepts in imaging-based surveillance could extend to other cardiovascular conditions. A PRS for abdominal aortic aneurysm (AAA) improved prediction beyond traditional risk factors and was independent and additive to a family history of AAA80. For conditions such as AAA, where family history is used to guide noninvasive screening programmes81, using PRS would be likely to improve the detection rate and potentially the cost-effectiveness of this strategy80. In addition, PRS can provide genetic risk information in situations where family history is not known by patients.
Early statin therapy
A single, abnormal quantitative risk factor — severely elevated LDL-C concentration (>190 mg/dl) — is sufficient to indicate statin therapy early in life11. Premature severe hypercholesterolemia portends very high risk for premature CAD, and risk is markedly attenuated with early LDL-C reduction. Among individuals with mutations for FH (that is, in LDLR, APOB or PCSK9 genes), retrospective analyses indicate an even greater relative clinical benefit from statins82. Beyond severe hypercholesterolemia, few situations exist in which statins would be guideline-indicated antecedent to a premature CAD event.
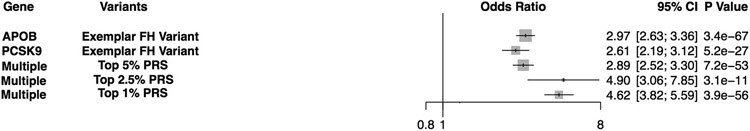
Extremely elevated PRS for CAD could provide a singular basis for early statin therapy. In a study of expanded PRS for five common diseases, including CAD, individuals within the highest 1–5% of PRS (depending on the disease) had a greater than threefold increase in the risk of developing the disease34. The risk was akin to that conferred by the presence of monogenic mutations for FH34 (FIG. 2). Similar results were later observed for incident venous thromboembolic events using data from the Women’s Health Initiative Study83, and subsequently for incident CAD events in the Malmö Diet and Cancer Study50. Taken together these findings suggest that, for cardiovascular syndromes and diseases where monogenic mutations inform clinical practice and guidelines for surveillance, extension to those with elevated PRS might be warranted. A PRS for CAD with a risk equivalency to mutations for FH could indicate the early use of statins. This notion is supported by study in preprint, which demonstrated an association between disclosure of a high PRS for CAD and the self-reported initiation of lipid-lowering therapy84.
Fig. 2 ∣. Effect estimate for polygenic risk scores.
Effect estimates for individuals at the right tail (the top 1–5%, depending on the disease) of the polygenic risk score distribution are often equivalent to, or exceed, those of well-described monogenic mutations34. In current clinical practice, the presence of a monogenic mutation can alter clinical decision making and treatment. APOB, apolipoprotein B-100; FH, familial hypercholesterolaemia; PCSK9, proprotein convertase subtilisin/kexin type 9.
Importantly, PRS represent the combination of many causal risk factors, rather than one single pathway that leads to disease. However, individuals at increased genetic risk of cardiovascular disease, whether polygenic or monogenic, will benefit from broad treatment and risk reduction strategies. Indeed, retrospective analyses indicate that both causal mutations for FH and PRS for CAD are predictive of increased relative and absolute clinical benefit from statins46,59,82. Whether these benefits extend to other preventive medicines requires further study. Such post hoc analyses are increasingly feasible as genotyping is becoming more common in clinical trial populations.
Prevention of recurrent CAD events
General consensus exists that all patients who have experienced symptomatic CAD should receive LDL-C lowering therapy. The absolute LDL-C thresholds vary across international guidelines by estimates of recurrent risk, even to <40 mg/dl in some scenarios11,85,86. The advent of costly, nonstatin LDL-C-reducing medications has prompted the use of additional factors to guide allocation. Among individuals with CAD, a high PRS is predictive of recurrent events59,87,88. In post hoc analyses of patients with appropriate indications for lipid-lowering therapy who were already receiving statins, those with a high PRS for CAD had both greater relative and absolute risk reduction from PCSK9 monoclonal antibodies when compared with placebo87,88. As described earlier, these findings are strikingly similar to LDL-C reduction with statins in primary prevention trials46,59, and suggest that PRS for CAD could be combined with traditional clinical risk factors to guide LDL-C thresholds.
Prospective validation of post hoc analyses
Several expanding, direct-to-consumer, highly phenotyped biobank genetic datasets have been created. Participants provide consent to be contacted, which enables resource-efficient, prospective identification of individuals. Such datasets require only single time-point genotyping for PRS to be applied to any heritable trait and could be leveraged to prospectively validate post hoc analyses performed within clinical trials.
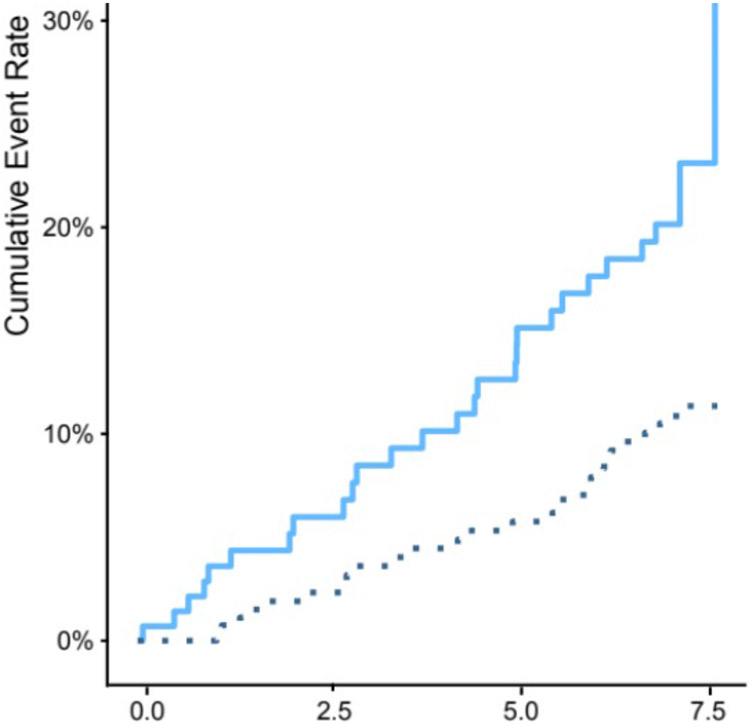
Retrospective genetic analysis of clinical trial data has demonstrated that individuals with elevated genetic risk (a high PRS) derive both greater relative and absolute risk reduction from LDL-C-lowering strategies. This finding suggests that fewer such patients would be required for inclusion in a clinical trial to reach a prespecified number of disease end points (FIG. 3). Extrapolations for post hoc analyses indicate that inclusion of patients in the top 20th percentile of PRS for CAD could improve trial efficiency by approximately threefold.
Fig. 3 ∣. Event rates for clinical trials of coronary artery disease.
Enriching clinical trials for patients with elevated polygenic risk scores (solid blue line), based on observations of absolute and relative risk reduction, would allow the prespecified number of disease end points to be reached more rapidly and, therefore, with reduced costs relative to a standard clinical trial population (dotted blue line).
Given the substantial costs involved in clinical trials driven by cardiovascular outcomes, the role of a trial to validate the interaction between PRS for CAD and LDL-C reduction merits discussion. Statins are safe and have been in widespread clinical use for several decades. In a clinical trial, disclosure of a high PRS for CAD to patients and clinicians was associated with greater LDL-C reduction61, and such disclosure has not been linked to adverse health perceptions or heightened anxiety in patients61,89. Prospective clinical trial testing should be performed to effectively evaluate the risks and costs linked to clinical disclosure of PRS for CAD, and retrospectively confirm reported outcomes. For context, it is worth noting that most clinical risk scores — including the Framingham Risk Score and Pooled Cohort Equations used to guide statin use — have not been prospectively validated in clinical trials driven by cardiovascular outcomes.
Design of clinical trials
The cost of running cardiovascular clinical trials is daunting. The mean cost of pivotal cardiovascular clinical trials for agents approved in 2015–2016 was US $157 million, which is six-times higher than the cost in other disease areas90. Reimbursement challenges and slow uptake of newly approved medicines can result in a frustratingly protracted investment cycle.
A major goal of biomedicine is to identify groups of individuals both at increased risk of disease and who are most likely to derive benefit from an intervention to reduce that risk. The ODYSSEY OUTCOMES trial88 showed that alirocumab, a monoclonal antibody targeting PCSK9, reduced cardiovascular disease risk by 15% compared with placebo among 18,924 individuals91. Among the 1,197 individuals with high PRS for CAD, a 37% reduction in cardiovascular disease risk with alirocumab was observed. The number of individuals at high genetic risk needed to treat to prevent one cardiovascular disease event was 17 compared with 64 individuals not at high genetic risk88. Similar observations were made for evolocumab, another monoclonal antibody targeting PCSK987,92. An a priori prediction and corresponding trial design could have led to a roughly fivefold reduction in trial size targeting a smaller, higher value subset of patients.
A common strategy in clinical trials of prevalent diseases, such as CAD, is to improve trial efficiency by including groups of individuals with multiple risk factors. Including individuals at high estimated risk leads to a greater number of primary outcome events over a shorter period of time. Assuming the therapy is not dependent on the enriching factors, trial efficiency is improved by having more events alone. Conventional clinical risk factors are typically used and treated similarly, meaning a set number of minimal enriching factors among a pool are required. Conventional clinical risk factors are also used because they are easily assessed, thereby maximizing screening efficiency. Although numerous novel biomarkers are associated with CAD risk, they are not typically measured and would impose additional costs. However, as noted above, increasingly large datasets are available in which clinical risk factors are included and a PRS for CAD can be calculated. Screening efficiency is maximized because an additional available enriching factor could be considered for inclusion. The power of a trial might be improved by enriching for events if all participants were required to have a high PRS for CAD. However, trial efficiency, anticipated treatment effect and market size to estimate value need to be considered before embarking on such a trial. In addition, the potential for ancestry-specific and socioeconomic disparities, as well as appropriately extrapolating effect estimates derived in patients with a high PRS to the general population, need careful consideration.
Maximizing relative treatment benefit remains a challenging concept. As discussed earlier, post hoc genetic analyses of completed clinical trials can provide a strong scientific premise for new prospective trials to validate the findings. Relative differences detected for novel or infrequently used medications would be likely to require a new trial PRS be proposed as the key including factor for an intervention. However, conventional subgroup analyses impose arbitrary dichotomies. Novel methods to assess treatment heterogeneity incorporating combinations of genetic and nongenetic factors simultaneously might better prioritize individuals benefitting from therapies in post hoc studies93, such as the evaluation of gene by therapy or gene by interaction with comorbidity. However, replicating such findings from a single completed trial is challenging because another complementary genotyped completed trial might not be available. Analyses could be considered for existing medications within genotyped healthcare associated biobanks, but would require additional consideration given the nonrandom allocation of medications.
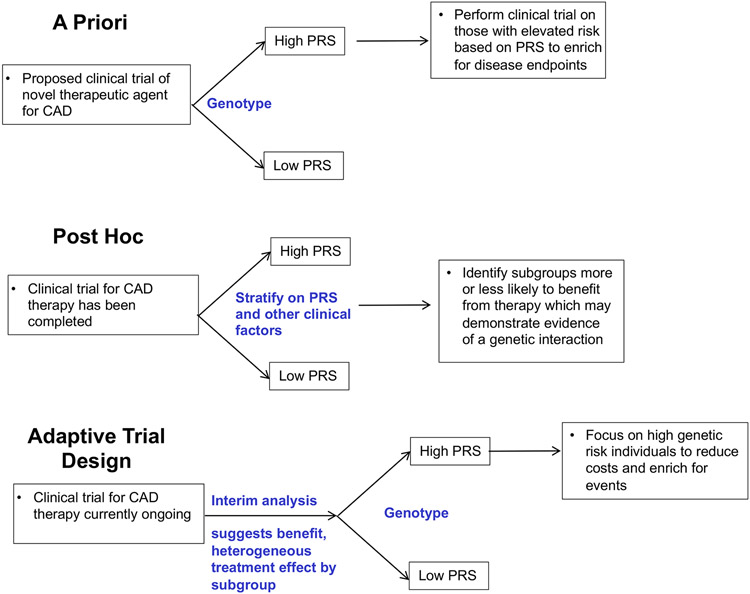
Adaptive trial designs could enable both elements of trial efficiency and discovery94. In conventional randomized clinical trials, participants are allocated to placebo or intervention on the basis of consistent inclusion criteria throughout the duration of the clinical trial enrolment period. In adaptive clinical trials, outcomes are analysed at prespecified interim time points and changes to the prespecified study design can be incorporated accordingly. As recommended by the 21st Center Cures Act of 2016, the FDA updated its guidance on adaptive designs to improve trial efficiency and reduce costs while maintaining its rigorous standards. Alterations to prespecified study design are typically guided by specific hypotheses enabling robust inference. If a PRS for CAD is used as one of the enriching factors, an adaptive protocol could allow additional participants to be enrolled if an interim analysis indicated potential benefit in patients with a high PRS. An adaptive approach would thereby obviate the need for a new trial only in patients with high PRS for CAD. A summary of the ways in which PRS might be leveraged in clinical trial design is shown in FIG. 4.
Fig. 4 ∣. Leveraging PRS for clinical trial design.
In an a priori trial design, individuals are chosen for participation based on their genotype prior to enrolment in an effort to enrich the trial for disease end points. In a post hoc design, individuals are stratified by genotype after completion of the study to delineate groups of individuals who are most likely to benefit, or derive the greatest benefit, from a therapy. In an adaptive trial design, heterogeneity observed at an interim analysis identifies individuals with a high PRS who are likely to derive the greatest benefit from a therapy. Following this, trial participants are selected based on genotype. CAD, coronary artery disease; PRS, polygenic risk scores.
As PRS for CAD become increasingly available to clinicians, incorporating treatment decisions made by clinicians into trial design could provide novel insights for existing paradigms. For example, a ‘delayed start’ trial design is being used to understand the effects of clinical PRS disclosure in Veterans Affairs Medical Centers (GenoVA NCT04331535)95. Pragmatic trials also enable the incorporation of clinician decisions in the context of parallel trial designs96.
Prioritizing patients for intervention
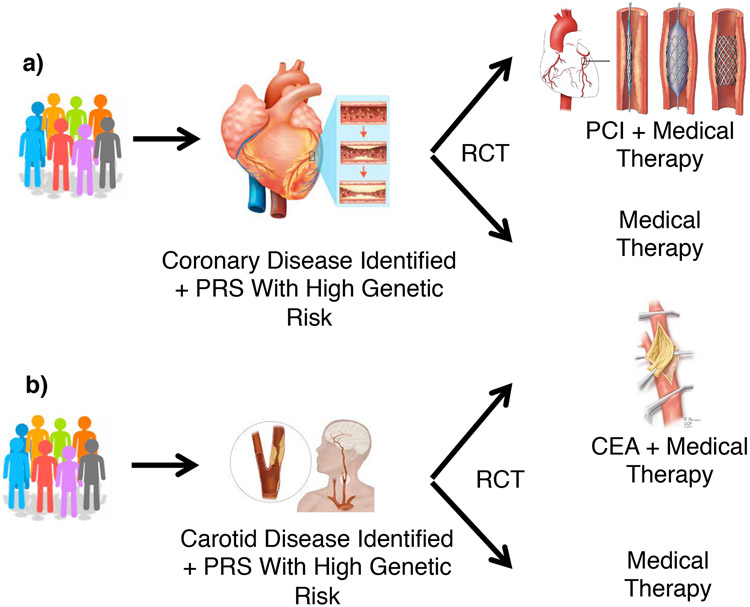
PRS data could be used to stratify patients who might benefit most from surgery or percutaneous intervention. Over the past decade, there has been substantial debate about whether the addition of revascularization to medical therapy is appropriate for patients with stable CAD. In general, contemporary trials indicate that routine revascularization is not superior to optimal medical therapy in this scenario97-99. However, traditional trial design has typically led to low enrolment rates and concerns over the extent to which findings can be generalized100. Concomitant, unremitting angina is a factor that typically favours revascularization101. Whether other conditions associated with a high risk of recurrence, such as type 2 diabetes mellitus, also favour a combined treatment strategy is not well understood. A high PRS for CAD is linked to risk of recurrence among individuals with CAD59,102. Pragmatic procedural clinical trials could flexibly incorporate clinician decisions into the study design103, enabling exploration of subgroups, such as patients with a high PRS for CAD, for intervention heterogeneity (FIG. 5a). Complementarily considering pharmacogenomic alleles used in the context of intracoronary stenting might also be well suited to this approach104,105.
Fig. 5 ∣. Prioritization of patients with high PRS for invasive therapy.
Clinical equipoise exists for the use of a ∣ coronary and b ∣carotid revascularization for specific patient populations. PRS could provide an opportunity to identifying patients likely to derive the greatest benefit from surgery or percutaneous intervention and alter the risk/benefit profile of surgery and anaesthesia. This strategy would require thorough evaluation in a RCT. CEA, carotid endarterectomy; PCI, percutaneous coronary intervention; PRS, polygenic risk scores. RCT, randomized controlled trial.
Another condition for which clinical equipoise remains is the optimal treatment selection for significant (>60–70%), asymptomatic extracranial carotid stenosis. Pivotal randomized, control trials initially demonstrated a benefit for carotid endarterectomy (CEA) over medical therapy in these patients106,107. However, medical therapy has improved dramatically since the early 2000s, and some trials have indicated that transfemoral or transcarotid carotid-artery stenting might have similar efficacy to CEA in stroke prevention108,109. By assessing polygenic risk for stroke among individuals with asymptomatic extracranial carotid stenosis at the time of diagnosis, a new trial could be designed to determine whether patients will derive greater benefit from CEA or percutaneous intervention (FIG. 5b). PRS screening could also, therefore, meaningfully alter the risk/benefit profiles of surgery and anaesthesia. The necessary infrastructure to conduct such clinical trials is increasingly available.
Considerations and challenges
An overview of current challenges and proposed solutions for the implementation of PRS in clinical practice are provided in Table 1.
Table 1 ∣.
Challenges and potential solutions for the implementation of PRS in clinical practice
| Current status of PRS | Associated challenges | Potential solutions |
|---|---|---|
| The majority of GWAS are performed in individuals of European ancestry | PRS predictions are poor in individuals of non-European ancestry and, in some cases, create potential for racial disparities | Expand discovery GWAS in non-European populations Fund cohorts with high proportions of non-European participants Improve GWAS methods for handling admixed populations Use multi-ancestry summary statistics for PRS construction |
| Lack of standard genotyping arrays and imputation panels for cohorts and large-scale biobanks | Difficulties in harmonizing PRS for broad application | Combine haplotypes across ancestries for larger imputation panels Decrease whole-genome sequencing costs to eliminate the need for future imputation |
| Summary statistics and outcome measures vary by study | Difficulties in comparing effect estimates across studies | Use a single set of PRS reporting standards for data display and publication |
| Various PRS exist for the same phenotype | Difficulties with PRS implementation | Consensus statement from societies on single PRS to be used prospectively |
| Provision of education on genetics in schools is suboptimal | Poor patient understanding of genetic risk for disease | Collaboration with genomics researchers, clinical specialists, and genetic counsellors to promote education on the genetic risk of disease |
| GINA does not cover life, disability or long-term care insurance | Potential for genetic information and PRS to be used against patients in insurance claims and in setting healthcare insurance premium costs | Additional legal protection and precedents for the handling of genetic information, risks and disclosure |
GINA, Genetic Information and Nondiscrimination Act; GWAS, genome-wide association studies; PRS, polygenic risk score.
PRS performance in diverse ancestries
Over the past decade, GWAS analyses and imputation reference panels were dominated by individuals of European genetic ancestry. Therefore, PRS construction and performance has been clearly biased towards this group110 and the predictive capacity of PRS are reduced in non-European populations80,111. For example, whereas a PRS for CAD has been shown to be associated with disease in multiple ancestry groups, predictive accuracy was unsurprisingly superior in European individuals54. Drivers of losses in predictive accuracy among non-European populations have largely been attributed to ancestry-specific differences in LD structure. A GWAS on CAD in Japanese participants suggests that a PRS constructed with transancestry summary statistics outperforms a population-specific PRS27, highlighting an approach to increasing the accuracy of current PRS for CAD in non-Europeans. Evidence from a 2021 preprint indicates that the stability of PRS predictions across ancestries is partly related to the variance explained by the discovery GWAS itself112. Another preprint suggests that novel Bayesian methods built upon the aforementioned methods could improve prediction for binary traits in non-European populations113.
In the future, the inclusion of individuals of diverse ancestries in large-scale GWAS will be integral in mitigating the risk of exacerbating disparities in PRS clinical implementation. Many US-based, federally-sponsored projects are working towards this goal, including the NHLBI Trans-Omics for Precision Medicine (TOPMed)114, VA Million Veterans Program (MVP)115, NIH AllOfUs116, and NHGRI Electronic Medical Records and Genomics (eMERGE)117. Strategies to minimize the risk of disparities in future PRS research include increasing power in GWAS of individuals with non-European ancestry to reveal novel associations118; utilization of multiancestry summary statistics for PRS construction27; increased funding and emphasis from national agencies and foundations into genetic research in non-European populations; and improvements in GWAS methodology for handling admixed populations in discovery analysis.
Assay development, validation and clinical decision support
Array platforms for genome-wide genotyping often vary by vendor, methodology and variant coverage. Imputation improves consistency across platforms, but systematic differences persist. Challenges associated with genome-wide PRS, which require millions of SNPs, across diverse array genotyping platforms could be addressed by cost-efficient, low-coverage whole-genome sequencing55. Further research in this area is warranted. In the absence of whole-genome sequencing, standards to harmonize genotyping and differences in imputation will be necessary119. However, this issue might be less important for common variants driving much of the signal and increasingly diverse imputation reference panels120.
Relative effects per normalized score standard deviation and for percentile cut-offs (defined based on the validation dataset) are represented in most studies of PRS for CAD. Some differences in relative effects across ancestries occur, although many biobanks with internally normalized scores demonstrate similar relative effects60. As ancestry is an important determinant of distribution in PRS for CAD, genetic ancestry deconvolution and adjustment could facilitate the use of these scores in admixed individuals121. Transparency of corresponding metadata into reporting of PRS for CAD will improve clinical interpretation. Absolute risk is more useful and better harmonized with CAD prevention actionability standards than relative effect. Absolute risk difference is a function of relative risk conferred by a test result and pre-test absolute risk estimation using other factors. Therefore, optimal reporting will also include nongenetic factors to guide actionability. Although PRS reporting standards for the scientific community have been proposed, the optimal approach to presenting PRS to clinicians and patients is not well understood. In addition, a multitude of PRS have been constructed over the past decade, and the lack of a single, standardized PRS for implementation makes reproducibility difficult. Standardization in reporting, diagnostics and sharing have been prioritized to address this issue42,122. Ongoing research will help clarify these outstanding questions, and a consensus around a single PRS for clinical use would allow for broad implementation, simple reporting and easy interpretation.
A dynamic reporting framework allows the incorporation of new information. GWAS continue to expand exponentially, particularly in individuals with non-European ancestry, with corresponding increases in the number of new associated alleles identified. As knowledge is gained about alleles while genotypes remain unchanged, approaches for scalable vetting, incorporating and re-reporting require several important technical and logistical considerations.
Costs
The cost of genome-wide genotyping is less than $40 per individual and continues to decrease. A single test enables PRS to be calculated for virtually any trait and does not need to be repeated. Nevertheless, initial investment in recruitment and genotyping for a healthcare-associated biobank is necessary. The direct cost of genotyping to a payer is modest, but additional costs are incurred for healthcare systems requiring storage, dynamic PRS calculation and meaningful reporting. Furthermore, the downstream costs to payers, health systems and patients are not well understood and warrant further study across multiple disciplines.
Genetic literacy
Genetic literacy for both the lay public and healthcare professionals is suboptimal. Notions of genetic determinism from basic Mendelian genetics often frame motivation and interpretation among those pursuing and interpreting genetic testing123. Given that the field of population genetics and PRS has largely accelerated over the last 20 years, public and professional education lags far behind interest and feasibility of implementation124. Collaboration with genomics researchers, clinical specialists and genetic counsellors, will be necessary, in combination with meaningful reports, to improve the interpretation of genetic risk. In addition, clinicians providing genetic information to patients need to be appropriately trained in analysis, interpretation and disclosure of genetic results.
Genetic nondiscrimination
The Genetic Information Nondiscrimination Act (GINA) of 2008 bars the use of genetic information in health insurance or employment. GINA does not cover life, disability or long-term care insurance. As proposed clinical PRS scenarios include the deployment within broadly genotyped cohorts in healthcare settings, these considerations need to be adequately communicated to patients prior to reporting. In addition, protection for patients through updating laws and legal precedents regarding handling genetic information, risks and disclosure would reduce apprehension associated with genetic testing.
Conclusions
Advances in human genetics have enabled the heritable quantification of CAD risk, in the form of a PRS. Research observations and the rapidly expanding number of individuals with available genetic data now offer tangible paths for genomic medicine to be applied broadly for CAD prevention. PRS offers the potential to further improve upon current CAD risk prediction and screening algorithms, better prioritize and select patients for a specific treatment or procedure, and update upon current prospective clinical trial designs.
Key points.
Genome-wide association studies (GWAS) demonstrate that multiple common genetic variants predispose to coronary artery disease (CAD).
Polygenic risk scores (PRS) are singular, quantitative metrics for genetic susceptibility to a disease, such as CAD.
The predictive performance of PRS for CAD is improved by incorporating evidence for association, linkage disequilibrium, anticipated functional impact and pleiotropy; transancestry PRS improve their use in diverse ancestry populations.
Post hoc analyses from completed clinical trials indicate that individuals with a high PRS for CAD derive the greatest relative and absolute benefit from cholesterol-lowering strategies.
PRS for CAD could identify individuals who would benefit from intensive lifestyle modification, imaging surveillance and early statin therapy.
PRS for CAD could identify high-risk individuals for efficient clinical trial enrolment, and evidence of heterogeneous treatment benefit could be assessed through innovative trial designs.
Glossary terms
- Elastic Net
Regularized regression method combining penalties of LASSO and ridge methods in a linear fashion
- Genotype imputation
A technique that leverages the linkage disequilibrium between genotyped and ungenotyped variants to statistically infer missing genotypes using a reference panel of genotyped individuals
- Genome-wide association study (GWAS)
A study that searches, in an agnostic fashion, for allelic association with a particular phenotype by genotyping tag single-nucleotide polymorphisms across the entire genome
- Genome-wide significance
A level of statistical significance required to establish association for a common variant in genome-wide association studies (P = 5 × 10−8)
- Haplotype
A combination of alleles transmitted together on a single chromosome
- Heritability
The proportion of the total phenotypic variation that can be attributed to additive genetic effects
- Least absolute shrinkage and selection operator (LASSO)
A modelling procedure for linear regression encouraging model sparsity
- Linkage disequilibrium (LD)
The nonrandom association of alleles at ≥2 loci because of infrequent recombination events between them
- Pleiotropy
A single gene or variant yielding two or more apparently unrelated effects
- Single-nucleotide polymorphism (SNP)
Specific substitution of a single nucleotide at a specific position in the genome
- Whole-genome sequencing
Identification of all base pairs for an individual, with subsequent mapping of contiguous reads to a reference genome sequence (for next-generation sequencing)
Related links
- Polygenic Score Catalog: https://www.pgscatalog.org/
Footnotes
Competing interests
The authors declare no competing interests.
References
- 1.White PD Genes, the heart and destiny. N. Engl. J. Med 256, 965–969 (1957). [DOI] [PubMed] [Google Scholar]
- 2.Gertler MM, Garn SM & White PD Young candidates for coronary heart disease. JAMA 147, 621–625 (1951). [DOI] [PubMed] [Google Scholar]
- 3.Marenberg ME, Risch N, Berkman LF, Floderus B & de Faire U Genetic susceptibility to death from coronary heart disease in a study of twins. N. Engl. J. Med 330, 1041–1046 (1994). [DOI] [PubMed] [Google Scholar]
- 4.Natarajan P. Polygenic risk scoring for coronary heart disease: the first risk factor. J. Am. Coll. Cardiol 72, 1894–1897 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Inouye M et al. Genomic risk prediction of coronary artery disease in 480,000 adults: implications for primary prevention. J. Am. Coll. Cardiol 72, 1883–1893 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Aragam KG & Natarajan P Polygenic scores to assess atherosclerotic cardiovascular disease risk: clinical perspectives and basic implications. Cir. Res 126, 1159–1177 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Khera AV et al. Diagnostic yield of sequencing familial hypercholesterolemia genes in patients with severe hypercholesterolemia. J. Am. Coll. Cardiol 67, 2578–2589 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hu P et al. Prevalence of familial hypercholesterolemia among the general population and patients with atherosclerotic cardiovascular disease: a systematic review and meta-analysis. Circulation 141, 1742–1759 (2020). [DOI] [PubMed] [Google Scholar]
- 9.Rader DJ, Cohen J & Hobbs HH Monogenic hypercholesterolemia: new insights in pathogenesis and treatment. J. Clin. Invest 111, 1795–1803 (2003). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Sturm AC et al. Clinical genetic testing for familial hypercholesterolemia: JACC Scientific Expert Panel. J. Am. Coll. Cardiol 72, 662–680 (2018). [DOI] [PubMed] [Google Scholar]
- 11.Grundy SM et al. 2018 AHA/ACC/AACVPR/AAPA/ABC/ACPM/ADA/AGS/APhA/ASPC/NLA/PCNA Guideline on the Management of Blood Cholesterol: A Report of the American College of Cardiology/American Heart Association Task Force on Clinical Practice Guidelines. J. Am. Coll. Cardiol 73, e285–e350 (2019). [DOI] [PubMed] [Google Scholar]
- 12.Pe'er I, Yelensky R, Altshuler D & Daly MJ Estimation of the multiple testing burden for genomewide association studies of nearly all common variants. Genet. Epidemiol 32, 381–385 (2008). [DOI] [PubMed] [Google Scholar]
- 13.Winkler TW et al. Quality control and conduct of genome-wide association meta-analyses. Nat. Protoc 9, 1192–1212 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Samani NJ et al. Genomewide association analysis of coronary artery disease. N. Engl. J. Med 357, 443–453 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.McPherson R et al. A common allele on chromosome 9 associated with coronary heart disease. Science 316, 1488–1491 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Helgadottir A et al. A common variant on chromosome 9p21 affects the risk of myocardial infarction. Science 316, 1491–1493 (2007). [DOI] [PubMed] [Google Scholar]
- 17.Liu M et al. Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat. Genet 51, 237–244 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Tobacco and Genetics Consortium. Genome-wide meta-analyses identify multiple loci associated with smoking behavior. Nat. Genet 42, 441–447 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Köttgen A et al. New loci associated with kidney function and chronic kidney disease. Nat. Genet 42, 376–384 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Hellwege JN et al. Mapping eGFR loci to the renal transcriptome and phenome in the VA Million Veteran Program. Nat. Commun 10, 3842 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Warren HR et al. Genome-wide association analysis identifies novel blood pressure loci and offers biological insights into cardiovascular risk. Nat. Genet 49, 403–415 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ehret GB et al. The genetics of blood pressure regulation and its target organs from association studies in 342,415 individuals. Nat. Genet 48, 1171–1184 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Teslovich TM et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature 466, 707–713 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Global Lipids Genetics Consortium et al. Discovery and refinement of loci associated with lipid levels. Nat. Genet 45, 1274–1283 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Klarin D et al. Genetics of blood lipids among ~300,000 multi-ethnic participants of the Million Veteran Program. Nat. Genet 50, 1514–1523 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gibson G. Rare and common variants: twenty arguments. Nat. Rev. Genet 13, 135–145 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Koyama S et al. Population-specific and trans-ancestry genome-wide analyses identify distinct and shared genetic risk loci for coronary artery disease. Nat. Genet 52, 1169–1177 (2020). [DOI] [PubMed] [Google Scholar]
- 28.Abraham G & Inouye M Genomic risk prediction of complex human disease and its clinical application. Curr. Opin. Genet. Dev 33, 10–16 (2015). [DOI] [PubMed] [Google Scholar]
- 29.Khera AV et al. Genetic risk, adherence to a healthy lifestyle, and coronary disease. N. Engl. J. Med 375, 2349–2358 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Jostins L & Barrett JC Genetic risk prediction in complex disease. Hum. Mol. Genet 20, R182–R188 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Euesden J, Lewis CM & O'Reilly PF PRSice: polygenic risk score software. Bioinformatics 31, 1466–1468 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Vilhjálmsson BJ et al. Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am. J. Hum. Genet 97, 576–592 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Mak TSH, Porsch RM, Choi SW, Zhou X & Sham PC Polygenic scores via penalized regression on summary statistics. Genet. Epidemiol 41, 469–480 (2017). [DOI] [PubMed] [Google Scholar]
- 34.Khera AV et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat. Genet 50, 1219–1224 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Pattee J & Pan W Penalized regression and model selection methods for polygenic scores on summary statistics. PLoS Comput. Biol 16, e1008271 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Márquez-Luna C, Loh PR & Price AL Multiethnic polygenic risk scores improve risk prediction in diverse populations. Genet. Epidemiol 41, 811–823 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Hu Y et al. Joint modeling of genetically correlated diseases and functional annotations increases accuracy of polygenic risk prediction. PLoS Genet. 13, e1006836 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Hu Y et al. Leveraging functional annotations in genetic risk prediction for human complex diseases. PLoS Comput. Biol 13, e1005589 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Ye Y et al. Interactions between enhanced polygenic risk scores and lifestyle for cardiovascular disease, diabetes, and lipid levels. Circ. Genom. Precis. Med 14, e003128 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Márquez-Luna C et al. LDpred-funct: incorporating functional priors improves polygenic prediction accuracy in UK Biobank and 23andMe data sets. Preprint at BioRxiv 10.1101/375337 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Amariuta T et al. Improving the trans-ancestry portability of polygenic risk scores by prioritizing variants in predicted cell-type-specific regulatory elements. Nat. Genet 52, 1346–1354 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Wand H et al. Improving reporting standards for polygenic scores in risk prediction studies. Nature 591, 211–219 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Choi SW, Mak TS & O'Reilly PF Tutorial: a guide to performing polygenic risk score analyses. Nat. Protoc 15, 2759–2772 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Riaz M et al. Effect of APOE and a polygenic risk score on incident dementia and cognitive decline in a healthy older population. Aging Cell e13384 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Neumann JT et al. Predictive performance of a polygenic risk score for incident ischemic stroke in a healthy older population. Stroke 52, 2882–2891 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Natarajan P et al. Polygenic risk score identifies subgroup with higher burden of atherosclerosis and greater relative benefit from statin therapy in the primary prevention setting. Circulation 135, 2091–2101 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Mars N et al. Polygenic and clinical risk scores and their impact on age at onset and prediction of cardiometabolic diseases and common cancers. Nat. Med 26, 549–557 (2020). [DOI] [PubMed] [Google Scholar]
- 48.Jiang X, Holmes C & McVean G The impact of age on genetic risk for common diseases. PLoS Genet. 17, e1009723 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Stone NJ et al. 2013 ACC/AHA guideline on the treatment of blood cholesterol to reduce atherosclerotic cardiovascular risk in adults: a report of the American College of Cardiology/American Heart Association Task Force on Practice Guidelines. Circulation 129, S1–S45 (2014). [DOI] [PubMed] [Google Scholar]
- 50.Hindy G et al. Genome-wide polygenic score, clinical risk factors, and long-term trajectories of coronary artery disease. Arterioscler. Thromb. Vasc. Biol 40, 2738–2746 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Elliott J et al. Predictive accuracy of a polygenic risk score-enhanced prediction model vs a clinical risk score for coronary artery disease. JAMA 323, 636–645 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Mosley JD et al. Predictive accuracy of a polygenic risk score compared with a clinical risk score for incident coronary heart disease. JAMA 323, 627–635 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Riveros-Mckay F et al. Integrated polygenic tool substantially enhances coronary artery disease prediction. Circ. Genom. Precis. Med 14, e003304 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Khera AV et al. Whole-genome sequencing to characterize monogenic and polygenic contributions in patients hospitalized with early-onset myocardial infarction. Circulation 139, 1593–1602 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Homburger JR et al. Low coverage whole genome sequencing enables accurate assessment of common variants and calculation of genome-wide polygenic scores. Genome Med. 11, 74 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Arnett DK et al. 2019 ACC/AHA Guideline on the primary prevention of cardiovascular disease: a report of the American College of Cardiology/American Heart Association Task Force on Clinical Practice Guidelines. J. Am. Coll. Cardiol 74, e177–e232 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Baigent C et al. Efficacy and safety of cholesterol-lowering treatment: prospective meta-analysis of data from 90,056 participants in 14 randomised trials of statins. Lancet 366, 1267–1278 (2005). [DOI] [PubMed] [Google Scholar]
- 58.Mihaylova B et al. The effects of lowering LDL cholesterol with statin therapy in people at low risk of vascular disease: meta-analysis of individual data from 27 randomised trials. Lancet 380, 581–590 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Mega JL et al. Genetic risk, coronary heart disease events, and the clinical benefit of statin therapy: an analysis of primary and secondary prevention trials. Lancet 385, 2264–2271 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Aragam KG et al. Limitations of contemporary guidelines for managing patients at high genetic risk of coronary artery disease. J. Am. Coll. Cardiol 75, 2769–2780 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Kullo IJ et al. Incorporating a genetic risk score into coronary heart disease risk estimates: effect on low-density lipoprotein cholesterol levels (the MI-GENES Clinical Trial). Circulation 133, 1181–1188 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Tada H et al. Risk prediction by genetic risk scores for coronary heart disease is independent of self-reported family history. Eur. Heart J 37, 561–567 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Ripatti P et al. Polygenic hyperlipidemias and coronary artery disease risk. Circ. Genom. Precis. Med 13, e002725 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.CARDIoGRAMplusC4D Consortium. A comprehensive 1,000 genomes-based genome-wide association meta-analysis of coronary artery disease. Nat. Genet 47, 1121–1130 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Aragam KG et al. Discovery and systematic characterization of risk variants and genes for coronary artery disease in over a million participants. Preprint at MedRxiv 10.1101/2021.05.24.21257377 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Karmali KN, Goff DC Jr., Ning H & Lloyd-Jones DM A systematic examination of the 2013 ACC/AHA pooled cohort risk assessment tool for atherosclerotic cardiovascular disease. J. Am. Coll. Cardiol 64, 959–968 (2014). [DOI] [PubMed] [Google Scholar]
- 67.Goldstein BA, Knowles JW, Salfati E, Ioannidis JP & Assimes TL Simple, standardized incorporation of genetic risk into non-genetic risk prediction tools for complex traits: coronary heart disease as an example. Front. Genet 5, 254 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Said MA, Verweij N & van der Harst P Associations of combined genetic and lifestyle risks with incident cardiovascular disease and diabetes in the UK Biobank Study. JAMA Cardiol. 3, 693–702 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Merino J et al. Interaction between type 2 diabetes prevention strategies and genetic determinants of coronary artery disease on cardiometabolic risk factors. Diabetes 69, 112–120 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Robinson CL et al. Disclosing genetic risk for coronary heart disease: effects on perceived personal control and genetic counseling satisfaction. Clin. Genet 89, 251–257 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Brown SN, Jouni H, Marroush TS & Kullo IJ Effect of disclosing genetic risk for coronary heart disease on information seeking and sharing: the MI-GENES Study (Myocardial Infarction Genes). Circ. Cardiovasc. Genet 10, e001613 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Grant RW et al. Personalized genetic risk counseling to motivate diabetes prevention: a randomized trial. Diabetes Care 36, 13–19 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Widén E et al. Communicating polygenic and non-genetic risk for atherosclerotic cardiovascular disease - An observational follow-up study. Preprint at MedRxiv 10.1101/2020.09.18.20197137 (2020). [DOI] [PubMed] [Google Scholar]
- 74.Detrano R et al. Coronary calcium as a predictor of coronary events in four racial or ethnic groups. N. Engl. J. Med 358, 1336–1345 (2008). [DOI] [PubMed] [Google Scholar]
- 75.McClelland RL, Chung H, Detrano R, Post W & Kronmal RA Distribution of coronary artery calcium by race, gender, and age: results from the Multi-Ethnic Study of Atherosclerosis (MESA). Circulation 113, 30–37 (2006). [DOI] [PubMed] [Google Scholar]
- 76.Miedema MD et al. Association of coronary artery calcium with long-term, cause-specific mortality among young adults. JAMA Netw. Open 2, e197440 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Paixao AR et al. Coronary artery calcium improves risk classification in younger populations. JACC Cardiovasc. Imaging 8, 1285–1293 (2015). [DOI] [PubMed] [Google Scholar]
- 78.Tota-Maharaj R et al. Coronary artery calcium for the prediction of mortality in young adults <45 years old and elderly adults >75 years old. Eur. Heart J 33, 2955–2962 (2012). [DOI] [PubMed] [Google Scholar]
- 79.Nasir K et al. Coronary artery calcification and family history of premature coronary heart disease: sibling history is more strongly associated than parental history. Circulation 110, 2150–2156 (2004). [DOI] [PubMed] [Google Scholar]
- 80.Klarin D et al. Genetic architecture of abdominal aortic aneurysm in the Million Veteran Program. Circulation 142, 1633–1646 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Owens DK et al. Screening for abdominal aortic aneurysm: US Preventive Services Task Force Recommendation Statement. JAMA 322, 2211–2218 (2019). [DOI] [PubMed] [Google Scholar]
- 82.Besseling J, Hovingh GK, Huijgen R, Kastelein JJP & Hutten BA Statins in familial hypercholesterolemia: consequences for coronary artery disease and all-cause mortality. J. Am. Coll. Cardiol 68, 252–260 (2016). [DOI] [PubMed] [Google Scholar]
- 83.Klarin D et al. Genome-wide association analysis of venous thromboembolism identifies new risk loci and genetic overlap with arterial vascular disease. Nat. Genet 51, 1574–1579 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Muse ED et al. Response to polygenic risk: results of the MyGeneRank Mobile Application-Based Coronary Artery Disease Study. Preprint at MedRxiv 10.1101/2021.04.26.21256141 (2021). [DOI] [Google Scholar]
- 85.Mach F et al. 2019 ESC/EAS Guidelines for the management of dyslipidaemias: lipid modification to reduce cardiovascular risk. Eur. Heart J 41, 111–188 (2020). [DOI] [PubMed] [Google Scholar]
- 86.Pearson GJ et al. 2021 Canadian Cardiovascular Society guidelines for the management of dyslipidemia for the prevention of cardiovascular disease in the adult. Can. J. Cardiol 37, 1129–1150 (2021). [DOI] [PubMed] [Google Scholar]
- 87.Marston NA et al. Predicting benefit from evolocumab therapy in patients with atherosclerotic disease using a genetic risk score: results from the FOURIER trial. Circulation 141, 616–623 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Damask A et al. Patients with high genome-wide polygenic risk scores for coronary artery disease may receive greater clinical benefit from alirocumab treatment in the ODYSSEY OUTCOMES trial. Circulation 141, 624–636 (2020). [DOI] [PubMed] [Google Scholar]
- 89.Knowles JW et al. Impact of a genetic risk score for coronary artery disease on reducing cardiovascular risk: a pilot randomized controlled study. Front. Cardiovasc. Med 4, 53 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Moore TJ, Zhang H, Anderson G & Alexander GC Estimated costs of pivotal trials for novel therapeutic agents approved by the US Food and Drug Administration, 2015-2016. JAMA Intern. Med 178, 1451–1457 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Schwartz GG et al. Alirocumab and cardiovascular outcomes after acute coronary syndrome. N. Engl. J. Med 379, 2097–2107 (2018). [DOI] [PubMed] [Google Scholar]
- 92.Sabatine MS et al. Evolocumab and clinical outcomes in patients with cardiovascular disease. N. Engl. J. Med 376, 1713–1722 (2017). [DOI] [PubMed] [Google Scholar]
- 93.Kent DM, Steyerberg E & van Klaveren D Personalized evidence based medicine: predictive approaches to heterogeneous treatment effects. BMJ 363, k4245 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Bothwell LE, Avorn J, Khan NF & Kesselheim AS Adaptive design clinical trials: a review of the literature and ClinicalTrials.gov. BMJ Open 8, e018320 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.D'Agostino RB Sr. The delayed-start study design. N. Engl. J. Med 361, 1304–1306 (2009). [DOI] [PubMed] [Google Scholar]
- 96.Tunis SR, Stryer DB & Clancy CM Practical clinical trials: increasing the value of clinical research for decision making in clinical and health policy. JAMA 290, 1624–1632 (2003). [DOI] [PubMed] [Google Scholar]
- 97.Maron DJ et al. Initial invasive or conservative strategy for stable coronary disease. N. Engl. J. Med 382, 1395–1407 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Al-Lamee R et al. Percutaneous coronary intervention in stable angina (ORBITA): a double-blind, randomised controlled trial. Lancet 391, 31–40 (2018). [DOI] [PubMed] [Google Scholar]
- 99.Boden WE et al. Optimal medical therapy with or without PCI for stable coronary disease. N. Engl. J. Med 356, 1503–1516 (2007). [DOI] [PubMed] [Google Scholar]
- 100.Franklin BA Lessons learned from the COURAGE trial: generalizability, limitations, and implications. Prev. Cardiol 10, 117–120 (2007). [DOI] [PubMed] [Google Scholar]
- 101.Weintraub WS et al. Effect of PCI on quality of life in patients with stable coronary disease. N. Engl. J. Med 359, 677–687 (2008). [DOI] [PubMed] [Google Scholar]
- 102.Levin MG et al. Genomic risk stratification predicts all-cause mortality after cardiac catheterization. Circ. Genom. Precis. Med 11, e002352 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Ford I & Norrie J Pragmatic trials. N. Engl. J. Med 375, 454–463 (2016). [DOI] [PubMed] [Google Scholar]
- 104.Pereira NL et al. Effect of genotype-guided oral P2Y12 inhibitor selection vs conventional clopidogrel therapy on ischemic outcomes after percutaneous coronary intervention: the TAILOR-PCI randomized clinical trial. JAMA 324, 761–771 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Pereira NL et al. Effect of CYP2C19 genotype on ischemic outcomes during oral P2Y(12) inhibitor therapy: a meta-analysis. JACC Cardiovasc. Interv 14, 739–750 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Executive Committee for the Asymptomatic Carotid Atherosclerosis Study. Endarterectomy for asymptomatic carotid artery stenosis. JAMA 273, 1421–1428 (1995). [PubMed] [Google Scholar]
- 107.Halliday A et al. Prevention of disabling and fatal strokes by successful carotid endarterectomy in patients without recent neurological symptoms: randomised controlled trial. Lancet 363, 1491–1502 (2004). [DOI] [PubMed] [Google Scholar]
- 108.Brott TG et al. Stenting versus endarterectomy for treatment of carotid-artery stenosis. N. Engl. J. Med 363, 11–23 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Schermerhorn ML et al. Association of transcarotid artery revascularization vs transfemoral carotid artery stenting with stroke or death among patients with carotid artery stenosis. JAMA 322, 2313–2322 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Claussnitzer M et al. A brief history of human disease genetics. Nature 577, 179–189 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Curtis D. Polygenic risk score for schizophrenia is more strongly associated with ancestry than with schizophrenia. Psychiatr. Genet 28, 85–89 (2018). [DOI] [PubMed] [Google Scholar]
- 112.Schultz LM et al. Stability of polygenic scores across discovery genome-wide association studies. Preprint at BioRxiv 10.1101/2021.06.18.449060 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Ruan Y et al. Improving polygenic prediction in ancestrally diverse populations. Preprint at MedRxiv 10.1101/2020.12.27.20248738 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Taliun D et al. Sequencing of 53,831 diverse genomes from the NHLBI TOPMed Program. Nature 590, 290–299 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Gaziano JM et al. Million Veteran Program: a mega-biobank to study genetic influences on health and disease. J. Clin. Epidemiol 70, 214–223 (2016). [DOI] [PubMed] [Google Scholar]
- 116.All of Us Research Program Investigators. The “All of Us” Research Program. N. Engl. J. Med 381, 668–676 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.McCarty CA et al. The eMERGE Network: a consortium of biorepositories linked to electronic medical records data for conducting genomic studies. BMC Med. Genomics 4, 13 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Vujkovic M et al. Discovery of 318 new risk loci for type 2 diabetes and related vascular outcomes among 1.4 million participants in a multi-ancestry meta-analysis. Nat. Genet 52, 680–691 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Chen SF et al. Genotype imputation and variability in polygenic risk score estimation. Genome Med. 12, 100 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Kowalski MH et al. Use of >100,000 NHLBI Trans-Omics for Precision Medicine (TOPMed) Consortium whole genome sequences improves imputation quality and detection of rare variant associations in admixed African and Hispanic/Latino populations. PLoS Genet. 15, e1008500 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Marnetto D et al. Ancestry deconvolution and partial polygenic score can improve susceptibility predictions in recently admixed individuals. Nat. Commun 11, 1628 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Lambert SA et al. The Polygenic Score Catalog as an open database for reproducibility and systematic evaluation. Nat. Genet 53, 420–425 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Carver RB, Castéra J, Gericke N, Evangelista NA & El-Hani CN Young adults' belief in genetic determinism, and knowledge and attitudes towards modern genetics and genomics: the PUGGS questionnaire. PLoS One 12, e0169808 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Kaufman DJ, Baker R, Milner LC, Devaney S & Hudson KL A survey of US adults' opinions about conduct of a Nationwide Precision Medicine Initiative® cohort study of genes and environment. PLoS One 11, e0160461(2016). [DOI] [PMC free article] [PubMed] [Google Scholar]