Abstract

Soluble signaling molecules and extracellular matrix (ECM) regulate cell dynamics in various biological processes. Wound healing assays are widely used to study cell dynamics in response to physiological stimuli. However, traditional scratch-based assays can damage the underlying ECM-coated substrates. Here, we use a rapid, non-destructive, label-free magnetic exclusion technique to form annular aggregates of bronchial epithelial cells on tissue-culture treated (TCT) and ECM-coated surfaces within 3 h. The cell-free areas enclosed by the annular aggregates are measured at different times to assess cell dynamics. The effects of various signaling molecules, including epidermal growth factor (EGF), oncostatin M, and interleukin 6, on cell-free area closures are investigated for each surface condition. Surface characterization techniques are used to measure the topography and wettability of the surfaces. Further, we demonstrate the formation of annular aggregates on human lung fibroblast-laden collagen hydrogel surfaces, which mimic the native tissue architecture. The cell-free area closures on hydrogels indicate that the substrate properties modulate EGF-mediated cell dynamics. The magnetic exclusion-based assay is a rapid and versatile alternative to traditional wound healing assays.
1. Introduction
Extracellular biochemical and biophysical cues guide cell behavior, including migration, proliferation, and differentiation.1 Soluble signaling molecules such as growth factors, cytokines, hormones, and neurotransmitters provide biochemical cues by binding to specific cell surface receptors that trigger a cascade of intracellular molecular events.2 Besides the soluble factors, the extracellular matrix (ECM) presents a variety of biochemical and biophysical cues to the cells.1,3,4 The ECM biochemical composition controls cell fate decisions by regulating integrin-mediated cell adhesion to ECM proteins.4,5 The ECM also sequesters and releases various soluble signaling molecules to influence cell behavior.6 In addition to its biochemical composition, the ECM mechanical properties provide biophysical cues to modulate cell phenotype and function.1,4 ECM stiffness affects the cytoskeletal framework within cells and drives changes in protein localization and gene expression.1,7 Moreover, the micro- and nano-scale topographical features of fibrous ECM proteins guide the cytoskeleton to influence cell motility.4,7
Growth factors are essential for efficient tissue repair and regeneration.2,8 Signaling molecules such as the epidermal growth factor (EGF) stimulate epithelial cell motility to re-establish the epithelial barrier disrupted by tissue injury.2,9 Besides growth factors, various cytokines are released in the wounded tissue that coordinate cellular processes during different healing stages, such as inflammation, regeneration, and tissue remodeling.2,10 Oncostatin M (OSM), a member of the interleukin 6 (IL6) family, is a pleiotropic cytokine that modulates cell differentiation, proliferation, and inflammatory networks.11,12 Besides homeostatic functions, OSM can act synergistically with various cytokines to promote immune cell infiltration and excessive ECM accumulation in chronic inflammatory conditions associated with skin, lungs, and joints.11,12 OSM has been reported to facilitate murine dermal wound closure by enhancing keratinocyte proliferation and migration.13 However, it is not well known whether OSM can have similar effects on epithelial cells of tissues other than the skin, such as the bronchial airway tract.
Wound healing assays are commonly used in vitro to study cell dynamics.14,15 The conventional 2D scratch-based assay involves scraping cells with a pipette tip or a pin tool to create a cell-free area in a cell monolayer.14−16 The changes in the cell-free area are monitored over time to assess cell migration from the surrounding intact regions. The scratch assay is straightforward and inexpensive, but scratching damages the underlying plastic surface or ECM substrate, which can significantly influence cell dynamics.14−16 While 2D monotypic assays provide fundamental insights into the molecular mechanisms of cell migration, they fail to mimic the in vivo tissue microenvironment.15 Recently, a 3D bronchial airway model of the lung added bronchial epithelial cell suspensions to ECM-coated 3D polydimethylsiloxane (PDMS) scaffolds to mimic the in vivo epithelium-lined tissue architecture.17 These cells attached to the scaffold surfaces and proliferated to form confluent monolayers.17
Label-free magnetic manipulation is a rapid, versatile, and scalable cell assembly technique. Cells are suspended in a medium with higher magnetic susceptibility and exposed to an inhomogeneous magnetic field. The susceptibility difference and magnetic field gradient result in a magnetophoretic force that drives the cells toward regions with the lowest magnetic field strength, where they form clusters.18 We previously demonstrated the formation of annular cell clusters using a label-free magnetic exclusion technique to study cell migration without significantly affecting the cell viability, metabolism, and transcriptome profiles.19 The magnetic exclusion technique is rapid, easy-to-use, and inexpensive and can be readily integrated into the existing multiwell plates. Moreover, the versatile label-free method can be used to pattern cellular structures on various substrates, including soft materials, where insert-based methods may be difficult to implement.
Here, we use the magnetic exclusion technique to form annular aggregates of human bronchial epithelial (HBEC3 KT) cells, enclosing cell-free areas. The cell-free area closures are observed over time to investigate the effects of OSM on HBEC3 KT dynamics. Since cell receptor complexes for OSM and IL6 have a common glycoprotein 130 (gp130) signal transduction chain, we stimulate HBEC3 KT with IL6 to determine whether this cytokine has similar effects on the cell dynamics as OSM.11 EGF is used as a positive control, as we have previously demonstrated enhanced HBEC3 KT migration due to EGF (0.8 ng mL–1) exposure.19 The annular aggregates are assembled on tissue culture-treated (TCT) and ECM-coated surfaces within 3 h to assess the impact of surfaces on cell-free area closures. Surface characterization techniques, i.e., atomic force microscopy (AFM) and tensiometry, are used to measure the ECM-coated surface properties. Further, we extend the magnetic method to demonstrate, for the first time, magnetic exclusion-based cell patterning on hydrogel surfaces. Annular HBEC3 KT aggregates are assembled on human lung fibroblast-laden collagen hydrogels to investigate the effects of 3D soft substrates on HBEC3 KT dynamics and determine whether hydrogel surfaces influence the signaling molecule-mediated cell-free area closures. Fibroblasts are introduced into the hydrogels to mimic the native multicellular tissue.
2. Results and Discussion
2.1. OSM and IL6 Bioactivity in HBEC3 KT
The binding of OSM and IL6 to their respective gp130 receptor complexes activates multiple intracellular signaling pathways, such as the Janus kinase (JAK)-signal transducer and activator of transcription (STAT) pathway.11,20 The JAK-STAT pathway involves the recruitment of latent transcription factors STATs that, following phosphorylation by tyrosine kinases JAKs, translocate to the nucleus to modulate gene expression.11,21 The OSM and IL6 ligand–receptor interactions are not well documented for HBEC3 KT. Therefore, we detect phosphorylated STAT3 (pSTAT3) levels, a common transcription factor regulated by OSM and IL6, in HBEC3 KT using immunoassay to determine cytokine bioactivity.11,21 The maximum OSM and IL6 concentration selected is 1 ng mL–1, i.e., the same order of magnitude used previously for in vitro stimulation.22 Since HBEC3 KT cells are exposed to 25 mM Gadavist solution to form annular aggregates within 3 h (Section 2.2), we investigate the impact of the Gadavist exposure on OSM and IL6 signaling. Figure 1 shows pSTAT3 absorbance values for HBEC3 KT stimulated with the cytokines for 12 h, with and without Gadavist pre-exposure. The figure shows that OSM (1 ng mL–1) and IL6 (1 ng mL–1) significantly increase pSTAT3 levels in HBEC3 KT over the control. Similar results are obtained for 25 mM Gadavist pre-exposed cells, which indicate that the Gadavist exposure for 3 h does not significantly affect the cytokine bioactivity.
Figure 1.

Phosphorylated STAT3 (pSTAT3) levels in HBEC3 KT cell lysates measured using an enzyme-linked immunosorbent assay (ELISA), with and without Gadavist exposure (N = 4). The cells are stimulated with OSM and IL6 for 12 h before the pSTAT3 detection. The statistically significant results are shown with asterisks. The signaling molecule concentrations in ng mL–1 are indicated in parentheses.
2.2. Annular HBEC3 KT Aggregate Formation Using Magnetic Exclusion
An array of coaxially arranged N52 grade
neodymium ring [12.7 mm (o.d.) × 6.35 mm (i.d.) × 6.35 mm]
and cylinder [3.175 mm (d) × 6.35 mm] magnets
(K&J Magnetics) is used to assemble the annular aggregates (Figure 2A). The magnetic
field distribution of the coaxial magnet arrangement consists of a
low magnetic field region in the annular space between the ring and
cylinder magnets (Figure 2B). HBEC3 KT cells suspended in 25 mM Gadavist solution are
seeded to the wells (7.4 × 7.4 mm) of a 96-well flat-bottom plate
placed on the magnet array. The cells experience a magnetic body force 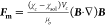 in the inhomogeneous magnetic field, where
(χc – χsol) denotes the magnetic susceptibility difference
between the cells and the solution, Vc denotes the cell volume, μo denotes the permeability of free space, and B represents magnetic flux density. For regular cell
culture medium, (χc – χsol) ≅ 0. Therefore, we add the Gadavist paramagnetic
agent to the culture medium to increase the susceptibility difference.
Since χc < χsol, the magnetic force Fm drives the cells toward the lowest magnetic field region on
the well surface, where they form clusters. The force balance equation
governing the cell motion in the suspension is
in the inhomogeneous magnetic field, where
(χc – χsol) denotes the magnetic susceptibility difference
between the cells and the solution, Vc denotes the cell volume, μo denotes the permeability of free space, and B represents magnetic flux density. For regular cell
culture medium, (χc – χsol) ≅ 0. Therefore, we add the Gadavist paramagnetic
agent to the culture medium to increase the susceptibility difference.
Since χc < χsol, the magnetic force Fm drives the cells toward the lowest magnetic field region on
the well surface, where they form clusters. The force balance equation
governing the cell motion in the suspension is
| 1 |
where mc and vc represent the mass and velocity of a cell, respectively. The viscous drag determined by Stokes’ law is Fd = −6πηavc, where η denotes the Gadavist solution viscosity and a denotes the cell radius. The net gravitational force Fg,net magnitude is (ρc – ρsol)gVc, where ρc denotes the cell density, ρsol denotes the Gadavist solution density, and g denotes the acceleration due to gravity. A mathematical analysis of the forces acting on cells during aggregate formation has been reported previously.23
Figure 2.
(A) Schematic of the magnet array consisting of coaxially arranged ring-cylinder magnets with the N–N orientation. A 96-well plate is positioned on the array such that the center of each ring-cylinder magnet aligns with a well center. (B) Magnetic flux density distribution of the ring-cylinder magnet arrangement. (C) Illustration of the steps to form an annular aggregate within 3 h. The magnetic force drives the HBEC3 KT cells toward the lowest magnetic field region (dark blue) on the well surface. (D) Fluorescence-stitched image of an annular HBEC3 KT monolayer. The cells are stained with a Qtracker labeling kit (orange). Here, t = 0 h denotes the time of adding the culture medium to the wells after the rinsing step. The initial cell-free area is marked with a white dotted line. The scale bar is 1 mm. (E) Fluorescence-stitched images of the cell-free areas on different surfaces for OSM (0.1 ng mL–1) at 12 and 18 h. The DECM concentrations in mg mL–1 are indicated in parentheses. The scale bar is 1 mm.
Figure 2C illustrates the steps for forming the annular HBEC3 KT aggregates. The wells containing the aggregates are rinsed with the regular culture medium after 3 h, which removes loose cell aggregates and results in a monolayer. The fluorescence image of an annular HBEC3 KT monolayer assembled on a TCT surface with the initial cell-free area marked by a white dotted line is shown in Figure 2D. Here, t = 0 h denotes the time when the culture medium is added to the wells after the washing step.
In addition to TCT, annular HBEC3 KT monolayers are formed on collagen- and decellularized matrix (DECM)-coated surfaces to investigate the influence of the underlying substrates on cell-free area closures. Since collagen type I and IV are among the major collagen subtypes that constitute lung ECM, we use mixtures of these subtypes to prepare the collagen-coated surfaces.24 Depending on the decellularization protocol, tissue-derived DECM comprises various ECM components such as collagen, elastin, fibronectin, and glycosaminoglycans (GAGs).25,26 We use human lung DECM to determine whether it can enhance cell-free area closures compared to only collagen. Porcine lung DECM (0.1 mg mL–1) has previously been used to investigate the effects of ECM proteins on epithelial barrier function.26 We use human lung DECM of the same concentration as the porcine lung DECM to coat the well surfaces. In addition, a 10-fold higher DECM concentration is used to investigate concentration-dependent effects on cell-free area closures. The average diameters (major and minor axes) of the initial cell-free regions enclosed by the annular monolayers are (∼3.70 ± 0.08 mm, ∼3.61 ± 0.07 mm), (∼3.45 ± 0.07 mm, ∼3.37 ± 0.08 mm), (∼3.49 ± 0.05 mm, ∼3.43 ± 0.04 mm), and (∼3.40 ± 0.06 mm, ∼3.33 ± 0.08 mm) for TCT, collagen-, DECM (0.1 mg mL–1)-, and DECM (1 mg mL–1)-coated surfaces, respectively [biological replicates (N) = 3, technical replicates = 6].
The annular HBEC3 KT monolayers formed on various surfaces are incubated with EGF (0.8 ng mL–1), OSM (0.1 and 1 ng mL–1), IL6 (0.1 and 1 ng mL–1), and combinations of EGF with OSM and IL6 to study the influence of these signaling molecules on cell behavior. In combination with other cytokines, OSM has synergistic effects on cellular pro-inflammatory responses.27,28 Hence, we combine EGF with OSM and IL6 to determine whether these signaling molecules can synergistically enhance cell-free area closures. Fluorescence images of cell-free area closures on different surfaces for OSM (0.1 ng mL–1) at 12 and 18 h are shown in Figure 2E.
2.3. Effects of Surfaces on Cell-Free Area Closures
The cell-free areas covered by HBEC3 KT on coated and uncoated TCT surfaces are compared at 12 and 18 h for each signaling molecule condition to analyze the surface effects on cell-free area closures (Figures 3A and S1). The collagen- and DECM-coated surfaces enhance cell-free area closures significantly more than TCT for the majority of the signaling molecule conditions. No significant differences in cell-free area coverage are observed between collagen- and DECM-coated surfaces. Noting that DECM contains collagen, the effects of collagen on cell-free area closures may be dominant over other DECM proteins. Moreover, the concentrations of non-collagen DECM proteins, post decellularization, may be inadequate to influence cell dynamics significantly. The binding of integrins, cell transmembrane receptors, to ligands such as collagen drives cell migration through integrin-actin cytoskeleton linkages.29,30 The cyclic formation and release of integrin-mediated adhesion to ligands are essential for cell migration.29 The maximum migration speed for most cell types occurs at intermediate ligand concentrations, where adhesion formation and release are efficient.29,31 The cell-free area closures do not increase monotonically with DECM concentration for HBEC3 KT. In contrast, DECM (0.1 mg mL–1) enhances cell-free area coverage more than DECM (1 mg mL–1) without exogenous signaling molecules (Figure 3Ai), which suggests that 0.1 mg mL–1 is closer to the optimum concentration for maximum closure rate.
Figure 3.
(A) Percentages of cell-free areas covered by HBEC3 KT on various surfaces for (i) control, (ii) EGF (0.8 ng mL–1), and (iii) OSM (1 ng mL–1) at 12 and 18 h (N ≥ 3). (B) Surface roughness profiles of (i) collagen-, (ii) DECM (0.1 mg mL–1)-, and (iii) DECM (1 mg mL–1)-coated surfaces. (C) Circularity values for (i) control, (ii) EGF (0.8 ng mL–1), and (iii) OSM (1 ng mL–1) at 0, 12, and 18 h (N ≥ 3). The DECM concentrations in mg mL–1 are indicated in parentheses. The statistically significant results are shown with asterisks. (D) (i) Fluorescence image of an annular HBEC3 KT monolayer section on TCT at 0 h. The cell front is shown with an arrow and (ii) ImageJ analysis of the same section shows the cell front with micro-scale irregularities. The scale bar is 500 μm.
The surface properties of ECM-coated substrates can regulate cell migration.4 We measure the wettability and surface topography of the substrates to identify factors that can affect closure rates. Surface wettability can influence cell adhesion to the substrate.32 Since wettability affects surface protein adsorption, the contact angles for all surfaces are measured after exposure to the cell culture medium for 3 h, i.e., the duration to form the annular aggregates. The contact angles for TCT, collagen-, DECM (0.1 mg mL–1)-, and DECM (1 mg mL–1)-coated surfaces are 5.73°, 7.73°, 6.20°, and 11°, respectively, indicating similar hydrophilicity of the surfaces. Unlike TCT plastic surface, cells in native tissues are exposed to topographical features, including nano-scale surface roughness, sub-micrometer-sized fibrils, and micrometer-sized protein bundles.1 We measure the topography of collagen- and DECM-coated surfaces using AFM. Figure 3B shows the surface roughness profiles of the coated surfaces. The average root-mean-square (RMS) roughnesses of collagen- and DECM (0.1 mg mL–1)-coated surfaces are ∼0.9 ± 0.04 and ∼1 ± 0.72 nm, respectively, i.e., the same order of magnitude, which may contribute to similar cell-free area coverage at 18 h without exogenous signaling molecules (Figure 3Ai). In contrast, the DECM (1 mg mL–1)-coated surfaces have a higher average RMS roughness of ∼18 ± 10.5 nm with fibrillar features. The fibrillar features may promote stronger cellular adhesions to the substrate and impede the release of adhesion complexes, resulting in slower cell migration without exogenous signaling molecules (Figure 3Ai).
Fluorescence
images of HBEC3 KT annular monolayers at 12 and 18
h (Figure 2E) indicate
surface-dependent uniformity of cell-free area closures. We use the
ImageJ dimensionless parameter Circularity = 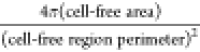 to assess the closure uniformity, where Circularity = 1 indicates a perfect circle. The Circularity
values for all surfaces and signaling molecule conditions at 0, 12,
and 18 h are shown in Figures 3C and S2. The cell-free areas are
more circular for collagen-coated surfaces than other surface conditions,
irrespective of the signaling molecules, which indicate higher closure
uniformity. The coefficient of variation of RMS roughnesses for collagen-,
DECM (0.1 mg mL–1)-, and DECM (1 mg mL–1)-coated surfaces are ∼4, ∼72, and ∼58%, respectively.
The higher closure uniformity on collagen surfaces may be attributed
to the lower variation in surface roughness, which allows uniform
integrin-mediated adhesions. In contrast, the surface roughness heterogeneity
of DECM-coated surfaces may be responsible for their lower Circularity values. The initial cell-free areas of the magnetically
assembled annular monolayers have a Circularity <
1 because the image processing method captures the micro-scale irregularities
of the cell front, as shown in Figure 3D.
to assess the closure uniformity, where Circularity = 1 indicates a perfect circle. The Circularity
values for all surfaces and signaling molecule conditions at 0, 12,
and 18 h are shown in Figures 3C and S2. The cell-free areas are
more circular for collagen-coated surfaces than other surface conditions,
irrespective of the signaling molecules, which indicate higher closure
uniformity. The coefficient of variation of RMS roughnesses for collagen-,
DECM (0.1 mg mL–1)-, and DECM (1 mg mL–1)-coated surfaces are ∼4, ∼72, and ∼58%, respectively.
The higher closure uniformity on collagen surfaces may be attributed
to the lower variation in surface roughness, which allows uniform
integrin-mediated adhesions. In contrast, the surface roughness heterogeneity
of DECM-coated surfaces may be responsible for their lower Circularity values. The initial cell-free areas of the magnetically
assembled annular monolayers have a Circularity <
1 because the image processing method captures the micro-scale irregularities
of the cell front, as shown in Figure 3D.
2.4. Effects of Signaling Molecules on Cell-free Area Closures
The percentages of cell-free areas covered
by HBEC3 KT in response to the signaling molecules are calculated
using 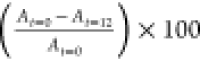 and
and  at 12 and 18 h, respectively, where At=0, At=12, and At=18 denote the cell-free areas at 0, 12, and 18 h (Figure 4), respectively.
The cell-free area covered due to EGF (0.8 ng mL–1) exposure is significantly higher than the control for all surfaces
at 12 h (Figure 4).
In contrast, OSM and IL6 fail to enhance cell-free area closures significantly,
except for DECM (1 mg mL–1)-coated surfaces, where
statistically significant closures are obtained for OSM (1 ng mL–1) and IL6 (1 ng mL–1) (Figure 4D). Similar to EGF
(0.8 ng mL–1), exposures to combinations of EGF
with OSM and IL6 result in statistically significant closures on all
surfaces at 12 h. Since no significant differences are observed between
the cell-free areas covered due to EGF and the signaling molecule
combinations, these results can be attributed to EGF (0.8 ng mL–1) alone. We hypothesize that the synergistic effects
of OSM on cellular responses occur only in combinations with pro-inflammatory
cytokines. Further experiments with other growth factors are required
to validate this conjecture.
at 12 and 18 h, respectively, where At=0, At=12, and At=18 denote the cell-free areas at 0, 12, and 18 h (Figure 4), respectively.
The cell-free area covered due to EGF (0.8 ng mL–1) exposure is significantly higher than the control for all surfaces
at 12 h (Figure 4).
In contrast, OSM and IL6 fail to enhance cell-free area closures significantly,
except for DECM (1 mg mL–1)-coated surfaces, where
statistically significant closures are obtained for OSM (1 ng mL–1) and IL6 (1 ng mL–1) (Figure 4D). Similar to EGF
(0.8 ng mL–1), exposures to combinations of EGF
with OSM and IL6 result in statistically significant closures on all
surfaces at 12 h. Since no significant differences are observed between
the cell-free areas covered due to EGF and the signaling molecule
combinations, these results can be attributed to EGF (0.8 ng mL–1) alone. We hypothesize that the synergistic effects
of OSM on cellular responses occur only in combinations with pro-inflammatory
cytokines. Further experiments with other growth factors are required
to validate this conjecture.
Figure 4.
Percentages of cell-free areas covered by HBEC3 KT on (A) TCT (N = 3), (B) collagen- (N = 4), (C) DECM (0.1 mg mL–1)- (N = 4), and (D) DECM (1 mg mL–1)-coated surfaces at 12 and 18 h in response to various signaling molecule conditions (N = 4). The statistically significant results are shown with asterisks. The signaling molecule concentrations in ng mL–1 are indicated in parentheses.
The repopulation of the cell-free areas can occur
due to cell proliferation
and migration from the neighboring high-cell-density regions. The
cell counts of the entire well (cw) and the initial cell-free region (ccf) are obtained for all surfaces and signaling molecule conditions
at 0 and 18 h to assess the contributions of cell proliferation and
migration to the cell-free area closures. The overall cell densities
of the wells and the localized densities within the initial cell-free
regions are determined using 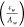 and
and 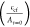 , where Aw and At=0 denote
the well and the initial cell-free areas, respectively. At 18 h, the
overall cell densities increase by ∼199 (control), ∼248
(0.8 ng mL–1 EGF), ∼284 (0.1 ng mL–1 OSM), ∼141 (1 ng mL–1 OSM), ∼162
(0.1 ng mL–1 IL6), and ∼189 cells mm–2 (1 ng mL–1 IL6) on TCT surfaces,
which are attributed to cell proliferation. In contrast, the localized
cell densities within the initial cell-free regions of TCT increase
to ∼706 (control), ∼928 (0.8 ng mL–1 EGF), ∼707 (0.1 ng mL–1 OSM), ∼801
(1 ng mL–1 OSM), ∼714 (0.1 ng mL–1 IL6), and ∼785 cells mm–2 (1 ng mL–1 IL6) at 18 h, i.e., the cell densities are approximately
4–5-fold greater than the overall increase in cell densities
due to proliferation. For collagen- and DECM-coated surfaces, the
cell densities in the initial cell-free regions are 5–10-fold
greater than the overall increase in cell densities at 18 h. These
results suggest that the cell-free area closures occur primarily by
cell migration.
, where Aw and At=0 denote
the well and the initial cell-free areas, respectively. At 18 h, the
overall cell densities increase by ∼199 (control), ∼248
(0.8 ng mL–1 EGF), ∼284 (0.1 ng mL–1 OSM), ∼141 (1 ng mL–1 OSM), ∼162
(0.1 ng mL–1 IL6), and ∼189 cells mm–2 (1 ng mL–1 IL6) on TCT surfaces,
which are attributed to cell proliferation. In contrast, the localized
cell densities within the initial cell-free regions of TCT increase
to ∼706 (control), ∼928 (0.8 ng mL–1 EGF), ∼707 (0.1 ng mL–1 OSM), ∼801
(1 ng mL–1 OSM), ∼714 (0.1 ng mL–1 IL6), and ∼785 cells mm–2 (1 ng mL–1 IL6) at 18 h, i.e., the cell densities are approximately
4–5-fold greater than the overall increase in cell densities
due to proliferation. For collagen- and DECM-coated surfaces, the
cell densities in the initial cell-free regions are 5–10-fold
greater than the overall increase in cell densities at 18 h. These
results suggest that the cell-free area closures occur primarily by
cell migration.
2.5. Annular HBEC3 KT Aggregate Formation and Cell-free Area Closures on Fibroblast-Laden Collagen Hydrogels
Substrate stiffness significantly affects cell dynamics.33,34 The cell migration speeds on ECM-coated substrates do not vary monotonically with substrate stiffness.34 Instead, the maximum speed occurs within a specific stiffness range depending on the cell type and ECM concentration.34 We form HBEC3 KT annular aggregates on hydrogels to investigate the effects of substrate stiffness on cell-free area closures. Since collagen and DECM have similar impacts on cell-free area closures, collagen hydrogels are used for this study. Human primary lung fibroblasts are added to the hydrogels to mimic the cellular composition of native tissues. Since collagen gels suffer from poor mechanical properties without covalent crosslinking, we use four-arm polyethylene glycol succinimidyl glutarate (PEG-SG) as a crosslinker.35 PEG-SG has previously been used to prepare cell-laden collagen gels with high mechanical stability and cell viability.35,36 The Young’s modulus of PEG-SG-crosslinked collagen gels is in the range of ∼0.5–1.5 kPa, i.e., approximately two orders of magnitude lower than collagen-coated surfaces (∼100–200 kPa).36,37 Due to the collagen solution meniscus, the hydrogels have curved surfaces with thicknesses of ∼800 μm at the well boundaries and ∼120 μm near the well center of a 96-well plate. HBEC3 KT annular aggregates formed on curved surfaces are difficult to image as cells are located at different focal planes. Therefore, to minimize the meniscus effect, we prepare the hydrogels in 24-well plates with larger wells (well diameter = ∼14 mm). Ring magnets with an outer diameter of ∼19 mm are used to account for the larger well size. The remaining dimensions of the ring-cylinder magnet arrangement are the same as the magnets used to form aggregates in 96-well plates. Due to the excess liquid content of hydrogels (>90%), the Gadavist concentration is adjusted to a final concentration of 25 mM in the wells.35 The incubation duration on the magnet array is increased from 3 to 6 h because of the larger well volume of the 24-well plates. Figure 5A shows the steps of forming HBEC3 KT annular aggregates on the fibroblast-laden hydrogels using the magnetic exclusion technique. Initially, the fibroblasts in the hydrogels have a globular morphology, which becomes more elongated over a period of 24 h (Figure 5B). Suspensions of HBEC3 KT in Gadavist are added to the wells at 24 h after forming the fibroblast-laden hydrogels, allowing the fibroblasts to adapt to the hydrogel environment.
Figure 5.
(A) Illustration of the steps to form an annular aggregate on a fibroblast-laden collagen hydrogel crosslinked with PEG-SG. (B) Brightfield images show the fibroblast morphologies in the collagen hydrogel at (i) 0 and (ii) 24 h. Here, t = 0 h denotes the time of adding the cell culture medium to the wells of a 24-well plate after forming the fibroblast-laden PEG-SG crosslinked collagen hydrogels. The scale bar is 200 μm. (C) (i) Fluorescence-stitched image of an annular HBEC3 KT aggregate assembled on a fibroblast-laden hydrogel using the magnetic exclusion technique. The cells are stained with a Qtracker labeling kit (orange) and (ii) a fluorescence-stitched image of the same annular aggregate (orange) with the fibroblasts stained with calcein AM (green). The figure shows the maximum intensity projection of the combined z-stack images of the hydrogel. The initial cell-free region is marked with a white dotted line. The scale bar is 1 mm. (D) Effect of Gadavist exposure on the metabolism of fibroblasts within the hydrogel, measured using a resazurin-based assay (technical replicates = 4). (E) Percentages of cell-free areas covered by HBEC3 KT on hydrogel surfaces at 12 and 18 h in response to the signaling molecules (N = 3). (F) Brightfield images show the HBEC3 KT morphologies in annular aggregates magnetically assembled on TCT and hydrogel surfaces. The cell fronts are indicated with red dotted lines. Here, t = 0 h denotes the time of adding the regular culture medium to the wells after the HBEC3 KT annular aggregate formation. The scale bar is 50 μm. (G) Fluorescence-stitched images of cell-free area closures on hydrogels in response to 25-fold higher signaling molecule concentrations. The scale bar is 1 mm. (H) Percentages of cell-free areas covered by HBEC3 KT on hydrogel surfaces at 12 and 18 h, stimulated with higher signaling molecule concentrations (N = 4). The signaling molecule concentrations in ng mL–1 are indicated in parentheses.
Figure 5Ci shows the fluorescence image of an annular HBEC3 KT aggregate assembled on a fibroblast-laden hydrogel using the magnetic exclusion technique. The fibroblasts within the hydrogel are stained with calcein AM-stained (green), and the maximum intensity projection of the combined z-stack images is shown in Figure 5Cii.
We have previously demonstrated that Gadavist exposure has insignificant effects on HBEC3 KT viability.19 The impact of Gadavist exposure on primary fibroblast metabolism, an indicator of viability, is investigated using a resazurin-based assay (Presto Blue HS). No significant difference in metabolic activity is observed between the control and Gadavist-exposed fibroblasts (Figure 5D). The annular HBEC3 KT aggregates are incubated with EGF (final concentration = 0.8 ng mL–1), OSM (final concentration = 1 ng mL–1), IL6 (final concentration = 1 ng mL–1), and combinations of EGF with OSM and IL6 to study the cell-free area closures on hydrogel surfaces at 12 and 18 h. The signaling molecule concentrations are adjusted to the final concentrations, accounting for the excess liquid content of hydrogels. Figure 5E shows that the cell-free area closures on collagen hydrogel surfaces are slower than the collagen-coated surface (Figure 4B), with no significant differences in area closures between the control and the signaling molecules at 12 and 18 h. The cell migration speed profile varies with substrate stiffness depending on the cell type.34 The delayed cell-free area closures on the hydrogels can be attributed to the lower substrate stiffness. Further, the higher collagen concentration (∼4 mg mL–1) of the hydrogels compared to the collagen-coated surfaces (∼0.1 mg mL–1) can decrease the closure rates by promoting stronger cell-substrate adhesions.31Figure 5F shows that the majority of the HBEC3 KT cells on the hydrogel surface have a globular morphology compared to the TCT surface at 0 h. The cells spread over 12 h to form compact monolayers on the hydrogels.
The HBEC3 KT cells are stimulated with higher concentrations of EGF, OSM, and IL6 to investigate the effects of signaling molecule concentrations on cell-free area closures on hydrogels. Since 25 ng mL–1 OSM has previously been used to stimulate human epithelial cells, the signaling molecule concentrations are increased 25-fold.13 Fluorescence images of cell-free area closures on hydrogel surfaces at 12 and 18 h in response to higher signaling molecule concentrations are shown in Figure 5G. While exposures to EGF (final concentration = 20 ng mL–1), OSM (final concentration = 25 ng mL–1), and IL6 (final concentration = 25 ng mL–1) increase the mean values of the cell-free areas covered on hydrogel surfaces, only OSM enhances cell-free area coverage significantly more than the control at 18 h (Figure 5H). In contrast to TCT and ECM-coated surfaces, the delayed cell-free area closures on hydrogels in response to EGF indicate that the substrate properties modulate EGF-mediated cell dynamics. Fluorescence z-stack images of the hydrogel at multiple locations are analyzed to determine whether the HBEC3 KT cells migrate within the hydrogel bulk. Figure S3 shows that the HBEC3 KT cells are located on the same focal plane of their respective locations on the hydrogel at 18 h, indicating that the cells migrate primarily on the hydrogel surfaces.
3. Conclusions
We use label-free magnetic exclusion to form HBEC3 KT annular aggregates and investigate the effects of various surface and signaling biomolecule conditions on the cell-free area closures. On TCT, collagen-, and DECM-coated surfaces, EGF (0.8 ng mL–1) enhances cell-free area closures significantly, much more than with OSM (0.1 and 1 ng mL–1) and IL6 (0.1 and 1 ng mL–1). The cell-free area closures occur more rapidly on collagen- and DECM-coated surfaces than on TCT. However, the closure rates do not increase monotonically with DECM concentration. Topography measurements reveal that collagen- and DECM (0.1 mg mL–1)-coated surfaces have RMS roughnesses of the same order of magnitude. Fibrillar features are observed on DECM (1 mg mL–1)-coated surfaces, resulting in a higher RMS roughness. Among all surfaces, cell-free areas become covered more uniformly on collagen surfaces. The magnetic technique also produces annular HBEC3 KT aggregates on fibroblast-laden collagen hydrogel surfaces. Cell-free area closures on hydrogels are slower than on coated surfaces, which is attributed to the lower stiffness and higher collagen concentration of the hydrogels. No significant differences are observed in the cell-free area coverage on hydrogel surfaces between the control and when signaling molecules are introduced, including EGF (0.8 ng mL–1). On increasing the concentration to 25 ng mL–1, OSM enhances cell-free area coverage on hydrogel surfaces significantly more than the control at 18 h. The cell-free area closures on hydrogel surfaces show that the substrate properties modulate the effects of EGF on cell dynamics. The HBEC3 KT-fibroblast wound-healing model can potentially be used to mimic pathological lung conditions and investigate intercellular crosstalk in an in vivo-like tissue environment.
4. Experimental Methods
4.1. Cell Culture
Immortalized human bronchial epithelial cells HBEC3 KT (ATCC) were cultured using keratinocyte serum-free medium (KSFM) supplemented with 0.8 ng mL–1 recombinant human EGF, 50 μg mL–1 bovine pituitary extract, and 1× penicillin/streptomycin (ThermoFisher Scientific). Human lung primary fibroblasts (passage 2) were received as a gift from the laboratory of Dr. Kjetil Ask (McMaster University). Tumor-free regions of lung tissue biopsy samples obtained from lung cancer patients were used to isolate the fibroblasts. All works using the human samples were conducted with patients’ consent in compliance with the ethical and scientific guidelines approved by the Hamilton Integrated Research Ethics Board (HiREB), #00–1839. The primary fibroblasts were cultured using Advanced Dulbecco’s Modified Eagle Medium (DMEM) supplemented with 5% fetal bovine serum, 1× GlutaMAX, and 1× penicillin/streptomycin (ThermoFisher Scientific). All cells were maintained at 37° C and 5% CO2 in a humidified atmosphere.
4.2. Phospho-STAT3 (Y705) Assay
HBEC3 KT cells were seeded into 12-well TCT plates at a density of ∼1.2 × 105 cells per well. After 24 h incubation, the cells were exposed to either 25 mM Gadavist or regular KSFM (control) for 3 h. Subsequently, the wells were rinsed, and cells were exposed to recombinant human OSM (0.1 and 1 ng mL–1; R&D Systems) and IL6 (0.1 and 1 ng mL–1; R&D Systems) for 12 h. The pSTAT3 levels were measured in the HBEC3 KT lysates using the pSTAT3 (Y705) DuoSet ELISA kit (R&D Systems), as per the manufacturer’s instructions.
4.3. Surface Preparations and Annular HBEC3 KT Aggregate Formation
Lung tissue samples were obtained from patients with a history of smoking and suspected lung cancer that required surgical interventions. The tissue samples were collected following informed consent from the patients, and the experimental protocol was approved by the HiREB, #5305-T. The lung DECM used in this study was devoid of any visible tumor cells, as determined by a certified pathologist. The tissue samples were decellularized as previously described.25 The decellularized tissue samples were lyophilized, milled, and mechanically homogenized for 4 h in a buffer containing pepsin (1 mg mL–1) dissolved in 0.01 M sterile HCL.(26) The resulting DECM stock solution (8 mg mL–1) was diluted to 1 and 0.1 mg mL–1 concentrations using 0.1 M acetic acid for coating TCT well surfaces of 96-well plates (IBIDI). The collagen solution was prepared by mixing rat tail collagen type I (final concentration = 0.1 mg mL–1; Advanced Biomatrix) and mouse collagen type IV (final concentration = 0.01 mg mL–1; Corning).26,38 The wells were coated with DECM and collagen for ∼24 h at 4° C on an orbital shaker and, subsequently, rinsed with phosphate-buffered saline. Annular HBEC3 KT aggregates were formed using the magnetic exclusion technique, as previously described.19 Briefly, ∼250 μL suspensions of HBEC3 KT in 25 mM Gadavist (∼2 × 105 cells mL–1) were added to the coated and uncoated wells placed on a ring-cylinder magnet array. The cells were incubated for 3 h (37° C and 5% CO2 in a humidified atmosphere) to form the annular aggregates. Subsequently, the well plate was taken off the array, and the wells were rinsed with regular KSFM to obtain annular HBEC3 KT monolayers. The cells were pre-stained using a Qtracker labeling kit (ThermoFisher Scientific), suitable for long-term cell tracking, following the manufacturer’s instructions. Fluorescence stitched images (excitation/emission = 540/605 nm) were taken using a 10× objective of a Nikon Eclipse Ti2 inverted microscope.
4.4. Surface Characterizations
The TCT, collagen-, and DECM-coated well surfaces were exposed to regular KSFM for 3 h and dried at room temperature before measuring the surface wettability. The contact angles were determined by the sessile drop method using an FTÅ200 tensiometer (First Ten Angstroms). The surface roughnesses of collagen- and DECM-coated substrates were measured using AFM (Dimension Icon, Bruker). Silicon nitride cantilevers with spring constant 0.35 N m–1 (SNL-10 probes, Bruker) were used, and the measurements were performed at a 1 Hz scan rate. NanoScope Analysis software was used to generate surface roughness profiles and measure RMS roughness at multiple locations.
4.5. Geometric Measurements and Cell Counting
The cell-free area dimensions were measured using ImageJ. The fluorescence image contrast was adjusted with the Enhance Local Contrast (CLAHE) function. We used the Sobel edge detection algorithm (Process–Find edges) to highlight the annular monolayer boundaries. Image smoothing was performed using Gaussian Blur (Process–Filter–Gaussian Blur), and the threshold was set automatically. The Fit ellipse function was used to determine the cell-free area diameters along the major and minor axes. The cell-free area circularity was measured using the Circularity shape descriptor function. The cell counts were obtained automatically from the fluorescence images using ImageJ. Individual cells in high cell-density regions were detected with the watershed segmentation algorithm (Process–Binary–Watershed).
4.6. Annular HBEC3 KT Aggregate Formation on Fibroblast-Laden Collagen Hydrogels
Bovine collagen type I solution in 0.01 M HCL (Advanced Biomatrix) was mixed with 10× Hanks’ Balanced Salt Solution (HBSS, Sigma-Aldrich), and the pH was adjusted using 1 M sterile NaOH (Sigma-Aldrich). Advanced DMEM was added to adjust the collagen concentration to ∼8 mg mL–1, and 4arm-PEG-SG (molecular weight 10,000; Sigma-Aldrich) was used as the crosslinker. ∼150 μL mixtures of collagen, PEG-SG, and primary lung fibroblasts were added to the TCT wells of 24-well plates (IBIDI) and incubated for 1.5 h at 37° C to form hydrogels. The final concentrations of collagen, PEG-SG, and fibroblasts in the hydrogel solution were ∼4 mg mL–1, ∼2 mg mL–1, and ∼5 × 105 cells mL–1, respectively. After 24 h, ∼300 μL suspensions of HBEC3 KT in Gadavist (∼2 × 105 cells mL–1) were added to the wells. The cells were incubated for 6 h to assemble annular HBEC3 KT aggregates on the hydrogel surfaces using the magnetic-exclusion technique. The fibroblasts and HBEC3 KT were exposed to co-culture media consisting of KSFM and serum-free Advanced DMEM mixed in a 1:1 volumetric ratio. Calcein AM (2 μM) solution was used to stain fibroblasts within the hydrogels. Fluorescence z-stack images (excitation/emission = 480/535 nm) were taken using a 4× objective of a Nikon Eclipse Ti2 inverted microscope.
4.7. Cell Metabolism Assay
Fibroblast-laden hydrogels in 24-well plates were incubated with either 25 mM Gadavist or regular Advanced DMEM (control) for 6 h. Subsequently, the wells were rinsed with the regular medium. Following 18 h incubation, the cell metabolic activity was measured using the resazurin-based Presto Blue assay (ThermoFisher Scientific), as per the manufacturer’s instructions. Blank wells containing only culture medium were used for background correction.
4.8. Statistical Analysis
All statistical tests were performed using the GraphPad Prism software (version 9.4.1). Two-way ANOVA with Tukey’s multiple comparisons test was used to analyze the effects of Gadavist exposure and cytokine stimulation on pSTAT3 levels in HBEC3 KT. The effects of signaling molecules on cell-free area closures at different times were analyzed using two-way ANOVA (repeated measures) with Tukey’s multiple comparisons test. The same statistical test was used to assess the effects of surfaces on cell dynamics and the Circularity values at different times. The fibroblast metabolism of the control and Gadavist groups were compared at 18 h using an unpaired t-test. A p-value < 0.05 was considered statistically significant.
4.9. Simulation
The Magnetic Fields, No Currents (mfnc) interface of COMSOL Multiphysics 5.3a was used to simulate the magnetic field distribution of the ring-cylinder magnet arrangement. A remanent flux density of 1.4 T was used to model the neodymium N52 grade magnets.
Acknowledgments
The authors thank Dr. Kjetil Ask (McMaster University) for providing human lung primary fibroblasts. This work was supported by the Natural Sciences and Engineering Research Council of Canada (NSERC) through a Discovery grant (RGPIN-2019-06571) and an Idea 2 Innovation grant (I2IPJ 555927-20).
Supporting Information Available
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acsomega.3c02052.
Percentages of cell-free areas covered by HBEC3 KT on various surfaces for OSM (0.1 ng mL–1), IL6 (0.1 ng mL–1), IL6 (1 ng mL–1), EGF (0.8 ng mL–1) with OSM (0.1 ng mL–1), EGF (0.8 ng mL–1) with OSM (1 ng mL–1), EGF (0.8 ng mL–1) with IL6 (0.1 ng mL–1), and EGF (0.8 ng mL–1) with IL6 (1 ng mL–1); Circularity values for OSM (0.1 ng mL–1), IL6 (0.1 ng mL–1), IL6 (1 ng mL–1), EGF (0.8 ng mL–1) with OSM (0.1 ng mL–1), EGF (0.8 ng mL–1) with OSM (1 ng mL–1), EGF (0.8 ng mL–1) with IL6 (0.1 ng mL–1), and EGF (0.8 ng mL–1) with IL6 (1 ng mL–1); and fluorescence z-stack images of the collagen hydrogel at different locations (PDF)
The authors declare no competing financial interest.
Supplementary Material
References
- Gasiorowski J. Z.; Murphy C. J.; Nealey P. F. Biophysical Cues and Cell Behavior: The Big Impact of Little Things. Annu. Rev. Biomed. Eng. 2013, 15, 155–176. 10.1146/annurev-bioeng-071811-150021. [DOI] [PubMed] [Google Scholar]
- Barrientos S.; Stojadinovic O.; Golinko M. S.; Brem H.; Tomic-Canic M. PERSPECTIVE ARTICLE: Growth factors and cytokines in wound healing: Growth factors and cytokines in wound healing. Wound Repair Regen 2008, 16, 585–601. 10.1111/j.1524-475X.2008.00410.x. [DOI] [PubMed] [Google Scholar]
- Muncie J. M.; Weaver V. M. The Physical and Biochemical Properties of the Extracellular Matrix Regulate Cell Fate. Curr. Top. Dev. Biol. 2018, 130, 1–37. 10.1016/bs.ctdb.2018.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Janson I. A.; Putnam A. J. Extracellular Matrix Elasticity and Topography: Material-Based Cues That Affect Cell Function via Conserved Mechanisms. J. Biomed. Mater. Res. Part A 2015, 103, 1246–1258. 10.1002/jbm.a.35254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moreno-Layseca P.; Streuli C. H. Signalling Pathways Linking Integrins with Cell Cycle Progression. Matrix Biol. 2014, 34, 144–153. 10.1016/j.matbio.2013.10.011. [DOI] [PubMed] [Google Scholar]
- Bonnans C.; Chou J.; Werb Z. Remodelling the Extracellular Matrix in Development and Disease. Nat. Rev. Mol. Cell Biol. 2014, 15, 786–801. 10.1038/nrm3904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun Y.; Chen C. S.; Fu J. Forcing Stem Cells to Behave: A Biophysical Perspective of the Cellular Microenvironment. Annu. Rev. Biophys. 2012, 41, 519–542. 10.1146/annurev-biophys-042910-155306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ren X.; Zhao M.; Lash B.; Martino M. M.; Julier Z. Growth Factor Engineering Strategies for Regenerative Medicine Applications. Front. Bioeng. Biotechnol. 2020, 7, 1–9. 10.3389/fbioe.2019.00469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bodnar R. J. Epidermal Growth Factor and Epidermal Growth Factor Receptor: The Yin and Yang in the Treatment of Cutaneous Wounds and Cancer. Adv. Wound Care 2013, 2, 24–29. 10.1089/wound.2011.0326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barrientos S.; Brem H.; Stojadinovic O.; Tomic-Canic M. Clinical Application of Growth Factors and Cytokines in Wound Healing. Wound Repair Regen 2014, 22, 569–578. 10.1111/wrr.12205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richards C. D. The Enigmatic Cytokine Oncostatin M and Roles in Disease. ISRN Inflamm 2013, 2013, 1–23. 10.1155/2013/512103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stawski L.; Trojanowska M. Oncostatin M and Its Role in Fibrosis. Connect. Tissue Res. 2019, 60, 40–49. 10.1080/03008207.2018.1500558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Das A.; Madeshiya A. K.; Biswas N.; Ghosh N.; Gorain M.; Rawat A.; Mahajan S. P.; Khanna S.; Sen C. K.; Roy S. Oncostatin M Improves Cutaneous Wound Re-Epithelialization and Is Deficient under Diabetic Conditions. J. Invest. Dermatol. 2022, 142, 679–691. 10.1016/j.jid.2021.04.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Riahi R.; Yang Y.; Zhang D. D.; Wong P. K. Advances in Wound-Healing Assays for Probing Collective Cell Migration. J. Lab. Autom. 2012, 17, 59–65. 10.1177/2211068211426550. [DOI] [PubMed] [Google Scholar]
- Kramer N.; Walzl A.; Unger C.; Rosner M.; Krupitza G.; Hengstschläger M.; Dolznig H. In vitro cell migration and invasion assays. Mutat. Res. Mutat. Res. 2013, 752, 10–24. 10.1016/j.mrrev.2012.08.001. [DOI] [PubMed] [Google Scholar]
- Hulkower K. I.; Herber R. L. Cell Migration and Invasion Assays as Tools for Drug Discovery. Pharmaceutics 2011, 3, 107–124. 10.3390/pharmaceutics3010107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nof E.; Artzy-Schnirman A.; Bhardwaj S.; Sabatan H.; Waisman D.; Hochwald O.; Gruber M.; Borenstein-Levin L.; Sznitman J.. Ventilation-Induced Epithelial Injury Drives Biological Onset of Lung Trauma in Vitro and Is Mitigated with Prophylactic Anti-Inflammatory Therapeutics. Bioeng. Transl. Med. 2022, 7(). 10.1002/btm2.10271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Akiyama Y.; Morishima K. Label-Free Cell Aggregate Formation Based on the Magneto-Archimedes Effect. Appl. Phys. Lett. 2011, 98, 163702. 10.1063/1.3581883. [DOI] [Google Scholar]
- Gupta T.; Gupta R.; Dabaghi M.; Sahu R. P.; Hirota J. A.; Puri I. K. Label-Free Cell Migration Assay Using Magnetic Exclusion. Adv. Mater. Technol. 2022, 7, 2101033. 10.1002/admt.202101033. [DOI] [Google Scholar]
- Su H.; Lei C. T.; Zhang C. Interleukin-6 Signaling Pathway and Its Role in Kidney Disease: An Update. Front. Immunol. 2017, 8, 1–10. 10.3389/fimmu.2017.00405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Balestrini J. L.; Niklason L. E. Extracellular Matrix as a Driver for Lung Regeneration. Ann. Biomed. Eng. 2015, 43, 568–576. 10.1007/s10439-014-1167-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wong S.; Botelho F. M.; Rodrigues R. M.; Richards C. D. Oncostatin M Overexpression Induces Matrix Deposition, STAT3 Activation, and SMAD1 Dysregulation in Lungs of Fibrosis-Resistant BALB/c Mice. Lab. Investig. 2014, 94, 1003–1016. 10.1038/labinvest.2014.81. [DOI] [PubMed] [Google Scholar]
- Ghosh D.; Gupta T.; Sahu R. P.; Das P. K.; Puri I. K. Three-Dimensional Printing of Diamagnetic Microparticles in Paramagnetic and Diamagnetic Media. Phys. Fluids 2020, 32, 072001. 10.1063/5.0012522. [DOI] [Google Scholar]
- Balestrini J. L.; Niklason L. E. Extracellular Matrix as a Driver for Lung Regeneration. Ann. Biomed. Eng. 2015, 43, 568–576. 10.1007/s10439-014-1167-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dabaghi M.; Saraei N.; Carpio M. B.; Nanduri V.; Ungureanu J.; Babi M.; Chandiramohan A.; Noble A.; Revill S. D.; Zhang B.; Ask K.; Kolb M.; Shargall Y.; Moran-Mirabal J.; Hirota J. A. A Robust Protocol for Decellularized Human Lung Bioink Generation Amenable to 2d and 3d Lung Cell Culture. Cells 2021, 10, 1538–1623. 10.3390/cells10061538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young B. M.; Shankar K.; Tho C. K.; Pellegrino A. R.; Heise R. L. Laminin-Driven Epac/Rap1 Regulation of Epithelial Barriers on Decellularized Matrix. Acta Biomater. 2019, 100, 223–234. 10.1016/j.actbio.2019.10.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barksby H. E.; Hui W.; Wappler I.; Peters H. H.; Milner J. M.; Richards C. D.; Cawston T. E.; Rowan A. D. Interleukin-1 in Combination with Oncostatin M up-Regulates Multiple Genes in Chondrocytes: Implications for Cartilage Destruction and Repair. Arthritis Rheum 2006, 54, 540–550. 10.1002/art.21574. [DOI] [PubMed] [Google Scholar]
- Guilloteau K.; Paris I.; Pedretti N.; Boniface K.; Juchaux F.; Huguier V.; Guillet G.; Bernard F.-X.; Lecron J.-C.; Morel F. Skin Inflammation Induced by the Synergistic Action of IL-17A, IL-22, Oncostatin M, IL-1α, and TNF-α Recapitulates Some Features of Psoriasis. J. Immunol. 2010, 184, 5263–5270. 10.4049/jimmunol.0902464. [DOI] [PubMed] [Google Scholar]
- Huttenlocher A.; Horwitz A. R. Integrins in Cell Migration. Cold Spring Harb. Perspect. Biol. 2011, 3, 0050744–a5116. 10.1101/cshperspect.a005074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brakebusch C. NEW EMBO MEMBER’S REVIEW: The Integrin-Actin Connection, an Eternal Love Affair. EMBO J. 2003, 22, 2324–2333. 10.1093/emboj/cdg245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwartz M. A.; Horwitz A. R. Integrating Adhesion, Protrusion, and Contraction during Cell Migration. Cell 2006, 125, 1223–1225. 10.1016/j.cell.2006.06.015. [DOI] [PubMed] [Google Scholar]
- Cai S.; Wu C.; Yang W.; Liang W.; Yu H.; Liu L. Recent Advance in Surface Modification for Regulating Cell Adhesion and Behaviors. Nanotechnol. Rev. 2020, 9, 971–989. 10.1515/ntrev-2020-0076. [DOI] [Google Scholar]
- Lo C. M.; Wang H. B.; Dembo M.; Wang Y. L. Cell Movement Is Guided by the Rigidity of the Substrate. Biophys. J. 2000, 79, 144–152. 10.1016/S0006-3495(00)76279-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Janmey P. A.; Fletcher D. A.; Reinhart-King C. A. Stiffness Sensing by Cells. Physiol. Rev. 2020, 100, 695–724. 10.1152/physrev.00013.2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sarrigiannidis S. O.; Rey J. M.; Dobre O.; González-García C.; Dalby M. J.; Salmeron-Sanchez M. A Tough Act to Follow: Collagen Hydrogel Modifications to Improve Mechanical and Growth Factor Loading Capabilities. Mater. Today Bio 2021, 10, 100098. 10.1016/j.mtbio.2021.100098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lotz C.; Schmid F. F.; Oechsle E.; Monaghan M. G.; Walles H.; Groeber-Becker F. Cross-Linked Collagen Hydrogel Matrix Resisting Contraction to Facilitate Full-Thickness Skin Equivalents. ACS Appl. Mater. Interfaces 2017, 9, 20417–20425. 10.1021/acsami.7b04017. [DOI] [PubMed] [Google Scholar]
- Vasse G. F.; Kühn P. T.; Zhou Q.; Bhusari S. A.; Reker-Smit C.; Melgert B. N.; van Rijn P. Collagen Morphology Influences Macrophage Shape and Marker Expression in Vitro. J. Immunol. Regen. Med. 2018, 1, 13–20. 10.1016/j.regen.2018.01.002. [DOI] [Google Scholar]
- Gilpin S. E.; Li Q.; Evangelista-Leite D.; Ren X.; Reinhardt D. P.; Frey B. L.; Ott H. C. Fibrillin-2 and Tenascin-C Bridge the Age Gap in Lung Epithelial Regeneration. Biomaterials 2017, 140, 212–219. 10.1016/j.biomaterials.2017.06.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.






