Abstract
Double-knotted peptides identified in venoms and synthetic bivalent peptide constructs targeting ion channels are emerging tools for the study of ion channel pharmacology and physiology. These highly complex and disulfide-rich peptides contain two individual cystine knots, each comprising six cysteines and three disulfide bonds. Until now, native double-knotted peptides, such as Hi1a and DkTx, have only been isolated from venom or produced recombinantly, whereas engineered double-knotted peptides have successfully been produced through enzymatic ligation using sortase A to form a seamless amide bond at the ligation site between two knotted toxins, and by alkyne/azide click chemistry, joining two peptide knots via a triazole linkage. To further pursue these double-knotted peptides as pharmacological tools or probes for therapeutically relevant ion channels, we sought to identify a robust methodology resulting in a high yield product that lends itself to rapid production and facile mutational studies. In this study, we evaluated the ligation efficiency of enzymatic (sortase A5°, butelase 1, wild-type OaAEP 1, C247A-OaAEP 1, and peptiligase) and mild chemical approaches (α-ketoacid-hydroxylamine, KAHA) for forming a native amide bond linking the toxins while maintaining the native disulfide connectivity of each pre-folded peptide. We used two NaV1.7 inhibitors: PaurTx3, a spider-derived gating modifier peptide, and KIIIA, a small cone snail-derived pore blocker peptide, which have previously been shown to increase affinity and inhibitory potency on hNaV1.7 when ligated together. Correctly folded peptides were successfully ligated in varying yields, without disulfide bond shuffling or reduction, with sortase A5° being the most efficient, resulting in 60% ligation conversion within 15 min. In addition, electrophysiology studies demonstrated that for these two peptides, the amino acid composition of the linker did not affect the activity of the double-knotted peptides. This study demonstrates the powerful application of enzymes in efficiently ligating complex disulfide-rich peptides, paving the way for facile production of double-knotted peptides.
Graphical Abstract

INTRODUCTION
Double-knotted peptides are increasingly being identified in spider venoms.1–3 Two double-knotted peptides, DkTx and Hi1a, were recently found to target the heat- and acid-sensitive tetrameric transient receptor potential ion channel TRPV1 and the acid-sensitive ion channel ASIC1a, respectively,1,2,4,5 whereas CpTx1 has been reported to damage membranes in a receptor-independent manner.3 DkTx and Hi1a comprise 79 and 75 amino acid residues, respectively, and contain two individual inhibitor cystine knot (ICK) domains, with three disulfide bonds each, linked together via a short amino acid chain.1,2 CpTx1, on the other hand, is a much larger 134 amino acid residue peptide containing two domains of four disulfide bonds each.3 In addition to these three native double-knotted venom peptides, identified using traditional assay-guided fractionation, a recent study using data-mining of the UniProtKB database identified more than 60,000 secreted cysteine-rich repeat peptides (SCREPs) for a range of putative targets including bovine pancreatic trypsin inhibitor/Kunitz domains, with 836 non-redundant SCREPs targeting ion channels.6 This highlights the prevalence of these multi-domain peptides in nature, their potential as pharmacological tools or drug leads for a range of therapeutic targets, and the importance of the development of robust synthetic pathways to further explore the pharmacology of this class of peptides.
For DkTx and Hi1a, although both domains are independently active to varying degrees, neither of the individual domains can reach the potency of their respective full-length peptide.1,2 This suggests that both domains must simultaneously bind to the receptors to promote full activity. Despite the similarity in the channel architecture between TRPV1 and NaVs,4,7–9 so far there are no reports of naturally occurring multi-domain peptides that target voltage-gated sodium channels, albeit ~10% of SCREPs are predicted to target NaVs.6 The first attempt to produce an artificial double-knotted peptide targeting NaVs was by Murray et al., who produced a homodimer of the gating modifier toxin (GMT) GpTx-1 by means of linking the two domains with a polyethylene glycol (PEG) linker using alkyne/azide click chemistry.10 Dimerization caused a 10-fold increase in potency and increased the peptide residence time on NaV1.7 compared to a single GpTx-1 molecule.10 In addition, a recent study from our group showed successful homo- and hetero-dimerization of GMTs CcoTx-1, CcoTx-2, gHwTx-IV, and D1la targeting NaV1.7 using the evolved sortase A5° (SrtA5°) and a four-residue glycine peptide linker.11 These peptides did not display improved affinity for NaV1.7, but the results suggested the presence of cooperative binding.11 These findings have emphasized the potential of homo- and hetero-bivalent neurotoxins targeting NaVs. In addition to studies focusing on homo- and hetero-ligation dimerization of two ICK GMTs targeting the voltage-sensing domains of NaV1.7, the pharmacological activity of hetero-bivalent peptides comprising the GMTs mHwTx-IV and PaurTx3 ligated to the NaV channel pore blocker μ-conotoxin KIIIA has also been explored.12,13
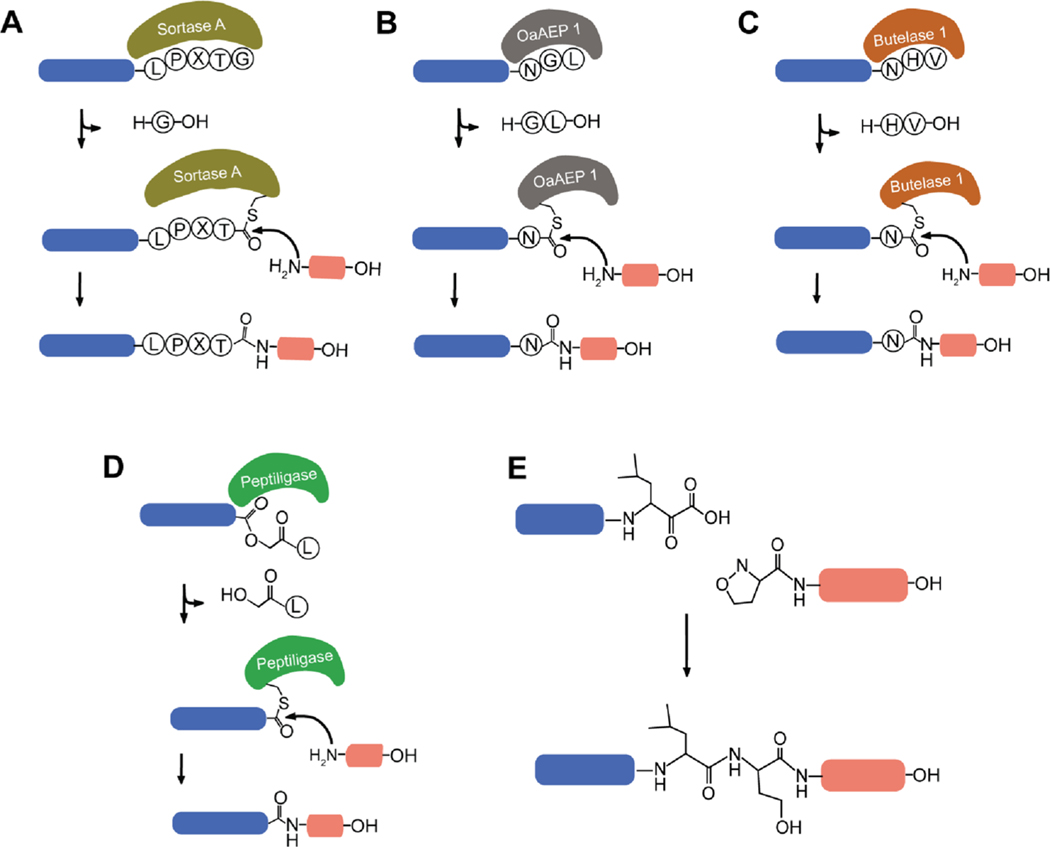
Although there are multiple ways to join two peptide fragments together using a variety of linkers and ligation strategies,10,11,14–19 our aim was to utilize a ligation method leaving a native amide bond to mimic bivalent peptides found in nature. The challenge in producing these bivalent peptides synthetically arises from our desire to fold each of the domains prior to ligation to ensure correct disulfide bond connectivity. This means that the chosen ligation method must be non-reducing to avoid disulfide bond shuffling during the ligation reaction. Popular chemical ligation strategies, including a variety of native chemical ligation (NCL) methods, which are elegant, rapid, and high yielding reactions, usually require reducing agents like tris(2-carboxyethyl)phosphine (TCEP) and thiol additives like 4-mercaptophenylacetic acid or sodium 2-mercaptoethanesulfonate20–23 and are thus not suitable for this study. On the other hand, enzymatic ligation utilizing enzymes including sortase A (SrtA) (Figure 1A),11,12,24–26 asparaginyl endopeptidases (OaAEP 1) (Figure 1B), butelase 1 (Figure 1C),27–34 and peptiligase (Figure 1D)35–39 or chemical ligation using α-ketoacid-hydroxylamine amide (KAHA) (Figure 1E)40–44 approaches has been shown to be able to form native amide bonds in the absence of reducing agents.
Figure 1.
Scheme showing enzymatic and chemical ligation approaches examined in this study. The recognition sequences of the enzymes are shown as single letter amino acid residue codes. The N-terminal peptide is shown in blue, and the C-terminal peptide is shown in orange. (A) SrtA, (B) OaAEP 1, (C) butelase 1, (D) peptiligase, and (E) KAHA ligation.
In this study, we sought to evaluate these ligation strategies using PaurTx3, a spider-derived GMT from Phrixotrichus auratus,45,46 and KIIIA, a μ-conotoxin NaV pore blocker isolated from the venom of Conus kinoshitai.47 These peptides were originally selected in a previous study due to their moderate and approximately equipotent activities at hNaV1.7, one of the best-validated pain targets, (PaurTx3 IC50 = 290 nM, KIIIA IC50 = 363 nM), allowing us to detect any potential improvements in activity upon ligation.12 Following evaluation of the two recently reported cryo-EM structures of KIIIA in complex with NaV1.248 and the structure of the gating modifier toxin ProTxII in complex with NaV1.7,49 measuring the distances between the two toxin binding sites, we applied an SrtA5°-mediated approach using polyglycine linkers to evaluate various linker lengths in order to optimize the NaV1.7 activity of ligated PaurTx3 to KIIIA.12 The synthetic bivalent peptide with an optimized 9-glycine linker length showed improved potency at NaV1.7 compared to either PaurTx3 or KIIIA individually with an IC50 = 58 nM.12 This work suggested that novel and improved NaV1.7 inhibitors can be designed by combining a pore blocker toxin and a gating modifier toxin to confer desired pharmacological properties from both the voltage sensing domain and the pore domain. However, SrtA5°-mediated ligation reactions resulted in hydrolysis products that proved to be challenging to separate from the starting material as they elute close to PaurTx3 by HPLC. In addition, the five-residue-long SrtA motif ultimately proved to be susceptible to degradation in human serum.12 Our aim with this study was therefore to evaluate alternative approaches to serve our purpose of creating bivalent disulfide-rich peptides via native amide bonds while maintaining their structure, potency, and stability under harsh proteolytic conditions encountered in vivo.
Herein, we present a comparative study evaluating the ability of both enzymatic (SrtA5°, OaAEP 1, butelase 1, and peptiligase) and chemical (KAHA) ligation approaches for creating bivalent peptides from disulfide-rich monomers of KIIIA and PaurTx3. Using NMR, we confirmed that the different ligation strategies did not compromise the overall fold in the individual domains, we evaluated the impact of different linker sequences on bivalent peptide activity using electrophysiology, and finally we evaluated the ligated peptide’s stability in serum.
RESULTS
In our previous study, we found that the linker length for connecting PaurTx3 and KIIIA to reach maximum potency at NaV1.7 was nine amino acid residues.12 We therefore maintained this linker length for all ligation strategies tested here in order to evaluate the efficiency of different ligation strategies in addition to assessing if different linker compositions affected activity, adding or subtracting Gly residues to accommodate the different ligation motives to achieve the desired linker length. Peptide sequences, enzyme recognition sequences, linkers, and theoretical and observed masses for peptides used in this study are summarized in Table 1 and Table S1. All results of SrtA5°-mediated ligation in this study were extracted from Tran et al.12 and are used for comparative purposes.
Table 1.
Summary of Peptide Sequences Synthesized for This Studya
| Strategy | Peptide | Sequence |
|---|---|---|
|
| ||
| PaurTx3 | DCLGFLWKCNPSNDKCCRPNLVCSRKDKWCKYQI | |
|
| ||
| KIIIA | CCNCSSKWCRDHSRCC* | |
|
| ||
| SrtA5° | PaurTx3[LPAT|GG] | DCLGFLWKCNPSNDKCCRPNLVCSRKDKWCKYQILPATGG |
|
| ||
| [G5]KIIIA | GGGGGCCNCSSKWCRDHSRCC * | |
|
| ||
| P[LPATG5]K | DCLGFLWKCNPSNDKCCRPNLVCSRKDKWCKYQILPATGGGGGCCNCSSKWCRDHSRCC * | |
|
| ||
| wt-OaAEP 1 | PaurTx3[N|GL] | DCLGFLWKCNPSNDKCCRPNLVCSRKDKWCKYQINGL |
| [G8]KIIIA | GGGGGGGGCCNCSSKWCRDHSRCC* | |
| Pr[NG8]K | DCLGFLWKCNPSNDKCCRPNLVCSRKDKWCKYQINGGGGGGGGCCNCSSKWCRDHSRCC* | |
|
| ||
| PaurTx3[N|GL] | DCLGFLWKCNPSNDKCCRPNLVCSRKDKWCKYQINGL | |
| [GVG6]KIIIA | GVGGGGGGCCNCSSKWCRDHSRCC* | |
| P[NGVG6]K | DCLGFLWKCNPSNDKCCRPNLVCSRKDKWCKYQINGVGGGGGGCCNCSSKWCRDHSRCC* | |
|
| ||
| C247A-OaAEP 1 | PaurTx3[N|GL] | DCLGFLWKCNPSNDKCCRPNLVCSRKDKWCKYQINGL |
| [GVG6]KIIIA |
GVGGGGGGCCNCSSKWCRDHSRCC* | |
| PrNGVG6lK | DCLGFLWKCNPSNDKCCRPNLVCSRKDKWCKYQINGVGGGGGGCCNCSSKWCRDHSRCC* | |
|
| ||
| Butelase 1 | PaurTx3[N|HV] | DCLGFLWKCNPSNDKCCRPNLVCSRKDKWCKYQINHV |
| [GVG6]KIIIA | GVGGGGGGCCNCSSKWCRDHSRCC* | |
| P[NGVG6]K | DCLGFLWKCNPSNDKCCRPNLVCSRKDKWCKYQINGVGGGGGGCCNCSSKWCRDHSRCC* | |
|
| ||
| KAHA | PaurTx3-ka# | DCLGFLWKCNPSNDKCCRPNLVCSRKDKWCKYQL-COOH |
| [OprG8]KIIIA | OprGGGGGGGGCCNCSSKWCRDHSRCC* | |
| P[hSG8]K | DCLGFLWKCNPSNDKCCRPNLVCSRKDKWCKYQLhSGGGGGGGGCCNCSSKWCRDHSRCC* | |
|
| ||
| Omniligase 1 | PaurTx3[AVRL|-OCamL] | DCLGFLWKCNPSNDKCCRPNLVCSRKDKWCKYQAVRL-OCamL |
| [FIG3]KIIIA | FIGGGCCNCSSKWCRDHSRCC* | |
|
| ||
| Thymoligase 1 | PaurTx3[ITTK|-OCamL] | DCLGFLWKCNPSNDKCCRPNLVCSRKDKWCKYQITTK-OCamL |
| [DLG3]KIIIA | DLGGGCCNCSSKWCRDHSRCC* | |
: enzyme cleavage site, red: C-terminal recognition motifs/mutations, blue: N-terminal polyglycine/recognition motif.
C: terminal amide
ka: Leu-a-ketocarboxylic acid, Opr: oxaproline
S: homoserine, and OCam: carboxyamidomethyl ester.
OaAEP 1 Ligation.
All peptide precursors used in this study were synthesized using fluorenylmethyloxycarbonyl (Fmoc) chemistry and cleaved individually. KIIIA was assembled with an eight-residue polyglycine at the N-terminus ([G8]KIIIA), while PaurTx3 was assembled with the OaAEP 1 recognition site, NGL, at its C-terminus (PaurTx3[NGL]) (Table 1). Under thermodynamic oxidizing conditions, PaurTx3[NGL] formed only one major product during oxidation (28% yield) (Figure S1A), while [G8]KIIIA formed two isomers as reported previously (Figure S2A).12 While μ-conotoxins typically display a disulfide connectivity of C1–C9, C2–C15, and C4–C16,50 full 3D structure characterization and direct mass spectrometric collision-induced dissociation fragmentation of KIIIA carried out by Khoo et al. confirmed that the major isomer observed following thermodynamic oxidation of KIIIA display a C1–C15, C2–C9, and C4–C16 disulfide bond connectivity.51 As reported previously,12 comparison of the 1D 1H NMR amide region of [G8]KIIIA with published KIIIA (PDB ID: 2lxg)51 (Figure S3A) and secondary Hα chemical shifts of both peptides suggests that the major isomer following folding of [G8]KIIIA exhibits a similar overall structure compared to the major KIIIA isomer with C1–C15, C2–C9, and C4–C16 connectivity reported by Khoo et al. (Figure S3B).51 The major oxidation isomer of [G8]KIIIA (yield: 38%) was subsequently used for ligation. Ligation conditions and reaction lengths were optimized for all strategies (Figure S4) and are reported in Table 2. Folded PaurTx3[NGL] and [G8]KIIIA were ligated in phosphate buffer using OaAEP1 (Table 2), with reaction progress monitored at 0, 2, 4, 6, 8, and 24 h (Figure S4A). The desired product was identified using ESI-MS and purified by reversed-phase high-performance liquid chromatography (RP-HPLC) at final yield of 12% (Figures 2A and 3A). Since it has been reported that a GV recognition site in the N-terminal region of the C-terminal fragment improves OaAEP 1 catalysis and that ligation with a GV recognition site is irreversible,52 we also synthesized [GVG6]KIIIA. Under thermodynamic oxidizing conditions, [GVG6]KIIIA, like [G8]KIIIA, also formed two isomers. The predominant [GVG6]KIIIA isomer (40% yield) (Figure S2B), evaluated by 1D and 2D 1H NMR (as described above) to confirm correct folding (Figure S3), was subsequently used for ligation. The ligation was conducted using both the wild-type and the C247A-OaAEP 1 enzyme. When treated with the wild-type enzyme, [GVG6]KIIIA (yield 16%) (Figure 2B) showed little improvement in ligation yield compared to [G8]KIIIA (yield: 12%) (Figure 2A). However, as expected, the reaction in which [GVG6]KIIIA was incubated with the C247A-OaAEP 1 mutant produced a higher yield (35% yield) (Figure 2C and Figure S4B). The 1D 1H NMR spectra of the ligated peptides superimposed well with the individual peptide domain spectra, thus suggesting that the disulfide bonds were intact during ligation (Figure S5A). Furthermore, no disulfide bond shuffling was observed during ligation, as evidenced by a single ligation peak in the HPLC chromatograms (Figure 2 and Figure S4). In addition, Hα secondary shift analysis derived from 2D NMR TOCSY and NOESY spectra of the bivalent peptides overlapped well with previously published PaurTx3 (PDB ID: 5we3)53 and KIIIA (PDB ID: 2lxg),51 confirming that the enzymatic ligation does not shuffle already formed disulfide bonds (Figure S5A,B and Figure 3D).
Table 2.
Summary of Ligation Conditions and Yields of each Oxidation and Ligation Approach
| strategy | KIIIA precursor oxidation yielda | PaurTx3 precursor oxidation yielda | conditionc | reaction time | ligation yieldb |
|---|---|---|---|---|---|
| SrtA5° | 40% | 50% | 100 μM PaurTx3[LPATGG]:300 μM [G8]KIIIA:30 μM enzyme 50 mM Tris:150 mM NaCl:10 mM CaCl2, pH 7.4, 37 °C |
15 min | 60% |
| wt-OaAEP 1 | 38% | 28% | 250 μM PaurTx3[NGL]:750 μM [G8]KIIIA:3 μM enzyme 0.1 M Na2HPO4:0.1 M NaCl, pH 6.5 |
4 h | 12% |
| C247A-OaAEP 1 | 40% | 18% | 20 μM PaurTx3[NGL]:60 μM [GVG6]KIIIA:75 nM enzyme 0.1 M Na2HPO4:0.1 M NaCl, pH 6.5 |
6 h | 35% |
| butelase 1 | 40% | 29% | 20 μM PaurTx3[NHV]:60 μM [GVG6]KIIIA:100 nM enzyme 0.1 M Na2HPO4:0.1 M NaCl, pH 6.5 |
6 h | 38% |
| KAHA | 25% | 28% | 22 mM PaurTx3-ka:15 mM [OprG8]-KIIIA | 6 h | 36% |
| peptiligased | 40% | n/d | 90% DMSO:10% H2O, 0.1 M oxalic acid, 60 °C 10 mM PaurTx3[linker]-OCamL:15 mM [linker]-KIIIA 10% DMSO in 1 M potassium phosphate, pH 8–8.5 (with/without 2 |
n/a | n/d |
Oxidation yield was calculated based on the precursor weight and isolated oxidized peptide weight.
Ligation yield was calculated by dividing the observed weight by the theoretical weight calculated from PaurTx3 analogs.
Reactions carried out at room temperature unless specified.
Ligation was not carried due to insolubility of peptide fragments.
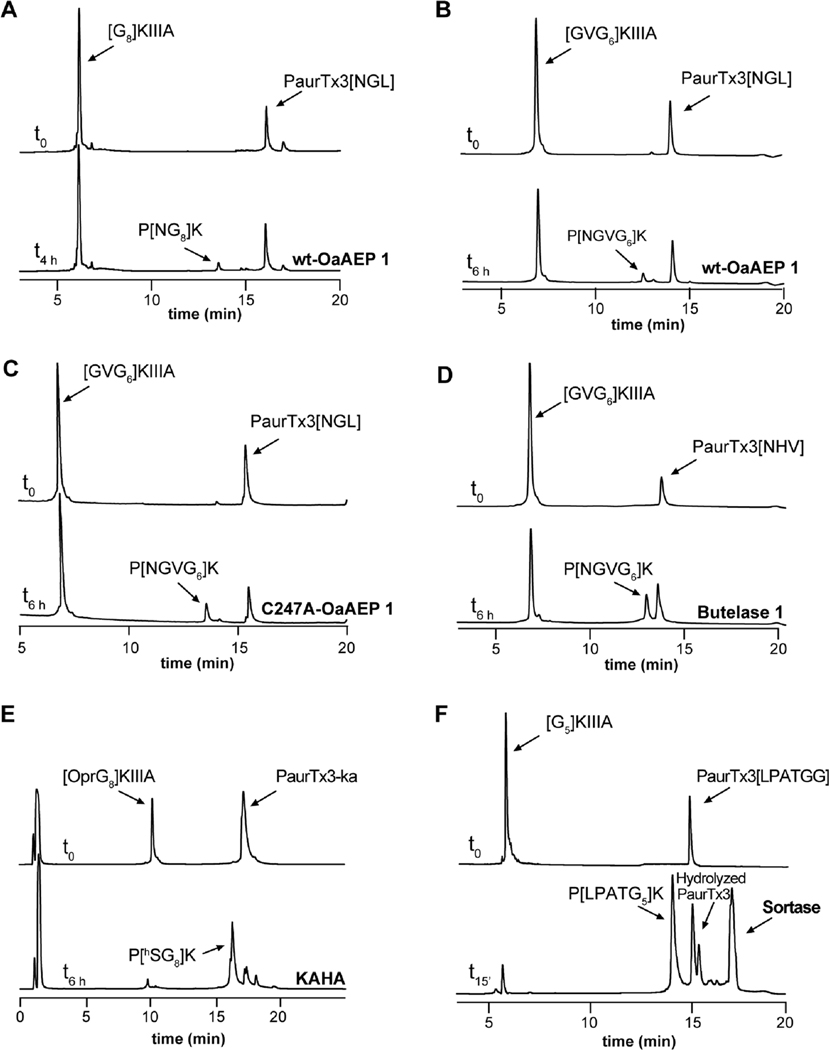
Figure 2.
Enzymatic and chemical ligation approaches of PaurTx3 and KIIIA. RP-HPLC chromatograms of PaurTx3 and KIIIA starting material (top) and ligated products (bottom) following completion of the ligation reaction. (A) Ligation of PaurTx3[NGL] and [G8]KIIIA using wt-OaAEP 1. (B) Ligation of PaurTx3[NGL] and [GVG6]KIIIA using wt-OaAEP 1. (C) Ligation of PaurTx3[NGL] and [GVG6]KIIIA using C247A-OaAEP 1. (D) Ligation of PaurTx3[NHV] and [GVG6]KIIIA using butelase 1. (E) KAHA ligation between PaurTx3-ka and [OprG8]KIIIA. (F) Ligation of PaurTx3[LPATGG] and [G5]KIIIA using SrtA5° (SrtA5° data adapted from Tran et al. and included for comparison purposes12).
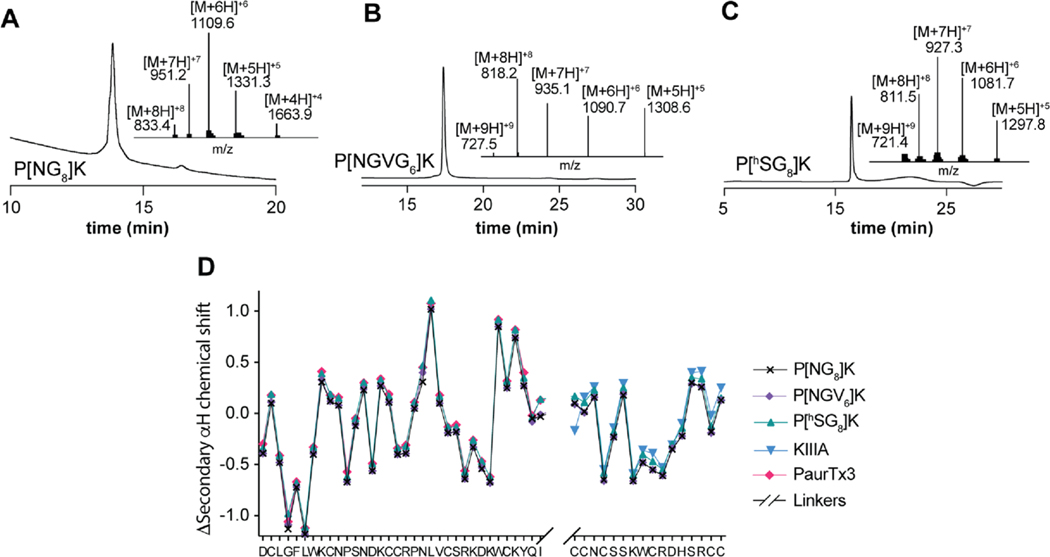
Figure 3.
Enzymatic and chemical ligation products following PaurTx3 and KIIIA ligation. Analytical RP-HPLC traces and observed masses (ESI-MS) of purified bivalent peptides. (A) Product from wt-OaAEP 1 ligation using PaurTx3[NGL] and [G8]KIIIA. (B) Product from wt-OaAEP 1 (identical product from C247A-OaAEP 1, not shown) and butelase 1 ligation using PaurTx3[NGL] and PaurTx3[NHV], respectively, and [GVG6]KIIIA. (C) Product from KAHA ligation using PaurTx3-ka and [OprG8]KIIIA. (D) Secondary Hα chemical shifts for P[NG8]K, P[NGVG6]K, and P[hSG8]K, overlaid with published KIIIA (from Khoo et al.),51 and native PaurTx3 (from Agwa et al.).53 The linker region was omitted for clarity.
Butelase 1 Ligation.
Butelase 1 is an asparaginyl endopeptidase like OaAEP 1 with a similar ligation mechanism and recognition motif, and therefore, [GVG6]KIIIA could again be used as the C-terminal fragment for butelase 1 ligation.31,33,34 PaurTx3 was assembled with the NHV butelase 1 recognition site at its C-terminus (PaurTx3[NHV]) (Table 1). Folding of PaurTx3[NHV] formed one primary oxidation product (29% yield) (Figures S1B and S3). The two fragments were dissolved in butelase 1 ligation buffer (Table 2), and ligation reaction was monitored by LC/MS at room temperature for 6 h (Figures 2D and 3B and Figure S4C). The desired product was formed at 38% yield and was examined by 1D and 2D NMR (Figure 3D and Figure S5C).
Peptiligase.
The substrate-tailored variants of peptiligase, omniligase 1, and thymoligase 1 were used for ligation of bivalent peptides. KIIIA was synthesized with three Gly residues followed by either FI or DL at the N-terminus ([FIG3]KIIIA or [DLG3]KIIIA) for omniligase and thymoligase, respectively. PaurTx3 was assembled with glycolic acid to yield the ester linkers ARVL-OCamL or ITTK-OCamL at its C-terminus (PaurTx3[ARVL-OCamL] or PaurTx3[ITTK-OCamL]) for omniligase and thymoligase, respectively (Table 1). Peptides were cleaved from the resin, isolated by HPLC, and oxidized individually. Under thermodynamic oxidizing conditions previously described, the KIIIA analogs [FIG3]KIIIA or [DLG3]KIIIA formed one major isomer each (Figure S2C,D). Unfortunately, while PaurTx3[ARVL-OCamL] and PaurTx3[ITTK-OCamL] displayed one main HPLC peak following thermodynamic oxidation using two different folding conditions (Figure S1C,D), these peptides appeared misfolded when examined using 1D 1H NMR (Figure S3). Furthermore, both the linear and the misfolded PaurTx3-OCamL esters (5–10 mM) precipitated in ligation buffer (0.1–1 M phosphate buffer pH 8.3, with or without 2 M urea, 1 M Gn·HCl, or DMSO). Thus, ligation of PaurTx3 and KIIIA using omniligase 1 or thymoligase 1 could not be attempted. Peptiligase ligation reaction require an optimal pH between 8.0 and 8.5 as at lower pH (<7.5), a significant amount of the N-terminal amine is protonated and thus unreactive.54 It is likely that both linear and oxidized PaurTx3 analogs precipitated in potassium phosphate ligation buffer as the theoretical pI of these PaurTx3 analogs is 8.8 and thus incompatible with these conditions.
KAHA Ligation.
The C-terminal KIIIA-fragment used for KAHA-ligation was synthesized with eight Gly residues followed by an oxaproline (Opr) at the N-terminus ([OprG8]-KIIIA), while the N-terminal PaurTx3-fragment was assembled with a Leu-α-ketocarboxylic acid (ka) at the C-terminal in place of Ile34 (PaurTx3-ka) (Table 1). Peptides were cleaved from the resin and oxidized individually. Under thermodynamic oxidizing conditions, both PaurTx3-ka and [OprG8]-KIIIA formed one primary product, with yields of 28 and 25%, respectively (Figures S1E, S2E, and S3). Folded peptides were ligated for 6 h in 90% DMSO/10% H2O and 0.1 M oxalic acid (Table 2), and the desired product was purified with a yield of 36% (Figures 2E and 3C and Figure S4D). Following ligation, treatment with NH4HCO3 resulted in a rightward shift in RP-HPLC retention time, indicating that O → N shift (Figure S4E) was completed and P[hSG8]K, containing a homoserine residue in its linker, was formed via a native amide bond (Figure 3C). The ligated peptide P[hSG8]K was examined by 1D and 2D NMR and compared to native PaurTx3 and KIIIA to ensure that the integrity of the disulfide bonds was intact and the overall fold was maintained (Figure S5D and Figure 3D).
In addition to generating the individual oxidized domains of PaurTx3-ka and [OprG8]-KIIIA and ligating them, we also produced the linear peptide P[hSG8]K using KAHA ligation prior to oxidation of the two individual domains (Figure S6A). Following successful ligation and purification of the linear peptide, we attempted to fold the full-length linear peptide. However, the desired folding of the peptide was not achieved using our standard PaurTx3 oxidation condition (0.1 M Tris/0.81 mM GSH/0.81 mM GSSG) nor the KIIIA oxidation condition (0.1 M NH4HCO3/100 equiv GSH/10 equiv GSSG, pH 8) (Figure S6B), highlighting the importance of our modular folding strategy.
Automated Patch-Clamp Electrophysiology.
To assess the impact of linker types on the activity of the bivalent peptides, all successfully ligated bivalent peptide constructs were tested for activity at human NaV1.7 (hNaV1.7) expressed in HEK293 cells using automated whole cell patch-clamp electrophysiology (Figure 4A–C and Figure S7).
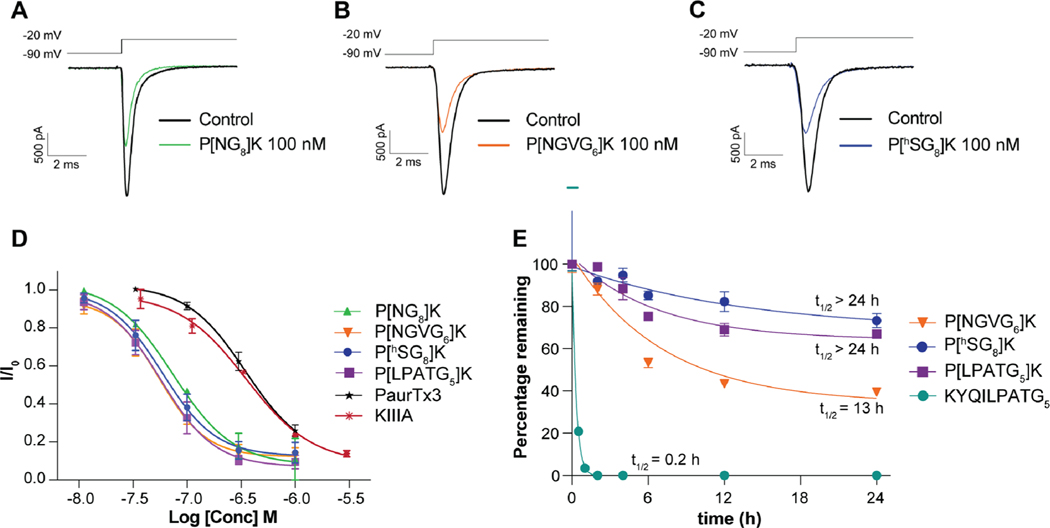
Figure 4.
Pharmacology at hNaV1.7 and stability of bivalent peptides. (A) Representative current trace before and after the addition of 100 nM P[NG8]K. (B) Representative current trace before and after the addition of 100 nM P[NG8]K. (C) Representative current trace before and after the addition of 100 nM P[hSG8]K. (D) Concentration–response curves of bivalent peptides on hNaV1.7 expressed in HEK293 cells assessed using automated whole cell patch-clamp electrophysiology (n = 3). (E) Serum stability of P[NGVG6]K (ligated with wt-OaAEP 1, C247A-OaAEP 1, or butelase 1), P[hSG8]K (KAHA), and P[LPATG5]K (ligated with SrtA5°) in human serum over 24 h. The linear linker sequence KYQILPATG5 (SrtA5° linker sequence) was used as the positive control. P[LPATG5]K and linear linker sequence data are adapted from Tran et al.12 for comparison purposes.
All bivalent peptides produced in this study performed better than PaurTx3 and KIIIA (Figure 4D) individually, with minor differences in the linker sequences resulting in negligible effects on activity at NaV1.7 (P[NG8]K IC50 = 74 ± 0.9 nM, P[NGVG6]K IC50 = 55 ± 0.8 nM, P[hSG8]K IC50 = 59 ± 0.8 nM, P[LPATG5]K IC50 = 57 ± 0.9 nM) (Figure 4D).
Serum Stability.
Proteolytic stability of P[NGVG6]K, P[hSG8]K, and P[LPATG5]K was determined by incubation in human serum and compared to the linear linker used in SrtA5° ligation (Figure 4E).12 Interestingly, incorporation of different linkers, arising from the different enzymes and ligation strategies employed in this study, gave rise to differences in serum stability. After 24 h of incubation, P[hSG8]K, chemically ligated using KAHA-ligation, proved to be the most proteolytically stable analog with 73.3 ± 3.3% of the peptide remaining, followed by P[LPATG5]K, ligated by SrtA5° with 66.9 ± 3.3% remaining,12 and P[NGVG6]K, ligated with either wt-OaAEP 1, C247A-OaAEP 1, or butelase 1, with 39.4 ± 1.0% peptide remaining (Figure 4E and Figure S8).
DISCUSSION
Peptide toxins that target NaVs are hailed as promising candidates for future analgesic treatments. However, despite extensive discovery and development efforts focused on potent and selective venom-derived gating-modifier toxins targeting hNaV1.7, this potential is yet to be realized.55–60 Recently discovered double-knotted spider toxins1–3 and enabling synthetic methods10,11,14–19 have paved a path for the development of an exciting new class of pharmacological tools and potential drug leads, involving conjugation of two potent NaV inhibitors with different modes of action to create synthetic bivalent constructs. However, the route to synthetically accessing this new class of bivalent peptides is not straightforward due to complications introduced by the presence of complex disulfide bond frameworks in each individual peptide domain, prohibiting the use of most standard chemical ligation techniques. We therefore evaluated a series of enzymatic and chemical strategies to delineate the most facile and high yielding method of producing these peptides.
By employing automated Fmoc solid-phase peptide synthesis protocols, individual peptide precursors, including the enzyme-specific recognition sequences incorporated at the N- or C-termini of the peptides designed for the different ligations evaluated in this study, were successfully assembled in high yield and purity. All peptide precursors, apart from PaurTx3- [ARVL-OCamL] and PaurTx3[ITTK-OCamL], were successfully oxidized, and structures of the individual peptides incorporating the different ligation recognition motifs were evaluated by 1D and 2D NMR and shown to correlate well with their native counterparts, confirming that the correct overall secondary structure of the peptides was maintained. Although none of the ligation approaches evaluated achieved complete conversion, four approaches, including SrtA5°, OaAEPs, butelase 1, and KAHA-ligation, proved their ability to ligate complex disulfide-rich peptides under mild conditions without any detectable disulfide bond reduction or shuffling.
OaAEP 1 ligation resulted in a clean reaction with the desired product and unreacted starting materials being easy to recover. Due to the removal of the hydrophobic Gly-Leu recognition motif in the C-terminal of P[GVG6]K following the ligation, the ligated product was easily separated from the more hydrophobic starting material PaurTx3[NGL] by RP-HPLC. Additionally, the small amount of added enzyme resulted in easy separation of the desired product during purification. Despite this, wt-OaAEP 1 reaction was the least efficient approach evaluated in this study. After several attempts to ligate PaurTx3[NGL] and [G8]KIIIA, a maximum yield of only 12% was achieved after 4 h. OaAEP 1 has been reported to be an effective catalyst when cyclizing cyclotide kB1, but it was also observed that the kinetic cyclization rate of OaAEP 1 on other peptides was inferior to that of kB1.27,61 The interaction between the peptide and the binding pocket of OaAEP 1 has been shown to play a vital role in catalytic efficiency, and peptides with an N-terminus containing GL or GV present better kinetics than other residues, including VF or VK.52,62,63 The C-terminal amino acids following N in the recognition motif have also been shown to be critical, and NGL, NGI, and NGV have been shown to produce more efficient cleavage sites.52,62,63 Thus, OaAEP 1 ligation was also attempted with PaurTx3[NGL] and [GVG6]KIIIA. This combination allowed ligation of the two fragments to proceed using less peptide starting material and enzyme, and the yield also increased slightly to 16%. When the ligation was performed using C247A-OaAEP 1 with the same peptide fragments, an improved yield of 35% was observed. This result agrees with the findings of Rehm et al.52 and Yang et al.28 that C247A-OaAEP 1 displays improved kinetics compared to wt-OaAEP 1, resulting in the more efficient ligation of peptides and proteins. Taken together, these results show that OaAEP 1 is a highly site-specific enzyme, but both N- and C-terminal Asn residues of peptide fragments interfere with OaAEP 1 activity, thus affecting the ligation rate of the peptide. Conversely, OaAEP 1 ligation leaves only an Asn residue footprint in the sequence following ligation. Several studies have demonstrated that the addition of small molecules can further improve the yields of ligated products through quenching the released nucleophiles when using OaAEP 1.64 This can be attempted in future strategies when ligating bivalent peptides with OaAEP 1.
Despite their sequence homology, butelase 1 and OaAEP 1 produced different ligation yields in this study. Butelase 1 ligation of PaurTx3[NHV] and [GVG6]KIIIA took about 6 h to reach the maximum yield (38%) and showed a similar kinetic rate to C247A-OaAEP 1, although butelase 1 has a reported ligation rate of 90 times wt-OaAEP 1.27,28 In our hands, the yield from butelase 1 ligation was only double that of the yield from using wt-OaAEP 1 when applied to the same fragments, and the reaction was slower, 6 h for butelase 1 compared to 4 h for wt-OaAEP 1.
The only chemical approach evaluated in this study, the KAHA ligation, was found to be a promising approach for the chemical conjugation of two peptides each comprising a complex disulfide bond network. With both peptides highly concentrated (15–25 mM), the ligation reaction produced a 36% yield after 6 h. Therefore, ligation using the KAHA method was comparable with respect to yields resulting from butelase 1 and C247A-OaAEP 1 ligations. The P[hSG8]K bivalent peptide produced using KAHA ligation was also the most stable in serum compared to those produced using the enzymatic ligations.
To demonstrate the importance of our modular ligation strategy, we produced linear P[hSerG8]K using KAHA ligation (Figure S6A) and attempted to fold the peptide using our standard PaurTx3 oxidation condition or KIIIA oxidation (Figure S6B), but the desired folding of the peptide was not achieved. While it was straightforward to oxidize each precursor domain in good yield, it was challenging for each domain to obtain its native fold following ligation of the linear peptides. This result emphasized the importance of pre-folding disulfide-rich peptide fragments into their correct structure before ligating them with their partners.
Overall, the SrtA5° approach proved to be highly efficient for ligating disulfide-rich peptide fragments. Despite ligated peptides containing the residual SrtA5° recognition sequence being vulnerable to hydrolysis by the enzyme, we were able to isolate conjugated peptide with a yield of 60% within 15 min. In addition, the synthesis of both SrtA5° precursor fragments produced more materials than other strategies (Table 2).
Evaluation of the activity of the ligated peptides on hNaV1.7 using automated whole cell patch-clamp electrophysiology demonstrated that the amino acid sequence in the linkers in this case is less important than having the correct linker length, which has been previously proven to be critical for optimal activity of ligated PaurTx3 and KIIIA at hNaV1.7.12 All successful ligation approaches delivered bivalent peptides with similar potency, suggesting that the linker sequence neither significantly contributes to channel interactions nor to the orientation of the two ICK domains. On the other hand, evaluation of serum stability of the bivalent peptides included in this study proved to be affected by the composition of amino acids in the linker sequence. The most stable peptide was found to be the chemically KAHA-ligated P[hSG8]K, with 73.3 ± 3.3% of the peptide remaining after 24 h. This is most likely due to the presence of an unnatural residue (homoserine) in the linker sequence of P[hSG8]K, leading to increased resistance to proteases present in human serum compared to the other peptides. While the SrtA5° [LPATG5] linker in P[LPATG5]K also displayed promising stability in serum with 66.9% ± 1.8% remaining after 24 h, the [GVG6] linker used for ligation with OaAEP 1 (wt and C247A) and butelase 1 of P[GVG6]K resulted in only 39.4% ± 1.0% of ligated peptide remaining after 24 h.
Although both OaAEP 1 and butelase 1 have previously been used to cyclize disulfide-rich peptides,27,29–34,52,61–63 to our knowledge, this is the first study to evaluate their ability to ligate two previously oxidized disulfide-rich peptides. In addition, we highlight the versatility of the KAHA-ligation strategy, being able to chemically ligate complex disulfide-rich peptides without reducing or scrambling the disulfide bonds. Although peptiligases, including thymoligase 1 and omniligase 1, have been proven to be promising enzymes for protein and peptide labeling and ligation in previous studies,35–39,52,54,62,65,66 due to the oxidation of PaurTx3[ARVL-OCamL] and PaurTx3[ITTK-OCamL] resulting in misfolded peptide and the pI of the peptides being too close to the required pH for enzyme activity, the peptide fragments precipitated and we could not investigate the efficiency of thymoligase 1 and omniligase 1 in ligating two folded disulfide-rich peptides.
All the ligation strategies used in this work are summarized in Table 3 and are suitable for peptide ligation and cyclization. However, they all come with their own specific advantages and disadvantages. Sortase A-mediated ligation is the most efficient when a more extended enzyme recognition motif in the ligation product is not an issue. The enzyme is straightforward to produce in high yield, resulting in good accessibility to any laboratory. For peptide cyclization, OaAEP 1 and butelase 1 are two of the most efficient enzymes to date, although application and use of these enzymes have been limited due to the poor accessibility, resulting from challenging recombinant expression and low yield during extraction from native materials.33,34 For traceless ligation and cyclization of peptides, peptiligases are considered the best option. The ligation reaction is fast with high ligation yield, and the enzymes are easy to produce. However, an active ester is required as the starting material, which can be challenging to synthesize. Chemical ligation using the KAHA methodology was not the most efficient ligation strategy in terms of conversion of starting materials, but the method requires a short motif and leaves only a homoserine as a footprint at the ligation site, making this ligation suitable for shorter linkers or for mutational studies of the linker itself. After considering all aspects of the ligation strategies investigated here, SrtA5° is, in our hands, the most efficient strategy for ligation of complex disulfide-rich peptides overall. Despite leaving a large footprint of LPATG at the ligation site, in terms of the straightforward production of enzyme, efficiency of ligation, final yield, and ease of purification of the desired product, SrtA5° is the best, provided that the linker is of sufficient length for the enzyme to access the recognition motifs. We envisage that easy synthetic access will open the door to future development of this hitherto underexplored class of molecules.
Table 3.
Advantages and Disadvantages of Ligation Strategies Applied in this Study
| ligation strategies | advantages | disadvantages |
|---|---|---|
| sortase A | ultrafast ligation, good accessibility to the enzyme, high folding/ligation yield | long enzyme motif, motif is intolerant to modification, susceptible to hydrolysis |
| OaAEP 1/butelase 1 | short enzyme motif, traceless ligation, ease of purification | medium ligation, yield poor, accessibility to enzyme challenging |
| peptiligase | traceless amount of used enzyme | synthesis of ester peptides is challenging; not suitable for long, hydrophobic peptides; ligation requires concentrated starting materials |
| KAHA | short recognition motif, homoserine footprint increases protease stability | limited access to the building blocks, ligation requires concentrated starting materials |
METHODS
Peptide Synthesis.
An automated microwave Liberty Prime synthesizer (CEM, USA) was used to assemble peptide precursors on Fmoc-polystyrene resin (RAPP Polymere, 0.72 mmol/g, 0.1 mmol scales) for peptide C-terminal carboxamides and either ketocarboxylic Leu on rink amide resin (Bode Lab, 0.28 mmol/g, 0.1 mmol scales) for PaurTx3-ka or pre-loaded Wang (LL) resin (CEM, 0.31 mmol/g, 0.1 mmol scales) for C-terminal peptide carboxylates. Couplings were performed in dimethylformamide (DMF) using 5 equivalents (equiv) of Fmoc-protected amino acid/5 equiv ethyl cyanohydroxyiminoacetate (Oxyma Pure)/10 equiv diisopropylcarbodiimide (DIC), relative to resin substitution, for 1 min at 105 °C. Pyrrolidine (25% v/v in DMF) was used to remove the Fmoc protecting group (40 s at 100 °C). Cleavage and simultaneous removal of sidechain protecting groups were accomplished by treatment with 95% trifluoracetic acid (TFA)/2.5% triisopropylsilane (TIPS)/2.5% H2O for 2 h at room temperature. Following filtration of cleavage solution, ice-cold diethyl ether (Et2O) was added to precipitate the crude peptide. The crude peptides were centrifuged (5000 rpm for 5 min × 3) in Et2O then lyophilized in 0.05% TFA/50% acetonitrile (ACN)/H2O.
Synthesis with Fmoc-Leu-OCam-Ester.38
Fmoc-glycolic acid (2 eq) was manually coupled at room temperature on Leu pre-loaded Rink amide resin (RAPP Polymere, 0.63 mmol/g) using DIC (5 equiv) and Oxyma pure (5 equiv) in DMF for 30 min. The Fmoc group was deprotected by 25% piperidine in DMF for 2 min × 2. The four amino acids after glycolic acid were coupled manually using 4 equiv of amino acid/4 equiv of DIC/4 equiv of Oxyma pure (20 min × 2). The treated resin was then transferred to Liberty Prime to complete the sequences. Couplings were performed in DMF using 5 equiv of Fmoc-protected amino acid/5 equiv of Oxyma pure/10 equiv of DIC for 2 min at 90 °C, and Fmoc was deprotected by 25% pyrrolidine in DMF 1 min at 90 °C.
Oxidative Folding.
Purified linear PaurTx3 analogs were dissolved in a minimum volume of aqueous ACN and then diluted to a peptide concentration of 0.1 mg/mL with 0.1 M Tris buffer at pH 8 containing 0.81 mM GSH/0.81 mM GSSG and stirred for 24 h at room temperature.53 The reaction was stopped by adding 1% TFA/H2O to pH ~2, and the correctly folded peptide was isolated by HPLC. Purity and correct mass were confirmed by RP-HPLC and ESI-MS, respectively, and 1D 1H NMR experiments were used to confirm the correct fold.
[OprG8]KIIIA was folded in 20% DMSO/H2O for 24 h.Unless otherwise specified, linear KIIIA analogs were folded with a peptide concentration of 0.3 mg/mL in a 0.1 M NH4HCO3 buffer at pH 8 containing 100 equiv of GSH/10 equiv of GSSG and stirred for 24 h at room temperature. Following addition of 1% TFA/H2O to lower the pH to ~2, the peptide was isolated by preparative HPLC and the correct fold was confirmed by ESI-MS and NMR.
Reversed-Phase High-Performance Liquid Chromatography.
Preparative and analytical HPLC were carried out on a Shimadzu LC-20AT system equipped with an SPD-20A Prominence UV/VIS detector, an SIL-20AHT auto-injector, and an FRC-10A fraction collector. An Eclipse XDB–C18 column (Agilent; 7 μm, 21.2 cm × 250 mm, 80 Å, flow rate 8 mL/min) with a gradient from 0 to 50% solvent B in 50 min was used to purify the peptides. A Hypersil GOLD–C18 column (Thermo Fisher; 3 μm, 2.1 × 100 mm, 175 Å, flow rate 0.3 mL/min) with a gradient 0–60% B over 30 min was used to monitor oxidation and ligation and to analyze the purity of the peptides. Absorbance was monitored at 214 and 280 nm. (Solvent A: 0.05% TFA in H2O; solvent B: 90% ACN/0.05% TFA in H2O).
Sortase Ligation.
Peptide ligation was performed using a procedure adapted from Agwa et al.11,67 Briefly, PaurTx3[LPATGG] and [G8]KIIIA were dissolved in SrtA ligation buffer containing 50 mM Tris, 150 mM NaCl, and 10 mM CaCl2 at pH 8 at concentrations of 100 and 300 μM, respectively. SrtA5° was added to this solution to a concentration of 30 μM. The mixture was incubated at 37 °C for 15 min and then acidified with 1% aqueous TFA to stop the reaction and purified by RP-HPLC.
OaAEP 1 Ligation.
For OaAEP 1 ligation, 250 μM PaurTx3[NGL] and 750 μM [G8]KIIIA were dissolved in ligation buffer containing 50 mM Tris, 150 mM NaCl, and 10 mM CaCl2 at pH 6. wt-OaAEP 1 was added to this solution to a concentration of 3 μM. The mixture was left to react at room temperature for 4 h and then acidified with 1% aqueous TFA to stop the reaction. PaurTx3[NGL] (20 μM) and 60 μM [GVG6]KIIIA were dissolved in ligation buffer containing 50 mM Tris, 150 mM NaCl, and 10 mM CaCl2 at pH 6, and wt-OaAEP 1 (or C274A-OaAEP 1) was added to this solution to a concentration of 75 nM. The mixture was left to react at RT for 6 h and then acidified with 1% aqueous TFA to stop the reaction, and the ligated peptide was purified by RP-HPLC.
Butelase 1 Ligation.
PaurTx3[NHV] and [GVG6]KIIIA were dissolved in ligation buffer containing 50 mM Tris, 150 mM NaCl, and 10 mM CaCl2 at pH 6 at concentrations of 20 and 60 μM, respectively. Butelase 1 was added to this solution to a concentration of 100 nM. The mixture was left to react for 6 h and then acidified with 1% aqueous TFA to stop the reaction, and the ligated peptide was purified by RP-HPLC.
Omniligase 1 and Thymoligase 1.
The protocol was previously described by Schmidt and Nuijens.54 Briefly, peptide fragments were dissolved in 50% ACN/H2O or DMSO before being added to enzymatic ligation buffer (1 M potassium phosphate, pH 8.2–8.4, with or without 2 M urea or 1 M Gn·HCl) to make final concentrations of 10 mM Pep-OCamL:15 mM H2N-Pep:8 μM omniligase 1 (or thymoligase 1).
KAHA Ligation.
Oxidized PaurTx3-ka (22 mM) and oxidized [OprG8]KIIIA (15 mM) were dissolved in 0.1 M oxalic acid in DMSO:H2O (9:1) and incubated at 60 °C for 6 h to yield the peptide ester. Similarly for the linear peptide ligation, 22 mM linear PaurTx3-ka and 15 mM linear [OprG8]KIIIA were dissolved in 0.1 M oxalic acid in DMSO:H2O (9:1) with 50 equiv of TCEP and incubated at 60 °C for 7 h to yield the peptide ester. The reactions were monitored by LC/MS, then diluted with buffer A, and purified by RP-HPLC. Lyophilized peptide ester was dissolved in 0.1 M NH4HCO3 at pH 9.5 for 2 h. The O → N shift was monitored by HPLC, then acidified by 1% TFA, and purified.
NMR.
Peptides were dissolved in 500 μL of MilliQ water (MilliPore, USA) and 50 μL of D2O (Cambridge Isotopes). A Bruker 600 MHz Avance III spectrometer equipped with a cryoprobe (Bruker, Billerica, MA, USA) was used to acquire 1D 1H NMR spectra, and 2D 1H-1H TOCSY (80 ms mixing time) and NOESY (200 ms mixing time) were collected,68,69 as described by Agwa et al.11 Spectra were processed using TopSpin 3.5 (Bruker), and CCPNMR Analysis 2.5.2 (CCPN, University of Cambridge, Cambridge, UK) was used to assign resonances.70,71 The chemical shift of water at 4.76 ppm was used as reference.
Serum Stability.
Serum stability assays were performed as previously described.12,72 Briefly, male human AB serum (Sigma) was thawed to room temperature and centrifuged at 17,000g for 15 min to separate and remove the lipid component. The supernatant was incubated at 37 °C for 15 min, and then each peptide was incubated in either 100% serum or PBS at 37 °C at 20 μM. Samples (40 μL) were removed at time points in triplicate (0, 2, 4, 6, 12, and 24 h). Each sample was quenched with 40 μL of 6 M urea and incubated at 4 °C for 10 min. Serum proteins were precipitated with 40 μL of 20% trichloroacetic acid (TCA) followed by another 10 min incubation at 4 °C. Samples were centrifuged at 17,000g for 10 min to remove precipitated proteins. The supernatant was analyzed by analytical HPLC with a linear 0– 50% solvent B gradient over 50 min. Peptides were quantified by integration at 215 nm and identified by mass spectrometry.
HEK 293 Cell Culture.
HEK 293 cells heterologously expressing human NaV1.7 and β1 subunit (SB Drug Discovery, Glasgow, UK) were used for all electrophysiology experiments. Cells were cultured in minimum essential medium Eagle supplemented with 10% (v/v) heat-inactivated fetal bovine serum (HI-FBS), 1% (v/v) GlutaMAX, 4 μg/mL blasticidin, and 0.6 mg/mL geneticin at 37 °C and 5% CO2. Cells were passaged every 3–4 days at 70–80% confluence using TrypLE Express (Thermo Fisher Scientific).
QPatch Electrophysiology.
Electrophysiology assays were performed as previously described using a QPatch-16 automated electrophysiology platform (Sophion Bioscience, Ballerup, Denmark).73 HEK293 cells expressing hNaV1.7/β1 were dissociated and suspended in Ham’s F12 containing 25 mM HEPES, 100 U/mL penicillin–streptomycin, and 0.04 mg/mL soybean trypsin inhibitor. The suspension was stirred for 30–60 min prior to use. The extracellular solution contained (in mM) 70 NaCl, 70 choline chloride, 4 KCl, 2 CaCl2, 1 MgCl2, 10 HEPES, and 10 glucose and was adjusted to pH 7.4 and osmolarity to 305 mOsm. The intracellular solution contained (in mM) 140 CsF, 1 EGTA, 5 CsOH, 10 HEPES, and 10 NaCl and was adjusted to pH 7.3 using CsOH and osmolarity to 320 mOsm.
As described previously,12,74 peptides were diluted in extracellular solution containing 0.1% bovine serum albumin. Each peptide concentration was incubated for 5 min, and the peak current was compared to the buffer control. Concentration–response curves were acquired using a holding potential of −90 mV and a 50 ms pulse to −20 mV every 20 s (0.05 Hz). Peak current post-peptide addition (I) was normalized to the buffer control (I0). IC50 values were determined by plotting the difference in peak current (I/I0) and log peptide concentration. Calculated IC50 values were compared across subtypes, and statistical differences were determined by ordinary one-way ANOVA. I–V curves were obtained with a holding potential of −90 mV followed by a series of 500 ms step pulses that ranged from −110 to +55 mV in 5 mV increments (repetition interval: 5 s). Conductance– voltage curves were obtained by calculating the conductance (G) at each voltage (V) using the equation G = I/(V – Vrev), where Vrev is the reversal potential and was fitted with a Boltzmann equation. Voltage dependence of steady-state fast inactivation was assessed using a 10 ms pulse of −20 mV immediately after the 500 ms step (described above) to assess the availability of non-inactivated channels.
Data Analysis.
GraphPad Prism, v8.0.2 was used for all data analysis and plotting. Concentration–response curves were fitted with a four-parameter Hill equation with variable Hill coefficient. Data are presented as mean ± SEM.
Supplementary Material
ACKNOWLEDGMENTS
The authors would like to thank Prof. Jeffery Bode at ETH Zurich for generous gifts of Fmoc-ketocarboxylic Leu resin and Boc-oxaproline; Dr. Leendert van den Bos at EnzyTag for Fmoc-glycolic acid, omniligase 1, thymoligase 1, and advice on chemical synthesis and ligation of peptides for peptiligase ligation; and Prof. David Liu at Natural Sciences and Chemical Biology at Harvard University for his kind gift of the SrtA5° plasmid.
Funding
This work was funded by the Australian National Health and Medical Research Council (NMHRC) through Project Grants (APP1080405 and 2020002103) awarded to C.I.S. and I.V., respectively, a Career Development Fellowship (APP1162503) awarded to I.V., an Early Career Fellowship (APP1139961) awarded to J.R.D., Australian Research Council grants (ARC; DP1501004430 and CE200100012) and an ARC Australian Laureate Fellowship (FL150100146) awarded to D.J.C., and an ARC Future Fellow (FT160100055) awarded to C.I.S. H.N.T.T., K.L.M. and P.T. were supported by the Australian Government Research Training Program Scholarships.
Footnotes
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.bioconjchem.1c00452.
Analytical RP-HPLC chromatograms of linear and oxidized C-terminal-modified PaurTx3 analogs, analytical RP-HPLC chromatograms of linear and oxidized N-terminal-modified KIIIA analogs, NMR spectra of individual oxidized peptide fragments prior to ligation, PaurTx3 and KIIIA ligation time course for different ligation strategies and enzymes used in this study, 1H NMR spectra of peptide precursors and ligated bivalent peptide constructs, HPLC trace and mass of linear bivalent synthesis, pharmacology at hNaV1.7 of bivalent peptide P[NGVG6]K on human NaV1.7 expressed in HEK 293 cells assessed using automated-patch clamp electrophysiology, HPLC traces of human serum stability assay on bivalent peptides, and theoretical and observed masses of individual peptide fragments and bivalent peptides (PDF).
Complete contact information is available at:
The authors declare no competing financial interest.
Contributor Information
Hue N. T. Tran, Institute for Molecular Bioscience, The University of Queensland, Brisbane, Queensland 4072, Australia
Poanna Tran, Institute for Molecular Bioscience, The University of Queensland, Brisbane, Queensland 4072, Australia.
Jennifer R. Deuis, Institute for Molecular Bioscience, The University of Queensland, Brisbane, Queensland 4072, Australia
Kirsten L. McMahon, Institute for Molecular Bioscience, The University of Queensland, Brisbane, Queensland 4072, Australia
Kuok Yap, Institute for Molecular Bioscience, The University of Queensland, Brisbane, Queensland 4072, Australia.
David J. Craik, Institute for Molecular Bioscience, The University of Queensland, Brisbane, Queensland 4072, Australia
Irina Vetter, Institute for Molecular Bioscience, The University of Queensland, Brisbane, Queensland 4072, Australia; School of Pharmacy, The University of Queensland, Woolloongabba, Queensland 4102, Australia.
Christina I. Schroeder, Institute for Molecular Bioscience, The University of Queensland, Brisbane, Queensland 4072, Australia Center for Cancer Research, National Cancer Institute, National Institutes of Health, Frederick, Maryland 21702, United States.
REFERENCES
- (1).Bohlen CJ; Priel A; Zhou S; King D; Siemens J; Julius D. A bivalent tarantula toxin activates the capsaicin receptor, TRPV1, by targeting the outer pore domain. Cell 2010, 834–845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (2).Chassagnon IR; McCarthy CA; Chin YKY; Pineda SS; Keramidas A; Mobli M; Pham V; de Silva TM; Lynch JW; Widdop RE; Rash LD; King GF Potent neuroprotection after stroke afforded by a double-knot spider-venom peptide that inhibits acid-sensing ion channel 1a. Proc. Natl. Acad. Sci. U. S. A 2017, 114, 3750–3755. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (3).Vassilevski AA; Fedorova IM; Maleeva EE; Korolkova YV; Efimova SS; Samsonova OV; Schagina LV; Feofanov AV; Magazanik LG; Grishin EV Novel class of spider toxin: active principle from the yellow sac spider Cheiracanthium punctorium venom is a unique two-domain polypeptide. J. Biol. Chem 2010, 285, 32293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (4).Gao Y; Cao E; Julius D; Cheng Y. TRPV1 structures in nanodiscs reveal mechanisms of ligand and lipid action. Nature 2016, 534, 347–351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (5).Bae C; Anselmi C; Kalia J; Jara-Oseguera A; Schwieters CD; Krepkiy D; et al. Structural insights into the mechanism of activation of the TRPV1 channel by a membrane-bound tarantula toxin. eLife 2016, 5, No. e11273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (6).Maxwell M; Undheim EAB; Mobli M. Secreted cysteine-rich repeat proteins “SCREPS:” A novel multi-domain architecture. Front. Pharmacol 2018, 9, 1333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (7).Liao M; Cao E; Julius D; Cheng Y. Structure of the TRPV1 ion channel determined by electron cryo-microscopy. Nature 2013, 504, 107–112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (8).Catterall WA; Goldin AL; Waxman SG Nomenclature and structure-function relationships of voltage-gated sodium channels. Pharmacol. Rev 2005, 57, 397–409. [DOI] [PubMed] [Google Scholar]
- (9).Payandeh J; Scheuer T; Zheng N; Catterall WA The crystal structure of a voltage-gated sodium channel. Nature 2011, 475, 353–358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (10).Murray JK; Biswas K; Holder JR; Zou A; Ligutti J; Liu D; Poppe L; Andrews KL; Lin FF; Meng SY; Moyer BD; McDonough SI; Miranda LP Sustained inhibition of the NaV1.7 sodium channel by engineered dimers of the domain II binding peptide GpTx-1. Bioorg. Med. Chem. Lett 2015, 25, 4866–4871. [DOI] [PubMed] [Google Scholar]
- (11).Agwa AJ; Blomster LV; Craik DJ; King GF; Schroeder CI Efficient enzymatic ligation of inhibitor cystine knot spider venom peptides: Using sortase A to form double-knottins that probe voltage-gated sodium channel NaV1.7. Bioconjugate Chem. 2018, 29, 3309–3319. [DOI] [PubMed] [Google Scholar]
- (12).Tran HNT; Tran P; Deuis JR; Agwa AJ; Zhang AH; Vetter I; Schroeder CI Enzymatic ligation of a pore blocker toxin and a gating modifier toxin: creating double-knotted peptides with improved sodium channel NaV1.7 inhibition. Bioconjugate Chem. 2020, 31, 64–73. [DOI] [PubMed] [Google Scholar]
- (13).Peschel A; Cardoso FC; Walker AA; Durek T; Stone MRL; Braga Emidio N; et al. Two for the price of one: Heterobivalent ligand design targeting two binding sites on voltage-gated sodium channels slows ligand dissociation and enhances potency. J. Med. Chem 2020, 63, 12773. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (14).Dirksen A; Hackeng TM; Dawson PE Nucleophilic catalysis of oxime ligation. Angew. Chem., Int. Ed 2006, 45, 7581. [DOI] [PubMed] [Google Scholar]
- (15).Fang GM; Li YM; Shen F; Huang YC; Li JB; Lin Y; et al. Protein chemical synthesis by ligation of peptide hydrazides. Angew. Chem., Int. Ed 2011, 50, 7645. [DOI] [PubMed] [Google Scholar]
- (16).Patil NA; Karas JA; Turner BJ; Shabanpoor F. Thiolcyanobenzothiazole ligation for the efficient preparation of peptide− PNA conjugates. Bioconjugate Chem. 2019, 30, 793–799. [DOI] [PubMed] [Google Scholar]
- (17).van Geel R; Wijdeven MA; Heesbeen R; Verkade JMM; Wasiel AA; van Berkel SS; van Delft FL Chemoenzymatic conjugation of toxic payloads to the globally conserved N-glycan of native mAbs provides homogeneous and highly efficacious antibodydrug conjugates. Bioconjugate Chem. 2015, 26, 2233–2242. [DOI] [PubMed] [Google Scholar]
- (18).Zhang Y; Xu C; Lam HY; Lee CL; Li X. Protein chemical synthesis by serine and threonine ligation. Proc. Natl. Acad. Sci. U. S. A 2013, 110, 6657–6662. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (19).Zheng J-S; Tang S; Qi Y-K; Wang Z-P; Liu L. Chemical synthesis of proteins using peptide hydrazides as thioester surrogates. Nat. Protoc 2013, 8, 2483. [DOI] [PubMed] [Google Scholar]
- (20).Loibl SF; Harpaz Z; Seitz O. A type of auxiliary for native chemical peptide ligation beyond cysteine and glycine junctions. Angew. Chem., Int. Ed 2015, 54, 15055. [DOI] [PubMed] [Google Scholar]
- (21).Dawson PE; Muir TW; Clark-Lewis I; Kent SB Synthesis of proteins by native chemical ligation. Science 1994, 776–779. [DOI] [PubMed] [Google Scholar]
- (22).Cistrone PA; Bird MJ; Flood DT; Silvestri AP; Hintzen JCJ; Thompson DA; Dawson PE Native chemical ligation of peptides and proteins. Curr. Protoc. Chem. Biol 2019, 11, e61–e61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (23).Mitchell NJ; Malins LR; Liu X; Thompson RE; Chan B; Radom L; Payne RJ Rapid additive-free selenocystine− selenoester peptide ligation. J. Am. Chem. Soc 2015, 137, 14011. [DOI] [PubMed] [Google Scholar]
- (24).Kwon S; Bosmans F; Kaas Q; Cheneval O; Conibear AC; Rosengren KJ; et al. Efficient enzymatic cyclization of an inhibitory cystine knot-containing peptide. Biotechnol. Bioeng 2016, 113, 2202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (25).Guimaraes CP; Witte MD; Theile CS; Bozkurt G; Kundrat L; Blom AEM; Ploegh HL Site-specific C-terminal and internal loop labeling of proteins using sortase-mediated reactions. Nat. Protoc 2013, 8, 1787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (26).Stanger K; Maurer T; Kaluarachchi H; Coons M; Franke Y; Hannoush RN Backbone cyclization of a recombinant cystineknot peptide by engineered Sortase A. FEBS Lett. 2014, 588, 4487–4496. [DOI] [PubMed] [Google Scholar]
- (27).Harris KS; Durek T; Kaas Q; Poth AG; Gilding EK; Conlan BF; Saska I; Daly NL; van der Weerden NL; Craik DJ; Anderson MA Efficient backbone cyclization of linear peptides by a recombinant asparaginyl endopeptidase. Nat. Commun 2015, 6, 10199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (28).Yang R; Wong YH; Nguyen GKT; Tam JP; Lescar J; Wu B. Engineering a catalytically efficient recombinant protein ligase. J. Am. Chem. Soc 2017, 139, 5351. [DOI] [PubMed] [Google Scholar]
- (29).Yap K; Du J; Looi FY; Tang SR; de Veer SJ; Bony AR; et al. An environmentally sustainable biomimetic production of cyclic disulfide-rich peptides. Green Chem. 2020, 5002–5016. [Google Scholar]
- (30).Yap K; Du J; Rehm FBH; Tang SR; Zhou Y; Xie J; et al. Yeast-based bioproduction of disulfide-rich peptides and their cyclization via asparaginyl endopeptidases. Nat. Protoc 2021, 16, 2021. [DOI] [PubMed] [Google Scholar]
- (31).Hemu X; Zhang X; Bi X; Liu CF; Tam JP Butelase 1mediated ligation of peptides and proteins. Methods Mol. Biol 2019, 2012, 83–109. [DOI] [PubMed] [Google Scholar]
- (32).James AM; Haywood J; Leroux J; Ignasiak K; Elliott AG; Schmidberger JW; Fisher MF; Nonis SG; Fenske R; Bond CS; Mylne JS The macrocyclizing protease butelase 1 remains autocatalytic and reveals the structural basis for ligase activity. Plant J. 2019, 98, 988–999. [DOI] [PubMed] [Google Scholar]
- (33).Nguyen GKT; Kam A; Loo S; Jansson AE; Pan LX; Tam JP Butelase 1: A versatile ligase for peptide and protein macrocyclization. J. Am. Chem. Soc 2015, 137, 15398. [DOI] [PubMed] [Google Scholar]
- (34).Nguyen GKT; Wang S; Qiu Y; Hemu X; Lian Y; Tam JP Butelase 1 is an Asx-specific ligase enabling peptide macrocyclization and synthesis. Nat. Chem. Biol 2014, 10, 732–738. [DOI] [PubMed] [Google Scholar]
- (35).Nuijens T; Toplak A; Mathijs BAC; Schmidt M; Michel G; Dick B; Quaedflieg P. Chemo-enzymatic peptide synthesis (CEPS) using omniligases and selective peptiligases. Efficient biocatalysts for assembling linear and cyclic peptides and protein conjugates. Chim. Oggi – Chem. Today 2016, 34, 16–19. [Google Scholar]
- (36).Nuijens T; Toplak A; Quaedflieg PJLM; Drenth J; Wu B; Janssen DB Engineering a diverse ligase toolbox for peptide segment condensation. Adv. Synth. Catal 2016, 358, 4041. [Google Scholar]
- (37).Schmidt M; Toplak A; Quaedflieg PJLM; Ippel H; Richelle GJJ; Hackeng TM; et al. Omniligase-1: A powerful tool for peptide head-to-tail cyclization. Adv. Synth. Catal 2017, 359, 2050. [Google Scholar]
- (38).Nuijens T; Toplak A; van de Meulenreek MBAC; Schmidt M; Goldbach M; Quaedflieg PJLM Improved solid phase synthesis of peptide carboxyamidomethyl (Cam) esters for enzymatic segment condensation. Tetrahedron Lett. 2016, 57, 3635–3638. [Google Scholar]
- (39).Schmidt M; Toplak A; Rozeboom HJ; Wijma HJ; Quaedflieg PJLM; van Maarseveen JH; et al. Design of a substrate-tailored peptiligase variant for the efficient synthesis of thymosin-α1. Org. Biomol. Chem 2018, 16, 609. [DOI] [PubMed] [Google Scholar]
- (40).Bode JW; Fox RM; Baucom KD Chemoselective amide ligations by decarboxylative condensations of N-alkylhydroxylamines and α-ketoacids. Angew. Chem., Int. Ed 2006, 45, 1248. [DOI] [PubMed] [Google Scholar]
- (41).Fukuzumi T; Bode JW A reagent for the convenient, solid-phase synthesis of n-terminal peptide hydroxylamines for chemoselective ligations. J. Am. Chem. Soc 2009, 131, 3864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (42).Ju L; Lippert AR; Bode JW Stereoretentive synthesis and chemoselective amide-forming ligations of C-terminal peptide αketoacids. J. Am. Chem. Soc 2008, 130, 4253. [DOI] [PubMed] [Google Scholar]
- (43).Pusterla I; Bode JW The mechanism of the α-ketoacid–hydroxylamine amide-forming ligation. Angew. Chem., Int. Ed 2012, 51, 513. [DOI] [PubMed] [Google Scholar]
- (44).Pusterla I; Bode JW An oxazetidine amino acid for chemical protein synthesis by rapid, serine-forming ligations. Nat. Chem 2015, 7, 668–672. [DOI] [PubMed] [Google Scholar]
- (45).Bosmans F; Rash L; Zhu S; Diochot S; Lazdunski M; Escoubas P; Tytgat J. Four novel tarantula toxins as selective modulators of voltage-gated sodium channel subtypes. Mol. Pharmaceutics 2006, 69, 419–429. [DOI] [PubMed] [Google Scholar]
- (46).Agwa AJ; Peigneur S; Chow CY; Lawrence N; Craik DJ; Tytgat J; et al. Gating modifier toxins isolated from spider venom: Modulation of voltage-gated sodium channels and the role of lipid membranes. J. Biol. Chem 2018, 293, 9041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (47).Bulaj G; West PJ; Garrett JE; Watkins M; Zhang M-M; Norton RS; Smith BJ; Yoshikami D; Olivera BM Novel conotoxins from conus striatus and conus kinoshitai selectively block TTX-resistant sodium channels. Biochemistry 2005, 44, 7259–7265. [DOI] [PubMed] [Google Scholar]
- (48).Pan X; Li Z; Huang X; Huang G; Gao S; Shen H; Liu L; Lei J; Yan N. Molecular basis for pore blockade of human Na+ channel NaV1.2 by the μ-conotoxin KIIIA. Science 2019, 1309–1313. [DOI] [PubMed] [Google Scholar]
- (49).Xu H; Li T; Rohou A; Arthur CP; Tzakoniati F; Wong E; Estevez A; Kugel C; Franke Y; Chen J; Ciferri C; Hackos DH; Koth CM; Payandeh J. Structural basis of NaV1.7 inhibition by a gating-modifier spider toxin. Cell 2019, 176, 702–715.e14. [DOI] [PubMed] [Google Scholar]
- (50).Corpuz GP; Jacobsen RB; Jimenez EC; Watkins M; Walker C; Colledge C; Garrett JE; McDougal O; Li W; Gray WR; Hillyard DR; Rivier J; McIntosh JM; Cruz LJ; Olivera BM Definition of the μ-conotoxin superfamily: Characterization of novel peptides from molluscivorous Conus venoms. Biochemistry 2005, 44, 8176–8186. [DOI] [PubMed] [Google Scholar]
- (51).Khoo KK; Gupta K; Green BR; Zhang M-M; Watkins M; Olivera BM; Balaram P; Yoshikami D; Bulaj G; Norton RS Distinct disulfide Isomers of μ-conotoxins KIIIA and KIIIB block voltage-gated sodium channels. Biochemistry 2012, 51, 9826–9835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (52).Rehm FBH; Harmand TJ; Yap K; Durek T; Craik DJ; Ploegh HL Site-specific sequential protein labeling catalyzed by a single recombinant ligase. J. Am. Chem. Soc 2019, 141, 17388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (53).Agwa AJ; Huang Y-H; Craik DJ; Henriques ST; Schroeder CI Lengths of the C-terminus and interconnecting loops impact stability of spider-derived gating modifier toxins. Toxins 2017, 9, 248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (54).Schmidt M; Nuijens T. Chemoenzymatic synthesis of Linearand Head-to-Tail cyclic peptides using omniligase-1, In Enzyme-Mediated Ligation Methods; (Nuijens T; Schmidt M, Eds.), Springer New York, New York, NY, 2019. pp. 43–61. [DOI] [PubMed] [Google Scholar]
- (55).Deuis JR; Dekan Z; Wingerd JS; Smith JJ; Munasinghe NR; Bhola RF; Imlach WL; Herzig V; Armstrong DA; Rosengren KJ; Bosmans F; Waxman SG; Dib-Hajj SD; Escoubas P; Minett MS; Christie MJ; King GF; Alewood PF; Lewis RJ; Wood JN; Vetter I. Pharmacological characterisation of the highly NaV1.7 selective spider venom peptide Pn3a. Sci. Rep 2017, 7, 40883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (56).Zeng X; Li P; Chen B; Huang J; Lai R; Liu J; Rong M. Selective closed-state NaV1.7 blocker JxTx-34 exhibits analgesic effects against pain. Toxins 2018, 10, 64–13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (57).Mueller A; Dekan Z; Kaas Q; Agwa AJ; Starobova H; Alewood PF; Schroeder CI; Mobli M; Deuis JR; Vetter I. Mapping the molecular surface of the analgesic NaV1.7-selective peptide Pn3a reveals residues essential for membrane and channel interactions. ACS Pharmacol. Transl. Sci 2020, 3, 535–546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (58).Mueller A; Starobova H; Morgan M; Dekan Z; Cheneval O; Schroeder CI; Alewood PF; Deuis JR; Vetter I. Antiallodynic effects of the selective NaV1.7 inhibitor Pn3a in a mouse model of acute postsurgical pain: evidence for analgesic synergy with opioids and baclofen. Pain 2019, 160, 1766–1780. [DOI] [PubMed] [Google Scholar]
- (59).Stanciu G-D; Luca A; Marza A; Alexa-Stratulat T; Tudorancea I; Bild W; Rezus E; Rezus C; Tamba BI Intracerebroventricular coadministration of Protoxin-II and trace elements in rats enhances the analgesic effect of the 1.7 voltage-gate sodium channel blocker. Biomed Res. Int 2019, 2019, 8057803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (60).Xiao Y; Blumenthal K; Jackson JO II; Liang S; Cummins TR The tarantula toxins ProTx-II and huwentoxin-IV differentially interact with human Nav1.7 voltage sensors to inhibit channel activation and inactivation. Mol. Pharmacol 2010, 78, 1124–1134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (61).Jackson MA; Gilding EK; Shafee T; Harris KS; Kaas Q; Poon S; Yap K; Jia H; Guarino R; Chan LY; Durek T; Anderson MA; Craik DJ Molecular basis for the production of cyclic peptides by plant asparaginyl endopeptidases. Nat. Commun 2018, 9, 2411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (62).Rehm FBH; Jackson MA; De Geyter E; Yap K; Gilding EK; Durek T; Craik DJ Papain-like cysteine proteases prepare plant cyclic peptide precursors for cyclization. Proc. Natl. Acad. Sci. U. S. A 2019, 116, 7831–7836. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (63).Harris KS; Guarino RF; Dissanayake RS; Quimbar P; McCorkelle OC; Poon S; Kaas Q; Durek T; Gilding EK; Jackson MA; Craik DJ; van der Weerden NL; Anders RF; Anderson MA A suite of kinetically superior AEP ligases can cyclise an intrinsically disordered protein. Sci. Rep 2019, 10820. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (64).Rehm FBH; Tyler TJ; Yap K; Durek T; Craik DJ Improved Asparaginyl-Ligase-Catalyzed Transpeptidation via Selective Nucleophile Quenching. Angew. Chem., Int. Ed 2021, 60, 4004. [DOI] [PubMed] [Google Scholar]
- (65).Pawlas J; Nuijens T; Persson J; Svensson T; Schmidt M; Toplak A; et al. Sustainable, cost-efficient manufacturing of therapeutic peptides using chemo-enzymatic peptide synthesis (CEPS). Green Chem. 2019, 21, 6451–6467. [Google Scholar]
- (66).Polgar L; Bender ML The reactivity of thiol-subtilisin, an enzyme containing a synthetic functional group. Biochemistry 1967, 6, 610–620. [DOI] [PubMed] [Google Scholar]
- (67).Agwa AJ, Craik DJ; Schroeder CI (2019) Cyclizing disulfide-rich peptides using sortase A, In Enzyme-Mediated Ligation Methods (Nuijens T; Schmidt M, Eds.), pp. 29–41, Springer New York, New York, NY. [DOI] [PubMed] [Google Scholar]
- (68).Jeener J; Meier BH; Bachmann P; Ernst RR Investigation of exchange processes by two-dimensional NMR spectroscopy. J. Chem. Phys 1979, 71, 4546. [Google Scholar]
- (69).Braunschweiler L; Ernst RR Coherence transfer by isotropic mixing: Application to proton correlation spectroscopy. J. Magn. Reson 1983, 53, 521. [Google Scholar]
- (70).Wüthrich K. NMR of proteins and nucleic acids; Wiley Interscience: New York, 1986. [Google Scholar]
- (71).Vranken WF; Boucher W; Stevens TJ; Fogh RH; Pajon A; Llinas M; et al. The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 2005, 59, 687–696. [DOI] [PubMed] [Google Scholar]
- (72).Gunasekera S; Foley FM; Clark RJ; Sando L; Fabri LJ; Craik DJ; Daly NL Engineering stabilized vascular endothelial growth factor-a antagonists: synthesis, structural characterization, and bioactivity of grafted analogues of cyclotides. J. Med. Chem 2008, 51, 7697. [DOI] [PubMed] [Google Scholar]
- (73).Deuis JR; Dekan Z; Inserra MC; Lee TH; Aguilar MI; Craik DJ; Lewis RJ; Alewood PF; Mobli M; Schroeder CI; Henriques ST; Vetter I. Development of a μO-conotoxin analogue with improved lipid membrane interactions and potency for the analgesic sodium channel NaV1.8. J. Biol. Chem 2016, 291, 11829–11842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (74).McMahon KL; Tran HNT; Deuis JR; Lewis RJ; Vetter I; Schroeder CI Discovery, pharmacological characterisation and NMR structure of the novel μ-conotoxin SxIIIC, a potent and irreversible NaV channel inhibitor. Biomedicines 2020, 8, 391–406. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.