Abstract
Saccharomyces cerevisiae DNA polymerase IV (Pol4) like its homolog, human DNA polymerase lambda (Polλ), is involved in Non-Homologous End-Joining and Microhomology-Mediated Repair. Using genetic analysis, we identified an additional role of Pol4 also in homology-directed DNA repair, specifically in Rad52-dependent/Rad51-independent direct-repeat recombination. Our results reveal that the requirement for Pol4 in repeat recombination was suppressed by the absence of Rad51, suggesting that Pol4 counteracts the Rad51 inhibition of Rad52-mediated repeat recombination events. Using purified proteins and model substrates, we reconstituted in vitro reactions emulating DNA synthesis during direct-repeat recombination and show that Rad51 directly inhibits Polδ DNA synthesis. Interestingly, although Pol4 was not capable of performing extensive DNA synthesis by itself, it aided Polδ in overcoming the DNA synthesis inhibition by Rad51. In addition, Pol4 dependency and stimulation of Polδ DNA synthesis in the presence of Rad51 occurred in reactions containing Rad52 and RPA where DNA strand-annealing was necessary. Mechanistically, yeast Pol4 displaces Rad51 from ssDNA independent of DNA synthesis. Together our in vitro and in vivo data suggest that Rad51 suppresses Rad52-dependent/Rad51-independent direct-repeat recombination by binding to the primer-template and that Rad51 removal by Pol4 is critical for strand-annealing dependent DNA synthesis.
Graphical Abstract
Graphical Abstract.
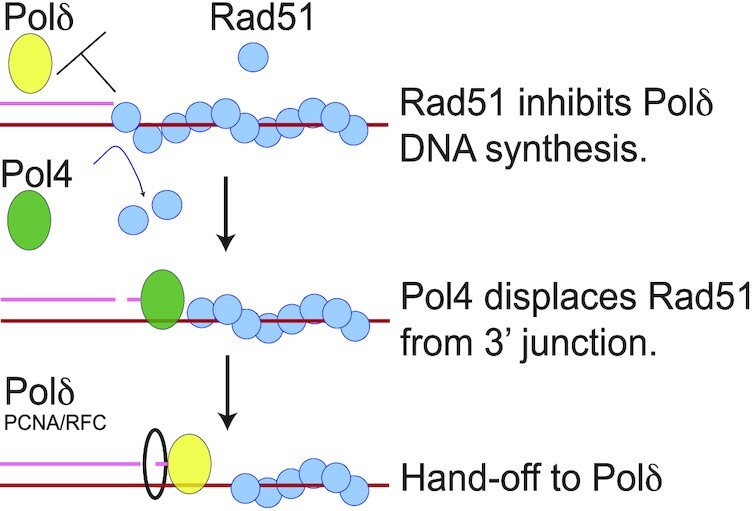
Rad51 inhibits DNA synthesis by Pold (use Greek delta for last letter) during Rad52-mediated repeat recombination. Poll (use Greek lambda for last letter) can displace Rad51 to allow PCNA loading and Pold (use Greel for last letter) DNA synthesis.
INTRODUCTION
Homology-directed repair (HDR) capitalizes on the accuracy of Watson–Crick base-pairing to maintain genomic stability (1). In the context of DNA double-strand break (DSB) repair, two major pathways are operational: homologous recombination (HR) in which Rad51 mediates homology search and DNA strand invasion, and single-strand annealing (SSA), in which Rad52 mediates the pairing of single-stranded DNAs bound by the ssDNA binding protein RPA (2–4). In yeast, both processes—HR and SSA—depend on Rad52, as this protein also serves as the mediator for Rad51 filament formation (5,6). In mammals and other organisms, BRCA2 is the key mediator protein for RAD51 in HR, and hence the process is largely RAD52-independent (7). HR typically utilizes the homology on the sister chromatid as a template, leading to high-fidelity repair (8,9). In contrast, SSA requires proximal repeated DNA sequences on the same chromatid or different chromosomes leading to associated mutations (deletion, translocation, inversion) depending on the exact position and orientation of the DNA repeats (3). SSA was discovered and the term coined studying DSB-associated recombination events of extrachromosomal DNA in mammalian cells (10,11). Both processes, HR and SSA, depend on DSB end resection, which generates 3’ending ssDNA tails of sufficient length to be bound by RPA (12). During HR, RPA is displaced by the Rad51 filament which conducts homology search and DNA strand invasion upon finding a suitable donor, forming a displacement loop (D-loop) (2). After extension of the invading 3’-OH end in the D-loop by DNA polymerase delta (Polδ), different pathways can lead to the formation of intact repaired chromosomes (4,13). Alternatively, SSA can also occur following DNA end resection but only when resection exposes complementary ssDNA with extensive homology. While spontaneous annealing is suppressed by RPA binding to ssDNA, RAD52 has the unique ability to reanneal RPA-coated ssDNA (14). SSA leads to the deletion of one repeat and the intervening sequence between the repeats (3). The occurrence of such SSA-mediated deletion events has been documented in the human genome generating functionally impactful small deletions, for example in the BRCA1 gene (15). SSA can also operate between repeats residing on different chromosomes/chromatids leading to genomic rearrangements (16,17). Similarly, HR can operate between repeated DNA sequences located across the genome, a process referred to as non-allelic homologous recombination, or engage multiple distinct repeats in a single pairing event to form a multi-invasion which can likewise lead to genome rearrangements (18–20). It is often difficult to infer from product analysis which HDR pathway was involved without additional information, for example on genetic dependencies.
HDR competes with end-joining processes for the repair of DSBs, and pathway competition is regulated to a large degree by the cell cycle-regulated control of DSB resection to ensure that HR takes place only in S/G2 when a sister chromatid is available (8,12). Classic non-homologous end-joining (NHEJ), the dominant end-joining pathway dependent on DNA ligase 4, is able to rejoin two ends of a DSB in the absence of resection and operates during all phase of the cell cycle (21). Alternative end-joining pathways were initially identified to operate when NHEJ is genetically impaired though recent studies demonstrate their importance also in NHEJ-proficient cells (21–25). One such pathway is DNA polymerase theta (Polθ)-mediated end-joining (TMEJ) which is of particular importance, as it can utilize resected DSB ends thereby assuming a critical function in HDR-deficient cells (24,26,27). This process often involves small homologies, and hence has also been termed micro-homology mediated end-joining (21,24). There is evidence for additional end-joining processes beyond TMEJ and NHEJ (21,24). Moreover, not all organisms have Polθ in their DNA polymerase repertoire; for example, the budding yeast Saccharomyces cerevisiae lacks Polθ but clearly shows NHEJ-independent repair involving microhomologies (13,28,29). For this reason, we utilize the term micro-homology-mediated repair (MHMR) here. How DSB repair pathway choice is regulated at resected DSBs is not fully understood (12,30). It is unclear but often presumed that resection for HDR is more extensive than for MHMR including TMEJ. Resection length is of importance, as RPA has a binding site size of about 30 nt for high-affinity binding of ssDNA, leaving the possibility of short overhangs that are devoid of RPA or engaged in a lower affinity binding mode (31–33). Recent evidence suggests that TMEJ is actively suppressed in the S/G2 phases and only active in M phase for one-sided DSBs induced in S-phase or IR-induced DSBs. Such cell cycle-specific regulation suppresses TMEJ-mediated chromosomal rearrangements (23,34).
HDR is also involved in the recovery of replication forks stalled at spontaneous DNA damage (35,36). Genetically this process can be monitored by direct repeat recombination (DRR) (37,38). DRR can be mediated by multiple mechanisms such as SSA, unequal sister chromatid recombination, intra-sister recombination, and others. While SSA requires an initiating DSB, other spontaneous lesions may induce the alternate mechanisms, and the initiating lesion is difficult to pinpoint. Based on genetic analysis of separation-of-function mutants in RAD52, it was shown that most spontaneous lesions inducing recombination are not DSBs (39). For spontaneous interhomolog crossovers, two independent lines of evidence support DSBs as the predominant initiating lesion (40,41). This suggests that the selected endpoint and assay system are differentially affected by the specific initiating lesion. Spontaneous DRR events are context dependent and can be fully or partially dependent on Rad51 or inhibited by Rad51, but in yeast all depend on Rad52 (37,38). The Rad52 requirement reflects its ability to anneal RPA-coated ssDNA which can occur in the context of DSB-initiated SSA or independent of DSBs (14). How stalled replication forks are processed is not fully understood but of great importance as fork stalling is a prevalent response to DNA damage and associated with genomic instability (36,42).
Most DNA repair processes require some form of DNA synthesis, and this step is critical for its accuracy (13,43). Although known for being highly accurate, DSB repair by HDR is far from error-free and associated with a 100–1000-fold increase in mutagenesis surrounding the DSB site (13,44,45). The various end-joining processes are even more error-prone (21,22,24,25,28). DNA polymerases are grouped into four different families: A, B, Y and X, according to their evolutionary conservation (46). DNA polymerases from all families have been reported to be involved in DSB repair, and their intrinsic error rates contribute to the overall associated mutagenesis (13,46). While evidence shows that the B family Polδ is central to HR-associated DNA synthesis, end-joining pathways are most commonly associated with DNA polymerases from the X family (13,46). Interestingly, Polδ is also involved in MHMR-associated DNA synthesis in budding yeast and human cells suggesting that multiple DNA polymerases cooperate during DSB repair events (47–49).
S. cerevisiae DNA polymerase IV (Pol4) is a member of the X family of DNA polymerases and an ortholog to DNA polymerase lambda (Polλ) in humans (46,50). The polymerases from the X family are characterized by their nucleotidyl transferase activity including a template-independent terminal deoxynucleotidyl transferase activity (51,52). Pol X family enzymes lack a proofreading subdomain, and their intrinsic DNA synthesis fidelity is much lower than replicative polymerases such as Polδ (50,52). Budding yeast Pol4 is a 68-kDa single-subunit protein first identified based on sequence similarity to Polβ (53,54). The purified protein exhibits distributive DNA synthesis activity on a 3′-recessed primer-template substrate and processive DNA synthesis on substrates that have a one to five nucleotide gap (53,54). Pol4 has a major function in NHEJ and MHMR, and in vivo studies revealed that this polymerase is not essential for cell viability (47,55–60). A possible function of yeast Pol4 and mammalian Polλ in HDR was inferred from their induced expression during meiotic prophase when meiotic recombination takes place (54,61–63). In addition, the yeast Pol4 mutant was described to exhibit a meiotic hyper-recombination phenotype (63). Consistent with a role in HR, the presence of the human Polλ allele R438W was associated with a defect in sister chromatid exchange and sensitivity to the topoisomerase inhibitor camptothecin in humans (64). Although a role in HDR appeared plausible, there are conflicting results regarding Pol4 function in the repair of MMS-induced DNA damage and the genetic analysis of POL4 still remains incomplete (63,65). Furthermore, there is no biochemical information on the function of Pol4 in reconstituted reactions that would support a role of Pol4 role in HDR.
In order to determine the role of Pol4 in HDR, we investigated its role in various genetic assays and identified a requirement for Pol4 in Rad52-dependent and Rad51-independent DRR. We found that Pol4 is only required when Rad51 is present and the need for Pol 4 is suppressed by the absence of Rad51. In vitro, we showed that Rad51 inhibits Polδ-mediated DNA synthesis and that Pol4 overcomes Rad51 inhibition of Polδ DNA synthesis, consistent with the genetic data. We observed a difference in the decrease in DRR frequencies in the absence of Pol4 or when using a catalytic deficient version of the polymerase. These results suggested a dual role of Pol4 in DRR, one dependent and one independent on DNA synthesis, for which we also provide biochemical evidence, identifying a stronger requirement for Pol4-mediated DNA synthesis with substrates containing a single nucleotide 3’ flap.
MATERIALS AND METHODS
Yeast strains and plasmids
Standard techniques for yeast growth and genetic manipulation were used (66). All yeast strains used in this study are in the W303-1A background and WT RAD5 (Supplementary Table S1). The pol4::KANMX (pol4Δ), pol4-D367E (pol4-CD) and rad52::TRP1 (rad52Δ) mutants were described previously (47,59). The rad51::NATMX (rad51Δ) mutant was generated by PCR amplification of NATMX using pWDH1018 as a template. See Supplementary Table S2 for all plasmids used. The primers contain 50 nucleotides on their 5’-end, which share homologies to sequences upstream and downstream of the RAD51 ORF with 21 nt homology to pWDH1018 to amplify NATMX4 (67) (olWDH2120, olWDH2121; see Supplementary Table S3 for all oligonucleotides and their sequences). The deletion of the RAD51 ORF was verified by PCR in the transformant. Plasmids bearing GST-fusions containing a thrombin site with either POL4, pol4-CD, hPOLL or hPOLL-CD were created by cloning the ORF of each allele into the empty vector pWDH403 (68). Both POL4 and pol4-CD were amplified using PCR from genomic DNA preparation with the appropriate genotype using olWDH924 and olWDH932 that contain a BamHI and XhoI restriction site, respectively. The POL4 or pol4-CD PCR fragments were digested with BamHI and XhoI, mixed separately with pWDH403 digested with BamHI and XhoI, ligated, and verified through sequencing to generate pWDH1145 (POL4) and pWDH1147 (pol4-CD). The hPOLL ORF cloned into pBluescript was purchased from ATCC (#10624945), PCR amplified using olWDH950 and olWDH952 containing XhoI and HindIII restriction sites, respectively. Creation of the hPOLL-CD allele was achieved using site-directed mutagenesis with the oligonucleotides olWDH957 and olWDH958, which was verified through DNA sequencing of the entire ORF. The hPOLL-CD allele was PCR amplified using olWDH950 and olWDH967 containing XhoI and BglII restriction sites, respectively. Finally, the hPOLL or hPOLL-CD PCR fragments were digested with XhoI and HindIII/BglII, mixed separately with pWDH403 digested with XhoI and HindIII/BglII, ligated, and verified through DNA sequencing to generate pWDH1146 (hPOLL) and pWDH1148 (hPOLL-CD).
Ionizing radiation sensitivity assay
Haploid cells of the appropriate genotype were selected from freshly dissected segregants and used to inoculate a minimum of three 5 mL YPD (1% yeast extract, 2% bacto peptone, and 2% dextrose) liquid cultures. Cultures were grown at 30°C until mid-log when cells were collected through centrifugation, washed, resuspended in distilled water, counted by hemocytometer, diluted, and approximately 500 cells were plated onto YPD. Cells were exposed to varying doses of ionizing radiation using a 137Cs source and allowed to grow at 30°C for 4 days before counting colonies. Percent survival was determined by dividing the number of colonies on each plate by 500, normalizing to the cells not exposed to ionizing radiation, and multiplying by 100. Extrachromosomal vectors containing POL4 (pWDH1145), pol4-CD (pWDH1147), hPOLL (pWDH1146), or hPOLL-CD (pWDH1148) or empty vector (WDH403) were transformed into freshly dissected segregants of the appropriate genotype and transformants were selected by growing cells on synthetic medium lacking uracil. Fresh transformants were used as indicated above to determine percent survival following exposure to varying ionizing radiation doses. This procedure was repeated a minimum of three times for each genotype and the mean ± SEM was determined using Prism (GraphPad, San Diego, CA, USA).
Interhomolog allelic recombination assay
The interhomolog recombination (IHR) rate was determined from haploid strains containing either ade2-a or ade2-n alleles on chromosome XV, which were crossed to generate ade2-a/ade2-n heterozygous diploids. A minimum of three independent diploids were generated for each genotype tested and single colonies were used to inoculate 5 mL of YPD medium. Cultures were grown overnight at 30°C before appropriate dilutions were used to plate onto YPD and synthetic medium lacking adenine to determine viability and IHR, respectively. Plates were incubated at 30°C for 4 days before counting colonies. IHR rates and ±95% confidence intervals were determined using the method of the median (69) from a minimum of nine independent cultures for each genotype.
Meiotic crossover frequency and distribution
The meiotic crossover frequency and distribution across chromosome III was determined as previously described (70). Briefly, diploids of the appropriate genotype were grown on sporulation medium a minimum of 48 h, asci were treated with zymolyase, and micro-dissected on YPD before marker analysis was performed. A minimum of one-hundred, four spore, tetrads were analyzed for each genotype and map distances (cM) were calculated for each genetic interval using Perkins equation: (T + 3NPD/total × 100). Standard error and statistical differences in each crossover interval between wildtype and pol4 were calculated using Stahl Lab Online Tools (https://elizabethhousworth.com/StahlLabOnlineTools/).
Unequal sister-chromatid recombination assay
The rates of unequal sister-chromatid recombination (USCR) were determined as described previously (71). At least nine cultures from freshly dissected segregants of the appropriate genotype were grown overnight at 30°C. Cell density was determined by OD600 using a dilution of the overnight culture, and a near equivalent density of cells for each culture and genotype were diluted and plated to YPD and synthetic medium lacking histidine to determine viable cells and His+ recombinants, respectively. Plates were incubated at 30°C for 4 days before counting colonies. Median rate of USCR and ± 95% confidence intervals was determined from a minimum of nine independent segregants for each genotype using the method of the median (69).
ADE2 direct-repeat recombination and gene conversion assay
Haploid cells of the appropriate genotype were selected from at least nine freshly dissected segregants, possessing the direct-repeat ade2 alleles, ade2-n and ade2-a, separated by URA3 and used to inoculate nine separate 5 ml YPD liquid cultures as described previously (72). Cultures were grown overnight at 30°C, cell density was determined by OD600 using a dilution of the overnight culture, and a near equivalent density of cells for each culture and genotype were diluted and plated to YPD and synthetic medium lacking uracil to determine viable cells and Ade+ recombinants, respectively. Plates were incubated at 30°C for 4 days before counting colonies. Cells that were white (Ade+), but Ura− were recorded as direct-repeat recombinants, while cells that were white (Ade+) and Ura+ were recorded as gene conversion. Median rates of direct-repeat recombination or gene conversion and ±95% confidence intervals were determined from a minimum of nine independent segregants for each genotype using the method of the median (69).
HIS3 direct-repeat recombination assay
Haploid cells of the appropriate genotype were selected from at least nine freshly dissected segregants, possessing the direct-repeat truncated his3 alleles sharing 103 bp or 415 bp of homology, and used to inoculate nine separate 5 ml YPD liquid cultures as described previously (73). Cultures were grown overnight at 30°C, cell density was determined by OD600 using a dilution of the overnight culture, and a near equivalent density of cells for each culture and genotype were diluted and plated to YPD and synthetic medium lacking histidine to determine viable cells and His+ recombinants, respectively. Plates were incubated at 30°C for 4 days before counting colonies. Median rate of direct-repeat recombination and ±95% confidence intervals was determined from a minimum of nine independent segregants for each genotype using the method of the median (69).
Interchromosomal SSA assay
Interhomolog SSA frequency was determined from diploid strains containing both his3Δ3’ and his3Δ5’ alleles sharing 311 bp homology on chromosome XV and III, respectively (16). A minimum of three independent diploids were generated for each genotype tested and single colonies were used to inoculate 2 ml of YP-Raffinose medium as described previously (74). Cultures were grown overnight at 30°C before the addition of galactose to a final concentration of 2% and incubated for an additional 4 h, which activated expression of HO endonuclease and DSB formation next to his3Δ3’. Following DSB induction, appropriate dilutions were used to plate onto YPD and synthetic medium lacking histidine to determine viability and translocation frequency, respectively. Plates were incubated at 30°C for four days before counting colonies. Translocation frequencies and ±95% confidence intervals were determined using the method of the median (69) from a minimum of nine independent cultures for each genotype.
Proteins
Native yeast Rad51 and RPA, and Rad52-His6 were purified as previously described (75,76). Yeast Polδ, His-tagged RFC and native PCNA were obtained as a generous gift from Dr Peter Burgers (76,77). N-terminally GST-tagged human DNA Polλ and yeast DNA Pol4 (WT, Pol4-CD) were purified as follows: S. cerevisiae strains carrying expression plasmids with GST + PreScission-fusion versions of Pol4, Pol4-CD and hPolλ in an inducible expression system were obtained by cloning an in-frame XhoI/BamHI digest of Pol4, Pol4-CD or hPolλ ORFs into a XhoI/BamHI digested expression plasmid (pWDH597) creating pWDH1172, pWDH1173, pWDH1174, respectively. Protein expression was induced by the addition of galactose to a final concentration of 2% within the culture followed by incubation at 30°C for 3 h. After that, cells were harvested by centrifugation and the pellet stored at –80°C. All purification steps were carried out at 4°C. The cell pellet was thawed in ice and cell lysate was obtained by bead beating in PBS buffer supplemented with 500 mM of NaCl and protease inhibitors (2 ml/l 0.1 PMSF, 1 ml/l 2 mM Leupeptin, 2 ml/l 1mM Pepstatin A, 350 ul/l β-mercaptoethanol). Cell lysate was cleared by centrifugation at 40 000 rpm for 45 min using the 50.2-T1 rotor in the Beckman Optima LE-80K ultracentrifuge. The clarified lysate was mixed with 20 ml of glutathione-sepharose 4B (GE) and the mix was incubated overnight in order to maximize protein-resin binding. After that, the mix was transferred into a column, and the resin-protein bound washed extensively in PBS supplemented with 500 mM of NaCl and protease inhibitors (2 ml/l 0.1 PMSF, 1 ml/l 2 mM Leupeptin, 2 ml/l 1 mM Pepstatin A, 350 ul/l β-mercaptoethanol). Proteins were eluted by cleavage using PreScission protease [109]. Briefly, 5 column volumes of PreScission buffer (50 mM Tris–HCl, pH 7.5, 100 mM NaCl, 1 mM EDTA, 1 mM DTT) was used to equilibrate the column prior to protease addition. 2 units/ul of PreScission protease was manually added to the column (closed flow) and incubated for 2 h. The cleavage product was eluted in elution buffer (1 mM DTT, 100 mM NaCl, 20 mM MES, pH 6, 5 mM EDTA) and this material was loaded onto a 1 ml mono S column (GE), washed with elution buffer containing 10% glycerol and then eluted with elution buffer/10% glycerol/0.1 to 1 M NaCl over a 50 ml volume. The eluted protein fractions were analyzed by SDS-PAGE and then pooled and concentrated in a centrifugal filter device (Millipore) in Pol4 buffer (20 mM Tris–HCl, pH 7.5, 350 mM NaCl, 1 mM DTT, 10% Glycerol) and flash frozen in liquid N2. Protein concentrations were determined using the BioRad protein assay kit, using BSA as a standard protein.
A specific antibody to yeast Pol4 was produced against a C-terminal Pol4 fragment (aa 225–583) using olWDH1015 and olWDH1016 as primers to amplify a 1058 bp fragment. This fragment was digested with NcoI and BamHI before insertion into a pET28-b expression vector (pWDH1151) containing a C-terminal 6xHIS-tag, and subsequent transformation into E. coli (WDH992, DH5α). Expression of the truncated Pol4 fragment in pET28-b was achieved through addition of IPTG to WDH992 cultures followed by purification using a Ni2+ affinity column. Following purification, the C-terminal Pol4 fragment was injected in rabbits and guinea pigs to generate polyclonal antibodies against Pol4. Anti-human Polλ antibody was obtained from Novus Biological (NB100-81664).
DNA substrates
The 35mer oligonucleotides (olWDH2122, 2124) were purified by 10% denaturing acrylamide gel electrophoresis. The 80mer oligonucleotide (olWDH2123) contained biotin on the 5’ and 3’ ends, while olWDH2134 contained biotin only on the 5’ end. All oligonucleotides were obtained from Integrated DNA Technologies. Supplementary Table S3 provides the sequence for all oligonucleotides.
DNA synthesis assays and Rad51 titration
Unless otherwise stated in the figure legends, DNA synthesis reactions were carried out in presence of 25 nM of substrate molecules (35mer pre-annealed with 25 nM of 80mer), with 12.5 nM Polδ and/or the indicated concentrations of Pol4 and Rad51 in Pol4 reaction buffer (25 mM Tris–acetate (pH 7.4), 0.1 mg/ml BSA, 2 mM DTT, 7 mM MgCl2) supplied with 2 mM ATP, 0.6 μM NeutrAvidin (Thermo Scientific) and 500 μM of dNTPs. For the Rad51 titrations, each DNA substrate was first incubated with Rad51 for 10 min at 30°C and DNA synthesis was started by the addition of Polδ and further incubation for 10 min at 30oC. For Pol4 synthesis, DNA was incubated in the presence or absence of Rad51 (1:1 Rad51:nt) for 10 min at 30oC, followed by the addition of 25nM of Pol4 or Pol4-CD and incubation for 10 min at 30oC. For the Pol4 titration reactions, DNA was incubated at 30°C with proteins as follow: Rad51 (1:1 Rad51:nt) for 10 min, Pol4 for 2 min and Polδ for 8 min. In the absence of one or more proteins, the same volume was replaced with the respective dilution buffer. Reactions were terminated by the addition of 1 mg/ml Proteinase K, 1% SDS and incubation for 1 h at 37oC or overnight at room temperature and then analyzed by 10% denaturing acrylamide gel electrophoresis. Gels were dried, imaged with a Molecular Dynamics PhosphoImager and quantified using ImageQuant software.
Annealing-dependent DNA synthesis assay
Unless otherwise stated in the figure legends, reactions were carried out in the presence of 25 nM of the 35mer (olWDH2122) as the primer to be extended and 25 nM of the 80mer (olWDH2123) as template, with 25 nM RFC, 25 nM PCNA, 12.5 nM Polδ and/or indicated concentrations of Pol4 in a Pol4 reaction buffer supplied with 2 mM ATP, 0.6 μM NeutrAvidin (Thermo Scientific) and 500 μM of dNTP. When present, RPA, Rad51 and Rad52 had a stoichiometry of 1:20 nt, 1:3 nt and 1:10 nt, respectively. Reactions were carried at 30°C as follows: first, the 35mer primer was incubated with Rad51 for 5 min followed by addition of RPA for 5 min. Next, Rad52 and the 80mer was added to the reaction and incubated for another 5 min. After that, DNA synthesis was started by the addition of Pol4 for 2 min followed by the addition of pre-mixed PCNA/RFC for 2 min and Polδ for 8 min. PCNA/RFC concentrations varied within the assays in order to preserve 1:1 stoichiometry in relation to the DNA substrate (as previously published, (78)). In the absence of one or more proteins, the same volume was replaced with the respective dilution buffer. Reactions with Rad51, RPA and Rad52 were incubated with the 35mer before addition of 80mer. Reactions were terminated by the addition of 1 mg/ml Proteinase K, 1% SDS and incubation for 1 h or overnight at 37°C and then analyzed by 10% denaturing acrylamide gel electrophoresis. Gels were dried, imaged with a Molecular Dynamics PhosphoImager and quantified using ImageQuant software.
DNA synthesis quantifications
DNA synthesis assays were quantified using ImageQuant software. For primer utilization quantifications, only the intensity of the unextended primer band was analyzed, and results were normalized to the control reaction (no protein added). For full length quantifications the intensity of bands corresponding to the addition of the 41st nt and more were quantified, and the results were normalized to reactions with Polδ only. For total DNA synthesis, the intensity of bands corresponding to the addition of the 1st nt and more were quantified, and the results were normalized to the respective control reaction.
Rad51 displacement assay
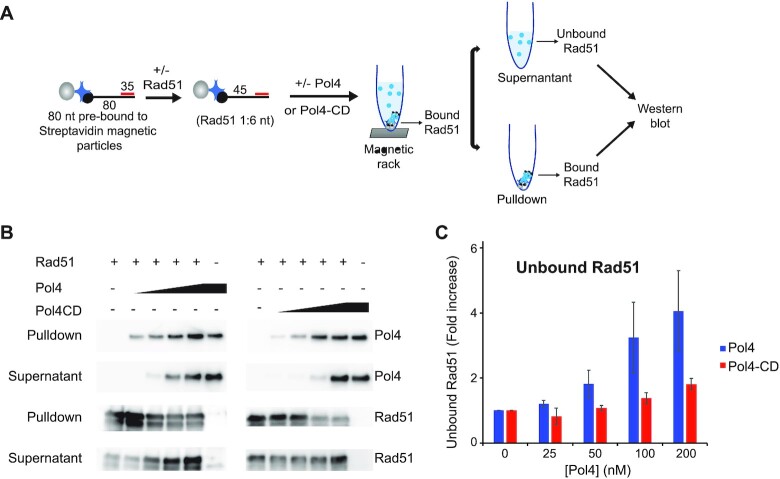
For the Rad51 displacement assay, the 80mer (olWDH2134) was pre-annealed with 35mer (olWDH2122) before immobilization onto magnetic streptavidin beads (Roche, #164177) following the manufactory protocol. For the experiments with Pol4 and Pol4-CD (Figure 6), 50 nM of tailed DNA (olWDH213/olWDH2122) was incubated with 375 nM Rad51 (1:6 nt of the ssDNA part) for 10 min at 30°C in Pol4 reaction buffer supplemented with 2 mM ATP, 500 μM dNTPs and an ATP-regenerating system containing 20 mM creatine phosphate and 0.1 mg/ml creatine kinase. Increasing amounts of Pol4 or Pol4-CD were added and incubated for 10 min before separating into pulldown and supernatant fractions. For Supplementary Figure S9, the 80mer (olWDH2123) was bound to magnetic streptavidin beads (Roche, #164177) following the manufacturer's protocol. Reactions were carried as follows: 100 nM of 80mer was incubated with Rad51 (1:3 nt) for 5 min at 30°C in Pol4 reaction buffer supplied with 2 mM ATP and with or without 500 μM of dNTPs. After Rad51 incubation, 100 nM of the 35mer (olWDH2122) and indicated concentrations of Pol4 were added for 10 min. Reactions were terminated by placing the reaction tubes in a magnetic rack and transferring the supernatant for new tubes containing Laemmli buffer. Reactions were boiled for 5 min and analyzed by immunoblotting using our Rad51 and Pol4 anti-sera.
Figure 6.
Pol4 displaces Rad51 from DNA. (A) Rad51 displacement assay. Black dots indicate biotin labels at the 3’ end of the 80mer. (B) Immunoblots of bound and unbound Rad51 and Pol4 or Pol4-CD. Reactions analyzed were carried out with 50 nM of the annealed tailed DNA substate, 375 nM of Rad51 (1:6 nt of the ssDNA part of the tailed DNA), and increasing amount of Pol4 or Pol4-CD (0, 25, 50, 100 and 200 nM). (C) Quantitation of results in (B) and additional experiments. Data show mean ± SEM of n = 3.
Statistical analysis
The 95% or better confidence intervals for the median were calculated using the information provided in Supplementary Table S4. This utilizes the formula: (n + 1)/2 ± 1.96 (√n/2) to provide the upper and lower limit of the 95% or better confidence interval from the rank order of values. As such, this method will generate an uneven ± confidence interval around the median. The p-values were always generated between two data sets using pairwise unpaired t-test with Welch's correction that does not assume equal standard deviations between data sets.
RESULTS
Pol4 acts in homology-directed direct repeat recombination
Pol4 plays a well-documented role in DSB repair via classic NHEJ and MHMR (47,55–60). Whether Pol4 is involved in HDR is an unsettled question (see Introduction). We identified roles of Pol4 in ionizing radiation (IR)-resistance and spontaneous DRR which suggest that Pol4 plays a specific role in a subset of HDR events.
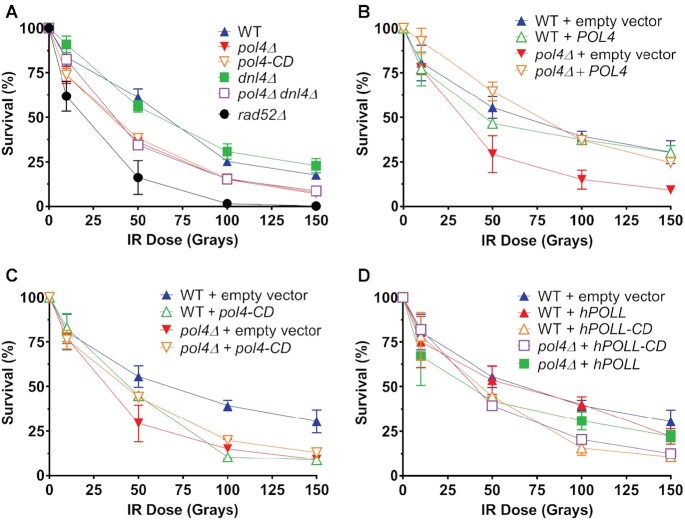
Examining the sensitivity of S. cerevisiae cells lacking Pol4, we found that exponentially growing pol4Δ cells are mildly but significantly and reproducibly hypersensitive to IR compared to wild type (WT) in W303, a strain background commonly used in studies of the DNA damage response (Figure 1). Previously, it was shown that Pol4 DNA synthesis is required for telomere fusions in Rap1-deficient cells and for MHMR but not for NHEJ (56,57). The function in IR resistance is also dependent on the polymerase activity of Pol4, since the catalytic deficient pol4-D367E (referred to as pol4-CD (60)) mutant shows IR sensitivity indistinguishable from the pol4Δ strain (Figure 1A–C). The Pol4-CD protein was shown to have no protein stability defect and accumulated to the same steady state level as wild type Pol4 (60). The pol4-CD mutant exhibited a dominant-negative phenotype when overexpressed in WT cells resulting in IR sensitivity equivalent to that of the pol4Δ mutant (Figure 1C). The deletion of the core NHEJ factor DNA ligase (DNL4) did not affect IR sensitivity, and the dnl4Δ mutation did not result in additional IR-sensitivity of pol4Δ cells under the conditions used (Figure 1A). These results support the conclusion that NHEJ does not contribute to IR-resistance in exponentially growing yeast cells and are consistent with earlier observations showing that yeast dnl4 mutants show mild IR sensitivity only when kept in stationary phase but not as exponential phase cells (79–81). Furthermore, the presence of WT human Polλ (hPOLL) but not its catalytic mutant, hPOLL-D427E (referred to as hPOLL-CD), could partially restore the IR sensitivity of yeast pol4-Δ mutants at the higher doses of 100 and 150 Gy (Figure 1D). Similar to the yeast pol4-CD mutant, expression of the human hPOLL-CD mutant resulted in a negative phenotype (Figure 1D). Both results suggest partial functional conservation between S. cerevisiae and human Polλ. We conclude that Pol4 contributes to IR resistance distinct from its role in NHEJ, consistent with a possible role in HDR.
Figure 1.
Loss of Pol4 activity confers ionizing radiation (IR) sensitivity. (A–D) Cultures of haploid WT and mutant cells were grown to mid-log, counted by hemocytometer, and spread onto YPD plates before exposure to 0, 10, 50, 100 or 150 Grays of ionizing radiation. Percent survival was determined for each genotype by counting the number of colonies, dividing by the number of cells plated, and multiplying by 100. The mean percent survival ± SEM was determined for each genotype from a minimum of 9 independent cultures, normalized to cells not exposed to IR, and plotted against radiation exposure in Grays. (A) Percent survival of pol4Δ (haploid segregant of WDHY2816), pol4-CD (haploid segregant of WDHY2895), dnl4Δ (haploid segregant of WDHY2829), and pol4Δ dnl4Δ (haploid segregant of WDHY3150) mutants compared to WT (haploid segregant of WDHY2990) and rad52 (WDHY2556) following varying levels of IR exposure. (B–D) IR survival curves for WT (haploid segregant of WDHY2990) and pol4Δ (haploid segregant of WDHY2816) mutant cells possessing an extrachromosomal vector with a functional yeast POL4 (pWDH1145) or human hPOLL (pWDH1146) DNA polymerase λ gene, a DNA polymerase defective allele from yeast, pol4-CD (pWDH1147) or human hPOLL-CD (pWDH1148), or without an insert, empty vector (pWDH403). The absence of error bars indicates that they are smaller than the plotting symbol.
To identify a possible role of Pol4 in HDR, we employed a series of assays that revealed no effect of pol4Δ on mitotic or meiotic interhomolog recombination (Supplementary Figures S1 and S2) nor on unequal sister-chromatid recombination (Supplementary Figure S3). Moreover, we identified no repair defect of the pol4Δ mutation in two SSA assays (Supplementary Figures S4 and S5). The assay system in Supplementary Figure S4 reports on interchromosomal SSA between two 311 bp repeats after DSB induction (16). The DRR assay system in Supplementary Figure S5 monitors spontaneous events between two 2241 bp repeats of ade2 genes: gene conversion as ADE+ URA+ colonies whereas ADE+ URA− colonies are indicative of SSA, unequal sister chromatid exchange, or intrachromatid recombination (72). We conclude that Pol4 is not involved in mitotic or meiotic allelic recombination nor in DSB-induced SSA.
The assay in Supplementary Figure S5 showed a small but significant effect of the pol4-CD mutation for ADE+ URA− events, which can result from SSA or other types of DRR. The effect of the pol4-CD mutant may suggest that the mutant protein obstructs access by alternate DNA polymerases that may act in partial redundancy with Pol4. Hence, we investigated the role of Pol4 in a separate DRR system which employs two truncated versions of the HIS3 gene oriented as a tandem direct-repeat with either 103 or 415 bp of homology (Figure 2A) (73). In this system no DSB is induced, and the HIS3 gene is restored spontaneously in a homology-dependent fashion (73). Our results show that pol4Δ mutants result in a 2.9-fold or 27-fold decrease of HIS3+ recombinants compared with WT cells when 103 bp or 415 bp of homology are used, respectively (Figure 2B, C). The rad51Δ mutants showed a statistically significant increase of about 3-fold in the frequencies of HIS3+ recombinants compared to WT for both substrates. This is consistent with previous results showing that Rad51 inhibits translocation events resulting from Rad52-mediated annealing to the same extent (16). Surprisingly, the deletion of RAD51 in a pol4Δ strain resulted in a statistically significant rescue of the phenotype of pol4Δ regardless of the length of the shared homology, suggesting Pol4 is only required in the presence of Rad51 (Figure 2B, C). For the 415 bp repeat, the suppression by rad51Δ in the pol4Δ and pol4-CD mutants reached the rate of the rad51Δ single mutant (comparison to rad51Δ: P = 0.79 and P = 0.67, respectively). For the 103 bp repeat, the suppression appeared incomplete, although only the difference between the rad51Δ and the rad51Δ pol4Δ was significant (P = 0.0194) while to the pol4-CD mutant was not (P = 0.88).
Figure 2.
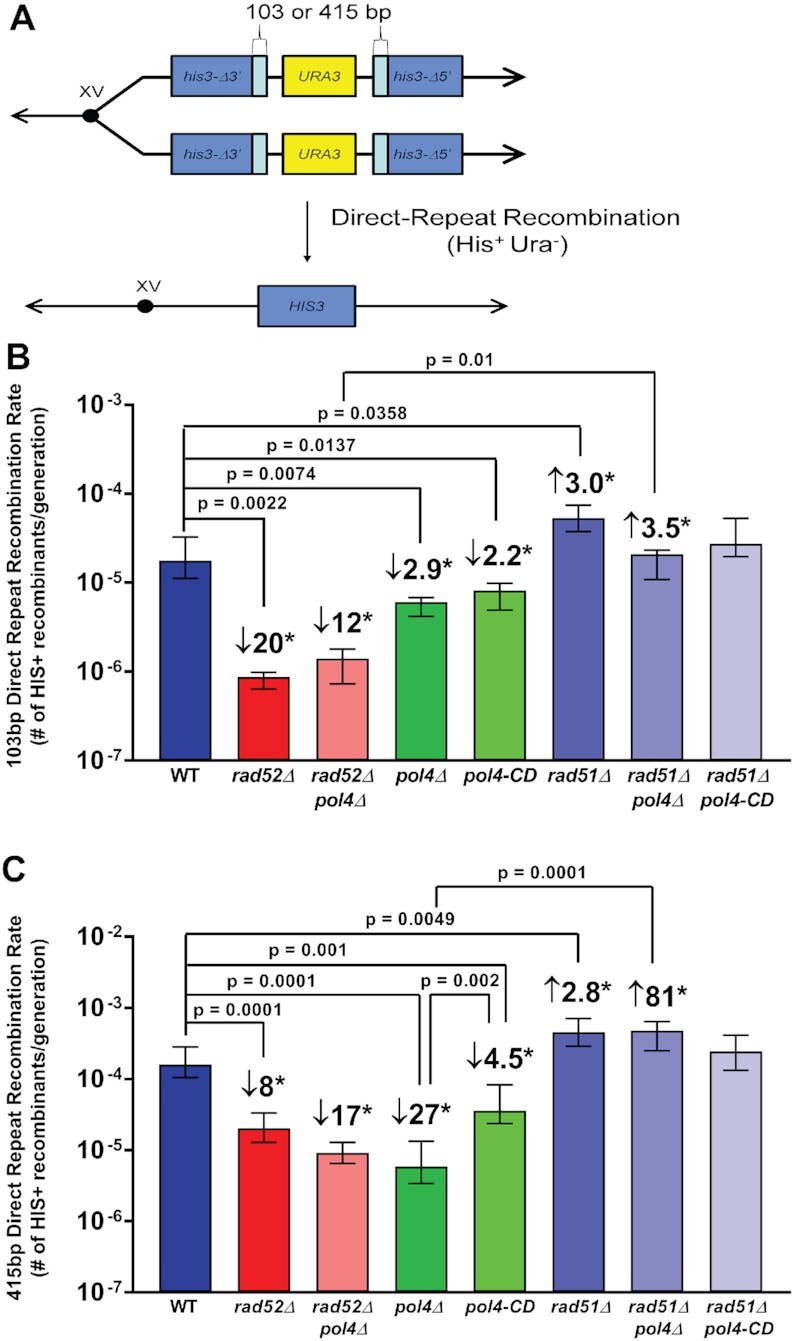
Direct-repeat recombination rate is influenced by Pol4 and Rad51. (A) Spontaneous direct-repeat recombination is measured as the rate of His+ recombinant formation in haploid cells possessing two truncated his3 alleles sharing 103 bp or 415 bp of homology. (B) Haploid wildtype (WT) cells (WDHY5901), rad52Δ (WDHY5905), rad52Δ pol4Δ (WDHY6173), pol4Δ (WDHY5909), pol4-CD (WDHY5843), rad51Δ (segregant of WDHY5227), rad51Δ pol4Δ (segregant of WDHY5226), and rad51Δ pol4-CD (segregant of WDHY5226) mutants show varying direct-repeat recombination rates using 103 bp of homology. (C) Haploid wildtype (WT) cells (WDHY5903), rad52Δ (WDHY5907), rad52Δ pol4Δ (WDHY6174), pol4Δ (WDHY5911), pol4-CD (WDHY5845), rad51Δ (segregant of WDHY5224), pol4Δ rad51Δ (segregant of WDHY5225), and pol4-CD rad51Δ (segregant of WDHY5225) mutants show varying direct-repeat recombination rates using 415 bp of homology. The median rate and ± 95% confidence interval of direct-repeat recombination was calculated from a minimum of nine independent cultures. * indicates significant differences as determined using pair-wise unpaired t-tests with P ≤ 0.05. The fold increases/decreases are given for the comparison indicated by the P-value with the exception of the comparison for pol4Δ and pol4-CD in C, for which no fold change is given.
The substitution of Pol4 by its catalytic deficient version Pol4-CD elicited a milder decrease in HIS3+ recombination frequency compared to the pol4Δ mutant, resulting in 2.2-fold or 4.5-fold decreases compared to WT with 103 bp or 415 bp of homology, respectively. Although only the difference between pol4Δ and pol4-CD with the 415 bp substrate was statistically significant (Figure 2B, C). This suggests that this DRR event is only partially dependent on Pol4 DNA synthesis. As expected, rad52 mutants significantly reduced the frequencies of HIS3+ recombinants compared to WT cells, demonstrating that the mechanism of HIS3+ recombination is independent of Rad51 but dependent of Rad52 consistent with an annealing-driven process (Figure 2B, C). The effect of the pol4Δ with the 415 bp repeats appeared stronger than that of rad52Δ, but the difference was not statistically significant. The defect in rad52Δ cells was significantly more pronounced than in pol4Δ mutants for the 103 bp repeat (P = 0.0001). The relatively small decrease in the rad52 mutant, especially with the 415 bp repeat (8-fold with a remaining rate of about 1 × 10−5), may suggest the existence of alternative annealing proteins that might become relevant in this particular context. Recent evidence showed that also other proteins than Rad52 can anneal RPA-coated ssDNA, such as human SMARCAL and ZRANB3 (82).
The results from these genetic assays suggest that Pol4 is involved in a distinct subset of HDR events that involve repeated DNA utilizing a mechanism that requires Rad52 but is independent of Rad51; in fact, the data show that such events are inhibited by Rad51. The Rad52 requirement implies a mechanism that involves annealing of ssDNA. However, this is unlikely to be classic DSB-induced SSA, as pol4Δ showed no defect in the interchromosomal SSA assay. This suggests that Pol4 is involved in DRR that is not associated with DSBs but possibly with stalled replication forks. It is unclear why pol4Δ does not show a defect in the DRR assay involving the repeated ade2 genes (Figure S5) but we note that these repeats are significantly longer (2241 bp) than those in the DRR assay of Figure 2 (103 or 415 bp). It is possible that homology length affects the mechanisms involved. Alternatively, the different sequence or chromosomal location may impact initiation or outcome of DRR events. Moreover, we note that the HIS3 repeat system of Figure 2 involves truncations with no requirement for mismatch repair to generate HIS + recombinant, whereas the ADE2 system in Supplementary Figure S5 involves point mutations requiring mismatch repair for ADE + formation. While with the 103 bp repeats the pol4Δ and pol4-CD mutants showed equivalent defects (Figure 2B), the pol4Δ mutant exhibited a stronger defect with the 415 bp repeats than the pol4-CD mutant which was statistically significant (Figure 2C). This suggests that some DRR events, at least with the 415 bp repeats, required additional Pol4 functions besides its catalytic DNA polymerase activity. Homology length may also explain the differences between the 103 and 415 bp repeats in the DRR assay of Figure 2.
Pol4 and Polδ cooperate to overcome Rad51 inhibition of DNA synthesis
To understand the role of Pol4 in DRR and its contribution to DNA synthesis during HDR, we purified S. cerevisiae WT Pol4, a catalytic deficient polymerase (Pol4-D367E = Pol4-CD), and the homologous human protein, hPolλ, to biochemically reconstitute DNA synthesis reactions (Supplementary Figure S6A). We used a primer-template substrate composed of an 80-mer template strand and a homologous 35-mer primer. The 80mer contained biotin at its 5’- and 3’ ends (indicated by the black dots in Supplementary Figure S6A, Figure 3A and subsequent DNA substrate illustrations) to be able to constrain the PCNA sliding clamp after loading (see experiments including PCNA/RFC in Figures 5, 7, S7, Supplementary Figure S8) (78). To achieve consistency between all DNA synthesis experiments, we used this same substrate throughout but varied the 35-mer primer. Consistent with previous observations (83,84), the presence of the biotin at the ends did not affect the DNA synthesis reactions. As previously observed (52), Pol4 exhibits limited DNA synthesis on primer-template substrates, adding primarily 1 nt but rarely more than 4–5 nt under optimal DNA extension reaction conditions. In the presence of super-saturating amounts of Rad51 to bind not only the ssDNA portion of the substrate but also the dsDNA portion, Pol4 is still able to add a single nt to the primer, albeit with diminished efficiency, and extension is strictly limited to + 1 nt (Supplementary Figure S6B, C). This suggests that even with excess Rad51 Pol4 has access to the primer template, can extend the primer by one nt but further extension is precluded by Rad51 binding. As expected, the Pol4-CD enzyme is completely defective in DNA synthesis (Supplementary Figure S6C). Thus, unless the DNA synthesis required is less than 4–5 nt, Pol4 is likely cooperating with other DNA polymerases in HDR.
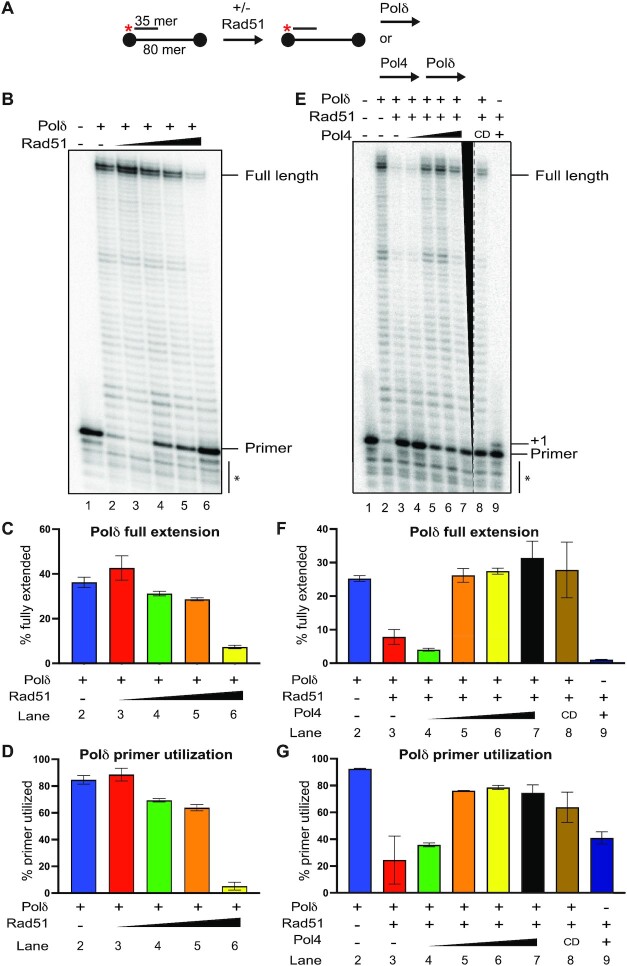
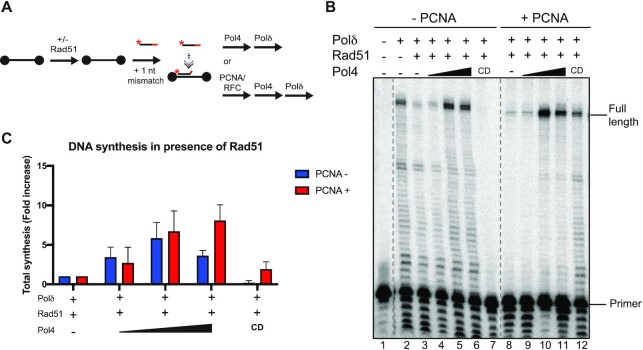
Figure 3.
Pol4 and Polδ cooperate to overcome Rad51 inhibition of DNA synthesis. (A) DNA synthesis assay using substrates olWDH2123 (80mer) and olWDH2124 (35mer). Black dots indicate biotin labels at the 5’ and 3’ ends of the 80mer. (B) Denaturing 10% acrylamide gel of DNA synthesis reactions. Reactions were carried out with 25 nM molecules substrate incubated without or with Rad51 (1:12 nt, 1:6 nt, 1:3 nt, 1:1 nt of the 80mer) and 12.5 nM of Polδ. (C and D) Quantitation of results in (B) and additional experiments. Data shows mean ± SEM of three independent experiments. (E) Denaturing 10% acrylamide gel of DNA synthesis reactions. Reactions were carried with 25 nM molecules substrate incubated without or with Rad51 (1:1 nt) or Pol4 (12.5, 25, 50,100 nM) or Pol4-CD (100 nM) or Polδ (12.5 nM). The dotted line and black triangle indicate that lanes 8 and 9 were taken from a different gel. (F and G) Quantitation of results in (E) and additional experiments. Data show mean ± SEM of n = 3. * denotes bands below the primer that derive from shorter oligonucleotide synthesis products and products of Polδ exonuclease activity.
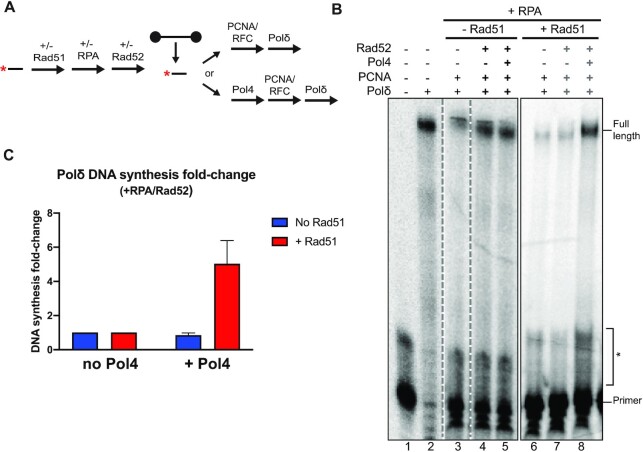
Figure 5.
Pol4 suppresses Rad51 inhibition of annealing-dependent DNA synthesis in the presence of Rad51 and PCNA/RFC. (A) Annealing-dependent DNA synthesis assay. (B) Denaturing 10% acrylamide gel of DNA synthesis reactions. The broken line indicates that the control lanes (1,2) were from the same gel with intervening lanes omitted. Lanes 6–8 were from a different gel. Reactions were carried in the presence of 50 nM of the 35mer substrate and 50nM of the 80mer. Proteins were added as follows: ± Rad51 (1:9 nt), ± RPA (1:60 nt), Rad52 (1:40 nt), ± Pol4 (25 nM), ± PCNA/RFC (50 nM each) and ± 25 nM of Pol δ and (C) Quantitation comparing the Pol4 effect on DNA synthesis in the presence of RPA and Rad52 in reactions without Rad51 (B, lanes 4 and 5) and with Rad51 (B, lanes 7 and 8) and additional replica experiments. The data reflect the increase (fold-change) in Polδ synthesis in the presence of Pol4 normalized by the respective reaction without Pol4. Data show mean ± SEM of n = 3. * marks bands that arise from secondary structures of the primer; see lane 1 in (B).
Figure 7.
Pol4 catalytic activity is required in the presence of a + 1 nt flap at the 3’-OH end. (A) Annealing-dependent DNA synthesis assay. Black dots indicate biotin labels at the 5’ and 3’ ends of the 80mer. (B) Denaturing 10% acrylamide gel of DNA synthesis reactions. Reactions were carried out in the presence of 25 nM of the 34 + 1mer substrate and 25 nM of the 80mer. Proteins were added as follows: ± Rad51 (1:3 nt), ± PCNA/RFC (25 nM each), ± Pol4 (12.5, 25, 50 nM) or Pol4-CD (50 nM) and ± 12.5 nM of Polδ. Broken lines indicate that some lanes from the gels were spliced out. (C) Quantitation of DNA total synthesis (see Material and Methods) from (B) and additional experiments. Data show mean ± SEM of 3 independent experiments.
We previously reported that Pol4 and Polδ cooperate in DSB repair by MHMR (47). Rad51 was shown to inhibit PCNA loading at D-loops and Polδ-mediated DNA synthesis in D-loops (85). We tested the effect of Rad51 in the DNA synthesis reaction facilitated by Pol4 and Polδ, individually and in combination (Figure 3A). We observed that Rad51 inhibited both primer utilization and extension by Polδ (Figure 3B–D), especially when present at a stoichiometry of 1 Rad51 protomer per nt, an amount that leads to binding of Rad51 to the entire substrate molecule including the dsDNA (86). Unlike RecA, Rad51 readily binds duplex DNA which has functions in replication fork protection (87). The small stimulation with the lowest concentration of Rad51 was not statistically significant. Interestingly, Pol4 incorporated 1 nt in the presence of the highest concentration of 1 Rad51 per nt in almost half of the available primers (Figure 3E, G), consistent with the result from Supplementary Figure S6C. Importantly, the addition of Pol4 prior to Polδ in the DNA synthesis reaction restored primer utilization and DNA extension carried out by Polδ (Figure 3E–G). This requires a stoichiometric amount of Pol4, as 12.5 nM Pol4 with 25 nM substrate had little effect but 25 nM Pol4 led to complete restoration of Polδ DNA synthesis indistinguishable from adding up to 100 nM (Figure 3E–G, lanes 3–7). The role of Pol4 in restoring Polδ activity in the presence of Rad51 appears to be partially independent of its polymerase activity since the catalytic deficient version of Pol4 (Pol4-CD) also restored significant Polδ DNA synthesis, although we noted some variability between experiments as indicated by the large error bars (Figure 3E–G, lane 8; see also Figure 4B, Supplementary Figure S7C, D). Surprisingly, Rad51 does not equally inhibit Polϵ under the same conditions, suggesting a specific functional interaction between Pol4, Polδ, and Rad51 (Supplementary Figure S6D–G). We conclude that Pol4 overcomes the inhibition of Polδ by Rad51 and that Pol4 DNA synthesis is not essential for this effect (Figure 3). This biochemical finding is consistent with the genetic data (Figure 2C) showing that Pol4 activities beyond DNA synthesis are required for full function in DRR.
Figure 4.
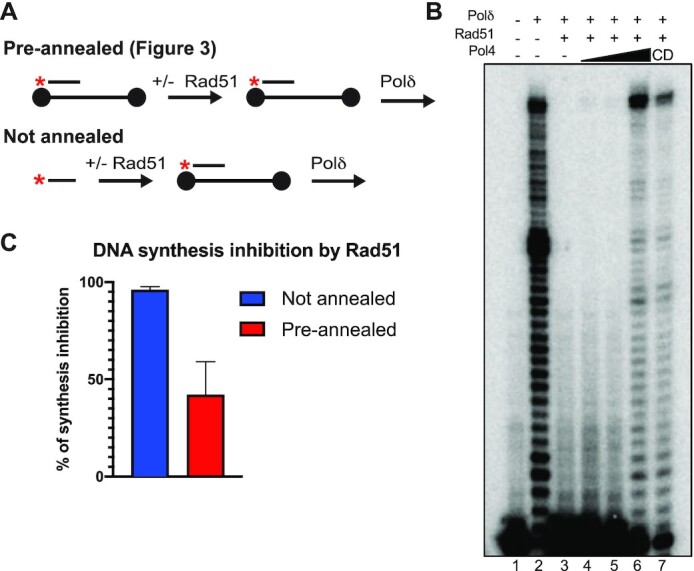
Rad51 inhibition of annealing-dependent DNA synthesis. (A) Reaction schemes for DNA synthesis with pre-annealed (see Figure 3) and not pre-annealed primer. Black dots indicate biotin labels at the 5’ and 3’ ends of the 80mer. (B) Reactions were carried out in the presence of 25 nM of the 35mer substrate and 25 nM of the 80mer. Black dots indicate biotin labels at the 5’ and 3’ ends of the 80mer. Proteins were added as follows: ± Rad51 (1:3 nt), ± Pol4 (12.5, 25, 50 nM) or Pol4-CD (50 nM) and ± 12.5 nM of Polδ. (C) Quantitation of DNA synthesis inhibition by Rad51 in reactions with pre-annealed primers (Figure 3B, lanes 2, 5), without pre-annealed primers (B, lanes 2, 3) and additional replica experiments. Reactions contained Rad51 at 1:3 nt. Data shows mean ± SEM of n = 3.
Rad51 abrogates annealing-dependent DNA synthesis by Polδ
To investigate the importance of DNA strand-annealing in the Pol4 and Polδ synthesis process described above, we designed an annealing-dependent DNA synthesis assay. In this assay, the reaction is staged with the complementary DNA strands being introduced consecutively during the reaction. The final product of the reaction (extended DNA) can only be formed if an intermediate step of strand-annealing occurs (Supplementary Figure S7A). Due to its known role as a processivity factor of DNA polymerases that orchestrates polymerase exchanges and to better emulate the DNA synthesis conditions for Polδ, we included the processivity clamp PCNA and its loader RFC to the reactions. While Polδ DNA synthesis is stimulated by PCNA/RFC, as expected (88), we did not observe a stimulation of Pol4-mediated DNA synthesis by PCNA/RFC or an increase in the number of nucleotides incorporated (Supplementary Figure S7B–D). Supporting the hypothesis that Pol4 is involved in DNA strand-annealing, we show that Pol4 and PCNA have additive effects in stimulating Polδ synthesis when DNA annealing is needed (Supplementary Figure S7C-D). In the absence of PCNA, the Pol4 wild type and Pol4-CD proteins did not stimulate Polδ DNA synthesis, whereas in the presence of PCNA, Pol4-CD stimulated slightly but significantly more than the wild type Pol4 protein, as indicated by the non-overlapping error bars (Supplementary Figure S7D). This may suggest a Pol4–PCNA interaction that does not affect Pol4 processivity but other functions. The human homolog of Pol4, Polλ has been shown to interact with PCNA and this interaction suppresses Polλ DNA synthesis but stimulates bypass of AP sites likely by increasing the affinity to primer template (89,90).
To further investigate the importance of DNA annealing, we used the values obtained for DNA synthesis from our previous analysis to generate a direct comparison of Polδ DNA synthesis when the substrate was pre-annealed (Figure 3) or not annealed (Figure 4A). The analysis reveals that under the same conditions as in the DNA synthesis reactions with pre-annealed primer-template substrates, Rad51 inhibition of Polδ was more pronounced when DNA annealing was required prior to DNA synthesis (Figure 4B, C), consistent with the ability of Rad51 to inhibit DNA annealing (91) and Polδ DNA synthesis (Figure 3). Also in this reaction scheme requiring strand annealing, addition of Pol4 overcame the inhibition and the catalytic deficient Pol4-CD protein exhibited partial but significant suppression (Figure 4B, lanes 6 and 7).
Rad51 inhibits Rad52-mediated annealing of RPA-coated complementary ssDNA suggesting that Rad51 could play a role in DNA repair pathway choice by inhibiting the SSA pathway thereby favoring HR (3,91). We tested whether Pol4 could also suppress Rad51 inhibition of Rad52-mediated strand-annealing in the annealing-dependent DNA synthesis assay. First, we investigate whether the proteins would behave as previously described in our annealing-dependent DNA synthesis assay (Figure 4A, Supplementary Figure S8A). Using saturating levels of proteins, we showed that while Pol4 is capable of alleviating Rad51 inhibition of Polδ synthesis (Figure 4B same as Supplementary Figure S8B), the addition of RPA to our assay resulted in DNA synthesis blockage (Supplementary Figure S8C). This result is consistent with previous studies that showed that RPA strongly inhibits DNA annealing (14,91,92). However, the addition of Rad52, known to promote the annealing of RPA-coated ssDNA (14), restored DNA synthesis by Polδ fully (Supplementary Figure S8D). In both experiments a difference in DNA synthesis was observed in the reactions with or without Pol4 (Supplementary Figure S8C, D). Pol4 becomes essential to restore Polδ activity in reactions containing Rad51, RPA and Rad52 (Supplementary Figure S8E). The ability of Pol4 to promote Polδ-mediated DNA synthesis even in the presence of Rad51, RPA and Rad52 proteins was particularly interesting due the role of these proteins in controlling DNA annealing and replication fork support and DNA repair pathways choices (91,92).
During DNA metabolic processes in cells, Rad51, RPA and Rad52 coexist and dynamically bind single-strand DNA. To emulate these conditions and ensure that more than one protein could bind the substrate, we also performed our assays using sub-saturating concentrations keeping the protein to DNA stoichiometry as previously shown to generate mixed protein-DNA filament formation (Figure 5A) (92). Under these conditions, the presence of RPA had a mild effect inhibiting DNA synthesis by Polδ (Figure 5B, lane 3) and the addition of Rad52 was able to increase Polδ synthesis (Figure 5B, lane 4). Furthermore, the addition of Pol4 did not alter Polδ synthesis in the presence of RPA and Rad52 (Figure 5B, lane 5). The addition of Rad51 strikingly reduced the ability of Polδ to carry out DNA synthesis in the presence of RPA (Figure 5B, lane 6), which was not improved by the addition of Rad52 (Figure 5B, lane 7). This suggests that Rad51 has a similar inhibitory effect in our annealing-dependent DNA synthesis assay as in the previously studied annealing assays (91). Remarkably, addition of Pol4 overcame the Rad51 inhibition of annealing and DNA synthesis almost completely (Figures 5B, lane 8, 3C).
We conclude that Pol4 overcomes the Rad51 inhibition of Polδ in DNA synthesis following DNA strand annealing by Rad52 in the presence or absence of PCNA/RFC and note that the Pol4 effect was not completely dependent on its polymerase activity.
Pol4 can displace Rad51 from DNA
Our results suggest that Pol4 has a non-catalytic role in displacing Rad51 from DNA. We hypothesize that Pol4 displaces Rad51 at the primer template junction just enough to grant Polδ access to the primer-template and developed a Rad51 displacement assay. In this assay, Rad51 bound to primer-template DNA was collected through a pulldown, and the amounts of Rad51 and Pol4 bound to DNA and in the supernatant were measured by immunoblotting (Figure 6A). Both Rad51 and Pol4 bind to the DNA (Figure 6B), precluding an approach using gel shift assays. The results show that Rad51 distributes from the DNA to the supernatant in response to the Pol4 titration (Figure 6B, C). The amount of Rad51 detected in the supernatant increased upon titrating Pol4, showing the ability of wild type Pol4 to displace Rad51 from DNA. The order of addition, Rad51 before Pol4, strongly suggests that Pol4 can displace Rad51 from DNA and not just compete with Rad51 DNA binding. The genetic data suggested that also the Pol4-CD mutant protein retains the ability to displace Rad51. Hence, we performed the same Rad51 displacement assay with Pol4-CD. The results show a diminished ability of Pol4-CD to displace Rad51 from the DNA substrate (Figure 6B, C). These results were confirmed analyzing Rad51 displacement from DNA in the presence and absence of the 4 dNTPs by the wild type Pol4 protein, showing reduced Rad51 displacement in the absence of dNTPs (Supplementary Figure S9). This suggests that dNTP binding induces a conformational change that reinforces DNA binding and subsequent Rad51 displacement and that Pol4-mediated DNA synthesis leads to further Rad51 displacement due to the ability of Pol4 to incorporate 1–5 nts displacing 1–2 extra Rad51 protomers when DNA synthesis is possible (Figure 6, Supplementary Figure S9). We conclude that Pol4 displaces Rad51 from DNA partly independent of its DNA polymerase activity. Since our IR results showed that hPolλ can partially complement the IR sensitivity of the pol4-Δ mutant in yeast at the high IR doses (Figure 1D), we tested whether hPolλ could also displace yeast Rad51. Our results revealed that also hPolλ is able to displace yeast Rad51 (Supplementary Figure S9D, E). This opens the possibility that hPolλ may play a similar role to Pol4 in human cells, although this conjecture requires future elaboration.
Pol4 synthesis is required in presence of +1nt mismatch at 3’-OH
Since our in vivo results revealed that the Pol4 catalytic activity is required in some cases, we tried to understand in which context Pol4 polymerase activity was essential for DNA repair. Pol4 has been shown to be important in extending primers containing mismatches, especially when located as a flap at the 3’-OH end (49,59,60,93). We tested if the presence of small 3’-OH flaps in the primer affected the need for Pol4 polymerase activity in extension reactions (Figure 7A). Our results revealed that Pol4 DNA synthesis becomes essential in the presence of a 1 nt 3’-flap in the absence of PCNA (Figure 7B). We also evaluated the effect of PCNA in this reaction. As observed before in our annealing-dependent DNA synthesis assays (Figure 4), the addition of PCNA appears to facilitate Polδ DNA synthesis at the higher concentrations of Pol4 (Figure 7B, C). Moreover, Polδ DNA synthesis became partially independent of the Pol4 DNA synthesis (Figure 7B, lane 12). This result suggests that without PCNA Pol4 competes with Polδ for the substrate, while the presence of PCNA could aid Polδ DNA synthesis by promoting polymerase exchange after Rad51 is displaced by Pol4 even in the absence of Pol4-mediated DNA synthesis.
DISCUSSION
S. cerevisiae Pol4, like its human counterpart Polλ, plays a specialized role in DNA repair during NHEJ and MHMR to fill gaps in a distributive fashion with very short insertion tracts (46,47,50,51,55–60). Pol4 is especially adept with primer-template substrates that contain mismatches (49,59,60,93). Previous analyses of Pol4 have hinted at a potential role of the enzyme in HDR, motivated by the highly induced expression of the yeast and mammalian enzymes during meiotic prophase at the time of meiotic recombination; however, no specific role had been defined (54,61–63).
Here, we identified a role of Pol4 in a subset of HDR events, namely spontaneous DRR likely triggered at stalled replication forks. This process is independent of the DNA strand invasion protein Rad51 but depends on Rad52-mediated reannealing. The key result is an unanticipated role of Pol4 in overcoming the inhibition of Polδ-mediated DNA synthesis by Rad51, a mechanism that explains the observed Rad51-dependent role of Pol4 in DRR in vivo. The effect of Rad51 on Polδ is specific, as Polϵ is not affected by Rad51 (Supplementary Figure S6E–G). This may relate to the role of Polδ during lagging strand replication, where Rad51 may assemble at post-replicative gaps. A similar ability to compete with RAD51 DNA binding was identified for human Polθ (26). Specifically, a fragment of Polθ lacking the DNA polymerase domain interacts with human RAD51 and competes with it for DNA binding dependent on the ATPase activity and the RAD51 interaction site of Polθ (26). However, the mechanisms involved appear different, as Pol4 and hPolλ can displace Rad51 from established Rad51-ssDNA filaments, while hPolθ cannot and only inhibits RAD51 DNA binding when added before RAD51 (26). As both, yeast Pol4 and hPolλ, can displace yeast Rad51, it appears unlikely that the mechanism involves specific protein-protein interactions but may be more related to the intrinsic DNA binding properties of the DNA polymerases. The effect of Rad51 in repressing spontaneous DRR may involve additional mechanisms besides the inhibiton of Polδ DNA synthesis. The complete suppression of the Pol4 requirement by deleting RAD51 suggests that the majority of the effect can be explained by the proposed mechanism. The results show that Pol4 DNA synthesis is strongly required for substrates with a single mismatch at the 3’-end, but much less so for perfectly matched primers. Our observation is consistent with the specificity of Pol4 for these substrate type in NHEJ as well as MHMR and may suggest general cooperation between Pol4 and Polδ that appears to be regulated by the processivity clamp PCNA (47,49,59,60,93).
Role of Pol4 in Rad51-independent recombination
Recombination independent of the central DNA strand invasion protein has been extensively studied in bacteria, and RecA-independent recombination between repeated DNA is a well-established phenomenon (94–102). Similar to our observations in yeast (Figure 2), RecA also inhibits RecA-independent recombination in bacteria (95), but genes similar to POL4 that are specifically required for RecA-independent recombination in RecA WT cells have not yet been identified. Recombination independent of the eukaryotic RecA homolog Rad51 has also been documented in eukaryotes including yeast, Arabidopsis thaliana, Drosophila melanogaster, Caenorhabditis elegans, and mammalian cells and involves various mechanisms (3). Most prominent is the SSA pathway in DSB repair, which requires a DSB either experimentally induced by an endonuclease or randomly by IR between two DNA repeats on the same chromatid, leading to a restored chromosome with a deletion of one repeat and the intervening sequence (3). The SSA assay in Supplemental Figure 5 represents a version with two DSBs located on different chromosomes that are adjacent to the DNA repeats, leading to a reciprocal translocation or a single translocation with loss of the other chromosomes (16). This SSA assay is independent of Rad51 but strongly dependent on Rad52 (16), and our data show independence of Pol4. Other mechanisms of repeat recombination do not rely on induced DSBs and are associated with DNA replication involving either DNA polymerase slipping between two repeats or recombination between repeats involving replication-associated DNA gaps exposing single-stranded DNA (98). A critical difference between bacteria and eukaryotes with regard to repeat-recombination at stalled replication forks is the organization of the replicons and respective presence or absence of defined replication-terminating regions. In bacteria with circular chromosomes, stalled forks are required to recover to complete DNA synthesis, whereas in eukaryotes, a stalled fork can wait for the converging fork providing additional opportunities for exposed ssDNA when both forks meet (37). In a process termed Inter-Fork Strand Annealing (IFSA), repeats at two converging stalled forks can engage via strand annealing leading to deletion of one repeat and the intervening DNA sequence on one sister chromatid (Supplementary Figure S10 (37)). While DNA polymerase slippage remains a formal possibility for the events observed here (Figure 2, Supplementary Figure S5), we favor the IFSA model by Morrow et al. (37), as it can explain additional features of the Pol4-dependent events.
IFSA describes the situation at two converging forks with DNA repeats present in replication-associated gaps or lagging strand and in a regressed fork (Supplementary Figure S10 (37)). We surmise the following steps to explain our experimental data. First, fork regression provides the opportunity for a DNA polymerase (maybe Polδ as this represents the lagging strand but it is currently unclear which polymerase is responsible) to generate a blunt DSB. Second, the blunt DSB constitutes a substrate for untemplated DNA synthesis and addition of a single nt to the 3’-end, which represents a plausible mechanism to generate ends with a single unmatched 3’-flap that specifically require Pol4-mediated DNA synthesis. Untemplated DNA synthesis leading to the addition of a single nucleotide to the 3’-OH end of a blunt DSB has been observed with many DNA polymerases, especially X family members such as Pol4, but also the B family yeast DNA Pol1 (103,104). It is currently not clear which DNA polymerase in yeast would carry out this step. The significant difference between the pol4Δ and the pol4-CD mutant (Figure 2) suggests that only a subset of ends contain a 1 nt extension. Third, after Exo1-mediated degradation of the 5’-ending strand, the +1 nt 3’ tail engages in Rad52-mediated annealing with the repeat at the converging stalled fork leading to a primer template with a single nt 3’-flap. Fourth, Rad51 has the potential to bind even small target gaps with a binding site size of 3 nt, which blocks extension by Polδ, requiring displacement by Pol4. Pol4-mediated extension of the single nt flap is accompanied by Rad51 displacement. Pol4 DNA synthesis is independent of PCNA (Supplementary Figure S6). Fifth, the distributive nature of Pol4 leads to the incorporation of one or few nucleotides. Sixth, PCNA-mediated polymerase exchange to Polδ positions Polδ for long-range DNA synthesis with the ability to proofread the error resulting from the + 1 nt flap. The limited Pol4 DNA synthesis ensures that the mismatch remains in the proofreading window of Polδ (105,106). While all substrates require Pol4 to displace Rad51 to allow Polδ access (and/or PCNA loading), +1 nt 3’-flaps specifically require Pol4 DNA synthesis. While Polδ efficiently proofreads such substrates in vitro and in vivo (107,108), it is precluded from acting before Pol4 displacement of Rad51, and Pol4 is devoid of proofreading activity (50). The alternate 3’-flap processing pathway by Rad1-Rad10 (human XPF-ERCC1) is likely inefficient to process such minimal flaps as it incises the single-stranded flap 7–8 nt away from the junction (109). The other 3’-flap endonuclease, Mus81-Mms4 incises the duplex DNA upstream of the junction but requires a 5’ end within 5–9 nt of the branch point (110,111). Hence, we infer that small flaps outside the context of small gap (<5 nt) required by Mus81-Mms4 are the biological substrate for Polδ proofreading, necessitating the more complex route of Pol4-mediated extension and subsequent Polδ proofreading on Rad51 bound substrates, on which Polδ cannot engage. Pol4 may also be involved in gap filling, and the partial requirement for the Pol4 catalytic activity may also reflect the size of the DNA gap involved.
It is currently unclear how these findings relate to the mild IR sensitivity documented for pol4 mutants (Figure 1). Besides DSBs, IR also induces ssDNA nicks and base damages (43), and it is possible that IR-induced base damage leads to fork stalling triggering the DRR events outlined in Supplementary Figure S10. While the IR-sensitivity between the pol4Δ and pol4-CD mutants was indistinguishable, the pol4Δ had a slightly stronger effect with 415 bp repeats but the same defect with 103 bp repeats (Figures 1, 2). It is unclear whether this difference is biologically significant that would hint at additional roles of Pol4 to mediate IR resistance. Our results do not provide an explanation for the meiotic induction of Pol4 transcription, as we failed to identify a role of Pol4 in meiotic recombination.
In sum, our data document a specific role of yeast Pol4 in a subset of HDR-mediated events that underline the collaboration between Pol4 and Polδ, which had been recognized in NHEJ and MHMR. The key ability of Pol4 to displace Rad51 is conserved in human Polλ, opening the potential for a similar role of this enzyme in human cells, consistent with phenotypes associated with a human POLL mutant (64). The overall process exploits the ability of Pol4 to extend mismatched primer-templates and its distributive DNA synthesis as well as the superior proofreading activity and high-fidelity processive DNA synthesis of Polδ to achieve higher overall fidelity.
DATA AVAILABILITY
The data that support the findings of this study are presented in the main text paper and in the online Supplementary Information file. Further information and requests for resources and reagents should be directed to and will be fulfilled by the corresponding authors, Wolf-Dietrich Heyer (wdheyer@ucdavis.edu).
Supplementary Material
ACKNOWLEDGEMENTS
We thank Diedre Reitz and Hang Phuong Le for helpful discussion and comments on the manuscript, Adam Bailis, Neil Hunter, and Lorraine Symington for providing strains, Stephen Kowalczykowski for providing Rad52 protein, and Peter Burgers for providing PCNA, RFC, and Polδ.
Notes
Present address: Paula G. Cerqueira, Inscripta Inc, 5720 Stoneridge Drive #300, Pleasanton, CA 94588, USA.
Present address: Damon Meyer, California Northstate University, College of Health Sciences, 2910 Prospect Park Drive, Rancho Cordova, CA 95670, USA.
Present address: Lilin Zhang, School of Life Science, Tianjin University, No. 92 Weijun Road, Nankai District, Tianjin, 300072, PR China.
Present address: Benjamin Mallory, Genome Sciences Graduate Program, University of Washington, Seattle, WA 98195, USA.
Present address: Becky Xu Hua Fu, ARC Institute, 3181 Porter Drive, Palo Alto, CA 94304, USA.
Contributor Information
Paula G Cerqueira, Department of Microbiology and Molecular Genetics, University of California, Davis, One Shields Avenue, Davis, CA 95616-8665, USA.
Damon Meyer, Department of Microbiology and Molecular Genetics, University of California, Davis, One Shields Avenue, Davis, CA 95616-8665, USA.
Lilin Zhang, Department of Microbiology and Molecular Genetics, University of California, Davis, One Shields Avenue, Davis, CA 95616-8665, USA.
Benjamin Mallory, Department of Microbiology and Molecular Genetics, University of California, Davis, One Shields Avenue, Davis, CA 95616-8665, USA.
Jie Liu, Department of Microbiology and Molecular Genetics, University of California, Davis, One Shields Avenue, Davis, CA 95616-8665, USA.
Becky Xu Hua Fu, Department of Microbiology and Molecular Genetics, University of California, Davis, One Shields Avenue, Davis, CA 95616-8665, USA.
Xiaoping Zhang, Department of Microbiology and Molecular Genetics, University of California, Davis, One Shields Avenue, Davis, CA 95616-8665, USA.
Wolf-Dietrich Heyer, Department of Microbiology and Molecular Genetics, University of California, Davis, One Shields Avenue, Davis, CA 95616-8665, USA; Department of Molecular and Cellular Biology, University of California, Davis, One Shields Avenue, Davis, CA 95616-8665, USA.
SUPPLEMENTARY DATA
Supplementary Data are available at NAR Online.
FUNDING
Resources from Peter Burgers were supported by NIH [GM32431 to P.B.]; P.C. was partially supported by a fellowship from the Science without Borders Program of Brazil; D.M. was partially supported by fellowship 15IB-0109 from the CBCRP and the T32 training program in Oncogenic Signals and Chromosome Biology [CA108459]; L.Z. was supported by a sabbatical scholarship under the State Scholarship Fund by China Scholarship Council; B.X.H.F. had partial support from a Presidential Undergraduate Fellowship from the University of California; National Institutes of Health [GM58015 and GM137751 to W.D.H.]; Cancer Center Core Support Grant NCI [P30CA093373]. Funding for open access charge: NIH [GM58015].
Conflict of interest statement. None declared.
REFERENCES
- 1. Kowalczykowski S.C., Hunter N., Heyer W.-D.. DNA Recombination. 2016; Cold Spring Harbor Laboratory Press. [Google Scholar]
- 2. Kowalczykowski S.C. An overview of the molecular mechanisms of recombinational DNA repair. Cold Spring Harbor Perspect. Biol. 2015; 7:a016410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Bhargava R., Onyango D.O., Stark J.M.. Regulation of single-strand annealing and its role in genome maintenance. Trends Genet. 2016; 32:566–575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Mehta A., Haber J.E.. Sources of DNA double-strand breaks and models of recombinational DNA repair. Cold Spring Harbor Perspect. Biol. 2014; 6:a016428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Krogh B.O., Symington L.S.. Recombination proteins in yeast. Annu. Rev. Genet. 2004; 38:233–271. [DOI] [PubMed] [Google Scholar]
- 6. Ivanov E.L., Sugawara N., Fishmanlobell J., Haber J.E.. Genetic requirements for the single-strand annealing pathway of double-strand break repair in Saccharomyces cerevisiae. Genetics. 1996; 142:693–704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Prakash R., Zhang Y., Feng W.R., Jasin M.. Homologous recombination and human health: the roles of BRCA1, BRCA2, and associated proteins. Cold Spring Harbor Perspect. Biol. 2015; 7:a016600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Heyer W.D., Ehmsen K.T., Liu J.. Regulation of homologous recombination in eukaryotes. Annu. Rev. Genet. 2010; 44:113–139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Kadyk L.C., Hartwell L.H.. Sister chromatids are preferred over homologs as substrates for recombinational repair in Saccharomyces cerevisiae. Genetics. 1992; 132:387–402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Lin F.-L., Sperle K., Sternberg N.. Model for homologous recombination during transfer of DNA into mouse L cells: role for DNA ends in the recombination process. Mol. Cell. Biol. 1984; 4:1020–1034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Lin F.-L., Sperle K., Sternberg N.. Intermolecular recombination between DNAs introduced into mouse L cells Is mediated by a nonconservative pathway that leads to crossover products. Mol. Cell. Biol. 1990; 10:103–112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Symington L.S., Gautier J.. Double-strand break end resection and repair pathway choice. Annu. Rev. Genet. 2011; 45:247–271. [DOI] [PubMed] [Google Scholar]
- 13. McVey M., Khodaverdian V.Y., Cerqueira P., Heyer W.-D.. Eukaryotic DNA polymerases in homologous recombination. Annu. Rev. Genet. 2016; 50:393–421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Sugiyama T., New J.H., Kowalczykowski S.C.. DNA annealing by Rad52 Protein is stimulated by specific interaction with the complex of replication protein A and single-stranded DNA. Proc. Natl. Acad. Sci. U.S.A. 1998; 95:6049–6054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Pavlicek A., Noskov V.N., Kouprina N., Barrett J.C., Jurka J., Larionov V.. Evolution of the tumor suppressor BRCA1 locus in primates: implications for cancer predisposition. Hum. Mol. Genet. 2004; 13:2737–2751. [DOI] [PubMed] [Google Scholar]
- 16. Manthey G.M., Bailis A.M.. Rad51 inhibits translocation formation by non-conservative homologous recombination in Saccharomyces cerevisiae. PLoS One. 2010; 5:e11889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Argueso J.L., Westmoreland J., Mieczkowski P.A., Gawel M., Petes T.D., Resnick M.A.. Double-strand breaks associated with repetitive DNA can reshape the genome. Proc. Natl. Acad. Sci. U.S.A. 2008; 105:11845–11850. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Piazza A., Heyer W.D.. Multi-invasion-induced rearrangements as a pathway for physiological and pathological recombination. Bioessays. 2018; 40:e1700249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Piazza A., Heyer W.-D.. Homologous recombination and the formation of complex genomic rearrangements. Trends Cell Biol. 2019; 29:135–149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Weckselblatt B., Rudd M.K.. Human structural variation: mechanisms of chromosome rearrangements. Trends Genet. 2015; 31:587–599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Zhao B.L., Rothenberg E., Ramsden D.A., Lieber M.R.. The molecular basis and disease relevance of non-homologous DNA end joining. Nat. Rev. Mol. Cell Biol. 2020; 21:765–781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Schimmel J., van Schendel R., den Dunnen J.T., Tijsterman M.. Templated insertions: a smoking gun for polymerase theta-mediated end joining. Trends Genet. 2019; 35:632–644. [DOI] [PubMed] [Google Scholar]
- 23. Llorens-Agost M., Ensminger M., Le H.P., Gawai A., Liu J., Cruz-Garcia A., Bhetawal S., Wood R.D., Heyer W.D., Lobrich M.. POL theta-mediated end joining is restricted by RAD52 and BRCA2 until the onset of mitosis. Nat. Cell Biol. 2021; 23:1095–1105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Ramsden D.A., Carvajal-Garcia J., Gupta G.P.. Mechanism, cellular functions and cancer roles of polymerase-theta-mediated DNA end joining. Nat. Rev Mol Cell Biol. 2022; 23:125–140. [DOI] [PubMed] [Google Scholar]
- 25. Sfeir A., Symington L.S.. Microhomology-mediated end joining: a back-up survival mechanism or dedicated pathway?. Trends Biochem. Sci. 2015; 40:701–714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Ceccaldi R., Liu J.C., Amunugama R., Hajdu I., Primack B., Petalcorin M.I.R., O’Connor K.W., Konstantinopoulos P.A., Elledge S.J., Boulton S.J.et al.. Homologous-recombination-deficient tumours are dependent on Pol theta-mediated repair. Nature. 2015; 518:258–262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Mateos-Gomez P.A., Gong F.D., Nair N., Miller K.M., Lazzerini-Denchi E., Sfeir A.. Mammalian polymerase theta promotes alternative NHEJ and suppresses recombination. Nature. 2015; 518:254–257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Sallmyr A., Tomkinson A.E.. Repair of DNA double-strand breaks by mammalian alternative end-joining pathways. J. Biol. Chem. 2018; 293:10536–10546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Boulton S.J., Jackson S.P.. Saccharomyces cerevisiae Ku70 potentiates illegitimate DNA double-strand break repair and serves as a barrier to error-prone DNA repair pathways. EMBO J. 1996; 15:5093–5103. [PMC free article] [PubMed] [Google Scholar]
- 30. Chapman J.R., Taylor M.R., Boulton S.J.. Playing the end game: DNA double-strand break repair pathway choice. Mol. Cell. 2012; 47:497–510. [DOI] [PubMed] [Google Scholar]
- 31. Chen R., Wold M.S.. Replication protein A: single-stranded DNA’s first responder. Bioessays. 2014; 36:1156–1161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Kim C.S., Paulus B.F., Wold M.S.. Interactions of human replication protein A with oligonucleotides. Biochemistry. 1994; 33:14197–14206. [DOI] [PubMed] [Google Scholar]
- 33. Blackwell L.J., Borowiec J.A.. Human replication protein a binds single-stranded DNA in two distinct complexes. Mol. Cell. Biol. 1994; 14:3993–4001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Petropoulos M., Halazonetis T.D.. Delayed DNA break repair for genome stability. Nat. Cell Biol. 2021; 23:1055–1057. [DOI] [PubMed] [Google Scholar]
- 35. Li X., Heyer W.D.. Homologous recombination in DNA repair and DNA damage tolerance. Cell Res. 2008; 18:99–113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Kolodner R.D., Putnam C.D., Myung K.. Maintenance of genome stability in Saccharomyces cerevisiae. Science. 2002; 297:552–557. [DOI] [PubMed] [Google Scholar]
- 37. Morrow C.A., Nguyen M.O., Fower A., Wong I.N., Osman F., Bryer C., Whitby M.C.. Inter-fork strand annealing causes genomic deletions during the termination of DNA replication. Elife. 2017; 6:1–15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Lambert S., Mizuno K., Blaisonneau J., Martineau S., Chanet R., Freon K., Murray J.M., Carr A.M., Baldacci G.. Homologous recombination restarts blocked replication forks at the expense of genome rearrangements by template exchange. Mol. Cell. 2010; 39:346–359. [DOI] [PubMed] [Google Scholar]
- 39. Lettier G., Feng Q., Antunez de Mayolo A., Erdeniz N., Reid R.J.D., Lisby M., Mortensen U.H., Rothstein R.. The role of DNA double-strand breaks in spontaneous homologous recombination in S. cerevisiae. PLoS Genetics. 2006; 2:e194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. St Charles J., Petes T.D.. High-resolution mapping of spontaneous mitotic recombination hotspots on the 1.1 Mb Arm of yeast chromosome IV. PLos Genet. 2013; 9:e1003434. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Hum Y.F., Jinks-Robertson S.. DNA strand-exchange patterns associated with double-strand break-induced and spontaneous mitotic crossovers in Saccharomyces cerevisiae. PLos Genet. 2018; 14:e1007302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Zellweger R., Dalcher D., Mutreja K., Berti M., Schmid J.A., Herrador R., Vindigni A., Lopes M.. Rad51-mediated replication fork reversal is a global response to genotoxic treatments in human cells. J. Cell Biol. 2015; 208:563–579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Friedberg E.C., Walker G.C., Siede W.. DNA Repair and Mutagenesis. 1995; Washington, DC: ASM Press. [Google Scholar]
- 44. Strathern J.N., Shafer B.K., Mcgill C.B.. DNA synthesis errors associated with double-strand-break repair. Genetics. 1995; 140:965–972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Yang Y., Sterling J., Storici F., Resnick M.A., Gordenin D.A.. Hypermutability of damaged single-strand DNA formed at double-strand breaks and uncapped telomeres in yeast Saccharomyces cerevisiae. PLoS Genet. 2008; 4:e1000264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. McCulloch S.D., Kunkel T.A.. The fidelity of DNA synthesis by eukaryotic replicative and translesion synthesis polymerases. Cell Res. 2008; 18:148–161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Meyer D., Fu B.X.H., Heyer W.D.. DNA polymerases delta and lambda cooperate in repairing double-strand breaks by microhomology-mediated end-joining in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. U.S.A. 2015; 112:E6907–E6916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Layer J.V., Debaize L., Van Scoyk A., House N.C., Brown A.J., Liu Y.P., Stevenson K.E., Hemann M., Roberts S.A., Price B.D.et al.. Polymerase delta promotes chromosomal rearrangements and imprecise double-strand break repair. Proc. Natl. Acad. Sci. U.S.A. 2020; 117:27566–27577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Chan C.Y., Galli A., R.H. S.. Pol3 is involved in nonhomologous end-joining in Saccharomyces cerevisiae. DNA Repair (Amst.). 2008; 7:1531–1541. [DOI] [PubMed] [Google Scholar]
- 50. Yamtich J., Sweasy J.B.. DNA polymerase Family X: function, structure, and cellular roles. Biochim. Biophys. Acta. 2010; 1804:1136–1150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Aravind L., Koonin E.V.. DNA polymerase beta-like nucleotidyltransferase superfamily: identification of three new families, classification and evolutionary history. Nucleic. Acids. Res. 1999; 27:1609–1618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Bebenek K., Garcia-Diaz M., Patishall S.R., Kunkel T.A.. Biochemical properties of Saccharomyces cerevisiae DNA polymerase IV. J. Biol. Chem. 2005; 280:20051–20058. [DOI] [PubMed] [Google Scholar]
- 53. Prasad R., Widen S.G., Singhal R.K., Watkins J., Prakash L., Wilson S.H.. Yeast open reading frame YCR14C encodes a DNA beta-polymerase-like enzyme. Nucleic. Acids. Res. 1993; 21:5301–5307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Shimizu K., Santocanale C., Ropp P.A., Longhese M.P., Plevani P., Lucchini G., Sugino A.. Purification and characterization of a new DNA polymerase from budding yeast Saccharomyces cerevisiae. A probable homolog of mammalian DNA polymerase beta. J. Biol. Chem. 1993; 268:27148–27153. [PubMed] [Google Scholar]
- 55. Bebenek K., Pedersen L.C., Kunkel T.A.. Structure-function studies of DNA polymerase lambda. Biochemistry. 2014; 53:2781–2792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Lee K., Lee S.E.. Saccharomyces cerevisiae Sae2- and Tell-dependent single-strand DNA formation at DNA break promotes microhomology-mediated end joining. Genetics. 2007; 176:2003–2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Pardo B., Ma E., Marcand S.. Mismatch tolerance by DNA polymerase Pol4 in the course of nonhomologous end joining in Saccharomyces cerevisiae. Genetics. 2006; 172:2689–2694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Picher A.J., Garcia-Diaz M., Bebenek K., Pedersen L.C., Kunkel T.A., Blanco L.. Promiscuous mismatch extension by human DNA polymerase lambda. Nucleic. Acids. Res. 2006; 34:3259–3266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Meyer D., Fu B.X.H., Chavez M., Loeillet S., Cerqueira P.G., Nicolas A., Heyer W.D.. Cooperation between non-essential DNA polymerases contributes to genome stability in Saccharomyces cerevisiae. DNA Repair (Amst.). 2019; 76:40–49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Wilson T.E., Lieber M.R.. Efficient processing of DNA ends during yeast nonhomologous end joining. Evidence for a DNA polymerase beta (Pol4)-dependent pathway. J. Biol. Chem. 1999; 274:23599–23609. [DOI] [PubMed] [Google Scholar]
- 61. Garcia-Diaz M., Dominguez O., Lopez-Fernandez L.A., de Lera L.T., Saniger M.L., Ruiz J.F., Parraga M., Garcia-Ortiz M.J., Kirchhoff T., del Mazo J.et al.. DNA polymerase lambda (Pol lambda), a novel eukaryotic DNA polymerase with a potential role in meiosis. J. Mol. Biol. 2000; 301:851–867. [DOI] [PubMed] [Google Scholar]
- 62. Sakamoto A., Iwabata K., Koshiyama A., Sugawara H., Yanai T., Kanai Y., Takeuchi R., Daikuhara Y., Takakusagi Y., Sakaguchi K.. Two X family DNA polymerases, lambda and mu, in meiotic tissues of the basidiomycete, Coprinus cinereus. Chromosoma. 2007; 116:545–556. [DOI] [PubMed] [Google Scholar]
- 63. Leem S.H., Ropp P.A., Sugino A.. The yeast Saccharomyces cerevisiae DNA polymerase IV: possible involvement in double strand break DNA repair. Nucleic Acids Res. 1994; 22:3011–3017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64. Capp J.P., Boudsocq F., Bergoglio V., Trouche D., Cazaux C., Blanco L., Hoffmann J.S., Canitrot Y.. The R438W polymorphism of human DNA polymerase lambda triggers cellular sensitivity to camptothecin by compromising the homologous recombination repair pathway. Carcinogenesis. 2010; 31:1742–1747. [DOI] [PubMed] [Google Scholar]
- 65. Sterling C.H., Sweasy J.B.. DNA polymerase 4 of Saccharomyces cerevisiae is important for accurate repair of methyl-methanesulfonate-induced DNA damage. Genetics. 2006; 172:89–98. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Burke D.J., Dawson D., Stearns T.. Methods in Yeast Genetics: A Cold Spring Harbor Laboratory Course Manual. 2000; Cold Spring Harbor Laboratory Press. [Google Scholar]
- 67. Goldstein A.L., McCusker J.H.. Three new dominant drug resistance cassettes for gene disruption in Saccharomyces cerevisiae. Yeast. 1999; 15:1541–1553. [DOI] [PubMed] [Google Scholar]
- 68. Solinger J.A., Lutz G., Sugiyama T., Kowalczykowski S.C., Heyer W.-D.. Rad54 protein stimulates heteroduplex DNA formation in the synaptic phase of DNA strand exchange via specific interactions with the presynaptic Rad51 nucleoprotein filament. J. Mol. Biol. 2001; 307:1207–1221. [DOI] [PubMed] [Google Scholar]
- 69. Lea D.E., Coulson C.A.. The distribution of the numbers of mutants in bacterial populations. J. Genet. 1949; 49:399–406. [DOI] [PubMed] [Google Scholar]
- 70. Lao J.P., Cloud V., Huang C.C., Grubb J., Thacker D., Lee C.Y., Dresser M.E., Hunter N., Bishop D.K.. Meiotic crossover control by concerted action of Rad51-Dmc1 in homolog template bias and robust homeostatic regulation. PLoS Genet. 2013; 9:e1003879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Fasullo M.T., Davis R.W.. Recombinational substrates designed to study recombination between unique and repetitive sequences in vivo. Proc. Natl. Acad. Sci. U.S.A. 1987; 84:6215–6219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Huang K.N., Symington L.S.. Mutation of the gene encoding protein kinase C1 stimulates mitotic recombination in Saccharomyces cerevisiae. Mol. Cell. Biol. 1994; 14:6039–6045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Maines S., Negritto M.C., Wu X.L., Manthey G.M., Bailis A.M.. Novel mutations in the RAD3 and SSL1 genes perturb genome stability by stimulating recombination between short repeats in Saccharomyces cerevisiae. Genetics. 1998; 150:963–976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Manthey G.M., Bailis A.M.. Multiple pathways promote short-sequence recombination in Saccharomyces cerevisiae. Mol. Cell. Biol. 2002; 22:5347–5356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75. Van Komen S., Macris M., Sehorn M.G., Sung P.. Purification and assays of Saccharomyces cerevisiae homologous recombination proteins. Meth Enzymol. 2006; 408:445–463. [DOI] [PubMed] [Google Scholar]
- 76. Binz S.K., Dickson A.M., Haring S.J., Wold M.S.. Functional assays for replication protein A (RPA). Methods Enzymol. 2006; 409:11–38. [DOI] [PubMed] [Google Scholar]
- 77. Bermudez V.P., Farina A., Raghavan V., Tappin I., Hurwitz J.. Studies on human DNA polymerase epsilon and GINS complex and their role in DNA replication. J. Biol. Chem. 2011; 286:28963–28977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78. Koc K.N., Stodola J.L., Burgers P.M., Galletto R.. Regulation of yeast DNA polymerase delta-mediated strand displacement synthesis by 5 '-flaps. Nucleic Acids Res. 2015; 43:4179–4190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79. Wilson T.E., Grawunder U., Lieber M.R.. Yeast DNA ligase IV mediates non-homologous DNA end joining. Nature. 1997; 388:495–498. [DOI] [PubMed] [Google Scholar]
- 80. Teo S.H., Jackson S.P.. Identification of Saccharomyces cerevisiae DNA ligase .4. Involvement in DNA double-strand break repair. EMBO J. 1997; 16:4788–4795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81. Herrmann G., Lindahl T., Schar P.. Saccharomyces cerevisiae LIF1: a function involved in DNA double-strand break repair related to mammalian XRCC4. EMBO J. 1998; 17:4188–4198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82. Halder S., Ranjha L., Taglialatela A., Ciccia A., Cejka P.. Strand annealing and motor driven activities of SMARCAL1 and ZRANB3 are stimulated by RAD51 and the paralog complex. Nucleic Acids Res. 2022; 50:8008–8022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83. Dovrat D., Stodola J.L., Burgers P.M., Aharoni A.. Sequential switching of binding partners on PCNA during in vitro Okazaki fragment maturation. Proc. Natl. Acad. Sci. U.S.A. 2014; 111:14118–14123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84. Mondol T., Stodola J.L., Galletto R., Burgers P.M.. PCNA accelerates the nucleotide incorporation rate by DNA polymerase δ. Nucleic Acids Res. 2019; 47:1977–1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85. Li J., Holzschu D.L., Sugiyama T.. PCNA is efficiently loaded on the DNA recombination intermediate to modulate polymerase delta, eta, and zeta activities. Proc. Natl. Acad. Sci. U.S.A. 2013; 110:7672–7677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86. Zaitseva E.M., Zaitsev E.N., Kowalczykowski S.C.. The DNA binding properties of Saccharomyces cerevisiae Rad51 protein. J. Biol. Chem. 1999; 274:2907–2915. [DOI] [PubMed] [Google Scholar]
- 87. Halder S., Sanchez A., Ranjha L., Reginato G., Ceppi I., Acharya A., Anand R., Cejka P.. Double-stranded DNA binding function of RAD51 in DNA protection and its regulation by BRCA2. Mol. Cell. 2022; 82:3553–3565. [DOI] [PubMed] [Google Scholar]
- 88. Burgers P.M.J., Gerik K.J.. Structure and processivity of two forms of Saccharomyces cerevisiae DNA polymerase delta. J. Biol. Chem. 1998; 273:19756–19762. [DOI] [PubMed] [Google Scholar]
- 89. Maga G., Villani G., Ramadan K., Shevelev I., Tanguy Le Gac N., Blanco L., Blanca G., Spadari S., Hubscher U.. Human DNA polymerase lambda functionally and physically interacts with proliferating cell nuclear antigen in normal and translesion DNA synthesis. J. Biol. Chem. 2002; 277:48434–48440. [DOI] [PubMed] [Google Scholar]
- 90. Shimazaki N., Yazaki T., Kubota T., Sato A., Nakamura A., Kurei S., Toji S., Tamai K., Koiwai O.. DNA polymerase lambda directly binds to proliferating cell nuclear antigen through its confined C-terminal region. Genes Cells. 2005; 10:705–715. [DOI] [PubMed] [Google Scholar]
- 91. Wu Y., Kantake N., Sugiyama T., Kowalczykowski S.C.. Rad51 protein controls Rad52-mediated DNA annealing. J. Biol. Chem. 2008; 283:14883–14892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92. New J.H., Kowalczykowski S.C.. Rad52 protein has a second stimulatory role in DNA strand exchange that complements replication protein-A function. J. Biol. Chem. 2002; 277:26171–26176. [DOI] [PubMed] [Google Scholar]
- 93. Tseng S.F., Gabriel A., Teng S.C.. Proofreading activity of DNA polymerase Pol2 mediates 3'-end processing during nonhomologous end joining in yeast. PLoS Genet. 2008; 4:e1000060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94. Romero Z.J., Armstrong T.J., Henrikus S.S., Chen S.H., Glass D.J., Ferrazzoli A.E., Wood E.A., Chitteni-Pattu S., van Oijen A.M., Lovett S.T.et al.. Frequent template switching in postreplication gaps: suppression of deleterious consequences by the Escherichia coli Uup and RadD proteins. Nucleic. Acids. Res. 2020; 48:212–230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95. Dutra B.E., Sutera V.A., Lovett S.T.. RecA-independent recombination is efficient but limited by exonucleases. Proc. Natl. Acad. Sci. U.S.A. 2007; 104:216–221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96. Lovett S.T., Gluckman T.J., Simon P.J., Sutera V.A., Drapkin P.T.. Recombination between repeats in Escherichia coli by a recA-independent, proximity-sensitive mechanism. Mol. Gen. Genet. 1994; 245:294–300. [DOI] [PubMed] [Google Scholar]
- 97. Lovett S.T., Drapkin P.T., Sutera V.A., Gluckmanpeskind T.J.. A sister-strand exchange mfor recA-independent deletion of repeated DNA sequences in Escherichia coli. Genetics. 1993; 135:631–642. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98. Lovett S.T. Template-switching during replication fork repair in bacteria. DNA Repair (Amst.). 2017; 56:118–128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99. Romero Z.J., Chen S.H., Armstrong T., Wood E.A., van Oijen A., Robinson A., Cox M.M.. Resolving Toxic DNA repair intermediates in every E. coli replication cycle: critical roles for RecG, Uup and RadD. Nucleic. Acids. Res. 2020; 48:8445–8460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100. Jain K., Wood E.A., Romero Z.J., Cox M.M.. RecA-independent recombination: dependence on the Escherichia coli RarA protein. Mol. Microbiol. 2021; 115:1122–1137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101. Cao Y., Kogoma T.. The mechanism of recA polA lethality: suppression by RecA-independent recombination repair activated by the lexA(Def) mutation in Escherichia coli. Genetics. 1995; 139:1483–1494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102. Ithurbide S., Bentchikou E., Coste G., Bost B., Servant P., Sommer S.. Single strand annealing plays a major role in RecA-independent recombination between repeated sequences in the radioresistant Deinococcus radiodurans bacterium. PLoS Genet. 2015; 11:e1005636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103. Clark J.M. Novel non-templated nucleotide addition reactions catalyzed by procaryotic and eucaryotic DNA polymerases. Nucleic. Acids. Res. 1988; 16:9677–9686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104. Loc’h J., Delarue M.. Terminal deoxynucleotidyltransferase: the story of an untemplated DNA polymerase capable of DNA bridging and templated synthesis across strands. Curr. Opin. Struct. Biol. 2018; 53:23–31. [DOI] [PubMed] [Google Scholar]
- 105. Jin Y.H., Garg P., Stith C.M.W., Al-Refai H., Sterling J.F., Murray L.J.W., Kunkel T.A., Resnick M.A., Burgers P.M., Gordenin D.A.. The multiple biological roles of the 3 ‘→ 5 ’ exonuclease of Saccharomyces cerevisiae DNA polymerase delta require switching between the polymerase and exonuclease domains. Mol. Cell. Biol. 2005; 25:461–471. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106. Anand R., Beach A., Li K., Haber J.. Rad51-mediated double-strand break repair and mismatch correction of divergent substrates. Nature. 2017; 544:377–381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107. Bauer G.A., Heller H.M., Burgers P.M.. DNA polymerase III from Saccharomyces cerevisiae. I. Purification and characterization. J. Biol. Chem. 1988; 263:917–924. [PubMed] [Google Scholar]
- 108. Pâques F., Haber J.E.. Two pathways for removal of nonhomologous DNA ends during double-strand break repair in Saccharomyces cerevisiae. Mol. Cell. Biol. 1997; 17:6765–6771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109. de Laat W.L., Appeldoorn E., Jaspers N.G., KHoeijmakers J.J.. DNA structural elements requried for ERCC1-XPF endonuclease activity. J. Biol. Chem. 1998; 273:7835–7842. [DOI] [PubMed] [Google Scholar]
- 110. Ehmsen K.T., Heyer W.D.. A junction branch point adjacent to a DNA backbone nick directs substrate cleavage by Saccharomyces cerevisiae Mus81-Mms4. Nucleic. Acids. Res. 2009; 37:2026–2036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111. Bastin-Shanower S.A., Fricke W.M., Mullen J.R., Brill S.J.. The mechanism of Mus81-Mms4 cleavage site selection distinguishes it from the homologous endonuclease Rad1-Rad10. Mol. Cell. Biol. 2003; 23:3487–3496. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The data that support the findings of this study are presented in the main text paper and in the online Supplementary Information file. Further information and requests for resources and reagents should be directed to and will be fulfilled by the corresponding authors, Wolf-Dietrich Heyer (wdheyer@ucdavis.edu).