Abstract
Next generation amplicon sequencing has created a plethora of data from human microbiomes. The accessibility to this scientific data and its corresponding metadata is important for its reuse, to allow for new discoveries, verification of published results, and serving as path for reproducibility. Dietary fiber consumption has been associated with a variety of health benefits that are thought to be mediated by gut microbiota. To enable direct comparisons of the response of the gut microbiome to fiber, we obtained 16S rRNA sequencing data and its corresponding metadata from 11 fiber intervention studies for a total of 2,368 samples. We provide curated and pre-processed genetic data and common metadata for comparison across the different studies.
Subject terms: Bacteria, Microbial communities
Background & Summary
Fiber is naturally present in plants, fungi, animals, bacteria, and can also be synthetically made1,2. Dietary fibers are carbohydrates that resist digestion by the small intestine and have physiological health benefits to humans3,4. High fiber diets show a risk reduction for or amelioration of various illnesses such as constipation, obesity, diabetes, high cholesterol, heart disease, allergies, among others5–9. Furthermore, they are associated with improving mineral absorption, insulin responses, gut barrier permeability, immune system defense, production of beneficial metabolites, and inducing changes in the gut microbiome1,10. Fiber can modify the gut microbiome by affecting host secretions and transit stool time. It also serves as fermentative substrate for specific microbes and in turn, alters microbial activity more broadly (e.g., through cross-feeding and competition)11.
To understand the influence of dietary fiber on the gut microbiota, researchers have performed dietary fiber interventions among both healthy and unhealthy individuals12. These studies usually take a fecal sample from a person before and after their dietary change to assess shifts in the composition of the gut microbiome. Currently, the most common approach to assess microbial taxonomic composition is amplicon sequencing of a portion of the universal bacterial 16S ribosomal RNA (rRNA) marker gene13 because of the relatively low cost of next generation sequencing and the variety of tools available for bioinformatic processing. However, it still is challenging to access and harmonize such data to compare across studies, especially when its corresponding metadata is missing or hard to decipher14.
Motivated by the investigation of fiber-induced shifts in microbiota and the potential for re-analyzing sequencing data, we screened more than 1,500 abstracts and obtained data from 11 fiber intervention studies performed in healthy human subjects, for a total of 2,368 samples from 488 subjects. The purpose of publishing this data descriptor is to provide a detailed description of these valuable datasets, allow others to re-use the data that was carefully curated, and to promote data accessibility. Here, we present 1) the next generation 16S rRNA amplicon sequencing data which have been pre-processed and checked for quality scores, 2) its corresponding metadata which has been harmonized across studies, and 3) the operational taxonomic unit (OTU) tables that contain the number of reads per sample for each taxonomic unit. The sequencing data was primarily produced by Illumina platforms, but also includes 454 and Ion Torrent technologies. All metadata was curated to include similar columns across studies that are clearly defined in the metadata dictionary. The availability of scientific data and its corresponding metadata in comparable and reusable forms will allow researchers to re-analyze and synthesize these data in new ways to better understand the role of fiber in gut health.
Methods
Data collection and harmonization
We conducted a keyword search of published literature through the PubMed search engine (keywords: dietary, fiber, and microbiome) under the Best Match algorithm recommended by PubMed on May 9th, 2020. The search yielded 977 abstract hits from 2010 to 2020 (https://pubmed.ncbi.nlm.nih.gov/). We also searched through all the records available in the database of open-source microbial management site Qiita15 (https://qiita.ucsd.edu) on April 7th, 2020 and found 528 microbiome studies including human and animal studies. From both sources, each abstract was carefully read to select studies with fiber interventions in healthy humans that included 16S rRNA amplicon sequencing data from fecal microbial communities (n = 34). We excluded studies in animals and unhealthy humans (Fig. 1). Corresponding authors and first authors were contacted up to 4 times requesting their sequencing data and metadata when not publicly available. We were able to obtain 16S rRNA amplicon sequencing data and their corresponding metadata from 11 studies (Table 1). Data was shared to us via accession number16–23 or, if not publicly available, via virtual box. For the studies that did not make their datasets available at the time of publication (Dahl_2016_V1V2, Hooda_2012_V4V6, and Morales_2016_V3V4), we received consent to deposit their data under the BioProject ID: PRJNA891951 to the NCBI Sequence Read Archive24. For these studies, we recommend downloading the raw data through the SRA Run Selector Tool that allows users to see the Library Name. Each Library Name includes the study name followed by an underscore and the Sample ID. These Sample IDs are described in the metadata files created for this manuscript (see Data Records and Harmonization of datasets for more information). All studies included in this data repository complied with their relevant ethical regulations and have consent from their human participants to collect and share the data. For more information regarding guidelines for study procedure and trial registration numbers we refer our readers to the individual studies referenced in Table 1 and Table 2. The naming scheme for each of the studies included in this data collection is the following: Last name of the first author in the publication, followed by the year the study was published, and ending with the amplified region of the 16S rRNA bacterial gene (e.g., Liu_2017_V4).
Fig. 1.
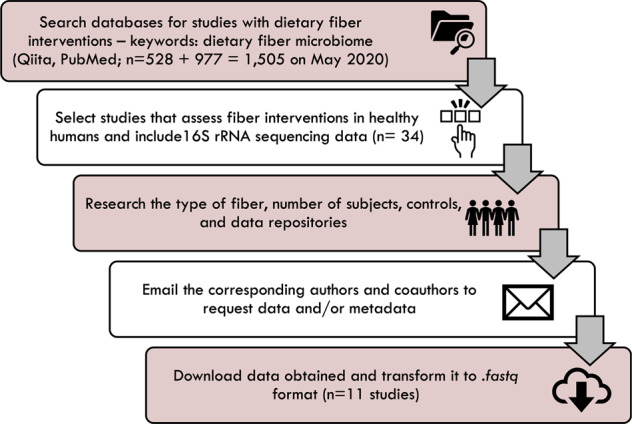
Data collection workflow.
Table 1.
Data collected and available for eleven fiber intervention studies.
| Study Name | Repository for raw data | Accession number for raw data | Sequencing platform used | Single- or paired-end data | Processed data in this manuscript deposited to Figshare repository include |
|---|---|---|---|---|---|
| Baxter_2019_V435 | NCBI Sequence Read Archive | SRP128128 | Illumina MiSeq | paired | cleaned reads, metadata, OTU tables |
| Dahl_2016_V1V236 | NCBI Sequence Read Archive | SRP403421 | Illumina MiSeq | paired | cleaned reads, metadata, OTU tables |
| Deehan_2020_V5V62 | NCBI Sequence Read Archive | SRP219296 | Illumina MiSeq | paired | cleaned reads, metadata, OTU tables |
| Healey_2018_V3V437 | NCBI Sequence Read Archive | SRP120250 | Illumina MiSeq | paired | cleaned reads, metadata, OTU tables |
| Hooda_2012_V4V638 | NCBI Sequence Read Archive | SRP403421 | 454/Roche pyrosequencing | single | cleaned reads, metadata, OTU tables |
| Kovatcheva_2015_V1V239 | NCBI Sequence Read Archive | SRP062889 | 454/Roche pyrosequencing | single | cleaned reads, metadata, OTU tables |
| Liu_2017_V440 | European Nucleotide Archive | PRJEB15149 | Ion Torrent | single | cleaned reads, metadata, OTU tables |
| Morales_2016_V3V441 | NCBI Sequence Read Archive | SRP403421 | Illumina MiSeq | paired | cleaned reads, metadata, OTU tables |
| Rasmussen_2017_V1V342 | NCBI Sequence Read Archive | SRP106361 | 454/Roche pyrosequencing | single | cleaned reads, metadata, OTU tables |
| Tap_2015_V3V443 | European Nucleotide Archive | PRJEB2165 | 454/Roche pyrosequencing | single | cleaned reads, metadata, OTU tables |
| Venkataraman_2016_V444 | NCBI Sequence Read Archive | SRP067761 | Illumina MiSeq | paired | cleaned reads, metadata, OTU tables |
Table 2.
Summary of data collected by study.
| Study Name | Number of interventions | Fibers used in intervention + control when applicable | Amount of fiber or control given in intervention (grams) | Duration of intervention (days) | Collection timepoints | Number of subjects | Number of samples |
|---|---|---|---|---|---|---|---|
| Baxter_2019_V435 | 4 | Resistant starch from potatoes (RPS), resistant starch from maize (RMS), inulin from chicory root, and an accessible corn starch control | 20–40 | 14 | 8 | 175 | 1,205 |
| Dahl_2016_V1V236 | 3 | RS-4-A, RS-4-B, RS-4-C - Resistant potato starches (RS type 4) | 30 | 14 | 4 | 53 | 212 |
| Deehan_2020_V5V62 | 4 | Tapioca, potato, and maize- Resistant starches (RS type 4) + corn starch control | increments 10–50 | 28 | 5 | 40 | 200 |
| Healey_2018_V3V437 | 2 | 50:50 inulin to fructo-oligosaccharide and maltodextrin control | 16 | 21 | 4 | 34 | 134 |
| Hooda_2012_V4V638 | 2 | Polydextrose and soluble corn fiber control | 21 | 21 | 3 | 10 | 28 |
| Kovatcheva_2015_V1V239 | 2 | Kernel-based bread (BKB) and white-wheat-bread (WWB) | 37.6 & 9.1 | 3 | 3 | 20 | 60 |
| Liu_2017_V440 | 2 | Fructooligosaccharides (FOS) and galactooligosaccharides (GOS) | 16 | 14 | 4 | 35 | 132 |
| Morales_2016_V3V441 | 2 | Oligofructose and maltodextrin control (extra treatments of Orlistat were also given) | 16 | 7 | 2 | 41 | 82 |
| Rasmussen_2017_V1V342 | 2 | Starch-entrapped microspheres and psyllium | 9 & 12 | 84 | 2 | 41 | 82 |
| Tap_2015_V3V443 | 1 | Dietary fiber meals | 10 & 40 | 5 | 4 | 19 | 76 |
| Venkataraman_2016_V444 | 1 | Resistant starch (unmodified potato starch; RS type 2) | 48 | 17 | 8 | 20 | 157 |
Shows the studies included in this data descriptor and their pertinent information such as fibers used, duration of intervention, number of subjects, etc.
We provide Table 2 with a summary of each of the studies which includes: number of interventions per study, fibers used and their amounts, length of interventions, number of colletion timepoints, subjects and total samples. Because the metadata available was heterogeneous across studies, we performed harmonization across the datasets, so that common variables across studies could be easily identified. The metadata dictionary (Table 3) contains the definition for the data collected across studies.
Table 3.
Metadata dictionary. Explains each column in the metadata files.
| Column Name | Description |
|---|---|
| sampleid | The name of the fastq file that corresponds to one fecal sample |
| study | Shows the last name of the first author of the study where the data came from |
| sample_id_2 | Original sampleID depicted in raw sequence reads |
| subject_id | The ID of the subject (person) that the sample was collected from |
| treatment | Shows whether the type of treatment administered was a dietary fiber (fiber) or a placebo (control) |
| timepoint | The time at which this sample was taken - before or after treatment |
| timepoint_numeric | Defines the time course the fecal sample was taken in chronological order (e.g., 1,2,3..) if coming from the same individual |
| timepoint_id | Description of the timepoint, including timepoint + timepoint_numeric: before versus after, with chronological number attached to it, in the case multiple samples were taken from the same individual |
| sample_name | Has the subject_id attached to timepoint_numeric |
| fiber_type | The specific type of fiber that was used in the treatment, and/or the name of the control compound administered |
| fiber_amount | Grams per day of the compound in the treatment administered, if known (e.g., 20 g/d of inulin) |
| time_days | The days that had passed since the intervention started, if known. Note that weeks were counted as 7 days, for instance if the intervention lasted 12 weeks, we converted that to 84 days. |
| number | Order in which samples were originally arranged by the metadata given by authors, should equal number of fecal samples collected |
| gender | The gender of the subject as reported by original authors (available only for the Healey study) |
| age | The age in years of the subject reported by original authors (available only for the Healey study) |
| sample-name-original | The name given to the sample in the original study |
To provide as much information on the dietary fiber interventions as possible, we investigated the specific fibers that were used in each study. Table 4 shows all the dietary fibers that were used in the interventions and their manufacturer or recipe (when available) including controls.
Table 4.
Fibers and placebos given in the interventions.
| Fiber type | Description/manufacturer | Study |
|---|---|---|
| Resistant starch from potatoes (RPS) | Bob’s Red Mill, Milwaukee, OR | Baxter_2019_V435 |
| Inulin from chicory root | Swanson Health Products, Fargo, ND | Baxter_2019_V435 |
| Hi-Maize 260 resistant corn starch (RMS) | Manufactured by Ingredion Inc., Westchester, IL, and distributed by myworldhut.com | Baxter_2019_V435 |
| Amylase-accessible corn starch (placebo) | Amioca powder; Skidmore Sales and Distribution, West Chester, OH | Baxter_2019_V435 |
| Resistant potato starch RS4-A | PenFibe® RO – 170; phosphorylated, soluble fibre with high viscosity - Penford Food Ingredients Inc., Denver, CO, USA | Dahl_2016_V1V236 |
| Resistant potato starch RS4-B | PenFibe® RO – 177; hydrolysed, phosphorylated, soluble fibre with low viscosity - Penford Food Ingredients Inc., Denver, CO, USA | Dahl_2016_V1V236 |
| Resistant potato starch RS4-C | PenFibe® RS; insoluble fibre with low viscosity - Penford Food Ingredients Inc., Denver, CO, USA | Dahl_2016_V1V236 |
| AMIOCA™ Powder TF (Placebo) | Ingredion Inc, Bridgewater, NJ 08807, USA | Deehan_2020_V5V62 |
| VERSAFIBE™ 2470 (Maize RS4) | Ingredion Inc, Bridgewater, NJ 08807, USA | Deehan_2020_V5V62 |
| VERSAFIBE™ 1490 (Potato RS4) | Ingredion Inc, Bridgewater, NJ 08807, USA | Deehan_2020_V5V62 |
| VERSAFIBE™ 3490 (Tapioca RS4) | Ingredion Inc, Bridgewater, NJ 08807, USA | Deehan_2020_V5V62 |
| Orafti® Synergy1–50:50 inulin to fructo-oligosaccharide mix | Beneo GmbH | Healey_2018_V3V437 |
| Glucidex® 29 Premium-digestible maltodextrin; placebo | Roquette Worldwide | Healey_2018_V3V437 |
| Polydextrose | PDX; Litesse II, Danisco | Hooda_2012_V4V638 |
| Soluble corn fiber (placebo) | SCF; PROMITOR, Tate and Lyle Ingredients | Hooda_2012_V4V638 |
| Kernel-based bread (KBB) | NA | Kovatcheva_2015_V1V239 |
| White-wheat-bread (WWB) | NA | Kovatcheva_2015_V1V239 |
| Fructooligosaccharide- FOS (QHT-Purity95%) | Source: Sucrose; Quantum Hi-Tech (China) Biological company, Guangdong, China | Liu_2017_V440 |
| Galactooligosaccharide- GOS (QHT- Purity95%) | Source: lactose; Quantum Hi-Tech (China) Biological company, Guangdong, China | Liu_2017_V440 |
| Maltodextrin (placebo) | NA | Morales_2016_V3V441 |
| Oligofructose | NA | Morales_2016_V3V441 |
| Starch-entrapped microspheres (SM) | A suspension of sodium alginate (2% w/v) and normal corn starch (9% w/v) was made in water through a special recipe | Rasmussen_2017_V1V342 |
| Psyllium | Natural Foods Inc (Toledo, OH) | Rasmussen_2017_V1V342 |
| Dietary fiber meals (different foods) | NA | Tap_2015_V3V443 |
| Raw unmodified potato starch | Bob’s Red Mill, Milwaukie, OR. This potato starch contains approximately 50% resistant starch (type 2) by weight. | Venkataraman_2016_V444 |
The description of the compound administered during the intervention as described by the original authors, when available.
Sequencing processing
Individual studies used different methods for sequencing processing and bioinformatic pipelines, and such differences can influence the diversity and composition of microorganisms detected in a sample as well as the variation observed across samples25. Thus, to compare the sequences directly across studies, we obtained the raw sequencing reads for each study and then processed them in a similar manner.
First, we assessed the quality of the 16S rRNA sequencing data using FastQC software26 (version 0.11.8). The sequencing reads were cleaned from poor quality sequences using the Fastp program27 (version 0.20.0). The cleaned sequences were imported into the QIIME2 platform28 (version 2020.11.1), and primers were removed using Cutadapt29 plugin when necessary. We then denoised the reads using DADA230 plugin, obtaining an OTU table depicting the number of reads per sample for each taxonomic unit (Fig. 2).
Fig. 2.
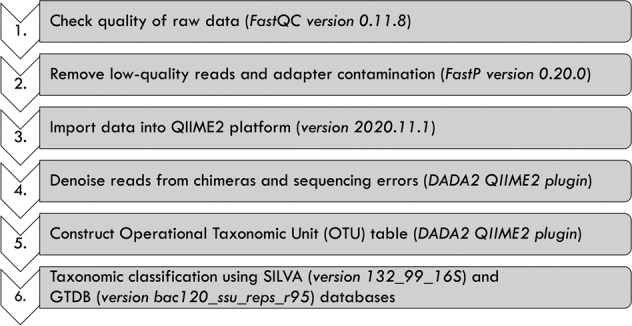
Bioinformatics pipeline for data processing.
Next, the taxonomic classification of the reads was also performed in the QIIME2 platform by training the SILVA31 (version 132_99_16S) and the Genome Taxonomy Database32 (GTDB; version bac120_ssu_reps_r95) databases to each respective study based on the primers that were originally used (Fig. 2). The SILVA database was used to remove chloroplast and mitochondrial DNA. Then, the cleaned reads were assigned to a final taxonomic group using the GTDB trained database. Reads that were not classified at least to the phylum level were removed from the analysis; sequences were classified to the finest level when possible (e.g., species and/or strain). The sequencing processing and taxonomic classification was performed with both the forward and reverse reads when paired-end data was available. We also repeated the analyses with only the forward reads, and found that both gave very similar results. We provide the OTU tables obtained with both procedures (e.g., baxter_OTU_table_paired_reads.tsv and baxter_OTU_table_forward_reads.tsv) to allow the reader to choose either option for further analysis.
Data Records
The following data have been deposited in the Figshare33 repository: 1) The compressed 16S rRNA sequencing reads (.fastq.gz) containing the amplicon data that were quality filtered as described above; 2) the metadata files per study in tab-delimited format (.txt) describing their corresponding samples serving as a reference to help identify and sort the DNA sequences by different metrics (e.g., timepoint, treatment, individual, etc.); 3) the OTU tables with taxonomic assignment per study (.tsv) presenting the number of reads per sample for each taxonomic unit. As mentioned in the Data collection section and in Table 1, the raw reads for the studies mentioned here can be found in publicly available databases16–23. For the studies that did not make their datasets available prior to this publication (Dahl_2016_V1V2, Hooda_2012_V4V6, and Morales_2016_V3V4), we received consent to deposit their data under the BioProject ID: PRJNA891951 to the NCBI Sequence Read Archive24.
Technical Validation
Data integrity
For quality assurance of the sequencing reads, we utilized the FastQC tool26 as it provides quality control statistics such as sequence length, per base quality scores, and adapter contamination34. We used the Fastp software27 to ensure data integrity: we removed low quality reads from all datasets, only keeping reads with an average quality score of 30, the average score of 25 was chosen in only two occasions (Rasmussen_2017_V1V3 and Liu_2017_V4) because read counts dropped dramatically with a higher threshold (−average_qual 30 or 25); we discarded sequences shorter than 100 bp (−length_required 100) to remove small sequences that could not complete 16S rRNA amplicon fragments. We only had to remove adapter contamination from one study (Deehan_2020_V5V6) using the detection of adapter correction tool in Fastp (−detect_adapter_for_pe). When paired-end data was available, we enabled base correction in overlapped regions of paired reads (−correction). When corrupted data, having characters that did not belong to the sequencing reads, was found (Hooda_2012_V4V6) we discarded those samples (n = 10).
Harmonization of datasets
To ensure the datasets were comparable, we converted sequencing reads from all studies into .fastq extension files (when necessary). Furthermore, we followed the same pipeline using consistent software and versions (Fig. 2) and cross-validated our results by visually inspecting the sequences after each clean-up step using Geneious prime (version 2020.2.4; https://www.geneious.com). For instance, after removing primers from reads using the Cutadapt plugin in QIIME2, we extracted the reads and imported them into Geneious to verify that sequences had been properly trimmed. Moreover, to ensure clarity and consistency of metadata across datasets, we created a metadata dictionary (Table 3) to explain the data type (categorical, numerical, text, etc.). In most cases, the metadata files available for the studies did not follow a consistent report of variables. For example, there was a big difference in how the timepoints were described (e.g., “before”/“after” vs “post”/“pre” vs numeric) and in most instances the fiber type and grams of fiber were not included. To remedy this, we carefully curated the data collected per sample across studies to have similar naming schemes.
Acknowledgements
We would like to thank all the authors of the studies mentioned here for making their data available for publication. We would also like to thank the members of the Martiny lab for their encouragement. This work was supported by a Faculty Mentor Program (FMP) fellowship, UC President’s Dissertation Year Fellowship, and Rose Hills Foundation Science & Engineering Fellowship to CIR.
Author contributions
J.B.H.M. and C.I.R. conceived the project, wrote the manuscript, and collected data. All other co-authors provided their data and helped with deciphering metadata categories. All listed authors read and approved the final manuscript.
Code availability
The parameters and step-by-step scripts used to clean up the data, remove chimeras, and assign taxonomy are available at https://github.com/cirodri1/fiber-data_records (e.g, trimming lengths, primers, databases, etc.).
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Carlson JL, Erickson JM, Lloyd BB, Slavin JL. Health effects and sources of prebiotic dietary fiber. Curr. Dev. Nutr. 2018;2:nzy005. doi: 10.1093/cdn/nzy005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Deehan EC, et al. Precision microbiome modulation with discrete dietary fiber structures directs short-chain fatty acid production. Cell Host Microbe. 2020;27:389–404.e6. doi: 10.1016/j.chom.2020.01.006. [DOI] [PubMed] [Google Scholar]
- 3.Jones JM. CODEX-aligned dietary fiber definitions help to bridge the ‘fiber gap’. Nutr. J. 2014;13:34. doi: 10.1186/1475-2891-13-34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Food and Drug Administration. Food Labeling: Revision of the Nutrition and Supplement Facts Labels. Federal Registerhttps://www.federalregister.gov/documents/2016/05/27/2016-11867/food-labeling-revision-of-the-nutrition-and-supplement-facts-labels (2016). [PubMed]
- 5.Yang J, Wang H-P, Zhou L, Xu C-F. Effect of dietary fiber on constipation: A meta analysis. World J. Gastroenterol. WJG. 2012;18:7378–7383. doi: 10.3748/wjg.v18.i48.7378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Hosseini-Esfahani F, et al. The interaction of fat mass and obesity associated gene polymorphisms and dietary fiber intake in relation to obesity phenotypes. Sci. Rep. 2017;7:18057. doi: 10.1038/s41598-017-18386-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Yao B, et al. Dietary fiber intake and risk of type 2 diabetes: a dose–response analysis of prospective studies. Eur. J. Epidemiol. 2014;29:79–88. doi: 10.1007/s10654-013-9876-x. [DOI] [PubMed] [Google Scholar]
- 8.Mirmiran P, Bahadoran Z, Khalili Moghadam S, Zadeh Vakili A, Azizi F. A prospective study of different types of dietary fiber and risk of cardiovascular cisease: tehran lipid and glucose study. Nutrients. 2016;8:686. doi: 10.3390/nu8110686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Folkerts J, et al. Effect of dietary diber and metabolites on mast cell activation and mast cell-associated diseases. Front. Immunol. 2018;9:1067. doi: 10.3389/fimmu.2018.01067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Zhou T, et al. Dietary fiber, genetic variations of gut microbiota-derived short-chain fatty acids, and bone health in UK biobank. J. Clin. Endocrinol. Metab. 2020;106:201–210. doi: 10.1210/clinem/dgaa740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Cantu-Jungles, T. M. & Hamaker, B. R. New view on dietary fiber selection for predictable shifts in gut microbiota. mBio11 (2020). [DOI] [PMC free article] [PubMed]
- 12.Sawicki CM, et al. Dietary fiber and the human gut microbiota: application of evidence mapping methodology. Nutrients. 2017;9:125. doi: 10.3390/nu9020125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Thompson LR, et al. A communal catalogue reveals Earth’s multiscale microbial diversity. Nature. 2017;551:457–463. doi: 10.1038/nature24621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Jurburg SD, Konzack M, Eisenhauer N, Heintz-Buschart A. The archives are half-empty: an assessment of the availability of microbial community sequencing data. Commun. Biol. 2020;3:1–8. doi: 10.1038/s42003-020-01204-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gonzalez A, et al. Qiita: rapid, web-enabled microbiome meta-analysis. Nat. Methods. 2018;15:796–798. doi: 10.1038/s41592-018-0141-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.2018. NCBI Sequence Read Archive. SRP128128
- 17.2020. NCBI Sequence Read Archive. SRP219296
- 18.2017. NCBI Sequence Read Archive. SRP120250
- 19.2015. NCBI Sequence Read Archive. SRP062889
- 20.ENA European Nucleotide Archivehttps://identifiers.org/ena.embl:PRJEB15149 (2017). [DOI] [PMC free article] [PubMed]
- 21.2017. NCBI Sequence Read Archive. SRP106361
- 22.ENA European Nucleotide Archivehttps://identifiers.org/ena.embl:PRJEB2165 (2013).
- 23.2016. NCBI Sequence Read Archive. SRP067761
- 24.2022. NCBI Sequence Read Archive. SRP403421
- 25.Marizzoni, M. et al. Comparison of bioinformatics pipelines and operating systems for the analyses of 16s rrna gene amplicon sequences in human fecal samples. Front. Microbiol. 11, (2020). [DOI] [PMC free article] [PubMed]
- 26.Andrews, S. Babraham Bioinformatics - FastQC a quality control tool for high throughput sequence data.https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2010).
- 27.Chen S, Zhou Y, Chen Y, Gu J. Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34:i884–i890. doi: 10.1093/bioinformatics/bty560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Bolyen E, et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 2019;37:852–857. doi: 10.1038/s41587-019-0209-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal. 2011;17:10–12. doi: 10.14806/ej.17.1.200. [DOI] [Google Scholar]
- 30.Callahan BJ, et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods. 2016;13:581–583. doi: 10.1038/nmeth.3869. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Quast C, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2013;41:D590–D596. doi: 10.1093/nar/gks1219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Parks DH, et al. A complete domain-to-species taxonomy for Bacteria and Archaea. Nat. Biotechnol. 2020;38:1079–1086. doi: 10.1038/s41587-020-0501-8. [DOI] [PubMed] [Google Scholar]
- 33.Rodriguez CI, 2023. Curated and harmonized gut microbiome 16S rRNA amplicon sequences, metadata, and OTU tables from dietary fiber intervention studies in humans. Figshare. [DOI] [PMC free article] [PubMed]
- 34.Research Technology Support Facility Team. Michigan State University. FastQC Tutorial & FAQ.https://rtsf.natsci.msu.edu/genomics/tech-notes/fastqc-tutorial-and-faq/ (2019).
- 35.Baxter NT, et al. Dynamics of human gut microbiota and short-chain fatty acids in response to dietary interventions with three fermentable fibers. mBio. 2019;10:e02566–18. doi: 10.1128/mBio.02566-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Dahl WJ, et al. Resistant potato starches (type 4 RS) exhibit varying effects on laxation with and without phylum level changes in microbiota: A randomised trial in young adults. J. Funct. Foods. 2016;23:1–11. doi: 10.1016/j.jff.2016.02.013. [DOI] [Google Scholar]
- 37.Healey G, et al. Habitual dietary fibre intake influences gut microbiota response to an inulin-type fructan prebiotic: a randomised, double-blind, placebo-controlled, cross-over, human intervention study. Br. J. Nutr. 2018;119:176–189. doi: 10.1017/S0007114517003440. [DOI] [PubMed] [Google Scholar]
- 38.Hooda S, et al. 454 pyrosequencing reveals a shift in fecal microbiota of healthy adult men consuming polydextrose or soluble corn fiber. J. Nutr. 2012;142:1259–1265. doi: 10.3945/jn.112.158766. [DOI] [PubMed] [Google Scholar]
- 39.Kovatcheva-Datchary P, et al. Dietary fiber-induced improvement in glucose metabolism is associated with increased abundance of prevotella. Cell Metab. 2015;22:971–982. doi: 10.1016/j.cmet.2015.10.001. [DOI] [PubMed] [Google Scholar]
- 40.Liu F, et al. Fructooligosaccharide (FOS) and galactooligosaccharide (GOS) increase bifidobacterium but reduce butyrate producing bacteria with adverse glycemic metabolism in healthy young population. Sci. Rep. 2017;7:11789. doi: 10.1038/s41598-017-10722-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Morales P, et al. Impact of dietary lipids on colonic function and microbiota: an experimental approach involving orlistat-induced fat malabsorption in human volunteers. Clin. Transl. Gastroenterol. 2016;7:e161. doi: 10.1038/ctg.2016.20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Rasmussen HE, et al. Starch-entrapped microsphere fibers improve bowel habit but do not exhibit prebiotic capacity in those with unsatisfactory bowel habits: a Phase I, randomized, double-blind, controlled human trial. Nutr. Res. N. Y. N. 2017;44:27–37. doi: 10.1016/j.nutres.2017.05.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Tap J, et al. Gut microbiota richness promotes its stability upon increased dietary fibre intake in healthy adults. Environ. Microbiol. 2015;17:4954–4964. doi: 10.1111/1462-2920.13006. [DOI] [PubMed] [Google Scholar]
- 44.Venkataraman A, et al. Variable responses of human microbiomes to dietary supplementation with resistant starch. Microbiome. 2016;4:33. doi: 10.1186/s40168-016-0178-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Citations
- 2018. NCBI Sequence Read Archive. SRP128128
- 2020. NCBI Sequence Read Archive. SRP219296
- 2017. NCBI Sequence Read Archive. SRP120250
- 2015. NCBI Sequence Read Archive. SRP062889
- 2017. NCBI Sequence Read Archive. SRP106361
- 2016. NCBI Sequence Read Archive. SRP067761
- 2022. NCBI Sequence Read Archive. SRP403421
- Rodriguez CI, 2023. Curated and harmonized gut microbiome 16S rRNA amplicon sequences, metadata, and OTU tables from dietary fiber intervention studies in humans. Figshare. [DOI] [PMC free article] [PubMed]
Data Availability Statement
The parameters and step-by-step scripts used to clean up the data, remove chimeras, and assign taxonomy are available at https://github.com/cirodri1/fiber-data_records (e.g, trimming lengths, primers, databases, etc.).


