Abstract
Cryptococcus is a fungal pathogen whose virulence relies on proliferation in and dissemination to host sites, and on synthesis of a defensive yet metabolically costly polysaccharide capsule. Regulatory pathways required for Cryptococcus virulence include a GATA-like transcription factor, Gat201, that regulates Cryptococcal virulence in both capsule-dependent and capsule-independent ways. Here we show that Gat201 is part of a negative regulatory pathway that limits fungal survival at alkaline pH. RNA-seq analysis found strong induction of GAT201 expression within minutes of transfer to RPMI media at alkaline pH. Microscopy, growth curves, and colony forming unit assays show that in RPMI at alkaline pH wild-type Cryptococcus neoformans yeast cells produce capsule but do not bud or maintain viability, while gat201Δ cells make buds and maintain viability, yet fail to produce capsule. GAT201 is required for transcriptional upregulation of a specific set of genes, the majority of which are direct Gat201 targets. Evolutionary analysis shows that Gat201 is in a subfamily of GATA-like transcription factors that is conserved within pathogenic fungi but absent in model yeasts. This work identifies the Gat201 pathway as controlling a trade-off between proliferation and production of defensive capsule. The assays established here will allow characterisation of the mechanisms of action of the Gat201 pathway. Together, our findings urge improved understanding of the regulation of proliferation as a driver of fungal pathogenesis.
Author Summary
Micro-organisms face trade-offs in adapting to their environments. For example, pathogens adapting to host niches must balance investing in proliferation – reproduction and growth – against investing in defense against the host immune system. Cryptococcus neoformans is an encapsulated fungal pathogen that can infect human airways and, in immunocompromised people, can move to the brain to cause life-threatening meningitis. It is well appreciated that fungal persistence in these sites depends on production of a sugar capsule that surrounds the cell, hiding it from host detection. However, in both the lung and brain, fungal proliferation through budding is also a major driver of pathogenesis: both cryptococcal pneumonia and meningitis are characterised by high yeast burden. This presents a trade-off between production of a metabolically costly capsule and cellular proliferation. The regulators of Cryptococcus proliferation are poorly understood, as they are distinct from other model yeasts at the level of cell cycle and morphogenesis. In this work, we study this trade off growing Cryptococcus under conditions that approximate the alkaline surface of human airways, and that restrict fungal growth. We identify a GATA-like transcription factor, Gat201, and its target, Gat204, that positively regulate capsule production and negatively regulate proliferation. The GAT201 pathway is conserved within pathogenic fungi but lost in other model yeasts. Together our findings reveal how a fungal pathogen regulates the balance between defense and proliferation and highlight the need for improved understanding of proliferation in non-model systems.
Introduction
Cryptococcus neoformans is an environmental saprophyte and a critical priority global human pathogen (World Health Organization 2022) that is a leading cause of death in HIV-positive individuals (Rajasingham et al. 2022; Park et al. 2009). C. neoformans is a basidiomycete fungus that is associated worldwide with bird guano and arboreal habitats (Nielsen, De Obaldia, and Heitman 2007; Springer, Mohan, and Heitman 2017; Litvintseva et al. 2011). C. neoformans can become an opportunistic pathogen following inhalation of airborne spores or desiccated yeast cells into mammalian airways (Walsh et al. 2019; Velagapudi et al. 2009; Powell et al. 1972; Giles et al. 2009). These infectious particles are initially metabolically inactive and must reactivate proliferation and simultaneously escape the immune system to infect host airways (Velagapudi et al. 2009; Ballou and Johnston 2017; Botts and Hull 2010; May et al. 2016). Proliferation of Cryptococcus in the upper airways or lung can be followed by dissemination through other host niches including to the central nervous system, causing fatal meningitis. Thus, both adaptation to diverse environmental niches and immune evasion are crucial for Cryptococcal virulence (Bose et al. 2003; Williamson 1997; A. Casadevall, Rosas, and Nosanchuk 2000; John R. Perfect 2006; Arturo Casadevall, Steenbergen, and Nosanchuk 2003).
However, proliferation and immune evasion can place competing metabolic demands on the fungal cell (J. Kronstad et al. 2012). For example, in the host, Cryptococcus cells produce a protective polysaccharide capsule. This unique and metabolically costly host evasion mechanism is dispensable for fungal proliferation and morphogenesis (García-Rodas et al. 2014), but is essential to Cryptococcus survival and dissemination in the host (O’Meara and Alspaugh 2012; Bose et al. 2003). By investigating conditions in which C. neoformans can either proliferate or protect itself, we can better understand the regulatory pathways controlling these competing demands.
We set out to probe this competition between proliferation and evasion during the initial switch to host-like conditions. Here, we reasoned that an informative model of initial infection would involve starting with stationary phase yeast cells that were then cultivated in “host-like” growth conditions, where different host-relevant conditions might give different insights into Cryptococcus adaptation. As the human airway surface becomes alkaline during each inhaled breath (Dusik Kim et al. 2021), we therefore investigated which genes are induced in C. neoformans stationary phase yeast cells directly after inoculation into in vitro conditions including host-like RPMI-1640 media at alkaline pH (RPMI). Our RNA-seq revealed strong induction of the GAT201 pathway in these conditions.
The GATA-like zinc finger transcription factor Gat201 is a key regulator of virulence that acts in capsule-dependent (Liu et al. 2008) and capsule-independent ways (Chun, Brown, and Madhani 2011). C. neoformans strains with GAT201 deleted have lower virulence in mouse models of infection (Liu et al. 2008) and reduced capsule (Jung et al. 2015; Gish et al. 2016; Jang et al. 2022). GAT201 deletion strains are also more readily taken up by mammalian macrophages independently of capsule production (Chun, Brown, and Madhani 2011). The genes targeted by Gat201p have been mapped by RNA-seq and ChIP-seq (Homer et al. 2016; Chun, Brown, and Madhani 2011), yet the nature of its capsule-independent contributions to virulence remain unexplained. One challenge is that gat201Δ exhibits relatively weak phenotypes in standard microbiological growth conditions, making it difficult to probe Gat201-regulated virulence (Jung et al. 2015).
We observed that WT cells inoculated into RPMI media at alkaline pH proliferated poorly, but that deletion of GAT201 dramatically improved proliferation and long-term viability. This suggests that poor growth in RPMI conditions is a consequence of regulated gene expression, controlled by the transcription factor Gat201, rather than a physiological response to nutrient starvation. Through RNA-seq, we identify GAT201-dependent transcriptional signatures of this phenotype and demonstrate that it is activated to suppress proliferation under alkaline conditions, independently of serum and of cyclic AMP. Correlating transcription factor activation to different microbial phenotypes can reveal signaling pathways required for adaptation to specific environments. Our findings of novel microbiological growth phenotypes dependent on GAT201 will enable future mechanistic studies of this virulence pathway.
Results
Cryptococcus neoformans rapidly induces media-specific growth programs upon reactivation from stationary phase.
We set out to determine which gene expression pathways are induced in C. neoformans stationary yeast cells when they reactivate in nutrient rich growth medium (YPD) versus host-like medium (RPMI-1640 + 10% Heat inactivated fetal calf serum; abbreviated RPMI) at 2 different temperatures (25°C or 37°C), and grown at 60 rpm shaking to mimic low-oxygen conditions. To investigate the early events of cellular reactivation, we used stationary phase cells incubated for 5 days in YPD at 30°C (Fig 1A). We sampled 2 biological replicates of cells prior to inoculation (0 minutes), and then sampled the cultures at 10, 30, 60, and 120 minutes following inoculation into either YPD or RPMI media at the two temperatures. We also collected a standard growth condition, mid-exponential phase cells reinoculated 1:30 into fresh YPD at 30°C for 180 min. Cells were examined for morphology (Fig 1B) and for transcriptional changes via RNA-seq (Fig 1C,D,E)
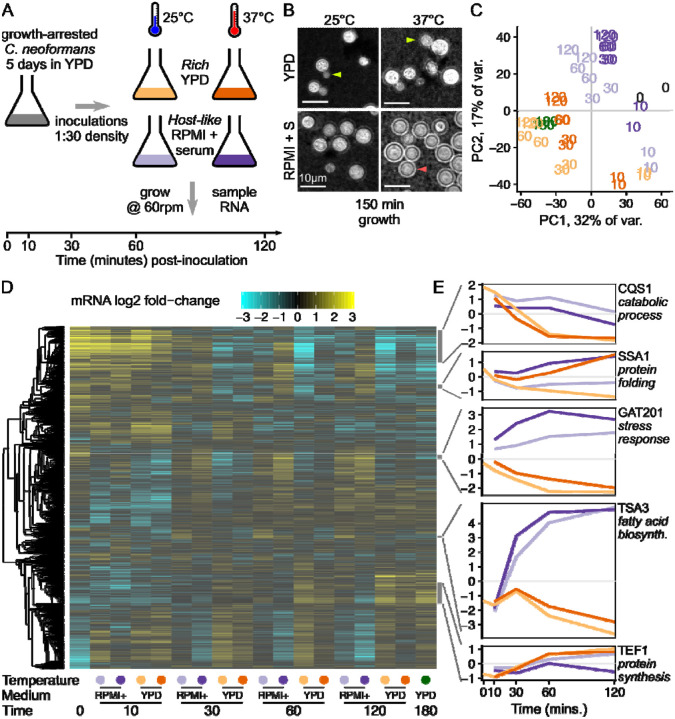
Fig 1: Cryptococcus neoformans rapidly induces media-specific growth programs upon reactivation from stationary phase.
A, Design of time-course experiment to measure the contribution of media and temperature on reactivation of stationary phase cells. B, Budding (yellow-green arrow) is seen in YPD at both temperatures (top row) while capsule (light red arrow) is primarily seen in RPMI + serum at 37°C (bottom right). Micrograph shows India Ink staining of cells taken from one replicate of cells used for the RNA-seq experiment, 150 minutes after inoculation. C, Time from inoculation and media dominate the overall variance in gene expression, shown by principal component analysis on the regularized log-counts per gene in every replicate. Each replicate is plotted and labeled by the timepoint, with colours as in Fig. 1A (grey - 0 min, yellow - YPD 25°C, orange - YPD 37°C, light purple - RPMI 25°C, dark purple - RPMI 37°C), additionally with dark green for exponential phase rich media 30°C. D, Distinct clusters of co-regulated genes respond to reactivation in different media, shown by clustered heatmap of log2 fold-change per gene calculated by DESeq2. Each row represents an individual gene and rows are clustered by co-expression patterns (see methods), while each column represents the regularised log2 fold-change estimate across both replicates in a single condition. E, Representative genes from different clusters show distinct expression patterns, again in regularised log2 fold-change per gene. Colours of growth conditions are as in Fig. 1A.
We observed that cells reactivating in rich medium resumed growth and cell division, with visible buds 150 minutes after inoculation (Fig 1B). However, cells reactivating in RPMI medium showed no visible budding, and when grown at 37°C also produced a polysaccharide capsule visible with India Ink staining (Fig 1B). The RPMI media here is buffered with sodium bicarbonate, grown in a standard aerobic shaking incubator at 60 rpm. We observed the pH of the media rise to become alkaline over the timecourse, resembling human airway surface liquid that is alkaline during inhalation dependent on its bicarbonate content (Dusik Kim et al. 2021). We return to discuss the role of alkaline pH on the phenotypes later.
Consistent with observed changes in cell morphology, we identified distinct gene expression programs in the different media and temperatures. Principal component analysis (PCA) on the regularized log-counts per gene shows clear separation between growth conditions (Fig 1C and Fig S1). Principal components (PCs) 1 and 2 separate stationary phase cells and early timepoints (0 min, 10 mins) from later timepoints. PCs 1 and 2 also separate YPD media and RPMI media at later timepoints, while the exponential phase samples grown in rich media at 30°C group with the other rich media samples. PCA groups both biological replicates together for all sample conditions, indicating a highly reproducible experiment. PC3 separates samples grown at different temperatures with a clear divide between 25°C and 37°C (Fig S1). PC4 largely separates the 0 min samples from all later timepoints (Fig S1), emphasizing that induction of gene expression pathways is detectable after only 10 minutes in either growth medium and independent of temperature.
Clustering genes by their expression patterns reveals the combinatorial impact of media and temperature over time (Fig 1D). One cluster of 484 genes are highly expressed in stationary phase and decline in rich media, marked by CQS1 that encodes Cryptococcus quorum sensing protein Qsp1 (Fig 1E, Table S1). CQS1 is the most abundant transcript in cells in stationary phase conditions and RPMI media, as previously noted (Homer et al. 2016). This cluster of stationary-phase upregulated genes are also enriched in catabolic process functions (GO:0006091 generation of precursor metabolites and energy; GO:0005975 carbohydrate metabolic process), indicating that cells are starving.
A second cluster of 60 co-regulated genes are upregulated at 37°C in both media, marked by SSA1, encoding a heat shock protein (Fig 1E, Table S1). This cluster of genes is enriched in protein folding chaperones (GO:0006457, protein folding), including HSP40, HSP60, HSP70, HSP90, and HSP104 family members, as expected from the conserved heat shock response that has been previously observed in Cryptococcus (Steen et al. 2002). A relatively small number of genes - 60 in this cluster - were unambiguously induced by increased temperature in both media (Table S1; and DGE analysis in Table S2).
A third cluster of 57 co-regulated genes is upregulated in RPMI compared to YPD, marked by the virulence-associated transcription factor GAT201 (Fig 1E, Table S1). This cluster of genes is also enriched in associations with the stress response (GO:0006950, response to stress). These include cell wall-related enzymes (chitin synthase CHS4, chitinase CHI2, glucan glucosidase EXG2, Endoglucanase LPI9), as well as genes associated with redox metabolism (catalase CAT2, glutathione transferase CNAG_03848). Differential expression analysis of media conditions (Table S2) shows that GAT201 is one of the most differentially expressed genes after 1 hour at both 37°C (~24-fold, p < 10 −19) and 25°C (~13-fold, p < 10−20). Gat201 acts through other key transcription factors Gat204 and Liv3 (Homer et al. 2016). Consistent with GAT201 induction 10 minutes after inoculation, we later observed induction of GAT204 and LIV3 expression in RPMI at 30 minutes and onwards (Fig S2).
A fourth cluster of 25 co-regulated genes is even more induced in RPMI, marked by TSA3, encoding a thiol-specific antioxidant protein, which is over 80-fold induced (Fig 1E). TSA3 has been reported to be strikingly induced by temperature and hydrogen peroxide when grown in YNB media (Missall, Pusateri, and Lodge 2004). This cluster is enriched in genes involved in fatty acid biosynthesis (GO:0006629, lipid metabolic process).
Lastly, a large cluster of 321 genes associated with growth is induced in YPD at both temperatures and in RPMI at 25°C only, marked by translation elongation factor TEF1 (Fig 1E). This cluster of genes is enriched for genes involved in protein synthesis, including ribosomal proteins (GO:0005840, ribosome), translation elongation factors, and amino acid production (GO:0006520, cellular amino acid metabolic process). The cluster is also enriched in genes involved in DNA segregation (GO:0000278, mitotic cell cycle) and mitochondrial biogenesis (GO:0005739, mitochondrion).
Overall, our data show that reactivating cells rapidly activate transcription and biosynthetic pathways, in distinct ways in different conditions. Cells induce both protein synthesis and cell cycle progression in YPD media, and proliferate by producing buds within 3 hours of inoculation. A different set of pathways are activated in RPMI-1640 + serum media, prominently including the GAT201 transcription factor, genes associated with oxidative stress responses, and many genes of unknown function. In RPMI at 37°C, cells make a polysaccharide capsule that is associated with cellular defense, but do not start budding. This suggests that cells are making a condition-dependent decision between proliferation and defense, and that this may be achieved via differential expression of transcription factors.
GAT201 determines cellular phenotypes during reactivation
We next tested the effect of GAT201 on reactivation phenotypes, reasoning that the strong induction of GAT201 and its transcriptional targets in RPMI made this pathway a good candidate regulator. We used two independently generated mutants: gat201Δm is a complete deletion of the reading frame from start codon to stop codon from the Madhani collection (Liu et al. 2008) and gat201Δb is a disruption of the protein from the Bahn collection (Jung et al. 2015). We verified the mutations to the GAT201 locus by PCR. We also made 2 independent complemented strains GAT201-C1 and GAT201-C2 by expressing GAT201 from its native promoter and terminator at a genomic safe haven locus in the gat201Δm background (Figs 2A and 2E). We initially characterize these strains in RPMI-1640 medium without serum, and later return to examining the effects of serum.
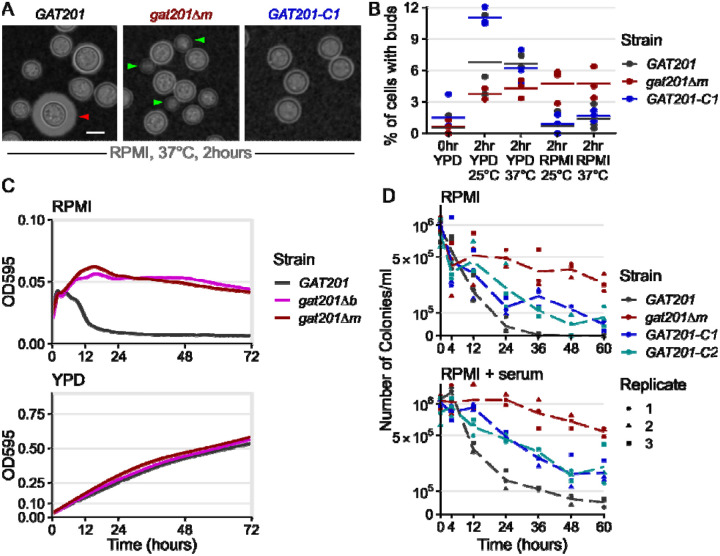
Fig 2: GAT201 represses the proliferation and viability of Cryptococcus neoformans during reactivation in RPMI medium.
A, GAT201 promotes capsule biosynthesis and represses budding in RPMI-1640 medium (without serum) at 37°C 2 hours after inoculation. Micrographs show GAT201 (H99), gat201Δm, and complemented GAT201-C1 strains, stained with India Ink, capsule highlighted with red arrow and buds highlighted with green arrows. GAT201-C1 complements the budding phenotype but does not clearly complement the capsule phenotype. B, Quantification of budding index at 2 hrs (% budded cells) shows that gat201Δm cells reactivate to produce buds in RPMI, (n= >100 cells per replicate, with 3 biological replicates per condition). Figs 2A and 2B are taken from the same experiment, and larger sets of representative cells are shown in Fig S3. C, GAT201 (H99) cell populations reactivating in RPMI show a fall in density after 10 hours growth, which is absent in gat201Δ strains and absent during growth in rich YPD media. Growth curves of optical density at 595 nm (OD595) were collected via plate reader from 7 biological replicates, 3 technical replicates each, at 37°C. Note the different y-axis limits in the subpanels, reflecting higher final OD in rich media. D. GAT201 (H99) cells reactivating in RPMI or RPMI + serum show a decline in viability after 12–24 hours, which is absent in gat201Δ and partially restored by complementing GAT201. The decline in viability is more severe in RPMI without serum than it is in RPMI with serum. Colony forming units per ml of culture were measured by serial dilution on plates, in 3 biological replicates; individual replicates are plotted as dots with a dotted line connecting the medians.
We observed that GAT201 represses proliferation during reactivation in RPMI medium. 2 hours after inoculation in RPMI at 37°C, wild-type GAT201 strains produce capsule and have few visible buds, while gat201Δ strains have visible buds and small capsule (Fig 2A, Fig S3). There is no difference in phenotype during growth in YPD (Fig S3). Genetic complementation of GAT201 again represses bud formation, but only partially restores capsule. GAT201 deletion mutants have previously been shown to be defective in capsule (Jang et al. 2022; Gish et al. 2016; Liu et al. 2008). The partial complementation of capsule in our strains with GAT201 integrated at a genomic safe haven locus is likely to be due to the reduced expression of GAT201 mRNA in this strain, measured by RT-qPCR as roughly ~10x lower than in wild-type (Fig S4). This RT-qPCR also confirms the GAT201-dependent expression of GAT204 and LIV3, which is partially restored in partially complemented strains and thus dependent on the abundance of GAT201 (Fig S4).
Given the observed transcriptional changes within 10 minutes of reactivation, we quantified the budding index in the same samples at 2 hours post reactivation (Fig 2B), which is long enough for C. neoformans yeast to complete a single cell cycle (Yamaguchi et al. 2007). This early time point was chosen because it allowed detection of reactivation from growth without the possibility of measuring repeated budding or proliferating daughter cells. This confirmed that, upon reactivation from stationary phase, 6–10% of wild-type GAT201 cells produce visible buds in rich media, but only 1–2% in RPMI media. Deletion of GAT201 may reduce budding during reactivation in rich media, but increases budding in RPMI, with roughly 5% of gat201Δ cells budding within 2 hours, a more than 2-fold increase. Genetic complementation of GAT201 reduces bud formation to near-wild-type levels.
Given this GAT201-dependent distinction in budding during reactivation, we investigated longerterm growth. While wild-type cells in RPMI medium did initially increase in optical density for 4 hours, density then rapidly declined within 10 hours (Fig 2C). In contrast, gat201Δ cultures for both mutants continually increased in cell density over 12 hours and maintained a higher OD595 of 0.05 over 3 days (Fig 2C). This effect is media-specific: GAT201 cells and gat201Δ mutants grew similarly in rich media, reaching a similar OD595 of around 0.5 after 3 days (Fig 2C).
To determine if the GAT201-dependent reduced cell density in RPMI represented a loss of viability, we quantified colony-forming units (CFUs) from cells grown in RPMI at 37°C (Fig 2D). All cultures started with approximately the same number of stationary cells per ml (1×106, OD595 = 0.1). By 24 hours the number of viable cells/ml in the gat201Δ mutant reduced by half, to approximately 5×105, compared to a 10-fold reduction to 4×104 in the wild-type, an order of magnitude less and a 25th of the starting cell number. By 36 hours viability in the gat201Δ mutant strain had decreased by slightly more than a third of the starting cell number (3.7×105 cells/ml) while GAT201 wildtype viability was 2 orders of magnitude less than the starting cell number (1×104 cells/ml). GAT201 viability dropped to zero by 48 hours, indicating these cells are inviable following growth in RPMI media at 37°C after 2 days. In contrast, the gat201Δ mutant strain remained viable for up to 60 hours post-inoculation, with just under a quarter of the starting cell number (2.7×105 cells/ml) still viable. Genetic complementation of GAT201 reduced viability, although not to wild-type levels.
We next asked if the addition of serum (a host-like component) would alter the GAT201-dependent loss of viability, we repeated the CFU assay in RPMI + 10% serum at 37°C (Fig 2D). A similar pattern to RPMI was observed in RPMI + serum, however the number of colonies/ml overall were greater in RPMI + serum in both strains. The order of magnitude difference was maintained between the wild-type and gat201Δ mutant strains, with 1/20th and half (5×104 and 5.4×105 cells/ml) of the starting cell number, respectively, at 60 hours post inoculation into fresh media. This indicates that the addition of serum to the media confers an overall benefit to cell viability in these conditions (Fig 2D), but that the effect is independent of GAT201 activity. Genetic complementation of GAT201 again partially restored the mutant phenotype.
Together, our data demonstrate that during reactivation GAT201 determines cellular phenotypes beyond capsule production. When grown in RPMI medium, GAT201 cells produce few buds within the first 2 hours of reactivation, lose optical density by 10 hours, and lose viability after 12–24 hours. The striking order-of-magnitude differences in viability, and qualitative differences in optical density, at longer timepoints, are prefigured by over 2-fold differences in budding index at earlier timepoints. Again, these experiments were conducted in an RPMI formulation buffered with sodium bicarbonate, grown under aerobic conditions so that the pH rose to become alkaline over the course of the experiment. We return to test the role of pH later, but first address the role of serum and GAT201 on gene expression.
How does serum affect the phenotype and gene expression?
Given the differential viability of wild-type and gat201Δ cells in RPMI media both with and without serum, we investigated the transcriptional pathways that might be responsible for these phenotypes. We designed an RNA-seq experiment with 2 deletion strains (gat201Δm and gat201Δb) matched with two congenic wild-type GAT201 strains (KN99 MATa and MATalpha) (Nielsen et al. 2003), each measured in 2 biological replicates. Thus, there are effectively 4 replicates per relevant genotype. We first checked the effects of serum on capsule and budding in GAT201 (KN99alpha) and gat201Δm. Microscopy confirmed that the major phenotypes are serum-independent: GAT201 cells produce capsule and few buds in both RPMI and RPMI + serum, while gat201Δm produce less capsule and more buds in both conditions (Fig 3A, Fig S5).
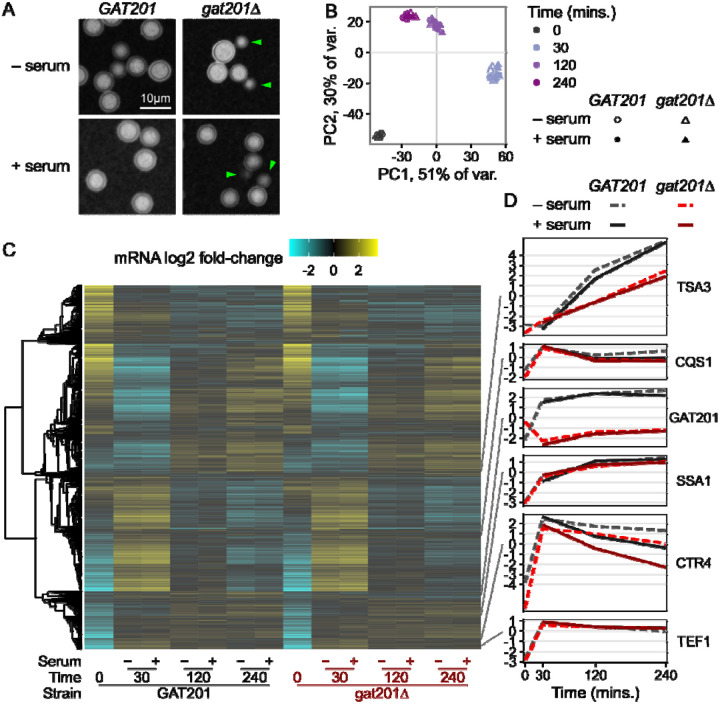
Fig 3: Serum is not the dominant driver of GAT201-dependent phenotypes in RPMI 1640 media.
A, GAT201 promotes capsule biosynthesis and represses budding in RPMI medium both with and without serum at 37°C, 2 hours after inoculation. Strains are GAT201 (KN99alpha) and gat201Δm. B, Time from inoculation dominates the overall variance in gene expression regardless of serum addition or GAT201 allele, shown by principal component analysis on the regularized log-counts per gene in every replicate. C, Only a small set of genes are differentially regulated by serum or by GAT201, shown by clustered heatmap of log2 fold-change per gene calculated by DESeq2. D, Representative genes show distinct expression patterns, again in log2 fold-change per gene.
We carried out RNA-seq of GAT201 and gat201Δ strains reactivating, from stationary phase, in RPMI and RPMI + serum at 30 min, 2 hrs, and 4 hrs. Principal Component Analysis on the regularized log-counts per gene revealed that time after inoculation is the dominant driver of variance in this dataset (Fig 3B). 81% of variance is attributed to the first two principal components, in which datasets cluster together at each timepoint from all strains grown with and without serum. Principal component 3 (4% of variance) distinguishes between GAT201 and gat201Δ strains (Fig S6), indicating that the RNA-seq detected a specific GAT201-dependent transcriptional program that we return to later. We were surprised not to detect a stronger serum-dependent effect, given previous work on the effect of serum on growth and capsule production in Cryptococcus, although this may reflect the short 4 hour time course (Zaragoza, Fries, and Casadevall 2003).
Consistent with serum not being the major driver of phenotype, we found few differences in the overall transcriptome-wide response to serum (Fig 3C, Table S3). Transcriptome-wide expression evolves over time with a large cluster of genes that are strongly upregulated at 30 minutes and still up at 2 and 4 hours. Conversely, a large cluster that is strongly downregulated at 30 minutes becomes less repressed at later timepoints. Expression of a set of representative genes selected from Fig 1 shows similar patterns to the previous dataset (Fig 3D). TSA3 is strongly induced over time in RPMI and RPMI + serum, dependent on GAT201 status. CQS1 is highly expressed at 0 minutes and further induced in RPMI and RPMI + serum. GAT201 itself is induced in RPMI and RPMI + serum in GAT201 cells. The RNA-seq protocol used here is a 3’-end targeted assay that detects a transcript fragment of GAT201 RNA in the two gat201Δ strains that have a truncated or deleted coding sequence; these transcript fragments do not encode a functional Gat201p. Both SSA1 and TEF1 are induced in RPMI and RPMI + serum as cells reactivate protein synthesis.
A small set of genes in wild-type cells differentially respond to the presence of serum at all time points. For example, copper transporter CTR4 (Fig 3D) and copper starvation-induced membrane protein BIM1 (Fig S7) are induced by 30 minutes growth in all media, and lower induction in media with added serum suggests differences in copper ion availability. Differential gene expression analysis with DESeq2 detected under 100 each of serum-upregulated and serum-downregulated genes at 5% FDR with at least 2-fold change, in GAT201 strains (Fig S9, Table S3). Loss of GAT201 dampened this response to serum, with about 4-fold more DEGs detected in GAT201 than in gat201Δ (Fig S8). This dampened response in gat201Δ strains corroborates previous studies that found an association between expression of GAT201 and downstream targets in alternative DMEM media with serum (Chun, Brown, and Madhani 2011).
Comparing our two RNA-seq datasets presented here indicates that the signature of GAT201 pathway activation is consistent. Direct comparison of differential gene expression between the matched conditions of 0 minutes (pre-inoculation) and 2 hours RPMI + serum in wild-type cells shows moderate correlation, R = 0.27 (Fig S9, Table S2, Table S3). The major trends that are the focus of this study are consistent: GAT201 and its targets are induced in RPMI + serum, as are many genes associated with protein synthesis. Despite this, there are some differences that may reflect local experimental conditions. For example, we observe that copper-dependent genes CTR4 and BIM1 are induced in RPMI in dataset 2 and not in dataset 1. This may reflect that the experiments were performed years apart by different experimentalists on different campuses. The datasets were also collected using different RNA-seq approaches. For dataset 1, we used RNATagSeq and rRNA depletion for full-length mRNA sequencing. For dataset 2, we used QuantSeq FWD for 3’-targeted mRNA sequencing.
Most GAT201-dependent DEGs are direct targets of GAT201.
A small and specific set of genes were found to be dependent on the expression of GAT201. After 5 days of growth in YPD (0 minute timepoint), there were only 82 2-fold differentially expressed genes at 5% FDR, up in wild-type GAT201 strain compared to mutant gat201Δ. GO analysis suggested enrichment for upregulated genes involved in microtubule-based movement (GO:0007018), and lipid metabolism (GO: 0006629). 35 genes were down in GAT201 compared to gat201Δ, with GO analysis highlighting probable changes in the cell wall (GO: 0016798, hydrolase activity acting on glycosyl bonds; GO:0005975, carbohydrate metabolic process) including the exoglucanase EXG104. After 30 minutes of incubation in RPMI or RPMI+serum, the GAT201-dependent differentially expressed genes are largely distinct from those observed in stationary phase. For example, EXG104 is upregulated in GAT201 after reactivation. The number of 2-fold differentially expressed genes at 5% FDR between GAT201 and gat201Δ increases from 30 minutes up to 240 minutes (Fig 4A). We observed over 200 differential-expressed genes in each direction in RPMI (without serum) at 240 minutes (Fig 4B). Also, the magnitude of each gene’s differential expression between strains tends to increase over time (Fig 4C). Below, we focus on analysis of the RPMI 240 minutes timepoint.
Fig 4. GAT201 specifically affects gene expression as cells reactivate, acting via its direct targets.
A. There is more GAT201-dependent differential gene expression at later time points in activation. Fig 4A shows the number of 2-fold differentially expressed (DE) genes at 5% FDR at each combination of growth condition and time point. Differential expression is calculated by DESeq2 using the Wald test as the average over 4 samples: 2 wild-type and 2 deletion strains, each strain measured in biological duplicate. B. GAT201 promotes upregulation of specific genes more than downregulation. Volcano plot of log2 fold-change and p-value, with differential expressed genes calculated and coloured as in panel A. Genes with extreme p-values or fold-changes are plotted at the edge of the panel area. C. GAT201-dependent differential gene expression is more extreme at later timepoints. The panel shows all the genes that are at least 8x differentially expressed in any combination of condition and time, ordered by their average fold-change at 4 hours. D. Over half of the upregulated differentially expressed genes are direct targets of GAT201. Venn diagram shows the number DEGs in RPMI at 4 hours (as in panel B) compared to Gat201 targets measured by ChIP-seq from Homer et al. 2016. This is approximately a 3-fold enrichment. E. GAT204, as well as GAT201, is required to repress growth of cells in RPMI.
Loss of activation of the Gat201 pathway in gat201Δ strains is confirmed by lower expression of known Gat201 targets during reactivation including BLP1 (Fig 4C), GAT204, and LIV3 (Fig S8). Functional enrichment analysis of genes up in GAT201 cells highlighted changes at the cell surface, including transmembrane transport (GO:0055085) and cell-wall related terms carbohydrate metabolic processes (GO:0005975) and glycosyl hydrolase activity (GO:0016798). Metabolic pathway analysis highlighted GAT201-upregulated genes required for capsule biosynthesis (PWY-5114__PK__MetaCyc, UDP-sugars interconversion), including UXS1/CNAG_03322 and UGD1/CNAG_04969 (O’Meara and Alspaugh 2012), consistent with the observed GAT201-dependent capsule synthesis during reactivation. Genes that are down in GAT201 strains are enriched for ribosome biogenesis (GO:0042254) and related terms, indicating downregulation of the core biosynthetic process of protein synthesis. Genes down in GAT201 strains were also enriched in carbohydrate metabolic processes (GO:0005975), indicating that different sets of carbohydrate-processing functions are upregulated and downregulated depending on GAT201. Overall, this implicates Gat201 in controlling carbohydrate metabolism and cell surface remodeling, and in repressing core biosynthetic processes related to growth.
We found that many of the GAT201-dependent genes are direct targets of Gat201. Previous measurements of binding of Gat201 to DNA by ChIP-seq found roughly 1200 enriched peaks out of approximately 6800 annotated Cryptococcal genes (Homer et al. 2016). We compared these targets to our list of differentially expressed genes at 240 minutes in RPMI (Fig 4D). Over 50% of the GAT201-upregulated genes here are direct targets (151/290), representing approximately 3x enrichment. The GAT201-downregulated genes at 240 minutes in RPMI are also about 2x enriched in direct targets (73/210). We see similar enrichment at earlier timepoints.
GAT204, as well as GAT201, is required to repress growth of cells in RPMI.
We next asked which GAT201 targets could be required for the GAT201-dependent repression of proliferation and growth in RPMI. We took a reverse genetics approach focusing on phenotypically important GAT201 targets and transcription factors, measuring growth curves of deletion mutants from the Madhani collection (Liu et al. 2008). Gat204 and Liv3 are transcription factors implicated in virulence that are Gat201 targets (Chun, Brown, and Madhani 2011) and whose direct target genes overlap with those of Gat201 (Homer et al. 2016). In RPMI medium, gat204Δ cells behave similarly to gat201Δ cells by increasing in density, unlike wild-type cells that decline in density within 10 hours (Fig 4E, Fig S10). Gat204 regulates approximately 30% of Gat201-regulated genes (Homer et al., 2016). An intermediate density is shown by liv3Δ cells (Fig 4E). Conversely, the barwin-like protein Blp1 that is required for the antiphagocytic function of Gat201 is dispensable for the growth phenotype: blp1Δ cells grow similarly to wild-type cells in RPMI (Fig 4E).
Of the genes that we have tested so far (Fig S10), only GAT201, GAT204, and LIV3 are required for growth restriction in RPMI medium. Deletion of other GAT201 targets, including transcription factors PDR802 and ECM2201, and metalloproteinase MEP1, did not relieve restriction of growth (Fig S11).
Furthermore, we tested GAT204 and LIV3 genetic interactions in RPMI medium. The growth phenotypes of gat204Δ, liv3Δ and the double mutant gat204Δ liv3Δ were similar, all growing better than wild-type in RPMI but not as well as gat201Δ (Fig S11). This result is consistent with GAT204 and LIV3 operating in the same pathway, as previously reported. Overall, these data argue for the existence of a specific Gat201/Gat204/Liv3 dependent pathway that restricts growth in specific media at alkaline pH.
GAT201 represses growth at alkaline pH but is required for growth in RPMI at neutral pH
We next examined the role of pH in determining the restricted growth phenotype. We had previously observed that, in RPMI buffered with sodium bicarbonate in our aerobic growth conditions, media alkalinization occurred regardless of the addition of serum or cells. However, in an alternative RPMI-like “CO2-independent media”, which is kept near neutral pH with a phosphate-based buffering agent, both GAT201 and gat201Δ cells continue to increase in density over a 72 hour period (Fig S12). This differential growth in media with otherwise identical nutrient composition suggested that the buffering agent and/or pH was responsible for the phenotype.
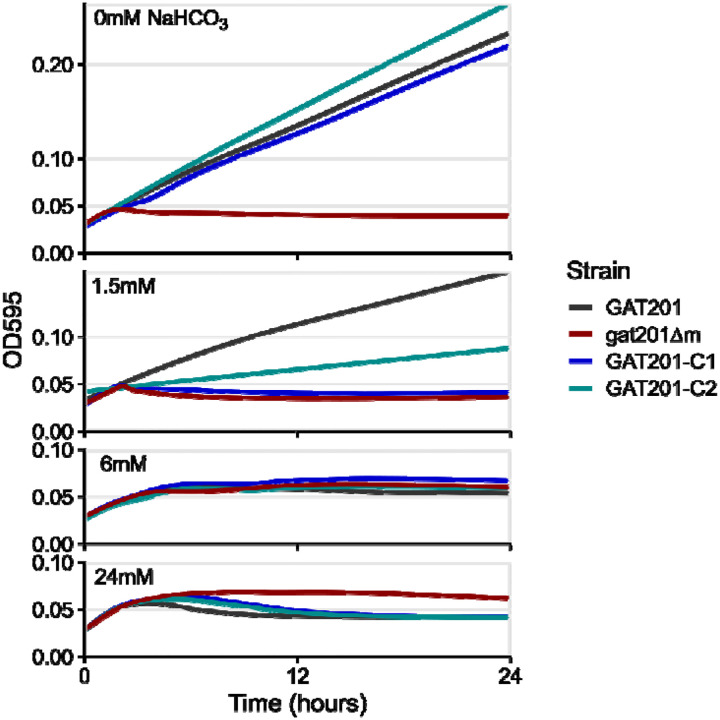
Accordingly, we next isolated the effect of sodium bicarbonate (NaHCO3) on growth, by growing cells in RPMI with varying concentrations of NaHCO3 in aerobic conditions. With 24mM NaHCO3, the same as in the standard RPMI formulation used previously, we again observed that wild-type GAT201 cells do not grow in the long-term but gat201Δ cells do grow (Fig 5, Fig S13). Surprisingly, at lower concentrations of NaHCO3 this effect changes, and in RPMI with no added NaHCO3 wild-type cells grow consistently while gat201Δ cells do not grow (Fig 5, Fig S13). Complementing GAT201 into deletion strains restored the wild-type growth phenotype of growth at 0mM NaHCO3 and arrest at 24mM NaHCO3.
Fig 5. The effect of GAT201 on growth depends on sodium bicarbonate (NaHCO3).
Starting with an RPMI formulation lacking NaHCO3, we added either 0mM, 1.5mM, 6mM, or 24mM NaHCO3 and grew Cryptococcus for 24 hours. Wild-type GAT201 cells grow in 0mM NaHCO3 but do not grow in 24 mM NaHCO3, while gat201Δ have opposite phenotypes of no growth in 0mM and growth at 24 mM. These cells have intermediate phenotypes at intermediate concentrations of NaHCO3, while complemented strains have growth phenotypes resembling wild-type. This figure shows the median of 3 technical replicates from a single biological replicate, and 2 further biological replicates are shown in Fig S13.
To interpret these data, note that bicarbonate ions in the media are in exchange with carbon dioxide in the air around the cultures (Michl, Park, and Swietach 2019). Addition of 24mM NaHCO3 leads to an equilibrium pH of about 7.5 in 5% CO2 (Michl, Park, and Swietach 2019), however at atmospheric CO2 of roughly 0.04% this media reaches pH ~9.5 within hours. Without any added NaHCO3, the media pH remains close to neutral. Thus, the major differences in growth phenotypes that we observe occur after equilibration to more alkaline pH. Further work will be needed to dissect the effect of pH, buffers, and exogenous CO2 on GAT201-dependent growth, but the phenotype that GAT201 promotes growth in RPMI media at near-neutral pH and represses growth at alkaline pH is reproducible in our hands.
Because cyclic-AMP dependent signaling through the Rim101 transcription factor also regulates growth at alkaline pH (Caza and Kronstad 2019; O’Meara et al. 2010), we tested if cyclic AMP signaling affects our observed GAT201-dependent phenotype. We found that addition of the exogenous cAMP does not substantially affect growth in RPMI with or without NaHCO3 added: again, wild-type GAT201 cells grow far more than gat201Δm in the absence of NaHCO3, and the phenotype is reversed in 24 mM NaHCO3 (Fig S14). This shows that the GAT201 pathway is largely independent of cAMP signaling, and thus of Rim101 signaling, indicating a distinct alkaline-responsive pathway controlled by GAT201.
C. neoformans Gat201 is homologous to other GATA-family zinc finger proteins that regulate fungal growth and environmental responses.
Lastly, we asked if Gat201 could be homologous to other fungal transcription factors that might indicate a conserved pathway. Gat201 is a 435 amino acid-long protein predicted to have only a single structured domain of 58 amino acids near the C-terminus, the GATA-like zinc finger domain (Fig 6A). This domain is found across a broad variety of transcription factors that integrate environmental signals and metabolism (Lowry and Atchley 2000). Searching for homologs of C. neoformans Gat201 by BLASTP (Sayers et al. 2022) detects many proteins with GATA-like domains and a variety of domain structures, consistent with the known diversity in GATA-like transcription factors (Lowry and Atchley 2000). Because there are multiple GATA-like domain proteins in each fungal species that we examined, we turned to more precise methods to search for true homologs of Gat201.
Fig 6. C. neoformans Gat201 is homologous to other GATA-family zinc finger proteins that regulate fungal growth and environmental responses.
A, Domain structure of Gat201 and 4 close homologs, with GATA-like zinc finger domain shown in red (Interpro IPR013088) and predicted unstructured regions in blue (MobiDB Lite consensus disorder), taken from Interpro (Blum et al. 2021). B, Multiple sequence alignment of the GATA-like zinc finger domains of homologs made with MUSCLE (Edgar 2004). Conserved cysteine residues typical of GATA-like zinc fingers are indicated with asterisks. An extended phylogeny and homology analysis is shown in Fig S15.
We turned to the PANTHERDB curated homology database, and selected the proteins closest to C. deneoformans Gat201 within family GATA transcription factor, PTHR45658 (Mi et al. 2019). This database groups Gat201 with fungal and amoebal proteins that have the same domain structure, including S. cerevisiae Gat2 and C. neoformans Gat204, as well as wc-2 transcription factors that are involved in light responses in filamentous fungi, that have an additional PAS sensing domain (Ballario et al. 1998). We then filtered to only homologs with the same domain structure as Gat201, performed a full-length protein alignment seeded with the GATA domain using MAFFT (Katoh and Standley 2013), and calculated a phylogenetic tree by Bayesian maximum likelihood using IQ-TREE (Minh et al. 2020).
Our analysis grouped C. neoformans Gat201 alongside a subset of other proteins with a single C-terminal GATA-like domain: Ustilago maydis UMAG_04076, Neurospora crassa Sub-1, Aspergillus fumigatus NsdD, and Candida albicans Brg1 (Fig 6A, Fig S15A). Several of these homologs have reported roles in regulating growth and environmental responses (Muñoz-Guzmán, Caballero, and Larrondo 2021; Lee et al. 2014; Cleary et al. 2012; Lu, Su, and Liu 2012). Fungal Gat201-like transcription factors are most closely related to amoeba homologs including gtaI, a GATA-family transcription factor that is required for morphological transitions in Dictyostelium discoideum (Katoh-Kurasawa et al. 2021). S. cerevisiae Gat2, Gat3, and Gat4 were grouped in a separate clade, while C. neoformans Gat204 was grouped into another clade close to a different set of amoebal homologs (Fig S15A). The zinc-finger domains of these Gat201 homologs are highly conserved (Fig 6B, Fig S15B), including the 4 cysteines that coordinate the zinc ion (Lowry and Atchley 2000). In addition, the AlphaFold2 protein structure database predicts a short unannotated alpha-helix-rich domain in C. neoformans Gat201, N. crassa Sub-1 and A. fumigatus NsdD (Varadi et al. 2022).
As an alternative method of searching for true homologs of GAT201, we looked for syntenic (gene-order conserved) homologs using the GenomicusFungi synteny browser (Nguyen et al. 2018). We searched against C. deneoformans GAT201 (CNC06330), which is syntenic to C. neoformans GAT201 across the locus. We found a set of basidiomycete syntenic homologs with a C-terminal GATA-like domain, including U. maydis UMAG_04076, that was identified as a close homolog in our phylogenetic analysis. Within the Agaricomycotina, the gene neighborhoods of these GATA-like domain proteins were dynamic, with extensive recombination. While there was limited preservation of wider gene synteny, GATA-like proteins within the Basidiomycota tend to be flanked upstream by genes encoding a Vms1-like protein (CNC06290, Ch3, UMAG_04964, Ch15), similar to S. cerevisiae YDR049W, (ChIV), and a lipid binding protein (CNC06300, Chr3; and UMAG_11290, Chr11) homologous to S. cerevisiae YJL036W (ChrX). There is limited longer range synteny downstream of GAT201, but the GAT201 gene is closely flanked by a DEAD box RNA helicase (CNC06360, Chr3; and UMAG_04080, Chr11). S. cerevisiae encodes two orthologous genes of the DEAD-box RNA helicase: DED1 and DBP1 (YOR204W and YPL119C, located on Chr XV and Chr XVI respectively). Examination of synteny around these genes in S. cerevisiae identified genes that are well conserved in Cryptococcus and Ustilago, including BEM3 (CNJ02560, Chr10; UMAG_11852, Chr2) and HIS3 (CNH01620 Chr8; UMAG_11859 Chr2), neither of which were syntenic with GAT201 in either Cryptococcus neoformans or Ustilago maydis, nor even located on the same chromosome (GAT201: CNC06330, Chr 3; UMAG_04076, Chr 11). Overall, this analysis does not support a syntenic relationship between GAT201 and other genes containing GATA-like domains beyond the Basidiomycota.
It remains possible that a more exhaustive analysis would detect more homologs or more precise recombination events involving related GATA-like domains. Short and less structured proteins like Gat201 contain less phylogenetic signal and are prone to “homology detection failure” (Weisman, Murray, and Eddy 2020). Still, the phylogenetic identification of a Gat201-like subfamily is supported by bootstrap analysis, by the similarity of the GATA domains, and by synteny analysis. Future work will be needed to assess which of Gat201’s predicted homologs have conserved molecular function or operate in a conserved pathway.
Discussion
Proliferation is a major driver of C. neoformans pathogenesis: cryptococcosis pathology is driven by the accumulation of yeast in diverse host niches, and high fungal burden is a strong correlate of poor outcomes (J. W. Kronstad et al. 2011; Ballou and Johnston 2017; Brouwer et al. 2004; Bicanic et al. 2009). To proliferate in the host and cause disease, C. neoformans yeast must rapidly adapt to the lung environment, characterised by nutrient limitation, high temperature (37°C), and transient high pH (>8.5) (Dusik Kim et al. 2021). In this study we modeled the early events of the fungal transition from stationary phase to conditions that support proliferation (rich media; 25°C or 37°C, pH 5.5) and those that stimulate defense through the formation of a polysaccharide capsule (RPMI+serum; 37°C, pH 8.5–9). At early time points, we observed the activation of cryptic pathways in host-like media (RPMI-1640 with or without serum) that correlated with expression of the virulence-associated transcription factor GAT201. Surprisingly, activation of these pathways was associated with restriction in budding and, later, with loss of viability. By studying both short term and extended growth in alkaline conditions, we identified a cryptic growth program not revealed by previous work. Future work will leverage these conditions to reveal the molecular mechanisms underpinning Alkaline-Restricted Growth (ARG).
C. neoformans is known to be extremely sensitive to alkaline pH, failing to grow above pH 8.5 (Howard 1961; Levitz et al. 1997). However, it is surprising that this alkaline-restricted growth phenotype is rescued by deletion of the GAT201 transcription factor, and that the same transcription factor blocks growth at neutral pH in otherwise identical media. The fact that deletion of a transcription factor restores budding, density, and viability indicates that the observed restricted growth of wild-type cells is a consequence of a regulated gene expression program. This genetic constraint on growth is not solely a physiological constraint due to alkaline pH or lack of nutrients.
Previous studies in C. neoformans have focused on genes whose loss further restricts growth at alkaline pH (Ost et al. 2015). Here, by contrast, we demonstrated that loss of the Gat201 pathway enables growth at alkaline pH. This restriction is apparent as early as 2 hours, when there is little yeast bud emergence and production of substantial polysaccharide capsule in wild-type cells. Deletion of GAT201 restores yeast cell budding and also disrupts the production of capsule, and this is consistent with microscopy data published in a previous study that focused on capsule rather than on proliferation (Gish et al. 2016). From 12 hours onwards, we found that wild-type C. neoformans becomes inviable, while viability was rescued by deletion of GAT201. These assays for Gat201 pathway activation will enable dissection of the pathway and its involvement in proliferation, that could shed light on Gat201’s role in promoting virulence.
We observed transcriptional activation of GAT201 within minutes of reactivation of C. neoformans in RPMI, independent of serum and temperature. Correspondingly, in these conditions we observed upregulation of many direct targets of GAT201, i.e. genes whose promoters are bound by Gat201, including transcriptional cofactors GAT204 and LIV3. We further observed that GAT204 and LIV3 are also required for alkaline-repressed growth. This Gat201-dependent transcriptional pathway appears to be independent of well-studied alkaline responsive pathways: RIM101 transcript expression does not depend on GAT201 (Fig S7), nor does GAT201 expression depend on RIM101 (O’Meara et al. 2013), and our comparison of Gat201-dependent and Rim101-dependent transcriptional profiles revealed no statistical enrichment for shared targets (data not shown). Rim101 acts downstream of the cAMP pathway, and exogenous cAMP does not change the GAT201-dependent growth phenotypes (Fig S14). Also, the expression of genes necessary for growth in alkaline conditions, such as PHO4 (Lev et al. 2017), ENA1 (Idnurm et al. 2009), ECA1 (Ariño, Ramos, and Sychrová 2010), CAN2, or CAC1 (Mogensen et al. 2006), were not dependent on GAT201. Although we did identify a subset of GAT201-dependent transcripts specific to serum within 4 hours, we did not observe an early phenotypic impact of serum on growth, suggesting that pH rather than serum is the dominant environmental signal.
The GAT201-dependent transcriptional profiles during alkaline-restricted growth provide some insight into the mechanisms of growth restriction. GAT201-dependent downregulation of ribosome biogenesis genes indicates that protein synthesis, a core pathway required for growth, is repressed downstream of GAT201 during reactivation in RPMI. By contrast, we observed ribosomal proteins to be strongly induced in wild-type cells reactivating in rich YPD media. This shows that Cryptococcus grown in alkaline RPMI represses biosynthetic and replicative processes while also upregulating capsule production. In addition, others have shown that capsule synthesis is restricted to the G1 phase of the cell cycle, while budding occurs in G2 (García-Rodas et al. 2014).
We propose 3 non-exclusive hypotheses for how GAT201 restricts growth. First, the GAT201 pathway could restrict growth directly or indirectly through transcriptional regulation of a core growth pathway. Second, the GAT201 pathway could promote capsule production, and the redirection of biosynthetic resources to capsule production could result in restricted growth. Third, some factors required for nutrient acquisition could be regulated downstream of GAT201, and a failure to acquire some essential nutrients would restrict growth. While nutrient acquisition could be a key contributor, we can exclude that nutrient content alone explains growth restriction, because GAT201 and gat201Δ cells behave differently in identical nutrient-rich RPMI media. Modulation of media pH and buffering agents changes these growth phenotypes, and in RPMI without sodium bicarbonate at near-neutral pH GAT201 instead promotes growth, again in media with otherwise identical nutrient composition. Alkaline conditions reduce the availability of H+ ions, which are important for the transport of nutrients across the cell membrane. We observed no GAT201-dependence for the expression of known H+ pumps required for alkaline growth, however several transmembrane transporters are upregulated in GAT201 cells compared to gat201Δ. Future work will need to investigate these mechanisms.
In contrast to C. neoformans, ascomycete pathogens are more alkaline tolerant: Aspergillus species are tolerant to pH 11 (Wheeler, Hurdman, and Pitt 1991), and Candida species can tolerate alkaline conditions ranging from pH 10, for C. albicans, to pH 13 for C. auris and C. parapsilosis (Heaney et al. 2020). In ascomycetes, alkaline growth is enabled by Rim101/PacC (Selvig and Alspaugh 2011). Likewise, Rim101 is required for C. neoformans growth at pH 8–8.5 (Ost et al. 2015). However, C. neoformans also employs Rim101-independent mechanisms to mediate alkaline tolerance, for example, the sterol homeostasis pathway regulated by the transcription factor Sre1 (Brown et al. 2020). Interestingly, the basidiomycete plant pathogen Ustilago maydis also exhibits alkaline-restricted growth that is Rim101-independent, but the causative pathway is unknown (Cervantes-Montelongo, Aréchiga-Carvajal, and Ruiz-Herrera 2016).
Our predictions of Gat201 homologous proteins in basidiomycetes and ascomycetes also suggest hypotheses for future investigation. It would be interesting to test if the syntenic homologous protein in Ustilago maydis, UMAG_04076, is involved in alkaline restricted growth (Cervantes-Montelongo, Aréchiga-Carvajal, and Ruiz-Herrera 2016). The predicted Neurospora homologous protein, Sub-1, co-regulates genes downstream of the light responsive white collar complex, connecting light responses and fungal development (Chen et al. 2009). The Aspergillus homologous protein, NsdD, is a crucial regulator of sexual development (Han et al. 2001; Szewczyk and Krappmann 2010). Brg1 in Candida albicans is required for hyphal growth, biofilm formation and virulence (Su et al. 2018). Collectively, this suggests that Gat201 may be part of a conserved family of GATA transcription factors that regulate proliferation and morphology in response to environmental stimuli. Defining the regulatory targets, co-factors, and upstream signaling pathways leading to Gat201 family activation in different species would reveal the degree of functional conservation.
In conclusion, we have found that GAT201 is part of an alkaline-restricted growth pathway that responds to environmental signals, including alkaline pH, to restrict cell proliferation and promote the synthesis of defensive capsule (Fig 7). The alkaline-restricted growth pathway is independent of the Rim101 pathway that promotes growth at weak alkaline pH, as well as independent of cAMP. We see over 10-fold increase in GAT201 mRNA abundance within 30 minutes of inoculation in RPMI media, suggesting the existence of fast-acting upstream pathway components that induce GAT201 transcription and/or stabilise the GAT201 transcript. Downstream, Gat201 regulates hundreds of targets, including other transcription factors that are implicated in Cryptococcus virulence, as well as many poorly characterised genes. So far, the only other genes that we know to be required for alkaline-restricted growth encode co-factors GAT204 and LIV3. Future work will map pathway components using forward genetic approaches that exploit growth conditions where the functional Gat201 pathway renders Cryptococcus inviable.
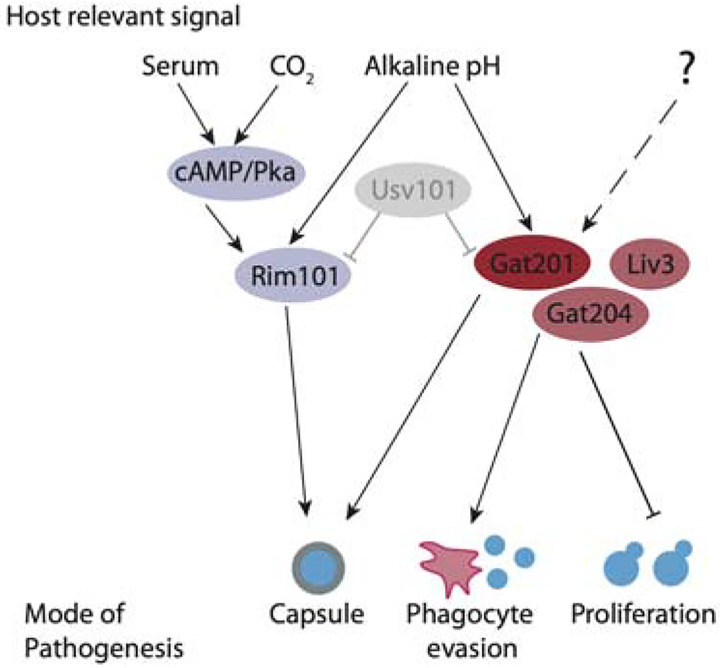
Fig 7. The Gat201 pathway promotes Cryptococcus virulence and represses proliferation.
Gat201 acts in parallel to serum-responsive cAMP/Pka pathway and the major pH-responsive Rim101 pathway. Gat201 requires mutual activators, Gat204 and Liv3, to suppress proliferation.
Materials and Methods
Strains, media, and growth conditions
RNA-seq Dataset 1
Wild-type C. neoformans H99 (J. R. Perfect, Lang, and Durack 1980) yeast were gifts from Andrew Alspaugh, Duke University, NC, USA and were maintained on YPD agar plates at room temperature. For stationary phase, cells were inoculated into multiple tubes of 5 ml liquid YPD (1% yeast extract, 2% Bactopeptone, 2% Dextrose) and incubated for 5 days at 200 rpm, 30°C. On day 5, the temperature was reduced to 25°C and the cells were allowed to adjust for >4 hours. 1mL of this culture was collected for RNA extraction as the “0 minute” timepoint. Cells were counted by hemocytometer at approximately 3.5 × 108 cells. Four aliquots of 3 ml of stationary phase culture were pelleted and resuspended in conditions: 25°C YPD, 37°C YPD (prewarmed), 25°C RPMI 1640 + 10% heat-inactivated fetal calf serum (HI-FCS), and 37°C RPMI + 10% HI-FCS (prewarmed), 100 ml each, so roughly a 1:30 dilution. The precise media used was RPMI 1640 with L-glutamine, sodium bicarbonate and pH indicator phenol red (Sigma R8758). Fetal calf serum (Biosera FB-1285) was heat-inactivated by incubating at 56°C for 30 minutes, then aliquoted and stored at −20°C until required. The pH was checked at time 0 and 120 minutes and confirmed to be within physiological range for the duration of the experiment (7.0 – 8.5). Cells were incubated at 60 rpm. Another stationary phase sample was inoculated into at YPD at 30°C and grown at 150 rpm for 180 min. The two biological replicates were collected on two successive days.
RNA-seq Dataset 2
Wild-type KN99a and KN99alpha C. neoformans yeast (Nielsen et al. 2003) were maintained on YPDA agar plates (1% yeast extract, 2% Bacto-peptone, 2% Dextrose, 0.002% Adenine, 2% agar) at room temperature or stored at −80°C in 15% glycerol. Gat201 mutants from the Bahn lab transcription factor disruption collection (Jung et al. 2015) and the Madhani lab C. neoformans deletion collection (Liu et al. 2008) were obtained from the Fungal Genetics Stock Center (Manhattan, KS, USA; https://fgsc.net). We verified the gene disruption/deletion by PCR and Sanger sequencing. For stationary phase, cells from a single colony were inoculated into 20 ml liquid YPDA (1% yeast extract, 2% Bacto-peptone, 2% Dextrose, 0.002% Adenine) and incubated for 5 days at 200 rpm, 30°C. On day 5, 3.5 ml from each sample was added to fresh media, pre-warmed RPMI or RPMI + 10% HI-FBS, to a total volume of 100 ml, and incubated at 37°C with 60 rpm shaking. Samples were collected at 30 min, 2 hours, and 4 hours. Two biological replicates were performed for each of the four strains. RPMI 1640 with L-glutamine, sodium bicarbonate and pH indicator phenol red (Sigma R8758) and heat-inactivated serum (HI-FBS; Sigma F9665, Lot# BCCB5091) was aliquoted and stored at −20°C until required.
RNA extraction and library preparation
RNA-seq Dataset 1
At each time point, 15 ml of the 100 ml culture (1 ml of stationary phase culture) was collected and immediately fixed in 7 ml Methanol (32% v/v final) in a 50 mL falcon tube on dry ice. Fixed cells were pelleted, transferred to a 1.5 ml screw-top tube in 1 mL ddH2O, and pelleted again. Pellets were lyophilized and mixed with 200 μL zirconium beads and ground on a bead-beater (Biospec 112011EUR) for 5 minutes. 1400 μL of buffer RLT (Qiagen) + 1% β-mercaptoethanol was added, and the mixture vortexed. RNA was then extracted by the Qiagen RNeasy Plant Mini Kit according to the manufacturer’s protocol for filamentous fungi, proceeding from step 4. RNA quality was checked by nanodrop and agilent bioanalyzer.
RNA-seq Dataset 2
At each time point, 15 ml of culture (2 ml of stationary phase culture) was collected and immediately fixed in 7 ml Methanol (32% v/v final) in a 50 mL falcon tube on dry ice. Fixed cells were pelleted, transferred to a 1.5 ml screw-top tube in 1 mL ddH2O, and pelleted again. Pellets were lyophilized, then mixed with 200 μL zirconium beads + 1 ml TRI reagent (Invitrogen) and incubated at room temperature for 5 mins followed by flash freezing in a dry ice/ethanol bath. Samples were thawed at room temperature and subjected to mechanical lysis by bead beating (using a PreCellys machine) for 3 × 10 seconds (6000 rpm) followed by a pause of 20 seconds and placed on ice for 1 minute. This was repeated 10 times. After mechanical disruption the zirconium beads were pelleted and the supernatant transferred to a QIA shredder spin column from the Qiagen RNeasy Plant Mini Kit (Qiagen, Valencia, CA, USA) and RNA was then extracted according to the manufacturer’s protocol, proceeding from step 4. Total RNA quantity and quality were assessed using nanodrop and the Fragment Analyser Automated Capillary Electrophoresis System (Agilent Technologies Inc, #5300) and the Standard Sensitivity RNA Analysis Kit, 15nt (#DNF-471).
For the experiments in Fig 1 (Dataset 1), 2 μg of RNA from each sample was used as input for RNA sequencing by the RNATagSeq protocol (Shishkin et al. 2015), with minor modifications including ribosomal RNA depleted using the Yeast RiboZero Gold kit (Illumina; now discontinued), and a random barcode added to the 2nd ligation primer, with 12 cycles of PCR. Libraries were sequenced on a Nextseq500 (Illumina).
For the experiments in Figs 3–4 (Dataset 2) 500 ng of RNA from each sample was used as input for cDNA library preparation using the QuantSeq FWD 3’ mRNA-Seq Library Prep Kit for Illumina platforms according to the manufacturer’s instructions. We spiked in 10 ng of Saccharomyces cerevisiae total RNA as a loading control, but did not use this spike-in information in the data analysis presented here. The QuantSeq protocol generates only one fragment per transcript, close to the 3’ end of the transcripts. cDNA fragments of ~300 bp were purified from each library and confirmed for quality by the Fragment Analyser Automated Capillary Electrophoresis System (Agilent Technologies Inc, #5300) and the Standard Sensitivity NGS 1–6000bp Kit (#DNF-473–33). Single read sequencing was performed using the NextSeq 500/550 High-Output v2.5 (75 cycles) Kit (#20024906) on the NextSeq 550 platform (Illumina Inc, #SY-415-1002). Libraries were combined in a single equimolar pool of 56 based on Qubit and Bioanalyser assay results and run across a High-Output v2.5 Flow Cell.
RNA-sequencing data are available on GEO under accession numbers GSE133067 (Dataset 1) and GSE217345 (Dataset 2).
Bioinformatic and statistical data analyses of RNA-seq data
Complete analysis code for the RNA-seq datasets from raw reads onwards is found on github, https://github.com/ewallace/CryptoWakeupRNASeq (Dataset 1) https://github.com/ewallace/CryptoGat201RNASeq (Dataset 2).
In summary, basic assessments of sequence data quality were performed using FastQC (Andrews and Others 2010) and MultiQC (Ewels et al. 2016). Raw sequencing reads were trimmed and filtered using cutAdapt (Martin 2011). Sequenced reads were aligned to the C. neoformans H99 reference sequence CNA3 (Janbon et al. 2014) using Hisat2 (Daehwan Kim et al. 2019). We used featureCounts (Liao, Smyth, and Shi 2014) to assign mapped reads per gene using the longest transcript per gene annotation from (Wallace et al. 2020). Gene expression was normalized using the regularized logarithm (rlog) function from DESeq2 (Love, Huber, and Anders 2014). We evaluated the gene expression differences using a test based on a negative binomial distribution, also in DESeq2 (Love, Huber, and Anders 2014), using a 5% false discovery rate calculated by the ‘p.adjust’ function in R using the Benjamin and Hochberg method (Haynes 2013).
Gene clustering was performed by using (1 - correlation) as a distance metric, then hierarchical clustering in UPGMA using R’s hclust function (Holmes and Huber 2019). After a lengthy iterative process in which many methods were evaluated, representative clusters of genes were selected from the hierarchical clusters using R’s cutree function with user-defined numbers of groups (k option). Full details and analysis code are in the repositories.
Differentially expressed genes and gene clusters were subjected to GO term enrichment analysis using the online resources at FungiDB (Basenko et al. 2018).
Strain Construction
To complement gat201Δ, theGAT201 gene including native promoter, terminator, and introns with a HYG selection marker (pGAT201-cGAT201-tGAT201-HYG) was integrated into a genomic safe haven locus 4 on chromosome 7 (Erpf, Stephenson, and Fraser 2019) in the Madhani lab gat201Δ strain. We made the integration constructs using modular cloning by Möbius assembly (Andreou and Nakayama 2018), the details of which we will explain in another publication. A full plasmid map is included in supplementary table S4. We integrated this construct using a Cryptococcus CRISPR-Cas9 system (Huang et al. 2022).
Microscopy
For stationary phase, cells were revived from −80°C glycerol stocks on YPD agar and within two days single colonies were inoculated into 10 ml liquid YPD (1% yeast extract, 2% Bactopeptone, 2% Dextrose) and incubated for 5 days at 200 rpm, 30°C. On day 5, the temperature was reduced to 25°C and the cells were allowed to adjust for >4 hours. Cells were counted by hemocytometer and 1 ml of the pellet (106 cells) was collected, washed 1x with PBS, and split into four tubes, then resuspended in the appropriate pre-warmed medium as indicated to a final volume of 10 ml each. Cells were incubated in the indicated condition for 120 minutes, and then the entire pellet was collected and fixed with 4% methanol free formaldehyde (Pierce) for 10 minutes, then washed 3x with PBS. India ink (Remel) slides were prepared and cells were imaged using an inverted Zeiss AxioObserver Z1 with a Plan-Neofluor 40X/1.3 numerical aperture (NA) oil immersion lens objective (Carl Zeiss) and a 16-bit CoolSNAP H2 charge-coupled-device (CCD) camera (Photometrics). For each figure, three biological replicates were initiated using independent stationary cultures and collected serially on the same day. The entire experiment was performed independently twice.
Growth Curves
Cells from a single colony were inoculated into 5 ml liquid YPDA (1% yeast extract, 2% Bacto-peptone, 2% Dextrose, 0.002% Adenine) and incubated for 20 hours at 180 rpm, 30°C. Cells were washed with ddH2O and resuspended in the required volume of the appropriate media to an initial OD 595 nm of 0.2. Wells in the microplate were filled with this suspension (200 μl in each well). The absorbance in each well was measured at 595 nm at given intervals (10 minutes) with shaking (300 rpm for 1 minute) directly prior to reading. Reference measurements were performed on the outer wells where 200 μl of media only was added. The microplate was Incubated in Tecan Infinite® 200 Pro plate reader at 37°C for 48 or 72 hours. Cells were grown as stated in RPMI 1640 (Sigma R8758) with or without heat-inactivated serum (Sigma F9665), YPDA, for figures except where noted. For Fig S12, we used CO2-independent media, buffered with mono and dibasic sodium phosphate and β-glycerophosphate (Gibco™ / ThermoFisher 18045088). For Figs 4, S13, and S14, we used RPMI media without phenol red and without NaHCO3 (Sigma R7855), adding dibutyryl cAMP (Sigma D0627) or NaHCO3 from aqueous stock solutions to the indicated final concentration. For these we were particularly careful to rapidly prepare media and then inoculate cells for growth curves, for reproducible pH of the growth media.
Colony forming unit assay
Colony forming unit assay is an in vitro cell survival assay based on the ability of a single cell to grow into a colony. Cells from a single colony were inoculated into 25 ml liquid YPDA (1% yeast extract, 2% Bacto-peptone, 2% Dextrose, 0.002% Adenine) and incubated for 5 days at 150 rpm, 30°C. Cells were washed with ddH2O and resuspended in a total volume of 20 ml RPMI at an OD 595 nm of 0.1 for each strain. Cultures were incubated at 37°C, 60rpm and samples were collected at 12 hour intervals (0–60 hours). Serial dilutions were prepared for each sample collected from each strain down to 10−4, 100 μl of dilutions 10−3 and 10−4 were plated onto YPDA agar plates and incubated at 30°C for 48 hours. Plates were imaged using an ImageQuant 800 (Amersham/Cytiva, settings: Colorimetric, OD measurement, Auto exposure, Capture area = 160 × 220 nm) and the resulting colonies were counted. Three biological replicates were collected for each strain.
RT-qPCR
Cells from a single colony were inoculated into 5 ml liquid YPDA (1% yeast extract, 2% Bacto-peptone, 2% Dextrose, 0.002% Adenine) and incubated for 20 hours at 180 rpm, 30°C. Cells were washed with ddH2O and resuspended in 100 ml RPMI (Sigma R8758) at an initial OD 595 nm of 0.1, incubated at 37°C, 150 rpm for 7 hours. Cells were fixed in methanol and RNA extracted using mechanical disruption in TRizol with zirconium beads followed by the Qiagen Plant and Fungal Extraction Kit. 100 ng of purified RNA was used from each sample to synthesize cDNA using Superscript IV Reverse Transcriptase (Invitrogen) and random primers (NEB). Samples were DNase treated prior to reverse transcription. QPCR was carried out using Brilliant III Ultra fast SYBR Green qPCR mix (Agilent) with appropriate target gene primers. mRNA expression of GAT201 (Forward primer: 5’-ACCACGAGTCTTGGGATAGA-3’, Reverse primer: 5’-CTGGGTGTTCGGGATAAAGTAG-3’), GAT204 (Forward primer: 5’-CCACCTCTTCCTTCCTTGTTAAA-3’, Reverse primer: 5’-GTCTGCCATCGTCGTACTAATG-3’), and LIV3 (Forward primer: 5’-CCTCTTCCACTTCCACATCAA-3’, Reverse primer: 5’GGTCTCGGCACAGCATATT-3’). Test genes were compared to 3 reference genes ACT1 (Forward primer: 5’-GTGGTTCTATCCTTGCCTCTTT-3’, Reverse primer: 5’-CACTTTCGGTGGACGATTGA-3’), GPD1 (Forward primer: 5’-TCGAGCAACGTCTTGGTATC3’, Reverse primer: 5’-GCTCTCCATCCTCCTTGTTT-3’) and SRP14 (Forward primer: 5’--3’, Reverse primer: 5’--3’). The data were analysed with tidyqpcr (Wallace and Haynes 2022).
Data analysis
Data analysis scripts and raw data for budding index assay, growth curve assay, CFU assay, and RT-qPCR are in the repository https://github.com/ewallace/CryptoGat201_2023_suppdata. Scripts and raw data for the homology analysis are in the repository https://github.com/ewallace/Gat201homology_2022/. Data were analysed in the statistical open-source language R (R Core Team 2023), making extensive use of the tidyverse for data manipulation (Wickham et al. 2019) and ggplot2 for figures (Wickham 2009). Additional figures were prepared in Inkscape (The Inkscape Team, https://inkscape.org/).
Supplementary Material
Acknowledgements
E.S.H. was supported by a Fellowship from the Daphne Jackson Memorial Trust, funded by the University of Edinburgh and the Biotechnology & Biological Sciences Research Council. E.R.B. and E.W.J.W. are each supported by Sir Henry Dale Fellowships jointly funded by the Wellcome Trust and the Royal Society (211241/Z/18/Z to E.R.B., 208779/Z/17/Z to E.W.J.W.). Collection of Dataset 1 was funded by a Research Incentive Grant from the Carnegie Trust for the Universities of Scotland to E.W.J.W., and by the European Union’s Horizon 2020 research and innovation programme under Marie Skłodowska-Curie grant agreement no. 661179 to E.W.J.W. We acknowledge funding from the MRC Centre for Medical Mycology at the University of Exeter (MR/N006364/2 and MR/V033417/1), and the NIHR Exeter Biomedical Research Centre. Additional work may have been undertaken by the University of Exeter Biological Services Unit. The views expressed are those of the author(s) and not necessarily those of the NIHR or the Department of Health and Social Care. We thank Hiten Madhani for gifts of deletion strains, supported by NIH grant (R01AI100272). We thank Andrew Cassidy for sequencing RNA-seq dataset 1 at the Tayside Centre for Genomic Analysis, University of Dundee. We thank Richard Clarke, Angie Fawkes, and Lee Murphy, for sequencing RNA-seq dataset 2 at the Genetics Core of the Edinburgh/Wellcome Trust Clinical Research Facility. We thank the Wallace lab and Ballou lab for comments and feedback on the project over the years.
References
- Andreou Andreas I., and Nakayama Naomi. 2018. “Mobius Assembly: A Versatile Golden-Gate Framework towards Universal DNA Assembly.” PloS One 13 (1): e0189892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andrews Simon, and et al. 2010. “FastQC: A Quality Control Tool for High Throughput Sequence Data.” Babraham Bioinformatics, Babraham Institute, Cambridge, United Kingdom. [Google Scholar]
- Ariño Joaquín, Ramos José, and Sychrová Hana. 2010. “Alkali Metal Cation Transport and Homeostasis in Yeasts.” Microbiology and Molecular Biology Reviews: MMBR 74 (1): 95–120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ballario P., Talora C., Galli D., Linden H., and Macino G.. 1998. “Roles in Dimerization and Blue Light Photoresponse of the PAS and LOV Domains of Neurospora Crassa White Collar Proteins.” Molecular Microbiology 29 (3): 719–29. [DOI] [PubMed] [Google Scholar]
- Ballou Elizabeth R., and Johnston Simon A.. 2017. “The Cause and Effect of Cryptococcus Interactions with the Host.” Current Opinion in Microbiology 40 (December): 88–94. [DOI] [PubMed] [Google Scholar]
- Basenko Evelina Y., Pulman Jane A., Shanmugasundram Achchuthan, Harb Omar S., Crouch Kathryn, Starns David, Warrenfeltz Susanne, et al. 2018. “FungiDB: An Integrated Bioinformatic Resource for Fungi and Oomycetes.” Journal of Fungi (Basel, Switzerland) 4 (1). 10.3390/jof4010039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bicanic Tihana, Muzoora Conrad, Brouwer Annemarie E., Meintjes Graeme, Longley Nicky, Taseera Kabanda, Rebe Kevin, et al. 2009. “Independent Association between Rate of Clearance of Infection and Clinical Outcome of HIV-Associated Cryptococcal Meningitis: Analysis of a Combined Cohort of 262 Patients.” Clinical Infectious Diseases: An Official Publication of the Infectious Diseases Society of America 49 (5): 702–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blum Matthias, Chang Hsin-Yu, Chuguransky Sara, Grego Tiago, Kandasaamy Swaathi, Mitchell Alex, Nuka Gift, et al. 2021. “The InterPro Protein Families and Domains Database: 20 Years on.” Nucleic Acids Research 49 (D1): D344–54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bose Indrani, Reese Amy J., Ory Jeramia J., Janbon Guilhem, and Doering Tamara L.. 2003. “A Yeast under Cover: The Capsule of Cryptococcus Neoformans.” Eukaryotic Cell 2 (4): 655–63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Botts Michael R., and Hull Christina M.. 2010. “Dueling in the Lung: How Cryptococcus Spores Race the Host for Survival.” Current Opinion in Microbiology 13 (4): 437–42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brouwer Annemarie E., Rajanuwong Adul, Chierakul Wirongrong, Griffin George E., Larsen Robert A., White Nicholas J., and Harrison Thomas S.. 2004. “Combination Antifungal Therapies for HIV-Associated Cryptococcal Meningitis: A Randomised Trial.” The Lancet 363 (9423): 1764–67. [DOI] [PubMed] [Google Scholar]
- Brown Hannah E., Telzrow Calla L., Saelens Joseph W., Fernandes Larissa, and Alspaugh J. Andrew. 2020. “Sterol-Response Pathways Mediate Alkaline Survival in Diverse Fungi.” mBio 11 (3). 10.1128/mBio.00719-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Casadevall A., Rosas A. L., and Nosanchuk J. D.. 2000. “Melanin and Virulence in Cryptococcus Neoformans.” Current Opinion in Microbiology 3 (4): 354–58. [DOI] [PubMed] [Google Scholar]
- Casadevall Arturo, Steenbergen Judith N., and Nosanchuk Joshua D.. 2003. “‘Ready Made’ Virulence and ‘Dual Use’ Virulence Factors in Pathogenic Environmental Fungi--the Cryptococcus Neoformans Paradigm.” Current Opinion in Microbiology 6 (4): 332–37. [DOI] [PubMed] [Google Scholar]
- Caza Mélissa, and Kronstad James W.. 2019. “The cAMP/Protein Kinase a Pathway Regulates Virulence and Adaptation to Host Conditions in Cryptococcus Neoformans.” Frontiers in Cellular and Infection Microbiology 9 (June): 212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cervantes-Montelongo Juan Antonio, Aréchiga-Carvajal Elva Teresa, and Ruiz-Herrera José. 2016. “Adaptation of Ustilago Maydis to Extreme pH Values: A Transcriptomic Analysis.” Journal of Basic Microbiology 56 (11): 1222–33. [DOI] [PubMed] [Google Scholar]
- Chen Chen-Hui, Ringelberg Carol S., Gross Robert H., Dunlap Jay C., and Loros Jennifer J.. 2009. “Genome-Wide Analysis of Light-Inducible Responses Reveals Hierarchical Light Signalling in Neurospora.” The EMBO Journal 28 (8): 1029–42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chun Cheryl D., Brown Jessica C. S., and Madhani Hiten D.. 2011. “A Major Role for Capsule-Independent Phagocytosis-Inhibitory Mechanisms in Mammalian Infection by Cryptococcus Neoformans.” Cell Host & Microbe 9 (3): 243–51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cleary Ian A., Lazzell Anna L., Monteagudo Carlos, Thomas Derek P., and Saville Stephen P.. 2012. “BRG1 and NRG1 Form a Novel Feedback Circuit Regulating Candida Albicans Hypha Formation and Virulence.” Molecular Microbiology 85 (3): 557–73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edgar Robert C. 2004. “MUSCLE: Multiple Sequence Alignment with High Accuracy and High Throughput.” Nucleic Acids Research 32 (5): 1792–97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Erpf Paige E., Stephenson Christina J., and Fraser James A.. 2019. “amdS as a Dominant Recyclable Marker in Cryptococcus Neoformans.” Fungal Genetics and Biology: FG & B 131 (October): 103241. [DOI] [PubMed] [Google Scholar]
- Ewels Philip, Magnusson Måns, Lundin Sverker, and Käller Max. 2016. “MultiQC: Summarize Analysis Results for Multiple Tools and Samples in a Single Report.” Bioinformatics 32 (19): 3047–48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- García-Rodas Rocío, Cordero Radames J. B., Trevijano-Contador Nuria, Janbon Guilhem, Moyrand Frédérique, Casadevall Arturo, and Zaragoza Oscar. 2014. “Capsule Growth in Cryptococcus Neoformans Is Coordinated with Cell Cycle Progression.” mBio 5 (3): e00945–14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giles Steven S., Dagenais Taylor R. T., Botts Michael R., Keller Nancy P., and Hull Christina M.. 2009. “Elucidating the Pathogenesis of Spores from the Human Fungal Pathogen Cryptococcus Neoformans.” Infection and Immunity 77 (8): 3491–3500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gish Stacey R., Maier Ezekiel J., Haynes Brian C., Santiago-Tirado Felipe H., Srikanta Deepa L., Ma Cynthia Z., Li Lucy X., et al. 2016. “Computational Analysis Reveals a Key Regulator of Cryptococcal Virulence and Determinant of Host Response.” mBio 7 (2): e00313–16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han K. H., Han K. Y., Yu J. H., Chae K. S., Jahng K. Y., and Han D. M.. 2001. “The nsdD Gene Encodes a Putative GATA-Type Transcription Factor Necessary for Sexual Development of Aspergillus Nidulans.” Molecular Microbiology 41 (2): 299–309. [DOI] [PubMed] [Google Scholar]
- Haynes Winston. 2013. “Benjamini–Hochberg Method.” In Encyclopedia of Systems Biology, edited by Dubitzky Werner, Wolkenhauer Olaf, Cho Kwang-Hyun, and Yokota Hiroki, 78–78. New York, NY: Springer New York. [Google Scholar]
- Heaney Helen, Laing Juliette, Paterson Linda, Walker Alan W., Gow Neil A. R., Johnson Elizabeth M., MacCallum Donna M., and Brown Alistair J. P.. 2020. “The Environmental Stress Sensitivities of Pathogenic Candida Species, Including Candida Auris, and Implications for Their Spread in the Hospital Setting.” Medical Mycology: Official Publication of the International Society for Human and Animal Mycology 58 (6): 744–55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holmes Susan, and Huber Wolfgang. 2019. Modern Statistics for Modern Biology. Cambridge University Press. [Google Scholar]
- Homer Christina M., Summers Diana K., Goranov Alexi I., Clarke Starlynn C., Wiesner Darin L., Diedrich Jolene K., Moresco James J., et al. 2016. “Intracellular Action of a Secreted Peptide Required for Fungal Virulence.” Cell Host & Microbe 19 (6): 849–64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Howard D. H. 1961. “Some Factors Which Affect the Initiation of Growth of Cryptococcus Neoformans.” Journal of Bacteriology 82 (3): 430–35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang Manning Y., Joshi Meenakshi B., Boucher Michael J., Lee Sujin, Loza Liza C., Gaylord Elizabeth A., Doering Tamara L., and Madhani Hiten D.. 2022. “Short Homology-Directed Repair Using Optimized Cas9 in the Pathogen Cryptococcus Neoformans Enables Rapid Gene Deletion and Tagging.” Genetics 220 (1). 10.1093/genetics/iyab180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Idnurm Alexander, Walton Felicia J., Floyd Anna, Reedy Jennifer L., and Heitman Joseph. 2009. “Identification of ENA1 as a Virulence Gene of the Human Pathogenic Fungus Cryptococcus Neoformans through Signature-Tagged Insertional Mutagenesis.” Eukaryotic Cell 8 (3): 315–26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Janbon Guilhem, Ormerod Kate L., Paulet Damien, Byrnes Edmond J. 3rd, Yadav Vikas, Chatterjee Gautam, Mullapudi Nandita, et al. 2014. “Analysis of the Genome and Transcriptome of Cryptococcus Neoformans Var. Grubii Reveals Complex RNA Expression and Microevolution Leading to Virulence Attenuation.” PLoS Genetics 10 (4): e1004261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jang Eun-Ha, Kim Ji-Seok, Yu Seong-Ryong, and Bahn Yong-Sun. 2022. “Unraveling Capsule Biosynthesis and Signaling Networks in Cryptococcus Neoformans.” Microbiology Spectrum, October, e0286622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jung Kwang-Woo, Yang Dong-Hoon, Maeng Shinae, Lee Kyung-Tae, So Yee-Seul, Hong Joohyeon, Choi Jaeyoung, et al. 2015. “Systematic Functional Profiling of Transcription Factor Networks in Cryptococcus Neoformans.” Nature Communications 6 (April): 6757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Katoh Kazutaka, and Standley Daron M.. 2013. “MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability.” Molecular Biology and Evolution 30 (4): 772–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Katoh-Kurasawa Mariko, Hrovatin Karin, Hirose Shigenori, Webb Amanda, Ho Hsing-I, Zupan Blaž, and Shaulsky Gad. 2021. “Transcriptional Milestones in Dictyostelium Development.” Genome Research 31 (8): 1498–1511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim Daehwan, Paggi Joseph M., Park Chanhee, Bennett Christopher, and Salzberg Steven L.. 2019. “Graph-Based Genome Alignment and Genotyping with HISAT2 and HISAT-Genotype.” Nature Biotechnology 37 (8): 907–15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim Dusik, Liao Jie, Scales Nathan B., Martini Carolina, Luan Xiaojie, Asmahan Abu-Arish Renaud Robert, et al. 2021. “Large pH Oscillations Promote Host Defense against Human Airways Infection.” The Journal of Experimental Medicine 218 (4). 10.1084/jem.20201831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kronstad James W., Attarian Rodgoun, Cadieux Brigitte, Choi Jaehyuk, D’Souza Cletus A., Griffiths Emma J., Geddes Jennifer M. H., et al. 2011. “Expanding Fungal Pathogenesis: Cryptococcus Breaks out of the Opportunistic Box.” Nature Reviews. Microbiology 9 (3): 193–203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kronstad Jim, Saikia Sanjay, Nielson Erik David, Kretschmer Matthias, Jung Wonhee, Hu Guanggan, Geddes Jennifer M. H., et al. 2012. “Adaptation of Cryptococcus Neoformans to Mammalian Hosts: Integrated Regulation of Metabolism and Virulence.” Eukaryotic Cell 11 (2): 109–18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee Mi-Kyung, Kwon Nak-Jung, Choi Jae Min, Lee Im-Soon, Jung Seunho, and Yu Jae-Hyuk. 2014. “NsdD Is a Key Repressor of Asexual Development in Aspergillus Nidulans.” Genetics 197 (1): 159–73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levitz S. M., Harrison T. S., Tabuni A., and Liu X.. 1997. “Chloroquine Induces Human Mononuclear Phagocytes to Inhibit and Kill Cryptococcus Neoformans by a Mechanism Independent of Iron Deprivation.” The Journal of Clinical Investigation 100 (6): 1640–46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lev Sophie, Kaufman-Francis Keren, Desmarini Desmarini, Juillard Pierre G., Li Cecilia, Stifter Sebastian A., Feng Carl G., et al. 2017. “Pho4 Is Essential for Dissemination of Cryptococcus Neoformans to the Host Brain by Promoting Phosphate Uptake and Growth at Alkaline pH.” mSphere 2 (1). 10.1128/mSphere.00381-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liao Yang, Smyth Gordon K., and Shi Wei. 2014. “featureCounts: An Efficient General Purpose Program for Assigning Sequence Reads to Genomic Features.” Bioinformatics 30 (7): 923–30. [DOI] [PubMed] [Google Scholar]
- Litvintseva Anastasia P., Carbone Ignazio, Rossouw Jenny, Thakur Rameshwari, Govender Nelesh P., and Mitchell Thomas G.. 2011. “Evidence That the Human Pathogenic Fungus Cryptococcus Neoformans Var. Grubii May Have Evolved in Africa.” PloS One 6 (5): e19688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Oliver W., Chun Cheryl D., Chow Eric D., Chen Changbin, Madhani Hiten D., and Noble Suzanne M.. 2008. “Systematic Genetic Analysis of Virulence in the Human Fungal Pathogen Cryptococcus Neoformans.” Cell 135 (1): 174–88. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Love Michael I., Huber Wolfgang, and Anders Simon. 2014. “Moderated Estimation of Fold Change and Dispersion for RNA-Seq Data with DESeq2.” Genome Biology 15 (12): 550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lowry J. A., and Atchley W. R.. 2000. “Molecular Evolution of the GATA Family of Transcription Factors: Conservation within the DNA-Binding Domain.” Journal of Molecular Evolution 50 (2): 103–15. [DOI] [PubMed] [Google Scholar]
- Lu Yang, Su Chang, and Liu Haoping. 2012. “A GATA Transcription Factor Recruits Hda1 in Response to Reduced Tor1 Signaling to Establish a Hyphal Chromatin State in Candida Albicans.” PLoS Pathogens 8 (4): e1002663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin Marcel. 2011. “Cutadapt Removes Adapter Sequences from High-Throughput Sequencing Reads.” EMBnet.journal 17 (1): 10–12. [Google Scholar]
- May Robin C., Stone Neil R. H., Wiesner Darin L., Bicanic Tihana, and Nielsen Kirsten. 2016. “Cryptococcus: From Environmental Saprophyte to Global Pathogen.” Nature Reviews. Microbiology 14 (2): 106–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Michl Johanna, Park Kyung Chan, and Swietach Pawel. 2019. “Evidence-Based Guidelines for Controlling pH in Mammalian Live-Cell Culture Systems.” Communications Biology 2 (April): 144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mi Huaiyu, Muruganujan Anushya, Ebert Dustin, Huang Xiaosong, and Thomas Paul D.. 2019. “PANTHER Version 14: More Genomes, a New PANTHER GO-Slim and Improvements in Enrichment Analysis Tools.” Nucleic Acids Research 47 (D1): D419–26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Minh Bui Quang, Schmidt Heiko A., Chernomor Olga, Schrempf Dominik, Woodhams Michael D., von Haeseler Arndt, and Lanfear Robert. 2020. “IQ-TREE 2: New Models and Efficient Methods for Phylogenetic Inference in the Genomic Era.” Molecular Biology and Evolution 37 (5): 1530–34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Missall Tricia Ann, Pusateri Mary Ellen, and Lodge Jennifer K.. 2004. “Thiol Peroxidase Is Critical for Virulence and Resistance to Nitric Oxide and Peroxide in the Fungal Pathogen, Cryptococcus Neoformans.” Molecular Microbiology 51 (5): 1447–58. [DOI] [PubMed] [Google Scholar]
- Mogensen Estelle Geweiss, Janbon Guilhem, Chaloupka James, Steegborn Clemens, Fu Man Shun, Moyrand Frédérique, Klengel Torsten, et al. 2006. “Cryptococcus Neoformans Senses CO2 through the Carbonic Anhydrase Can2 and the Adenylyl Cyclase Cac1.” Eukaryotic Cell 5 (1): 103–11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muñoz-Guzmán Felipe, Caballero Valeria, and Larrondo Luis F.. 2021. “A Global Search for Novel Transcription Factors Impacting the Neurospora Crassa Circadian Clock.” G3, April. 10.1093/g3journal/jkab100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nguyen Nga Thi Thuy, Vincens Pierre, Crollius Hugues Roest, and Louis Alexandra. 2018. “Genomicus 2018: Karyotype Evolutionary Trees and on-the-Fly Synteny Computing.” Nucleic Acids Research 46 (D1): D816–22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nielsen Kirsten, Cox Gary M., Wang Ping, Toffaletti Dena L., Perfect John R., and Heitman Joseph. 2003. “Sexual Cycle of Cryptococcus Neoformans Var. Grubii and Virulence of Congenic a and Alpha Isolates.” Infection and Immunity 71 (9): 4831–41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nielsen Kirsten, De Obaldia Anna L., and Heitman Joseph. 2007. “Cryptococcus Neoformans Mates on Pigeon Guano: Implications for the Realized Ecological Niche and Globalization.” Eukaryotic Cell 6 (6): 949–59. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O’Meara Teresa R., and Alspaugh J. Andrew. 2012. “The Cryptococcus Neoformans Capsule: A Sword and a Shield.” Clinical Microbiology Reviews 25 (3): 387–408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O’Meara Teresa R., Holmer Stephanie M., Selvig Kyla, Dietrich Fred, and Alspaugh J. Andrew. 2013. “Cryptococcus Neoformans Rim101 Is Associated with Cell Wall Remodeling and Evasion of the Host Immune Responses.” mBio 4 (1). 10.1128/mBio.00522-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O’Meara Teresa R., Norton Diana, Price Michael S., Hay Christie, Clements Meredith F., Nichols Connie B., and Alspaugh J. Andrew. 2010. “Interaction of Cryptococcus Neoformans Rim101 and Protein Kinase A Regulates Capsule.” PLoS Pathogens 6 (2): e1000776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ost Kyla S., O’Meara Teresa R., Huda Naureen, Esher Shannon K., and Alspaugh J. Andrew. 2015. “The Cryptococcus Neoformans Alkaline Response Pathway: Identification of a Novel Rim Pathway Activator.” PLoS Genetics 11 (4): e1005159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park Benjamin J., Wannemuehler Kathleen A., Marston Barbara J., Govender Nelesh, Pappas Peter G., and Chiller Tom M.. 2009. “Estimation of the Current Global Burden of Cryptococcal Meningitis among Persons Living with HIV/AIDS.” AIDS 23 (4): 525–30. [DOI] [PubMed] [Google Scholar]
- Perfect John R. 2006. “Cryptococcus Neoformans: The Yeast That Likes It Hot.” FEMS Yeast Research 6 (4): 463–68. [DOI] [PubMed] [Google Scholar]
- Perfect J. R., Lang S. D., and Durack D. T.. 1980. “Chronic Cryptococcal Meningitis: A New Experimental Model in Rabbits.” The American Journal of Pathology 101 (1): 177–94. [PMC free article] [PubMed] [Google Scholar]
- Powell K. E., Dahl B. A., Weeks R. J., and Tosh F. E.. 1972. “Airborne Cryptococcus Neoformans: Particles from Pigeon Excreta Compatible with Alveolar Deposition.” The Journal of Infectious Diseases 125 (4): 412–15. [DOI] [PubMed] [Google Scholar]
- Rajasingham Radha, Govender Nelesh P., Jordan Alexander, Loyse Angela, Shroufi Amir, Denning David W., Meya David B., Chiller Tom M., and Boulware David R.. 2022. “The Global Burden of HIV-Associated Cryptococcal Infection in Adults in 2020: A Modelling Analysis.” The Lancet Infectious Diseases 22 (12): 1748–55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- R Core Team. 2023. “R: A Language and Environment for Statistical Computing.” /ra-Language-and-Environment-Forstatistical-Computing. Vienna, Austria: R Foundation for Statistical Computing. https://www.R-project.org/. [Google Scholar]
- Sayers Eric W., Bolton Evan E., Brister J. Rodney, Canese Kathi, Chan Jessica, Comeau Donald C., Connor Ryan, et al. 2022. “Database Resources of the National Center for Biotechnology Information.” Nucleic Acids Research 50 (D1): D20–26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Selvig Kyla, and Alspaugh J. Andrew. 2011. “pH Response Pathways in Fungi: Adapting to Host-Derived and Environmental Signals.” Mycobiology 39 (4): 249–56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shishkin Alexander A., Giannoukos Georgia, Kucukural Alper, Ciulla Dawn, Busby Michele, Surka Christine, Chen Jenny, et al. 2015. “Simultaneous Generation of Many RNA-Seq Libraries in a Single Reaction.” Nature Methods 12 (4): 323–25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Springer Deborah J., Mohan Rajinikanth, and Heitman Joseph. 2017. “Plants Promote Mating and Dispersal of the Human Pathogenic Fungus Cryptococcus.” PloS One 12 (2): e0171695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steen Barbara R., Lian Tian, Zuyderduyn Scott, MacDonald William Kim, Marra Marco, Jones Steven J. M., and Kronstad James W.. 2002. “Temperature-Regulated Transcription in the Pathogenic Fungus Cryptococcus Neoformans.” Genome Research 12 (9): 1386–1400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Su Chang, Yu Jing, Sun Qiangqiang, Liu Qian, and Lu Yang. 2018. “Hyphal Induction under the Condition without Inoculation in Candida Albicans Is Triggered by Brg1-Mediated Removal of NRG1 Inhibition.” Molecular Microbiology 108 (4): 410–23. [DOI] [PubMed] [Google Scholar]
- Szewczyk Edyta, and Krappmann Sven. 2010. “Conserved Regulators of Mating Are Essential for Aspergillus Fumigatus Cleistothecium Formation.” Eukaryotic Cell 9 (5): 774–83. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Varadi Mihaly, Anyango Stephen, Deshpande Mandar, Nair Sreenath, Natassia Cindy, Yordanova Galabina, Yuan David, et al. 2022. “AlphaFold Protein Structure Database: Massively Expanding the Structural Coverage of Protein-Sequence Space with High-Accuracy Models.” Nucleic Acids Research 50 (D1): D439–44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Velagapudi Rajesh, Hsueh Yen-Ping, Geunes-Boyer Scarlett, Wright Jo Rae, and Heitman Joseph. 2009. “Spores as Infectious Propagules of Cryptococcus Neoformans.” Infection and Immunity 77 (10): 4345–55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wallace Edward W. J., and Haynes Samuel J.. 2022. “Tidyqpcr: Quantitative PCR Analysis in the Tidyverse.” Journal of Open Source Software 7 (74): 4507. [Google Scholar]
- Wallace Edward W. J., Maufrais Corinne, Sales-Lee Jade, Tuck Laura R., de Oliveira Luciana, Feuerbach Frank, Moyrand Frédérique, Natarajan Prashanthi, Madhani Hiten D., and Janbon Guilhem. 2020. “Quantitative Global Studies Reveal Differential Translational Control by Start Codon Context across the Fungal Kingdom.” Nucleic Acids Research 48 (5): 2312–31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walsh Naomi M., Botts Michael R., McDermott Andrew J., Ortiz Sébastien C., Wüthrich Marcel, Klein Bruce, and Hull Christina M.. 2019. “Infectious Particle Identity Determines Dissemination and Disease Outcome for the Inhaled Human Fungal Pathogen Cryptococcus.” PLoS Pathogens 15 (6): e1007777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weisman Caroline M., Murray Andrew W., and Eddy Sean R.. 2020. “Many, but Not All, Lineage-Specific Genes Can Be Explained by Homology Detection Failure.” PLoS Biology 18 (11): e3000862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wheeler K. A., Hurdman B. F., and Pitt J. I.. 1991. “Influence of pH on the Growth of Some Toxigenic Species of Aspergillus, Penicillium and Fusarium.” International Journal of Food Microbiology 12 (2–3): 141–49. [DOI] [PubMed] [Google Scholar]
- Wickham Hadley. 2009. ggplot2: Elegant Graphics for Data Analysis. Springer Science & Business Media. [Google Scholar]
- Wickham Hadley, Averick Mara, Bryan Jennifer, Chang Winston, Lucy McGowan Romain François, Grolemund Garrett, et al. 2019. “Welcome to the Tidyverse.” Journal of Open Source Software 4 (43): 1686. [Google Scholar]
- Williamson P. R. 1997. “Laccase and Melanin in the Pathogenesis of Cryptococcus Neoformans.” Frontiers in Bioscience: A Journal and Virtual Library 2 (November): e99–107. [DOI] [PubMed] [Google Scholar]
- World Health Organization. 2022. WHO Fungal Priority Pathogens List to Guide Research, Development and Public Health Action. Geneva: World Health Organization. [Google Scholar]
- Yamaguchi Masashi, Ohkusu Misako, Biswas Sondip Kumar, and Kawamoto Susumu. 2007. “Cytological Study of Cell Cycle of the Pathogenic Yeast Cryptococcus Neoformans.” Nihon Ishinkin Gakkai Zasshi = Japanese Journal of Medical Mycology 48 (4): 147–52. [DOI] [PubMed] [Google Scholar]
- Zaragoza Oscar, Fries Bettina C., and Casadevall Arturo. 2003. “Induction of Capsule Growth in Cryptococcus Neoformans by Mammalian Serum and CO(2).” Infection and Immunity 71 (11): 6155–64. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.