Abstract
Transcriptional enhanced associate domain (TEAD) proteins together with their transcriptional coactivator yes-associated protein (YAP) and transcriptional coactivator with the PDZ-binding motif (TAZ) are important transcription factors and cofactors that regulate gene expression in the Hippo pathway. In mammals, the TEAD families have four homologues: TEAD1 (TEF-1), TEAD2 (TEF-4), TEAD3 (TEF-5), and TEAD4 (TEF-3). Aberrant expression and hyperactivation of TEAD/YAP signaling have been implicated in a variety of malignancies. Recently, TEADs were recognized as being palmitoylated in cells, and the lipophilic palmitate pocket has been successfully targeted by both covalent and noncovalent ligands. In this report, we present the medicinal chemistry effort to develop MYF-03–176 (compound 22) as a selective, cysteine-covalent TEAD inhibitor. MYF-03–176 (compound 22) significantly inhibits TEAD-regulated gene expression and proliferation of the cell lines with TEAD dependence including those derived from mesothelioma and liposarcoma.
Graphical Abstract

INTRODUCTION
The Hippo pathway is an evolutionarily conserved signaling cascade with >30 components that plays a pivotal role in organ size control, tissue homeostasis, stem cell renewal, cell proliferation, angiogenesis, and tumorigenesis.1 Dysregulation of the Hippo pathway through merlin/neurofibromin-2 (NF2) loss, large tumor suppressor kinase 1 (LATS1) fusion, yes- associated protein (YAP)/TAZ fusions, and YAP/TAZ amplification has been tightly linked to the occurrence and progression of tumor malignancies including mesothelioma, meningioma, lung cancer, liver cancer, and other solid tumors.2 Activation of the YAP signaling pathway has also been frequently observed as a resistance mechanism to targeted therapies. For example, in EGFR-mutant lung cancer, treatment with osimertinib and trametinib induces senescence-like tumor dormancy via transcriptional enhanced associate domain (TEAD)/YAP corepressor function.3 Pharmacological inhibition of TEAD/YAP by small molecules or through genetic depletion of YAP enhances apoptosis and reduces the number of dormant cells.3 In ER-positive cancers, TEAD/YAP-mediated overexpression of CDK6 in the absence of FAT1 leads to the resistance to a CDK4/6 inhibitor.4 Similarly, emerging evidence suggests that activation of TEAD/YAP is associated with tumor relapse in pancreatic cancer when KRAS function is inhibited.5
Although pharmacological regulation of the Hippo pathway has significant therapeutic potential, direct targeting of the Hippo pathway has been difficult. The functions of core kinases involved in the Hippo pathway are mainly as tumor suppressors such as MST1/2 and LATS1/2 or scaffolding proteins such as salvador homologue 1 and Mps1 binding protein, and hence it remains challenging to develop small molecules that activate those kinases. Alternatively, targeting TEAD-YAP protein−protein interface with peptides or small molecules has been explored.6 Indeed, recent reports on TEAD/YAP interface disruptors such as CPD3.1,7 compound 6,8 and IAG9339 demonstrates the potential of this strategy and offers an inspiration for medicinal chemists to target TEAD/YAP interfaces.
Interestingly, a reversible post-translational modification, palmitoylation on TEAD, has been identified to regulate TEAD stability and function, which represents a potential “Achilles heel” in TEAD-YAP/TAZ dominated maligancies.10 Therefore, there is growing enthusiasm in both industry and academia to develop TEAD palmitate-binding pocket (PBP) inhibitors as an alternate way to modulate TEAD-YAP signaling. A deluge of TEAD investigational inhibitors has been disclosed including VT-107,11 MGH-CP-1,12 TM2,13 compound 2,14 and MSC-410615 and cysteine-directing covalent inhibitors such as TED-347,16 DC-TEADin02,17 DC-TEAD3in03,18 kojic acid analogues,19 MYF-01–037,3 and K-97520 (Figure 1). Moreover, three TEAD inhibitors IK930, VT3989, and IAG933 have progressed into clinical trials NCT05228015, NCT04665206, and NCT04857372) as monotherapy in subjects with advanced solid tumors. However, very limited information has been disclosed regarding the structure−activity relationships (SARs) required to bind to and inhibit TEAD-dependent transcriptional activity. Also, since YAP-binding domain (YBD) domains in all TEAD isoforms (TEAD1−4) are highly conserved, it may be challenging to engineer isoform-selective TEAD PBP inhibitors which would be needed to investigate TEAD isoform selective pharmacology. Here, we detailed our medicinal chemistry efforts toward the development of a series of Y-shaped TEAD covalent inhibitors. We also biochemically investigated TEAD isoform selectivity among this series of compounds and validated TEAD dependency in liposarcoma by using MYF-03–176 (compound 22), which provides impetus to further TEAD-directed therapeutics for the treatment for this cancer.
Figure 1.

Selective examples of TEAD PBP inhibitors.
RESULTS AND DISCUSSION
Previously, we reported MYF-01–037 as a suboptimal covalent TEAD binder identified from MS-based screening with covalent fragments using TEAD2 YAP-binding domain protein.3 With MYF-01–037 as a starting point, we conducted covalent docking into the palmitoyl pocket of TEAD2, which enabled us to identify a hydrophilic pocket adjacent to the lipid pocket.21 Therefore, the modification of MYF-01–037 was mostly focused on introducing an additional substitution to the 4-carbon of the pyrrolidine ring to generate Y-shaped molecules that could gain access to this hydrophilic pocket (Figure 2). To experimentally verify the docking predictions, we solved a cocrystal of TEAD1209−424 with one of MYF-01–037 analogues, namely 1 (MYF-03–69, PDB ID: 7LI5). The crystal structure confirmed that the acrylamide warhead forms a covalent bond with Cys359 (TEAD1) and that the carbonyl interacts with backbone-NH of Cys359 through a hydrogen bond; the para-trifluoromethyl benzyl group is deeply buried inside the palmitoyl pocket, thereby forming extensive interactions with the surrounding residues F239, L294, L369, and L382; the 3-pyridinyl group faces toward F221 and the pyridinyl nitrogen interacts through a hydrogen bond with S310 and Y312 which is mediated by a water molecule22 (Figure 2). This binding mode allowed us to design a series of analogues to construct a comprehensive SAR with respect to TEAD binding.
Figure 2.
Evolution of MYF-01–037. The binding conformation of 1 in complex with TEAD1-YBD was retrieved from the PDB database (PDB ID: 7LI5). Hydrogen bonds are highlighted with yellow dashes. 1 is depicted in green stick.
To test this series of compounds, we established a TR-FRET biochemical assay for all TEADs using a FAM-labeled probe (WZJ-10) which was built off a known reversible pan-TEAD inhibitor VT-10711 (Figures 3A,B, S1A, S2, and S3). Conjugation of FAM to VT-107 decreased the binding affinity as evidenced by TEAD4 gel-based ABPP assays (Figure S1B). Second, we developed a TEAD reporter gene assay employing a pan-TEAD response element in the promotor region to drive luciferase expression in NCI-H226 cells to investigate whether the biochemical activity correlated with the TEAD-dependent transcription (Figure S4A,B). Finally, the active compounds from TR-FRET and reporter gene assays were progressed to an antiproliferation assay with mesothelioma cells which are known to depend on the TEAD-YAP activity.
Figure 3.
(A) Schematic representation of TEAD TR-FRET assays. The figure was created with BioRender.com. (B) Representative curve of inhibitory activities of VT107 in the established TR-FRET biochemical assay. IC50 values are reported as mean ± SEM of two experiments. Each experiment has three independent biological replicates.
We first tested 1 using the TEAD1–4 TR-FRET assay in which we picked a fixed time point of 5 h to evaluate and rank the compounds. As shown in Table 1, 1 displays a low nanomolar IC50s for all TEADs and an EC50 of 106 ± 30 nM in the TEAD reporter assay. As expected, introducing a polar group to this section was much less favored, and compounds 2 and 3 lost 6- and 21-fold activities, respectively. Disubstitution on the phenyl ring with Cl and CF3 (compounds 4 and 5) does not increase the hydrophobic interaction, and on the contrary, it impairs the binding, indicating that the hydrophobic pocket is likely to be narrow and linear. Finally, replacing the phenyl ring with aliphatic rings such as gem-difluorocyclohexane (compound 6) and adamantane (compound 7) suggests that TEAD2 and TEAD3 could be selectively inhibited than other TEADs, although its cellular potency is less than that observed for 1, which might be due to reduced cell permeability.
Table 1.
Substitution in the hydrophilic pocket

|
IC50 values are reported as mean ± SEM of two experiments. Each experiment has three independent biological replicates. NE means no effect (IC50 > 10,000 nM).
We next fixed the trifluoromethyl benzyl group and turned toward the modification to the hydrophilic site to understand if a favorable engagement in this tunnel could improve activity (Table 2). Inspired by the cocrystal structure where the pyridine group on 1 forms a water-bridged hydrogen bond network with the surrounding residues, we then questioned if the pyrimidine substituent could also retain or even improve this interaction. Compound 8 was synthesized, which indeed demonstrates a negative response to TEAD4 inhibition while not having much impact on other TEADs and the reporter gene assay. Interestingly, a truncated triazole analogue (compound 9) does not lose any potency, indicating that the water-mediated interaction observed in the cocrystal with TEAD1-YBD might not exist or has a minimal contribution to inhibition. Further design was focused on screening a small panel of polar groups, as depicted in Table 2. In general, this hydrophilic pocket has a high tolerance for a variety of functional groups including triazole (compound 9), pyrazole (compound 10), pyridazine-3-one (compound 11), 4-methyl- sulfonyl benzyl group (compound 12), methylene-pyridine (compound 13), and pyrimidin-2-amine (compound 14). Of note, the potency was significantly improved by compound 14 against TEADs and in the reporter gene assay with IC50 values of 14 ± 4 nM, respectively.
Table 2.
Substitution in the hydrophilic pocket

|
IC50 values are reported as mean ± SEM of two experiments. Each experiment has three independent biological replicates.
Next, we expanded the pyrrolidine core to larger rings: piperidine and 1,4-diazeopane and synthesized compounds 16 and 17. As shown in Table 3, compound 15, a fluorinated version of 14 shows a similar potency. Interestingly, a piperidine-based analogue 16 significantly lost activity for TEAD2, 3, and 4 while only slightly diminishing TEAD1 inhibition. A similar trend was observed with 1,4-diazepane- containing compound 17, implying that a selective TEAD1 inhibitor could be designed by tuning the size and/or conformation of the core ring. Of note, in the reporter gene assay, both 16 and 17 still maintain an IC50 of 171 ± 88 and 522 ± 263 nM, respectively, indicating TEAD1 as one of the potential main contributors to TEAD transcription in NCI-H226 cells.
Table 3.
Expansion of the pyrrolidine core

|
IC50 values are reported as mean ± SEM of two experiments. Each experiment has three independent biological replicates. NE means no effect (IC50 > 10000 nM).
Finally, we incorporated a set of covalent warheads with different electrophilic reactivity or conformational accessibility to the cysteine (Table 4). First, we shielded acrylamide by introducing methylene dimethylamine to β-carbon of the double bond, which is known to tune electrophilicity and improve drug properties. Unfortunately, 19 became inactive on all TEADs and did not inhibit TEAD transcription. Replacing acrylamide with propynamide (compound 18) selectively impairs TEAD4 binding while maintaining decent inhibition of the other TEADs, which translates well to cellular inhibition. Inspired by the KRAS covalent inhibitor MRTX849 with a fluorinated acrylamide as warhead, which resulted in a higher resistance to GSH conjugation compared to its nonfluorination version, we designed compound 21.23 Encouragingly, 21 displayed comparable activity with its fluorine-free counterpart for TEAD1–3 and a slight weaker inhibition of TEAD4. Interestingly, the activity on TEAD4 was restored in compound 22, where a smaller amino-pyrimidine group was introduced to the hydrophilic pocket rather than triazolepyrimidine. Moreover, compound 22 exhibits a potent inhibition on TEAD transcription with an IC50 value of 17 ± 5 nM. To further establish that the compounds form a covalent complex with TEAD, the recombinant TEAD1-YBD protein was incubated with 10-fold molar excess of compound 22 for 1 h at rt. We observed a mass shift consistent with compound 22 demonstrating its covalency (Figure S5).
Table 4.
Warhead exploration

|
IC50 values are reported as mean ± SEM of two experiments. Each experiment has three independent biological replicates. NE means no effect (IC50 > 10000 nM).
Viability Assay.
To further investigate whether the observed transcriptional inhibition in the TEAD reporter gene assay can be translated to antiproliferation activity in TEAD-dependent cell lines, we conducted a 5-day proliferation experiment in NCI-H226 mesothelioma cells with selected compounds in reporter assays. As expected, top-ranked TEAD inhibitors from the TEAD reporter assay all displayed a potent inhibition of cell proliferation in NF2-deficient NCI-H226 mesothelioma cells among which compound 14 and compound 22 are best-of-class with IC50 values of 16 and 24 nM, respectively, while they exhibited limited effect in NF2- sufficient NCI-H28 mesothelioma cells (Figures 4A and S6). In addition, we also plotted the IC50s from the reporter assay against the antiproliferation IC50s from the NCI-H226 proliferation assay, which demonstrates a reasonable correlation (Figure 4B).
Figure 4.
(A) Five-day antiproliferation activity of TEAD inhibitors in NCI-H226 cells. Data are presented as mean ± SD for n = 3 biological independent samples. (B) Correlation between antiproliferation IC50 values and 3-day reporter IC50 values.
Finally, to understand TEAD-YAP dependency in a larger panel of cancer cells, we performed PRISM screening with a DNA-barcoded cancer cell library (930 cell lines) with compound 22.24 The area under the curve (AUC) was plotted against YAP1 dependency from DepMap (https://depmap.org/). Encouragingly, mesothelioma cells (MSTO-211H, NCI-H2052), well-differentiated liposarcoma (94T778 and 95T1000), and liver cancer SKHEP-1 ranked most highly (Figure 5A,B). Correlation analysis demonstrated that the knockout of TEAD1, YAP1, and WWTR1 (TAZ) is positively correlated with the compound cytotoxicity profile, while NF2, TAOK1, LATS2, and YAP-binding partner angiomotin like 2 (AMOTL2), which are well-characterized negative regulators on TEAD/YAP activity in the Hippo pathway, are negatively correlated (Figure 5C, Table S1).25 Pearson correlation analysis with portal data (drug sensitivity PRISM repurposing primary screen) indicates that compound 22 has a high correlation of XAV939, a known tankyrase inhibitor that can suppress YAP activity by the stabilization of AMOT family proteins26 (Table S2).
Figure 5.
PRISM study of 22. (A) Compound AUC was plotted against YAP1 dependency from DepMap database. (B) Most sensitive cancer cell lines (Top 10) in the PRISM study ranked by AUC. (C) Dependency correlation with the CRISPR KO data set from DepMap.
To confirm compound activity in additional lineages identified from PRISM, we choose 94T778 liposarcoma cells for viability assays (Figures 6A and S6). A similar response was observed with high IC50 correlation (R-square of 0.62) between two TEADi-sensitive cell lines (Figure 6A,B). In addition, according to the DepMap database, the gene dependency scores for TEAD1/2/3/4 in 94TT78 are −1.61, 0.05, −0.36, and −0.32, respectively, which suggests that TEAD1 inhibition may be the primary driver of compound efficacy. Collectively, these PRISM results support 22 as a highly selective TEAD inhibitor with a high therapeutic potential in TEAD-driven malignancies.
Figure 6.
(A) Five-day antiproliferation activity of TEAD inhibitors in 94T778 cells. Data are presented as mean ± SD of n = 3 biological independent samples. (B) Correlation between antiproliferation IC50 values in mesothelioma cells NCI-H226 and liposarcoma cells 94T778. IC50 curves for other compounds are provided in the Supporting Information.
CONCLUSIONS
TEAD has emerged as a druggable target spurring extensive medicinal chemistry efforts to develop TEAD inhibitors, and two compounds have entered clinical evaluation. Our approach to inhibit TEAD is through covalent engagement of a palmitoylated cysteine. We first employed mass spectrometry-enabled protein labeling to screen covalent fragment library and identified MYF-01–037 as the lead compound. Lead optimization advanced MYF-01–037 as a linear molecule that can occupy the lipid-binding pocket to a series of Y-shape molecules that exploit an adjacent hydrophilic pocket. TR-FRET and TEAD reporter assays demonstrate that overall the Y-shaped scaffold improves the activity to an IC50 below 50 nM. Interestingly, TEAD isoforms respond differently to certain modifications. For example, TEAD2 has a higher tolerance for aliphatic rings than the other TEADs, as exemplified by compounds 6 and 7. In contrast, TEAD1 accommodates a larger aliphatic heterocycle than pyrrolidine and thus exhibits greater sensitivity to compounds 16 and 17 than TEAD2/3/4. Our chemistry optimization eventually resulted in the development of a potent pan-TEAD inhibitor 22 (MYF-03–176). We show that MYF-03–176 inhibits TEAD in biochemical assays, diminishes TEAD-coordinate transcriptional responses in a gene reporter assay, and has a potent effect on cell viability in TEAD-dependent mesothelioma cells. Barcoded cell line screens identified additional indications including liposarcoma and liver cancers. In addition, 22 and its analogues also provide a chemistry starting point to develop an isoform-specific inhibitor, which might have a wider therapeutic window than a pan-TEAD inhibitor.
Chemistry.
Synthetic routes for the preparation of compounds are outlined in Schemes 1−6. Commercially available tert-butyl 6-oxa-3-azabicyclo[3.1.0]hexane-3-carboxylate S1 underwent epoxide ring opening reaction (either in a racemic way with sodium azide as the nucleophile or in an asymmetric way with TMSN3 as the nucleophile and chromium complex as the chiral auxiliary) to afford intermediate S2. Subsequent copper-catalyzed click chemistry with 3-ethynylpyridine gave intermediate S3. Nucleophilic substitution reaction between S3 and various haloalkanes or alkyl triflate provided intermediate S4, which was followed by deprotection to afford intermediate S5. Coupling of S5 with acryloyl chloride provided the corresponding final compounds (±)-2−7 (Scheme 1). Compounds (±)-1, 8, and 9 were prepared via a slightly different route (Scheme 2). SN2 reaction of S2 with 1-(bromomethyl)-4-(trifluoromethyl)benzene afforded intermediate S6. Click chemistry of S6 with various ethynylarenes afforded intermediate S7, and subsequent deprotection and installation of acrylamide warhead resulted in compounds (±)-1, 8 and 9. Similarly, the ring opening reaction of epoxide with other nucleophilic reagents gave intermediate S9, which enabled us to synthesize compounds (±)-10—13 (Scheme 3). Reduction of azide S6 by using PPh3 provided amine S11. Nucleophilic aromatic substitution of S11 with 2-bromopyrimidine or 2-bromo-5-fluoropyrimidine afforded S12, following deprotection and installation of the warhead, providing compounds (±)-14 and 15. Compound (±)-16 was prepared from the starting material tert-butyl 7- oxa-3-azabicyclo[4.1.0]heptane-3-carboxylate S13 via a similar procedure with (±)-15 (Scheme 4). In particular, the synthesis of compound (±)-17 started with the PMB protection of ethane-1,2-diamine S17. Subsequent cyclization and deprotection of PMB led to intermediate S20. Further, the 2-step reaction afforded final product (±)-17 (Scheme 5). Finally, to explore the effect of various Cys-reactive warheads, compounds (3R,4R)-18−22 were synthesized via coupling of key optically pure intermediates S8–1 or S12–2 with various warhead precursors (Scheme 6).
Scheme 1.
Synthesis of Compounds (±)-2−7a
aReagents and conditions: (a) NaN3, NH4Cl, MeOH, 50−60 °C, 60% yield, or TMSN3, (1S,2S)-(+)-[1,2-cyclohexanediamino-N,N′-bis(3,5-di-t-butylsalicylidene)]chromium(III) chloride, K2CO3, MeOH, rt 71% yield. (b) 3-Ethynylpyridine, CuSO4, sodium L-ascorbate, THF, BuOH, H2O, 70 °C, 45% yield; (c) haloalkane or alkyl triflate, NaH, DMF, 0 °C; (d) TFA, DCM, rt >90% yield. (e) Acryloyl chloride, Et3N, DCM, 0 °C.
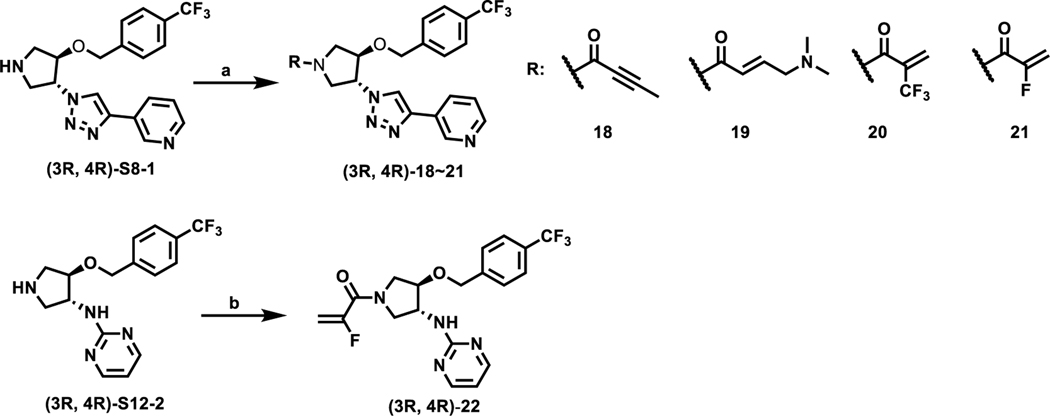
Scheme 6.
Synthesis of Compounds (3R, 4R)-18−22a
aReagents and conditions: (a) acid or acyl chloride, coupling condition; (b) 2-fluoroacrylic acid, HATU, DIPEA, DMF, rt.
Scheme 2.
Synthesis of Compounds (±)-1,8 and 9a
aReagents and conditions: (a) 1-(bromomethyl)-4-(trifluoromethyl)benzene, NaH, DMF, 0 °C; (b) ethynylarene, CuSO4, sodium L-ascorbate, THF, BuOH, H2O, 70 °C; (c) TFA, DCM, rt >90% yield. (d) Acryloyl chloride, Et3N, DCM, 0 °C; (e) TBAF, THF.
Scheme 3.
Synthesis of Compounds (±)-10−13a
aReagents and conditions: (a) nucleophile, base, solvent, 0 °C; (b) 1-(bromomethyl)-4-(trifluoromethyl)benzene, NaH, DMF, 0 °C; (c) TFA, DCM, rt >90% yield; (d) acryloyl chloride, Et3N, DCM, 0 °C.
Scheme 4.
Synthesis of Compounds (±)-14−16a
aReagents and conditions: (a) PPh3, H2O, THF, 70 °C; (b) TMSN3, (1S,2S)-(+)-[1,2-cyclohexanediamino-N,N′-bis(3,5-di-t-butylsalicylidene)]- chromium(III)chloride, K2CO3, MeOH, rt; (c) 1-(bromomethyl)-4-(trifluoromethyl)benzene, NaH, DMF, 0 °C; (d) DIPEA, BuOH, 70 °C; (e) TFA, DCM, rt >90% yield; (f) acryloyl chloride, Et3N, DCM, 0 °C.
Scheme 5.
Synthesis of Compounds (±)-17a
aReagents and conditions: (a) 4-methoxybenzaldehyde, NaBH4, MeOH; (b) 1,3-dibromopropan-2-ol, Et3N, toluene, 120 °C; (c) 1-(bromomethyl)-4-(trifluoromethyl)benzene, NaH, DMF, 0 °C; (d) TFA, 90 °C; (e) 2-chloropyrimidine, DIPEA, DMSO, 90 °C; (f) Acryloyl chloride, Et3N, DCM, 0 °C.
EXPERIMENTAL SECTION
Compound Synthesis.
Unless otherwise noted, reagents and solvents were obtained from commercial suppliers and were used without further purification. 1H NMR and 13C NMR spectra were recorded on a 500 MHz (Bruker A500) spectrometer, and chemical shifts are reported in parts per million (ppm, d) downfield from tetramethylsilane (TMS). Coupling constants (J) are reported in Hz. Spin multiplicities are described as s (singlet), br (broad singlet), d (doublet), t (triplet), q (quartet), and m (multiplet). Mass spectra were obtained on a Waters Micromass ZQ instrument. Preparative HPLC was performed on a Waters Sunfire C18 column (19 × 50 mm, 5 μM) using a gradient of 15−95% methanol in water containing 0.05% trifluoroacetic acid (TFA) over 22 min (28 min run time) at a flow rate of 20 mL/min. Purities of assayed compounds were in all cases greater than 95%, as determined by reverse-phase HPLC analysis. The HPLC traces have been provided in Figure S7.
General Procedure A for the Synthesis of Compounds 2−7.
Step 1: Racemic way: A mixture of S1 (10 g, 54 mmol), NaN3 (7 g, 108 mmol), and NH4Cl (2.8 g, 54 mmol) in MeOH (120 mL) and H2O (20 mL) was stirred at 60 °C under N2 overnight. The reaction mixture was concentrated in vacuum, and the residue was extracted with ethyl acetate (300 mL × 3). The combined organic was washed with water (200 mL × 2), dried over anhydrous Na2SO4, and concentrated and purified by flash column chromatography on silica gel (ethyl acetate in petroleum ether = 20% v/v) to obtain racemic S2 (11 g) as a oil.
Asymmetric way: A mixture of S1 (4 g, 21.6 mmol), TMSN3 (2.664 g, 23.2 mmol), and chiral catalyst (1S,2S)-(−)-[1,2-cyclo- hexanediamino-N,N′-bis(3,5-di-t-butylsalicylidene)]chromium(III) chloride (328 mg, 0.42 mmol) was stirred at rt under N2 overnight. The reaction mixture was treated with MeOH (60 mL) and K2CO3 (1.788 g, 12.8 mmol), and stirring was continued at rt for 5 h. The reaction mixture was diluted with ethyl acetate (300 mL), washed with water (300 mL × 2), dried over anhydrous Na2SO4, and concentrated and purified by flash column chromatography on silica gel (ethyl acetate in petroleum ether = 20% v/v) to obtain S2 (3.5 g, 96% ee) as a clear oil.
Step 2: A mixture of S2 (1000 mg, 4.38 mmol), 3-ethynylpyridine (451 mg, 4.38 mmol), CuSO4 (654 mg, 2.6 mmol), and sodium L-ascorbate (257 mg, 1.3 mmol) in THF (3 mL), H2O (3 mL), and nBuOH (3 mL) was stirred at 70 °C overnight. The mixture was diluted with water (50 mL) and extracted with ethyl acetate (60 mL). The extract was washed with water (40 mL), dried over anhydrous Na2SO4, and concentrated and purified by flash column chromatography on silica gel (ethyl acetate in petroleum ether = 80% v/v) to obtain S3 as a yellow solid (650 mg). LC−MS (ESI) m/z: 332 [M + H]+.
Step 3: To a mixture of S3 (100 mg, 0.3 mmol) in DMF (10 mL) was added NaH (14 mg, 0.33 mmol). The mixture was stirred at rt for 10 min, and then haloalkane (0.3 mmol) was added. The mixture was stirred at rt under N2 overnight. The mixture was diluted with water (50 mL) and extracted with ethyl acetate (60 mL × 2). The extract was washed with water (40 mL), dried over anhydrous Na2SO4, and concentrated and purified by flash column chromatography on silica gel to obtain intermediate S4 as a solid.
Step 4: A mixture of S4 (0.1 mmol) and TFA (1 mL) in DCM (3 mL) was stirred at rt for 2 h. The mixture was concentrated to afford crude intermediate S5 which was used in the next step without further purification.
Step 5: To a mixture of S5 (0.1 mmol) and Et3N (0.2 mmol) in DCM (5 mL) was added acryloyl chloride (0.1 mmol). The mixture was stirred at 0 °C for 2 h and then concentrated and purified by prep-HPLC to obtain the final compounds.
General Procedure B for the Synthesis of Compounds 1, 8, and 9.
Step 1: A mixture of S2 (3 g, 13.1 mmol), 1-(bromomethyl)-4-(trifluoromethyl)benzene (3.1 g, 13.1 mmol), and 60% NaH (0.6 g, 15.7 mmol) in DMF (20 mL) was stirred at rt under N2 protection for 6 h. The reaction mixture was diluted with water (100 mL) and extracted with ethyl acetate (100 mL × 2). The combined organic was washed with water (100 mL), dried over anhydrous Na2SO4, and concentrated and purified by flash column chromatography on silica gel (ethyl acetate in petroleum ether = 30% v/v) to obtain S6 (3.8 g) as an oil.
Step 2: A mixture of S6 (100 mg, 0.44 mmol), 3-ethynylpyridine (45 mg, 0.44 mmol), CuSO4 (65 mg, 0.25 mmol), and sodium L-ascorbate (25 mg, 0.13 mmol) in THF (0.5 mL), H2O (0.5 mL), and nBuOH (0.5 mL) was stirred at 70 °C overnight. The mixture was diluted with water (10 mL) and extracted with ethyl acetate (20 mL × 2). The extract was washed with water (20 mL), dried over anhydrous Na2SO4, and concentrated and purified by flash column chromatog- raphy on silica gel to obtain S7.
Step 3: A mixture of S7 (0.1 mmol) and TFA (1 mL) in DCM (3 mL) was stirred at rt for 2 h. The mixture was concentrated to give crude intermediate S8 which was used in the next step without further purification.
Step 4: To a mixture of S8 (0.1 mmol) and Et3N (0.2 mmol) in DCM (5 mL) was added acryloyl chloride (0.1 mmol) at 0 °C. The mixture was stirred at rt for 2 h and then concentrated and purified by prep-HPLC to obtain the final compounds.
General Procedure C for the Synthesis of Compounds 10—13.
Step 1: To a suspension of 60% NaH (32 mg, 0.8 mmol) in THF (2 mL) was added the solution of nucleophile compound (0.8 mmol) in THF (1 mL). The mixture was stirred at 0 °C under N2 for 30 min, and then S1 (100 mg, 0.54 mmol) was added. The resulting mixture was heated at 60 °C under N2 for 16 h. After cooling down to rt, the mixture was diluted with ethyl acetate (10 mL × 2) and washed with water (10 mL). The organic layer was dried over anhydrous Na2SO4, filtered, and concentrated and purified by flash column chromatog- raphy on silica gel to afford intermediate S9.
Step 2: A mixture of S9 (0.2 mmol), 1-(bromomethyl)-4-(trifluoromethyl)benzene (0.2 mmol), and 60% NaH (0.3 mmol) in DMF (1 mL) was stirred at rt under N2 protection for 6 h. The reaction mixture was diluted with water (10 mL) and extracted with ethyl acetate (10 mL × 2). The combined organic was washed with water (10 mL), dried over anhydrous Na2SO4, and concentrated and purified by flash column chromatography on silica gel to afford intermediate S10.
Step 3: A mixture of S10 (0.1 mmol) and TFA (1 mL) in DCM (3 mL) was stirred at rt for 2 h. The mixture was concentrated to give the crude intermediate which was dissolved in DCM (5 mL), and then Et3N (0.2 mmol) and acryloyl chloride (0.1 mmol) were added at 0 °C. The resulting mixture was stirred at rt for 2 h and then concentrated and purified by prep-HPLC to obtain the final compounds.
General Procedure D for the Synthesis of Compounds 14−16.
S15 was prepared starting from S13 following a similar racemic procedure as S6.
Step 1: A mixture of compound S6 (300 mg, 0.77 mmol), PPh3 (240 mg, 0.93 mmol), and H2O (280 mg, 0.15 mmol) in THF (5 mL) was stirred at 70 °C under N2 for 5 h. The mixture was diluted with water (100 mL) and extracted with ethyl acetate (100 mL × 2). The organic was washed with water (100 mL), dried over anhydrous Na2SO4, and concentrated and purified by prep-HPLC to obtain intermediate S11 as a yellow oil.
Step 2: A mixture of S11 (0.4 mmol), 2-chloropyrimidine or 2-chloro-5-fluoropyrimidine (0.4 mmol), and DIPEA (1.2 mmol) in BuOH (3 mL) was stirred at 70 °C under N2 overnight. The mixture was concentrated and purified by prep-HPLC to obtain compound S12 as an oil.
Step 3: A mixture of S12 (0.1 mmol) and TFA (1 mL) in DCM (3 mL) was stirred at rt for 2 h. The mixture was concentrated to give the crude intermediate which was dissolved in DCM (5 mL), and then Et3N (0.2 mmol) and acryloyl chloride (0.1 mmol) were added. The resulting mixture was stirred at rt for 2 h and then concentrated and purified by prep-HPLC to obtain the final compounds.
1-(trans-3-(4-(Pyridin-3-yl)-1H-1,2,3-triazol-1-yl)-4-((4-(trifluoromethyl)benzyl)oxy)pyrrolidin-1-yl)prop-2-en-1-one (Compound (±)-1).
The title compound was prepared according to the general procedure B.
1H NMR (400 MHz, CD3OD): δ (ppm) 8.91 (d, J = 1.8 Hz, 1 H), 8.50 (d, J = 4.9 Hz, 1 H), 8.43 (d, J = 4.9 Hz, 1 H), 8.18 (dt, J = 8.0, 1.8 Hz, 1 H), 7.51 (d, J = 8.0 Hz, 2 H), 7.48−7.38 (m, 3 H), 6.60−6.49 (m, 1 H), 6.23 (dd, J = 16.8, 1.9 Hz, 1 H), 5.75−5.67 (m, 1 H), 5.41−5.30 (m, 1 H), 4.68 (d, J = 4.0 Hz, 2 H), 4.60−4.49 (m, 1 H), 4.32−4.16 (m, 1 H), 4.15−3.87 (m, 2 H), 3.85−3.63 (m, 1 H). 13C NMR (126 MHz, DMSO-d6): δ 164.2, 149.5, 146.9, 144.3, 143.0, 132.9, 129.4, 128.5, 128.3 (q, J = 32.5 Hz), 127.9, 126.9, 125.6, 124.5, 124.4 (q, J = 272.2 Hz), 122.2, 81.1, 70.5, 62.6, 50.2, 49.5. LC−MS (ESI) m/z: 444 [M + H]+.
1-(trans-3-(4-(Pyridin-3-yl)-1H-1,2,3-triazol-1-yl)-4-((5-(trifluoromethyl)pyridin-2-yl)methoxy)pyrrolidin-1-yl)prop-2-en-1-one (Compound (±)-2).
The title compound was prepared according to the general procedure A.
1H NMR (400 MHz, CD3OD): δ (ppm) 9.03 (d, J = 1.8 Hz, 1H), 8.82 (s, 1H), 8.65 (d, J = 6.2 Hz, 1H), 8.58−8.51 (m, 1H), 8.30 (d, J = 8.0 Hz, 1H), 8.13 (dd, J = 8.2, 1.9 Hz, 1H), 7.70 (d, J = 8.2 Hz, 1H), 7.54 (dd, J = 7.9, 4.9 Hz, 1H), 6.68 (dd, J = 16.8, 10.4 Hz, 1H), 6.36 (dd, J = 16.8, 1.4 Hz, 1H), 5.82 (ddd, J = 10.5, 3.7, 1.9 Hz, 1H), 5.58−5.47 (m, 1H), 4.91 (d, J = 3.5 Hz, 2H), 4.82−4.72 (m, 1H), 4.48−3.82 (m, 4H). 13C NMR (126 MHz, DMSO-d6): δ 164.2, 162.7, 149.5, 146.9, 146.7, 144.3, 134.8, 132.9, 129.4, 128.0, 126.9, 125.1 (q, J = 33.4 Hz), 124.1, 124.0 (q, J = 272.2 Hz), 122.3 (d, J = 4.1 Hz), 121.8, 81.53, 71.87, 62.59, 50.29, 49.50. LC−MS (ESI) m/z: 445 [M + H]+.
1-(trans-3-(4-(Pyridin-3-yl)-1H-1,2,3-triazol-1-yl)-4-((6-(trifluoromethyl)pyridin-3-yl)methoxy)pyrrolidin-1-yl)prop-2-en-1-one (Compound (±)-3).
The title compound was prepared according to the general procedure A.
1H NMR (400 MHz, CD3OD): δ (ppm) 9.04 (d, J = 2.2 Hz, 1H), 8.76−8.50 (m, 3H), 8.31 (dt, J = 7.9, 1.9 Hz, 1H), 8.03 (d, J = 8.2 Hz, 1H), 7.80 (d, J = 8.1 Hz, 1H), 7.56 (dd, J = 8.0, 4.9 Hz, 1H), 6.67 (ddd, J = 16.8, 10.4, 3.5 Hz, 1H), 6.36 (dd, J = 16.8, 1.7 Hz, 1H), 5.83 (ddd, J = 10.5, 5.1, 1.9 Hz, 1H), 5.58−5.44 (m, 1H), 4.97 (s, 2H), 4.81−4.68 (m, 1H), 4.48−4.03 (m, 3H), 4.02−3.80 (m, 1H). 13C NMR (126 MHz, DMSO-d6): δ 164.2, 149.7, 149.5, 146.9, 146.2 (q, J = 33.5 Hz), 144.3, 137.8, 137.7, 132.9, 129.4, 128.0, 126.9 (d, J = 2.8 Hz), 124.5, 122.2, 122.2 (q, J = 272.2 Hz), 120.9, 81.2, 68.3, 62.6, 50.2, 49.5. LC−MS (ESI) m/z: 445 [M + H]+.
1-(trans-3-(3-Chloro-5-(trifluoromethyl)benzyloxy)-4-(4-(pyridine-3-yl)-1H-1,2,3-triazol-1-yl)pyrrolidin-1-yl)prop-2-en-1-one (Compound (±)-4).
The title compound was prepared according to the general procedure A.
1H NMR (400 MHz, CD3OD): δ (ppm) 9.03 (d, J = 1.1 Hz, 1H), 8.62 (d, J = 4.9 Hz, 1H), 8.55 (d, J = 4.9 Hz, 1H), 8.31 (dt, J = 8.0, 1.8 Hz, 1H), 7.68−7.51 (m, 4H), 6.67 (ddd, J = 16.7, 10.4, 4.4 Hz, 1H), 6.36 (d, J = 16.8 Hz, 1H), 5.83 (ddd, J = 10.4, 5.6, 1.9 Hz, 1H), 5.56−5.40 (m, 1H), 4.82−4.63 (m, 3H), 4.40−4.05 (m, 3H), 3.97− 3.76 (m, 1H). 13C NMR (126 MHz, DMSO-d6): δ 164.2, 149.5, 146.9, 144.3, 142.3, 134.5, 132.9, 131.8, 131.3 (q, J = 33.5 Hz), 129.4, 127.9 (d, J = 2.3 Hz), 126.9 (d, J = 3.2 Hz), 124.8, 124.5, 123.8 (q, J = 272.2 Hz), 123.3, 122.1, 81.0, 69.8, 62.6, 50.1, 49.4. LC−MS (ESI) m/z: 478 [M + H]+.
1-(trans-3-(2-Chloro-5-(trifluoromethyl)benzyloxy)-4-(4-(pyridine-3-yl)-1H-1,2,3-triazol-1-yl)pyrrolidin-1-yl)prop-2-en-1-one (Compound (±)-5).
The title compound was prepared according to the general procedure A.
1H NMR (400 MHz, CD3OD): δ (ppm) 9.03−8.98 (m, 1H), 8.64 (d, J = 5.2 Hz, 1H), 8.55−8.50 (m, 1H), 8.31−8.26 (m, 1H), 7.80 (s, 1H), 7.61 (s, 2H), 7.55−7.51 (m, 1H), 6.70−6.61 (m, 1H), 6.37−6.30 (m, 1H), 5.83−5.77 (m, 1H), 5.57−5.45 (m, 1H), 4.86 (s, 2H), 4.79−4.68 (m, 1H), 4.42−3.80 (m, 4H). 13C NMR (126 MHz, DMSO-d6): δ 164.2, 149.5, 146.9, 144.3, 137.1, 137.0, 132.9, 130.7, 129.4, 129.3, 128.5 (q, J = 33.5 Hz), 127.9, 126.9, 126.6, 126.4, 124.4 (d, J = 2.3 Hz), 124.2 (q, J = 272.2 Hz), 122.2 (d, J = 3.3 Hz), 81.4, 68.3, 62.6, 50.1, 49.5. LC−MS (ESI) m/z: 478 [M + H]+.
1-(trans-3-((4,4-Difluorocyclohexyl)methoxy)-4-(4-(pyridin-3-yl)-1H-1,2,3-triazol-1-yl)pyrrolidin-1-yl)prop-2-en-1-one (Compound (±)-6).
The title compound was prepared according to the general procedure A.
1H NMR (400 MHz, CD3OD): δ (ppm) 8.91 (d, J = 1.2 Hz, 1H), 8.55−8.40 (m, 2H), 8.18 (dt, J = 8.0, 1.9 Hz, 1H), 7.43 (dd, J = 8.0, 4.9 Hz, 1H), 6.55 (ddd, J = 16.8, 10.4, 2.5 Hz, 1H), 6.28−6.20 (m, 1H), 5.70 (ddd, J = 10.5, 4.7, 1.9 Hz, 1H), 5.29−5.18 (m, 1H), 4.43−4.32 (m, 1H), 4.27−4.16 (m, 1H), 4.07−3.37 (m, 5H), 1.91 (ddd, J = 13.9, 7.0, 3.5 Hz, 2H), 1.76−1.54 (m, 5H), 1.26−1.13 (m, 2H). 13C NMR (126 MHz, DMSO-d6): δ 164.2, 149.5, 146.9, 144.2, 132.9, 129.5, 127.9, 126.9, 124.8 (t, J = 240.6 Hz), 124.5, 122.3, 81.2, 73.6, 62.5, 50.3, 49.4, 35.5, 32.8 (t, J = 23.8 Hz), 25.8 (d, J = 9.6 Hz). LC− MS (ESI) m/z: 418 [M + H]+.
1-(trans-3-((Adamantan-1-yl)methoxy)-4-(4-(pyridin-3-yl)-1H-1,2,3-triazol-1-yl)pyrrolidin-1-yl)prop-2-en-1-one (Compound (±)-7).
The title compound was prepared according to the general procedure A.
1H NMR (400 MHz, CD3OD): δ (ppm) 9.17 (d, J = 1.2 Hz, 1H), 8.85 (d, J = 5.0 Hz, 1H), 8.76 (d, J = 5.0 Hz, 1H), 8.12 (dd, J = 4.4, 1.2 Hz, 1H), 7.63 (d, J = 8.1 Hz, 2H), 7.53 (d, J = 8.1 Hz, 2H), 6.66 (dd, J = 16.8, 10.4 Hz, 1H), 6.35 (d, J = 16.8 Hz, 1H), 5.82 (ddd, J = 10.4, 5.4, 1.9 Hz, 1H), 5.60−5.44 (m, 1H), 4.79 (d, J = 3.4 Hz, 2H), 4.73−4.62 (m, 1H), 4.46−3.74 (m, 4H). 13C NMR (126 MHz, DMSO-d6): δ 164.2, 159.4, 158.7, 156.8, 145.7, 143.0, 129.4, 128.5, 128.3 (q, J = 32.5 Hz), 128.0, 125.6 (d, J = 4.4 Hz), 125.5, 124.4 (q, J = 272.2 Hz), 116.8, 80.9, 70.5, 62.8, 50.3, 49.4. LC−MS (ESI) m/z: 445 [M + H]+.
1-((trans)-3-(1H-1,2,3-Triazol-1-yl)-4-((4-(trifluoromethyl)-benzyl)oxy)pyrrolidin-1-yl)prop-2-en-1-one (Compound (±)-9).
The title compound was prepared according to the general procedure B.
1H NMR (400 MHz, DMSO-d6): δ (ppm) 8.28 (dd, J = 5.6, 1.1 Hz, 1H), 7.80 (dd, J = 3.8, 1.1 Hz, 1H), 7.70 (d, J = 7.9 Hz, 2H), 7.52 (d, J = 8.0 Hz, 2H), 6.61 (ddd, J = 16.8, 10.4, 1.5 Hz, 1H), 6.18 (ddd, J = 16.8, 3.0, 2.3 Hz, 1H), 5.72 (ddd, J = 10.8, 8.7, 2.3 Hz, 1H), 5.44 (ddt, J = 35.9, 7.6, 4.1 Hz, 1H), 4.78−4.67 (m, 2H), 4.52 (ddt, J = 30.3, 5.8, 3.7 Hz, 1H), 4.24 (dd, J = 11.5, 7.0 Hz, 0.5H), 4.11 (dd, J = 11.5, 4.5 Hz, 0.5H), 4.07−3.99 (m, 1H), 3.92 (dd, J = 13.2, 4.1 Hz, 0.5H), 3.87−3.77 (m, 1H), 3.58 (dd, J = 13.2, 3.6 Hz, 0.5H). 13C NMR (126 MHz, DMSO-d6): δ 164.1, 143.0, 134.1, 129.4, 128.7 (q, J = 32.7 Hz), 128.4 (d, J = 3.3 Hz), 127.9, 125.6 (t, J = 4.2 Hz), 125.0, 124.7 (q, J = 272.2 Hz), 80.9, 70.4, 62.1, 50.3, 49.6. LC−MS (ESI) m/z: 367 [M + H]+.
1-(trans)-3-(1H-Pyrazol-1-yl)-4-((4-(trifluoromethyl)benzyloxy)-pyrrolidin-1-yl)prop-2-en-1-one (Compound (±)-10).
The title compound was prepared according to the general procedure C.
1H NMR (400 MHz, CD3OD): δ (ppm) 7.74 (dd, J = 5.2, 2.4 Hz, 1H), 7.63 (d, J = 8.0 Hz, 2H), 7.56 (d, J = 1.6 Hz, 1H), 7.47 (d, J = 8.1 Hz, 2H), 6.66−6.59 (dd, J = 16.8, 10.4 Hz, 1H), 6.36−6.30 (m, 2H), 5.85−5.74 (m, 1H), 5.18−5.02 (m, 1H), 4.68 (d, J = 5.8 Hz, 2H), 4.51−4.42 (m, 1H), 4.28−4.08 (m, 2H), 4.01−3.93 (m, 1H), 3.83−3.65 (m, 1H). 13C NMR (126 MHz, DMSO-d6): δ 164.1, 143.3, 139.9, 130.4, 129.5, 128.5 (q, J = 32.5 Hz), 128.3, 127.7, 125.6 (d, J = 3.1 Hz), 124.4 (q, J = 272.2 Hz), 106.1 (d, J = 5.6 Hz), 81.2, 70.3, 63.2, 55.3, 51.1. LC−MS (ESI) m/z: 366 [M + H]+.
2-(trans-1-Acryloyl-4-((4-(trifluoromethyl)benzyl)oxy)pyrrolidin-3-yl)pyridazin-3(2H)-one (Compound (±)-11).
The title compound was prepared according to the general procedure C.
1H NMR (400 MHz, CD3OD): δ (ppm) 7.94 (dd, J = 3.8, 1.6 Hz, 1H), 7.65 (d, J = 8.0 Hz, 2H), 7.57 (d, J = 7.9 Hz, 2H), 7.50−7.40 (m, 1H), 7.02 (dt, J = 9.5, 1.6 Hz, 1H), 6.64 (ddd, J = 16.8, 12.6, 10.5 Hz, 1H), 6.32 (dd, J = 16.8, 1.5 Hz, 1H), 5.84−5.76 (m, 1H), 5.69−5.59 (m, 1H), 4.82 (dd, J = 19.8, 7.6 Hz, 2H), 4.42−4.29 (m, 1H), 4.24−4.09 (m, 1H), 4.05−3.99 (m, 1H), 3.99−3.83 (m, 1H), 3.77 (d, J = 3.7 Hz, 1H). 13C NMR (126 MHz, DMSO-d6): δ 163.9, 161.5, 147.0, 132.6, 132.5, 128.5 (q, J = 32.5 Hz), 127.1 (d, J = 4.4 Hz), 126.9, 124.4 (q, J = 272.2 Hz), 122.9, 122.4, 117.2, 80.9, 70.4, 62.1, 50.3, 49.6. LC−MS (ESI) m/z: 394 [M + H]+.
1-(trans-3-(4-(Methylsulfonyl)phenyl)-4-(4-(trifluoromethyl)-benzyloxy)pyrrolidin-1-yl)prop-2-en-1-one (Compound (±)-12).
The title compound was prepared according to the general procedure C.
1 H NMR (400 MHz, DMSO-d6): δ (ppm) 7.90 (dd, J = 8.3, 2.6 Hz, 2H), 7.69 (d, J = 8.3, 2.6 Hz, 2H), 7.60 (d, J = 8.4 Hz, 2H), 7.48 (d, J = 8.0 Hz, 2H), 6.69−6.58 (m, 1H), 6.22−6.15 (m, 1H), 5.75−5.67 (m, 1H), 4.66 (d, J = 3.4 Hz, 2H), 4.33−4.22 (m, 1H), 4.13−3.99 (m, 1H), 3.95−3.40 (m, 4 H), 3.21 (s, 3 H). 13C NMR (126 MHz, DMSO-d6): δ 164.0, 146.2, 143.4, 139.9, 129.7, 129.3, 128.9, 128.5 (q, J = 32.5 Hz), 128.3 (d, J = 5.2 Hz), 127.8 (d, J = 2.4 Hz), 125.9, 124.7 (q, J = 272.2 Hz), 82.4, 70.4, 50.6, 49.5, 47.5, 43.9. LC− MS (ESI) m/z: 454 [M + H]+.
1-(trans)-3-(Pyridin-2-ylmethyl)-4-((4-(trifluoromethyl)-benzyloxy)pyrrolidin-1-yl)prop-2-en-1-one (Compound (±)-13).
The title compound was prepared according to the general procedure C.
1H NMR (400 MHz, CD3OD): δ (ppm) 8.55−8.42 (m, 1H), 7.84−7.72 (m, 1H), 7.61 (d, J = 8.1 Hz, 2H), 7.48−7.25 (m, 4H), 6.60 (ddd, J = 16.7, 10.4, 5.0 Hz, 1H), 6.29 (dd, J = 16.8, 1.9 Hz, 1H), 5.76 (ddd, J = 10.4, 4.4, 1.9 Hz, 1H), 4.61 (d, J = 6.2 Hz, 2H), 4.07− 3.40 (m, 5H), 3.01−2.73 (m, 3H). 13C NMR (126 MHz, DMSO-d6): δ 164.1, 159.6, 149.6, 143.6, 136.9, 129.8, 128.5 (q, J = 33.5 Hz), 128.3, 127.3, 125.5 (p, J = 3.6 Hz), 124.7 (q, J = 272.2 Hz), 123.8, 121.9 (d, J = 4.4 Hz), 80.9, 69.6, 50.4, 49.3, 42.0, 39.2. LC−MS (ESI) m/z: 391 [M + H]+.
1-(trans-3-(Pyrimidin-2-ylamino)-4-((4-(trifluoromethyl)benzyl)-oxy)pyrrolidin-1-yl)prop-2-en-1-one (Compound (±)-14).
The title compound was prepared according to the general procedure D.
1H NMR (400 MHz, CD3OD): δ (ppm) 8.33 (d, J = 4.8 Hz, 2H), 7.63 (dd, J = 28.8, 8.2 Hz, 4H), 6.74−6.55 (m, 2H), 6.31 (dd, J = 16.8, 1.9 Hz, 1H), 5.78 (dd, J = 10.4, 1.7 Hz, 1H), 4.85 (t, J = 6.6 Hz, 2H), 4.61 (dd, J = 20.0, 6.0 Hz, 1H), 4.27−4.17 (m, 1H), 4.11−3.67 (m, 4H). 13C NMR (126 MHz, DMSO-d6): δ 164.2, 161.9, 158.4, 143.6, 129.8 (d, J = 5.7 Hz), 128.6 (q, J = 33.5 Hz), 128.2, 127.1, 125.5 (d, J = 3.6 Hz), 124.7 (q, J = 272.2 Hz), 111.2, 80.6, 69.8, 54.6, 50.2, 49.7. LC−MS (ESI) m/z: 393 [M + H]+.
1-(trans-3-((5-Fluoropyrimidin-2-yl)amino)-4-((4-(trifluoromethyl)benzyl)oxy)pyrrolidin-1-yl)prop-2-en-1-one (Compound (±)-15).
The title compound was prepared according to the general procedure D.
1H NMR (400 MHz, DMSO-d6): δ (ppm) 8.41 (d, J = 0.9 Hz, 2H), 7.75−7.52 (m, 5H), 6.56 (ddd, J = 17.2, 10.3, 7.2 Hz, 1H), 6.21−6.08 (m, 1H), 5.67 (ddd, J = 10.3, 4.6, 2.4 Hz, 1H), 4.82−4.68 (m, 2H), 4.37 (dt, J = 18.8, 6.0 Hz, 1H), 4.15−4.01 (m, 1H), 3.87 (ddd, J = 24.3, 11.4, 5.2 Hz, 1H), 3.77−3.60 (m, 2H), 3.60−3.51 (m, 1H). 19F NMR (376 MHz, DMSO-d6): δ 60.90, 60.91, 156.07, 156.08. 19F NMR (376 MHz, DMSO-d6): δ −60.9, −156.1. 13C NMR (126 MHz, DMSO-d6): δ 164.2, 159.1, 152.4 (d, J = 245.2 Hz), 145.9, 143.6 (d, J = 5.1 Hz), 129.9 (d, J = 6.5 Hz), 128.6 (q, J = 33.5 Hz), 128.3, 127.1, 125.5 (q, J = 4.3, 3.9 Hz), 124.7 (q, J = 272.2 Hz), 80.5, 69.7, 55.2, 50.2, 49.7. LC−MS (ESI) m/z: 411 (M + H)+.
1-(trans-4-(5-Fluoropyrimidin-2-ylamino)-3-(4-(trifluoromethyl)-benzyloxy)piperidin-1-yl)prop-2-en-1-one (Compound (±)-16).
The title compound was prepared according to the general procedure D.
1H NMR (400 MHz, CD3OD): δ (ppm) 8.21 (s, 2H), 7.51 (dt, J = 15.9, 8.4 Hz, 4H), 6.76 (ddd, J = 20.8, 16.8, 10.7 Hz, 1H), 6.19 (t, J = 15.5 Hz, 1H), 5.73 (dd, J = 14.7, 11.0 Hz, 1H), 4.76−4.55 (m, 2H), 4.41−3.42 (m, 6H), 2.13 (s, 1H), 1.61 (dt, J = 14.7, 6.8 Hz, 1H). 13C NMR (126 MHz, DMSO-d6): δ 164.9, 159.7, 152.2 (d, J = 247.6 Hz), 146.0, 144.3, 128.6 (q, J = 33.5 Hz), 128.1, 128.1, 127.7, 125.4, 124.7 (q, J = 272.2 Hz), 77.5, 69.7, 52.7, 47.0, 43.6, 29.5. LC−MS (ESI) m/ z: 425 [M + H]+.
1-(4-(Pyrimidin-2-yl)-6-(4-(trifluoromethyl)benzyloxy)-1,4-diaze-pan-1-yl)prop-2-en-1-one (Compound (±)-17).
Step 1: A mixture of 4-methoxybenzaldehyde (2 g, 14.7 mmol), ethane-1,2-diamine S16 (440 mg, 7.35 mmol), and NaBH4 (558 mg, 14.7 mmol) in MeOH (30 mL) was stirred at rt under N2 overnight. The mixture was diluted with water (100 mL) and extracted with EtOAc (3 × 50 mL). The combined organic was washed with brine (100 mL), dried over anhydrous Na2SO4, filtered, and concentrated to afford crude intermediate S17 as an oil (1.5 g). LC−MS (ESI) m/z: 301[M + H]+.
Step 2: A mixture of crude S17 (1000 mg, 3.0 mmol), 1,3-dibromopropan-2-ol (650 mg, 3.0 mmol), and Et3N (600 mg, 6.0 mmol) in toluene (20 mL) was stirred at 120 °C under N2 for 4 days. The mixture was concentrated and purified by prep-HPLC to obtain S18 as a brown solid (300 mg). LC−MS (ESI) m/z: 357 [M + H]+.
Step 3: To a solution of S18 (400 mg, 1.1 mmol) in DMF (5 mL) was added NaH (136 mg, 3.2 mmol) at 0 °C. The mixture was stirred at rt for 10 min, and then 1-(bromomethyl)-4-(trifluoromethyl)-benzene (268 mg, 1.1 mmol) was added. The resulting mixture was stirred at rt under N2 overnight. The mixture was diluted with water (100 mL) and extracted with EtOAc (50 mL × 3). The combined organic was washed with brine (100 mL), dried over anhydrous Na2SO4, and concentrated and purified by prep-HPLC to obtain compound S19 as a yellow oil (300 mg). LC−MS (ESI) m/z: 515 [M + H]+.
Step 4: A mixture of S19 (250 mg, 0.48 mmol) and TFA (20 mL) was stirred at 90 °C under N2 for 4 days. The mixture was concentrated to afford crude compound S20 as a white solid (100 mg). LC−MS (ESI) m/z: 275 [M + H]+.
Step 5: A mixture of S20 (200 mg, 0.72 mmol), 2-chloropyrimidine (80 mg, 0.72 mmol) and DIPEA (280 mg, 2.16 mmol) in DMSO (15 mL) was stirred at 90 °C under N2 for 3 h. The mixture was diluted with water (100 mL) and extracted with EtOAc (50 mL × 3). The combined organic was concentrated, and the residue was purified by prep-HPLC to obtain S21 as a yellow oil (50 mg). LC−MS (ESI) m/ z: 353 [M + H]+.
Step 6: A mixture of S21 (50 mg, 0.14 mmol), acryloyl chloride (13 mg, 0.14 mmol), and Et3N (28 mg, 0.28 mmol) in DCM (15 mL) was stirred at 0 °C for 2 h. The mixture was concentrated and purified by prep-HPLC to obtain compound 17 as a yellow oil (11 mg).
1H NMR (400 MHz, CD3OD): δ (ppm): 8.38−8.25 (m, 2H), 7.69−7.40 (m, 4H), 6.93−6.56 (m, 2H), 6.30−5.58 (m, 2H), 4.87− 3.39 (m, 10H), 3.15–3-08 (m, 1H). 13C NMR (126 MHz, DMSO-d6): δ 166.0, 161.9, 161.3, 158.5, 145.4, 129.1 (q, J = 33.5 Hz), 128.3, 126.1 (q, J = 3.8 Hz), 124.2 (q, J = 272.2 Hz), 110.9, 110.7, 74.8, 69.9, 50.2, 48.6, 47.3, 46.7. LC−MS (ESI) m/z: 407 [M + H]+.
1-((3R,4R)-3-(4-(Pyridin-3-yl)-1H-1,2,3-triazol-1-yl)-4-(4(trifluoromethyl)benzyloxy)pyrrolidin-1-yl)but-2-yn-1-one (Compound 18).
A mixture of 3-(1-((3R,4R)-4-((4-(trifluoromethyl)-benzyl)oxy)pyrrolidin-3-yl)-1H-1,2,3-triazol-4-yl)pyridine (100 mg, 0.25 mmol), but-2-ynoic acid (30 mg, 0.25 mmol), HATU (140 mg, 0.37 mmol), and Et3N (50 mg, 0.5 mmol) in DCM (10 mL) was stirred at rt for 2 h. The mixture was concentrated and purified by prep-HPLC to obtain compound 18 as a white solid (30 mg).
1H NMR (400 MHz, CD3OD): δ (ppm) 9.09−8.98 (m, 1H), 8.60 (d, J = 4.4 Hz, 1H), 8.57−8.53 (m, 1H), 8.33−8.27 (m, 1H), 7.70−7.50 (m, 5H), 5.52−5.40 (m, 1H), 4.79 (d, J = 8.3 Hz, 2H), 4.67−4.62 (m, 1H), 4.44−3.69 (m, 4H), 2.06 (d, J = 7.2 Hz, 3H). 13C NMR (126 MHz, DMSO-d6): δ 152.4, 149.5, 146.9, 144.3, 142.9, 132.9, 128.6 (q, J = 33.5 Hz), 128.5 (d, J = 2.3 Hz), 126.9, 125.6 (d, J = 3.9 Hz), 124.7 (q, J = 272.2 Hz), 124.5, 122.30, 89.14, 80.96, 74.29, 70.56, 62.47, 51.43, 49.09, 3.71. LC−MS (ESI) m/z: 456 [M + H]+.
(E)-4-(Dimethylamino)-1-((3R,4R)-3-(4-(pyridin-3-yl)-1H-1,2,3-triazol-1-yl)-4-(4-(trifluoromethyl)benzyloxy)pyrrolidin-1-yl)but-2-en-1-one (Compound 19).
A mixture of 3-(1-((3R,4R)-4-((4-(trifluoromethyl)benzyl)oxy)pyrrolidin-3-yl)-1H-1,2,3-triazol-4-yl)-pyridine (100 mg, 0.25 mmol), (E)-4-(dimethylamino)but-2-enoic acid (385 mg, 0.25 mmol), HATU (142 mg, 0.37 mmol), and TEA (50 mg, 0.5 mmol) in DCM (5 mL) was stirred at rt for 2 h. The mixture was concentrated and purified by prep-HPLC to obtain compound 19 as a white solid (60 mg).
1H NMR (400 MHz, CD3OD): δ (ppm) 9.03 (d, J = 2.1 Hz, 1H), 8.65−8.51 (m, 2H), 8.35−8.25 (m, 1H), 7.66−7.50 (m, 5H), 6.91 (dt, J = 15.2, 6.5 Hz, 1H), 6.51 (d, J = 15.3 Hz, 1H), 5.53−5.38 (m, 1H), 4.80 (d, J = 6.1 Hz, 2H), 4.67 (dd, J = 24.5, 5.4 Hz, 1H), 4.44− 3.99 (m, 3H), 3.96−3.75 (m, 1H), 3.21 (dd, J = 8.7, 3.3 Hz, 2H), 2.30 (d, J = 5.5 Hz, 6H). 13C NMR (126 MHz, DMSO-d6): δ 164.2, 149.5, 146.9, 144.3, 142.9, 142.8, 132.9, 128.7 (q, J = 33.5 Hz), 128.4 (d, J = 4.2 Hz), 126.9 (d, J = 1.9 Hz), 125.6 (q, J = 3.8 Hz), 124.6 (q, J = 272.2 Hz), 124.4 (d, J = 2.3 Hz), 123.4, 122.1, 81.0, 70.5, 62.7, 60.3, 50.2, 49.5, 45.5. LC−MS (ESI) m/z: 501 [M + H]+.
1-((3R,4R)-3-(4-(Pyridin-3-yl)-1H-1,2,3-triazol-1-yl)-4-(4-(trifluoromethyl)-benzyloxy)pyrrolidin-1-yl)-2-(trifluoromethyl)-prop-2-en-1-one (Compound 20).
A mixture of 3-(1-((3R,4R)-4-((4-(trifluoromethyl)benzyl)oxy)pyrrolidin-3-yl)-1H-1,2,3-triazol-4-yl)pyridine (50 mg, 0.12 mmol), 2-(trifluoromethyl)acrylic acid (20 mg, 0.12 mmol), HATU (70 mg, 0.18 mmol), and TEA (25 mg, 0.25 mmol) in DCM (6 mL) was stirred at rt for 2 h. The mixture was concentrated and purified by prep-HPLC to obtain compound 20 as a white solid (5 mg).
1H NMR (400 MHz, CD3OD): δ (ppm) 9.03 (s, 1H), 8.60 (d, J = 10.4 Hz, 1H), 8.56 (d, J = 4.4 Hz, 1H), 8.30 (d, J = 7.5 Hz, 1H), 7.71−7.49 (m, 5H), 6.33 (d, J = 10.3 Hz, 1H), 6.12 (s, 1H), 5.52−5.40 (m, 1H), 4.84−4.58 (m, 3H), 4.38−4.03 (m, 3H), 3.88−3.76 (m, 1H). 13C NMR (126 MHz, DMSO-d6): δ 162.3, 149.5, 146.9, 144.3, 142.9, 133.2, 132.9, 129.1 (q, J = 33.2 Hz), 128.5, 128.44, 127.2 (q, J = 33.2 Hz), 125.6 (d, J = 4.0 Hz), 125.6 (q, J = 272.2 Hz), 124.5, 122.3 (d, J = 8.1 Hz), 122.1 (q, J = 275.6 Hz), 80.9, 70.5, 62.5, 52.2, 49.8. LC−MS (ESI) m/z: 512.2 [M + H]+.
2-Fluoro-1-((3R,4R)-3-(4-(pyridin-3-yl)-1H-1,2,3-triazol-1yl)4(4-(trifluoromethyl)-benzyloxy)pyrrolidin-1-yl)prop-2-en-1-one (Compound 21).
A mixture of 3-(1-((3R,4R)-4-((4-(trifluoromethyl)-benzyl)oxy)pyrrolidin-3-yl)-1H-1,2,3-triazol-4-yl)pyridine (100 mg, 0.25 mmol), 2-fluoroacrylic acid (30 mg, 0.25 mmol), HATU (140 mg, 0.37 mmol), and Et3N (50 mg, 0.5 mmol) in DCM (10 mL) was stirred at rt for 2 h. The mixture was concentrated and purified by prep-HPLC to obtain compound 21 as a yellow oil (44 mg).
1H NMR (400 MHz, CD3OD): δ (ppm) 9.03 (s, 1H), 8.61 (s, 1H), 8.55 (dd, J = 5.2, 1.2 Hz, 1H), 8.30 (d, J = 7.8 Hz, 1H), 7.70−7.50 (m, 5H), 5.56 (dd, J = 47.2, 3.2 Hz, 1H), 5.51−5.42 (m, 1H), 5.31 (dd, J = 16.5, 3.5 Hz, 1H), 4.81 (s, 2H), 4.70−4.60 (m, 1H), 4.52−3.79 (m, 4H). 13C NMR (126 MHz, DMSO-d6): δ 159.7, 156.9 (d, J = 267.9 Hz), 149.5, 146.9, 144.3, 142.9, 132.9, 128.7 (q, J = 33.2 Hz), 128.4 (d, J = 4.8 Hz), 126.9, 125.6 (d, J = 3.9 Hz), 124.7 (q, J = 272.2 Hz), 124.4, 122.3, 101.2 (d, J = 15.3 Hz), 80.7, 70.5, 62.5, 51.5, 50.5. LC−MS (ESI) m/z: 462 [M + H]+.
2-Fluoro-1-((3R,4R)-3-(pyrimidin-2-ylamino)-4-((4-(trifluoromethyl)benzyl)oxy)pyrrolidin-1-yl)prop-2-en-1-one (Compound 22).
A mixture of 3-(1-((3R,4R)-4-((4-(trifluoromethyl)-benzyl)oxy)pyrrolidin-3-yl)-1H-1,2,3-triazol-4-yl)pyridine (200 mg, 0.58 mmol), 2-fluoroacrylic acid (60 mg, 0.69 mmol), HATU (256 mg, 0.69 mmol), and DIPEA (224 mg, 1.74 mmol) in DMF (5 mL) was stirred at rt overnight under N2 protection. The reaction mixture was monitored by LC−MS. The reaction mixture was concentrated and purified by p-HPLC to obtain compound 22 as a white solid (150 mg). 1H NMR (400 MHz, MeOD): δ 8.44 (s, 2 H), 7.82−7.50 (m, 4 H), 6.83 (dd, J = 8.3, 4.9 Hz, 1 H), 5.50 (dt, J = 47.1, 3.3 Hz, 1 H), 5.27 (dt, J = 16.5, 3.4 Hz, 1 H), 4.82 (dd, J = 13.3, 8.8 Hz, 2 H), 4.68−4.57 (m, 1 H), 4.28−3.67 (m, 5 H). 13C NMR (126 MHz, DMSO-d6): δ 161.6, 159.6, 158.3, 156.9 (d, J = 265.7 Hz), 143.4, 128.5, (q, J = 33.2 Hz), 128.1 (d, J = 9.0 Hz), 125.3 (d, J = 3.9 Hz), 124.5 (q, J = 272.2 Hz), 111.1 (d, J = 4.9 Hz), 100.2 (dd, J = 18.7, 15.5 Hz), 79.8, 69.5, 54.0, 51.2, 50.6. LC−MS (ESI) m/z: 411 [M + H]+.
Cell Lines.
NCI-H226 (ATCC, #CRL-5826) mesothelioma cells, NCI-H28 (ATCC, #CRL-5820) mesothelioma cells, 94T778 (ATCC, #CRL-3043) well-differentiated liposarcoma cells, and NCI-H226 reporter cells were cultured in RPMI1640 media supplemented with 10% FBS and 1% penicillin/streptomycin. Cells were passaged for no longer than 6 weeks for all experiments and tested negative for Mycoplasma using the MycoAlert mycoplasma detection kit (Lonza, #LT07–318).
Cell Proliferation.
NCI-H226 cells were seeded in 384 white plates (Corning, #3570) at the density of 100 cells per well with the total volume of 50 μL. Seeded cells were treated with TEAD inhibitors using the HP D300 digital dispenser (Tecan) and incubated for 5 days. On day 5, 12.5 μL of CellTiter-Glo reagent (Promega, #G7570) was added to the plate and incubated for 20 min. The luminescent signal was read using a PHERAstar FSX plate reader (BMG Labtech), and data were normalized to DMSO-treated samples.
TEAD Gene Reporter Assay.
TEAD transcriptional assay was conducted as previously described.19 NCI-H226 cells were infected by 150 μL of TEAD luciferase reporter lentivirus (BPS Bioscience, #79833) in the presence of 8 μg/mL polybrene. After 3 days of infection, cells were selected by 1 μg/μL puromycin. For testing TEADi, the selected cells were seeded in two 384 white plates (Corning, #3570) at a density of 1000 cells per well with the total volume of 40 μL. Seeded cells were treated with TEAD inhibitors using the HP D300 digital dispenser (Tecan). After 3 day treatment, the luciferase activity was measured by One-Glo luciferase reagents (Promega, #E6110). which was normalized by cell viability using CellTiter-Glo reagent (Promega, #G7570).
PRISM.
PRISM study (MTS019) was conducted as previously described.22 The compound was tested at an 8-point dose for 5-day treatment, with 902 cancer cells passing QC. Correlation and biomarker analyses were performed using data from DepMap (https://depmap.org/portal/download/all/).
Protein Purification.
TEAD2/3/4-YBD protein purification was conducted as previously described.19 TEAD1-YBD (209−424) expression vector was cloned using ligation-independent cloning with a 6×His tag at the N-terminus of the inserted gene. Protein expression in Rosetta(DE3) pLysS competent cells (Sigma, #70956) was inducted using 0.4 mM in isopropyl-1-thio-D-galactopyranoside at 18 °C for 16 h. Cell pellets were harvested and lysed in lysis buffer (20 mM HEPES pH 7.5, 800 mM NaCl, 10 mM imidazole pH 8, 10% glycerol, and 2 mM 2-mercaptoethanol) and purified using Cobalt resin. Then, elutes were further purified using gel filtration in the final buffer (20 mM Tris−HCl pH 8, 150 mM NaCl, and 1 mM TCEP). Protein was frozen down in the presence of 5% glycerol.
TR-FRET.
TR-FRET experiments were conducted in the assay buffer (50 mM HEPES pH 7.5, 200 mM NaCl, and 0.1% Pluronic F- 68 solution) as previously described.27 The synthesis of TR-FRET tracer WZJ10 has been described in the Supporting Information. In the assay, 100 nM His-TEAD-YBD recombinant protein was pretreated with TEAD inhibitors for 5 h in a 384 low-volume black plate (corning, #4514), and then 800 nM WZJ-10 was added to the plate followed by the addition of MAb Anti-6HIS Tb cryptate Gold HTRF (PerkinElmer, #61HI2TLA) with the final concentration of 50 ng/mL. The TR-FRET signal (490/520 nm) was collected using a PHERAstar FSX plate reader (BMG Labtech).
Intact Mass Spectrometry Analysis.
TEAD1 protein was incubated with DMSO or a 10-fold molar excess of compound 22 for 1 h at rt. Reactions were then analyzed by LC−MS using a Shimadzu autosampler and LC (Marlborough, MA) interfaced to an LTQ ion trap mass spectrometer (Thermo Fisher Scientific, San Jose, CA). Reactions were injected onto a self-packed column (0.5 mm I.D., packed 5 cm POROS 50R2 from Applied Biosystems, Framingham, MA), desalted for 4 min with 100% A (A = 0.2 M acetic acid in water), gradient eluted (0−100% B in 1 min; A = 0.2 M acetic acid in water, B = 0.2 M acetic acid in acetonitrile), and introduced to the mass spectrometer by electrospray ionization (spray voltage = 4.5 kV). The mass spectrometer was programmed to acquire full scan MS data (m/z 300−2000). Mass spectra were deconvoluted using MagTran version 1.03b2.28
Supplementary Material
ACKNOWLEDGMENTS
S.M.C. is supported by the National Institutes of Health (K08CA230220). B.N. is supported by the National Cancer Institute (K22CA258805).
ABBREVIATIONS
- ABPP
activity-based protein profiling
- AMOTL2
angiomotin like 2
- AUC
area under the curve
- FAM
fluorescein amidites
- IPTG
isopropyl-1-thio-D-galactopyranoside
- LATS1
large tumor suppressor kinase 1
- MOA
mechanism of action
- MOB1
Mps1 binding protein
- NF2
Neurofibromin-2
- PBP
palmitate-binding pocket
- PPI
protein−protein interaction
- SAR
structure−activity relationship
- SAV1
Salvador homolog 1
- TAZ
transcriptional coactivator with the PDZ-binding motif
- TEAD
transcriptional enhanced-associated domain
- TR-FRET
time-resolved fluorescence energy transfer
- YAP
yes-associated protein
- YBD
YAP binding domain
Footnotes
ASSOCIATED CONTENT
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.jmedchem.2c01548.
Gel-based ABPP data for WZJ-10 and VT-107; PRISM data for compound 22; TR-FRET data and reporter gene data for compounds 1−22; intact mass data for TEAD1-YBD with compound 22; antiproliferation data of TEAD inhibitors in NCI-H28 mesothelioma cells and 94T778 liposarcoma cells; HPLC traces for compounds 1−22; and synthesis procedure of WZJ-10 (PDF)
Molecular formula strings (CSV)
Complete contact information is available at: https://pubs.acs.org/10.1021/acs.jmedchem.2c01548
The authors declare the following competing financial interest(s): N.S.G. is a Scientific Founder, member of the SAB and equity holder in C4 Therapeutics, Syros, Soltego (board member), B2S/Voronoi, Allorion, Lighthorse, Cobroventures, GSK, Shenandoah (board member), Larkspur (board member) and Matchpoint. The Gray lab receives research funding from Springworks and Interline. B.N. has previously received speaking honoraria from Tocris Bio-Techne and Cell Signaling Technology. T.Z is a scientific founder, equity holder and consultant in Matchpoint. J.C. is a scientific founder for Matchpoint and M3 bioinformatics & technology Inc, consultant and equity holder for Soltego, Allorion, Matchpoint. J.A.M. is a founder, equity holder, and advisor to Entact Bio, serves on the SAB of 908 Devices, and receives sponsored research funding from Vertex, AstraZeneca, Taiho, Spring- works and TUO Therapeutics. TEAD inhibitors developed in this manuscript are licensed to a start-up (Lighthorse) where Gray has a financial interest. N.S.G. is one of the inventors on TEAD inhibitor patents (WO2022232088 A1).
Contributor Information
Wenchao Lu, Department of Chemical and Systems Biology, Chem-H and Stanford Cancer Institute, Stanford School of Medicine, Stanford University, Stanford, California 94305, United States.
Mengyang Fan, Hangzhou Institute of Medicine (HIM), Chinese Academy of Sciences, Hangzhou, Zhejiang 310022, China.
Wenzhi Ji, Department of Chemical and Systems Biology, Chem-H and Stanford Cancer Institute, Stanford School of Medicine, Stanford University, Stanford, California 94305, United States.
Jason Tse, Department of Chemical and Systems Biology, Chem-H and Stanford Cancer Institute, Stanford School of Medicine, Stanford University, Stanford, California 94305, United States.
Inchul You, Department of Chemical and Systems Biology, Chem-H and Stanford Cancer Institute, Stanford School of Medicine, Stanford University, Stanford, California 94305, United States.
Scott B. Ficarro, Department of Cancer Biology, Blais Proteomics Center, Center for Emergent Drug Targets, Dana- Farber Cancer Institute, Boston, Massachusetts 02215, United States
Isidoro Tavares, Department of Cancer Biology, Blais Proteomics Center, Center for Emergent Drug Targets, Dana- Farber Cancer Institute, Boston, Massachusetts 02215, United States.
Jianwei Che, Center for Protein Degradation, Dana-Farber Cancer Institute, Boston, Massachusetts 02215, United States.
Audrey Y. Kim, Department of Chemical and Systems Biology, Chem-H and Stanford Cancer Institute, Stanford School of Medicine, Stanford University, Stanford, California 94305, United States
Xijun Zhu, Department of Chemistry, Stanford University, Stanford, California 94305, United States.
Andrew Boghossian, Broad Institute of MIT and Harvard, Cambridge, Massachusetts 02142, United States.
Matthew G. Rees, Broad Institute of MIT and Harvard, Cambridge, Massachusetts 02142, United States
Melissa M. Ronan, Broad Institute of MIT and Harvard, Cambridge, Massachusetts 02142, United States
Jennifer A. Roth, Broad Institute of MIT and Harvard, Cambridge, Massachusetts 02142, United States
Stephen M. Hinshaw, Department of Chemical and Systems Biology, Chem-H and Stanford Cancer Institute, Stanford School of Medicine, Stanford University, Stanford, California 94305, United States
Behnam Nabet, Human Biology Division, Fred Hutchinson Cancer Center, Seattle, Washington 98109, United States.
Steven M. Corsello, Department of Medicine and Stanford Cancer Institute, Stanford School of Medicine, Stanford University, Stanford, California 94305, United States
Nicholas Kwiatkowski, Center for Protein Degradation, Dana-Farber Cancer Institute, Boston, Massachusetts 02215, United States.
Jarrod A. Marto, Department of Cancer Biology, Blais Proteomics Center, Center for Emergent Drug Targets, Dana- Farber Cancer Institute, Boston, Massachusetts 02215, United States; Department of Pathology, Brigham and Women’s Hospital and Harvard Medical School, Boston, Massachusetts 02215, United States
Tinghu Zhang, Department of Chemical and Systems Biology, Chem-H and Stanford Cancer Institute, Stanford School of Medicine, Stanford University, Stanford, California 94305, United States.
Nathanael S. Gray, Department of Chemical and Systems Biology, Chem-H and Stanford Cancer Institute, Stanford School of Medicine, Stanford University, Stanford, California 94305, United States
REFERENCES
- (1).Meng Z; Moroishi T; Guan KL Mechanisms of Hippo pathway regulation. Genes Dev. 2016, 30, 1–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (2).(a) Miyanaga A; Masuda M; Tsuta K; Kawasaki K; Nakamura Y; Sakuma T; Asamura H; Gemma A; Yamada T. Hippo pathway gene mutations in malignant mesothelioma: revealed by RNA and targeted exon sequencing. J. Thorac. Oncol. 2015, 10, 844–851. [DOI] [PubMed] [Google Scholar]; (b) Moroishi T; Hansen CG; Guan K-L The emerging roles of YAP and TAZ in cancer. Nat. Rev. Cancer 2015, 15, 73–79. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (3).Kurppa KJ; Liu Y; To C; Zhang T; Fan M; Vajdi A; Knelson EH; Xie Y; Lim K; Cejas P; et al. Treatment-Induced Tumor Dormancy through YAP-Mediated Transcriptional Reprogramming of the Apoptotic Pathway. Cancer Cell 2020, 37, 104–122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (4).Li Z; Razavi P; Li Q; Toy W; Liu B; Ping C; Hsieh W; Sanchez-Vega F; Brown DN; Da Cruz Paula AF; et al. Loss of the FAT1 Tumor Suppressor Promotes Resistance to CDK4/6 Inhibitors via the Hippo Pathway. Cancer Cell 2018, 34, 893–905. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (5).Kapoor A; Yao W; Ying H; Hua S; Liewen A; Wang Q; Zhong Y; Wu CJ; Sadanandam A; Hu B; et al. Yap1 activation enables bypass of oncogenic Kras addiction in pancreatic cancer. Cell 2014, 158, 185–197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (6).(a) Zhou Z; Hu T; Xu Z; Lin Z; Zhang Z; Feng T; Zhu L; Rong Y; Shen H; Luk JM; et al. Targeting Hippo pathway by specific interruption of YAP-TEAD interaction using cyclic YAP-like peptides. FASEB J 2015, 29, 724–732. [DOI] [PubMed] [Google Scholar]; (b) Jiao S; Wang H; Shi Z; Dong A; Zhang W; Song X; He F; Wang Y; Zhang Z; Wang W; et al. A peptide mimicking VGLL4 function acts as a YAP antagonist therapy against gastric cancer. Cancer Cell 2014, 25, 10066–180. [DOI] [PubMed] [Google Scholar]
- (7).Smith SA; Sessions RB; Shoemark DK; Williams C; Ebrahimighaei R; McNeill MC; Crump MP; McKay TR; Harris G; Newby AC; et al. Antiproliferative and Antimigratory Effects of a Novel YAP-TEAD Interaction Inhibitor Identified Using in Silico Molecular Docking. J. Med. Chem. 2019, 62, 1291–1305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (8).Sturbaut M; Bailly F; Coevoet M; Sileo P; Pugniere M; Liberelle M; Magnez R; Thuru X; Chartier-Harlin MC; Melnyk P; et al. Discovery of a cryptic site at the interface 2 of TEAD - Towards a new family of YAP/TAZ-TEAD inhibitors. Eur. J. Med. Chem. 2021, 226, 113835. [DOI] [PubMed] [Google Scholar]
- (9).Furet P; Bordas V; Le Douget M; Salem B; Mesrouze Y; Imbach-Weese P; Sellner H; Voegtle M; Soldermann N; Chapeau E; et al. The First Class of Small Molecules Potently Disrupting the YAP-TEAD Interaction by Direct Competition. ChemMedChem 2022, 17, No. e202200303. [DOI] [PubMed] [Google Scholar]
- (10).(a) Chan P; Han X; Zheng B; DeRan M; Yu J; Jarugumilli GK; Deng H; Pan D; Luo X; Wu X. Autopalmitoylation of TEAD proteins regulates transcriptional output of the Hippo pathway. Nat. Chem. Biol. 2016, 12, 282–289. [DOI] [PMC free article] [PubMed] [Google Scholar]; (b) Pobbati AV; Hong W. A combat with the YAP/TAZ-TEAD oncoproteins for cancer therapy. Theranostics 2020, 10, 3622–3635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (11).Tang TT; Konradi AW; Feng Y; Peng X; Ma M; Li J; Yu FX; Guan KL; Post L. Small Molecule Inhibitors of TEAD Auto-palmitoylation Selectively Inhibit Proliferation and Tumor Growth ofNF2-deficient Mesothelioma. Mol. Cancer Ther. 2021, 20, 986–998. [DOI] [PubMed] [Google Scholar]
- (12).Li Q; Sun Y; Jarugumilli GK; Liu S; Dang K; Cotton JL; Xiol J; Chan PY; DeRan M; Ma L; et al. Lats1/2 Sustain Intestinal Stem Cells and Wnt Activation through TEAD-Dependent and Independent Transcription. Cell Stem Cell 2020, 26, 675–692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (13).Hu L; Sun Y; Liu S; Erb H; Singh A; Mao J; Luo X; Wu X. Discovery of a new class of reversible TEA domain transcription factor inhibitors with a novel binding mode. Elife 2022, 11, No. e80210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (14).Holden JK; Crawford JJ; Noland CL; Schmidt S; Zbieg JR; Lacap JA; Zang R; Miller GM; Zhang Y; Beroza P; et al. Small Molecule Dysregulation of TEAD Lipidation Induces a Dominant-Negative Inhibition of Hippo Pathway Signaling. Cell Rep. 2020, 31, 107809. [DOI] [PubMed] [Google Scholar]
- (15).Heinrich T; Peterson C; Schneider R; Garg S; Schwarz D; Gunera J; Seshire A; Kötzner L; Schlesiger S; Musil D; et al. Optimization of TEAD P-Site Binding Fragment Hit into In Vivo Active Lead MSC-4106. J. Med. Chem. 2022, 65, 9206. [DOI] [PubMed] [Google Scholar]
- (16).Bum-Erdene K; Zhou D; Gonzalez-Gutierrez G; Ghozayel MK; Si Y; Xu D; Shannon HE; Bailey BJ; Corson TW; Pollok KE; et al. Small-Molecule Covalent Modification of Conserved Cysteine Leads to Allosteric Inhibition of the TEAD· Yap Protein-Protein Interaction. Cell Chem. Biol. 2019, 26, 378–389. [DOI] [PubMed] [Google Scholar]
- (17).Lu W; Wang J; Li Y; Tao H; Xiong H; Lian F; Gao J; Ma H; Lu T; Zhang D; et al. Discovery and biological evaluation of vinylsulfonamide derivatives as highly potent, covalent TEAD autopalmitoylation inhibitors. Eur. J. Med. Chem. 2019, 184, 111767. [DOI] [PubMed] [Google Scholar]
- (18).Lu T; Li Y; Lu W; Spitters T; Fang X; Wang J; Cai S; Gao J; Zhou Y; Duan Z; et al. Discovery of a subtype-selective, covalent inhibitor against palmitoylation pocket of TEAD3. Acta Pharm. Sin. B 2021, 11, 3206–3219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (19).Karatas H; Akbarzadeh M; Adihou H; Hahne G; Pobbati AV; Yihui Ng E; Guéret SM; Sievers S; Pahl A; Metz M; et al. Discovery of Covalent Inhibitors Targeting the Transcriptional Enhanced Associate Domain Central Pocket. J. Med. Chem. 2020, 63, 11972–11989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (20).Kaneda A; Seike T; Danjo T; Nakajima T; Otsubo N; Yamaguchi D; Tsuji Y; Hamaguchi K; Yasunaga M; Nishiya Y; et al. The novel potent TEAD inhibitor, K-975, inhibits YAP1/TAZ- TEAD protein-protein interactions and exerts an anti-tumor effect on malignant pleural mesothelioma. Am. J. Cancer Res. 2020, 10, 4399–4415. [PMC free article] [PubMed] [Google Scholar]
- (21).Liberelle M; Toulotte F; Renault N; Gelin M; Allemand F; Melnyk P; Guichou JF; Cotelle P. Toward the Design of Ligands Selective for the C-Terminal Domain of TEADs. J. Med. Chem. 2022, 65, 5926–5940. [DOI] [PubMed] [Google Scholar]
- (22).Fan M; Lu W; Che J; Kwiatkowski NP; Gao Y; Seo HS; Ficarro SB; Gokhale PC; Liu Y; Geffken EA; et al. Covalent disruptor of YAP-TEAD association suppresses defective hippo signaling. Elife 2022, 11, No. e78810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (23).Fell JB; Fischer JP; Baer BR; Blake JF; Bouhana K; Briere DM; Brown KD; Burgess LE; Burns AC; Burkard MR; et al. Identification of the Clinical Development Candidate MRTX849, a Covalent KRASG12C Inhibitor for the Treatment of Cancer. J. Med. Chem. 2020, 63, 6679–6693. [DOI] [PubMed] [Google Scholar]
- (24).Corsello SM; Nagari RT; Spangler RD; Rossen J; Kocak M; Bryan JG; Humeidi R; Peck D; Wu X; Tang AA; et al. Discovering the anticancer potential of non-oncology drugs by systematic viability profiling. Nat. Cancer 2020, 1, 235–248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (25).Zhao B; Li L; Lu Q; Wang LH; Liu CY; Lei Q; Guan KL Angiomotin is a novel Hippo pathway component that inhibits YAP oncoprotein. Genes Dev. 2011, 25, 51–63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (26).Wang W; Li N; Li X; Tran MK; Han X; Chen J. Tankyrase Inhibitors Target YAP by Stabilizing Angiomotin Family Proteins. Cell Rep. 2015, 13, 524–532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (27).Nowak RP; DeAngelo SL; Buckley D; He Z; Donovan KA; An J; Safaee N; Jedrychowski MP; Ponthier CM; Ishoey M; et al. Plasticity in binding confers selectivity in ligand-induced protein degradation. Nat. Chem. Biol. 2018, 14, 706–714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (28).Zhang Z; Marshall AG A universal algorithm for fast and automated charge state deconvolution of electrospray mass-to-charge ratio spectra. J. Am. Soc. Mass Spectrom. 1998, 9, 225–233. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.