Abstract
We utilized positron emission tomography (PET) imaging in vivo to map the spatiotemporal biodistribution/expression of class-IIa histone deacetylases (class-IIa HDACs) in the central nervous system (CNS). Herein we report an improved radiosynthesis of [18F]NT160 using 4-hydroxy-TEMPO which led to a significant improvement in radiochemical yield and molar activity. PET imaging with [18F]NT160, a highly potent class-IIa HDAC inhibitor, led to high-quality and high-contrast images of the brain. [18F]NT160 displayed excellent pharmacokinetic and imaging characteristics: brain uptake is high in gray matter regions, tissue kinetics are appropriate for a 18F-tracer, and specific binding for class-IIa HDACs is demonstrated by self-blockade. Higher uptake with [18F]NT160 was observed in the hippocampus, thalamus, and cortex while the uptake in the cerebellum was relatively low. Overall, our current studies with [18F]NT160 will likely facilitate the development and clinical translation of PET tracers for imaging of class-IIa HDACs biodistribution/expression in cancer and the CNS.
Introduction
The histone deacetylases (HDACs) are a family of 18 enzymes, and the class-IIa HDACs are a subfamily of four members (HDACs: 4, 5, 7, and 9). The dysregulation of class-IIa HDAC in the brain has been implicated in the disorders of the central nervous system (CNS) such as stroke1−4 and Huntington’s5−7 and Alzheimer’s diseases.8−10 Furthermore, there is an emerging role for class-IIa HDAC inhibition in cancer therapies. For example, treatment of a murine model of breast cancer with TMP195, a small molecule inhibitor of class-IIa HDACs, resulted in reduced primary tumor size and decreased pulmonary metastases thus verifying the potential therapeutic value of class-IIa HDACs in human cancers.11−13 Currently, noninvasive imaging biomarkers for quantifying the class-IIa HDAC expression in tumors and in the CNS are lacking. Therefore, we are targeting class-IIa HDACs as a novel theranostic for molecular imaging and targeted therapy. The aim of our ongoing work is to develop positron emission tomography (PET) radiotracers specific to class-IIa HDAC for noninvasive quantification and mapping of the biodistribution of class-IIa HDAC protein expression in cancer and brain diseases.
Initial efforts to develop radiolabeled HDAC inhibitors were hampered by poor blood–brain barrier (BBB) permeability14−17 which limited their utility for brain imaging. [11C]Martinostat was the first successful brain penetrant class-I HDAC (HDAC1, -2, and -3) inhibitor-based tracer.18−20 Also, PET imaging in rodent, nonhuman primates, and human subjects was reported with a brain penetrant and semiselective inhibitor of HDAC6 (class-IIb HDAC).21,22 However, class-IIa HDACs are a distinctive family of proteins and class-I/IIb HDAC targeting tracers do not bind to class-IIa HDACs; therefore, they do not provide information on the expression or distribution of class-IIa HDACs in health or disease. Therefore, there is an unmet need to quantify the level of class-IIa HDAC expression accurately and specifically in health and disease states.
Furthermore, class-IIa HDAC substrate-based radiotracers exhibited fast unspecific degradation and poor pharmacokinetics in vivo that were detrimental to their clinical utility as PET radiotracers.23 The main limitation to the utility of the substrate-based radiotracers is that the radiolabeled metabolites were taken up by glial cells24,25 and the metabolite based radiotracers did not remain localized to the point of catabolism by class-IIa HDACs,20 thus confounding the PET signal and leading to an inaccurate map of biodistribution of class-IIa HDACs in the brain. Therefore, the substrate-based radiotracer inform neither on expression nor on the activity of class-IIa HDACs thus precluding their practical utility for quantitative imaging. To overcome these limitations, we developed an inhibitor-type radiotracer thus enabling accurate quantification of class-IIa HDAC protein expression/density in the brain by PET imaging.
This publication provides the first accurate and quantitative information on the biodistribution/expression (protein density) of class-IIa HDACs in the CNS. Class-IIa HDACs are known to be highly expressed in various regions of the brain, and their overexpression has been indicated in various brain disorders.26,27 Therefore, mapping the biodistribution of class-IIa HDACs in the brain will pave the way for quantitative measurement of their expression in the healthy brain compared to the disease state.
Results and Discussion
We recently reported the design, synthesis, and structure–activity relationship (SAR) study of trifluoromethyl-oxadiazole (TFMO)-containing molecules and identified highly selective and potent small molecule inhibitors of class-IIa HDACs suitable for PET tracer development.28 We also reported a late stage radiosynthesis of the TFMO moiety with [18F]-fluoride,29 and we utilized the methodology to radiosynthesize [18F]TMP195 and [18F]NT160 (Figure 1) and demonstrated their brain entry in mice.28 However, the low spatial resolution of the microPET (μPET) does not permit obtaining quantitative spatiotemporal biodistribution of class-IIa HDACs in the brain. Therefore, to overcome this limitation, we performed PET imaging studies, biodistribution, and metabolite studies with [18F]NT160 in Sprague–Dawley (SD) rats.
Figure 1.
Chemical structures of radiolabeled class-IIa HDAC inhibitors.
Improved Radiosynthesis of [18F]NT160 with the Aid of 4-Hydroxy-TEMPO
The radiosynthesis of [18F]NT160 was performed similar to that described in our previous reports28,29 with modifications as shown in Scheme 1. As shown in Table 1, performing the radiochemical reaction at 175 °C with the addition of a small quantity (0.3–0.5 mg) of 4-hydroxy-2,2,6,6-tetramethylpiperidine-1-oxyl (known as 4-hydroxy-TEMPO or TEMPOL) led to a significant improvement of the radiochemical yield to 7.5 ± 1.0% (decay corrected) and the molar activity of 0.74–1.51 GBq/μmol (20–41 mCi/μmol) compared to the previous work: the radiochemical yield was 2–5% (decay corrected), and the molar activity was 0.33–0.49 GBq/μmol (8.9–13.4 mCi/μmol).28
Scheme 1. Improved Radiosynthesis of [18F]NT160.
Table 1. Radiochemical Reaction Conditions, Radiochemical Yields (RCY), and Molar Activity.
The use of 4-hydroxy-TEMPO allowed for performing the reaction at 175 °C, a relatively higher temperature than 150 °C used in our previous report.28,29 Interestingly, increasing the radiochemical reaction temperature to 200 °C produced similar radiochemical yields but with significantly reduced molar activity. In contrast, performing the radiochemical reaction at 150 °C led to a significantly reduced radiochemical yield. We initially hypothesized that 4-hydroxy-TEMPO protected the bromo-precursor by quenching the free radicals present or released into the reaction mixture at high reaction temperature. However, remarkably, the use of TEMPO, which is also a free radical quencher, did not produce improvement in the radiochemical yield or the molar activity which suggests that another mechanism might be involved in improving the radiochemical yield and the molar activity. The bromo-precursor remained mostly intact during the radiosynthesis despite the high temperature which suggests that 4-hydroxy-TEMPO is key to protect reactants from decomposition at high temperatures. Overall, this improvement is highly significant since it cut the amount of nonradioactive mass dose of the NT160 by >50% thus enabling performing blocking (target engagement) studies in vivo and will likely enable clinical translation of [18F]NT160 by starting with larger amounts of radioactivity (i.e., >3.0 Ci). We also recommend the general utility of 4-hydroxy-TEMPO for improving the yields of radiochemical reactions that require high temperatures with specific emphasis on the 18F-trifluoromethyl moiety.
Formulation and In Situ Stability of [18F]NT160
[18F]NT160 was formulated in 20% ethanol, 20% polysorbate 80, and 60% solution of saline that contains sodium ascorbate (5 mg/mL). [18F]NT160 remained intact (stable) with >98% radiochemical purity up to 4 h postformulation based on the analysis with radioactivity high performance liquid chromatography (radio-HPLC).
PET/CT Imaging
PET and PET/CT fusion with [18F]NT160 produced high-quality and high-contrast images among various brain regions. [18F]NT160 displayed excellent pharmacokinetic and imaging characteristics: brain uptake is high in gray matter regions, leading to high-quality PET images; tissue kinetics are appropriate for an 18F-tracer and specific binding for class-IIa HDACs is demonstrated by self-blockade. Higher uptake with [18F]NT160 was observed in the hippocampus, thalamus, and cortex, with lower uptake in the striatum and cerebellum.
High contrast is demonstrated between the brain and the surrounding tissues (Figure 2A–F). Our PET data indicate that [18F]NT160 exhibited excellent in vivo characteristics, with rapid BBB penetration and observable washout pharmacokinetics. The brain uptake in the brain peaked at ∼1.25%ID/cc (SUV = 3.2) in the first five min thus exceeding the benchmark (>0.1%ID/cc) for successful CNS PET tracers.30 Remarkably, no indication of defluorination (no radioactive uptake in skull bone) was observed (Figure 2B,D). Defluorination leads to the fluoride ion accumulating in the bone of the skull, which can lead to the signal “spilling” into the brain thus confounding the quantification of changes in signal in cortical regions.31,32
Figure 2.
Representative PET and PET/CT (fusion) images with [18F]NT160 of rat brain (summed 5–30 min): (A and B) coronal, (C and D) sagittal, and (E and F) transverse, respectively.
High brain-to-heart and brain-to-muscle ratios were observed with [18F]NT160 at 10 min postinjection and were maintained for the rest of the scan time (Figure 3A and Table 2). Time–activity curves generated from the brain and heart signals showed that [18F]NT160 cleared rapidly from the heart and that the tracer uptake stabilized at 10 min postinjection with an SUV = 0.4 (Figure 3A). Additionally, [18F]NT160 exhibited relatively low uptake in the muscle, which peaked and plateaued at an SUV = 0.6. The uptake of [18F]NT160 in muscle is likely specific and was displaceable (Figure S4), which is consistent with the previous reports that class-IIa HDACs are expressed in the skeletal muscle.33,34 However, the uptake in the periphery is likely confounded by the nonspecific uptake from the radiometabolite fraction in the blood which constitutes the major radioactive fraction in the blood (discussed in the radiometabolism section 2.7). Therefore, it is unclear as to whether the radiometabolite in the blood also contributed to the uptake in the muscle. Nevertheless, our imaging data demonstrate for the first time that the protein density/expression of class-IIa HDACs in normal/healthy muscle is low when compared to the brain.
Figure 3.
Time–activity curves of [18F]NT160 obtained from dynamic imaging for 60 min. (A) Brain, muscle, and heart; (B) hippocampus (Hip), cortex (Cortx), thalamus (TH), striatum (Str), and cerebellum (Cer).
Table 2. Biodistribution of [18F]NT160 (SUV ± SEM) in Rat Brain, Heart, and Muscle.
| time
after tracer injection |
||||
|---|---|---|---|---|
| 1 min | 10 min | 30 min | 60 min | |
| brain | 2.96 ± 0.95 | 2.40 ± 0.44 | 1.57 ± 0.29 | 1.11 ± 0.26 |
| heart | 3.71 ± 2.29 | 0.51 ± 0.27 | 0.52 ± 0.16 | 0.48 ± 0.18 |
| muscle | 0.40 ± 0.29 | 0.59 ± 0.17 | 0.59 ± 0.15 | 0.59 ± 0.15 |
| hippocampus | 2.37 ± 0.74 | 2.86 ± 0.28 | 1.90 ± 0.27 | 1.34 ± 0.23 |
| cerebellum | 3.01 ± 1.08 | 1.91 ± 0.89 | 1.08 ± 0.25 | 0.76 ± 0.18 |
| hippocampus-to-cerebellum ratio | 0.79 ± 0.91 | 1.50 ± 0.59 | 1.76 ± 0.26 | 1.76 ± 0.21 |
| brain-to-heart ratio | 0.80 ± 1.62 | 4.71 ± 0.35 | 3.01 ± 0.23 | 2.29 ± 0.22 |
| brain-to-muscle ratio | 7.40 ± 0.62 | 4.01 ± 0.31 | 2.66 ± 0.22 | 1.88 ± 0.21 |
Furthermore, the time–activity curves (Figure 3B) generated from dynamic PET imaging of the brain regions of interest showed that [18F]NT160 displayed heterogeneous uptake in the brain with higher tracer uptake in the hippocampus, thalamus, cortex, and brainstem compared to the cerebellum. The high contrast was observed among brain regions at 10 min postinjection and was sustained for 60 min postinjection. Altogether, these data strongly indicate that [18F]NT160 displayed an excellent pharmacokinetic profile in vivo with high brain-to-heart, brain-to-muscle, and hippocampus-to-cerebellum ratios maintained throughout the entire 60 min of the scan (Table 2). Based on the time activity curves (Figure 2B), the highest contrast among brain regions can be obtained by performing the PET scan in a 10–60 min time frame.
Spatiotemporal Biodistribution of [18F]NT160 as a Surrogate Biomarker of Class-IIa HDAC Expression (Protein Density) in the Brain
PET imaging (Figure 4) with [18F]NT160 showed a high contrast among various regions of the brain with higher uptake observed in the hippocampus, thalamus, brainstem, and cortex and relatively lower uptake in the striatum and cerebellum. Our in vivo findings provide the first accurate and quantitative map of the biodistribution/expression (protein density) patterns of class-IIa HDACs in the CNS in its entirety.
Figure 4.
Representative coronal PET images (summed, 5–30 min) with [18F]NT160 with the corresponding rat brain regions. Crt: Cortex, Cer: cerebellum, Th: thalamus, Hip: hippocampus, Str: striatum. The alignment for the images with the rat brain atlas was used to identify these regions of interest.35,36
Our data are consistent with previous reports on the heterogeneous biodistribution of class-IIa HDACs in the rodents’ brain.37,38 However, the main limitation of the previous reports is the reliance on semiquantitative or qualitative data to delineate the class-IIa HDAC biodistribution in the brain. HDAC4 was reported to be highly expressed in some mouse brain regions than in others and HDAC4 was absent from the white matter.38 Moreover, in situ hybridization, which was used to determine the HDAC isoforms expression in >50 rat brain regions,37 showed that HDAC11, -3, -5, and -4 are expressed most highly, and HDAC10, -9, and -7 have the lowest expression levels throughout the brain. In agreement with our imaging data in rats, the expression of class-IIa HDAC4 and -5 was high in the rat cerebral cortex, thalamus, and hippocampus. In the cerebellum, HDAC4 and -5 expressions were limited solely to the granule cell layer which can explain the relatively low tracer accumulation in the cerebellum. PET does not detect subcellular distribution of proteins but rather the protein density distribution in a specific region. Therefore, it is likely that the protein density class-IIa HDAC is less in the cerebellum compared to the hippocampus, thalamus, and brainstem. These results are in line with our in vitro autoradiography and histology with HDAC4 (Figure 5C). It is important to emphasize that PET is quantitative and more accurate than the available data on class-IIa HDAC expression in the brain which underscore the significance of our approach to noninvasively provide quantitative information on the level of class-IIa HDAC in vivo in real time.
Figure 5.
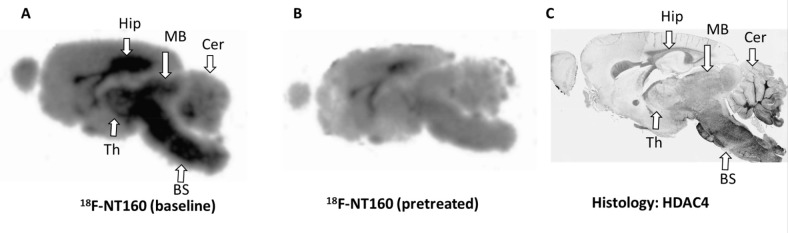
(A) In vitro autoradiography with [18F]NT160 at baseline and (B) after pretreatment with NT160 (1.0 μM). (C) Brain histology obtained with HDAC4 antibody.
In Vitro Autoradiography and Histology
We performed in vitro autoradiography with [18F]NT160 and histology with HDAC4 (Figure 5). In vitro autoradiography with [18F]NT160 showed that the highest uptake in the brain was observed in the hippocampus, midbrain, and brainstem with moderate uptake observed in the rest of the brain. Brain uptake was significantly reduced by competitive blocking with NT160 (1.0 μM). Significant reduction in the tracer uptake was observed in the hippocampus, midbrain, and brainstem. Also, the tracer uptake was reduced to a lesser extent in other brain regions. This data strongly indicate the brain uptake [18F]NT160 was displaceable and specific to class-IIa HDACs. We also performed histology with HDAC4 antibody which also confirmed high HDAC4 expression in the hippocampus, midbrain, and brainstem (Figure 5). It is important to note that NT160 is somewhat selective for HDAC4 (∼15 folds) over other class-IIa HDAC isoforms. Therefore, it is not surprising that the in vitro autoradiography with [18F]NT160 is skewed toward HDAC4 and likely is representative of HDAC4 biodistribution in the brain. Preliminary histology with HDAC5 antibody showed high expression in the cortex and brainstem and lower expression was observed in the hippocampus.
Specific Binding/Occupancy of [18F]NT160 In Vivo
[18F]NT160 exhibited inherently moderate specific activity associated with labeling the [18F]-trifluoromethyl moiety. The low specific activity means the tracer contains a high mass dose of the NT160 that could occupy the target causing self-blocking and may also produce pharmacologic effects. Brain uptake was also reduced in vivo (Figure 6) when comparing time–activity curves obtained from the tracer administered at the baseline with images obtained from the tracer coadmistered with 1.0 mg/kg of the nonradioactive NT160 (0.5 mg/kg in addition to 0.5 mg/kg calculated from the specific activity). Self-blocking with the unlabeled NT160 resulted in a significant decrease in the tracer uptake in the whole brain (30% reduction in the SUV) which demonstrates that the binding/occupancy of [18F]NT160 to class-IIa HDACs in vivo is specific and displaceable (Figure 6A). However, the overall decrease in radioactivity in the whole brain is rather simplistic; therefore, kinetic modeling of the PET data will be needed to accurately quantify the decrease in the nondisplaceable binding potential (BPND) which is more informative of the extent of tracer binding/displacement. Also, as was discussed above, the expression/density of class-IIa HDAC in the brain is heterogeneous, and therefore regional displacement is more informative on the extent of class-IIa HDAC biodistribution in the brain than the displacement in the whole brain.
Figure 6.
Time–activity curves obtained from dynamic imaging for 60 min: (A) whole brain; (B) hippocampus and cerebellum at baseline and after self-blocking. Cer: cerebellum, Hip: hippocampus.
The highest tracer uptake and consequently the largest reduction in tracer uptake due to blocking was observed in the hippocampus thus indicating high expression/density of class-IIa HDACs (Figure 6B). In contrast, the uptake of [18F]NT160 was lowest in the cerebellum and the uptake was somewhat displaceable, however to a lesser ratio than the hippocampus (Figure 6B). Previous literature reports point to expression of class-IIa HDACs in the cerebellum localized to the Purkinje cells in the granule cell layer.37,38 The higher reduction in tracer uptake in the hippocampus by blocking is reflective of high occupancy and high density of class-IIa HDACs in the hippocampus. In contrast, the lower uptake and lower reduction in tracer uptake in the cerebellum is reflective of the low density of class-IIa HDACs in the cerebellum. Our in vivo findings are supported with in vitro autoradiography which showed high [18F]NT160 accumulation in the hippocampus relative to the cerebellum (Figure 5A). Also, histology with HDAC4 antibody further supports the in vivo findings. Therefore, it is unclear whether the cerebellum can be utilized as a reference region for future kinetic modeling and quantification of the PET data. The tracer uptake in the cerebellum seems to be heterogeneous which opens the possibility for selecting a subregion with nondisplaceable tracer uptake as a reference region. Future studies in larger animals will likely identify additional regions suitable for utility as a reference region for kinetic modeling. Otherwise, arterial blood sampling will be needed to obtain an input function to support compartmental modeling methods.
Radiometabolism of [18F]NT160
The extent of radiometabolism of [18F]NT160 in the rat brain and plasma at 60 min postinjection is shown in the radio-HPLC chromatogram (Figure 7). Analytical radio-HPLC revealed that the parent [18F]NT160 remained mostly intact (83.0 ± 3%) in the brain during the entire scan time of 60 min postinjection (Figure 7A). In addition to the parent [18F]NT160, two polar radiometabolites M1 (6.0 ± 1%) and M2 (11.0 ± 3%) were also present in brain homogenates albeit in relatively smaller quantities (Figure 7A). Radio-HPLC analysis of the plasma at 60 min postinjection showed a smaller fraction of the parent [18F]NT160 (30 ± 2%) and a single polar radiometabolite (70 ± 2%) in which the elution time matches the M1 radiometabolite observed in the brain (Figure 7B). The lack of M2 in the plasma suggests that M2 was produced by radiometabolism in the brain. This also indicates that M1 likely originated in the blood and crossed the BBB and accumulated in the brain. Therefore, it was imperative to investigate and decipher the identity of M1 to ensure that the presence of M1 does not confound the PET data quantification by contributing to the class-IIa HDAC signal.
Figure 7.
Analytical HPLC chromatograms of [18F]NT160 obtained from (A) brain homogenates and (B) plasma. (C) Proposed blood metabolic pathway for M1 (radiometabolism of [18F]NT160 leads to [18F]1 (M1). (D and E) Inhibition of HDAC activity by compound 1 (M1) in HT-29 cells: (D) Inhibition of class-IIa HDAC and (E) inhibition of class-I/IIb. Compound 1 was screened side-by-side with SAHA and NT160.
We surmised that M1 arises from the cleavage of the amide bond thus producing 4-(5-[18F]-(trifluoromethyl)-1,2,4-oxadiazol-3-yl)benzoic acid ([18F]1) (Figure 7A). We then investigated whether the nonradioactive 4-(5-(trifluoromethyl)-1,2,4-oxadiazol-3-yl)benzoic acid (1) can inhibit HDACs using the cell-based assay similar to that described in our previous report.28 Our data demonstrate that the IC50 of compound 1 is >10.0 μM (Figure 6D,E and Figure S3) for all HDAC classes, and thus compound 1 is not an effective inhibitor of HDACs. Compound 1 exhibited poor inhibition (∼10%) of class-I, class-IIb, and IIa HDAC activities (Figure 7B,C) when compared side-by-side with SAHA (panHDAC inhibitor with higher selectivity to class-I and class-IIb HDACs39) and the parent NT160 (selective class-IIa HDAC).28 The weak affinity of compound 1 to class-I and class-IIb HDACs is highly desirable, since off target binding to these proteins that are expressed in the brain18,37 may complicate the PET data quantification.
Next, we produced [18F]1 in two radiosynthetic steps similar to the radiosynthesis of [18F]NT160 and that described in our previous report29 (details are provided in the SI). Dynamic PET imaging obtained with [18F]1 for 60 min revealed that [18F]1 can cross the BBB albeit in relatively low brain uptake of 0.2%ID/cc (SUV= 0.4) (Figure 8) compared to [18F]NT160. Critically, [18F]1 displayed slow pharmacokinetics and uniform uptake in the brain and therefore is not expected to contribute to the regional biodistribution of [18F]NT160;; however, [18F]1 (M1) is expected to slightly increase the overall volume of distribution (VT) in the whole brain. The time–activity curve of [18F]1 (M1) further confirmed the relatively low brain uptake of [18F]1 compared to the parent [18F]NT160. Radio-HPLC and radioactive thin layer chromatography (R-TLC) analysis showed that parent [18F]1 alone was present in the brain homogenates at 30 min postinjection. Further analysis by R-TLC indicates that [18F]1 is in fact the polar radiometabolite (M1) that was observed in brain homogenates of [18F]NT160 (Figure S6). Deciphering the structures of M1 and M2 will be critical to design the next generation of class-IIa HDAC PET tracers by selecting additional radiolabeling sites (i.e., repositioning of the [18F]fluoride or by incorporating 11C radionuclide).
Figure 8.
Representative PET, CT, and PT/CT images (summed, 0–60 min) with [18F]1: (A and B) coronal, (C and D) sagittal, and (E and F) transverse, respectively. (G) Time–activity curve of [18F]1 in rat brain.
The above data indicate that [18F]1 (M1) is not expected to contribute to the specific binding of class-IIa HDAC or the nonspecific and off target binding to other HDACs in vivo. [18F]1 (M1) displayed uniform uptake across the brain, and therefore the presence of [18F]1 (M1) in the brain is of less concern, as a reference tissue model can be reliably applied.40 Moreover, [18F]1 displayed a uniform distribution in the brain, despite containing both the TFMO and the linker pharmacophores of [18F]NT160, thus further demonstrating and supporting the specificity of [18F]NT160 to class-IIa HDACs in the brain. The identity of M2 has not been determined yet; however, it is likely to have arisen in part from metabolism by demethylation of the methylamine of [18F]NT160. Further studies are needed to confirm this speculation. However, the absence of M2 in the plasma and the presence of M2 solely in the brain in relatively smaller quantity indicate that M2 is unlikely to confound the PET data quantification.
Conclusions
In summary, we reported a 4-hydroxy-TEMPO-assisted radiosynthesis of [18F]NT160, a radioligand for molecular imaging of class-IIa HDACs, a key component of the epigenetic machinery. PET imaging studies with [18F]NT160, a highly potent and selective class-IIa HDAC inhibitor, demonstrated a favorable profile for in vivo imaging and is a useful CNS lead PET tracer. [18F]NT160 displayed excellent pharmacokinetic and imaging characteristics: brain uptake is high in gray matter regions, leading to high-quality PET images; tissue kinetics are appropriate for an 18F-tracer, and specific binding for class-IIa HDAC is demonstrated by self-blockade in vivo and in vitro using autoradiography. In vivo molecular imaging with [18F]NT160 provided the first map of distribution of class-IIa HDAC density in the brain, and therefore our studies will likely pave the way for accurate comparison of class-IIa expression in disease and health states. However, the moderate specific activity of [18F]NT160 is a limitation due to the presence of a high mass dose of NT160 that may affect occupancy of class-IIa HDAC in the brain and the periphery and thus can impact the overall brain uptake and may elicit pharmacological response. Therefore, further modifications of [18F]NT160 by repositioning the radiolabel to overcome the inherent moderate specific activity are ongoing and will be reported in due course.
Experimental Section
General Information
Solvents and starting material were obtained from commercial sources and were used as received. Radio-HPLC was performed with a 1260 series pump (Agilent Technologies, Stuttgart, Germany) with a built-in UV detector operated at 254 nm and a radioactivity detector with a single-channel analyzer (labLogic) using a semipreparative C18 reverse-phase column (00G-4041-N0: Luna 5 μm, 250 × 10 mm, Phenomenex) and an analytical C18 column (5 μm, 150 × 4.6 mm, ASCENTIS RP-AMIDE, Sigma). An acetonitrile/ammonium acetate buffer (MeCN/NH4OAc: 20 mM) solvent was used for quality control analyses at a flow of 1 mL/min. R-TLC was performed using TLC aluminum sheets and silica gel 60 matrix with fluorescent indicator.
Chemistry and Radiochemistry
The synthesis NT160 was performed according to our previous report.29 The purity of NT160 was >95%, and the radiochemical purity of [18F]NT160 was >97% (HPLC).
Radiosynthesis of [18F]NT160
The [18F]F– was trapped on a QMA cartridge and then eluted to a V-vial (Wheaton) with 80% acetonitrile and 20% water (1.0 mL) solution that contained kryptofix2.2.2 (12.0 mg) and Cs2CO3 (1.0 mg). The solvent was removed under a stream of argon at 110 °C. Water residue was removed azeotropically with the addition of acetonitrile (3 × 1.0 mL) and repeated drying under a stream of argon at 110 °C.
The bromo-precursor (6–8 mg) mixed with 4-hydroxy-TEMPO (0.3–0.5 mg) was dissolved in DMSO (0.4 mL) by heating at 110 °C for 2 min. The solution was added to the dried Cs[18F]F/kryptofix2.2.2, and the mixture was heated at 150, 175, or 200 °C for 20 min. The reaction mixture was cooled at room temperature in a water bath for 2 min, diluted with 30% methanol in dichloromethane (2.5 mL), and passed through a silica gel cartridge (Waters, 900 mg). The silica cartridge was further washed with 1.0 mL of methanol, and the solvent was evaporated under a stream of argon at 60–80 °C. The residue was redissolved in a solution (1.5 mL) containing 30% acetonitrile and 70% ammonium acetate (NH4OAc: 20 mM).
Purification was performed using a semipreparative radio-HPLC system with a 1260 series pump (Agilent Technologies) with a built-in UV detector operated at 250 nm and a radioactivity detector with a single-channel analyzer (labLogic). [18F]NT160 was eluted from a C18 reverse-phase column (00G-4041-N0: Luna 5 μm, 100 Å, 250 × 10 mm, Phenomenex) using 67% acetonitrile:33% ammonium acetate buffer (NH4OAc: 20 mM) at flow rate of 4.0 mL/min. [18F]NT160 was eluted and collected at 19–21 min postinjection. Water (15.0 mL) was added to the solution, and the solution was trapped on a C-18 light cartridge, eluted with ethanol (0.3 mL), and formulated for in vivo and in vitro studies.
Radiosynthesis of [18F]1
The details of the radiosynthesis of [18F]1 are provided in the Supporting Information.
HDAC Inhibitor Biochemical Assay
The details of the cell-based assay are provided in the Supporting Information. The cell-based HDAC inhibitor assays were performed in HT-29 cells (ATCC). Briefly, cells were grown in DMEM media (Gibco) supplemented with 10% FBS (Thermo Fisher Scientific) and 1X Anti-Anti (Gibco) in a humidified incubator at 37 °C with 5% CO2. The day before the experiment, cells were washed with PBS and dissociated using 0.25% trypsin–EDTA (Gibco). Trypsinization was stopped with media containing serum, and cells were collected by centrifugation. The cells were then washed with HDAC Assay Buffer (RPMI 1640 media without phenol red (Gibco) containing 0.1% FBS), recollected by centrifugation, resuspended in HDAC Assay Buffer, and counted. Two hundred thousand cells were seeded into 96-well plates and placed into the incubator overnight. HT-29 cells (obtained from ATCC) with the addition of either 100 μM Boc-Lys-TFA (class-IIa selective substrate) or 200 μM Boc-Lys-Ac (class-I/IIb selective substrate) were plated into 96-well plates at 200 000 cells/well in 45 μL of cellular assay buffer (RPMI without phenol red, 0.1% fetal bovine serum) and were incubated for 3 h at 37 °C. The deacetylation achieved by the addition of 50 μL of HDAC developer solution (2.5 mg/mL trypsin in DMEM without fetal bovine serum and 10% tween 80), followed by incubation for 1 h to sensitize the substrate and to lyse the cells. Fluorescent counts were read with a microplate reader at an excitation wavelength of 360 nm and detection of emitted light of 460 nm.
PET Imaging Procedures in Animals
All studies were performed under a protocol approved by the Institutional Animal Care and Use Committee of Stony Brook University. In vivo microPET/CT imaging studies were performed in healthy rats as described below. Anesthetized Sprague–Dawley (SD) female rats (200–450 g, N = 6) were placed in the Inveon uPET (Siemens, Knoxville, TN) in the supine position with the skull positioned in the center of the field of view. [18F]NT160 (18.5–25.9 MBq/animal) was administered via the tail-vein injection in a total volume of 0.5–1.2 mL. Dynamic PET images were obtained over 60 min followed by CT imaging (15 min scan). Images were reconstructed with attenuation correction using an ordered subset expectation maximization (OSEM2D) algorithm with 16 subsets and 4 iterations. PET image analysis was performed using the AMIDE software by using manual segmentation of regions of interest (ROI), including whole brain, cortex, hippocampus, thalamus, cerebellum, and brainstem. Rat Brain Atlas was used for alignment and identification of specific anatomical markers of the brain.35,36 The time–activity curve (TAC) was generated from the region of interest (ROI) by plotting the radioactivity/cc vs time. Levels of accumulation of the radiotracer in tissues were expressed as standard uptake values (SUV) that were calculated for the regions of interest (ROI) using the AMIDE software. The SUV is defined as the ratio of the tissue radioactivity concentration C (e.g., expressed as Bq/g tissue) at given time point postinjection T and the injected dose (e.g., in Bq, decay-corrected to the same time T) and normalized by the body weight in grams. GraphPad Prism (Graph Pad Software, La Jolla, CA) was used for image data analysis.
In Vitro Autoradiography
SD female rats (200–450 g, N = 3) were sacrificed, and the brains were fixed in 4% PFA paraffin overnight and then embedded into the paraffin block. Brains were sectioned into slices (10 μm). Then the slices were deparaffinized in xylene (three times) for 30 min (three times) followed by rehydration with 100% ethanol, 95% ethanol, 70% ethanol, and water for 30 min followed by rinsing in RO water for at least 30 min. The brain slides used for blocking studies were incubated with NT-160 in 20 mL of a 1 μM solution of NT-160 for 1 h, preincubated, and washed with binding buffer (50 mM Tris base, 2 mM MgCl2, pH 7.4) with 0.1% BSA for 10 min. 18F-NT160 (110.0 μCi) was added in 20 mL binding buffer and incubated at room temperature for 120 min. Subsequently, the slides were washed twice for 10 min in ice-cold binding buffer and dipped in ice-cold water before being dried. The slides were placed onto the storage phosphor screen in the dark overnight
Histology
SD female rats (200–450 g, N = 3) brains were fixed in 4% PFA paraffin overnight and then embedded into the paraffin block. Brains were sectioned into slices (10 μm). The slices were deparaffinized with xylene for 30 min three times and rehydrated with 10% ethanol, 90% ethanol, 70% ethanol, and water consecutively. Antigen retrieval was performed with citrate buffer (pH = 6) overnight at 60 °C followed by washing with PBS (three times). The slices were permeabilized with 0.1% Triton X-100 in 2.5% goat serum and stained with primer HDAC4 antibody (ABCAM, ab123513) overnight at 4 °C followed by washing with PBS (three times). The slices were incubated with biotinylated secondary antibody for 1 h and then were washed with PBS (three times). The slices were incubated with ABC agent for 30 min. The color was developed with DAB for 2 min followed by washing with water (three times). The staining was counted with hematoxylin for 1 min.
Radiometabolite Studies with [18F]NT160
The rat brains (n = 3) from the above imaging studies were excised and hemisected, and the half brain was homogenized in 1.0 mL of acetonitrile. The resulting suspension was centrifuged, the clear supernatant was vortexed and injected into an analytical radio-HPLC, and the extent of metabolism in the brain was determined. The radioactive peaks were compared to the retention time of the parent [18F]NT160. Also, samples were collected at a fraction of 1.0 min and counted using a gamma counter. The %radioactivity of the new peaks compared to the %radioactivity of the peak of the parent tracer was used to determine the extent of radiolabeled metabolites in the brain.
Blood was collected, and plasma was obtained by centrifugation. Protein-free plasma was obtained by addition of acetonitrile followed by centrifugation. The acetonitrile fraction was injected into analytical radio-HPLC (67% acetonitrile:ammonium acetate buffer (20 mM) solution). The radioactive peaks were compared to the retention time of the parent [18F]NT160 (∼6.9 min).28 Also, samples were collected at a fraction of 1.0 min and counted using a gamma counter. The %radioactivity of the new peaks compared to the %radioactivity of the peak of the parent tracer was used to determine the extent of radiolabeled metabolites in the blood.
Supporting Information Available
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.jmedchem.2c02064.
This work was funded by the National Institutes of Health/National Institute on Aging (grant no. R01AG067417).
The authors declare no competing financial interest.
Special Issue
Published as part of the Journal of Medicinal Chemistry virtual special issue “Diagnostic and Therapeutic Radiopharmaceuticals”.
Supplementary Material
References
- Kassis H.; Shehadah A.; Li C.; Zhang Y.; Cui Y.; Roberts C.; Sadry N.; Liu X.; Chopp M.; Zhang Z. G. Class IIa histone deacetylases affect neuronal remodeling and functional outcome after stroke. Neurochemistry international 2016, 96, 24–31. 10.1016/j.neuint.2016.04.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Faraco G.; Pancani T.; Formentini L.; Mascagni P.; Fossati G.; Leoni F.; Moroni F.; Chiarugi A. Pharmacological inhibition of histone deacetylases by suberoylanilide hydroxamic acid specifically alters gene expression and reduces ischemic injury in the mouse brain. Molecular pharmacology 2006, 70 (6), 1876–1884. 10.1124/mol.106.027912. [DOI] [PubMed] [Google Scholar]
- Ren M.; Leng Y.; Jeong M.; Leeds P. R.; Chuang D. M. Valproic acid reduces brain damage induced by transient focal cerebral ischemia in rats: potential roles of histone deacetylase inhibition and heat shock protein induction. Journal of neurochemistry 2004, 89 (6), 1358–1367. 10.1111/j.1471-4159.2004.02406.x. [DOI] [PubMed] [Google Scholar]
- Kim H. J.; Rowe M.; Ren M.; Hong J. S.; Chen P. S.; Chuang D. M. Histone deacetylase inhibitors exhibit anti-inflammatory and neuroprotective effects in a rat permanent ischemic model of stroke: multiple mechanisms of action. J. Pharmacol Exp Ther 2007, 321 (3), 892–901. 10.1124/jpet.107.120188. [DOI] [PubMed] [Google Scholar]
- Steffan J. S.; Bodai L.; Pallos J.; Poelman M.; McCampbell A.; Apostol B. L.; Kazantsev A.; Schmidt E.; Zhu Y. Z.; Greenwald M.; et al. Histone deacetylase inhibitors arrest polyglutamine-dependent neurodegeneration in Drosophila. Nature 2001, 413 (6857), 739–743. 10.1038/35099568. [DOI] [PubMed] [Google Scholar]
- Bürli R. W.; Luckhurst C. A.; Aziz O.; Matthews K. L.; Yates D.; Lyons K. A.; Beconi M.; McAllister G.; Breccia P.; Stott A. J.; et al. Design, synthesis, and biological evaluation of potent and selective class IIa histone deacetylase (HDAC) inhibitors as a potential therapy for Huntington’s disease. J. Med. Chem. 2013, 56 (24), 9934–9954. 10.1021/jm4011884. [DOI] [PubMed] [Google Scholar]
- Mielcarek M.; Benn C. L.; Franklin S. A.; Smith D. L.; Woodman B.; Marks P. A.; Bates G. P. SAHA decreases HDAC 2 and 4 levels in vivo and improves molecular phenotypes in the R6/2 mouse model of Huntington’s disease. PloS one 2011, 6 (11), e27746. 10.1371/journal.pone.0027746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shen X.; Chen J.; Li J.; Kofler J.; Herrup K. Neurons in Vulnerable Regions of the Alzheimer’s Disease Brain Display Reduced ATM Signaling. eNeuro 2016, 3 (1), ENEURO.0124-15.2016. 10.1523/ENEURO.0124-15.2016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Neuner S. M.; Wilmott L. A.; Hoffmann B. R.; Mozhui K.; Kaczorowski C. C. Hippocampal proteomics defines pathways associated with memory decline and resilience in normal aging and Alzheimer’s disease mouse models. Behavioural brain research 2017, 322 (Pt B), 288–298. 10.1016/j.bbr.2016.06.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sen A.; Nelson T. J.; Alkon D. L. ApoE4 and Aβ Oligomers Reduce BDNF Expression via HDAC Nuclear Translocation. Journal of neuroscience: the official journal of the Society for Neuroscience 2015, 35 (19), 7538–7551. 10.1523/JNEUROSCI.0260-15.2015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guerriero J. L.; Sotayo A.; Ponichtera H. E.; Castrillon J. A.; Pourzia A. L.; Schad S.; Johnson S. F.; Carrasco R. D.; Lazo S.; Bronson R. T.; et al. Class IIa HDAC inhibition reduces breast tumours and metastases through anti-tumour macrophages. Nature 2017, 543 (7645), 428–432. 10.1038/nature21409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Usami M.; Kikuchi S.; Takada K.; Ono M.; Sugama Y.; Arihara Y.; Hayasaka N.; Nakamura H.; Ikeda Y.; Hirakawa M.; et al. FOXO3a Activation by HDAC Class IIa Inhibition Induces Cell Cycle Arrest in Pancreatic Cancer Cells. Pancreas 2020, 49 (1), 135–142. 10.1097/MPA.0000000000001462. [DOI] [PubMed] [Google Scholar]
- Cai J. Y.; Xu T. T.; Wang Y.; Chang J. J.; Li J.; Chen X. Y.; Chen X.; Yin Y. F.; Ni X. J. Histone deacetylase HDAC4 promotes the proliferation and invasion of glioma cells. Int. J. Oncol. 2018, 53 (6), 2758–2768. 10.3892/ijo.2018.4564. [DOI] [PubMed] [Google Scholar]
- Hendricks J. A.; Keliher E. J.; Marinelli B.; Reiner T.; Weissleder R.; Mazitschek R. In vivo PET imaging of histone deacetylases by 18F-suberoylanilide hydroxamic acid (18F-SAHA). J. Med. Chem. 2011, 54 (15), 5576–5582. 10.1021/jm200620f. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zeglis B. M.; Pillarsetty N.; Divilov V.; Blasberg R. A.; Lewis J. S. The synthesis and evaluation of N1-(4-(2-[18F]-fluoroethyl)phenyl)-N8-hydroxyoctanediamide ([18F]-FESAHA), a PET radiotracer designed for the delineation of histone deacetylase expression in cancer. Nuclear medicine and biology 2011, 38 (5), 683–696. 10.1016/j.nucmedbio.2010.12.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hooker J. M.; Kim S. W.; Alexoff D.; Xu Y.; Shea C.; Reid A.; Volkow N.; Fowler J. S. Histone deacetylase inhibitor, MS-275, exhibits poor brain penetration: PK studies of [C]MS-275 using Positron Emission Tomography. ACS chemical neuroscience 2010, 1 (1), 65–73. 10.1021/cn9000268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seo Y. J.; Muench L.; Reid A.; Chen J.; Kang Y.; Hooker J. M.; Volkow N. D.; Fowler J. S.; Kim S. W. Radionuclide labeling and evaluation of candidate radioligands for PET imaging of histone deacetylase in the brain. Bioorganic & medicinal chemistry letters 2013, 23 (24), 6700–6705. 10.1016/j.bmcl.2013.10.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang C.; Schroeder F. A.; Wey H. Y.; Borra R.; Wagner F. F.; Reis S.; Kim S. W.; Holson E. B.; Haggarty S. J.; Hooker J. M. In vivo imaging of histone deacetylases (HDACs) in the central nervous system and major peripheral organs. J. Med. Chem. 2014, 57 (19), 7999–8009. 10.1021/jm500872p. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wey H. Y.; Wang C.; Schroeder F. A.; Logan J.; Price J. C.; Hooker J. M. Kinetic Analysis and Quantification of [11C]Martinostat for in Vivo HDAC Imaging of the Brain. ACS Chem. Neurosci. 2015, 6 (5), 708–715. 10.1021/acschemneuro.5b00066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reid A. E.; Hooker J.; Shumay E.; Logan J.; Shea C.; Kim S. W.; Collins S.; Xu Y.; Volkow N.; Fowler J. S. Evaluation of 6-([(18)F]fluoroacetamido)-1-hexanoicanilide for PET imaging of histone deacetylase in the baboon brain. Nuclear medicine and biology 2009, 36 (3), 247–258. 10.1016/j.nucmedbio.2008.12.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strebl M. G.; Campbell A. J.; Zhao W. N.; Schroeder F. A.; Riley M. M.; Chindavong P. S.; Morin T. M.; Haggarty S. J.; Wagner F. F.; Ritter T.; et al. HDAC6 Brain Mapping with [(18)F]Bavarostat Enabled by a Ru-Mediated Deoxyfluorination. ACS central science 2017, 3 (9), 1006–1014. 10.1021/acscentsci.7b00274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Celen S.; Rokka J.; Gilbert T. M.; Koole M.; Vermeulen I.; Serdons K.; Schroeder F. A.; Wagner F. F.; Bleeser T.; Hightower B. G.; et al. Translation of HDAC6 PET Imaging Using [(18)F]EKZ-001-cGMP Production and Measurement of HDAC6 Target Occupancy in Nonhuman Primates. ACS chemical neuroscience 2020, 11 (7), 1093–1101. 10.1021/acschemneuro.0c00074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bonomi R.; Mukhopadhyay U.; Shavrin A.; Yeh H. H.; Majhi A.; Dewage S. W.; Najjar A.; Lu X.; Cisneros G. A.; Tong W. P.; et al. Novel Histone Deacetylase Class IIa Selective Substrate Radiotracers for PET Imaging of Epigenetic Regulation in the Brain. PloS one 2015, 10 (8), e0133512. 10.1371/journal.pone.0133512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lear J. L.; Ackermann R. F. Evaluation of radiolabeled acetate and fluoroacetate as potential tracers of cerebral oxidative metabolism. Metabolic brain disease 1990, 5 (1), 45–56. 10.1007/BF00996977. [DOI] [PubMed] [Google Scholar]
- Ponde D. E.; Dence C. S.; Oyama N.; Kim J.; Tai Y. C.; Laforest R.; Siegel B. A.; Welch M. J. 18F-fluoroacetate: a potential acetate analog for prostate tumor imaging--in vivo evaluation of 18F-fluoroacetate versus 11C-acetate. Journal of nuclear medicine 2007, 48 (3), 420–428. [PubMed] [Google Scholar]
- Bisagno V.; Cadet J. L. Histone Deacetylases and Immediate Early Genes: Key Players in Psychostimulant-Induced Neuronal Plasticity. Neurotoxicity research 2021, 39 (6), 2134–2140. 10.1007/s12640-021-00420-3. [DOI] [PubMed] [Google Scholar]
- Parra M.; Verdin E. Regulatory signal transduction pathways for class IIa histone deacetylases. Current opinion in pharmacology 2010, 10 (4), 454–460. 10.1016/j.coph.2010.04.004. [DOI] [PubMed] [Google Scholar]
- Turkman N.; Liu D.; Pirola I. Design, synthesis, biochemical evaluation, radiolabeling and in vivo imaging with high affinity class-IIa histone deacetylase inhibitor for molecular imaging and targeted therapy. Eur. J. Med. Chem. 2022, 228, 114011. 10.1016/j.ejmech.2021.114011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Turkman N.; Liu D.; Pirola I. Novel late-stage radiosynthesis of 5-[18F]-trifluoromethyl-1,2,4-oxadiazole (TFMO) containing molecules for PET imaging. Sci. Rep 2021, 11 (1), 10668. 10.1038/s41598-021-90069-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van de Bittner G. C.; Ricq E. L.; Hooker J. M. A philosophy for CNS radiotracer design. Acc. Chem. Res. 2014, 47 (10), 3127–3134. 10.1021/ar500233s. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ryu Y. H.; Liow J. S.; Zoghbi S.; Fujita M.; Collins J.; Tipre D.; Sangare J.; Hong J.; Pike V. W.; Innis R. B. Disulfiram inhibits defluorination of (18)F-FCWAY, reduces bone radioactivity, and enhances visualization of radioligand binding to serotonin 5-HT1A receptors in human brain. J. Nucl. Med. 2007, 48 (7), 1154–1161. 10.2967/jnumed.107.039933. [DOI] [PubMed] [Google Scholar]
- Hostetler E. D.; Walji A. M.; Zeng Z.; Miller P.; Bennacef I.; Salinas C.; Connolly B.; Gantert L.; Haley H.; Holahan M.; et al. Preclinical Characterization of 18F-MK-6240, a Promising PET Tracer for In Vivo Quantification of Human Neurofibrillary Tangles. J. Nucl. Med. 2016, 57 (10), 1599–1606. 10.2967/jnumed.115.171678. [DOI] [PubMed] [Google Scholar]
- Luo L.; Martin S. C.; Parkington J.; Cadena S. M.; Zhu J.; Ibebunjo C.; Summermatter S.; Londraville N.; Patora-Komisarska K.; Widler L.; et al. HDAC4 Controls Muscle Homeostasis through Deacetylation of Myosin Heavy Chain, PGC-1α, and Hsc70. Cell Rep 2019, 29 (3), 749–763.e712. 10.1016/j.celrep.2019.09.023. [DOI] [PubMed] [Google Scholar]
- McGee S. L.; Hargreaves M. Histone modifications and skeletal muscle metabolic gene expression. Clin. Exp. Pharmacol. Physiol. 2010, 37 (3), 392–396. 10.1111/j.1440-1681.2009.05311.x. [DOI] [PubMed] [Google Scholar]
- Papp E. A.; Leergaard T. B.; Calabrese E.; Johnson G. A.; Bjaalie J. G. Waxholm Space atlas of the Sprague Dawley rat brain. Neuroimage 2014, 97, 374–386. 10.1016/j.neuroimage.2014.04.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Papp E. A.; Leergaard T. B.; Calabrese E.; Johnson G. A.; Bjaalie J. G. Addendum to “Waxholm Space atlas of the Sprague Dawley rat brain” [NeuroImage 97 (2014) 374–386]. Neuroimage 2015, 105, 561–562. 10.1016/j.neuroimage.2014.10.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Broide R. S.; Redwine J. M.; Aftahi N.; Young W.; Bloom F. E.; Winrow C. J. Distribution of histone deacetylases 1–11 in the rat brain. J. Mol. Neurosci 2007, 31 (1), 47–58. 10.1007/BF02686117. [DOI] [PubMed] [Google Scholar]
- Darcy M. J.; Calvin K.; Cavnar K.; Ouimet C. C. Regional and subcellular distribution of HDAC4 in mouse brain. J. Comp Neurol 2010, 518 (5), 722–740. 10.1002/cne.22241. [DOI] [PubMed] [Google Scholar]
- Zhang Y.; Yang P.; Chou C. J.; Liu C.; Wang X.; Xu W. Development of N-Hydroxycinnamamide-Based Histone Deacetylase Inhibitors with Indole-Containing Cap Group. ACS Med. Chem. Lett. 2013, 4 (2), 235–238. 10.1021/ml300366t. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pike V. W. Considerations in the Development of Reversibly Binding PET Radioligands for Brain Imaging. Curr. Med. Chem. 2016, 23 (18), 1818–1869. 10.2174/0929867323666160418114826. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.