Abstract
We aimed to develop a whole-genome sequencing (WGS)-based copy number variant (CNV) calling algorithm with the potential of replacing chromosomal microarray assay (CMA) for clinical diagnosis. JAX-CNV is thus developed for CNV detection from WGS data. The performance of this CNV calling algorithm was evaluated in a blinded manner on 31 samples and compared to the 112 CNVs reported by clinically validated CMAs for these 31 samples. The result showed that JAX-CNV recalled 100% of these CNVs. Besides, JAX-CNV identified an average of 30 CNVs per individual, respresenting an approximately seven-fold increase compared to calls of clinically validated CMAs. Experimental validation of 24 randomly selected CNVs showed one false positive, i.e., a false discovery rate (FDR) of 4.17%. A robustness test on lower-coverage data revealed a 100% sensitivity for CNVs larger than 300 kb (the current threshold for College of American Pathologists) down to 10× coverage. For CNVs larger than 50 kb, sensitivities were 100% for coverages deeper than 20×, 97% for 15×, and 95% for 10×. We developed a WGS-based CNV pipeline, including this newly developed CNV caller JAX-CNV, and found it capable of detecting CMA-reported CNVs at a sensitivity of 100% with about a FDR of 4%. We propose that JAX-CNV could be further examined in a multi-institutional study to justify the transition of first-tier genetic testing from CMAs to WGS. JAX-CNV is available at https://github.com/TheJacksonLaboratory/JAX-CNV.
Keywords: Copy number variant, Chromosomal microarray assay, Whole-genome sequencing, JAX-CNV, Genetic testing
Introduction
Copy number variants (CNVs) are known to play key roles in human evolution, genomic diversity, and disease susceptibility [1], [2], [3], [4], [5]. In addition, copy number changes have been reported to cause microdeletion and microduplication syndromes, such as Williams syndrome, Prader-Willi syndrome, Angelman syndrome, and DiGeorge syndrome [5], [6], [7], [8], [9]. Various technologies, including fluorescence in situ hybridization (FISH), PCR-based assays, chromosomal microarray assays (CMAs), and next-generation sequencing (NGS), have been developed in research and clinical laboratories for CNV detection. Since 2010, CMAs have been considered the first-tier test for patients with unexplained developmental delay or intellectual disability, autism spectrum disorders, and congenital anomalies [10], [11].
Over the past decade, advances in NGS technologies have brought unprecedented improvements in the throughput, speed, and cost of DNA sequencing. These improvements make whole-genome sequencing (WGS) feasible for broad use in research, with its ability to detect many types of genetic variations, and promise to offer the potential of a single test that captures nearly all genomic variations in an unbiased manner [12], [13], [14]. Although several WGS-based CNV calling algorithms were developed [15], [16], [17], [18], [19], [20], [21], none of them has been widely accepted for clinical applications, because callsets of those algorithms are not highly concordant with the current running assay (i.e., CMA).
In addition to developing new CNV calling algorithms, integrative pipelines that combine multiple CNV calling algorithms to improve accuracy and overcome limitations of individual performance are commonly used. For example, Zhou et al. [22] developed a method to integrate callsets of CNVnator [19] and Lumpy [16]. Noll et al. proposed SKALD [23], which is based on consensus, filtered calls from BreakDancer [24] and GenomeSTRiP [25]. Trost et al. [26] developed a pipeline that employs CNVnator and ERDS [27] for CNV identification. However, most of those pipelines are not open source, and apply filters that are specifically developed for individual projects but not standardized. Thus, it is unknown whether the sensitivity and specificity of those pipelines are comparable to the standards in clinical diagnosis. Given current limitations, an algorithmic pipeline with sufficient sensitivity and specificity for clinical application is demanded.
Here, we present JAX-CNV, a newly developed WGS-based CNV calling algorithm. Its performance was evaluated on WGS data from 31 patient samples and compared to the callsets of the clinically validated CMAs at the Jackson Laboratory for Genomic Medicine (JAX-GM CMA). The result suggests that JAX-CNV has a high sensitivity (100%) necessary for diagnostic decisions and a low false discovery rate (FDR; 4%). This algorithm could serve as a basis for the use of WGS, as a replacement for array-based clinical genetic testing.
Method
WGS analysis workflow
Pre-processing
The pre-processing step for a given reference genome (such as GRCh37, GRCh38, or other versions of human reference genome) includes BWA index (v0.7.15) and Jellyfish [28] count (v2.2.6). BWA index creates the required files for BWA alignment, while Jellyfish calculates the counts of each 25-mer of the reference genome and generates k-mer database (k-mer DB in Figure 1A) to indicate mappabilities of regions. This is similar to the concept proposed in CLImAT-HET [29], using k-mer for low mappability region identification. Theoretically, a larger k will increase the uniqueness of a k-mer against the given reference genome. However, that also requires larger computational resources to process larger k-mers. An experiment of different k-mers (k from 5 to 40) shows that a 25-mer achieves 76% unique k-mer (File S1; k-mer selection) and maintains the final sensitivity of 100% that is the result from the pipeline of multiple steps. The final sensitivity does not depend on k-mer selection solely. To efficiently compress the file size, each number in Jellyfish k-mer DB is converted to a character by calculating its log2 value added by 33 because ASCII code 33 in decimal is the first printable and visual character. For example, if a 25-mer has only one position in the genome, the log2 (1) value is zero. For converting the zero to the first printable ASCII code, 33 is added to the original log2 value (k-mer FASTA in Figure 1A). BWA index, Jellyfish count, and k-mer DB conversion may take 190 min, 105 min, and 403 min, respectively.
Figure 1.
WGS analysis workflow
A. An overview of the CNV calling pipeline consisting of three major steps: pre-processing, alignment, and CNV calling. B.–J. Details of CNV calling approach. WGS, whole-genome sequencing; CNV, copy number variant; DB, database; HMM, hidden Markov model.
Alignment
The analysis could start as raw FASTQ files or BWA aligned BAM/CRAM files. If FASTQ is given, FastQC (v0.11.5) would be applied additionally in the pipeline for quality control. Then, BWA mem (v0.7.15) is employed for mapping reads against a given human reference genome. Once alignments are obtained (from our pipeline or given by users), alignments are sorted by SAMtools and used as the input file of JAX-CNV. Figure 1B illustrates read depths after alignment.
CNV calling
Coverage calculation
JAX-CNV uses the log2 (25-mer) FASTA-format file from the pre-processing step to scan the high-confident mapping (unique) regions for an accurate coverage calculation. A region is considered as a high-confident mapping region when each 25-mer count in the region is one, and the size of the region is larger than 20 kb. For each chromosome, we calculate an average coverage based on those 20 high-confident regions on a given BAM/CRAM of a sample. Once average coverages of all chromosomes are obtained, the interquartile range is applied to filter outliers. Outliers could indicate trisomy, monosomy, and other gross chromosome number anomalies of a sample. Then, an overall coverage of the sequenced sample is calculated based on average coverages of all chromosomes excluding the outliers. Using the interquartile range method, we are able to detect aneuploidies. For those aneuploidies, we will not detect any smaller CNVs on the respective chromosomes in the further steps.
Hidden Markov model
The overall coverage of the sequenced sample is used as the baseline and set to the 50th percentile (Figure 1C). Next, we scan the BAM/CRAM by shifting bins (the default bin size is set to 50 bp, and it is user-adjustable by “--bin”) and assign a percentile for each bin according to the difference between the overall coverage and the read depth of the bin. The percentiles normally range from 0% to 180%, and 180%–200% are reserved for copy numbers larger than two. For example, if the overall coverage is 50× and a read depth of a bin is 100×, the percentile of the bin will be 100% [(100/50) × 50%; Figure 1C]. Then, a hidden Markov model (HMM) with a Poisson distribution [30] is applied to convert the percentile of each bin to one of the five HMM CNV statuses: CN = 0 (loss), CN = 1 (loss), CN = 2 (normal), CN = 3 (gain), and CN > 3 (gain), (Figure 1D). Afterward, if the CNV statuses of the two adjacent bins are the same, we merge them into a segment (Figure 1E).
CNV merging
Since the default bin size is set to 50 bp, which is smaller than the CNVs that we expect to detect, oscillations of CNV statuses could happen frequently as illustrated in Figure 1E. There is a small CN = 1 status inside a large CN = 0 region. Using larger bins may resolve oscillations, but it decreases sensitivity. To maintain high sensitivity, we set the default bin size to be 50 bp, which is user-adjustable by “--bin”, followed by a merging step. The merging step is necessary to mitigate oscillations caused by the uneven read depth problem which is common in WGS alignments. To show oscillations of CNV statuses and uneven read depth problems, Figure S1 shows coverage, log2 (25-mer), and the ratio of low-quality alignments for the 112 truth CNVs.
If the length of a CNV status is shorter than 5 kb, then it will be absorbed by the previous status (Figure 1F) to form a cluster regardless of the CNV type (loss or gain). This status consolidation may be too aggressive, namely merging too many other CNV types together into a cluster, leading to incorrect results. Thus, for each status consolidation, if the original CNV type of a cluster covers less than 80% of the total length of the current cluster, the merging stops and the CNV calling status reinstates the original status (Figure 1G). This approach also prevents small statuses from merging into large statuses.
After the completion of merging, all CNV regions are sorted by their respective lengths (Figure 1H). From the largest to the smallest, each CNV region checks other CNVs of the same type (loss or gain) on their downstream and upstream coordinates from the nearest to the farthest for further clustering (Figure 1I). The aforementioned procedure stops when encountering a CNV with a different type (loss or gain). This step allows larger CNVs to cross normal status (CN = 2) and to merge smaller CNVs nearby. Starting from the largest CNV enables larger CNVs to have more opportunities to cross normal statuses and merge with other CNVs of the same type. Candidate CNVs are then generated.
CNV filtering
For each candidate CNV obtained from the previous step, CNV calling: CNV merging, we divide it into ten bins of equal length. Each bin is assigned a unique value corresponding to the count of unique k-mers (i.e., the number of 25-mers that have a unique position in the reference genome). Starting from the bin with the smallest coordinate (the most left) to the bin with the largest coordinate, we filter a bin if the uniqueness value is low (percentage of unique k-mers is lower than 60% by default; user-adjustable by “--unique_kmer”). The procedure stops after encountering a low uniqueness bin (Figure 1J). The aforementioned procedure is repeated from the bin with the largest coordinate (the most right) to the bin with the smallest coordinate. This approach helps us trim the tails of CNVs in low-confident mapping regions.
Then, the density-based spatial clustering of applications with noise (DBSCAN) algorithm [31] is employed to cluster the candidate CNV fragments as the final step to making a CNV call. For this final step, using DBSCAN helps us to have a global view of CNV regions and consolidate them better. Specifically, we first sort the candidate CNV fragments based on their coordinates. Then, we separate the fragments into different raw clusters by two conditions: 1) the distance of any two continuous fragments is <3 Mb, and 2) the types (loss or gain) of all fragments are located in the raw cluster region. Next, for each raw cluster, we calculate the distance of every continuous fragment pair. The mean distance of the raw cluster is also calculated. The DBSCAN function (in DBSCAN R package) is applied to the distance matrix of each raw cluster. The procedure ends when the cluster results are not changed (File S1; DBSCAN). We report CNVs for a given individual in a BED-format file.
Assessment of pathogenicity of a CNV
The criteria and guidelines used for the classification and interpretation of CNVs were published previously [10], [11], [32]. In general, a CNV is classified as pathogenic if 1) it overlaps genomic coordinates for a well-known deletion or duplication syndrome; or 2) it contains disease genes reported in GeneReviews (https://www.ncbi.nlm.nih.gov/books/NBK1116/), OMIM (https://www.omim.org/), or Decipher (https://decipher.sanger.ac.uk/disorders#syndromes/overview).
Droplet digital PCR validation
Droplet digital PCR (ddPCR) assays were performed to examine the accuracy of the genomic aberrations detected by the JAX-GM CMA platform (File S1; ddPCR validation) and the JAX-CNV algorithm. The customized assays utilized primers designed by Primer3Plus [33], based on the GRCh38 assembly. All primer pairs were tested for their uniqueness across the human genome using in silico PCR from UCSC Genome Browser. The BLAST-Like Alignment Tool (BLAT) search was also performed at the same time to make sure all primer candidates only hit one site of the human genome. Lastly, the NCBI 1000 Genomes Browser was used to check if there were any single nucleotide variations (SNVs) in the primer or probe-binding region. All primers and probes used in this study are listed in Table S1.
Results
Dataset
Currently, the microarray proficiency test offered by the College of American Pathologists (CAP) requests participating clinical laboratories to report CNVs larger than 300 kb [32]. We selected 31 samples associated with various constitutional disorders (i.e., DiGeorge syndrome, Williams syndrome, Cri-du-chat syndrome, Smith-Magenis syndrome, Wolf-Hirschhorn syndrome, Miller-Dieker Lissencephaly syndrome, Tetralogy of Fallot syndrome, 1p deletion syndrome, 18p deletion syndrome, and Angelman syndrome) from the Corriell Institute. For 22 of these 31 samples, the Corriell Institute reports a total of 45 CNVs (25 deletions and 20 duplications, ranging from 101 kb to 94 Mb in size) (Table 1, Table S2).
Table 1.
Detection of the 45 Corriell-registered CNV calls by JAX-CNV pipeline
| Corriell ID | Corriell description | CNV region (length) | CNV type | Pathogenic annotation |
JAX-CNV |
||||
|---|---|---|---|---|---|---|---|---|---|
| Original_coverage (30×–48×) | 30× | 20× | 15× | 10× | |||||
| GM02820 | Chromosome aberration | 9p24.3p13.3 (34.5 Mb) | DUP | G/M | + | + | + | + | + |
| 12q24.32 q24.33 (7.3 Mb) | DEL | G/M | + | + | + | + | + | ||
| GM03997 | Derivative chromosome | 5q35.1 (130 kb) | DUP | M | + | + | + | + | + |
| 12p13.33p12.2 (20.8 Mb) | DUP | G/D/M | + | + | + | + | + | ||
| 12q24.33 (623 kb) | DEL | G/M | + | + | + | + | + | ||
| GM05876 | DiGeorge syndrome | 22q11.21 (1.4 Mb) | DEL | G/D/M | + | + | + | + | + |
| GM09025 | Ring chromosome | 16q24.2 (383 kb) | DUP | G/M | + | + | + | + | + |
| 22q13.31q13.33 (2.9 Mb) | DEL | G/D/M | + | + | + | + | + | ||
| GM09209 | Miller-Dieker Lissencephaly syndrome | 17p13.3 (5.9 Mb) | DEL | G/D/M | + | + | + | + | + |
| GM09687 | Recombinant chromosome | 16p13.3 (1.1 Mb) | DEL | G/D/M | + | + | + | + | + |
| 16q22.1q24.3 (20 Mb) | DUP | G/M | + | + | + | + | + | ||
| GM09711 | Dicentric chromosome | 2q13 (140 kb) | DUP | G/M | + | + | + | + | * |
| 13q11q34 (94 Mb) | DUP | G/M | + | + | + | + | + | ||
| 13q34 (1.7 Mb) | DEL | M | + | + | + | + | + | ||
| GM10946 | Recombinant chromosome | 6p21.2p21.1 (964 kb) | DUP | G/M | + | + | + | + | + |
| 6p12.3 (780 kb) | DUP | + | + | + | + | + | |||
| 6q14.1q16.3 (25 Mb) | DEL | G/M | + | + | + | + | + | ||
| GM11428 | Duplicated chromosome | 3p26.3p26.2 (5.3 Mb) | DEL | G/M | + | + | + | + | + |
| 3q22.1q26.1 (29.8 Mb) | DUP | G/D/M | + | + | + | + | + | ||
| 3q26.1 (112.8 kb) | DEL | + | + | + | + | + | |||
| 3q26.1q29 (35.2 Mb) | DUP | G/M | + | + | + | + | + | ||
| GM11516 | Angelman syndrome | 15q11.2q13.1 (7 Mb) | DEL | G/D/M | + | + | + | + | + |
| GM13480 | Williams syndrome | 7q11.23 (1.6 Mb) | DEL | G/D/M | + | + | + | + | + |
| 9p24.1 (107.6 kb) | DUP | + | + | + | + | + | |||
| GM13590 | Duplicated chromosome | 2q11.2q21.1 (33.6 Mb) | DUP | G/M | + | + | + | + | + |
| 2q37.3 (119 kb) | DEL | + | + | + | + | + | |||
| 4q31.22 (101.5 kb) | DEL | M | + | + | + | + | + | ||
| 9p13.3 (120.3 kb) | DUP | G/M | + | + | + | * | * | ||
| 17q11.1 (101 kb) | DUP | M | + | + | + | + | + | ||
| GM13946 | Williams syndrome | 7q11.23q11.23 (1.6 Mb) | DEL | G/D/M | + | + | + | + | + |
| GM14164 | Tetralogy of fallot | 13q14.2 (47.9 Mb) | DEL | G/M | + | + | + | + | + |
| 22q11.21 (148.8 kb) | DUP | M | + | + | + | + | No | ||
| GM16580 | 18p deletion syndrome | 18p11.32 (1.6 Mb) | DEL | M | + | + | + | + | + |
| 18q21.33q23 (13.5 Mb) | DUP | M | + | + | + | + | + | ||
| 18q23 (4.0 Mb) | DEL | G/M | + | + | + | + | + | ||
| GM16593 | Cri-du-chat syndrome | 5p15.3 (14.7 Mb) | DEL | G/M | + | + | + | + | + |
| 14q24.3 (2.7 Mb) | DEL | M | + | + | + | + | + | ||
| GM18828 | Chromosome aberration | 1q31.3 (118 kb) | DUP | G/M | + | + | + | + | No |
| 4p16.1 (140 kb) | DUP | M | + | + | + | + | + | ||
| GM20200 | Isodicentric chromosome | 1q31.3 (103 kb) | DEL | G/M | + | + | + | + | + |
| 15q11.1q13.1 (8.5 Mb) | DUP | G/D/M | + | + | + | + | + | ||
| GM20375 | Angelman syndrome | 15q11.2q13.1 (4.9 Mb) | DEL | G/D/M | + | + | + | + | + |
| GM20743 | Smith-Magenis syndrome | 17p11.2 (2.1 Mb) | DEL | G/D/M | + | + | + | + | + |
| GM22569 | 1p deletion syndrome | 1p36.33 (5.5 Mb) | DEL | G/M | + | + | + | + | + |
| GM22601 | Wolf-Hirschhorn syndrome | 4p16.3 (25.0 Mb) | DEL | G/D/M | + | + | + | + | + |
Note: In the “pathogenic annotation” column, ‘G’, ‘D’, and ‘M’ mean annotations from GeneReviews, Decipher, and OMIM databases, respectively. In the “JAX-CNV” column, ‘+’ denotes CNVs captured by the methods/coverages; ‘*’ denotes that CNVs are not 50% reciprocal overlapping but recovered in manual review; and ‘No’ means no call. DEL, deletion; DUP, duplication.
These 31 samples were also examined with a clinically validated Affymetrix CytoScan HD platform (Affymetrix, Santa Clara, CA) for the detection of chromosomal imbalances following the standard operating procedures of the CLIA-certified laboratory at JAX-GM. The CMA data analysis was performed using the software supplied by the vendor (ChAS v3.3; methods: Affymetrix CytoScan HD analysis flow). The JAX-GM CMA platform reported an additional 67 CNVs among these 31 individuals, using a size cutoff of 50 kb (Figure 2; Table S3). In total, these 112 CNVs (65 deletions and 47 duplications, ranging from 51 kb to 94 Mb in size) were used as the truth set and set an initial baseline for sensitivity analysis. Of note, 70 of the 112 CNVs were considered to be pathogenic based on the previously published criteria and guidelines [32], [34], including 41Corriell-registered CNVs and 29 CNVs detected by the JAX-GM CMA.
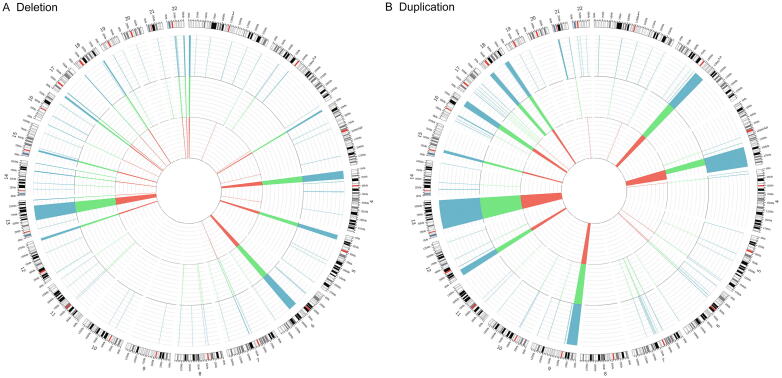
Figure 2.
JAX-CNV accurately detected all CNVs in the 31 testing samples
A. Deletion. B. Duplication. The 31 testing samples were examined by three different methods, including the Corriell Institute CMA (red inner circle), the JAX-GM CMA (green middle circle), and the JAX-CNV algorithm (blue outer circle).
JAX-CNV showed a 100% recall rate of CNVs detected by arrays
We performed WGS on these 31 samples by Illumina paired-end sequencing at 30×–48× coverage with the read length of 2 × 150 bp (Table S4) in a blinded study manner. BWA was applied for short read alignment against the GRCh38 human reference genome (Chr1–22, X, Y, and mitochondria), followed by CNV identification using the JAX-CNV algorithm. JAX-CNV accurately detected all 112 CNVs of the truth set from WGS data (Figure 2; Table 1). Figure 2 shows the aggregated CNVs of all samples, and the locations of CNVs of each sample are given in Figure S2 and Table S5. Of note, due to the different resolutions of CMAs and WGS, there were three deletions ranging from 104 kb to 291 kb in size (one from GM20375 and the other two from GM20743) and three duplications ranging from 105 kb to 292 kb in size (GM09687, GM13480, and GM20743) that did not meet the benchmark of 50% reciprocal overlap with the JAX-CNV calls, but they were still located in the same regions with either smaller or larger size ones (Figure S3).
Next, the CNV calls from JAX-CNV compared with the truth set. As shown in Figure 3A, JAX-CNV detected an additional 747 more CNVs than the array-based technology, and 89% of these calls were less than 300 kb in size, and 50% were less than 100 kb in size. Figure 3B further summarizes CNV calls of each sample. JAX-CNV used 119 calls to identify 112 CNVs in the truth set. For example, for a 20.8-Mb duplication at the chromosome region 12p13–12p12 in GM03997, JAX-CNV made two CNV calls (12.3 Mb and 8.5 Mb) across the 20.8-Mb duplicated region (Figure S4).
Figure 3.
JAX-CNV detected a number of unique CNV calls compared to the array-based method
A. Bar chart summarizing the unique JAX-CNV calls (in red) and CNV calls overlapping with the truth set (in green) as a function of CNV sizes. B. Concordance of CNV calls per sample.
JAX-CNV presented a 100% sensitivity and a 4.17% FDR for CNV calls larger than 50 kb
As described above, JAX-CNV detected 747 more CNVs (across 31 individuals), which resulted in an approximately seven-fold increase of CNV calls compared to the truth set (Table S5). To assess the accuracy of those additional calls, we randomly selected two samples, GM05876 and GM09209, for experimental validation. Compared to the truth set, there were 8 concordant CNV calls and 16 novel CNV calls in the two samples. Among the 16 novel CNV calls, 14 were validated by ddPCR (Table S6). For the two invalidated CNV calls, one (1450.3 kb loss at 21p11) was inconclusive due to an unclear separation of positive and negative droplet clusters in the ddPCR assay; the other one (duplication at 16p11; encompassing a segmental duplication and a simple repeat-rich region) was a false positive from JAX-CNV, which was detected in 28 of the 31 samples by JAX-CNV due to signal noise in this region. However, we did not simply filter this region because JAX-GM CMA identified a duplication in this region for GM16580. As a result, there were 15 CNVs (out of 24, larger than 50 kb in size) that were missed by the clinically validated CMAs and one false alspositive CNV call among the 24 randomly chosen JAX-CNV-based CNV calls. Together, these results represent a 100% sensitivity for CNV calls larger than 50 kb with a FDR of 4.17% (1/24) for JAX-CNV calls.
JAX-CNV maintained a high sensitivity at 10× WGS coverage
Although the cost for NGS has dropped rapidly, its price still remains a big concern when WGS is considered as a first-tier assay in clinical diagnostics. To address this issue and assess the ability of JAX-CNV to accurately detect CNVs on low-coverage WGS datasets, we down-sampled the WGS data and assessed CNV calling sensitivity of JAX-CNV. These 31 testing samples were originally sequenced at coverages ranging from 30× to 48× (Table S4). The simulation of different coverages was performed by SAMBAMBA [35] on the aligned BAM files. The test series consisted of down-sampling to 30×, 20×, 15×, and 10× from the original 30×-to-48× coverage. JAX-CNV was then applied on WGS data at different coverages. The assessments on WGS data at 9× and 8× coverages were also performed. At 9× coverage, JAX-CNV failed to identify one deletion and 20 duplications (loss of sensitivity, 42%), and at 8× coverage, JAX-CNV failed to identify one deletion and 24 duplications (loss of sensitivity, 50%). Thus, 10× WGS coverage is the lowest coverage that JAX-CNV could maintain the sensitivity.
Among the 112 CNVs of the truth set, 50 were larger than 300 kb (CAP standard cutoff size). Even when the coverage was reduced to 10×, JAX-CNV remained 100% sensitive for detecting CNVs larger than 300 kb (Figure 4). At 15× coverage, JAX-CNV failed to identify a 79.9-kb duplication at the chromosome region 17q21.31 of GM05876 (Figure S5). At 10× coverage, JAX-CNV additionally failed to identify a 55-kb deletion at the chromosome region 11p11.12 of GM09687 and five duplications ranging from 52 kb to 204 kb in size (Figure 4, Figure S6). In summary, for deletions larger than 50 kb, the sensitivities were 100% at 30×, 20×, and 15× coverages and 98.5% at 10× coverage, while for duplications larger than 50 kb, the sensitivities are 100% at 30× and 20× coverages, 97.9% at 15× coverage, and 89.4% at 10× coverage.
Figure 4.
JAX-CNV maintained a high sensitivity for CNV detection at 10× WGS coverage
The top two panels are the truth set using 300 kb as the cutoff. Red-colored bars indicate deletions, while blue-colored bars indicate duplications.
JAX-CNV matches clinical needs better than other CNV calling algorithms
We compared JAX-CNV to other CNV calling algorithms, Manta [15], Lumpy [16], Delly [17], CNVnator [19], and cn.MOPS [18], using the same WGS data. The scripts of performing those algorithms can be found in File S1. We also tested combined methods (pipelines) for detecting CNVs, such as FusorSV [36] and MetaSV [37]; however, the sensitivity assessments of these two pipelines failed, since both of them do not fully support GRCh38.
For the 112 CNVs of the truth set, Manta, Lumpy, Delly, CNVnator, and cn.MOPS identified 66, 61, 77, 100, and 20 of these CNVs, yielding sensitivities of 59%, 54%, 69%, 89%, and 18%, respectively (Figure S7). Some of the algorithms also incorrectly identified the CNV types (e.g., a deletion instead of a duplication). In general, read-depth-based algorithms, such as JAX-CNV and CNVnator, have greater sensitivities in detecting genomic imbalances. Other algorithms that primarily use paired-end, split-read, or combinations of these strategies clearly show lower sensitivities in identifying large chromosomal imbalances. Moreover, Manta, Lumpy, and Delly identified tens of thousands of CNVs larger than 50 kb in size (39781, 35886, and 350183, respectivley) (Figure S7). Such big amount of identified CNVs poses a huge challenge for efficiently conducting clinical diagnosis and decision-making with these CNV algorithms.
With respect to speed, JAX-CNV took less than 1 h to finish CNV detection for a given sample (ranging from 29 min to 47 min for the 31 testing samples) by using a single thread. Comparatively, Manta, Lumpy, Delly, CNVnator, and cn.MOPS took 6 h, 6.1 h, 8.2 h, 1.1 h, and 1 h, respectively, to finish CNV calling for a single sample. Moreover, analyses with JAX-CNV were completed using 4.5 GB of memory.
Discussion
Since WGS-based assays are able to detect all types of genetic variations [e.g., SNVs, insertions and deletions (InDels), and structural variants (SVs)], it has the potential to eventually supplant karyotyping, CMAs, and exome sequencing for disease diagnosis. Despite the growing applications of WGS-based assays for detecting SNVs and InDels in clinical settings [38], [39], [40], [41], it still remains challenging to reliably detect CNVs for clinical diagnostics. While a number of CNV detection pipelines have been developed for research studies and further employed for clinical applications [26], [34], [42], none of them has been widely used for clinical diagnosis.
For research purpose pipelines, sacrificing specificity for sensitivity is beneficial to avoid missing potential CNVs. However, reporting hundreds or even thousands of chromosomal aberrations is impractical for clinical diagnosis both due to time constraints and the complexity of evaluating a large number of CNVs with unknown significance. Without careful clinical considerations, it is impossible for those tools to meet the sensitivity, specificity, and turnaround time requirements for disease diagnosis. Thus, there is still a need to develop novel or optimize existing bioinformatics tools and/or pipelines to improve the accuracy and turnaround time of the WGS-based assays for clinical applications. Hence, we develop a WGS-based CNV caller, JAX-CNV, showing a 100% sensitivity for detecting CNVs larger than 50 kb.
Currently, the HMM in JAX-CNV has five statuses of CNVs: CN = 0 (loss), CN = 1 (loss), CN = 2 (normal), CN = 3 (gain), and CN > 3 (gain). Thus, JAX-CNV can only indicate that a CNV status is >3 if it is. To indicate the exact copy number that is >3, JAX-CNV needs to check BAMs/CRAMs for read depth of the detected CNV region and detect the exact copy number in this region.
Since pathogenic chromosomal abnormalities are not limited to CNVs. Other SVs, including translocations and inversions, can also cause diseases [32]. We are currently developing new modules based on the established pipeline to accurately identify translocations and inversions. Detecting translocations and inversions is more difficult than detecting CNVs, and a major reliance on the read depth signal as employed in the current CNV caller will not be effective. Thus, we are considering the inclusion of paired-end alignment distance and orientation as inputs for identifying translocations and inversions in the next version of this pipeline. Breakpoints of inversions may be associated with deletions and thus further increase the detection difficulty. Nevertheless, previous studies have successfully detected translocations and inversions in WGS data [43]. Thus, we believe that with a careful design that reflects advanced knowledge of SVs, our future pipeline will become a comprehensive SV caller for clinical applications.
JAX-CNV is a newly developed WGS-based CNV algorithm for detecting deletions and duplications that are larger than 50 kb. The results obtained from the 31 Corriell samples demonstrated a 100% concordance between JAX-CNV calls and the calls registered by Corriell and detected by JAX-GM CMA. In addition to the high sensitivity and specificity, JAX-CNV is easy-to-use, stable, robust, and fast for detecting CNVs in WGS data. JAX-CNV requires 4.5 GB of memory and finishes CNV detection for a single sample in less than 1 h. JAX-CNV meets the sensitivity, specificity, reproducibility, and speed requirements necessary for clinical applications, and demonstrates the potential to supplant CMA-based methods as the first-tier diagnostic assay.
Code availability
JAX-CNV is available at https://github.com/TheJacksonLaboratory/JAX-CNV.
CRediT author statement
Wan-Ping Lee: Conceptualization, Methodology, Software, Formal analysis, Investigation, Data curation, Writing - original draft, Writing - review & editing, Visualization. Qihui Zhu: Validation, Data curation, Writing - original draft, Writing - review & editing. Xiaofei Yang: Methodology, Software. Silvia Liu: Methodology, Visualization. Eliza Cerveira: Validation, Data curation, Writing - original draft. Mallory Ryan: Validation, Data curation. Adam Mil-Homens: Validation, Data curation. Lauren Bellfy: Validation, Data curation. Kai Ye: Resources, Supervision. Charles Lee: Conceptualization, Methodology, Resources, Writing - original draft, Writing - review & editing, Supervision, Project administration, Funding acquisition. Chengsheng Zhang: Conceptualization, Methodology, Validation, Resources, Writing - original draft, Writing - review & editing, Supervision, Project administration. All authors have read and approved the final manuscript.
Competing interests
The authors declare that they have no competing interests.
Acknowledgments
We would like to thank Drs. Zirui Dong and Yukyung Jun for their work on Figures 2 and 4. We would also like to thank Drs. Jee Young Kwon and Won Yeong Kang for their work on the manuscript revision. This study is supported in part by the operational funds from The First Affiliated Hospital of Xi’an Jiaotong University, China. WL is supported by the National Institutes of Health, USA (Grant Nos. U24AG041689 and U54AG052427). XY and KY are supported by the National Natural Science Foundation of China (Grant Nos. 61702406 and 31671372), the National Science and Technology Major Project of China (Grant No. 2018ZX10302205), the National Key R&D Program of China (Grant Nos. 2018YFC0910400 and 2017YFC0907500), and the General Financial Grant from the China Postdoctoral Science Foundation (Grant No. 2017M623178). CL was a distinguished Ewha Womans University Professor supported in part by the Ewha Womans University Research, South Korea (Grant No. 2018-2019). CZ was supported in part by the Connecticut Bio-Innovative Fund, USA.
Handled by Xiang Chen
Footnotes
Peer review under responsibility of Beijing Institute of Genomics, Chinese Academy of Sciences / China National Center for Bioinformation and Genetics Society of China.
Supplementary data to this article can be found online at https://doi.org/10.1016/j.gpb.2021.06.003.
Contributor Information
Wan-Ping Lee, Email: wan-ping.lee@pennmedicine.upenn.edu.
Chengsheng Zhang, Email: cszhang99@xjtu.edu.cn.
Supplementary material
The following are the Supplementary data to this article:
Supplementary methods
Profiles of 112 CNVs in the truth set presented by Read Depth, log2(k-mer count) and Ratio of low-quality alignment, respectively Each CNV consists of three panels that are, from the top to bottom, (1) Read Depth colored gray, (2) log2(k-mer_count) colored red, and (3) Ratio of low-quality alignment colored blue.
Common and unique CNVs in the 31 testing samples detected by three different methods Each sub figure shows reported CNVs, from the inner circle to the outer circle, are from (1) Corriell Institute, (2) JAX-GM CMA, and (3) JAX-CNV on WGS. Deletions and duplications are colored red and blue, respectively.
JAX-CNV detected CNVs that were less than 50% reciprocal overlap with the ones detected by aCGH Three deletions and three duplications that did not meet the benchmark of 50% reciprocal overlap with the JAX-CNV calls, but they were still located in the same regions with either smaller and larger size ones.
JAX-CNV detected a huge duplication with an arm-level size (20.8 Mb) A 20.8 Mb duplication at the p arm of the chromosome 12 in the sample GM03997 was detected by JAX-CNV with two calls to cover it entirely.
A false negative result was observed with JAX-CNV at 15x WGS coverage One duplication of 79.919 kb located in q21.31 region was detected by aCGH and JAX-CNV at 30x, 20x and 10x WGS coverages, but was missed by JAX-CNVat 15x WGS coverage data, suggesting a false negative result.
Decreased sensitivity of JAX-CNV at 10x WGS coverage JAX-CNV was unable to detect one deletion and six duplications at 10x WGS coverage data.
The sensitivity and specificity of JAX-CNV are better than other calling algorithms A. The recalled CNVs in the truth set of each algorithm. B. Numbers of total calls reported by each algorithm.
Primers used for the ddPCR validation
CNV calls reported from the Corriell Institute
CNV calls reported by JAX-GM CMA
Sequence coverage
CNV calls reported by JAX-CNV
ddPCR validation result
References
- 1.Bailey J.A., Gu Z., Clark R.A., Reinert K., Samonte R.V., Schwartz S., et al. Recent segmental duplications in the human genome. Science. 2002;297:1003–1007. doi: 10.1126/science.1072047. [DOI] [PubMed] [Google Scholar]
- 2.Lee C., Scherer S.W. The clinical context of copy number variation in the human genome. Expert Rev Mol Med. 2010;12:e8. doi: 10.1017/S1462399410001390. [DOI] [PubMed] [Google Scholar]
- 3.McCarroll S.A., Altshuler D.M. Copy-number variation and association studies of human disease. Nat Genet. 2007;39:S37–S42. doi: 10.1038/ng2080. [DOI] [PubMed] [Google Scholar]
- 4.Inoue K., Lupski J.R. Molecular mechanisms for genomic disorders. Annu Rev Genomics Hum Genet. 2002;3:199–242. doi: 10.1146/annurev.genom.3.032802.120023. [DOI] [PubMed] [Google Scholar]
- 5.Chen Y., Liu Y.J., Pei Y.F., Yang T.L., Deng F.Y., Liu X.G., et al. Copy number variations at the Prader-Willi syndrome region on chromosome 15 and associations with obesity in whites. Obesity. 2011;19:1229–1234. doi: 10.1038/oby.2010.323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Clayton-Smith J., Webb T., Cheng X.J., Pembrey M.E., Malcolm S. Duplication of chromosome 15 in the region 15q11-13 in a patient with developmental delay and ataxia with similarities to Angelman syndrome. J Med Genet. 1993;30:529–531. doi: 10.1136/jmg.30.6.529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Potocki L., Chen K.S., Park S.S., Osterholm D.E., Withers M.A., Kimonis V., et al. Molecular mechanism for duplication 17p11.2- the homologous recombination reciprocal of the Smith-Magenis microdeletion. Nat Genet. 2000;24:84–87. doi: 10.1038/71743. [DOI] [PubMed] [Google Scholar]
- 8.Scambler P.J. The 22q11 deletion syndromes. Hum Mol Genet. 2000;9:2421–2426. doi: 10.1093/hmg/9.16.2421. [DOI] [PubMed] [Google Scholar]
- 9.Schubert R., Viersbach R., Eggermann T., Hansmann M., Schwanitz G. Report of two new cases of Pallister-Killian syndrome confirmed by FISH: tissue-specific mosaicism and loss of i(12p) by in vitro selection. Am J Med Genet. 1997;72:106–110. doi: 10.1002/(sici)1096-8628(19971003)72:1<106::aid-ajmg21>3.0.co;2-u. [DOI] [PubMed] [Google Scholar]
- 10.Miller D.T., Adam M.P., Aradhya S., Biesecker L.G., Brothman A.R., Carter N.P., et al. Consensus statement: chromosomal microarray is a First-Tier clinical diagnostic test for individuals with developmental disabilities or congenital anomalies. Am J Hum Genet. 2010;86:749–764. doi: 10.1016/j.ajhg.2010.04.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Manning M., Hudgins L. Array-based technology and recommendations for utilization in medical genetics practice for detection of chromosomal abnormalities. Genet Med. 2010;12:742–745. doi: 10.1097/GIM.0b013e3181f8baad. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Wang H., Naghavi M., Allen C., Barber R.M., Bhutta Z.A., Carter A., et al. Global, regional, and national life expectancy, all-cause mortality, and cause-specific mortality for 249 causes of death, 1980–2015: a systematic analysis for the global burden of disease study 2015. Lancet. 2016;388:1459–1544. doi: 10.1016/S0140-6736(16)31012-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Stavropoulos D.J., Merico D., Jobling R., Bowdin S., Monfared N., Thiruvahindrapuram B., et al. Whole genome sequencing expands diagnostic utility and improves clinical management in pediatric medicine. NPJ Genomic Med. 2016;1:15012. doi: 10.1038/npjgenmed.2015.12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Schwarze K., Buchanan J., Taylor J.C., Wordsworth S. Are whole-exome and whole-genome sequencing approaches cost-effective? A systematic review of the literature. Genet Med. 2018;20:1122–1130. doi: 10.1038/gim.2017.247. [DOI] [PubMed] [Google Scholar]
- 15.Chen X., Schulz-Trieglaff O., Shaw R., Barnes B., Schlesinger F., Källberg M., et al. Manta: rapid detection of structural variants and indels for germline and cancer sequencing applications. Bioinformatics. 2016;32:1220–1222. doi: 10.1093/bioinformatics/btv710. [DOI] [PubMed] [Google Scholar]
- 16.Layer R.M., Chiang C., Quinlan A.R., Hall I.M. LUMPY: a probabilistic framework for structural variant discovery. Genome Biol. 2014;15:R84. doi: 10.1186/gb-2014-15-6-r84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Rausch T., Zichner T., Schlattl A., Stütz A.M., Benes V., Korbel J.O. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics. 2012;28:i333–i339. doi: 10.1093/bioinformatics/bts378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Klambauer G, Schwarzbauer K, Mayr A, Clevert DA, Mitterecker A, Bodenhofer U, et al. cn.MOPS: mixture of Poissons for discovering copy number variations in next-generation sequencing data with a low false discovery rate. Nucleic Acids Res 2012;40:e69. [DOI] [PMC free article] [PubMed]
- 19.Abyzov A., Urban A.E., Snyder M., Gerstein M. CNVnator: an approach to discover, genotype, and characterize typical and atypical CNVs from family and population genome sequencing. Genome Res. 2011;21:974–984. doi: 10.1101/gr.114876.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Yuan X., Yu J., Xi J., Yang L., Shang J., Li Z., et al. CNV_IFTV: an isolation forest and total variation-based detection of CNVs from short-read sequencing data. IEEE/ACM Trans Comput Biol Bioinform. 2021;18:539–549. doi: 10.1109/TCBB.2019.2920889. [DOI] [PubMed] [Google Scholar]
- 21.Dharanipragada P., Vogeti S., Parekh N. iCopyDAV: integrated platform for copy number variations-detection, annotation and visualization. PLoS One. 2018;13:e0195334. doi: 10.1371/journal.pone.0195334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Zhou B., Ho S.S., Zhang X., Pattni R., Haraksingh R.R., Urban A.E. Whole-genome sequencing analysis of CNV using low-coverage and paired-end strategies is efficient and outperforms array-based CNV analysis. J Med Genet. 2018;55:735–743. doi: 10.1136/jmedgenet-2018-105272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Noll A.C., Miller N.A., Smith L.D., Yoo B., Fiedler S., Cooley L.D., et al. Clinical detection of deletion structural variants in whole-genome sequences. NPJ Genom Med. 2016;1:16026. doi: 10.1038/npjgenmed.2016.26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Fan X, Abbott TE, Larson D, Chen K. BreakDancer: identification of genomic structural variation from paired-end read mapping. Curr Protoc Bioinformatics 2014;45:15.6.1–11. [DOI] [PMC free article] [PubMed]
- 25.Handsaker R.E., Van Doren V., Berman J.R., Genovese G., Kashin S., Boettger L.M., et al. Large multiallelic copy number variations in humans. Nat Genet. 2015;47:296–303. doi: 10.1038/ng.3200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Trost B., Walker S., Wang Z., Thiruvahindrapuram B., MacDonald J.R., Sung W.W.L., et al. A comprehensive workflow for read depth-based identification of copy-number variation from whole-genome sequence data. Am J Hum Genet. 2018;102:142–155. doi: 10.1016/j.ajhg.2017.12.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zhu M., Need A.C., Han Y., Ge D., Maia J.M., Zhu Q., et al. Using ERDS to infer copy-number variants in high-coverage genomes. Am J Hum Genet. 2012;91:408–421. doi: 10.1016/j.ajhg.2012.07.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Marçais G., Kingsford C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics. 2011;27:764–770. doi: 10.1093/bioinformatics/btr011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Yu Z., Li A., Wang M. CLImAT-HET: detecting subclonal copy number alterations and loss of heterozygosity in heterogeneous tumor samples from whole-genome sequencing data. BMC Med Genomics. 2017;10:1–11. doi: 10.1186/s12920-017-0255-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Mccallum K.J., Wang J.P. Quantifying copy number variations using a hidden Markov model with inhomogeneous emission distributions. Biostatistics. 2013;14:600–611. doi: 10.1093/biostatistics/kxt003. [DOI] [PubMed] [Google Scholar]
- 31.Ester M., Kriegel H.P., Sander J., Xu X. Proceedings of the Second International Conference on Knowledge Discovery and Data Mining; 1996. A density-based algorithm for discovering clusters in large spatial databases with noise; pp. 226–231. [Google Scholar]
- 32.Brothman A.R., Dolan M.M., Goodman B.K., Park J.P., Persons D.L., Saxe D.F., et al. College of American Pathologists/American College of Medical Genetics proficiency testing for constitutional cytogenomic microarray analysis. Genet Med. 2011;13:765–769. doi: 10.1097/GIM.0b013e31821d3165. [DOI] [PubMed] [Google Scholar]
- 33.Untergasser A., Nijveen H., Rao X., Bisseling T., Geurts R., Leunissen J.A.M. Primer3Plus, an enhanced web interface to Primer3. Nucleic Acids Res. 2007;35:W71–W74. doi: 10.1093/nar/gkm306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Dong Z., Zhang J., Hu P., Chen H., Xu J., Tian Q., et al. Low-pass whole-genome sequencing in clinical cytogenetics: a validated approach. Genet Med. 2016;18:940–948. doi: 10.1038/gim.2015.199. [DOI] [PubMed] [Google Scholar]
- 35.Tarasov A., Vilella A.J., Cuppen E., Nijman I.J., Prins P. Sambamba: fast processing of NGS alignment formats. Bioinformatics. 2015;31:2032–2034. doi: 10.1093/bioinformatics/btv098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Becker T., Lee W.P., Leone J., Zhu Q., Zhang C., Liu S., et al. FusorSV: an algorithm for optimally combining data from multiple structural variation detection methods. Genome Biol. 2018;19:38. doi: 10.1186/s13059-018-1404-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Mohiyuddin M., Mu J.C., Li J., Bani Asadi N., Gerstein M.B., Abyzov A., et al. MetaSV: an accurate and integrative structural-variant caller for next generation sequencing. Bioinformatics. 2015;31:2741–2744. doi: 10.1093/bioinformatics/btv204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Balloux F., Brønstad Brynildsrud O., van Dorp L., Shaw L.P., Chen H., Harris K.A., et al. From theory to practice: translating whole-genome sequencing (WGS) into the clinic. Trends Microbiol. 2018;26:1035–1048. doi: 10.1016/j.tim.2018.08.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Mousallem T, Urban TJ, McSweeney KM, Kleinstein SE, Zhu M, Adeli M, et al. Clinical application of whole-genome sequencing in patients with primary immunodeficiency. J Allergy Clin Immunol 2015;136:476–9.e6. [DOI] [PMC free article] [PubMed]
- 40.Arnold C., Edwards K., Desai M., Platt S., Green J., Conway D. Setup, validation, and quality control of a centralized whole-genome-sequencing laboratory: Lessons learned. J Clin Microbiol. 2018;56:e00261–e00318. doi: 10.1128/JCM.00261-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Colman R.E., Mace A., Seifert M., Hetzel J., Mshaiel H., Suresh A., et al. Whole-genome and targeted sequencing of drug-resistant mycobacterium tuberculosis on the iSeq100 and MiSeq: a performance, ease-of-use, and cost evaluation. PLoS Med. 2019;16:e1002823. doi: 10.1371/journal.pmed.1002794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Eisfeldt J., Nilsson D., Andersson-Assarsson J.C., Lindstrand A. AMYCNE: confident copy number assessment using whole genome sequencing data. PLoS One. 2018;13:e0189710. doi: 10.1371/journal.pone.0189710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Dong Z., Wang H., Chen H., Jiang H., Yuan J., Yang Z., et al. Identification of balanced chromosomal rearrangements previously unknown among participants in the 1000 genomes project: implications for interpretation of structural variation in genomes and the future of clinical cytogenetics. Genet Med. 2018;20:697–707. doi: 10.1038/gim.2017.170. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary methods
Profiles of 112 CNVs in the truth set presented by Read Depth, log2(k-mer count) and Ratio of low-quality alignment, respectively Each CNV consists of three panels that are, from the top to bottom, (1) Read Depth colored gray, (2) log2(k-mer_count) colored red, and (3) Ratio of low-quality alignment colored blue.
Common and unique CNVs in the 31 testing samples detected by three different methods Each sub figure shows reported CNVs, from the inner circle to the outer circle, are from (1) Corriell Institute, (2) JAX-GM CMA, and (3) JAX-CNV on WGS. Deletions and duplications are colored red and blue, respectively.
JAX-CNV detected CNVs that were less than 50% reciprocal overlap with the ones detected by aCGH Three deletions and three duplications that did not meet the benchmark of 50% reciprocal overlap with the JAX-CNV calls, but they were still located in the same regions with either smaller and larger size ones.
JAX-CNV detected a huge duplication with an arm-level size (20.8 Mb) A 20.8 Mb duplication at the p arm of the chromosome 12 in the sample GM03997 was detected by JAX-CNV with two calls to cover it entirely.
A false negative result was observed with JAX-CNV at 15x WGS coverage One duplication of 79.919 kb located in q21.31 region was detected by aCGH and JAX-CNV at 30x, 20x and 10x WGS coverages, but was missed by JAX-CNVat 15x WGS coverage data, suggesting a false negative result.
Decreased sensitivity of JAX-CNV at 10x WGS coverage JAX-CNV was unable to detect one deletion and six duplications at 10x WGS coverage data.
The sensitivity and specificity of JAX-CNV are better than other calling algorithms A. The recalled CNVs in the truth set of each algorithm. B. Numbers of total calls reported by each algorithm.
Primers used for the ddPCR validation
CNV calls reported from the Corriell Institute
CNV calls reported by JAX-GM CMA
Sequence coverage
CNV calls reported by JAX-CNV
ddPCR validation result