Abstract
In Kazakhstan, there is insufficient data on genetic epilepsy, which has its own clinical and management implications. Thus, this study aimed to use whole genome sequencing to identify and evaluate genetic variants and genetic structure of early onset epilepsy in the Kazakhstani pediatric population. In this study, for the first time in Kazakhstan, whole genome sequencing was carried out among epilepsy diagnosed children. The study involved 20 pediatric patients with early onset epilepsy and no established cause of the disease during the July–December, 2021. The average age at enrolment was 34.5 months, with a mean age at seizure onset of 6 months. Six patients (30%) were male, and 7 were familial cases. We identified pathogenic and likely pathogenic variants in 14 (70%) cases, among them, 6 novel disease gene variants (KCNQ2, CASK, WWOX, MT-CO3, GRIN2D, and SLC12A5). Other genes associated with the disease were SCN1A (x2), SLC2A1, ARX, CACNA1B, PCDH19, KCNT1, and CHRNA2. Identification of the genetic causes in 70% of cases confirms the general structure of the etiology of early onset epilepsy and the necessity of using NGS in diagnostics. Moreover, the study describes new genotype-phenotypic correlations in genetic epilepsy. Despite certain limitations of the study, it can be concluded that the genetic etiology of pediatric epilepsy in Kazakhstan is very broad and requires further research.
Supplementary Information
The online version contains supplementary material available at 10.1007/s12035-023-03346-3.
Keywords: Epilepsy, Genetics, Whole-genome sequencing, Early onset epilepsy, Next-generation sequencing
Introduction
Epilepsy is one of the most common neurological disorders, which affects people of all ages, races, social classes, and geographic locations. Epilepsy is a chronic disease of the brain characterized by a persistent predisposition to the onset of epileptic seizures with neurobiological, cognitive, psychological, and social consequences [1, 2]. In a systematic review and meta-analysis of incidence studies, the pooled incidence of epilepsy was 61.4 per 100,000 person-years, with an overall prevalence of 7.60 per 1000 population; incidence was higher in low/middle-income countries than in high-income countries. Moreover, the incidence of epilepsy is higher in the youngest and oldest age groups, with a rate of 86 per 100,000 per year in the first year of life [3]. Despite a decrease in the prevalence of the disease from 1990 to 2016, epilepsy remains an important cause of disability and death [4].
In the pediatric population, epilepsy is a common neurological disorder affecting approximately 0.5 to 1% of children [5]. According to Jaxybayeva et al. [6] in Kazakhstan, as of October 2020, epilepsy was diagnosed in 15,769 children (aged 0 to 18). Among 450 (32% of total hospitalizations) patients with epilepsy hospitalized in the Department of Pediatric Neurology of the National Scientific Center for Maternal and Child Health of “University Medical Center”; Corporate Found (CF) between 2013 and 2014, 170 of them were diagnosed with epileptic encephalopathy [7]. Early onset epilepsy is associated with poor long-term psychosocial outcomes, and the effects persist into adulthood. People with childhood-onset epilepsy have higher unemployment rates, lower educational attainment, and lower socioeconomic status [5, 8, 9]. According to Lepessova and Myrzaliyeva [10], a high prevalence of epilepsy was observed in regions with unfavorable environmental conditions, which emphasizes the socio-ecological component in the epidemiology of this disease among the pediatric population of Kazakhstan.
Epilepsy is considered a multifactorial disease. The genetic basis of some forms of epilepsy has been put forward for decades and was confirmed by gene mapping and the identification of specific mutations associated with epilepsy syndromes in the 1990s [11–14]. Available data suggests that 70–80% of epilepsy cases have a genetic cause, with the remaining 20–30% associated with acquired conditions such as ischemia, brain injury, tumors, and autoimmune diseases [13, 15]. A review by Wang et al. [16] identified 977 genes associated with epilepsy. At the same time, recent studies have found that genetic causes account for approximately 30% of cases reviewed in the pediatric population [17]. For this reason, genetic testing is widely used today in the practice by epileptologists [18].
Various genetic methods could potentially be applied in the diagnosis of epilepsy. In general, methods such as cytogenetic tests, comparative genomic hybridization (array-CGH), and multiplex ligase-dependent amplification (MLPA) make it possible to diagnose about 10% of pediatric epilepsies and 5% of epileptic encephalopathies [19, 20], since most epilepsies are associated with mutations in individual genes. Currently, traditional Sanger sequencing, which allows direct determination of the nucleotide sequence of a region of a single gene, has been replaced by next-generation sequencing (NGS), which allows simultaneous sequencing of many genes at a relatively low cost [21, 22]. Cohort study conducted in Munich, Germany between 2013 and 2017 indicates that NGS allowed physicians to change the clinical management of 63% of patients with epilepsy [23]. Continuing decline in sequencing costs, the use of WGS is an effective strategy for the clinical diagnosis of early onset epileptic encephalopathy [24]. Another group of researchers from the USA and the UK described two new genes KCNT1 and PIGQ that are pathogenic in Ohtahara syndrome using the WGS method [25].
The first data on whole-genome sequencing in the Kazakh population was published by Akilzhanova et al. [26]. Later, a study conducted among 350 children with early epileptic encephalopathies accompanied by intellectual retardation showed the need for additional funding in healthcare aimed at genetic research of patients with epilepsy; thus, only 15 (4.3%) study participants were able to undergo NGS in the other countries due to financial possibilities, among which 12 (80%) had various genetic mutations and variants associated with the etiology of the disease and resistance to therapy [7].
Thus, this study aims to identify and evaluate genetic variants based on whole-genome sequencing in children with early epilepsy in Kazakhstan.
Materials and Methods
Study design — retrospective single-center study.
Ethical Issues
The study was approved by the local commission on bioethics of the “UMC” CF, an extract from Protocol No. 1 dated 06/29/2021.
Study Participants
The study involved a total of 20 children from unrelated marriages with epilepsy of no known cause and early onset of the disease (according to criteria if the International League Against Epilepsy), hospitalized in the Department of Pediatric Neurology of the National Scientific Center for Maternal and Child Health of the CF “UMC” from July to December 2021, and meeting the following inclusion criteria:
•The onset of a seizures before the age of 3 years;
•The presence of epileptiform discharges on electroencephalography (EEG);
•The absence of structural changes in magnetic resonance imaging of the brain (MRI), which may be the cause of epilepsy;
•Delayed psychomotor development and/or the presence of resistance to antiepileptic drugs and/or the presence of severe epileptic encephalopathy;
•Absence of anomalies found in previous genetic studies (karyotyping);
The criterion for exclusion from the study is the diagnosis in patients before or during the study period of epilepsy with an etiology that could explain the epileptic syndrome. Examples of such etiologies are the brain and meningeal infections, neonatal hypoxic-ischemic encephalopathy, neoplasms, a history of moderate to severe brain injury, and autoimmune diseases affecting the nervous system.
Whole-Genome Sequencing
Genomic DNA was extracted from whole blood samples using the PromegaTM kit (USA) according to the manufacturer’s protocols. DNA libraries were prepared from 300 ng of genomic DNA using Illumina DNA PCR-Free Library Prep, Tagmentation protocol, with the IDT Indexes for Illumina DNA/RNA UD Set A. DNA libraries were validated using the Qubit ssDNA Assay on the Qubit Fluorometer according to the standard kit protocol. Sequencing was performed on the NovaSeq 6000 high throughput platform using the NovaSeq 6000 S4 Reagent Kit v1.5 (300 cycles) at the National Laboratory Astana.
Raw Data Preprocessing and Bioinformatics Analysis
Raw data files obtained from the Illumina NovaSeq sequencing platform in binary base call (bcl) format were converted to the fastq file format using the bcl2fastq tool. The quality of the generated sequences has been evaluated using FastQC v.0.11.9 [27] and MultiQC v.1.12 [28]. Sequencing reads were aligned to the human reference genome (NCBI GRCh37, hg19) using Burrows–Wheeler Aligner v.0.7.12 [29]. Picard tools v.2.27.4 used for sorting and marking reads duplicates. Genome Analysis Toolkit (GATK) v.3.8 has been used for genomic variant calling [30]. Genomic variants were annotated using ANNOVAR [31].
Interpretation of Genetic Variants
Identified genetic variants are described following the nomenclature guidelines of the Human Genome Variation Society (http://www.hgvs.org/mutnomen). Variant interpretation followed the 5-level classification system recommended by the American College of Medical Genetics and Genomics and the Association for Molecular Pathology (ACMG/AMP) and was conducted on the Franklin platform (Franklin by Genoox, Genoox, USA). All possible options identified during the study were evaluated using a three-stage analysis:
1.Variants filtered by their frequency in population control databases;
2.Literature and database review to search for the role of each identified variants in the etiology and course of the disease;
3.Evaluation of the pathogenicity of all identified variants in individual clinical cases.
The following databases are used for variant annotation: OMIM, Human Gene Mutation Database (HGMD), and ClinVar. The pathogenicity of the variants will be predicted using the Polymorphism Phenotyping v2 (PolyPhen-2), MutationTaster, and MutationAssessor prediction algorithms.
Results
DNAs from total of twenty patients were sequenced on Illumina NovaSeq 6000 platform and the total number of sequenced base pairs yielded from 68,8 to 202,7 Gb with the an average 116 Gb per sample. The mean genome coverage for all samples is 35X. On average, 99.15% of sequencing reads have been properly mapped on a reference genome.
Twenty pediatric patients of the Pediatric Neurology Department of the “University Medical Center” СF with an early onset of epilepsy and an unclear cause of the disease were included in the study. Six (30%) of the patients were male. The age of study participants at the time of recruitment ranged from 4 months to 13 years, with an average age of 34.5 months. Seizure onset ranged from the neonatal period to 3 years of age, with an average of 6 months. Among the probands, a burdened family history of epilepsy was noted in 7 cases.
Of 20 patients recruited in the current study, 7 (35%) presented with focal seizures, 5 (25%) with tonic-clonic seizures, 5 (25%) with generalized seizures, 2 (10%) with polymorphic seizures, and 1 (5%) with myoclonic seizures. Five (25%) patients in the cohort developed therapy-resistant seizures, among them, 3 patients were resistant to valproate, 1 — to phenobarbital, and 1 patient was resistance both to phenobarbital and carbamazepine. Epileptic encephalopathy was diagnosed in 5 patients. Thirteen (65%) patients had a psychomotor delay and 10 were diagnosed with speech delay.
After the whole genome sequencing, pathogenic and likely-pathogenic genetic variants were identified in 14 (70%) out of 20 cases. Clinical, phenotype, and genotype data of 14 patients with genetic etiology of epilepsy are presented in Tables 1 and 2. Clinical and phenotypic data, as well as row data of 6 patients without clinically significant variants are presented in supplementary materials.
Table 1.
Genetic diagnosis and phenotype in five familial cases
| Case # (gender) | Gene (Ref Seq) | Genetic variant | Inheritance | Seizures, type | Age of onset, months | Phenotype | EEG data | Brain MRI | Additional information |
|---|---|---|---|---|---|---|---|---|---|
| 1 (f) | KCNQ2 (NM_172107.4) |
c.796_797insGTGCTGTTTCTGCCCTGCCCTCCCTGCCTGGGCTGGAACTCAATAA (p.Asp266GlyfsTer80) |
AD, heterozygous | Focal, tonic | Infancy | PMD | Diffuse generalized discharge of epileptiform spike-wave activity of a continued nature along the left temporal leads with spread to the right homologous leads, also in the anterior-central regions | Expansion of the lateral ventricles, a cyst of the velum interpositum | Aunt (II-1) and grandmother (I-1) on the proband’s mother’s side, sibling (III-3) with epilepsy |
| 2 (f) | WWOX (NM_016373.4) |
c.911C>A (p.Ser304Tyr) c.230+1G>T (Splice donor) |
AR, compound heterozygous | Focal | 1 | PMD, spastic ataxia | Spike-slow wave complexes in the right parietotemporal leads | Moderate mixed hydrocephalus, atrophy of the cerebral hemisphere, post-hypoxic encephalopathy | Sibling (II-3) with epilepsy and spastic ataxia, died at the age of 11 months |
| 3 (f) | CACNA1B (NM_000718.4) |
c.390+1_390+2insACGACACGGAGCCCTATTTCATCGGGATCTTTTGCT (Splice Donor) c.501C>G (p.Asn167Lys) |
AR, compound heterozygous | Generalized, tonic | Infancy | PMD, SD | Diffuse delta activity | - | Sibling (III-2) and cousin (III-1) with epilepsy. The proband’s mother (II-4) had seizures during pregnancy. The proband’s grandfather (I-2) had several episodes of seizures after a traumatic brain injury in a car accident |
| 4 (f) | PCDH19 (NM_001184880.2) | c.1018A>G (p.Asn340Asp) | XL, heterozygous | Clonic-tonic | 11 | PMD, SD, aggression, resistance to valproate | Spike-slow wave complexes in the central-frontal-temporal region bilaterally | - | Three female siblings (II-2-4) with epilepsy |
| 5 (f) | SLC12A5 (NM_020708.5) | c.2930A>G (p.Gln977Arg) | AD, heterozygous | Generalized | 36 | PMD, SD | Periodic diffuse slow waves, low-amplitude single and grouped spike-slow wave complexes in the right central and temporal leads, rarely spreading to the right parts of the brain | Minimal post-hypoxic changes in the white matter of the cerebral hemispheres | Two female siblings (III-1,2) with epilepsy, one of them died at the age of 3 y 4 m (III-2). Paternal cousins (III-4,5) died in infancy from cerebral palsy |
Note: AD, autosomal dominant; AR, autosomal recessive; XL, X-linked; PMD, psychomotor delay; SD, speech delay; f, female
Table 2.
Genetic diagnosis and phenotype in 9 non-familial cases
| Case # (gender) | Gene (Ref Seq) | Genetic variant | Inheritance | Seizures, type | Age of onset, months | Phenotype | EEG data | Brain MRI |
|---|---|---|---|---|---|---|---|---|
| 6 (m) | CASK (NM_001367721.1) |
c.2301T>G c.2300T>G (p.Phe767Gly) |
XLD/XLR, hemizygous | Generalized, tonic | 6 | PMD, EE | Diffuse, bilateral-synchronous and asynchronous peak/spike-slow wave complexes | Diffuse changes in the white matter of the cerebral hemisphere (post-hypoxic genesis), mixed hydrocephalus, retrocerebellar cyst, cyst of the right choroidal fissure |
| 7 (f) | SLC2A1 (NM_006516.4) | c.988C>T (p.Arg330*) | AD, heterozygous | Generalized, clonic-tonic | 3 | PMD, SD, EE, microcephaly | Central leads, spike-slow wave complexes | Diffuse changes in the white matter of the cerebral hemisphere |
| 8 (m) | ARX (NM_139058.3) | c.1612A>G (p.Lys538Glu) | XLR, hemizygous | Polymorphic | 6 | PMD, SD, microcephaly, hypotony | Low-amplitude hypsarrhythmia — constant multifocal, asynchronous, epileptiform activity in the form of spike-slow wave complexes with a focus in the frontal, separately in the temporal and central leads | Post-hypoxic changes in the cerebral hemispheres, mixed hydrocephalus |
| 9 (f) | MT-CO3 | c.172G>A (p.Trp58*) | Mi, homoplasmy | Generalized, tonic | 1 | PMD, SD, acidosis, excess subcutaneous fat | Grouped complexes of a spike-slow wave in the frontal leads with spread to the middle-temporal leads bilaterally | Symmetrical focal changes in the structure of the thalamus of a metabolic/neurodegenerative nature (Leigh syndrome), moderate combined hydrocephalus, retrocerebellar cyst |
| 10 (f) | GRIN2D (NM_000836.4) | c.3550G>A (p.Ala1184Thr) | AD, heterozygous | Polimorphic | 7.5 | SD | Lateralized to the right leads diffuse rhythmic theta-like wave with the beginning in the parietal and temporal leads gradually turning into low-frequency delta waves and spreading to the left parts of the brain | Cysts of the septum pellucidum |
| 11 (f) | KCNT1 (NM_020822.3) | c.1421G>A (p.Arg474His) | AD, heterozygous | Clonic-tonic | 1 | PMD, SD, resistance to fenobarbital, carbamazepine | Rhythmic beta/alpha-like waves in the right occipital and separately in some seizures in the right temporal leads, with diffuse propagation, gradually turning into theta waves | Post-hypoxic changes in the white matter of the cerebral hemispheres, moderate external hydrocephalus |
| 12 (m) | CHRNA2 (NM_000742.4) | c.612G>A (p.Trp204*) | AD, heterozygous | Focal, clonic | Infancy | SD, status epilepticus, resistance to fenobarbital | Bursts of delta waves with a maximum amplitude in the central and temporal leads | Subatrophy of the cerebral hemispheres, bilateral convexital subdural hygroma more pronounced on the right |
| 13 (f) | SCN1A (NM_001165963.4) | c.3706-2A>C (Splice acceptor) | AD, heterozygous | Focal, clonic | 7 | PMD, SD | Continued deceleration in the right frontal leads | Intracranial hypertension |
| 14 (m) | SCN1A (NM_001165963.4) | c.3982T>C (p.Ser1328Pro) | AD, heterozygous | Focal, clonic | 4 | Resistance to levetiracetam | An increase in the index of diffuse slow delta waves, single sharp waves were also recorded in the occipital, separately in the frontal leads | - |
Note: AD, autosomal dominant; XLD, X-linked dominant; XL, XLR, X-linked recessive; Mi, mitochondrial; PMD, psychomotor delay; SD, speech delay; EE, epileptic encephalopathy; m, male; f, female
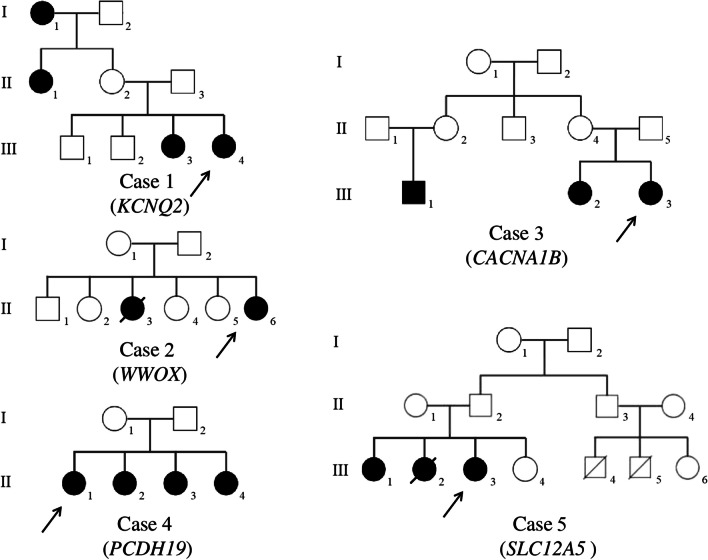
Among 14 patients with identified pathogenic/likely pathogenic genetic variants, a family history of epilepsy was in 5 cases (cases # 1 to 5). Clinical, phenotype, and genotype data of five cases with family history of epilepsy are presented in Table 1; pedigree charts of these cases are presented in Fig. 1.
Fig. 1.
Pedigree charts of the cases with family epilepsy history (created with BioRender.com)
Previously undescribed variants were found in cases 1, 2, 5, 6, 9, and 10 (KCNQ2, WWOX, SLC12A5, CASK, MT-CO3, and GRIN2D genes, respectively). Case 1 is a girl, aged 2 years at the time of genetic analysis. For the first time, convulsions were registered on the second day of life. In this patient, the study revealed a pathogenic variant in the KCNQ2 gene. This variant was a 46 bp insertion followed by a frameshift in the exon 5.
Case 2 is a girl who had her first seizures at the age of 1 month. During counseling, complaints were of seizures up to 30 times a day for 30 s. This patient was found to have compound heterozygous variants in the WWOX gene. Both genetic variants have not previously been described.
Case 5 is a girl with onset of seizures at 3 years old; however, epileptic activity on the EEG was noted earlier. The genetic variant in the proband was represented by a missense mutation in the SLC12A5 gene in the heterozygous state.
Case 6 is a boy with an onset of seizures at 6 months. Two nucleotide substitutions (missense mutations) were found in the CASK gene, leading to a change in the amino acid sequence at position Phe767Gly.
Case 9 is a girl with no family history of epilepsy and a previously undescribed variant in the mitochondrial gene, the cytochrome C oxidase 3 (MT-CO3) gene. This girl also had excess subcutaneous adipose tissue, opticopathy, and partial atrophy of the optic disc. Seizures in this patient were managed by the admission of valproate.
Case 10 is a girl whose first seizure was at 7.5 months on the background of elevated temperature. This patient had previously been screened for microdeletion/microduplication syndrome by MLPA which was negative. In this girl, a previously undescribed and pathogenic variant was identified in the GRIN2D gene.
Resistance to at least one of the antiepileptic drugs was found in cases 4, 11, 12, and 14. Case 4 had resistance to valproate, but seizures were partially controlled by oxcarbazepine. Case 11, with a pathogenic variant in the KCNT1 gene, was resistant to phenobarbital and carbamazepine, while a partial positive response was achieved with topiramate and vigabatrin. Case 12 was resistant to phenobarbital as well as to low doses of valproate, while seizure relief was achieved by administration of 150 mg/day of valproate in combination with levetiracetam at a dosage of 100 mg/day. Case 14 initially received levetiracetam, but the frequency of seizures was taken into account until the onset of status epilepticus. This patient is currently receiving a combination of valproate and carbamazepine.
Discussion
The study describes the results of the first whole genome sequencing among children with early onset epilepsy in Kazakhstan. Pathogenic and likely pathogenic genetic variants were identified in 70% of cases during the study (14/20). This indicator was higher than in previous studies. So, in one study, researchers found the presence of clinically significant genetic mutations in 20% of patients (among 118 children with epilepsy) in Sweden [32]. In other studies from South Korea and Scotland, with more detailed inclusion criteria, NGS detected genetic abnormalities in 37.8% and 24% of patients with early-onset epilepsy, respectively [33, 34]. The higher percentage of clinically relevant genetic variants identified in this study may be explained by the strict inclusion criteria and the small sample size.
The following genes have been identified as epilepsy etiology: ARX, CACNA1B, CASK, CHRNA2, GRIN2D, KCNQ2, KCNT1, MT-CO3, PCDH19, SCN1A (x2), SLC2A1, SLC12A5, and WWOX. In an early study, Jaxybayeva et al. [6] identified a genetic cause in 80% (12/15) of children with early seizures using whole-exome sequencing in Kazakhstan (CDKL5 (x3), SCN1A (x2), MECP (x2), STXBP1, UBE3A, PCDH19, FOLR1, PNPO). Summing up the results from the present and the above mentioned studies, which is acceptable due to relatively similar inclusion criteria, the overall detection rate for the genetic etiology of early-onset epilepsy was 74.3% (26/35). And most often clinically significant genetic variants have been identified in CDKL5 (x3), SCN1A (x3), MECP (x2), and PCDH19 (x2) genes.
KCNQ2-related neonatal-onset developmental and epileptic encephalopathy is characterized by mostly tonic seizures beginning in the first week of life [35], which coincided with the clinical presentation of case 1. GLUT1 deficiency syndrome caused by mutations of the SLC2A1 gene is characterized by early infantile epilepsy, developmental delay, microcephaly, complex movement disorders, and various paroxysmal neurological phenomena [36, 37], which also corresponds to the indicated case 7. This study also expands the understanding of the clinical features of mutations in the ARX gene. X-linked infantile spasms syndrome, West syndrome, Ohtahara syndrome, and myoclonic epilepsy syndromes may be associated with ARX gene mutations and characterized by pediatric epilepsy, intellectual disability, developmental and speech delay, intractable seizures, hypotonia, psychiatric abnormalities, brain malformations, and ambiguous genitalia [38–41]. Previously described variants associated with MT-CO3 (COX) gene were represented by the following clinical characteristics: MELAS syndrome, rhabdomyolysis, and mitochondrial myopathy with lactic acidosis and one with Leigh syndrome ([42–46]. In this study, we expand the clinical manifestation of the MT-CO3-related Leigh syndrome. Moreover, current research confirms and expands the genotype-phenotypic correlation of the following conditions: PCDH19-related epilepsy [47], GRIN2D-related developmental and epileptic encephalopathy [48], KCNT1-related epilepsy [49], CHRNA2-related autosomal dominant sleep-related hypermotor epilepsy [50], SCN1A-related epilepsy [51], SLC12A5-related epilepsy [52].
At the same time, case 6 differed from the previously described phenotypic manifestations. So, variants in the CASK gene were mainly the cause of epilepsy with microcephaly, pontine, and cerebellar hypoplasia, and typically affect females [53]. The previously described genotype-phenotypic correlation of neurological diseases associated with variants in the WWOX gene was associated with spinocerebellar ataxia type 12, early infantile epileptic encephalopathy, and autism spectrum disorder [54, 55]. In current study, WWOX-related epilepsy was characterized by psychomotor retardation, spastic ataxia, and encephalopathy of the cerebral hemispheres on the MRI picture. Individuals affected by CACNA1B variants presented with epileptic encephalopathy, severe neurodevelopmental delay, hyperkinetic movement disorder (myoclonus-dystonia syndrome), postnatal microcephaly, and hypotony [56, 57]. However, case 6 with compound heterozygous variants in CACNA1B had generalized tonic seizures, psychomotor, and speech delay.
One of the important results of identifying the etiology of epilepsy is the application of precision medicine based on genetic testing results. Among the identified 14 cases of genetic epilepsy, targeted therapy was indicated in 8 (57%) cases [22, 58]. The results obtained confirm the importance of using genetic diagnostic methods, especially NGS, for the management of patients with epilepsy.
Undoubtedly, the study was not population-based, had a limited number of participants (n=20), and had strict inclusion criteria. In this regard, the obtained high percentage of detection of genetic etiology does not reflect the genetic structure of epilepsy in Kazakhstan. However, the inclusion criteria adapted during the study makes it possible to determine a cost-effective algorithm for diagnosing epilepsy.
Conclusion
The data obtained indicate the high significance of NGS in the diagnosis and management of patients with early-onset epilepsy. Moreover, the study complements the existing knowledge about the genotype-phenotype correlation in epilepsy and highlights the need for the introduction and expansion of NGS diagnostics in Kazakhstan.
Supplementary Information
Author Contribution
Conceptualization: MB, NMT, and DSar. Methodology: MB, AN, AB, AA, NMT, and DSar. Data collection: AB, AN, and AM. Formal analysis and investigation: UKa, AB, SR, NS, DSam, and UKo. Writing — original draft preparation: AB. Writing — review and editing: MB, AA, and UK. Supervision: MB, NMT, AA, and DSar.
Funding
This study was funded and carried out as part of a joint pilot project between the “UMC” CF and the “NLA” PI (contract DPMRV-1986/161-20 dated 09.12.2021) and by the Science Committee of the Ministry of Science and Higher Education of the Republic of Kazakhstan (Grant No. BR18574184).
Data Availability
All data available by request to corresponding author. Sequencing data for individual samples have been deposited at National Center for Biotechnology Information Sequence Read Archive under accession number PRJNA954058 (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA954058).
Code Availability
Not applicable.
Declarations
Ethics Approval
The study was approved by the local commission on bioethics of the UMC CF, an extract from Protocol No. 1 dated 29.06.2021.
Consent to Participate
Written informed consent and permission to publish data was obtained from parents for all research subjects.
Consent for Publication
Written informed consent and permission to publish data was obtained from parents for all research subjects.
Conflict of Interest
The authors declare no competing interests.
Footnotes
Highlights
• In this study, for the first time in Kazakhstan, whole genome sequencing was carried out in epilepsy diagnosed children.
• The genetic etiology of childhood epilepsy was in 70% of cases.
• Six new genetic variants associated with the development of epilepsy have been found.
The original online version of this article was revised: The correct surname should be written as "Samatkyzy".
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Change history
10/13/2024
The original online version of this article was revised: The correct surname should be written as "Samatkyzy".
Change history
10/12/2024
A Correction to this paper has been published: 10.1007/s12035-024-04544-3
References
- 1.Beghi E (2020) The epidemiology of epilepsy. Neuroepidemiology 54(Suppl. 2):185–191. 10.1159/000503831 [DOI] [PubMed] [Google Scholar]
- 2.Fisher RS, van Emde Boas W, Blume W, Elger C, Genton P, Lee P, Engel J Jr (2005) Epileptic seizures and epilepsy: definitions proposed by the International League Against Epilepsy (ILAE) and the International Bureau for Epilepsy (IBE). Epilepsia 46(4):470–472. 10.1111/j.0013-9580.2005.66104.x [DOI] [PubMed] [Google Scholar]
- 3.Fiest KM, Sauro KM, Wiebe S, Patten SB, Kwon CS, Dykeman J, Pringsheim T, Lorenzetti DL et al (2017) Prevalence and incidence of epilepsy: a systematic review and meta-analysis of international studies. Neurology 88(3):296–303. 10.1212/WNL.0000000000003509 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Beghi E, Giussani G, Nichols E, Abd-Allah F, Abdela J, Abdelalim A, Abraha HN, Adib MG et al (2019) Global, regional, and national burden of epilepsy, 1990–2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet Neurol. 18(4):357–375. 10.1016/s1474-4422(18)30454-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Record EJ, Bumbut A, Shih S, Merwin S, Kroner B, Gaillard WD (2021) Risk factors, etiologies, and comorbidities in urban pediatric epilepsy. Epilepsy & behav: E&B 115:107716. 10.1016/j.yebeh.2020.107716 [DOI] [PubMed] [Google Scholar]
- 6.Jaxybayeva A, Nauryzbayeva A, Khamzina A, Takhanova M, Abilhadirova A, Rybalko A, Jamanbekova K (2021) Genomic investigation of infantile and childhood epileptic encephalopathies in Kazakhstan: an urgent priority. Front. Neurol. 12:639317. 10.3389/fneur.2021.639317 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Sarsenbayeva U, Tekebayeva L, Baigazieva L, Kazhaparova D, Kenzhegulova R, Jaxybayeva A (2015) P18 – 2422: the incidence of epileptic encephalopathy in children from birth up to 5 years old in Kazakhstan. Eur. J. Paediatr. Neurol. 19:S99. 10.1016/s1090-3798(15)30331-7 [Google Scholar]
- 8.Berg AT, Baca CB, Rychlik K, Vickrey BG, Caplan R, Testa FM, Levy SR (2016) Determinants of social outcomes in adults with childhood-onset epilepsy. Pediatrics 137(4):e20153944. 10.1542/peds.2015-3944 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wakamoto H, Nagao H, Hayashi M, Morimoto T (2000) Long-term medical, educational, and social prognoses of childhood-onset epilepsy: a population-based study in a rural district of Japan. Brain Dev. 22(4):246–255. 10.1016/s0387-7604(00)00121-2 [DOI] [PubMed] [Google Scholar]
- 10.Lepessova M, Myrzaliyeva B (2013) Clinical and epidemiological characteristics of epilepsy in children in Kazakhstan. J. Neurol. Sci. 333:e23–e24. 10.1016/j.jns.2013.07.092 [Google Scholar]
- 11.Annegers JF, Hauser WA, Anderson VE, Kurland LT (1982) The risks of seizure disorders among relatives of patients with childhood onset epilepsy. Neurology 32(2):174–179. 10.1212/wnl.32.2.174 [DOI] [PubMed] [Google Scholar]
- 12.Jallon, P., Loiseau, P., & Loiseau, J. (2001). Newly diagnosed unprovoked epileptic seizures: presentation at diagnosis in CAROLE study. Coordination Active du Réseau Observatoire Longitudinal de l' Epilepsie. Epilepsia, 42(4), 464–475. 10.1046/j.1528-1157.2001.31400.x [DOI] [PubMed]
- 13.Myers CT, Mefford HC (2015) Advancing epilepsy genetics in the genomic era. Genome Med. 7(1):91. 10.1186/s13073-015-0214-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Scheffer IE, Berkovic SF (1997) Generalized epilepsy with febrile seizures plus. A genetic disorder with heterogeneous clinical phenotypes. Brain : A J Neurol 120(Pt 3):479–490. 10.1093/brain/120.3.479 [DOI] [PubMed] [Google Scholar]
- 15.Myers KA, Johnstone DL, Dyment DA (2019) Epilepsy genetics: current knowledge, applications, and future directions. Clin. Genet. 95(1):95–111. 10.1111/cge.13414 [DOI] [PubMed] [Google Scholar]
- 16.Wang J, Lin ZJ, Liu L, Xu HQ, Shi YW, Yi YH, He N, Liao WP (2017) Epilepsy-associated genes. Seizure 44:11–20. 10.1016/j.seizure.2016.11.030 [DOI] [PubMed] [Google Scholar]
- 17.Gogou M, Cross JH (2022) Seizures and epilepsy in childhood. Continuum (Minneapolis, Minn.) 28(2):428–456. 10.1212/CON.0000000000001087 [DOI] [PubMed] [Google Scholar]
- 18.Lesca G, Baumgartner T, Monin P, De Dominicis A, Kunz WS, Specchio N (2022) Genetic causes of rare and common epilepsies: what should the epileptologist know? Eur. J. Med. Genet. 65(9):104570. 10.1016/j.ejmg.2022.104570 [DOI] [PubMed] [Google Scholar]
- 19.Dunn P, Albury CL, Maksemous N, Benton MC, Sutherland HG, Smith RA, Haupt LM, Griffiths LR (2018) Next generation sequencing methods for diagnosis of epilepsy syndromes. Front. Genet. 9:20. 10.3389/fgene.2018.00020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Orsini A, Zara F, Striano P (2018) Recent advances in epilepsy genetics. Neurosci. Lett 667:4–9. 10.1016/j.neulet.2017.05.014 [DOI] [PubMed] [Google Scholar]
- 21.Hamdan FF, Myers CT, Cossette P, Lemay P, Spiegelman D, Laporte AD, Nassif C, Diallo O et al (2017) High rate of recurrent de novo mutations in developmental and epileptic encephalopathies. Am. J. Hum. Genet. 101(5):664–685. 10.1016/j.ajhg.2017.09.008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Striano P, Minassian BA (2020) From genetic testing to precision medicine in epilepsy. Neurotherapeutics: J Am Soc Exp NeuroTher 17(2):609–615. 10.1007/s13311-020-00835-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Edge KA, Darling J (2020) Cylinder pressure transients in oil hydraulic pumps with sliding plate valves. Proc Inst Mech Eng, Part B: Manag Eng Manuf 51(1):45–54. 10.1243/PIME_PROC_1986_200_047_02 [Google Scholar]
- 24.Ostrander B, Butterfield RJ, Pedersen BS, Farrell AJ, Layer RM, Ward A, Miller C, DiSera T et al (2018) Whole-genome analysis for effective clinical diagnosis and gene discovery in early infantile epileptic encephalopathy. NPJ Genom. Med. 3:22. 10.1038/s41525-018-0061-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Martin HC, Kim GE, Pagnamenta AT, Murakami Y, Carvill GL, Meyer E, Copley RR, Rimmer A et al (2014) Clinical whole-genome sequencing in severe early-onset epilepsy reveals new genes and improves molecular diagnosis. Hum. Mol. Genet. 23(12):3200–3211. 10.1093/hmg/ddu030 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Akilzhanova A, Kairov U, Rakhimova S, Molkenov A, Rhie A, Kim JI, Seo JS, Zhumadilov Z (2014) The first Kazakh whole genomes: the first report of NGS data. Cent Asian J Glob health 3(Suppl):146. 10.5195/cajgh.2014.146 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Andrews S (2010) FastQC: a quality control tool for high throughput sequence data. Available online at: http://www.bioinformatics.babraham.ac.uk/projects/fastqc
- 28.Ewels P, Magnusson M, Lundin S, Käller M (2016 Oct 1) MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics. 32(19):3047–3048. 10.1093/bioinformatics/btw354 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Li H, Durbin R (2010) Fast and accurate long-read alignment with Burrows-Wheeler Transform. Bioinformatics 26:589–595. 10.1093/bioinformatics/btp698 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Mckenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A et al (2010) The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20:1297–1303. 10.1101/gr.107524.110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Wang K, Li M, Hakonarson H (2010) ANNOVAR: functional annotation of genetic variants from next-generation sequencing data. Nucleic Acids Res. 38:e164. 10.1093/nar/gkq603 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Stödberg T, Tomson T, Barbaro M, Stranneheim H, Anderlid BM, Carlsson S, Åmark P, Wedell A (2020) Epilepsy syndromes, etiologies, and the use of next-generation sequencing in epilepsy presenting in the first 2 years of life: a population-based study. Epilepsia 61(11):2486–2499. 10.1111/epi.16701 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Rim JH, Kim SH, Hwang IS, Kwon SS, Kim J, Kim HW, Cho MJ, Ko A et al (2018) Efficient strategy for the molecular diagnosis of intractable early-onset epilepsy using targeted gene sequencing. BMC Med. Genomics 11(1):6. 10.1186/s12920-018-0320-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Symonds JD, Zuberi SM, Stewart K, McLellan A, O'Regan M, MacLeod S, Jollands A, Joss S et al (2019) Incidence and phenotypes of childhood-onset genetic epilepsies: a prospective population-based national cohort. Brain J. Neurol. 142(8):2303–2318. 10.1093/brain/awz195 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Miceli, F., Soldovieri, M. V., Weckhuysen, S., Cooper, E., & Taglialatela, M. (2010). KCNQ2-related disorders. In M. P. Adam (Eds.) et. al., GeneReviews®. University of Washington, Seattle.
- 36.Hu Q, Shen Y, Su T, Liu Y, Xu S (2021) Clinical and genetic characteristics of chinese children with GLUT1 deficiency syndrome: case report and literature review. Front. Genet. 12:734481. 10.3389/fgene.2021.734481 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kolic I, Radic Nisevic J, Vlasic Cicvaric I, Butorac Ahel I, Lah Tomulic K, Segulja S, Baraba Dekanic K, Serifi S et al (2021) GLUT1 deficiency syndrome-early treatment maintains cognitive development? (Literature review and case report). Genes 12(9):1379. 10.3390/genes12091379 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Olivetti PR, Noebels JL (2012) Interneuron, interrupted: molecular pathogenesis of ARX mutations and X-linked infantile spasms. Curr. Opin. Neurobiol. 22(5):859–865. 10.1016/j.conb.2012.04.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Poeta L, Malacarne M, Padula A, Drongitis D, Verrillo L, Lioi MB, Chiariello AM, Bianco S et al (2022) Further delineation of duplications of ARX locus detected in male patients with varying degrees of intellectual disability. Int. J. Mol. Sci. 23(6):3084. 10.3390/ijms23063084 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Sharma S, Prasad AN (2013) Genetic testing of epileptic encephalopathies of infancy: an approach. Can J Neurol Sci / J Canadien Des Sciences Neurologiques 40(1):10–16. 10.1017/s0317167100012889 [DOI] [PubMed] [Google Scholar]
- 41.Velíšek L, Velíšková J (2020) Modeling epileptic spasms during infancy: are we heading for the treatment yet? Pharmacol. Ther. 212:107578. 10.1016/j.pharmthera.2020.107578 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Hanna MG, Nelson IP, Rahman S, Lane RJM, Land J, Heales S, Cooper MJ, Schapira AHV et al (1998) Cytochrome c oxidase deficiency associated with the first stop-codon point mutation in human mtDNA. Am. J. Hum. Genet. 63(1):29–36. 10.1086/301910 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Horváth R, Scharfe C, Hoeltzenbein M et al (2002) Childhood onset mitochondrial myopathy and lactic acidosis caused by a stop mutation in the mitochondrial cytochrome c oxidase III gene [2]. J Med Genet. 10.1136/jmg.39.11.812 [DOI] [PMC free article] [PubMed]
- 44.Marotta R, Chin J, Kirby DM, Chiotis M, Cook M, Collins SJ (2011) Novel single base pair COX III subunit deletion of mitochondrial DNA associated with rhabdomyolysis. J. Clin. Neurosci. 18(2):290–292. 10.1016/j.jocn.2010.06.001 [DOI] [PubMed] [Google Scholar]
- 45.Tiranti V, Corona P, Greco M, Taanman JW, Carrara F, Lamantea E, Nijtmans L, Uziel G et al (2000) A novel frameshift mutation of the mtDNA COIII gene leads to impaired assembly of cytochrome c oxidase in a patient affected by Leigh-like syndrome. Hum. Mol. Genet. 9(18):2733–2742. 10.1093/hmg/9.18.2733 [DOI] [PubMed] [Google Scholar]
- 46.Wang W, Sun Y, Lin Y, Xu X, Zhao D, Ji K, Li W, Zhao Y et al (2021) A novel nonsense variant in MT-CO3 causes MELAS syndrome. Neuromuscul. Disord. 31(6):558–565. 10.1016/j.nmd.2021.02.020 [DOI] [PubMed] [Google Scholar]
- 47.Dell'Isola GB, Mencaroni E, Fattorusso A, Tascini G, Prontera P, Imperatore V, Di Cara G, Striano P, Verrotti A (2022) Expanding the genetic and clinical characteristics of Protocadherin 19 gene mutations. BMC Medical Genomics 15(1):181. 10.1186/s12920-022-01313-w [DOI] [PMC free article] [PubMed]
- 48.Platzer K, Krey I, Lemke JR (2022) GRIN2D-related developmental and epileptic encephalopathy. In: Adam MP, Mirzaa GM, Pagon RA et al (eds) GeneReviews® [Internet]. University of Washington, Seattle, WA, pp 1993–2023. Available from: https://www.ncbi.nlm.nih.gov/books/NBK582335/
- 49.Lin Z, Sang T, Yang Y, Wu Y, Dong Y, Ji T, Zhang Y, Wu Y et al (2022) Efficacy of anti-seizure medications, quinidine, and ketogenic diet therapy for KCNT1-related epilepsy and genotype-efficacy correlation analysis. Front. Neurol. 12:834971. 10.3389/fneur.2021.834971 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Villa C, Colombo G, Meneghini S, Gotti C, Moretti M, Ferini-Strambi L, Chisci E, Giovannoni R et al (2019) CHRNA2 and nocturnal frontal lobe epilepsy: identification and characterization of a novel loss of function mutation. Front. Mol. Neurosci. 12:17. 10.3389/fnmol.2019.00017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Ceska K, Danhofer P, Horak O, Spanelova K, Kolar S, Oslejskova H, Aulicka S (2022) Phenotypic spectrum of the SCN1A mutation (from febrile seizures to Dravet syndrome). Bratisl. Lek. Listy 123(7):483–486. 10.4149/BLL_2022_076 [DOI] [PubMed] [Google Scholar]
- 52.McTague A, Kurian MA (2019) SLC12A5-related epilepsy of infancy with migrating focal seizures. In: Adam MP, Mirzaa GM, Pagon RA et al (eds) GeneReviews® [Internet]. University of Washington, Seattle, WA, pp 1993–2023. Available from: https://www.ncbi.nlm.nih.gov/books/NBK537476/
- 53.Dubbs H, Ortiz-Gonzalez X, Marsh ED (2022) Pathogenic variants in CASK: expanding the genotype-phenotype correlations. Am. J. Med. Genet. A 188(9):2617–2626. 10.1002/ajmg.a.62863 [DOI] [PubMed] [Google Scholar]
- 54.Aldaz CM, Hussain T (2020) WWOX loss of function in neurodevelopmental and neurodegenerative disorders. Int. J. Mol. Sci. 21(23):8922. 10.3390/ijms21238922 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Piard J, Hawkes L, Milh M, Villard L, Borgatti R, Romaniello R, Fradin M, Capri Y et al (2019) The phenotypic spectrum of WWOX-related disorders: 20 additional cases of WOREE syndrome and review of the literature. Genet Med: Off Journal Am Coll Med Gene 21(6):1308–1318. 10.1038/s41436-018-0339-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Gorman KM, Meyer E, Grozeva D, Spinelli E, McTague A, Sanchis-Juan A, Carss KJ, Bryant E et al (2019) Bi-allelic loss-of-function CACNA1B mutations in progressive epilepsy-dyskinesia. Am. J. Hum. Genet. 104(5):948–956. 10.1016/j.ajhg.2019.03.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Groen JL, Andrade A, Ritz K, Jalalzadeh H, Haagmans M, Bradley TE, Jongejan A, Verbeek DS et al (2015) CACNA1B mutation is linked to unique myoclonus-dystonia syndrome. Hum. Mol. Genet. 24(4):987–993. 10.1093/hmg/ddu513 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Zimmern V, Minassian B, Korff C (2022) A review of targeted therapies for monogenic epilepsy syndromes. Frontiers in neurology 13:829116. 10.3389/fneur.2022.829116 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All data available by request to corresponding author. Sequencing data for individual samples have been deposited at National Center for Biotechnology Information Sequence Read Archive under accession number PRJNA954058 (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA954058).
Not applicable.