Abstract
Plasma pharmacokinetic (PK) data are required as an input function for graphical analysis of single positron emission computed tomography/computed tomography (SPECT/CT) and positron emission tomography/CT (PET/CT) data to evaluate tissue influx rate of radiotracers. Dynamic heart imaging data are often used as a surrogate of plasma PK. However, accumulation of radiolabel in the heart tissue may cause overprediction of plasma PK. Therefore, we developed a compartmental model, which involves forcing functions to describe intact and degraded radiolabeled proteins in plasma and their accumulation in heart tissue, to deconvolve plasma PK of 125I-amyloid beta 40 (125I-Aβ40) and 125I-insulin from their dynamic heart imaging data. The three-compartment model was shown to adequately describe the plasma concentration-time profile of intact/degraded proteins and the heart radioactivity time data obtained from SPECT/CT imaging for both tracers. The model was successfully applied to deconvolve the plasma PK of both tracers from their naïve datasets of dynamic heart imaging. In agreement with our previous observations made by conventional serial plasma sampling, the deconvolved plasma PK of 125I-Aβ40 and 125I-insulin in young mice exhibited lower area under the curve than aged mice. Further, Patlak plot parameters extracted using deconvolved plasma PK as input function successfully recapitulated age-dependent plasma-to-brain influx kinetics changes. Therefore, the compartment model developed in this study provides a novel approach to deconvolve plasma PK of radiotracers from their noninvasive dynamic heart imaging. This method facilitates the application of preclinical SPECT/PET imaging data to characterize distribution kinetics of tracers where simultaneous plasma sampling is not feasible.
SIGNIFICANCE STATEMENT
Knowledge of plasma pharmacokinetics (PK) of a radiotracer is necessary to accurately estimate its plasma-to-brain influx. However, simultaneous plasma sampling during dynamic imaging procedures is not always feasible. In the current study, we developed approaches to deconvolve plasma PK from dynamic heart imaging data of two model radiotracers, 125I-amyloid beta 40 (125I-Aβ40) and 125I-insulin. This novel method is expected to minimize the need for conducting additional plasma PK studies and allow for accurate estimation of the brain influx rate.
Introduction
Preclinical characterization of plasma pharmacokinetics (PK) and biodistribution of drugs is required to make crucial decisions in the drug development process related to formulation optimization, establishing the PK/pharmacodynamics (PD) relationship, and determining the first-in-human dose in clinical trials. In vivo small animal imaging is widely used to assess biodistribution of drugs that could be tagged with a contrast agent, and drug distribution to tissues of interest could be assessed using various imaging modalities (Rudin and Weissleder, 2003; Gomes et al., 2011; Ding and Wu, 2012). Specifically, nuclear imaging modalities such as positron emission tomography (PET) and single photon emission computed tomography (SPECT) coupled with computed tomography (CT) are highly sensitive. The dynamic SPECT/CT imaging allows us to estimate rate of drug influx into specific tissues when used with graphical analysis methods like Palak plots (Gjedde, 1982; Patlak et al., 1983). In our previous publications, we used Patlak plots to calculate blood-to-brain influx clearance (Ki) of 125I-insulin and 125I-amyloid beta (125I-Aβ) peptides using dynamic SPECT/CT imaging (Swaminathan et al., 2018; Sharda et al., 2021). However, in these studies, plasma PK and imaging studies were conducted on separate cohorts of animals due to the infeasibility of plasma sampling during in vivo imaging. Limitations of this method are 1) additional reagent, animal, and labor costs and 2) biased estimation of the Patlak plot slope, which denotes the plasma-to-brain influx clearance (Ki). It is imperative to develop novel, low-cost methods to obtain both plasma PK and tissue distribution kinetics information from the same subject.
Dynamic heart imaging data has been reported as a surrogate for plasma PK and has been used as the input function for kinetic analysis of various tracers, including small-molecule 18F-FHBG (Green et al., 2004) and large molecule fluorescent antibody-drug conjugates (Giddabasappa et al., 2016). Bao et al. compared noninvasive heart fluorescence molecular tomography (FMT) and serial blood sampling to determine blood PK of probes ranging from 2 to 150 kDa (Bao et al., 2019). They found excellent agreement in kinetic profiles and associated PK parameters between these two methods and further applied FMT to estimate glomerular filtration rate under various pathologic conditions (Bao et al., 2019). However, for molecules such as insulin and Aβ, which have been the focus of research in our laboratory, this method can overpredict plasma levels because both proteins, especially Aβ, exhibit considerable accumulation in heart tissue (Troncone et al., 2016; Martinez-Naharro et al., 2018). The imaging signal from heart includes not only intravascular tracers but also those in extravascular space. Accordingly, any later analysis could be erroneous if the longitudinal heart dataset is used as a replacement for plasma PK.
To resolve this issue, we have separated the signal coming from blood and extravascular space by constructing a three-component PK model and have successfully deconvolved the plasma PK from dynamic heart imaging data. The model was calibrated using plasma concentration-time and dynamic heart imaging data for two different tracers, 125I-Aβ40 and 125I-insulin. Further, the calibrated model was applied to naïve datasets to derive plasma PK parameters and generate Patlak plots determining brain influx rate of these two tracers. The advantage of this kinetic modeling approach is the ability to infer plasma PK parameters from dynamic heart imaging data, thus eliminating the need for additional plasma PK studies. Moreover, heart dynamic imaging data are obtained from the same subjects in which brain imaging is performed, thereby yielding more accurate estimation of plasma-to-brain influx clearance.
Materials and Methods
Materials
Aβ40 peptide was procured from AAPPTec, LLC (Louisville, KY). Carrier-free Na125I radionuclides were obtained from PerkinElmer Life and Analytical Sciences (Boston, MA). Insulin (Novolin or Humulin) was purchased from Eli Lilly (Indianapolis, IN).
Animals
B6SJLF1/J mice were purchased from Jackson Laboratory (Bar Harbor, ME). Mice were housed in the Mayo Clinic animal care facility with food and water ad libitum under 12-hour light/dark cycles. Experiments were conducted in adherence with the guide for care and use of laboratory animals provided by National Institutes of Health. All protocols were approved by the Mayo Clinic Institutional Animal Care and Use Committee.
Radioiodination of Insulin and Aβ40
Insulin and Aβ40 were radiolabeled with 125I by the chloramine-T procedure as described in our previous publications (Kandimalla et al., 2005). Free 125I was removed from the radiolabeled protein by dialysis overnight in 0.01-M phosphate-buffered saline (PBS). Purity of the 125I-labeled proteins was determined by trichloroacetic acid (TCA) precipitation. Specific activities of 125I-insulin and 125I-Aβ40 were determined to be in the range of 45 − 48 μCi/μg.
Plasma Pharmacokinetics of 125I-Insulin and 125I-Aβ40
The femoral vein and artery of each animal were catheterized under general anesthesia with a mixture of isoflurane (1.5%) and oxygen (4 L/min). A single intravenous injection (100 μCi) of 125I-insulin (n = 12, 8- to 12-month-old females) or 125I-Aβ40 (n = 12, 5- to 8-month-old females) was administered in the femoral vein, and blood (20 μL) was sampled from the femoral artery using heparinized capillary tubes at various time points. Blood samples were diluted with saline and centrifuged to separate the plasma. TCA precipitation of plasma was conducted to separate intact and degraded 125I-insulin or 125I-Aβ40. The supernatant consisted of degraded protein, whereas the precipitate was assumed to contain intact protein. Total radioactivity counts in both supernatant and precipitate were analyzed using a gamma counter (Cobra II; Amersham Biosciences, Piscataway, NJ). A two-compartment model was fitted to plasma concentration-time data of intact proteins using Phoenix WinNonlin Version 8.3 (Certara USA, Inc., Princeton, NJ). Plasma concentration-time data of both degraded tracers were fitted by empirical functions using GraphPad Prism 9.1 (GraphPad Software, La Jolla, CA).
Dynamic SPECT/CT Studies of 125I-Insulin and 125I-Aβ40
To assess dynamic radioactivity changes in heart and brain, mice were administered 500 μCi of 125I-insulin (n = 8, 9- to 12-month-old females) or 125I-Aβ40 (n = 9, 7- to 8-month-old females) through the femoral vein. The animals were immediately imaged for the following 42 minutes at 1-minute intervals by dynamic SPECT/CT (Gamma Medica, Northridge, CA) as described previously (Jaruszewski et al., 2014). Both CT and SPECT images were reconstructed once the scan was acquired. A Feldkamp reconstruction algorithm was used for CT reconstruction, whereas the three-dimensional reprojection reconstruction (3DRP) method was used for SPECT images. The reconstructed images were then loaded into PMOD software (PMOD Technologies, Switzerland), and regions of interest (ROIs) were drawn using CT images.
Determination of Heart Cavity Volume
The volume of the heart cavity (Vh) was calculated separately for 125I-insulin and 125I-Aβ40 by the following equation:
 |
where is the total heart radioactivity at the end of imaging (t = 42 minutes), is the total heart radioactivity after transcardial perfusion, and and are the predicted plasma concentration of intact and degraded proteins at 42 minutes, respectively.
Model Development
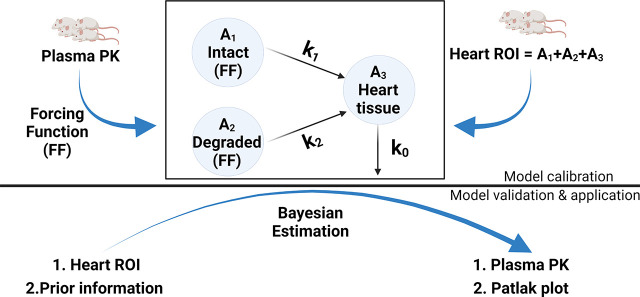
A three-compartment model, depicted in Fig. 2, was constructed, including intact radiolabeled protein (compartment 1), degraded radiolabeled protein (compartment 2) in plasma, and the accumulation of radiolabel in heart tissue (compartment 3). Plasma concentrations of intact proteins followed biexponential decay kinetics, and the heart radioactivity, , was described by:
Fig. 2.
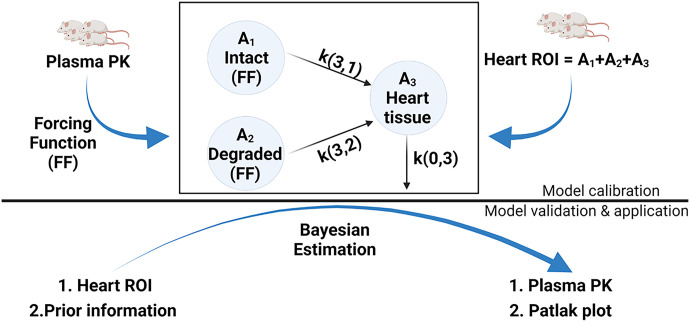
Model scheme and workflow. A three-compartment model was constructed, including intact and degraded radiolabeled proteins in plasma (defined by forcing functions) and their accumulation in heart tissue. The model was fitted to the heart radioactivity-time data, and transfer rate constants were estimated using SAAM II software. The model predictions were further verified using data from naïve animal group. Compartment 1 is intact protein (precipitate in TCA assay) and compartment 2 is degraded protein (supernatant in TCA assay) in the plasma. Compartment 3 is accumulation of radiotracers in the heart tissue. The parameters k1, k2, and k0 are first-order constants that describe transfer rates from compartments.
Plasma radioactivity associated with degraded protein in the heart cavity, , was described by:
where is an empirical function that depends on the protein of interest:
for degraded 125I-insulin and
 |
for degraded 125I-Aβ40.
The amount of radioactivity in heart tissue, , was described using a one-compartment model with the two input rates, and , representing transfer of radioactivity from plasma, and , representing first-order transfer out of the tissue. Thus,
 |
Total heart radioactivity, , was represented by the summation of three components:
The parameters A, α, B, β, C0, C1, YM, Y0, and k were predicted by fitting eq. 3 to concentration-time data of intact protein as well as eqs. 4 and 5 to concentration-time data of degraded 125I-insulin and 125I-Aβ40, respectively. Afterward, the whole model was fitted to heart radioactivity-time data, and the transfer rate constants k1, k2, and k0 were estimated using SAAM II (version 2.3; SAAM Institute, University of Washington, Seattle, WA). The final model was evaluated by diagnostic plots, including weighted residual versus time, weighted residual versus model prediction, and model prediction versus observation plots. Further, the goodness of fit was assessed by Akaike information criterion (AIC) and Bayesian information criterion (BIC), whereas standard error of the parameter estimates was used to evaluate the precision of estimation.
Model Validation and Application
The developed model was validated using independent naïve datasets obtained from three young and three aged animals. To deconvolve plasma PK from the heart imaging data, the model was fitted to heart radioactivity-time data. After fixing the parameters k1, k2, and k0, plasma PK parameters were inferred by the Bayesian estimation method, where prior estimates of population means and standard deviations of each parameter (A, B, α, β), obtained from our previous 125I-insulin (n = 22, 6- to 12-month-old females) and 125I-Aβ40 (n = 27, 6- to 11-month-old females) plasma PK studies, were employed. Heart tissue accumulation (μCi) at the last time point (42 minutes) for each independent animal, including young (n = 3, 3-month-old) and aged (n = 3, 24-month-old) mice, was also predicted and compared with observed heart radioactivity of both 125I-insulin and 125I-Aβ40 after transcardial perfusion. The plasma PK parameters, including maximal concentration (Cmax), clearance (CL), and area under the curve (AUC), were then predicted using the deconvolved plasma concentration-time datasets.
Further, the deconvolved plasma PK was used as input function to construct the Patlak plot by plotting:
 |
where is brain radioactivity at time t, is plasma AUC from the time of injection, and is the predicted plasma concentration of intact tracer at time t. The slope of the linear portion of the Patlak plot is referred to as the plasma-to-brain influx clearance, Ki.
Results
Comparison of Apparent Heart PK and Plasma PK of 125I-Insulin and 125I-Aβ40
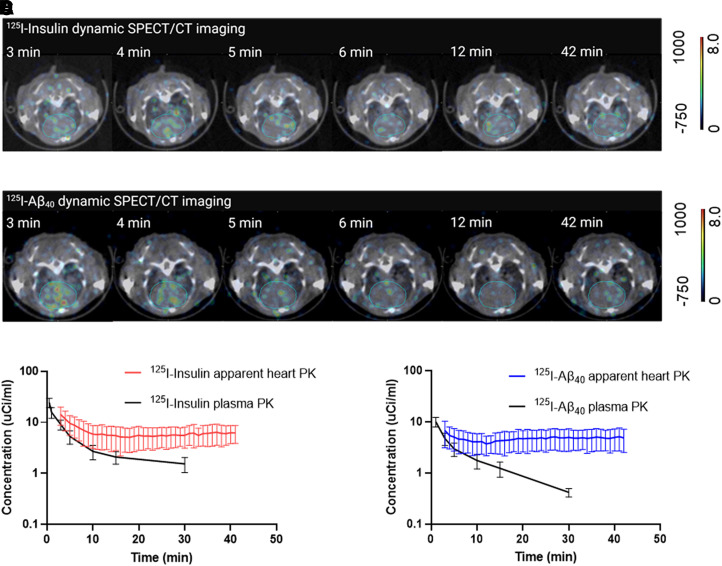
After intravenous injection, the radioactivity signal in the heart detected by dynamic SPECT/CT imaging declined rapidly at early time points but became stable at the later phase (Fig. 1, A and B). Radioactivity was quantified by ROI analysis, and apparent heart concentration was obtained by taking the ratio of radioactivity (μCi) and heart cavity volume (mL; 0.036 mL for 125I-insulin and 0.099 mL for 125I-Aβ40). Notably, the apparent heart concentration (μCi/mL) was found to be higher than the observed plasma concentration of 125I-insulin (Fig. 1C, n = 8) or 125I-Aβ40 (Fig. 1D, n = 9) at all time points.
Fig. 1.
Apparent heart pharmacokinetics obtained by dynamic SPECT/CT imaging overestimates plasma pharmacokinetics of radiolabeled tracers 125I-insulin and 125I-Aβ40. Female B6SJLF1/J mice were intravenously injected with around 500 μCi 125I-insulin (n = 8, 9 to 12 months old) or 125I-Aβ40 (n = 9, 7 to 8 months old), and the heart of each animal was imaged up to 42 minutes by dynamic SPECT/CT. (A, B) Representative images of mice after injection with (A) 125I-insulin and (B) 125I-Aβ40 at 3, 4, 5, 6, 12, and 42 minutes and example regions of interest for data sampling are shown. (C, D) Comparison of apparent heart PK and plasma PK of (C) 125I-insulin and (D) 125I-Aβ40. Data are presented as mean ± S.D. Apparent heart PK was calculated as the ratio of heart radioactivity (μCi) and heart cavity volume Vh (mL; 0.036 mL for 125I-insulin and 0.099 mL for 125I-Aβ40) as described in the Materials and Methods section.
Model Fit of 125I-Insulin and 125I-Aβ40 Heart Radioactivity-Time Profile
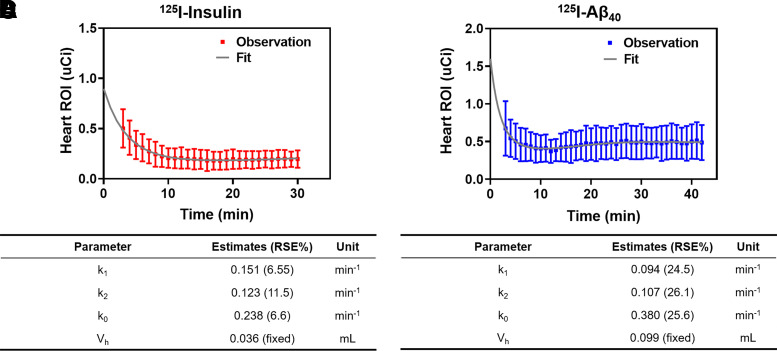
A three-compartment model was constructed, describing the radioactivity (μCi) of intact and degraded radiolabeled proteins in plasma and their accumulation in the heart tissue (Fig. 2). Both intact 125I-insulin (Fig. 3A, n = 12) and 125I-Aβ40 (Fig. 3C, n = 12) exhibited biexponential disposition in plasma. The simple linear eq. 4 adequately described the concentration-time data of degraded 125I-insulin (Fig. 3B, n = 12), whereas the nonlinear eq. 5 was used to fit the concentration-time data of degraded 125I-Aβ40 (Fig. 3D, n = 12). Parameter estimates for all four forcing functions are summarized in Table 1. After fixing the parameters describing plasma disposition of the intact and degraded proteins, the model was fitted to the heart radioactivity-time data as determined by dynamic SPECT/CT imaging after intravenous injection of 125I-insulin (Fig. 4A, n = 8) or 125I-Aβ40 (Fig. 4B, n = 9). The model satisfactorily predicted heart tissue radioactivity, determined after transcardial perfusion, of both 125I-insulin and 125I-Aβ40 with relatively low prediction error (Table 2). The goodness-of-fit plots for heart radioactivity data are displayed for 125I-insulin (Supplemental Fig. 1) and 125I-Aβ40 (Supplemental Fig. 2). Individual predictions were in good agreement with observed values. Weighted residuals were found to be randomly distributed along the zero-ordinate line. Parameter estimates and their relative standard error (RSE%) are presented in Fig. 4, C and D. Relatively low RSE% values associated with the parameter estimates indicated goodness of the model fit.
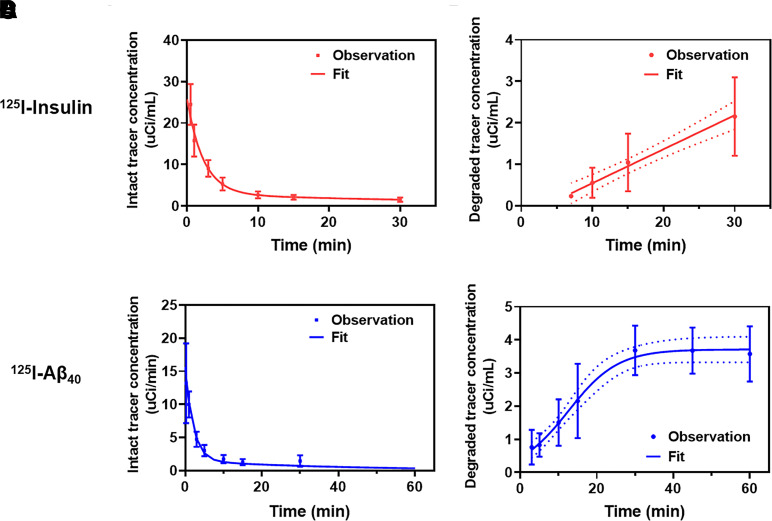
Fig. 3.
Forcing functions describing plasma concentration-time profile of intact and degraded 125I-insulin or 125I-Aβ40. Female B6SJLF1/J mice (n = 12, 9–12 months old for 125I-insulin; n = 12, 7 to 8 months old for 125I-Aβ40) were intravenously injected with 100 μCi of 125I-insulin or 125I-Aβ40. Time series plasma samples were taken at regular intervals and are subjected to TCA precipitation. The plasma concentration (μCi/ml) in precipitate (intact) and supernatant (degraded) were determined by gamma counter. Forcing functions were fitted to the plasma concentration-time data. (A) Intact 125I-insulin; (B) degraded 125I-insulin; (C) intact 125I-Aβ40; (D) degraded 125I-Aβ40. Data are presented as mean ± S.D.
TABLE 1.
Parameter estimates of forcing functions describing plasma concentration-time profiles of intact proteins (A, B, α, β), degraded 125I-insulin (C0, C1), and 125I-Aβ40 (YM, Y0, k) after intravenous bolus injection.
| 125I-Insulin | 125I-Aβ40 | ||
|---|---|---|---|
| Parameters | Estimates (Standard Error) | Parameters | Estimates (Standard Error) |
| A | 22.86 (2.46) | A | 11.95 (0.41) |
| B | 2.90 (0.57) | B | 3.50 (0.14) |
| α | 0.43 (0.060) | α | 0.62 (0.031) |
| β | 0.021 (0.0088) | β | 0.070 (0.002) |
| C0 | −0.27 (0.083) | YM | 3.714 (0.196) |
| C1 | 0.082 (0.0047) | Y0 | 0.463 (0.114) |
| k | 0.155 (0.027) | ||
Fig. 4.
Model fit of the dynamic heart radioactivity data. The model shown in Fig. 2 was fitted to heart radioactivity-time data of (A) 125I-insulin and (B) 125I-Aβ40. Data are presented as mean ± S.D. Parameter estimates and relative standard error were presented in (C) 125I-insulin and (D) 125I-Aβ40. Volume of heart cavity Vh was determined separately as described in the Materials and Methods section and fixed during model fitting. k1, k2, and k0 are the first-order rate constants between the respective compartments.
TABLE 2.
Comparison of model-predicted heart accumulation and observed post-transcardial perfusion heart radioactivity in training and validation datasets.
| Animal | Post-Transcardial Perfusion Heart Radioactivity (μCi) | Predicted Heart Tissue Radioactivity (μCi) | Prediction Error | |
|---|---|---|---|---|
| 125I-Insulin Training Dataset | 0.074 | 0.083 | 12.2% | |
| 125I-Insulin Validation Dataset | Young #1 | 0.114 | 0.081 | −29.3% |
| Young #2 | 0.086 | 0.135 | 57.2% | |
| Young #3 | 0.072 | 0.120 | 65.6% | |
| Aged #1 | 0.164 | 0.113 | −30.8% | |
| Aged #2 | 0.069 | 0.097 | 40.2% | |
| Aged #3 | 0.064 | 0.098 | 53.8% | |
| 125I-Aβ40 Training Dataset | 0.098 | 0.108 | 9.91% | |
| 125I-Aβ40 Validation Dataset | Young #1 | 0.083 | 0.127 | 53.1% |
| Young #2 | 0.083 | 0.132 | 59.8% | |
| Young #3 | 0.081 | 0.107 | 32.6% | |
| Aged #1 | 0.120 | 0.099 | −17.5% | |
| Aged #2 | 0.102 | 0.091 | −10.7% | |
| Aged #3 | 0.069 | 0.082 | 19.4% | |
Model Validation
The model thus developed was applied to deconvolve the plasma PK of both tracers from naïve dynamic heart imaging datasets. The model was found to adequately describe heart radioactivity-time datasets of both 125I-insulin (Fig. 5A) and 125I-Aβ40 (Fig. 6A) for six independent animals, including three young (3-month-old) and three aged (24-month-old) mice. Further, plasma PK parameters, including maximal concentration (Cmax), clearance (CL), and area under the curve (AUC), were predicted using the deconvolved plasma concentration-time datasets. In line with our previous report (Zhou et al., 2022), aged mice have significantly lower clearances compared with young mice for both 125I-insulin (Fig. 5C) and 125I-Aβ40 (Fig. 6C). The model was also validated by comparing model-predicted tracer accumulation in heart tissue and observed heart radioactivity after transcardial perfusion. As shown in Table 2, the predicted values were generally consistent with the observed heart radioactivity.
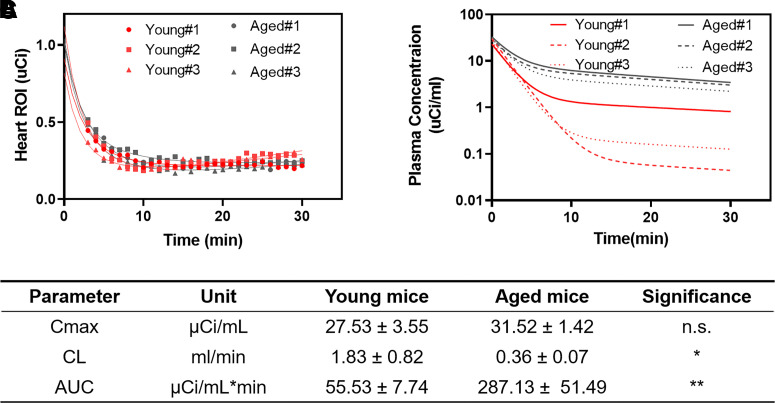
Fig. 5.
Prediction of 125I-insulin plasma PK in young and old mice from independent dynamic heart radioactivity datasets using the calibrated model. (A) Curve fitting of the dynamic heart radioactivity data and (B) deconvolved plasma PK profile for each independent animal. (C) Comparison of various PK parameters derived from deconvolved plasma PK profile in young (n = 3, 3-month-old females) and old (n = 3, 24-month-old females) mice. *P < 0.05, **P < 0.01, student’s t-test. Data are presented as mean ± S.D.
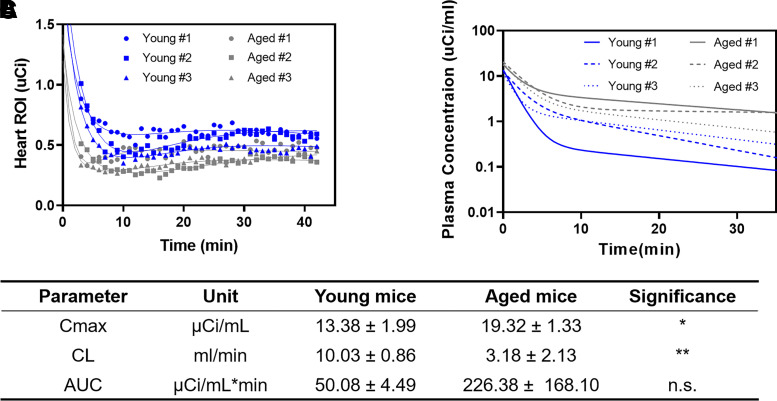
Fig. 6.
Prediction of 125I-Aβ40 plasma PK in young and old mice from independent dynamic heart radioactivity datasets using the calibrated model. (A) Curve fitting of the dynamic heart radioactivity data and (B) deconvolved plasma PK profile for each independent animal. (C) Comparison of various PK parameters derived from deconvolved plasma PK profile in young (n = 3, 3-month-old females) and old (n = 3, 24-month-old females) mice. *P < 0.05, **P < 0.01, student’s t-test. Data are presented as mean ± S.D.
Model Application in Patlak Plot Analysis
Individually predicted plasma PK data from the developed model was employed to construct Patlak plots, and the predicted plasma-to-brain influx clearance (Ki) values were compared between young and aged mice. The representative Patlak plots for both 125I-insulin and 125I-Aβ40 were generated by three methods: using observed plasma PK (Figs. 7A and 8A), deconvolved plasma PK (Figs. 7B and 8B), or apparent heart PK (Figs. 7C and 8C) as the input function. As summarized in Figs. 7D and 8D, methods employing observed plasma PK and deconvolved plasma PK were able to reproduce the Ki difference between young and aged mice, as reported previously (Zhou et al., 2022). The Ki values predicted by these two methods were not significantly different. Using the deconvolved plasma PK, the Ki value (mean ± S.D.) of 125I-insulin was estimated to be 0.0018 ± 0.0008 mL/min in young mice and 0.0003 ± 0.0001 mL/min in aged mice, whereas the Ki values were predicted as 0.0013 ± 0.0005 mL/min in young mice and 0.0005 ± 0.0001 mL/min in aged mice using observed plasma PK. In the case of 125I-Aβ40, the estimated Ki value using deconvolved plasma PK was 0.0047 ± 0.0013 mL/min in young mice and 0.0025 ± 0.0004 mL/min in aged mice, whereas using observed plasma PK, the Ki value was estimated to be 0.0044 ± 0.0014 mL/min in young mice and 0.0018 ± 0.0003 mL/min in aged mice. However, analysis using apparent heart PK cannot capture age-dependent Ki differences. Moreover, the Ki values in aged mice were significantly lower than those analyzed using observed plasma PK.
Fig. 7.
Patlak plot analysis of 125I-insulin in young and aged mice using different input functions. (A–C) Representative Patlak plot using (A) observed plasma PK (B) deconvolved plasma PK and (C) apparent heart PK as input function. (D) Comparison of predicted plasma-to-brain influx clearance values (Ki, mean ± S.D.) from linear regression of respective Patlak plot. *P < 0.05, **P < 0.01, two-way ANOVA followed by multiple comparisons using two-stage set-up method of Benjamini, Krieger, and Yekutieli.
Fig. 8.
Patlak plot analysis of 125I-Aβ40 in young and aged mice using different input functions. (A–C) Representative Patlak plot using (A) observed plasma PK, (B) deconvolved plasma PK, and (C) apparent heart PK as input function. (D) Comparison of predicted plasma-to-brain influx clearance values (Ki, mean ± S.D.) from linear regression of respective Patlak plot. **P < 0.01, ***P < 0.001, two-way ANOVA followed by multiple comparisons using two-stage set-up method of Benjamini, Krieger, and Yekutieli.
Discussion
In nuclear medicine, plasma PK is required as an input function for graphic analyses like Patlak and Logan plots to evaluate the influx kinetics of radiotracers from plasma to peripheral tissue (Patlak et al., 1983; Patlak and Blasberg, 1985; Logan, 2000). Although longitudinal imaging data of heart has been used as a surrogate for plasma PK of various tracers (Green et al., 2004; Giddabasappa et al., 2016), our results demonstrate that apparent heart PK overestimates 125I-insulin (Fig. 1C) and 125I-Aβ40 (Fig. 1D) plasma levels, with larger error in the terminal phase. This observed overprediction was most likely due to substantial accumulation of these tracers, especially Aβ, in the heart tissue. Heart accumulation of amyloid-forming proteins has been widely reported in the literature to cause cardiac amyloidosis (Martinez-Naharro et al., 2018).
In the present study, we developed a three-compartment pharmacokinetic model to deconvolve the plasma PK of radiotracers, 125I-insulin and 125I-Aβ40, from their heart dynamic SPECT/CT imaging data. This procedure provides a novel approach to characterize plasma PK of proteins and peptides by noninvasive imaging and enable more accurate estimation of tissue uptake kinetics of macromolecules under various pathophysiological conditions. Moreover, it allows for substantial reduction in the number of animals needed to reliably estimate the tissue influx rates.
The PK model enables separation of the vascular and extravascular contributions to the total radioactivity in the heart. Additionally, vascular radioactivity could consist of the radioactivity of intact protein (compartment 1) and the degradation products that carry the radiolabel (compartment 2) (Fig. 2). Both insulin (Duckworth et al., 1998) and Aβ (Xiang et al., 2015) metabolites have been shown to circulate in plasma, and each may have distinct disposition profiles from their parent molecules. Therefore, incorporating a separate compartment representing degraded metabolites could significantly improve the predictive ability of a model describing their disposition.
Vascular radioactivity of intact and degraded protein was described by a forcing function that is the product of plasma concentration and volume of the heart cavity. Only plasma instead of whole blood concentration was considered because neither insulin nor Aβ exhibits significant uptake by blood cells based on our previous studies. Therefore, vascular radioactivity was assumed to be substantially contributed by tracers in plasma. In this study, a biexponential equation was found to adequately describe the concentration-time data of both intact 125I-insulin and 125I-Aβ40 peptides (Fig. 3, A and C). These results were consistent with previous reports where two-compartment models were used to fit plasma PK data of these two peptides in mouse (Kandimalla et al., 2005; Zhou et al., 2022).
Unlike intact proteins, the concentrations of degradation products, characterized by supernatant counts after TCA precipitation, were found to increase over time for both 125I-insulin and 125I-Aβ40, possibly due to inefficient clearance of metabolites from systemic circulation. Notably, within the timeframe of the experiment, increase of degraded 125I-insulin was found to be linear (Fig. 3B) whereas 125I-Aβ40 metabolite appearance was nonlinear and reached a plateau at later time points (Fig. 3D). Interestingly, the intercept (C0) of the linear equation describing 125I-insulin metabolite concentration-time profile was found to be negative (Table 1). One possible reason for this outcome is that there is a lag time for 125I-insulin metabolite to appear in the systemic circulation.
Volume of heart cavity (Vh) was separately determined using post-perfusion radioactivity data obtained from heart dynamic imaging. The Vh value for 125I-insulin (35 μl) was found to be lower than that of 125I-Aβ40 (99 μL). One plausible explanation is the facile association of 125I-Aβ40 to the cavity wall and/or to the heart tissue that could be washed off during perfusion. Thus, greater loss of postperfusion radioactivity may apparently increase the estimate of heart cavity volume. This hypothesis was supported by literature reports pertaining to Aβ disposition in the heart. For instance, Aβ receptors are shown to be expressed in various heart cells, including endothelial cells, cardiomyocytes, and fibroblasts (Ramasamy and Schmidt, 2012; Potere et al., 2019). Further, Aβ aggregates have been found in hearts of patients with Alzheimer’s disease and affect myocardial function (Troncone et al., 2016; Martinez-Naharro et al., 2018). Although insulin receptors are ubiquitously expressed and are also highly enriched in the heart (Riehle and Abel, 2016; Abel, 2021), deposition of insulin in heart tissue has not been reported. Nevertheless, the calculated Vh values were comparable to reported anatomic volume of the heart cavity in mice. Previous studies have reported that the stroke volume, which pertains to the volume of blood pumped out of the left ventricle, in mice ranges from 0.025 to 0.04 ml (Barbee et al., 1992). Considering an ejection fraction of around 50% in anesthetized mice (Vinhas et al., 2013), the total blood volume of heart should be around 50–80 μl. After successfully identifying forcing functions of plasma PK and cavity volume, these parameters were fixed and the transfer rate constants (k1, k2, k0) were inferred by fitting the model to the heart radioactivity-time data. As shown in Fig. 4, A and C, the fitted heart radioactivity of both 125I-insulin and 125I-Aβ40 was in excellent agreement with observed data, which was also indicated by the goodness-of-fit plots (Supplemental Figs. 1 and 2). Moreover, all the rate constant estimates were shown to have low RSE%, confirming the precision of the parameter estimation from our data.
The developed model was applied to deconvolve plasma PK profiles of 125I-insulin and 125I-Aβ40 from independent heart radioactivity-time datasets. To further validate the predictive ability of our model, we used heart imaging datasets obtained from both young and aged mice, and the model is expected to reproduce age-dependent plasma PK changes as we reported previously. Specifically, increased plasma exposure (AUC) with aging was observed for both 125I-insulin and 125I-Aβ40 (Zhou et al., 2022). To avoid overparameterization, the Bayesian estimation method, widely used in population PK/PD analysis (Callegari et al., 2002; Bland et al., 2018), was used during curve fitting. The predicted plasma PK profile demonstrated significantly decreased clearance and thus increased AUC in aged mice compared with young mice for both tracers, which is consistent with our previously published results (Zhou et al., 2022). In contrast, when apparent heart PK was used as the input function, the difference between young and old mice was lost due to inaccurate estimation of real plasma PK. Taken together, these results indicated that our modeling approach could successfully deconvolve plasma PK of 125I-insulin and 125I-Aβ40 from heart imaging data under different pathophysiological conditions and reliably capture age-dependent plasma PK changes.
Excellent agreement was observed between model-predicted heart tissue accumulation and observed heart radioactivity after post-transcardial perfusion. However, for 8 out of 12 animals in the validation group, the model appeared to overestimate radioactivity in the heart tissue (Table 2), most likely due to loss of tracers from heart tissue during perfusion, which could yield lower radioactivity than the model prediction.
The modeling approach for deconvolution of plasma PK from dynamic heart imaging data are very useful because: 1) plasma PK analysis and imaging studies can be conducted in a single cohort of animals, substantially reducing the number of animals and conserving resources, and 2) it improves the accuracy and precision of Patlak plot parameter estimates since the plasma PK and dynamic imaging data are obtained from the same mouse. Although bulk heart imaging data has been found to agree with plasma PK of probes that are restricted to the vascular space (Bao et al., 2019), it is not applicable to molecules that are substantially taken up by the heart tissue. For example, a previous study reported that heart ROI time-activity curves (TACs) can overestimate the actual blood concentration of 18F-FDG, especially at later time points after injection. Consequently, the brain influx parameter estimated using heart ROI TAC shows a larger error (Green et al., 1998). Our modeling approach could address this issue by separating the vascular and extravascular radioactivity of the whole heart and serves as a better surrogate for plasma PK. The agreement between model predictions and experimental observations in young versus old mice established the feasibility of our modeling method to investigate plasma-to-tissue influx clearance (Ki) without conducting extra plasma PK studies. Notably, the brain and heart imaging was obtained from the same animal so that the estimated Ki is expected to be more accurate. Successful application of the modeling method established in this study will allow us to infer distribution kinetics of other tracers in both health and disease. Indeed, we have employed this approach to evaluate Ki of a newly developed PET probe 68Ga-NOTA-insulin in wild-type and Alzheimer’s transgenic mice (Taubel et al., 2022).
In conclusion, we have developed a simple compartmental modeling approach to deconvolve plasma PK of radiotracers from noninvasive dynamic heart imaging data obtained from mice. Successful prediction of age-dependent changes in the plasma pharmacokinetics of 125I-insulin and 125I-Aβ40 confirms its utility in assessing systemic disposition of various macromolecular tracers. Moreover, application of this method to generate Patlak plots allows for accurate prediction of tissue influx kinetics of radiotracers, especially when simultaneous plasma sampling is not feasible.
Acknowledgments
The authors would like to acknowledge Ronald J. Sawchuk for his hands-on instructions on the SAAM II software. The graphic abstract/Fig. 2 was generated using BioRender.
Data Availability
The authors declare that all of the data supporting the findings of this study are available within the paper and its Supplemental Data.
Abbreviations
- Aβ
amyloid beta
- AUC
area under the curve
- CT
computed tomography
- RSE
relative standard error
- Ki
influx clearance
- PET
positron emission tomography
- PK
pharmacokinetics
- ROI
region of interest
- SPECT
single positron emission computed tomography
- TCA
trichloroacetic acid
- Vh
volume of heart cavity
Author Contributions
Participated in research design: Z. Wang, L. Wang, Ebbini, Kandimalla.
Conducted experiments: Z. Wang, L. Wang, Curran.
Contributed new reagents or analytic tools: Min, Lowe.
Performed data analysis: Z. Wang, L. Wang.
Wrote or contributed to the writing of the manuscript: Z. Wang, L. Wang, Ebbini, Siegel, Lowe, Kandimalla.
Footnotes
This work was supported by the Minnesota Partnership for Biotechnology and Medical Genomics [MNP#15.31] and National Institutes of Health National Institute of Neurologic Disorders and Stroke [Grant R01NS125437].
Dr. Lowe reported consulting for Bayer Schering Pharma, Piramal Life Sciences, Life Molecular Imaging, Eisai Inc., AVID Radiopharmaceuticals, and Merck Research and receiving research support from GE Healthcare, Siemens Molecular Imaging, AVID Radiopharmaceuticals, and National Institutes of Health National Institute on Aging and National Cancer Institute. The other authors declared no potential conflicts of interests with respect to the research, authorship, and/or publication of this article.
Z.W. and L.W. contributed equally to this work.
A preprint of this article was deposited in bioRxiv [https://doi.org/10.1101/2022.11.17.517003].
 This article has supplemental material available at jpet.aspetjournals.org.
This article has supplemental material available at jpet.aspetjournals.org.
References
- Abel ED (2021) Insulin signaling in the heart. Am J Physiol Endocrinol Metab 321:E130–E145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bao B, Vasquez KO, Ho G, Zhang J, Delaney J, Rajopadhye M, Peterson JD (2019) Blood pharmacokinetics imaging by noninvasive heart fluorescence tomography and application to kidney glomerular filtration rate assessment. J Pharmacol Exp Ther 370:288–298. [DOI] [PubMed] [Google Scholar]
- Barbee RW, Perry BD, Ré RN, Murgo JP (1992) Microsphere and dilution techniques for the determination of blood flows and volumes in conscious mice. Am J Physiol 263:R728–R733. [DOI] [PubMed] [Google Scholar]
- Bland CM, Pai MP, Lodise TP (2018) Reappraisal of contemporary pharmacokinetic and pharmacodynamic principles for informing aminoglycoside dosing. Pharmacotherapy 38:1229–1238. [DOI] [PubMed] [Google Scholar]
- Callegari T, Caumo A, Cobelli C (2002) Generalization of map estimation in SAAM II: validation against ADAPT II in a glucose model case study. Ann Biomed Eng 30:961–968. [DOI] [PubMed] [Google Scholar]
- Ding H, Wu F (2012) Image guided biodistribution and pharmacokinetic studies of theranostics. Theranostics 2:1040–1053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duckworth WC, Bennett RG, Hamel FG (1998) Insulin degradation: progress and potential. Endocr Rev 19:608–624. [DOI] [PubMed] [Google Scholar]
- Giddabasappa A, Gupta VR, Norberg R, Gupta P, Spilker ME, Wentland J, Rago B, Eswaraka J, Leal M, Sapra P (2016) Biodistribution and targeting of anti-5T4 antibody-drug conjugate using fluorescence molecular tomography. Mol Cancer Ther 15:2530–2540. [DOI] [PubMed] [Google Scholar]
- Gjedde A (1982) Calculation of cerebral glucose phosphorylation from brain uptake of glucose analogs in vivo: a re-examination. Brain Res 257:237–274. [DOI] [PubMed] [Google Scholar]
- Gomes CM, Abrunhosa AJ, Ramos P, Pauwels EKJ (2011) Molecular imaging with SPECT as a tool for drug development. Adv Drug Deliv Rev 63:547–554. [DOI] [PubMed] [Google Scholar]
- Green LA, Gambhir SS, Srinivasan A, Banerjee PK, Hoh CK, Cherry SR, Sharfstein S, Barrio JR, Herschman HR, Phelps ME (1998) Noninvasive methods for quantitating blood time-activity curves from mouse PET images obtained with fluorine-18-fluorodeoxyglucose. J Nucl Med 39:729–734. [PubMed] [Google Scholar]
- Green LA, Nguyen K, Berenji B, Iyer M, Bauer E, Barrio JR, Namavari M, Satyamurthy N, Gambhir SS (2004) A tracer kinetic model for 18F-FHBG for quantitating herpes simplex virus type 1 thymidine kinase reporter gene expression in living animals using PET. J Nucl Med 45:1560–1570. [PubMed] [Google Scholar]
- Jaruszewski KM, Curran GL, Swaminathan SK, Rosenberg JT, Grant SC, Ramakrishnan S, Lowe VJ, Poduslo JF, Kandimalla KK (2014) Multimodal nanoprobes to target cerebrovascular amyloid in Alzheimer’s disease brain. Biomaterials 35:1967–1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kandimalla KK, Curran GL, Holasek SS, Gilles EJ, Wengenack TM, Poduslo JF (2005) Pharmacokinetic analysis of the blood-brain barrier transport of 125I-amyloid beta protein 40 in wild-type and Alzheimer’s disease transgenic mice (APP,PS1) and its implications for amyloid plaque formation. J Pharmacol Exp Ther 313:1370–1378. [DOI] [PubMed] [Google Scholar]
- Logan J(2000) Graphical analysis of PET data applied to reversible and irreversible tracers. Nucl Med Biol 27:661–670. [DOI] [PubMed] [Google Scholar]
- Martinez-Naharro A, Hawkins PN, Fontana M (2018) Cardiac amyloidosis. Clin Med (Lond) 18 (Suppl 2):s30–s35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patlak CS, Blasberg RG (1985) Graphical evaluation of blood-to-brain transfer constants from multiple-time uptake data. Generalizations. J Cereb Blood Flow Metab 5:584–590. [DOI] [PubMed] [Google Scholar]
- Patlak CS, Blasberg RG, Fenstermacher JD (1983) Graphical evaluation of blood-to-brain transfer constants from multiple-time uptake data. J Cereb Blood Flow Metab 3:1–7. [DOI] [PubMed] [Google Scholar]
- Potere N, Del Buono MG, Mauro AG, Abbate A, Toldo S (2019) Low density lipoprotein receptor-related protein-1 in cardiac inflammation and infarct healing. Front Cardiovasc Med 6:51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramasamy R, Schmidt AM (2012) Receptor for advanced glycation end products (RAGE) and implications for the pathophysiology of heart failure. Curr Heart Fail Rep 9:107–116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Riehle C, Abel ED (2016) Insulin signaling and heart failure. Circ Res 118:1151–1169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rudin M, Weissleder R (2003) Molecular imaging in drug discovery and development. Nat Rev Drug Discov 2:123–131. [DOI] [PubMed] [Google Scholar]
- Sharda N, Ahlschwede KM, Curran GL, Lowe VJ, Kandimalla KK (2021) Distinct uptake kinetics of Alzheimer disease amyloid-β 40 and 42 at the blood-brain barrier endothelium. J Pharmacol Exp Ther 376:482–490. [DOI] [PubMed] [Google Scholar]
- Swaminathan SK, Ahlschwede KM, Sarma V, Curran GL, Omtri RS, Decklever T, Lowe VJ, Poduslo JF, Kandimalla KK (2018) Insulin differentially affects the distribution kinetics of amyloid beta 40 and 42 in plasma and brain. J Cereb Blood Flow Metab 38:904–918. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taubel JCNelson NRBansal ACurran GLWang LWang ZBerg HMVernon CJMin HKLarson NB, et al. (2022) Design, synthesis, and preliminary evaluation of [68Ga]Ga-NOTA-insulin as a PET probe in an Alzheimer’s disease mouse model. Bioconjug Chem 33:892–906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Troncone L, Luciani M, Coggins M, Wilker EH, Ho CY, Codispoti KE, Frosch MP, Kayed R, Del Monte F (2016) Aβ amyloid pathology affects the hearts of patients with Alzheimer’s disease: mind the heart. J Am Coll Cardiol 68:2395–2407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vinhas M, Araújo AC, Ribeiro S, Rosário LB, Belo JA (2013) Transthoracic echocardiography reference values in juvenile and adult 129/Sv mice. Cardiovasc Ultrasound 11:12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiang YBu XLLiu YHZhu CShen LLJiao SSZhu XYGiunta BTan JSong WH, et al. (2015) Physiological amyloid-beta clearance in the periphery and its therapeutic potential for Alzheimer’s disease. Acta Neuropathol 130:487–499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou AL, Sharda N, Sarma VV, Ahlschwede KM, Curran GL, Tang X, Poduslo JF, Kalari KR, Lowe VJ, Kandimalla KK (2022) Age-dependent changes in the plasma and brain pharmacokinetics of amyloid-β peptides and insulin. J Alzheimers Dis 85:1031–1044. [DOI] [PMC free article] [PubMed] [Google Scholar]