Abstract
The expansion of genetic engineering has brought a new dimension for synthetic immunology. Immune cells are perfect candidates because of their ability to patrol the body, interact with many cell types, proliferate upon activation, and differentiate in memory cells. This study aimed at implementing a new synthetic circuit in B cells, allowing the expression of therapeutic molecules in a temporally and spatially restricted manner that is induced by the presence of specific antigens. This should enhance endogenous B cell functions in terms of recognition and effector properties. We developed a synthetic circuit encoding a sensor (a membrane-anchored B cell receptor targeting a model antigen), a transducer (a minimal promoter induced by the activated sensor), and effector molecules. We isolated a 734-bp-long fragment of the NR4A1 promoter, specifically activated by the sensor signaling cascade in a fully reversible manner. We demonstrate full antigen-specific circuit activation as its recognition by the sensor induced the activation of the NR4A1 promoter and the expression of the effector. Overall, such novel synthetic circuits offer huge possibilities for the treatment of many pathologies, as they are completely programmable; thus, the signal-specific sensors and effector molecules can be adapted to each disease.
Keywords: MT: Delivery Strategies, B cells, synthetic circuits, reprogramming, cell therapy, autologous, personalized medicine
Graphical abstract

Cosset et al. reprogrammed B cells with a lentiviral vector encoding a synthetic circuit, comprising a membrane sensor targeting a specific antigen, a promoter inducible by this sensor and controlling an effector transgene. Upon target antigen detection by the sensor, the effector protein was specifically expressed in a reversible manner.
Introduction
The expansion of techniques for genetic engineering has recently brought a new dimension for synthetic biology approaches. Synthetic biology relies on the design of genetic parts and biological blocks that are assembled in the target cells to create new genetic networks.1 For instance, synthetic bio-sensing circuits are composed of sensor elements that bind the signal molecule and transducer modules, which can lead to specific cellular responses. The input or inducing signal should be specific to the disease, such as an induced antigen or a dysregulation of the microenvironment, and is recognized by a dedicated receptor molecule. This recognition triggers a signaling cascade and the integration of information, leading to an output response from the reprogrammed cells. Such circuits are of valuable importance from a therapeutic point of view, notably for regulated drug delivery in vivo upon biomarker sensing.
Synthetic biology approaches have been implemented in immune cells and represent a new field called synthetic immunology, which holds great promises for the treatment of many diseases as cells of the immune system play a crucial role in detecting and responding to pathological deviations.2 Several novel functions can be conferred by genetic reprogramming of immune cells to enhance their endogenous properties. Indeed, immune cells recognize pathological signals, such as those induced by pathogens, abnormal cells, or inflammation, and trigger responses to restore homeostasis; yet, their recognition capacity can be potentiated by adding new receptors to more precisely distinguish a pathological environment from a normal environment. Additionally, it is also possible to augment their effector functions, for instance, by forcing the expression of therapeutic molecules.
Several characteristics of immune cells make them the perfect candidates for synthetic biology approaches. First, these cells move freely in the body to patrol and infiltrate various tissues. Because of their global distribution in the body, as well as their endogenous functions, immune cells can act as intermediaries and communicate with other cell types, thus leading to the modification of the immune responses on a broader scale. Second, they naturally expand when stimulated and differentiate into long-lived memory cell subsets, which can be especially useful during disease flares. In addition, immune cells either protect or directly contribute to many pathologies, such as cancer or autoimmune disease.3,4 Finally, immune cells can be easily collected and modified.
Most synthetic immunology approaches have used T cells, the best-known example being the successful chimeric antigen receptor (CAR)-T cells that were notably developed in cancer immunotherapy.5 Yet, B cells also offer unique opportunities for synthetic immunology. Indeed, a novel first-in-man phase I/IIa clinical trial in patients was recently initiated to evaluate the safety and tolerability of adoptively transferred donor B cells.6 The B cells were manufactured under Good Manufacturing Practices conditions and their transfer was well tolerated without any acute adverse reactions during the 4-month follow-up after adoptive transfer.6,7
Here, we developed a new synthetic circuit that was implemented in B cells upon lentiviral vector transduction. This synthetic circuit encodes a sensor (i.e., a B cell receptor [BCR], targeting a model antigen), a transducer (i.e., a part of the NR4A1 promoter), and an effector molecule. Upon binding of disease biomarkers, here recognized as s signal inducer for the synthetic circuit, to the ectopic BCR, the NR4A1 promoter was specifically activated, driving the expression of therapeutic molecules in a temporally and spatially restricted manner.8
Results
Generation of a small BCR-inducible ectopic NR4A1 promoter construct
The Nur77 (NR4A1) promoter and protein are specifically induced in B cells following BCR stimulation.8,9,10 Based on this property, we isolated several fragments of the NR4A1 promoter ending at 109 base pairs after the transcription start site and placed them upstream of the GFP reporter gene in a lentiviral vector (Figure 1A). Of note, these fragments contain binding sequences for the NFAT and nuclear factor κB transcription factors that are involved in the BCR signaling cascade.
Figure 1.
Isolation and characterization of a small ectopic BCR-inducible promoter construct
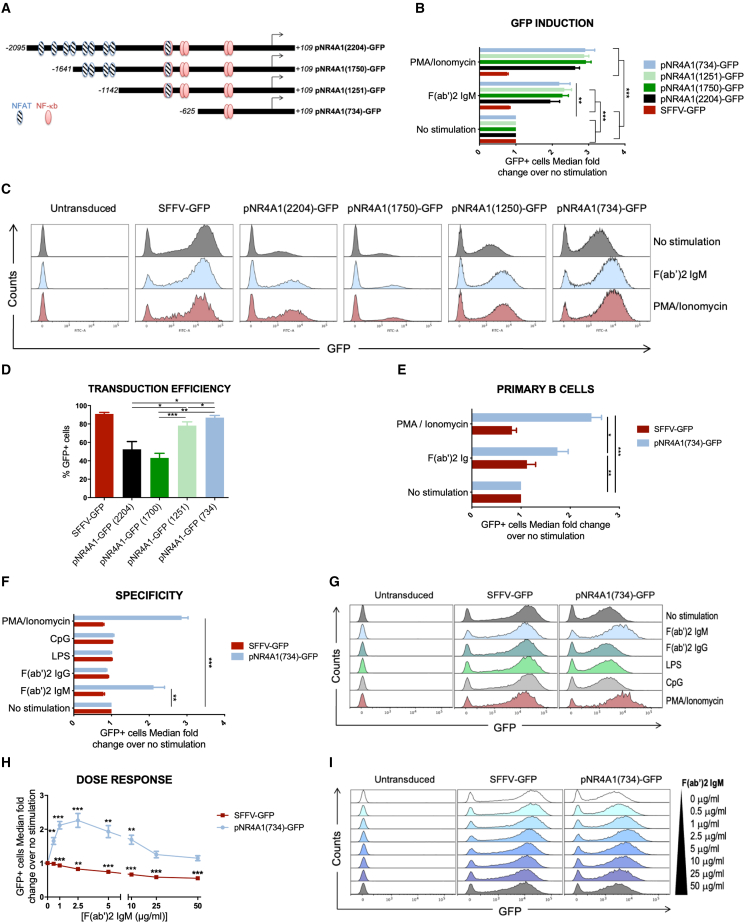
(A) Schematic representation of the NR4A1 reporter constructs with putative binding domains of the NF-κB and NFAT transcription factors. Different LVs encoding a reporter gene (GFP) with several promoter sizes (2,204, 1,750, 1,251, or 7,34 bp) have been constructed. (B) GFP induction of the reporter constructs after 24 h of BCR stimulation. BJAB cells were transduced with LVs encoding the reporter constructs (NR4A1 or SFFV as a constitutive control) before stimulation with F(ab’)2 IgM or PMA (combined with ionomycin for 24 h). The median GFP expression of GFP+ cells after stimulation was normalized by the non-stimulated condition. (C) Representative flow cytometry histograms of GFP expression are presented for one replicate. (D) Transduction efficiency of BJAB cells by the reporter constructs. The percentage of GFP+ cells after transduction was assessed for each construct 4 days after transduction. (E) Induction of the promoter in primary human B cells. Primary human B cells isolated from peripheral blood were transduced with LVs encoding the GFP under the 734-bp inducible reporter or the SFFV constitutive reporter before 24-h stimulation with F(ab’)2 Ig or PMA combined with ionomycin. GFP median expression was assessed by flow cytometry in GFP+ cells. The induction fold was computed as the median GFP intensity in GFP+ cells after stimulation over the median of non-stimulated GFP+ cells. (F) Specificity of the induction through the BCR. BJAB cells were transduced with LVs encoding the reporter constructs (734-bp NR4A1 or SFFV as a constitutive control) before stimulation with F(ab’)2 IgM, F(ab’)2 IgG, LPS, or CpG or PMA combined with ionomycin for 24 h. The median GFP expression of GFP+ cells after stimulation was normalized by the non-stimulated condition. (G) Representative flow cytometry histograms are presented for one replicate. (H) Dose-response of the 734-bp NR4A1 reporter construct with increasing amounts of F(ab’)2 IgM. BJAB cells were transduced with LVs encoding the 734-bp inducible reporter or the SFFV constitutive reporter before 24 h of stimulation with F(ab’)2 IgM. GFP median expression of DAPI- GFP+ cells relative to unstimulated cells was assessed. I. Representative histograms of GFP expression are presented. Data in (B, D, E, F and H) are representative of four independent experiments (n = 4, ANOVA 2 and multiple t-tests with Sidak Bonferroni correction [alpha = 0.05]). Error bars represent SEM. p values under 0.05 were considered statistically significant and the following denotations were used: ∗∗∗, p<0.001; ∗∗, p<0.01; ∗, p<0.05.
To assess the effect of BCR stimulation on promoter inducibility, BJAB cells, from a human B cell line, were transduced with the same MOI by lentiviral vectors encoding a GFP reporter construct under control of either the inducible promoters or a constitutively active promoter (from the Spleen Focus Forming Virus [SFFV]11) before a 24-h stimulation through the endogenous IgM BCR,12 using an F(ab’)2 IgM molecule. We detected an increase of approximately 2.5-fold of GFP expression for all inducible constructs, while GFP expression from the SFFV promoter was slightly downregulated. The induction was in the same range than that induced by ionomycin combined with phorbol myristate acetate (PMA), which activates the BCR signaling cascade without requiring BCR direct linkage (Figures 1B and 1C). The promoter length did not impact the magnitude of the induction after BCR stimulation, but influenced the percentage of transduction, which increased while decreasing promoter size, with the 734-bp promoter leading to more than 90% of transduction (Figure 1D). This result was expected as smaller cassettes have been shown to be more easily incorporated in viral particles and integrated into the genome.13,14 Moreover, the leakage of promoter activity, assessed by the median GFP expression of GFP-positive cells without stimulation, was 1.7-fold higher for the 734-bp construct as compared with the 2,204-bp construct (Figure S1). This increase was not caused by the lack of the regulatory sequences present on the distal parts of the NR4A1 promoter, but rather reflected a higher copy number of vectors integrated in the genome when using smaller promoter constructs (Figures 1A and S1). Of note, a minimal expression was expected; almost all promoters exhibit a physiological background activity.15
Characterization of the inducible 734-bp ectopic NR4A1 promoter
Given the similar levels of inducibility of the constructs and taking in to account the limited packaging capacity of lentiviral vectors, we focused on the smaller promoter construct (NR4A1[734]) for subsequent experiments.
Importantly, we found that the NR4A1 reporter construct was specifically induced upon BCR signaling but not upon Toll-like receptor (TLR)-4 (lipopolysaccharide [LPS]) or TLR-9 (CpG) stimulation (Figures 1F and 1G), confirming the specificity of activation of our ectopic minimal promoter through the BCR pathway. We established the dose-response curve of activation of this promoter fragment after 24 h of stimulation with F(ab’)2 IgM molecules, which displayed a peak at a concentration of 2.5 μg/mL (Figures 1H and 1I) Of note, higher concentrations of F(ab’)2 IgM molecules dramatically decreased cell viability (Figure S2), likely because of the triggering of activation-induced cell death, a process leading to cell apoptosis after over-stimulation of the BCR or stimulation without co-signal molecules, such as CD40L.16
While these experiments were carried out in BJAB cells, we also assessed the inducibility of the promoter in alternative B cell lines that express IgM BCRs (Figure S3). Similar to BAJB cells, a 2.5-fold induction of the NR4A1 reporter construct was observed in BL-2 cells after BCR linkage, but not in other cell lines, although GFP expression was fully induced after culture with PMA and ionomycin. Interestingly, all responsive cell lines were negative for Epstein-Barr virus (EBV), while all non-responsive cell lines were positive for EBV (Figure S3). Indeed, EBV infection latency might explain the unresponsiveness after BCR stimulation, since the lmp2a viral protein, expressed in the EBV positive cell lines, has been shown to inhibit BCR signaling (Figure S3).17,18 Nevertheless, since only a small fraction of B cells are EBV-positive in infected patients, approximately 1–50 cells per 106 cells, this should not affect the functionality of the ectopic NR4A1 promoter.19,20 Importantly, this short promoter was also responsive to BCR stimulation in transduced human primary B cells (Figures 1E and S4).
Induction, extinction, and reversibility kinetics of the ectopic NR4A1 promoter
To better characterize the kinetics of the inducible promoter cassette, we inserted a destabilized TurboGFP marker (TurboGFPdes) after the NR4A1(734) promoter, which has both a shorter maturation time and a shorter half-life than the classical GFP reporter.21 First, we assessed the kinetics of promoter induction after 0 to 24 h of continuous stimulation (Figures 2A, 2B, and S5). The induction was fast since we detected a 2-fold increase in TurboGFPdes after a 4-h stimulation, which reached 12-fold induction upon 24 h of stimulation. Of note, this induction was slightly higher after stimulation through the IgM as compared with PMA plus ionomycin. Second, we investigated the extinction kinetics to assess if induction was reversed when the stimulus is removed (Figures 2A and 2C). Cells transduced with lentiviral vectors (LVs) encoding the TurboGFPdes under control of constitutive or inducible promoters were stimulated through their BCRs for 1, 4, 8, or 24 h before removing the IgM BCR-stimulating agents. While at 1 day after stimuli removal, the levels of TurboGFPdes remained high, they were restored at the basal values of non-stimulated cells, thus demonstrating the reversibility of the induction. Finally, we assessed whether the inducible promoter cassette could be induced multiple times (Figures 2A and 2D). Three rounds of stimulation were performed on transduced cells by alternating 8 h of stimulation (gray) and 88 h of rest (white). We found that, while the induction-fold slightly increased over stimulation rounds, the induction was fully reversible even after three cycles of stimulation.
Figure 2.
Kinetics characterization of the inducible 734-bp NR4A1 promoter
(A) Scheme of the kinetics characterization set-up. (B) Kinetics of induction of the promoter. BJAB cells were transduced with LVs encoding the TurboGFPdes under the 734-bp inducible reporter before 0 to 24 h of stimulation with F(ab’)2 IgM or PMA combined with ionomycin. TurboGFPdes median expression in each condition (after the indicated stimulation duration) was assessed and normalized by TurboGFPdes median of unstimulated cells (n = 3). (C) Kinetics of extinction of the promoter. Transduced BJAB cells were stimulated for 0, 1, 4, 8, and 24 h with F(ab’)2 IgM or PMA combined with ionomycin. Cells were then washed three times and kept in culture. TurboGFPdes median expression was assessed at days 1, 3, and 6 after washing and normalized by TurboGFPdes median of unstimulated cells (n = 3). (D) Reversibility of the inducible promoter. Reversibility of pNR4A1(734)-responsive TurboGFPdes expression was assessed by culturing transduced BJAB cells while alternating 8 h of stimulation (gray) and 88 h of rest (white) three times with F(ab’)2 IgM. TurboGFPdes median expression was assessed before and after stimulation and normalized by TurboGFPdes median of unstimulated cells (n = 3). Error bars represent SEM.
Construction of a vector encoding a membrane-anchored BCR receptor as a signal sensor
Having validated the BCR-inducible promoter, we then sought to develop a complete synthetic circuit for B cell reprogramming. Hence, a specific sensor was generated using a membrane-anchored BCR that displays the variable regions of monoclonal antibodies directed against either the hepatitis B surface (HBs) protein of hepatitis B virus (FAM0-ADRI)22 or the ovalbumin (OVA) protein (FAM0-OVA),13 along with the constant IgG/kappa human immunoglobulin domains fused to the transmembrane domains. Of note, the intronic regions that allow the conditional secretion of immunoglobulins upon B cell activation were removed to express only the membrane-anchored form of the immunoglobulin.24 The B cell-specific FEEK promoter was used to control BCR expression.25 To validate this circuit component, both in terms of expression and functionality, LVs encoding either anti-HBs and anti-OVA BCR sensors were used to transduce BJAB cells, which endogenously express IgM immunoglobulins, but are negative for IgG immunoglobulins (Figure 3A). After LV transduction, IgG BCRs were detected by surface cytometry staining in approximately 40%–50% of cells, hence validating the membrane expression of the sensor component of the circuit (Figure 3A). The expression of an ectopic IgG BCR at cell surface correlated with a decrease in the expression of the endogenous BCR, which has also been previously observed.24 This might be caused by a defect in the addressing process, as immunoglobulins are assembled with Igα and Igβ molecules in the endoplasmic reticulum (ER) and then transported together onto the cell surface. If the amount of Igα and Igβ is limiting, a competition for pairing between ectopic and endogenous BCRs can occur, and the non-assembled BCR components will be retained in the ER by a quality control system.26
Figure 3.
Design and validation of a synthetic sensor
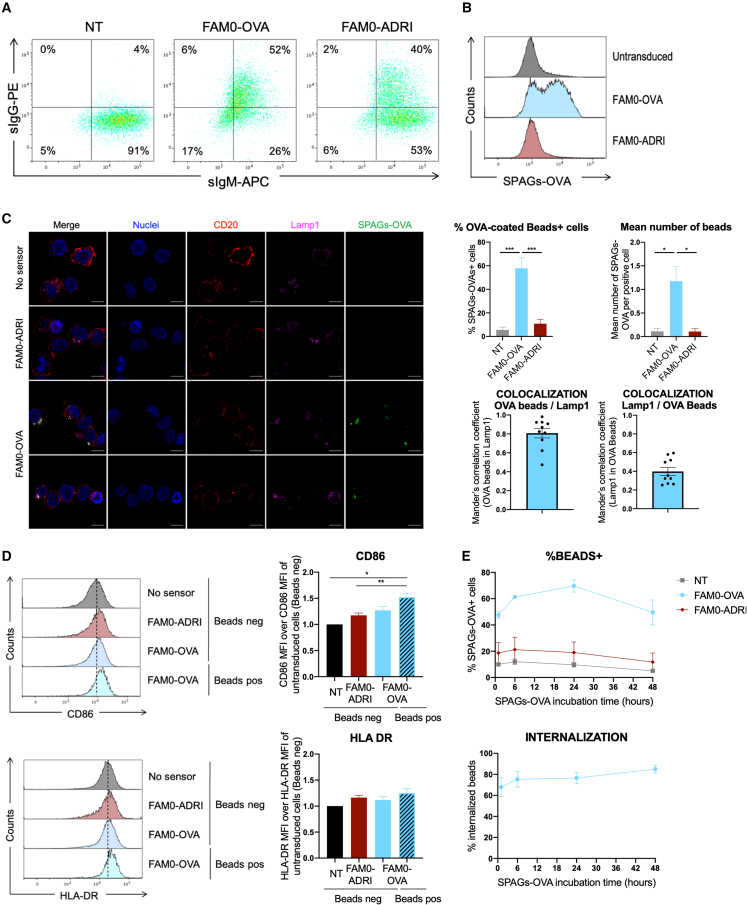
(A) Validation of sensor expression. BJAB cells (IgM+, IgG–) were transduced with LVs encoding sensors (membrane anchored IgG) recognizing either OVA (FAM0-OVA) or the HbS glycoprotein of HBV (FAM0-ADRI) 5 days before staining of IgG by flow cytometry. (B and C) Specific recognition of OVA by membrane-anchored BCR directed against OVA. BJAB cells were transduced with LVs encoding membrane-anchored BCR targeting either OVA or HbS. Five days after transduction, these cells were incubated for 24 h with OVA-coated fluorescent beads. Binding of fluorescent beads was assessed by flow cytometry (B) or immunofluorescence (C). For immunofluorescence, transduced cells were incubated overnight with OVA-coated fluorescent beads before being washed, fixed, and permeabilized for staining with an anti-Lamp1 antibody and Hoechst. Images were acquired on a confocal microscope. Scale bar, 10 μm. Four fields for each condition were quantified to assess the percentage of cells with beads as well as the median number of beads per cells. The Mander’s correlation coefficients of OVA beads and Lamp1 colocalization were computed. (D) Signaling and cell activation after SPAGs-OVA binding. BJAB cells transduced with membrane BCRs recognizing HbS or OVA were incubated for 24 h with SPAGs-OVA before staining with an anti-CD86 or an anti-HLA-DR antibody. One representative overlay is presented along with the median fluorescence intensity quantified from three experiments for both markers (ANOVA1 and multiple comparison with Tukey correction, n = 3). (E) Internalization of SPAGs-OVA specifically by the OVA sensors. BJAB cells transduced with membrane BCRs recognizing HbS or OVA were incubated for different times (1 h, 6 h, 24 h, or 48 h) with SPAGs-OVA before staining with an anti-OVA antibody (surface staining). The percentage of cells positive for SPAGs-OVA as well as the percentage of internalization computed as the percentage of SPAGs-OVA+/OVA-cells over the percentage of SPAGs-OVA+ cells are presented (n = 3). p values under 0.05 were considered statistically significant and the following denotations were used: ∗∗∗, p<0.001; ∗∗, p<0.01; ∗, p<0.05.
Next, synthetic particulate antigen (SPAGs) beads consisting of 400-nm fluorescent beads coated with OVA molecules were incubated with transduced cells to validate antigen-specific recognition of the sensor.27 After 24 h of SPAGs-OVA beads incubation, only cells transduced with the OVA-specific sensor were positively stained (Figure 3B). The percentage of the SPAGs-OVA beads positive cells as determined by FACS analysis, as well as the mean number of beads per positive cells assessed by immunofluorescence staining were, respectively, approximately 50% and 1 (Figures 3A–3C).
Then, to validate sensor functionality and signaling after antigen-specific recognition, we quantified the expression of activation markers by cytometry in sensor-transduced cells after 24 h of incubation with SPAGs-OVA beads (Figure 3D). We found that the CD86 costimulatory molecule was upregulated in B cell lines expressing the OVA sensor as compared with non-transduced cells or to cells expressing an irrelevant sensor, suggesting an antigen-specific activation.
After antigen binding, the BCR-antigen complex naturally undergoes endocytosis and intra-cellular processing, leading the antigen presentation on major histocompatibility complex (MHC) class II complexes. No upregulation of human leukocyte antigen DR-1 (HLA-DR) (MHC II) was observed after antigen ligation on FAM0-OVA sensors (Figure 3D), although colocalization of SPAG-OVA beads and the LAMP1 endosomal marker suggested antigen internalization upon binding to cells expressing the FAM0-OVA BCR sensor (Figure 3C). To confirm FAM0-OVA beads internalization, BJAB cells transduced with LVs encoding the sensors were incubated for different times with SPAGs-OVA beads before surface staining of OVA molecules present at cell surface (Figures 3E and S6). The global percentage of beads-positive cells did not increase over incubation time, suggesting that all sensors were rapidly saturated.
Finally, we computed the internalization percentage as the ratio of cells positive for SPAGs-OVA beads to cells positive for SPAGs-OVA beads but negative for OVA surface staining (Figure 3E). Indeed, cells that are positive for SPAGs-OVA beads but negative for OVA surface staining correspond with cells that have internalized the OVA antigen, as no more OVA molecules are present (and thus stained) at the cell surface. In contrast, cells that are double positive for SPAGs-OVA beads as well as for surface OVA staining will not have internalized the BCR-antigen complex. This internalization percentage was already high after 1 h of incubation (approximately 80%) and slightly increased over incubation time (≤90%), hence validating the internalization of the sensor-antigen complex.
Assembly of sensor, transducer, and effector components into a functional synthetic circuit
We next sought to integrate the different part of the synthetic circuit, namely the sensor (membrane anchored BCR), the transducer (the 734-bp NR4A1-inducible promoter) and the effector (here, the TurboGFPdes). These components were introduced together in BJAB cells via a double transduction with two LVs encoding either the sensor in a constitutive manner or the TurboGFPdes under the control of the BCR-inducible promoter. Upon stimulation of double transduced cells through their ectopic sensor with anti-IgG molecules, we detected a specific upregulation of TurboGFPdes expression only in cells co-expressing the sensors and the BCR-inducible promoter (Figure 4A). Of note, the slight decrease of GFP expression in cells transduced with the inducible promoter along with sensors compared with cells without sensors after endogenous IgM BCR stimulation might be explained by the decrease of IgM expression in cells expressing an ectopic IgG BCR (Figure 3A).
Figure 4.
Antigen-specific activation of the synthetic circuit
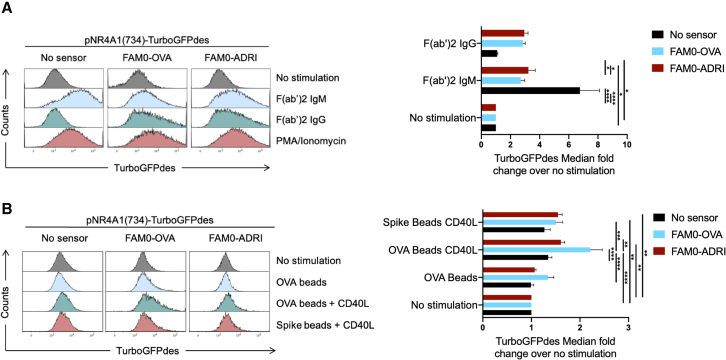
BJAB cells were transduced with LVs encoding pNR4A1(734)-TurboGFPdes 1 week before transduction with LVs encoding a membrane-anchored BCR directed against OVA (FAM0-OVA) or against HBs (FAM0-ADRI). Double transduced cells were then stimulated either with F(ab’)2 directed against IgM or IgG, or PMA/ionomycin as a control (A) or OVA/Spike-RBD-coated beads with or without CD40L (B) for 24 h before detection of TurboGFPdes fluorescence by flow cytometry. One representative overlay is presented along with the median fluorescence intensity quantified from five or six experiments (ANOVA 2 post hoc comparison with Tukey correction). p values under 0.05 were considered statistically significant and the following denotations were used: ∗∗∗, p<0.001; ∗∗, p<0.01; ∗, p<0.05.
Finally, we sought to validate the antigen-specific induction of the whole synthetic circuit by using OVA-coated beads alone or combined with the CD40L costimulatory molecule. We found that, after 24 h of co-stimulation, the circuit was specifically activated, by approximately 3-fold, in an antigen-specific manner since no activation was detected after stimulation with control spike RBD-coated beads or in cells expressing an irrelevant sensor (Figure 4B). Of note, the addition of CD40L was mandatory for the full activation of B cells, as it mimics the second stimulatory signal.
Development of a self-amplifying all-in-one vector
To transpose this approach toward a clinical setting that will be based on human primary B cells, we developed an all-in-one vector encoding all components within the same LV construct. Indeed, as the efficiency of transduction of human primary B cells remains relatively low at this time,28 such all-in-one LVs may increase the number of cells co-expressing all circuit components, thus decreasing variability. As a first generation of all-in-one-vectors, we constructed vectors with different combinations, in which the position and orientation of these promoters differed from each other (Figure S7A). For all LV designs, the B cell-specific FEEK promoter was used to drive the sensor expression, and the inducible promoter controlled the expression of the effector. However, we detected no expression of the sensor and no induction of the pNR4A1 promoter, which could be caused by promoter interferences (Figure S7).
To counteract this drawback, we next constructed a self-amplifying all-in-one-vector in which the inducible NR4A1 short promoter drives both the effector and sensor transgenes, using a T2A sequence between the two coding sequences (Figure 5A). Indeed, we expected a leakage activity of the inducible promoter, which could induce a basal sensor expression that is required to launch the circuit upon further stimulation (Figure S1). Thus, upon specific sensor stimulation, the inducible promoter would be fully activated, leading to effective expression of both the effector and sensor molecules, which would amplify the synthetic circuit response, hence creating a positive feedforward loop. Note that the insertion of the T2A motif after the effector and before the sensor open reading frames resulted in the addition of four amino acids fused to the TurboGFPdes marker, which may impair its destabilization and hence turnover. This could explain the higher basal fluorescence level in cells transduced with the self-amplifying construct as compared with cells expressing only the effector (Figure S8).29
Figure 5.
All-in-one vectors encoding a self-amplifying circuit
(A) Structure of self-amplifying vector. The inducible pNR4A1-734bp promoter fragment drives both the TurboGFPdes and the FAM0-OVA sensor transgenes with a T2A sequence in between. (B and E) Self-amplifying vector induction of effector expression in transduced BJAB cells after 24 h (B) or 48 h (E) stimulation with F(ab’)2 IgM or F(ab’)2 IgG. (C and F) Self-amplifying vector induction of sensor expression in transduced BJAB cells after 24 h (C) or 48 h (F) stimulation with F(ab’)2 IgM or F(ab’)2 IgG. The fold change of the mRNAs encoding the sensor was assessed by RT-qPCR after stimulation. (D and G) Self-amplifying vector induction of effector expression in transduced BJAB cells stimulated with OVA/Spike RBD-coated beads with or without CD40L for 24 h (D) or 48 h (G) before detection of TurboGFPdes fluorescence by flow cytometry. The median expression of TurboGFPdes was assessed by flow cytometry normalized by the median of unstimulated cells. ANOVA 2 post hoc comparison with Tukey correction (n = 5/6 for [B, D, E, G] and n = 3 for [C, F]). Error bars represent SEM. p values under 0.05 were considered statistically significant and the following denotations were used: ∗∗∗, p<0.001; ∗∗, p<0.01; ∗, p<0.05.
After 24 h of stimulation with anti-IgG molecules, we found that expression of both the TurboGFPdes and the sensor were upregulated in transduced BJAB cells, by 2- and 6-fold, respectively (Figures 5B and 5C). Importantly, we showed the antigen-specific induction of this self-amplifying synthetic circuit using OVA-coated beads (Figure 5D). Indeed, the levels of induction upon OVA-coated beads combined with CD40L stimulation was approximately 2-fold after 24 h and reached 4-fold after 48 h (Figures 5D–5G). Similarly, the upregulation of the effector and sensor lasted after 48 h of stimulation with anti-BCR antibodies (Figures 5E and 5F). The effector expression remained stable after 48 h of stimulation through the IgM BCR, but increased after IgG BCR stimulation as compared with 24 h of stimulation (from 4-fold after 24 h to 5-fold after 48 h), suggesting the initiation of a self-amplification (Figures 5E–5G). Yet, this induction was reversible; upon removal of sensor stimuli, the circuit was shut down in less than 3 days (Figure S9).
Overall, we demonstrate that the self-regulated construct allows specific expression of the effector upon sensor stimulation. Paving the way for clinical translational, we successfully introduced the self-regulated cassette in human primary B cells, but its antigen-specific induction remains to be confirmed (Figure S10).
Discussion
Here, we developed a new synthetic circuit for B cell reprogramming, using an ectopic sensor targeting a signal molecule whose binding induces the specific activation of an inducible promoter, driving the expression of effector proteins by modified B cells in a temporally restricted manner. In addition, this circuit exploits the endogenous properties of B cells, such as the BCR signaling cascade and their antigen presentation capacity, which enhances both their recognition and effectors functions.
We isolated a fragment of the NR4A1 promoter, which was successfully regulated upon BCR stimulation in a fully reversible manner (Figures 1 and 2). This reversibility would be extremely useful for the treatment of chronic diseases, which can have relapse episodes. Indeed, it is important to ensure that the expression of therapeutic molecules stops when the disease resolves, but also that the circuit can be reactivated in case of flares. This reactivation would be allowed by the persistence of B cells, which would have differentiated in memory cells. Hypothetically, this circuit should install a negative feedback loop where, upon disease clearance, the reprogrammed B cells are no longer exposed to antigens and, consequently, no effector is produced (Figure S8). Going further, another level of regulation was implemented by placing the sensor gene under the regulation of an inducible promoter (Figure 5). This is consistent with results obtained for Tet-ON and Tet-OFF regulation systems, as the use of a unique promoter to drive the regulatory module and the effector exhibited lower background activation and faster kinetics of induction as compared with the use of two distinct promoters.30,31,32 Yet, while this dynamic difference is expected, it is important to highlight that more complex genetic circuits with multiple inputs and outputs may increase the specificity of signal recognition and thus safety. For instance, such synthetic circuits have been implemented to trigger the expression of therapeutic molecules only if two defined pathological signals are detected, which leads to a more accurate discrimination of normal and pathological conditions.33,34 Similarly, placing the sensor under the regulation of an ectopic promoter that is specifically activated upon sensing of another pathological deviation could also be considered.35,36 Conversely, parts of our synthetic circuit may also be combined with other existing synthetic circuits or immunotherapy strategies such as CAR-T cells. For instance, the Nur77 protein is specifically expressed in T cells after T cell stimulation8 and after 24 h of T cell receptor stimulation with CD3 and CD28 coated beads, a 3-fold amplification in GFP expression could be observed in Jurkat T cells transduced with a LV encoding the GFP under the control of NR4A1 promoter (Figure S11). This opens the path for combination with CAR constructs for T cell therapies. Indeed, in the fourth generation of CAR T cells, therapeutic molecules are placed under an NFAT-sensitive promoter that is activated when the CAR binds its target.37,38,39 The NFAT promoter might be advantageously replaced by the NR4A1 ectopic promoter, which may ensure a higher induction specificity and a higher induction fold.
The induction fold and the leakage activity of the inducible promoter are key parameters to consider to fully implement synthetic circuits in the future. Although we observed a basal activity of the NR4A1 promoter with the GFP reporter, it was greatly decreased when implementing the TurboGFPdes as an effector. Likewise, the induction fold after BCR stimulation was five times higher with the TurboGFPdes as compared to the GFP. These discrepancies probably reflect the half-life differences of the two proteins, which is approximately 26 h for the GFP and 6 h for the TurboGFPdes (Figure S7). Indeed, of obvious importance for in vivo applications, is the half-life and concentration of the therapeutic effector proteins, as this is likely to determine the potential impact of the promoter leakage activity. Moreover, addressing the required local concentration of the effector would help to contextualize the number of modified cells to infuse to obtain a clinical effect. Of note, the stimulation of the ectopic BCR by its antigen should not only promote self-amplification of the circuit, but also reprogrammed B cell proliferation; thus, even if each B cell secretes a low amount of effector protein, the global concentration will increase.
Although our report demonstrates the functionality of our reprogramming synthetic circuit, its design can still be improved, notably at the level of the sensor component. While the introduction of an ectopic BCR decreased the expression of the endogenous BCR loci (Figure 3), there remain risks of signaling cross-talk and of chimeras between endogenous and ectopic light and heavy BCR chains. Of concern, these chimeras might have a different specificity and may target self-antigens, which could lead to an autoimmune reaction. To address this drawback, the CRISPR-Cas9 system could be used either to disrupt or to edit the endogenous BCR loci.40,41,42,43 Besides addressing the issue of chimera generation, editing of the BCR loci would also allow another level of regulation for our circuit. Indeed, upon BCR sensor stimulation, a part of edited mature B cells would differentiate into plasma cells and the sensor would become secreted, since the ratio of membrane-anchored immunoglobulins to secreted immunoglobulins decreases during B cell maturation.
The circuit described here is directed against a model antigen, but it can be switched to antigens in various applications, such as cancers, autoimmune disorders, transplantation rejection, allergies, and infectious diseases. Indeed, the main advantage of this approach lies in its full programmability in terms of recognized signals and output functions, which can be adapted to targeted diseases toward the same goal of continuous sensing of disease-specific biomarkers and triggering of physiological expression of therapeutic molecules in vivo. For instance, to treat cancers, a sensor targeting a tumor-specific antigen can be used to drive the expression of pro-inflammatory cytokines, such as IL-18, which will help to counteract the immunosuppressive tumor microenvironment and promote the differentiation of effector cells.44,45 Alternatively, Fas ligand could be used as effector molecule to directly induce cell death of CD95+ pathological cells. In addition, the presentation of tumor peptides after recognition and ectopic BCR/antigen internalization will drive the activation of antigen-specific T cells, which will help to destroy cancer cells.46,47 To promote T cell activation, co-stimulatory molecules, such as CD80 or CD86, might be implemented as effectors. Conversely, for the treatment of autoimmune disorders, such as rheumatoid arthritis, the signal molecule can be a self-peptide, and instead of secreting a pro-inflammatory cytokine, an anti-inflammatory mediator, such as IL-10, can be implemented as effector molecule. Locally, this protein may induce an immunosuppressive state, inhibiting immune responses and promoting the differentiation of regulatory cells.48 Paving the way for a preclinical proof of concept, a humanized mouse model platform has been recently developed to test B cell reprogramming therapies.49 Besides helping in the evaluation of the therapeutic effect, such an in vivo model will be of particular importance to assess the spatial pattern of the circuit activation as well as B cell homing to ensure that therapeutic molecules are only secreted locally, which would avoid the issues related to systemic drug delivery.
In the future, should such reprogramming circuits become routinely used in the clinics, libraries with different sensors or effectors might become available to adapt the treatment easily and at reduced costs to each patient. Moreover, implementing several sensors targeting different pathological signals would help to counteract resistances, which can be experienced with the current immune cell therapies.50,51,52 Finally, toward clinical applications, safety switches or suicide genes should be inserted in the construct to destroy infused cells in case of serious adverse events, such as cytokine release syndromes that are often experienced after modified immune cell transfer.53,54
Materials and methods
Construction of plasmids encoding the synthetic circuit components
The NR4A1 promoter fragments were amplified from human genomic DNA and cloned into the pHRSIN vector55 within EcoRI and BamHI restriction sites, to replace the SFFV promoter initially controlling a GFP transgene. The following primers were used.
| -2095 PrNR4A1 EcoRI | 5′-GATACGAATTCCAGTTCA AACGTGTTGTTCAAGGG-3′ |
| −1641 PrNR4A1 EcoRI | 5′-GATACGAATTCCAGGAGCA GAGATTAGGTGGCA-3′ |
| −1142 PrNR4A1 EcoRI | 5′-GATACGAATTCCAGGTGAT AACTGGTCAGAGCTG-3′ |
| −625 PrNR4A1 EcoRI | 5′-GATACGAATTCTGTGC ACTAGCTGCGCCTA-3′ |
| +109 PrNR4A1 BamHI | 5′-GATACGGATCCGGTC CCGCGTGCGCT-3′ |
The destabilized Turbo GFP was a kind gift of Dr. Mangeot and was cloned between BamHI and SbfI restriction sites after the promoters to replace GFP in the modified pHRSIN vectors above mentioned.
Fragments encoding the synthetic circuit transgenes for the all-in-one vectors and the self-amplifying vector were ordered from Genscript before being inserted by restriction cloning into in the original pHRSIN vector.55
For sensor cloning, fragments encoding the light chain (both the variable and constant domains) along with the variable domains of the heavy chain were synthesized by Genscript. The sequences encoding the variables region of monoclonal antibodies targeting either OVA or HBs (OBI sequence23 and ADRI-2F3 sequence22) were implemented. These fragments were then inserted in the original FAM0 construct to replace the variable regions initially targeting the glycoprotein of hepatitis C virus.24
Cell lines and primary cells
Jurkat cells (ACC-282) and Burkitt lymphoma cell lines
Namalwa cells subtype PNT (ACC-69), Raji (ACC-319) and BL-2 (ACC-625) were purchased from DSMZ (German Collection of Microorganisms and cell cultures GmbH). BJAB (ACC-757) and Ramos/B (ACC-603) cells, also originating from Burkitt lymphomas, were a kind gift of Pr. Belot and Dr. Gruffat. B cell lines were grown in culture flasks in RPMI-1640 medium containing 50 μg/mL penicillin and streptomycin supplemented with 10%–20% heat-inactivated fetal calf serum (FCS) at 37°C in a humidified atmosphere of 95% air/5% CO2 as recommended on the DSMZ website. We grew 293T cells (human kidney epithelium) in Dulbecco’s modified Eagle medium (Gibco, Invitrogen) supplemented with 10% FCS.
Peripheral blood mononuclear cells were isolated from peripheral blood of healthy human donors by Ficoll-gradient. Human primary B cells were then isolated by a positive selection with anti-CD19-conjugated magnetic beads (Miltenyi). They were grown in complete RPMI-1640 medium supplemented with 2 mM glutamine, 100 IU/mL penicillin, 100 mg/mL streptomycin, 55 μM B-ME, 1% HEPES, and 10% FCS at 1 × 106 cells/mL. Cross-linked CD40L (2 μg/mL, Miltenyi), IL-4 (2 ng/mL, Preprotech), and B cell-activating factor (BAFF, 10 ng/mL, Myltenyi) were added to the media.
Characterization of the EBV status
Total RNA from cells was extracted with TRI Reagent according to the manufacturer’s instructions (Molecular Research Center). RNAs were reverse-transcribed with SuperScript III reverse transcriptase kit (Invitrogen). The reverse-transcribed cDNA products were used to perform a PCR using the following specific oligonucleotides to amplify the lmp2a genome: forward (Lmp2aF: 5′- GGCGGTCACAACGGTCCTAACT) and reverse (Lmp2aR: 5′- CTACTCTCCACGGGATGZCTCAT) primers and HPRT, a housekeeping gene, with the following oligonucleotides: forward (5′-TCAGGCAGTATAATCCAAAGATGGT) and reverse (5′-AGTCTGGCTTATATCCAACACTTCG).
The PCR bands were visualized on propidium iodide-stained agarose gels
As an internal control of extraction, in vitro-transcribed exogenous RNAs from the linearized Triplescript plasmid pTRI-Xef (Invitrogen) were added into the samples before RNA extraction and quantified with specific primers (Xef-1a 970L20: 5′-CGACGTTGTCACCGGGCACG and Xef-1a 864U24: 5′-ACCAGGCATGGTGGTTACCTTTGC).
LV production and B cell transduction
LVs were generated by transient transfection of 293T cells through calcium phosphate precipitation. For pseudotyping of LVs with either the VSV-G and BRL glycoproteins, 2.7 μg and 7 μg envelope plasmid were respectively transfected together with 8.6 μg gagpol packaging plasmid (psPAX2, Addgene plasmid # 12260) and a plasmid encoding the LV construct (8.6 μg). Eighteen hours after transfection, the medium was replaced by Opti-MEM supplemented with 10 mM HEPES and 1% PenStrep (Gibco). Viral supernatants were harvested 48 h after transfection and filtered on 40-μm filters. Low-speed concentration was performed by overnight centrifugation of the viral supernatants at 3,000×g at 4°C.
LV titration
LVs were titrated by adding serial dilutions of the LVs to 293T target cells. Ten days after transduction, genomic DNA was extracted from target cells (Macherey Nagel) for qPCR analysis of viral genome copy number. The quantitative PCR was performed using 5 μL DNA on a StepOnePlus system with specific primers for detection of the integrated LV: primer F 5′-TGT GTG CCC GTC TGT TGT GT, primer R 5′-GAG TCC TGC GTC GAG AGA GC, and probe 5′-CAG TGG CGC CCG AAC AGG GA. Genomic vector copies in each sample were normalized to human actin gene copies using specific primers: primer F 5′ TCC GTG TGG ATC GGC GGC TCC A, primer R 5′-CTG CTT GCT GAT CCA CAT CTG, and probe CCT GGC CTC GCT GTC CAC CTT CCA. as previously described.24 The titers were normalized to human actin gene copies. Two control samples were processed in parallel for each run.
B cell transduction
For in vitro transduction of B cell lines, cells were cultured in supplemented RPMI-1640 medium and transduced at a MOI of 10. For transduction of primary human B cells, protamine sulfate was also added (8 μg/mL, Sigma Aldrich) to the media.
Cell stimulation
We stimulated 1.5 × 105 to 2.0 × 105 B cells in 96-well plates in supplemented RPMI-1640 medium with antibodies or immunostimulants during the indicated times. Stimuli included CpG oligodeoxynucleotides (ODN 2006) for B cells (InvivoGen), LPS from Escherichia coli (Sigma-Aldrich), anti-human F(ab’)2 IgM and F(ab’)2 IgG (Southern Biotech), ionomycin (EMD Millipore), PMA (Cell Signaling Technology), and cross-linked human CD40L (Miltenyi). After stimulation, cells were washed and resuspended in PBS before flow cytometry analysis. For antigen-specific stimulation, 100 antigen-coated beads per B cell were added in the supernatant.
For Jurkat T cells stimulation, 2 × 106 cells were plated in 96-well plates and stimulated with 1 μg/mL CD3/CD8 antibodies (Invitrogen) or with TransAct (Myltenyi Biotech) according to the manufacturer’s instructions. For kinetics assessments, to remove stimulant molecules, plates were washed three times with 200 μL PBS.
Generation of SPAG beads coated with OVA
SPAG beads were generated as previously described.56 Briefly, 0.4 μm-Flashred streptavidin beads (Bangslabs) were incubated with monobiotinylated OVA (Sigma) before being washed twice with PBS containing 2% BSA and filtered through 0.1-μm and 0.65-μm columns (Durapore, Merck Millipore) to remove respectively unbound molecules and bead aggregates. Beads concentration was assessed with a standard curve by reading FlashRed fluorescence on a Tecan plate reader.
Internalization assay
We plated 1 × 105 cells in 96-well plates and incubated for 1 h, 6 h, 24 h, or 48 h with 100 SPAGs-OVA per B cells. After incubation, cells were washed in PBS 2% FCS before surface staining with an anti-OVA antibody (Cell Signaling).
Confocal microscopy analysis
B cells loaded with SPAG beads were cultured overnight. We plated 1 × 106 B cells on 17-mm glass coverslips (Zeiss) preincubated for 4 h with 0.01% poly-L-lysine (Sigma). Cells were permeabilized with 0.1% Triton, then incubated for 30 min at room temperature with blocking solution (PBS-1% BSA) and stained with eFluor570 anti-B220 (clone RA3-6B2, BD) and AlexaFluor 488-conjugated anti-LAMP1 (clone H4A3, BD) mAbs for 1 h at room temperature. After three washes with PBS, cells were stained with Hoechst (1 μg/mL, Life Technology) for 5 min. After three additional washes, coverslips were mounted on glass slides with mowiol mounting medium. Confocal three-dimensional image stacks were acquired with confocal spectral LM610 (Zeiss). Images were analyzed with FIJI software.
Live microscopy
For live cell visualization, the medium was replaced with transparent RPMI containing 10% FCS and 1% penicillin-streptomycin in glass-bottom 96-well plates (Greiner). The chambered cover glass was placed in a temperature and CO2 controlled microscope stage insert upon a Spinning Disk Yokogawa CSU X1 confocal microscope equipped with a Q-Imaging Rolera EM-C2 camera (ENS de Lyon) with a 10× objective lens and illumination/filters appropriate for GFP visualization. Imaging was initiated as rapidly as possible following the addition of modulators and a picture was taken every 30 min for 72 h. Median fluorescence for each field was calculated for each time point using Fidji software.
Quantification of NanoLuciferase
B cells were stimulated as previously described. After stimulation, cells were counted using Trypan blue to exclude dead cells. Cells were lysed using passive lysis buffer (Promega) during 15 min at rom temperature before centrifugation at 400×g for 5 min. Supernatants were then mixed with assay buffer containing the NanoLuciferase substrate (from the NanoGlo kit, Promega) in p96 white plate and luminescence was read using a Mithras apparatus.
Flow cytometry staining and analysis
For staining, 2 × 105 cells were resuspended in PBS containing 2% FCS and incubated with an optimal dilution of fluorochrome conjugated antibodies anti-IgM-APC (Miltenyi), anti-IgG-PE (Miltenyi), anti-CD86-VB (Miltenyi), anti-HLA-DR-APC-Vio7 (Miltenyi), and anti-OVA-FITC (Cell Signaling Technology) for 30 min at 4°C before being washed with PBS complemented with 2% FCS.
Data were acquired on the FACSCantoII (BD Biosciences) and analyzed on FlowLogic.
Statistical analysis
Statistical significance was evaluated using Prism software. Data are expressed as means ± standard mean error and differences were considered as significant (∗p < 0.05), very significant (∗∗p < 0.01), or highly significant (∗∗∗p < 0.001).
Acknowledgments
We are grateful to all the members of our laboratory for their support and encouragement, especially Fouzia Amirache, Jean-François Bruxelles, and Clélia Giraudot. We thank Emily Sible for proofreading. We acknowledge the contribution of SFR Biosciences (UAR3444/CNRS, US8/Inserm, ENS de Lyon, UCBL) facilities: Vectorology, Platim, and Cytometry platforms. Our work is supported by funds from Janssen Horizon, THERA-B (PEPR Bioproduction), and PULSALYS. A.P. is supported by a fellowship from the ENS de Lyon and from the ARC Foundation.
Author contributions
A.P., F.F., and F.-L.C. designed the project. A.P. coordinated the project, designed, performed experiments, wrote the initial draft and revised subsequent drafts. M.D. and D.N. performed experiments. C.C. produced LVs. F.F. and F.-L.C. reviewed and edited the manuscript before submission. All authors have read and agreed to the published version of the manuscript.
Declaration of interests
The authors declare no conflict of interest.
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.omtn.2023.05.024.
Supplemental information
Data availability statement
The authors confirm that the data supporting the findings of this study are available within the article and its supplemental information.
References
- 1.Sedlmayer F., Aubel D., Fussenegger M. Synthetic gene circuits for the detection, elimination and prevention of disease. Nat. Biomed. Eng. 2018;2:399–415. doi: 10.1038/s41551-018-0215-0. [DOI] [PubMed] [Google Scholar]
- 2.Roybal K.T., Lim W.A. Synthetic immunology: hacking immune cells to expand their therapeutic capabilities. Annu. Rev. Immunol. 2017;35:229–253. doi: 10.1146/annurev-immunol-051116-052302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bugatti S., Vitolo B., Caporali R., Montecucco C., Manzo A. B cells in rheumatoid arthritis: from pathogenic players to disease biomarkers. BioMed Res. Int. 2014;2014:681678–681714. doi: 10.1155/2014/681678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Yuen G.J., Demissie E., Pillai S. B lymphocytes and cancer: a love–hate relationship. Trends Cancer. 2016;2:747–757. doi: 10.1016/j.trecan.2016.10.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Chabannon C., Bouabdallah R., Fürst S., Granata A., Saillard C., Vey N., Mokart D., Fougereau E., Lemarie C., Mfarrej B., et al. CAR-T cells : lymphocytes exprimant un récepteur chimérique à l’antigène. Rev. Med. Interne. 2019;40:545–552. doi: 10.1016/j.revmed.2018.12.002. [DOI] [PubMed] [Google Scholar]
- 6.Winkler J., Tittlbach H., Roesler W., Strobel J., Zimmermann R., Maas S., Spriewald B., Wolff D., Repp R., Kordelas L., et al. Adoptive transfer of purified donor-B-lymphocytes after allogeneic stem cell transplantation: results from a phase I/IIa clinical trial. Blood. 2016;128:502. doi: 10.1182/blood.V128.22.502.502. [DOI] [Google Scholar]
- 7.Tittlbach H., Schneider A., Strobel J., Zimmermann R., Maas S., Gebhardt B., Rauser G., Mach M., Mackensen A., Winkler T.H., Winkler J. GMP-production of purified human B lymphocytes for the adoptive transfer in patients after allogeneic hematopoietic stem cell transplantation. J. Transl. Med. 2017;15:228. doi: 10.1186/s12967-017-1330-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Ashouri J.F., Weiss A. Endogenous Nur77 is a specific indicator of antigen receptor signaling in human T and B cells. J.I. 2017;198:657–668. doi: 10.4049/jimmunol.1601301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Moran A.E., Holzapfel K.L., Xing Y., Cunningham N.R., Maltzman J.S., Punt J., Hogquist K.A. T cell receptor signal strength in Treg and iNKT cell development demonstrated by a novel fluorescent reporter mouse. J. Exp. Med. 2011;208:1279–1289. doi: 10.1084/jem.20110308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Tan C., Hiwa R., Mueller J.L., Vykunta V., Hibiya K., Noviski M., Huizar J., Brooks J.F., Garcia J., Heyn C., et al. NR4A nuclear receptors restrain B cell responses to antigen when second signals are absent or limiting. Nat. Immunol. 2020;21:1267–1279. doi: 10.1038/s41590-020-0765-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Winiarska M., Nowis D., Firczuk M., Zagozdzon A., Gabrysiak M., Sadowski R., Barankiewicz J., Dwojak M., Golab J. Selection of an optimal promoter for gene transfer in normal B cells. Mol. Med. Rep. 2017;16:3041–3048. doi: 10.3892/mmr.2017.6974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Girard-Gagnepain A., Amirache F., Costa C., Lévy C., Frecha C., Fusil F., Nègre D., Lavillette D., Cosset F.-L., Verhoeyen E. Baboon envelope pseudotyped LVs outperform VSV-G-LVs for gene transfer into early-cytokine-stimulated and resting HSCs. Blood. 2014;124:1221–1231. doi: 10.1182/blood-2014-02-558163. [DOI] [PubMed] [Google Scholar]
- 13.al Yacoub N., Romanowska M., Haritonova N., Foerster J. Optimized production and concentration of lentiviral vectors containing large inserts. J. Gene Med. 2007;9:579–584. doi: 10.1002/jgm.1052. [DOI] [PubMed] [Google Scholar]
- 14.Kumar V., Chaudhary N., Garg M., Floudas C.S., Soni P., Chandra A.B. Current diagnosis and management of immune related adverse events (irAEs) induced by immune checkpoint inhibitor therapy. Front. Pharmacol. 2017;8:49. doi: 10.3389/fphar.2017.00049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Huang L., Yuan Z., Liu P., Zhou T. Effects of promoter leakage on dynamics of gene expression. BMC Syst. Biol. 2015;9:16. doi: 10.1186/s12918-015-0157-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Donjerković D., Scott D.W. Activation-induced cell death in B lymphocytes. Cell Res. 2000;10:179–192. doi: 10.1038/sj.cr.7290047. [DOI] [PubMed] [Google Scholar]
- 17.Satoh T., Fukuda M., Sairenji T. Distinct patterns of mitogen-activated protein kinase phosphorylation and Epstein-Barr virus gene expression in Burkitt’s lymphoma cell lines versus B lymphoblastoid cell lines. Virus Gene. 2002;25:15–21. doi: 10.1023/A:1020113906966. [DOI] [PubMed] [Google Scholar]
- 18.Hasler P., Zouali M. Subversion of B lymphocyte signaling by infectious agents. Gene Immun. 2003;4:95–103. doi: 10.1038/sj.gene.6363941. [DOI] [PubMed] [Google Scholar]
- 19.Babcock G.J., Decker L.L., Volk M., Thorley-Lawson D.A. EBV persistence in memory B cells in vivo. Immunity. 1998;9:395–404. doi: 10.1016/S1074-7613(00)80622-6. [DOI] [PubMed] [Google Scholar]
- 20.Khan G., Miyashita E.M., Yang B., Babcock G.J., Thorley-Lawson D.A. Is EBV persistence in vivo a model for B cell homeostasis? Immunity. 1996;5:173–179. doi: 10.1016/S1074-7613(00)80493-8. [DOI] [PubMed] [Google Scholar]
- 21.Day R.N., Davidson M.W. The fluorescent protein palette: tools for cellular imaging. Chem. Soc. Rev. 2009;38:2887–2921. doi: 10.1039/b901966a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Cerino A., Bremer C.M., Glebe D., Mondelli M.U. A human monoclonal antibody against hepatitis B surface antigen with potent neutralizing activity. PLoS One. 2015;10 doi: 10.1371/journal.pone.0125704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Dougan S.K., Ogata S., Hu C.-C.A., Grotenbreg G.M., Guillen E., Jaenisch R., Ploegh H.L. IgG1+ ovalbumin-specific B-cell transnuclear mice show class switch recombination in rare allelically included B cells. Proc. Natl. Acad. Sci. USA. 2012;109:13739–13744. doi: 10.1073/pnas.1210273109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Fusil F., Calattini S., Amirache F., Mancip J., Costa C., Robbins J.B., Douam F., Lavillette D., Law M., Defrance T., et al. A lentiviral vector allowing physiologically regulated membrane-anchored and secreted antibody expression depending on B-cell maturation status. Mol. Ther. 2015;23:1734–1747. doi: 10.1038/mt.2015.148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Luo X.M., Maarschalk E., O’Connell R.M., Wang P., Yang L., Baltimore D. Engineering human hematopoietic stem/progenitor cells to produce a broadly neutralizing anti-HIV antibody after in vitro maturation to human B lymphocytes. Blood. 2009;113:1422–1431. doi: 10.1182/blood-2008-09-177139. [DOI] [PubMed] [Google Scholar]
- 26.Gottwick C., He X., Hofmann A., Vesper N., Reth M., Yang J. A symmetric geometry of transmembrane domains inside the B cell antigen receptor complex. Proc. Natl. Acad. Sci. USA. 2019;116:13468–13473. doi: 10.1073/pnas.1907481116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Sicard A., Levings M.K., Scott D.W. Engineering therapeutic T cells to suppress alloimmune responses using TCR s, CAR s, or BAR s. Am. J. Transplant. 2018;18:1305–1311. doi: 10.1111/ajt.14747. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Frecha C., Costa C., Lévy C., Nègre D., Russell S.J., Maisner A., Salles G., Peng K.-W., Cosset F.-L., Verhoeyen E. Efficient and stable transduction of resting B lymphocytes and primary chronic lymphocyte leukemia cells using measles virus gp displaying lentiviral vectors. Blood. 2009;114:3173–3180. doi: 10.1182/blood-2009-05-220798. [DOI] [PubMed] [Google Scholar]
- 29.Szymczak A.L., Vignali D.A.A. Development of 2A peptide-based strategies in the design of multicistronic vectors. Expet Opin. Biol. Ther. 2005;5:627–638. doi: 10.1517/14712598.5.5.627. [DOI] [PubMed] [Google Scholar]
- 30.Centlivre M., Zhou X., Pouw S.M., Weijer K., Kleibeuker W., Das A.T., Blom B., Seppen J., Berkhout B., Legrand N. Autoregulatory lentiviral vectors allow multiple cycles of doxycycline-inducible gene expression in human hematopoietic cells in vivo. Gene Ther. 2010;17:14–25. doi: 10.1038/gt.2009.109. [DOI] [PubMed] [Google Scholar]
- 31.Huang Y., Zhen R., Jiang M., Yang J., Yang Y., Huang Z., Lin Y. Development of all-in-one multicistronic Tet-On lentiviral vectors for inducible co-expression of two transgenes: all-in-One Multicistronic Tet-On Lentiviral Vectors. Biotechnol. Appl. Biochem. 2015;62:48–54. doi: 10.1002/bab.1239. [DOI] [PubMed] [Google Scholar]
- 32.Ogueta S.B., Yao F., Marasco W.A. Design and in vitro characterization of a single regulatory module for efficient control of gene expression in both plasmid DNA and a self-inactivating lentiviral vector. Mol. Med. 2001;7:569–579. doi: 10.1007/BF03401863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Schukur L., Geering B., Charpin-El Hamri G., Fussenegger M. Implantable synthetic cytokine converter cells with AND-gate logic treat experimental psoriasis. Sci. Transl. Med. 2015;7:318ra201. doi: 10.1126/scitranslmed.aac4964. [DOI] [PubMed] [Google Scholar]
- 34.Roybal K.T., Williams J.Z., Morsut L., Rupp L.J., Kolinko I., Choe J.H., Walker W.J., McNally K.A., Lim W.A. Engineering T cells with customized therapeutic response programs using synthetic notch receptors. Cell. 2016;167:419–432.e16. doi: 10.1016/j.cell.2016.09.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Morsut L., Roybal K.T., Xiong X., Gordley R.M., Coyle S.M., Thomson M., Lim W.A. Engineering customized cell sensing and response behaviors using synthetic notch receptors. Cell. 2016;164:780–791. doi: 10.1016/j.cell.2016.01.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Daringer N.M., Dudek R.M., Schwarz K.A., Leonard J.N. Modular extracellular sensor architecture for engineering mammalian cell-based devices. ACS Synth. Biol. 2014;3:892–902. doi: 10.1021/sb400128g. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Chmielewski M., Abken H. TRUCKs: the fourth generation of CARs. Expet Opin. Biol. Ther. 2015;15:1145–1154. doi: 10.1517/14712598.2015.1046430. [DOI] [PubMed] [Google Scholar]
- 38.Webster B., Xiong Y., Hu P., Wu D., Alabanza L., Orentas R.J., Dropulic B., Schneider D. Self-driving armored CAR-T cells overcome a suppressive milieu and eradicate CD19+ Raji lymphoma in preclinical models. Mol. Ther. 2021;29:2691–2706. doi: 10.1016/j.ymthe.2021.05.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Uchibori R., Teruya T., Ido H., Ohmine K., Sehara Y., Urabe M., Mizukami H., Mineno J., Ozawa K. Functional analysis of an inducible promoter driven by activation signals from a chimeric antigen receptor. Mol. Ther. Oncolytics. 2019;12:16–25. doi: 10.1016/j.omto.2018.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Voss J.E., Gonzalez-Martin A., Andrabi R., Fuller R.P., Murrell B., McCoy L.E., Porter K., Huang D., Li W., Sok D., et al. Reprogramming the antigen specificity of B cells using genome-editing technologies. Elife. 2019;8 doi: 10.7554/eLife.42995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Moffett H.F., Harms C.K., Fitzpatrick K.S., Tooley M.R., Boonyaratanakornkit J., Taylor J.J. B cells engineered to express pathogen-specific antibodies protect against infection. Sci. Immunol. 2019;4 doi: 10.1126/sciimmunol.aax0644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Greiner V., Bou Puerto R., Liu S., Herbel C., Carmona E.M., Goldberg M.S. CRISPR-mediated editing of the B cell receptor in primary human B cells. iScience. 2019;12:369–378. doi: 10.1016/j.isci.2019.01.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Nahmad A.D., Raviv Y., Horovitz-Fried M., Sofer I., Akriv T., Nataf D., Dotan I., Carmi Y., Burstein D., Wine Y., et al. Engineered B cells expressing an anti-HIV antibody enable memory retention, isotype switching and clonal expansion. Nat. Commun. 2020;11:5851. doi: 10.1038/s41467-020-19649-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Suzuki N., Chen N.-J., Millar D.G., Suzuki S., Horacek T., Hara H., Bouchard D., Nakanishi K., Penninger J.M., Ohashi P.S., Yeh W.C. IL-1 receptor-associated kinase 4 is essential for IL-18-mediated NK and Th1 cell responses. J. Immunol. 2003;170:4031–4035. doi: 10.4049/jimmunol.170.8.4031. [DOI] [PubMed] [Google Scholar]
- 45.Zhou T., Damsky W., Weizman O.-E., McGeary M.K., Hartmann K.P., Rosen C.E., Fischer S., Jackson R., Flavell R.A., Wang J., et al. IL-18BP is a secreted immune checkpoint and barrier to IL-18 immunotherapy. Nature. 2020;583:609–614. doi: 10.1038/s41586-020-2422-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Wennhold K., Thelen M., Schlößer H.A., Haustein N., Reuter S., Garcia-Marquez M., Lechner A., Kobold S., Rataj F., Utermöhlen O., et al. Using antigen-specific B cells to combine antibody and T cell–based cancer immunotherapy. Cancer Immunol. Res. 2017;5:730–743. doi: 10.1158/2326-6066.CIR-16-0236. [DOI] [PubMed] [Google Scholar]
- 47.Page A., Hubert J., Fusil F., Cosset F.-L. Exploiting B cell transfer for cancer therapy: engineered B cells to eradicate tumors. IJMS. 2021;22:9991. doi: 10.3390/ijms22189991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Mauri C., Gray D., Mushtaq N., Londei M. Prevention of arthritis by interleukin 10–producing B cells. J. Exp. Med. 2003;197:489–501. doi: 10.1084/jem.20021293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Page A., Laurent E., Nègre D., Costa C., Pierre V., Defrance T., Cosset F.-L., Fusil F. Efficient adoptive transfer of autologous modified B cells: a new humanized platform mouse model for testing B cells reprogramming therapies. Cancer Immunol. Immunother. 2021;71:1771–1775. doi: 10.1007/s00262-021-03101-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Aldea M., Andre F., Marabelle A., Dogan S., Barlesi F., Soria J.-C. Overcoming resistance to tumor-targeted and immune-targeted therapies. Cancer Discov. 2021;11:874–899. doi: 10.1158/2159-8290.CD-20-1638. [DOI] [PubMed] [Google Scholar]
- 51.Conde E., Vercher E., Soria-Castellano M., Suarez-Olmos J., Mancheño U., Elizalde E., Rodriguez M.L., Glez-Vaz J., Casares N., Rodríguez-García E., et al. Epitope spreading driven by the joint action of CART cells and pharmacological STING stimulation counteracts tumor escape via antigen-loss variants. J. Immunother. Cancer. 2021;9 doi: 10.1136/jitc-2021-003351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Sokolove J., Bromberg R., Deane K.D., Lahey L.J., Derber L.A., Chandra P.E., Edison J.D., Gilliland W.R., Tibshirani R.J., Norris J.M., et al. Autoantibody epitope spreading in the pre-clinical phase predicts progression to rheumatoid arthritis. PLoS One. 2012;7 doi: 10.1371/journal.pone.0035296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Kotter B., Engert F., Krueger W., Roy A., Rawashdeh W.A., Cordes N., Drees B., Webster B., Werchau N., Lock D., et al. Titratable pharmacological regulation of CAR T cells using zinc finger-based transcription factors. Cancers. 2021;13:4741. doi: 10.3390/cancers13194741. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Li H.-S., Wong N.M., Tague E., Ngo J.T., Khalil A.S., Wong W.W. High-performance multiplex drug-gated CAR circuits. Cancer Cell. 2022 doi: 10.1016/j.ccell.2022.08.008. S1535610822003725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Demaison C., Parsley K., Brouns G., Scherr M., Battmer K., Kinnon C., Grez M., Thrasher A.J. High-level transduction and gene expression in hematopoietic repopulating cells using a human imunodeficiency virus type 1-based lentiviral vector containing an internal spleen Focus forming virus promoter. Hum. Gene Ther. 2002;13:803–813. doi: 10.1089/10430340252898984. [DOI] [PubMed] [Google Scholar]
- 56.Sicard A., Koenig A., Graff-Dubois S., Dussurgey S., Rouers A., Dubois V., Blanc P., Chartoire D., Errazuriz-Cerda E., Paidassi H., et al. B cells loaded with synthetic particulate antigens: a versatile platform to generate antigen-specific helper T cells for cell therapy. Nano Lett. 2016;16:297–308. doi: 10.1021/acs.nanolett.5b03801. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The authors confirm that the data supporting the findings of this study are available within the article and its supplemental information.