Abstract
Esophageal cancer is one of the major life-threatening diseases in the world. RNA methylation is the most common post-transcriptional modification and a wide-ranging regulatory system controlling gene expression. Numerous studies have revealed that dysregulation of RNA methylation is critical for cancer development and progression. However, the diverse role of RNA methylation and its regulators in esophageal cancer remains to be elucidated and summarized. In this review, we focus on the regulation of major RNA methylation, including m 6A, m 5C, and m 7G, as well as the expression patterns and clinical implications of its regulators in esophageal cancer. We systematically summarize how these RNA modifications affect the “life cycle” of target RNAs, including mRNA, microRNA, long non-coding RNA, and tRNA. The downstream signaling pathways associated with RNA methylation during the development and treatment of esophageal cancer are also discussed in detail. Further studies on how these modifications function together in the microenvironment of esophageal cancer will draw a clearer picture of the clinical application of novel and specific therapeutic strategies.
Keywords: esophageal cancer, RNA methylation, post-transcriptional modification, N6-methyladenosine, 5-methylcytosine, 7-methylguanosine, tumor microenvironment
Introduction
Esophageal cancer represents a significant global health problem and is one of the most common causes of cancer-related deaths worldwide, accounting for approximately 600,000 new cases and over 500,000 deaths in 2020 [1]. Although the geographic variation in the incidence of esophageal cancer substantially differs between the two most common histologic subtypes, i.e., esophageal squamous cell carcinoma (ESCC) and esophageal adenocarcinoma (EAC), most esophageal cancers occur in Asia, particularly in China, with ESCC accounting for the majority [ 2, 3] . In addition, the incidence and mortality rate of esophageal cancer are 2‒3 times higher in men than in women. As a result, it is of great significance to understand the specific mechanism underlying the progression of esophageal cancer and to find new therapeutic targets for cancer treatment and prognosis evaluation.
Epigenetic modifications are usually reversible and dynamically regulated in eukaryotic cells. It involves regulating gene expression without altering the nucleotide sequence [4], including DNA methylation, histone modifications, chromatin remodeling, and RNA modifications [ 5– 7] . These modifications drive transcriptional and post-transcriptional programs, exerting significant impacts on cellular functions, embryonic development, and diseases [ 8, 9] . RNAs can be modified post-transcriptionally by over 170 modifications [10]. Due to RNA modifications, RNA no longer simply acts as a participant or intermediate product in the protein synthesis process. RNA modifications affect the life cycle of various RNAs and thus become a key component of the gene expression regulatory system, profoundly influencing the process of gene expression [11].
Accounting for 60% of all RNA modifications, RNA methylation is the most studied, which includes N6-methyladenosine (m 6A), 5-methylcytosine (m 5C), N7-methylguanosine (m 7G), and N1-methyladenosine (m 1A) [12]. The presence of m 6A was first established in the 1970s [13]. It refers to methylation at the N6 position of adenosine in various RNAs, including mRNA, tRNA, rRNA, microRNA (miRNA), long noncoding RNA (lncRNA), and circular RNA (circRNA) [ 14, 15] . m 6A is the most common RNA methylation and is regulated by methyltransferases (“writers”), demethylases (“erasers”), and reader proteins (“readers”). m 6A methylation is catalyzed by the m 6A writer complex, which includes methyltransferase-like 3/14/16 (METTL3/14/16), WT1 associated protein (WTAP), vir like m6A methyltransferase associated (KIAA1429), RNA binding motif protein 15/15B (RBM15/15B), CBLL1, and ZC3H13. Demethylases act as erasers to remove m 6A modifications in RNA, including fat-mass and obesity-associated protein (FTO) and a-ketoglutarate-dependent dioxygenase alkB homolog 5 (ALKBH5). Reader proteins can recognize and bind to m6A-modified RNA, thereby regulating the stability, splicing, export, translation, and biogenesis of RNA [ 16– 20] . Readers include YTH domain-containing family protein 1/2/3 (YTHDF1/2/3), YTH domain containing 1/2 (YTHDC1/2), insulin-like growth factor 2 mRNA-binding protein 1/2/3 (IGF2BP1/2/3), and heterogeneous nuclear ribonucleoprotein C/G/A2B1 (HNRNPC/G/A2B1) ( Table 1) [21].
Table 1 RNA methylation regulators in esophageal cancer
|
RNA methylation type |
Category |
RNA methylation regulator |
Expression in esophageal cancer* |
Prognostic value in esophageal cancer* |
|
m 6A |
Writers |
METTL3 |
Upregulated |
Unfavorable |
|
METTL14 |
Downregulated |
Favorable |
||
|
METTL16 |
|
|||
|
WTAP |
Upregulated |
Unfavorable |
||
|
KIAA1429 |
|
|||
|
RBM15/15B |
|
|||
|
CBLL1 |
|
|||
|
ZC3H13 |
|
|||
|
Erasers |
FTO |
Upregulated |
Unfavorable |
|
|
ALKBH5 |
Downregulated |
Favorable |
||
|
Readers |
YTHDF1/2/3 |
|
||
|
YTHDC1/2 |
|
|||
|
IGF2BP2 |
Upregulated |
|
||
|
IGF2BP3 |
Upregulated |
Unfavorable |
||
|
IGF2BP1 |
|
|||
|
HNRNPC |
Upregulated |
Unfavorable |
||
|
HNRNPA2B1 |
Upregulated |
Unfavorable |
||
|
HNRNPG |
|
|||
|
m 5C |
Writers |
NSUN2 |
Upregulated |
Unfavorable |
|
NSUN1/3/4/5/6/7 |
|
|||
|
DNMT2 |
|
|||
|
Erasers |
TET1 |
Downregulated |
|
|
|
TET2 |
Downregulated |
|
||
|
TET3 |
Upregulated |
Unfavorable |
||
|
ALKBH1 |
|
|||
|
Readers |
YBX1 |
|
||
|
ALYREF |
|
|||
|
m 7G |
Writers |
METTL1 |
Upregulated |
Unfavorable |
|
WDR4 |
Upregulated |
|
||
|
WBSCR22 |
Unfavorable |
|||
|
TRMT112 |
|
|||
|
RNMT |
|
|||
|
RAM |
|
*The expression and prognostic value of these RNA regulators have been experimentally validated.
First discovered in 1950, m 5C is a conserved and prevalent mark in RNA in all life domains. It refers to a methyl group inserted into the carbon atom at the fifth position of cytidine [22]. Similar to m 6A, m 5C is regulated by writer, eraser, and reader proteins. Methyltransferases of m 5C include DNA methyltransferase 2 (DNMT2) and NOP2/Sun RNA methyltransferase family member 1-7 (NSUN1-7) [ 23, 24] . Demethylases include the Ten-eleven translocation 1-3 (TET1-3) and ALKBH1 [ 25, 26] . m 5C reader proteins include Y-box binding protein 1 (YBX1) and Aly/REF export factor (ALYREF), which are associated with the nuclear export and stability of methylated RNA ( Table 1) [27].
m 7G methylation can mediate the metabolism and function of RNA and act as a molecule handler to alter the expression of target genes [28]. m 7G is usually located at the 5′ caps and internal positions within eukaryotic mRNA or the inside of rRNA and tRNA of all species [ 29– 31] . The methyltransferase-like 1/ WD repeat domain 4 (METTL1/WDR4) complex and Williams-Beuren syndrome chromosome region 22/ tRNA methyltransferase activator subunit 11-2 (WBSCR22/TRMT112) complex function as m 7G writers [32]. RNA guanine-7-methyltransferase (RNMT) and RNMT-activating mini protein (RAM) bind to each other to install m 7G modifications ( Table 1) [33].
As research progressed, more and more studies revealed the key role of RNA methylation in regulating the fate and function of mRNAs. Dysregulation in RNA methylation is critical for cancer development and progression [8]. In this study, we reviewed and summarized the emerging role of major types of RNA methylation in esophageal cancer, including m 6A, m 5C, and m 7G. We highlighted the expression, clinical implications, and biological functions of their regulators and aimed to improve the understanding of RNA methylation’s role and clinical application in esophageal cancer ( Tables 1 and 2).
Table 2 Functional roles of RNA methylation and their regulators in esophageal cancer
|
RNA methylation type |
RNA methylation regulator |
Target RNA |
Downstream molecular mechanism |
Phenotype |
Role in esophageal cancer |
Reference |
|
m 6A |
METTL3, YTHDF2 |
APC |
Wnt/β-catenin pathway |
Aerobic glycolysis, tumorigenesis |
Oncogene |
|
|
METTL3, YTHDF3 |
EGR1 |
Snail signaling |
Metastasis |
Oncogene |
||
|
METTL3 |
TNFR1 |
MAPK & NF-κB pathway |
Tumorigenesis, proliferation, migration, invasion |
Oncogene |
||
|
METTL3 |
GAS5 |
miR3912-5p/RALYL |
Tumorigenesis |
Oncogene |
||
|
METTL3 |
pri-miR-20a-5p |
miR-20a-5p/NFIC |
Metastasis, invasion, migration |
Oncogene |
||
|
METTL3 |
NOTCH1 |
Notch signaling pathway |
Invasion, migration, proliferation, tumorigenesis |
Oncogene |
||
|
METTL3 |
GLS2 |
Metastasis |
Oncogene |
|||
|
METTL3 |
AKT signaling pathway |
Proliferation, invasion |
Oncogene |
|||
|
METTL3 |
p21-axis |
Proliferation |
Oncogene |
|||
|
METTL3 |
COL12A1/MAPK signaling pathway |
Tumorigenesis |
Oncogene |
|||
|
METTL3 |
PI3K/AKT signaling pathway |
Proliferation, invasion, migration |
Oncogene |
|||
|
METTL14 |
miR-99a-5p |
TRIB2/HDAC2/AKT/mTOR/S6K1 |
Stemness, radioresistance |
Oncogene |
||
|
WTAP, YTHDF2 |
CPSF4 |
Proliferation, invasion, migration |
Oncogene |
|||
|
WTAP |
ki67 and Snail |
Proliferation, migration |
Oncogene |
|||
|
ALKBH5 |
CASC8 |
hnRNPL/Bcl2/caspase3 pathway |
Proliferation, chemoresistance |
Oncogene |
||
|
ALKBH5 |
pri-miR-194-2 |
RAI1/Hippo signaling pathway |
Tumorigenesis, metastasis, migration |
Tumor suppressor gene |
||
|
ALKBH5 |
LINC00278 |
AR signaling pathway |
Proliferation |
Oncogene |
||
|
ALKBH5 |
miR-193a-3p |
Proliferation, migration, invasion |
Tumor suppressor gene |
|||
|
ALKBH5 |
Proliferation, invasion |
Tumor suppressor gene |
||||
|
ALKBH5 |
Migration, invasion, tumorigenesis |
Tumor suppressor gene |
||||
|
FTO, YTHDF2 |
LINC00022 |
p21 |
Tumorigenesis |
Oncogene |
||
|
FTO, YTHDF1 |
HSD17B11 |
Lipid metabolism |
Tumorigenesis, migration, invasion |
Oncogene |
||
|
FTO, YTHDF1 |
ERBB2 |
Metastasis, tumorigenesis |
Oncogene |
|||
|
FTO |
MMP13 |
Proliferation, migration |
Oncogene |
|||
|
YTHDF1 |
HK2 |
Aerobic glycolysis |
Oncogene |
|||
|
IGF2BP2 |
TK1 |
Migration, invasion |
Oncogene |
|||
|
HNRNPA2B1 |
ACLY/ACC1 |
Proliferation, invasion, migration |
Oncogene |
|||
|
YTHDC2 |
Several cancer-related signaling pathways |
Proliferation |
Tumor suppressor gene |
|||
|
m 5C |
NSUN2 |
NMR |
BPTF/MMP3/MMP10 |
Chemoresistance, migration, invasion, proliferation |
Oncogene |
|
|
NSUN2 |
GRB2 |
PI3K/AKT and ERK/MAPK |
Tumorigenesis, invasion, migration |
Oncogene |
||
|
NSUN2 |
Cancer-related genes |
Proliferation, radiochemotherapy resistance |
Oncogene |
|||
|
YBX1 |
c-Myc |
Proliferation, metastasis |
Oncogene |
|||
|
TET3 |
HOXB2/c-MYC/NANOG pathway |
Stemness |
Oncogene |
|||
|
TET2 |
Proliferation, migration |
Tumor suppressor gene |
||||
|
TET1 |
Migration, invasion |
Tumor suppressor gene |
||||
|
TET1 |
Proliferation |
Tumor suppressor gene |
||||
|
m 7G |
METTL1/WDR4 |
tRNA |
RPTOR-pULK1 |
Proliferation, tumorigenesis |
Oncogene |
Role of m 6A in Esophageal Cancer
Expression and clinical significance of m 6A regulators
m 6A methylation within mRNA is increased overall in ESCC tissues compared with that in paracancerous tissues [36]. The mRNA and protein levels of METTL3, an m 6A writer, are consistently upregulated in esophageal cancer tissues [ 34, 35, 71] . Emerging evidence has shown that patients with high METTL3 expression have poorer overall survival than those with low METTL3 expression. METTL14 is downregulated in ESCC, and low METTL14 abundance predicts unfavorable overall survival in ESCC patients [45]. WTAP expression is significantly elevated in ESCC tissues compared to that in paracancerous tissues. It is strongly associated with high tumor-node-metastasis (TNM) stage and poor prognosis [47]. High RBM15 and KIAA1429 expressions are strongly correlated with better survival of ESCC patients by bioinformatics analysis [72] ( Table 1).
For m 6A readers, HNRNPA2B1 and HNRNPC are both upregulated in ESCC tissues [ 73, 74] . High expression of HNRNPA2B1 is closely associated with distant metastasis, higher lymph node stage, and worse overall survival in ESCC patients, and high expression of HNRNPC suggests a poor prognosis. By analyzing data from TCGA, researchers demonstrated that YTHDF1-3 expression is higher in esophageal cancer samples than in normal samples and that YTHDF1 is significantly associated with multiple clinical features of patients [ 75– 77] . Yang et al. [61] found that YTHDC2 mRNA level is significantly lower in ESCC tumor tissues than in paired normal tissues by analyzing RNA-seq data. The protein expressions of IGF2BP2 and IGF2BP3 are substantially elevated in ESCC tissues compared with those in normal tissues [ 78, 79] . High IGF2BP2 expression is strongly associated with high TNM stage, lymphatic infiltration, and lymph node metastasis. High IGF2BP3 expression level also correlates with poor prognosis in ESCC. Researchers also constructed prognostic signatures consisting of HNRNPC/ G/ A2B1 and ALKBH5 based on the TCGA ESCA cohort, which are strongly associated with esophageal cancer outcomes [ 73, 76] ( Table 1).
For m 6A erasers, FTO expression in esophageal cancer tissues is significantly increased compared to that in paracancerous tissues. The overall survival of patients with high FTO expression is significantly reduced compared to that of patients with low expression [ 42, 55] . ALKBH5 is considered to be a favorable prognostic factor for patients and is downregulated in ESCC tissue [ 49, 51, 53] . High ALKBH5 expression also indicates less local lymph node metastasis, lower TNM stage, and less recurrence after surgery [49] ( Table 1).
Expression modulation of m 6A regulators
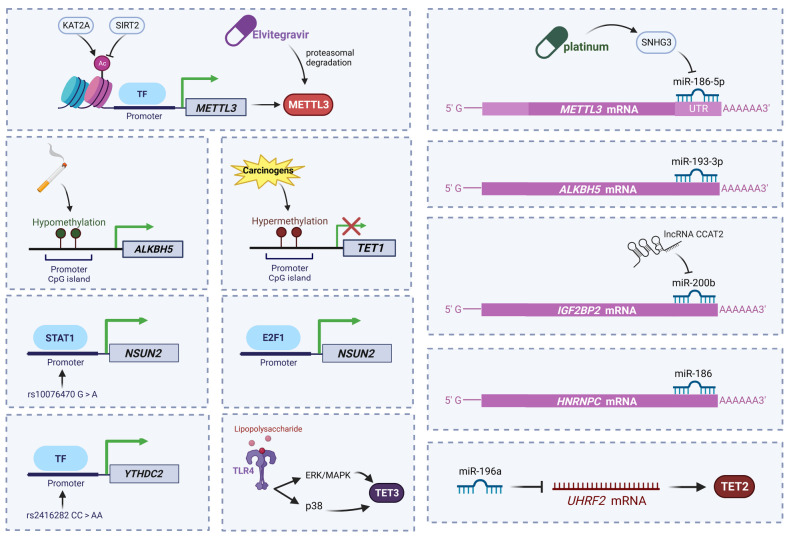
The expression of m 6A regulators is modulated through multiple ways in esophageal cancer. It has been reported that the histone acetylase KAT2A mediates H3K27 acetylation in the promoter region of METTL3, promoting METTL3 transcription. In contrast, the deacetylase SIRT2 acts as a suppressor of METTL3 expression [35]. In addition, this study identified a novel use of the antiretroviral drug elvitegravir through molecular conformational analysis of METTL3. The study found that elvitegravir can target METTL3 and promote STUB1-mediated proteasomal degradation. In lung cancer cells, METTL3 expression can be suppressed by miRNAs such as miR-338-5p [80]. Similarly, in esophageal cancer, Zhang et al. [81] demonstrated that miR-186-5p acts as a negative regulator of METTL3 by binding to the 3′ untranslated region (UTR) of mRNA. In addition, platinum treatment upregulates SNHG3 expression and causes its interaction with miR-186-5p to suppress the expression of the latter, which in turn upregulates METTL3 expression. Wu et al. [50] found that ALKBH5 promoter CpG island is hypomethylated after cigarette smoke condensate treatment, leading to upregulated ALKBH5 expression in esophageal cells. Xue et al. [51] elucidated that miR-193-3p targets ALKBH5 and suppresses its expression. Yang et al. [61] identified rs2416282 as a single nucleotide polymorphism in the promoter of YTHDC2 by analyzing genome-wide association studies. rs2416282 is an expression quantitative trait locus (eQTL) of YTHDC2 and can impair the transcriptional activity of YTHDC2. LncRNA CCAT2 can bind to miR-200b and reduce its expression. As an m 6A reader, IGF2BP2 is a target of miR-200b, and inhibition of miR-200b results in the upregulation of IGF2BP2 in esophageal cancer tissues [59]. According to the analysis of the ciBioPortal database, most m 6A regulators are amplified, deleted, and mutated in esophageal cancer, among which IGF2BP2 has the highest incidence rate (11%) [77]. Li et al. [74] demonstrated the potential binding site of miR-186 in HNRNPC. miR-186 can target HNRNPC and suppress its expression ( Figure 1).
Figure 1 .
Expression modulation of RNA methylation regulators in esophageal cancer
The expressions of m 6A regulators METTL3, ALKBH5, IGF2BP2, HNRNPC, and YTHDC2, and m 5C regulators NSUN2 and TET1-3 are modulated by drugs, carcinogens, transcription factors (TFs), miRNAs and many other modalities.
Biological function of m 6A
As an m 6A reader, METTL3 can upregulate the m 6A level of APC mRNA, which in turn recruits YTHDF2 and promotes the degradation of APC mRNA [34]. Downregulation of APC promotes aerobic glycolysis and in vivo tumorigenesis in esophageal cancer. METTL3 upregulates the m 6A level of EGR1, which in turn promotes the stability of EGR1 mRNA through YTHDF3 [35]. The subsequent activation of the EGR1/snail pathway promotes esophageal cancer metastasis. Furthermore, METTL3-mediated m 6A methylation of pri-miR-20a-5p induces its binding with DGCR8 [38]. DGCR8 binding promotes the maturation of miR-20a-5p and consequently promotes epithelial-mesenchymal transition (EMT), invasion, and metastasis in esophageal cancer. Using human m 6A epitranscriptomic microarrays, Jia et al. [37] found that m 6A-modified lncRNA GAS5 competitively absorbs miR-3912-5p and that RALYL is the downstream molecule of miR-3912-5p in ESCC. As a result, they presumed that METTL3 promotes the pathogenesis of ESCC by mediating m 6A methylation of lncRNA GAS5. Li et al. [36] revealed that TNFR1 is also a target gene of METTL3. ATXN2 binds to m 6A-modified TNFR1 mRNA and enhances translation of TNFR1, which in turn promotes tumor progression. Han et al. [39] found that METTL3 could prevent the degradation of NOTCH1 mRNA by mediating its m 6A modification. The upregulation of NOTCH1 activates the Notch signaling pathway and promotes ESCC initiation and progression. METTL3 promotes COL12A1 expression to activate the MAPK signaling pathway, but its exact mechanism is unclear [43]. METTL3 also promotes GLS2 expression by increasing m 6A methylation level at the 3′UTRs, resulting in a malignant phenotype of the cells [40]. METTL3 activates AKT signaling pathways and promotes the invasion and proliferation of esophageal cancer cells [41]. In contrast, silencing of METTL3 suppresses the malignant phenotypes of ESCC cells via downregulating PI3K/AKT signaling activity [44]. In 2021, Liu et al. [45] highlighted that METTL14/miR-99a-5p/TRIB2 constitutes a positive feedback loop and leads to radioresistance in ESCC. They found that METTL14 mediates m 6A modification of pri-miR-99a and promotes DGCR8-dependent processing, which upregulates miR-99a-5p expression. miR-99a-5p in turn induces TRIB2 mRNA degradation, while TRIB2 induces ubiquitin-mediated proteasomal degradation of METTL14. After WTAP inhibition, the cell proliferation and migration indicators, ki67 and Snail, are downregulated, while the apoptosis-associated proteins, Bax and Caspase-3, are upregulated [47]. Luo et al. [46] identified that WTAP-based m 6A modification of CPSF4 mRNA reduces the stability of CPSF4 by YTHDF2 binding ( Table 2, Figure 2).
Figure 2 .
Diverse effect of m 6A writers on the RNA life cycle and cellular function in esophageal cancer
m 6A writers METTL3, METTL14, and WTAP mediate m 6A modification of target RNAs and promote esophageal cancer progression.
As an m 6A demethylase, FTO was found to demethylate LINC00022 in esophageal carcinoma, leading to the inhibition of YTHDF2-mediated degradation of m 6A-modified LINC00022. This process in turn promotes the ubiquitinated degradation of the cell cycle inhibitor protein p21 and induces tumor proliferation [54]. Interestingly, another study found that as an m 6A methyltransferase, the expression of METTL3 also negatively correlates with the expression of p21 in ESCC, implying that METTL3 may modulate the cell cycle in a p21-dependent manner, but the exact mechanism involved is unclear [42]. FTO can bind to HSD17B11 mRNA and reduce its m 6A methylation level while relying on YTHDF1 to affect its translation. Elevated HSD17B11 level then promotes lipid droplet formation and facilitates the progression of esophageal carcinoma [55]. Moreover, FTO positively regulates the mRNA and protein levels of MMP13, which in turn promotes ESCC proliferation and migration [57]. Zhao et al. [56] found that FTO promotes esophageal cancer proliferation and metastasis through the demethylation of ERBB2 mRNA. Additionally, many studies have revealed the tumor-suppressive function of ALKBH5 in esophageal cancer. For example, Li et al. [52] found that overexpression of ALKBH5 reduces the overall m 6A methylation level in cells and inhibits malignant proliferation and invasion. Chen et al. [49] elucidated that the m 6A modification of pri-miR-194-2 allows its binding with DGCR8 and promotes its biogenesis. miR-194-2 subsequently suppresses RAI1 expression, resulting in inhibition of the Hippo pathway. However, the m 6A demethylase ALKBH5 can enhance the inhibitory effect of RAI1 on YAP/TAZ translocation to the nuclei by demethylating pri-miR-194-2, thereby suppressing the malignancy of esophageal cancer. Overexpression of ALKBH5 leads to G1-phase arrest and reduces ESCC migration and invasion, indicating that ALKBH5 is a tumor suppressor [53]. In addition, ALKBH5 is negatively correlated with miR-193a-3p expression and can also prevent DGCR8-mediated maturation of pri-miR-193a-3p by demethylating it, forming a positive feedback loop that leads to the growth and metastasis of ESCC [51]. Interestingly, another role of the ALKBH5-mediated m 6A demethylation pattern in tumors was found in 2020 [50]. In male ESCC patients, research has shown that smoking leads to the demethylation of LINC00278, which induces the activation of the androgen receptor signaling pathway and inhibits the apoptosis of ESCC cells. In 2022, Wu et al. [48] reported the pro-carcinogenic role of ALKBH5 in esophageal cancer. ALKBH5 mediates the m 6A demethylation and stabilization of CASC8, which promotes the proliferation and chemoresistance of ESCC ( Table 2, Figure 3A).
Figure 3 .
Diverse effect of m 6A erasers and readers on the RNA life cycle and cellular function in esophageal cancer
(A) m 6A erasers FTO and ALKBH5 mediate the demethylation of target RNAs and affect the biological function of esophageal cancer. (B) m 6A readers YTHDF1 and IGF2BP2 exert different effects on m 6A-modified RNAs and mediate ESCC progression.
As an m 6A reader, YTHDF1 can directly interact with HCP5 and enhance its own binding to m 6A-modified HK2 mRNA [58]. Subsequently, increased stability of HK2 mRNA leads to enhanced aerobic glycolysis in ESCC cells. HNRNPA2B1 positively correlates with m 6A level in esophageal cancer. It promotes lipid accumulation by upregulating the expression of the fatty acid synthesis enzymes ACLY and ACC1, thereby contributing to tumor progression [60]. Furthermore, HNRNPA2B1’s positive correlation with the miR-17-92 cluster suggests that HNRNPA2B1 may influence the prognosis of esophageal cancer through these microRNAs [73]. IGF2BP2 recognizes and binds to the m 6A modification site on TK1 mRNA and stabilizes the mRNA. Upregulation of TK1 then facilitates the development of ESCC [59]. Since YTHDC2 is a tumor suppressor of ESCC, impairment of its expression due to the genetic variant promotes the proliferation of ESCC cells by activating several cancer-related signaling pathways [61] ( Table 2, Figure 3B).
m 6A regulators are also involved in immune infiltration in esophageal cancer. For example, Guo et al. [82] revealed that PD-L1 expression in ESCC tissues increases significantly and is negatively correlated with the expressions of YTHDF2, METTL14, and KIAA1429. Pu et al. [72] developed a prognostic signature based on the expressions of YTHDF3, RBM15, KIAA1429, and ALKBH5 and found that the high-risk cohort is associated with a higher proportion of CD8 + T cells and higher expression of PD-L1. Using data from the TIMER database, Zhao et al. [77] found that WTAP positively correlates with the infiltration of cancer-associated fibroblasts, myeloid dendritic cells, CD4 + T cells, neutrophils, CD8 + T cells, and macrophages and negatively correlates with the infiltration of regulatory T cells. IGF2BP3 is positively correlated with the infiltration of cancer-associated fibroblasts and myeloid dendritic cells. YTHDC2 is significantly associated with the infiltration of CD4 + T cells, neutrophils, and CD8 + T cells. HNRNPA2B1 is positively associated with macrophage infiltration. KIAA1429 is positively associated with the infiltration of myeloid dendritic cells, cancer-associated fibroblasts, CD4 + T cells, and neutrophils. HNRNPC is positively associated with myeloid dendritic cells and cancer-associated fibroblasts and negatively correlates with neutrophil infiltration. Using the TCGA cohort and TIMER database, Liu et al. [75] found that YTHDF1 expression is positively correlated with B cell and macrophage infiltration and negatively associated with dendritic cell infiltration in esophageal cancer. By analyzing data from the TISIDB website, investigators concluded that immune infiltration of effective memory CD8 + T cells, natural killer cells, and neutrophils is highly related to METTL3 expression [83].
Several other studies have highlighted the role of m 6A in esophageal cancer. Chen et al. [84] analyzed the differentially expressed genes for m 6A and autophagy by using HADd and TCGA databases. The prognostic score model constructed using six m 6A-related autophagy genes ( ITPR1, MAP1LC3C, DAPK2, DIRAS3, TP53, and EIF2AK3) is an adverse prognostic factor for ESCC. By bioinformatics prediction, Qian et al. [85] revealed that circ-SLC7A5 has the maximum m 6A modification structure, indicating its potential in detecting ESCC. To explore the combination of computed tomography radiomic features and m 6A regulators for the characterization of esophageal cancer patients, Brancato et al. [86] performed radiogenomic analyses between clinical outcome-associated radiomic features and transcriptomic signatures of m 6A regulators. Body mass index assessment revealed that the expressions of METTL3, YTHDF1, YTHDF2, and YTHDC1 had a significant correlation with ten radiomic features, among which three belong to the LoG sigma group, and five belong to the wavelet group. For stage assessment, radiogenomic results revealed that the expressions of HRNPC, WTAP, METTL3, YTHDF1, YTHDF2, and YTHDC1 had a significant correlation with ten radiomic features of the wavelet group.
Role of m 5C in Esophageal Cancer
Expression and clinical significance of m 5C
Several studies have confirmed that the protein and mRNA expression of NSUN2 are elevated in esophageal cancer tissue [ 63, 87] . Patients with higher ESCC stages show higher NSUN2 expression than those with lower ESCC stages, and high NSUN2 expression indicates poor clinical outcomes. Li et al. [69] revealed that TET1 expression is downregulated in esophageal cancer. Shi et al. [88] found a significant decrease in RNA expressions of TET2 and TET3 in tumor tissues compared with those in paired non-tumor tissues. TET2 is epigenetically regulated in ESCC and is mutated in nearly 6% of the Japanese ESCC cohort [89]. According to this cohort, TET2 mutations can be considered negative prognostic indicators for patient survival. Interestingly, Xu et al. [66] reported that both RNA and protein levels of TET3 are elevated in ESCC tissues compared to those in paracancerous tissue, and TET3 expression negatively correlates with the survival of ESCC patients ( Table 1).
Expression modulation of m 5C regulators
Through screening of public databases and experimental validation, Su et al. [63] found that NSUN2 positively correlates with E2F1 expression and that the RNA and protein levels of NSUN2 are significantly downregulated after silencing of E2F1. Subsequently, the investigators demonstrated that E2F1 acts as a transcription factor (TF) for NSUN2, making its expression significantly upregulated in esophageal cancer. Researchers found an rs10076470 G to A mutation in the NSUN2 gene in some ESCC patients, which results in the formation of a cis-eQTL. The cis-eQTL leads to the binding of STAT1 as a transcription factor that promotes the expression of NSUN2 [64]. The TET1 promoter and exon 1 region contain a typical methylated CpG island, and CpG methylation induced by carcinogens most likely suppresses TET1 expression in esophageal cells [69]. Hu et al. [67] elucidated that TET2 expression can be downregulated by the miR-196a/UHRF2 axis. TET3 expression is elevated in ESCC through the TLR4/p38/ERK-MAPK pathway under lipopolysaccharide stimulation [66] ( Figure 1).
Biological function of m 5C
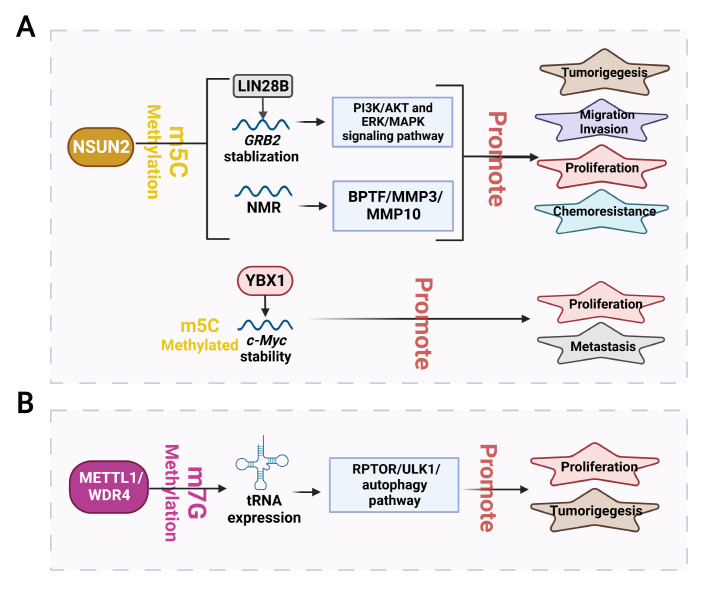
NSUN2 activates many cancer-related genes through mRNA m 5C methylation, leading to radiochemotherapy resistance in patients [64]. NSUN2 mediates the m 5C modification of GRB2 and induces the binding of LIN28B, a novel m 5C mediator other than YBX1 and ALYREF. LIN28B stabilizes GRB2 mRNA and consequently activates the PI3K/AKT and ERK/MAPK pathways [63]. Li et al. [62] found that lncRNA NMR is a methylation target of NSUN2 and plays various roles in esophageal cancer. m 5C-methylated NMR can increase chemoresistance in ESCC cells and bind to the chromatin regulator BPTF to mediate tumor progression. Interestingly, methylated NMR may also competitively inhibit the methylation of some mRNAs related to cell migration and invasion, such as LAMB1, PLOD3, HSPG2, and COL4A5 ( Table 2 and Figure 4A).
Figure 4 .
Diverse effect of m 5C and m 7G regulators on the RNA life cycle and cellular function in esophageal cancer
(A) m 5C writer NSUN2 and reader YBX1 exert different effects on m 5C-modified RNAs and mediate ESCC progression. (B) m 7G writers METTL1/WDR4 promote tRNA expression through m 7G modification and mediate ESCC progression.
YBX1 can bind to the 3′UTR of c-Myc mRNA with the help of the “scaffold” linc02042. The stability of c-Myc mRNA is subsequently enhanced, which leads to the proliferation and metastasis of ESCC [65]. Although the function of the TET family as RNA methylation regulators has not been discussed explicitly in published research, its relationship with esophageal cancer has already been investigated in several studies. LncRNA ZNF667-AS1 activates the expressions of its target genes, ZNF667 and E-cadherin, by interacting with and recruiting TET1 to the genes, thereby inhibiting ESCC progression [68]. Suppression of TET1 demethylase is correlated with low chromosomal copy-number alterations (CNAs) in HPV (+) ESCC [90]. Li et al. [69] elucidated that loss of TET1 expression promotes tumor suppressor gene inactivation in tumor pathogenesis. Knockdown of TET2 contributes to the malignant phenotypes of ESCC by enhancing cell proliferation, migration, and invasion [ 67, 89] . Moreover, overexpression of TET3 has been reported to induce the stemness of ESCC cells [66] ( Table 2 and Figure 4A).
Role of m 7G in Esophageal Cancer
In 2022, Han et al. [70] demonstrated that the m 7G methyltransferases METTL1 and WDR4 are consistently elevated in ESCC tissues and are both negative indicators for ESCC prognosis. METTL1/WDR4 regulates the m 7G modification and expression of tRNA, thereby affecting the translation efficiency of a large subset of mRNAs, including RPTOR, which upregulates the ULK1/autophagy pathway and mediates ESCC progression. High WBSCR22 expression predicts poor prognosis in ESCC [91]. Zhao et al. [92] developed a robust and potent m 7G regulator-related lncRNA prognostic signature that shows better predictive value than traditional clinical risk factors. Their results also revealed that the expression levels of m 7G-related lncRNAs AC025754.2, AL451165.2, and AL513550.1 were significantly elevated in ESCC tissue, while AC007566.1, SRP14-AS1, SNHG7 and HAND2-AS1 were downregulated in ESCC tissue ( Tables 1, 2 and Figure 4B).
Conclusions and Perspectives
In recent years, many studies have reported the dynamic role of RNA methylation in the pathology of esophageal cancer. In addition to m 6A, the novel roles of m 5C and m 7G are gradually being revealed. The biological functions of m 1A will also be elucidated in the future. In this review, we focused on the roles of m 6A, m 5C, and m 7G in esophageal cancer. First, we briefly introduced the three types of RNA methylation and their regulators. Then, we highlighted the function and clinical significance of RNA methylation regulators in esophageal cancer. Importantly, we also summarized the expression modulation of these regulators in esophageal cancer through a systematic review of published studies. In addition, the pathophysiological functions and dysregulation pathways of m 6A, m 5C, and m 7G in esophageal cancer were discussed in detail.
With the illustration of the role of RNA methylation in tumor progression, therapies targeting the methylation of RNA have become emerging targets for cancer treatment. The development of inhibitors or activators for RNA methylation regulators could provide valuable drug candidates and valuable experience in cancer management [93]. In 2021, STM2457, a highly specific and effective inhibitor of the methyltransferase METTL3, was discovered, which showed a significant therapeutic effect in a preclinical acute myeloid leukemia (AML) model [ 94, 95] . Using a hybrid virtual screening strategy, Du et al. [96] identified an METTL3 inhibitor from natural products and validated that it can inhibit the proliferation of tumor cells. Excitingly, STC-15, another METTL3 inhibitor, has been approved for a Phase 1, multi-center, open-label, and first-in-human study (NCT05584111). This study was designed to systematically assess the safety and tolerability, pharmacokinetics, pharmacodynamics, and clinical activity of STC-15 in adult subjects with advanced malignancies. In addition, Xie et al. [97] developed a specific inhibitor of the m 6A demethylase activity of FTO, namely 18097 which significantly decreases the overall m 6A level and suppresses the in vivo growth of breast cancer cells. Other inhibitors of FTO, including meclofenamic acid (MA), R-2HG and FB23-2, also exhibit good anti-tumor activity [ 98– 100] . Weng et al. [101] identified CWI1-2, a small-molecule compound targeting IGF2BP2, which shows promising potential in leukemia treatment. Li et al. [102] identified ALKBH5 as a potential therapeutic target in melanoma and colorectal cancer through the use of ALKBH5 inhibitor ALK-04. Fang et al. [103] discovered the compound 20m as a potent and specific inhibitor of ALKBH5. Although there is still a relatively long way to go before these drugs enter clinical use and their effectiveness in esophageal and many other malignancies has not been validated, we can foresee good potential for their future application.
Interestingly, it is worth noting that as an inhibitor of METTL3, STM2457 suppresses the development of AML and induces the differentiation and apoptosis of tumor cells [94]. FB23-2, an inhibitor of demethylase FTO, remarkably suppresses proliferation and stimulates the differentiation and apoptosis of AML cells [100]. Thus, methylation and demethylation sometimes seem to play conflicting roles. Furthermore, Cui et al. [98] reported that the growth and self-renewal are restrained after overexpressing METTL3 or inhibiting FTO in glioblastoma stem cells (GSCs), while Visvanathan et al. [104] reported that elevated METTL3 levels observed in GSCs are critical for GSC maintenance and radioresistance. The phenotypic variations could partly be interpreted by the different dependence on m 6A-modified RNAs and the genetic heterogeneity in cancer cells. However, we believe that there must be some balance between methylation and demethylation that affects the malignant phenotype.
Immunotherapy has become an effective and prominent treatment strategy for esophageal cancer, therefore, it is of great value to target RNA methylation in the tumor microenvironment of esophageal cancer. A growing body of evidence suggests that a thorough comprehension of the molecular mechanism of esophageal cancer calls for attention to not only tumor cells but also the tumor microenvironment [105]. Recently, researchers have revealed the critical role of m 6A in modulating tumor immune microenvironment [ 106– 108] . Li et al. [102] found that targeting m 6A may be involved in regulating anti-PD-1 therapy response by modulating immunosuppressive cell infiltration in melanoma. Liu et al. [107] revealed the possibility of improving the efficacy of anti-PD-1 treatment in NSCLC by targeting m 6A modification. Wan et al. [106] reported that m 6A modification of PD-L1 mRNA promotes its stability and inhibits immune surveillance in breast cancer. In addition, since an increasing number of studies have revealed the potential of targeting m 6A and m 5C to reverse chemo- and radio-resistance in esophageal cancer and other malignancies, targeting RNA methylation in esophageal cancer and thereby promoting the efficacy of various therapeutic modalities, such as immunotherapy, chemotherapy and radiotherapy, is promising and awaits future applications [ 48, 62, 64, 104, 109– 111] .
COMPETING INTERESTS
The authors declare that they have no conflict of interest.
Funding Statement
This work was supported by the grants from the National Natural Science Foundation of China (Nos. 82072557 and 81871882), the National Key Research and Development Program of China (No. 2021YFC2500900), the Shanghai Municipal Education Commission-Gaofeng Clinical Medicine Grant (No. 20172005, the 2nd round of disbursement), and the Program of Shanghai Academic Research Leader from Science and Technology Commission of Shanghai Municipality (No. 20XD1402300).
References
- 1.Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. . 2021;71:209–249. doi: 10.3322/caac.21660. [DOI] [PubMed] [Google Scholar]
- 2.Morgan E, Soerjomataram I, Rumgay H, Coleman HG, Thrift AP, Vignat J, Laversanne M, et al. The global landscape of esophageal squamous cell carcinoma and esophageal adenocarcinoma incidence and mortality in 2020 and projections to 2040: new estimates from GLOBOCAN 2020. Gastroenterology. . 2022;163:649–658.e2. doi: 10.1053/j.gastro.2022.05.054. [DOI] [PubMed] [Google Scholar]
- 3.Li S, Chen H, Man J, Zhang T, Yin X, He Q, Yang X, et al. Changing trends in the disease burden of esophageal cancer in China from 1990 to 2017 and its predicted level in 25 years. Cancer Med. . 2021;10:1889–1899. doi: 10.1002/cam4.3775. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Dawson MA, Kouzarides T. Cancer epigenetics: from mechanism to therapy. Cell. . 2012;150:12–27. doi: 10.1016/j.cell.2012.06.013. [DOI] [PubMed] [Google Scholar]
- 5.Esteller M. Cancer epigenomics: DNA methylomes and histone-modification maps. Nat Rev Genet. . 2007;8:286–298. doi: 10.1038/nrg2005. [DOI] [PubMed] [Google Scholar]
- 6.Jones PA. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat Rev Genet. . 2012;13:484–492. doi: 10.1038/nrg3230. [DOI] [PubMed] [Google Scholar]
- 7.Wiener D, Schwartz S. The epitranscriptome beyond m 6A . Nat Rev Genet. . 2021;22:119–131. doi: 10.1038/s41576-020-00295-8. [DOI] [PubMed] [Google Scholar]
- 8.Hogg SJ, Beavis PA, Dawson MA, Johnstone RW. Targeting the epigenetic regulation of antitumour immunity. Nat Rev Drug Discov. . 2020;19:776–800. doi: 10.1038/s41573-020-0077-5. [DOI] [PubMed] [Google Scholar]
- 9.Chen Y, Hong T, Wang S, Mo J, Tian T, Zhou X. Epigenetic modification of nucleic acids: from basic studies to medical applications. Chem Soc Rev. . 2017;46:2844–2872. doi: 10.1039/C6CS00599C. [DOI] [PubMed] [Google Scholar]
- 10.Machnicka MA, Milanowska K, Osman Oglou O, Purta E, Kurkowska M, Olchowik A, Januszewski W, et al. MODOMICS: a database of RNA modification pathways—2013 update. Nucleic Acids Res. . 2013;41:D262–D267. doi: 10.1093/nar/gks1007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Barbieri I, Kouzarides T. Role of RNA modifications in cancer. Nat Rev Cancer. . 2020;20:303–322. doi: 10.1038/s41568-020-0253-2. [DOI] [PubMed] [Google Scholar]
- 12.Hopfinger MC, Kirkpatrick CC, Znosko BM. Predictions and analyses of RNA nearest neighbor parameters for modified nucleotides. Nucleic Acids Res. . 2020;48:8901–8913. doi: 10.1093/nar/gkaa654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Desrosiers R, Friderici K, Rottman F. Identification of methylated nucleosides in messenger RNA from Novikoff hepatoma cells. Proc Natl Acad Sci USA. . 1974;71:3971–3975. doi: 10.1073/pnas.71.10.3971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Zaccara S, Ries RJ, Jaffrey SR. Reading, writing and erasing mRNA methylation. Nat Rev Mol Cell Biol. . 2019;20:608–624. doi: 10.1038/s41580-019-0168-5. [DOI] [PubMed] [Google Scholar]
- 15.Shi H, Wei J, He C. Where, when, and how: context-dependent functions of rna methylation writers, readers, and erasers. Mol Cell. . 2019;74:640–650. doi: 10.1016/j.molcel.2019.04.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Zhao X, Yang Y, Sun BF, Shi Y, Yang X, Xiao W, Hao YJ, et al. FTO-dependent demethylation of N6-methyladenosine regulates mRNA splicing and is required for adipogenesis. Cell Res. . 2014;24:1403–1419. doi: 10.1038/cr.2014.151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Xiao W, Adhikari S, Dahal U, Chen YS, Hao YJ, Sun BF, Sun HY, et al. Nuclear m(6)A reader YTHDC1 regulates mRNA splicing. Mol Cell. . 2016;61:507–519. doi: 10.1016/j.molcel.2016.01.012. [DOI] [PubMed] [Google Scholar]
- 18.Spitale RC, Flynn RA, Zhang QC, Crisalli P, Lee B, Jung JW, Kuchelmeister HY, et al. Structural imprints in vivo decode RNA regulatory mechanisms . Nature. . 2015;519:486–490. doi: 10.1038/nature14263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Roundtree IA, Luo GZ, Zhang Z, Wang X, Zhou T, Cui Y, Sha J, et al. YTHDC1 mediates nuclear export of N(6)-methyladenosine methylated mRNAs. Elife. 2017, 6: e31311 . [DOI] [PMC free article] [PubMed]
- 20.Wang X, Zhao BS, Roundtree IA, Lu Z, Han D, Ma H, Weng X, et al. N6-methyladenosine modulates messenger RNA translation efficiency. Cell. . 2015;161:1388–1399. doi: 10.1016/j.cell.2015.05.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Jiang X, Liu B, Nie Z, Duan L, Xiong Q, Jin Z, Yang C, et al. The role of m6A modification in the biological functions and diseases. Sig Transduct Target Ther. . 2021;6:74. doi: 10.1038/s41392-020-00450-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Wyatt GR. Occurrence of 5-Methyl-Cytosine in nucleic acids. Nature. . 1950;166:237–238. doi: 10.1038/166237b0. [DOI] [PubMed] [Google Scholar]
- 23.Bujnicki JM. Sequence-structure-function studies of tRNA:m5C methyltransferase Trm4p and its relationship to DNA:m5C and RNA:m5U methyltransferases. Nucleic Acids Res. . 2004;32:2453–2463. doi: 10.1093/nar/gkh564. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Motorin Y, Lyko F, Helm M. 5-methylcytosine in RNA: detection, enzymatic formation and biological functions. Nucleic Acids Res. . 2010;38:1415–1430. doi: 10.1093/nar/gkp1117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Fu L, Guerrero CR, Zhong N, Amato NJ, Liu Y, Liu S, Cai Q, et al. Tet-mediated formation of 5-hydroxymethylcytosine in RNA. J Am Chem Soc. . 2014;136:11582–11585. doi: 10.1021/ja505305z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Shen Q, Zhang Q, Shi Y, Shi Q, Jiang Y, Gu Y, Li Z, et al. Tet2 promotes pathogen infection-induced myelopoiesis through mRNA oxidation. Nature. . 2018;554:123–127. doi: 10.1038/nature25434. [DOI] [PubMed] [Google Scholar]
- 27.Chen X, Li A, Sun BF, Yang Y, Han YN, Yuan X, Chen RX, et al. 5-methylcytosine promotes pathogenesis of bladder cancer through stabilizing mRNAs. Nat Cell Biol. . 2019;21:978–990. doi: 10.1038/s41556-019-0361-y. [DOI] [PubMed] [Google Scholar]
- 28.Chen Y, Lin H, Miao L, He J. Role of N7-methylguanosine (m7G) in cancer. Trends Cell Biol. . 2022;32:819–824. doi: 10.1016/j.tcb.2022.07.001. [DOI] [PubMed] [Google Scholar]
- 29.Furuichi Y. Discovery of m 7G-cap in eukaryotic mRNAs . Proc Jpn Acad Ser B-Phys Biol Sci. . 2015;91:394–409. doi: 10.2183/pjab.91.394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Alexandrov A, Martzen MR, Phizicky EM. Two proteins that form a complex are required for 7-methylguanosine modification of yeast tRNA. RNA. . 2002;8:1253–1266. doi: 10.1017/S1355838202024019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Malbec L, Zhang T, Chen YS, Zhang Y, Sun BF, Shi BY, Zhao YL, et al. Dynamic methylome of internal mRNA N7-methylguanosine and its regulatory role in translation. Cell Res. . 2019;29:927–941. doi: 10.1038/s41422-019-0230-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Zhang LS, Liu C, Ma H, Dai Q, Sun HL, Luo G, Zhang Z, et al. Transcriptome-wide mapping of internal N7-Methylguanosine methylome in mammalian mRNA. Mol Cell. . 2019;74:1304–1316.e8. doi: 10.1016/j.molcel.2019.03.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Cowling VH. Enhanced mRNA cap methylation increases cyclin D1 expression and promotes cell transformation. Oncogene. . 2010;29:930–936. doi: 10.1038/onc.2009.368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Wang W, Shao F, Yang X, Wang J, Zhu R, Yang Y, Zhao G, et al. METTL3 promotes tumour development by decreasing APC expression mediated by APC mRNA N6-methyladenosine-dependent YTHDF binding. Nat Commun. . 2021;12:3803. doi: 10.1038/s41467-021-23501-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Liao L, He Y, Li SJ, Zhang GG, Yu W, Yang J, Huang ZJ, et al. Anti-HIV drug elvitegravir suppresses cancer metastasis via increased proteasomal degradation of m6A methyltransferase METTL3. Cancer Res. . 2022;82:2444–2457. doi: 10.1158/0008-5472.CAN-21-4124. [DOI] [PubMed] [Google Scholar]
- 36.Li R, Zeng L, Zhao H, Deng J, Pan L, Zhang S, Wu G, et al. ATXN2-mediated translation of TNFR1 promotes esophageal squamous cell carcinoma via m6A-dependent manner. Mol Ther. . 2022;30:1089–1103. doi: 10.1016/j.ymthe.2022.01.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Jia X, Zhang Z, Wei R, Li B, Chen Y, Li J. Comprehensive analysis of transcriptome-wide m6A methylome in intermediate-stage Esophageal squamous cell carcinoma. Pathol-Res Pract. . 2022;237:154055. doi: 10.1016/j.prp.2022.154055. [DOI] [PubMed] [Google Scholar]
- 38.Liang X, Zhang Z, Wang L, Zhang S, Ren L, Li S, Xu J, et al. Mechanism of methyltransferase like 3 in epithelial-mesenchymal transition process, invasion, and metastasis in esophageal cancer. Bioengineered. . 2021;12:10023–10036. doi: 10.1080/21655979.2021.1994721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Han H, Yang C, Zhang S, Cheng M, Guo S, Zhu Y, Ma J, et al. METTL3-mediated m6A mRNA modification promotes esophageal cancer initiation and progression via Notch signaling pathway. Mol Ther Nucleic Acids. . 2021;26:333–346. doi: 10.1016/j.omtn.2021.07.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Chen X, Huang L, Yang T, Xu J, Zhang C, Deng Z, Yang X, et al. METTL3 promotes esophageal squamous cell carcinoma metastasis through enhancing GLS2 expression. Front Oncol. . 2021;11:667451. doi: 10.3389/fonc.2021.667451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Hou H, Zhao H, Yu X, Cong P, Zhou Y, Jiang Y, Cheng Y. METTL3 promotes the proliferation and invasion of esophageal cancer cells partly through AKT signaling pathway. Pathol Res Pract. . 2020;216:153087. doi: 10.1016/j.prp.2020.153087. [DOI] [PubMed] [Google Scholar]
- 42.Zou J, Zhong X, Zhou X, Xie Q, Zhao Z, Guo X, Duan Y. The M6A methyltransferase METTL3 regulates proliferation in esophageal squamous cell carcinoma. Biochem Biophys Res Commun. . 2021;580:48–55. doi: 10.1016/j.bbrc.2021.05.048. [DOI] [PubMed] [Google Scholar]
- 43.Li J, Li Z, Xu Y, Huang C, Shan B. METTL3 facilitates tumor progression by COL12A1/MAPK signaling pathway in esophageal squamous cell carcinoma. J Cancer. . 2022;13:1972–1984. doi: 10.7150/jca.66830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Hu W, Liu W, Liang H, Zhang C, Zou M, Zou B. Silencing of methyltransferase-like 3 inhibits oesophageal squamous cell carcinoma. Exp Ther Med. 2020, 20: 138 . [DOI] [PMC free article] [PubMed]
- 45.Liu Z, Wu K, Gu S, Wang W, Xie S, Lu T, Li L, et al. A methyltransferase‐like 14/miR‐99a‐5p/tribble 2 positive feedback circuit promotes cancer stem cell persistence and radioresistance via histone deacetylase 2‐mediated epigenetic modulation in esophageal squamous cell carcinoma. Clin Transl Med. . 2021;11 doi: 10.1002/ctm2.545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Luo Q, Zhan X, Kuang Y, Sun M, Dong F, Sun E, Chen B. WTAP promotes oesophageal squamous cell carcinoma development by decreasing CPSF4 expression in an m6A-dependent manner. Med Oncol. . 2022;39:231. doi: 10.1007/s12032-022-01830-9. [DOI] [PubMed] [Google Scholar]
- 47.Luo GM, Qi Y, Lei ZY, Shen XY, Chen MM, Du LL, Wu CX, et al. A potential biomarker of esophageal squamous cell carcinoma WTAP promotes the proliferation and migration of ESCC. Pathol Res Pract. . 2022;238:154114. doi: 10.1016/j.prp.2022.154114. [DOI] [PubMed] [Google Scholar]
- 48.Wu Q, Zhang H, Yang D, Min Q, Wang Y, Zhang W, Zhan Q. The m6A-induced lncRNA CASC8 promotes proliferation and chemoresistance via upregulation of hnRNPL in esophageal squamous cell carcinoma. Int J Biol Sci. . 2022;18:4824–4836. doi: 10.7150/ijbs.71234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Chen P, Li S, Zhang K, Zhao R, Cui J, Zhou W, Liu Y, et al. N6-methyladenosine demethylase ALKBH5 suppresses malignancy of esophageal cancer by regulating microRNA biogenesis and RAI1 expression. Oncogene. . 2021;40:5600–5612. doi: 10.1038/s41388-021-01966-4. [DOI] [PubMed] [Google Scholar]
- 50.Wu S, Zhang L, Deng J, Guo B, Li F, Wang Y, Wu R, et al. A novel micropeptide encoded by Y-Linked LINC00278 links cigarette smoking and ar signaling in male esophageal squamous cell carcinoma. Cancer Res. . 2020;80:2790–2803. doi: 10.1158/0008-5472.CAN-19-3440. [DOI] [PubMed] [Google Scholar]
- 51.Xue J, Xiao P, Yu X, Zhang X. A positive feedback loop between AlkB homolog 5 and miR-193a-3p promotes growth and metastasis in esophageal squamous cell carcinoma. Hum Cell. . 2021;34:502–514. doi: 10.1007/s13577-020-00458-z. [DOI] [PubMed] [Google Scholar]
- 52.Li J, Liu H, Dong S, Zhang Y, Li X, Wang J. ALKBH5 is lowly expressed in esophageal squamous cell carcinoma and inhibits the malignant proliferation and invasion of tumor cells. Comput Math Methods Med. 2021: 1001446 . [DOI] [PMC free article] [PubMed] [Retracted]
- 53.Xiao D, Fang TX, Lei Y, Xiao SJ, Xia JW, Lin TY, Li YL, et al. m6A demethylase ALKBH5 suppression contributes to esophageal squamous cell carcinoma progression. Aging. . 2021;13:21497–21512. doi: 10.18632/aging.203490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Cui Y, Zhang C, Ma S, Li Z, Wang W, Li Y, Ma Y, et al. RNA m6A demethylase FTO-mediated epigenetic up-regulation of LINC00022 promotes tumorigenesis in esophageal squamous cell carcinoma. J Exp Clin Cancer Res. . 2021;40:294. doi: 10.1186/s13046-021-02096-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Duan X, Yang L, Wang L, Liu Q, Zhang K, Liu S, Liu C, et al. m6A demethylase FTO promotes tumor progression via regulation of lipid metabolism in esophageal cancer. Cell Biosci. . 2022;12:60. doi: 10.1186/s13578-022-00798-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Zhao F, Ge F, Xie M, Li Z, Zang C, Kong L, Pu Y, et al. FTO mediated ERBB2 demethylation promotes tumor progression in esophageal squamous cell carcinoma cells. Clin Exp Metastasis. . 2022;39:623–639. doi: 10.1007/s10585-022-10169-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Liu S, Huang M, Chen Z, Chen J, Chao Q, Yin X, Quan M. FTO promotes cell proliferation and migration in esophageal squamous cell carcinoma through up-regulation of MMP13. Exp Cell Res. . 2020;389:111894. doi: 10.1016/j.yexcr.2020.111894. [DOI] [PubMed] [Google Scholar]
- 58.Wang Y, Yu Z, Shi W, Shen J, Guan Y, Ni F. HLA complex P5 upregulation is correlated with poor prognosis and tumor progression in esophageal squamous cell carcinoma. Bioengineered. . 2022;13:9302–9312. doi: 10.1080/21655979.2022.2051854. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Wu X, Fan Y, Liu Y, Shen B, Lu H, Ma H. Long non-coding RNA CCAT2 promotes the development of esophageal squamous cell carcinoma by inhibiting miR-200b to upregulate the IGF2BP2/TK1 axis. Front Oncol. . 2021;11:680642. doi: 10.3389/fonc.2021.680642. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Guo H, Wang B, Xu K, Nie L, Fu Y, Wang Z, Wang Q, et al. m6A reader HNRNPA2B1 promotes esophageal cancer progression via up-regulation of ACLY and ACC1. Front Oncol. . 2020;10:553045. doi: 10.3389/fonc.2020.553045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Yang N, Ying P, Tian J, Wang X, Mei S, Zou D, Peng X, et al. Genetic variants in m6A modification genes are associated with esophageal squamous-cell carcinoma in the Chinese population. Carcinogenesis. . 2020;41:761–768. doi: 10.1093/carcin/bgaa012. [DOI] [PubMed] [Google Scholar]
- 62.Li Y, Li J, Luo M, Zhou C, Shi X, Yang W, Lu Z, et al. Novel long noncoding RNA NMR promotes tumor progression via NSUN2 and BPTF in esophageal squamous cell carcinoma. Cancer Lett. . 2018;430:57–66. doi: 10.1016/j.canlet.2018.05.013. [DOI] [PubMed] [Google Scholar]
- 63.Su J, Wu G, Ye Y, Zhang J, Zeng L, Huang X, Zheng Y, et al. NSUN2-mediated RNA 5-methylcytosine promotes esophageal squamous cell carcinoma progression via LIN28B-dependent GRB2 mRNA stabilization. Oncogene. . 2021;40:5814–5828. doi: 10.1038/s41388-021-01978-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Niu X, Peng L, Liu W, Miao C, Chen X, Chu J, Yang X, et al. A cis-eQTL in NSUN2 promotes esophageal squamous-cell carcinoma progression and radiochemotherapy resistance by mRNA-m5C methylation. Sig Transduct Target Ther. . 2022;7:267. doi: 10.1038/s41392-022-01063-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Du J, Zhang G, Qiu H, Yu H, Yuan W. A novel positive feedback loop of linc02042 and c-Myc mediated by YBX1 promotes tumorigenesis and metastasis in esophageal squamous cell carcinoma. Cancer Cell Int. . 2020;20:75. doi: 10.1186/s12935-020-1154-x. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 66.Xu F, Liu Z, Liu R, Lu C, Wang L, Mao W, Zhu Q, et al. Epigenetic induction of tumor stemness via the lipopolysaccharide-TET3-HOXB2 signaling axis in esophageal squamous cell carcinoma. Cell Commun Signal. . 2020;18:17. doi: 10.1186/s12964-020-0510-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Hu C, Peng J, Lv L, Wang X, Huo J, Liu D. MiR-196a promotes the proliferation and migration of esophageal cancer via the UHRF2/TET2 axis. Mol Cell Biochem. . 2022;477:537–547. doi: 10.1007/s11010-021-04301-3. [DOI] [PubMed] [Google Scholar]
- 68.Dong Z, Li S, Wu X, Niu Y, Liang X, Yang L, Guo Y, et al. Aberrant hypermethylation-mediated downregulation of antisense lncRNA ZNF667-AS1 and its sense gene ZNF667 correlate with progression and prognosis of esophageal squamous cell carcinoma. Cell Death Dis. . 2019;10:930. doi: 10.1038/s41419-019-2171-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Li L, Li C, Mao H, Du Z, Chan WY, Murray P, Luo B, et al. Epigenetic inactivation of the CpG demethylase TET1 as a DNA methylation feedback loop in human cancers. Sci Rep. . 2016;6:26591. doi: 10.1038/srep26591. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Han H, Yang C, Ma J, Zhang S, Zheng S, Ling R, Sun K, et al. N7-methylguanosine tRNA modification promotes esophageal squamous cell carcinoma tumorigenesis via the RPTOR/ULK1/autophagy axis. Nat Commun. . 2022;13:1478. doi: 10.1038/s41467-022-29125-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Xia TL, Yan SM, Yuan L, Zeng MS. Upregulation of METTL3 expression predicts poor prognosis in patients with esophageal squamous cell carcinoma. Cancer Manag Res. . 2020;12:5729–5737. doi: 10.2147/CMAR.S245019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Pu Y, Lu X, Yang X, Yang Y, Wang D, Li M, Guan W, et al. Estimating the prognosis of esophageal squamous cell carcinoma based on The Cancer Genome Atlas (TCGA) of m6A methylation-associated genes. J Gastrointest Oncol. . 2022;13:1–12. doi: 10.21037/jgo-21-686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Li K, Chen J, Lou X, Li Y, Qian B, Xu D, Wu Y, et al. HNRNPA2B1 affects the prognosis of esophageal cancer by regulating the miR-17-92 cluster. Front Cell Dev Biol. . 2021;9:658642. doi: 10.3389/fcell.2021.658642. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Li L, Xie R, Wei Q. Network analysis of miRNA targeting m6A-related genes in patients with esophageal cancer. PeerJ. . 2021;9:e11893. doi: 10.7717/peerj.11893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Liu XS, Kui XY, Gao Y, Chen XQ, Zeng J, Liu XY, Zhang Y, et al. Comprehensive analysis of YTHDF1 immune infiltrates and ceRNA in human esophageal carcinoma. Front Genet. . 2022;13:835265. doi: 10.3389/fgene.2022.835265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Xu L, Pan J, Pan H. Construction and validation of an m6A RNA methylation regulators-based prognostic signature for esophageal cancer. Cancer Manag Res. . 2020;12:5385–5394. doi: 10.2147/CMAR.S254870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Zhao H, Xu Y, Xie Y, Zhang L, Gao M, Li S, Wang F. m6A regulators is differently expressed and correlated with immune response of esophageal cancer. Front Cell Dev Biol. . 2021;9:650023. doi: 10.3389/fcell.2021.650023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Lu F, Chen W, Jiang T, Cheng C, Wang B, Lu Z, Huang G, et al. Expression profile, clinical significance and biological functions of IGF2BP2 in esophageal squamous cell carcinoma. Exp Ther Med. . 2022;23:252. doi: 10.3892/etm.2022.11177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Wakita A, Motoyama S, Sato Y, Nagaki Y, Fujita H, Terata K, Imai K, et al. IGF2BP3 expression correlates with poor prognosis in esophageal squamous cell carcinoma. J Surg Res. . 2021;259:137–144. doi: 10.1016/j.jss.2020.10.024. [DOI] [PubMed] [Google Scholar]
- 80.Wu H, Li F, Zhu R. miR-338-5p inhibits cell growth and migration via inhibition of the METTL3/m6A/c-Myc pathway in lung cancer. Acta Biochim Biophys Sin. . 2021;53:304–316. doi: 10.1093/abbs/gmaa170. [DOI] [PubMed] [Google Scholar]
- 81.Zhang M, Bai M, Wang L, Lu N, Wang J, Yan R, Cui M, et al. Targeting SNHG3/miR-186-5p reverses the increased m6A level caused by platinum treatment through regulating METTL3 in esophageal cancer. Cancer Cell Int. . 2021;21:114. doi: 10.1186/s12935-021-01747-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Guo W, Tan F, Huai Q, Wang Z, Shao F, Zhang G, Yang Z, et al. Comprehensive analysis of PD-L1 expression, immune infiltrates, and m6A RNA methylation regulators in esophageal squamous cell carcinoma. Front Immunol. . 2021;12:669750. doi: 10.3389/fimmu.2021.669750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Zhou Y, Guo S, Li Y, Chen F, Wu Y, Xiao Y, An J. METTL3 is associated with the malignancy of esophageal squamous cell carcinoma and serves as a potential immunotherapy biomarker. Front Oncol. . 2022;12:824190. doi: 10.3389/fonc.2022.824190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Chen F, Gong E, Ma J, Lin J, Wu C, Chen S, Hu S. Prognostic score model based on six m6A ‐related autophagy genes for predicting survival in esophageal squamous cell carcinoma. J Clin Lab Anal. . 2022;36:e24507. doi: 10.1002/jcla.24507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Wang Q, Liu H, Liu Z, Yang L, Zhou J, Cao X, Sun H. Circ-SLC7A5, a potential prognostic circulating biomarker for detection of ESCC. Cancer Genet. . 2020;240:33–39. doi: 10.1016/j.cancergen.2019.11.001. [DOI] [PubMed] [Google Scholar]
- 86.Brancato V, Garbino N, Mannelli L, Aiello M, Salvatore M, Franzese M, Cavaliere C. Impact of radiogenomics in esophageal cancer on clinical outcomes: A pilot study. World J Gastroenterol. . 2021;27:6110–6127. doi: 10.3748/wjg.v27.i36.6110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Okamoto M, Hirata S, Sato S, Koga S, Fujii M, Qi G, Ogawa I, et al. Frequent increased gene copy number and high protein expression of tRNA (cytosine-5-)-methyltransferase (NSUN2) in human cancers. DNA Cell Biol. . 2012;31:660–671. doi: 10.1089/dna.2011.1446. [DOI] [PubMed] [Google Scholar]
- 88.Shi X, Yu Y, Luo M, Zhang Z, Shi S, Feng X, Chen Z, et al. Loss of 5-Hydroxymethylcytosine is an independent unfavorable prognostic factor for esophageal squamous cell carcinoma. PLoS One. . 2016;11:e0153100. doi: 10.1371/journal.pone.0153100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Sawada G, Niida A, Uchi R, Hirata H, Shimamura T, Suzuki Y, Shiraishi Y, et al. Genomic landscape of esophageal squamous cell carcinoma in a japanese population. Gastroenterology. . 2016;150:1171–1182. doi: 10.1053/j.gastro.2016.01.035. [DOI] [PubMed] [Google Scholar]
- 90.Campbell JD, Yau C, Bowlby R, Liu Y, Brennan K, Fan H, Taylor AM, et al. Genomic, pathway network, and immunologic features distinguishing squamous carcinomas. Cell Rep. . 2018;23:194–212.e6. doi: 10.1016/j.celrep.2018.03.063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Li C, Wang Q, Ma J, Shi S, Chen X, Yang H, Han J. Integrative pathway analysis of genes and metabolites reveals metabolism abnormal subpathway regions and modules in esophageal squamous cell carcinoma. Molecules. . 2017;22:1599. doi: 10.3390/molecules22101599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Zhao F, Dong Z, Li Y, Liu S, Guo P, Zhang D, Li S. Comprehensive analysis of molecular clusters and prognostic signature based on m7G-related LncRNAs in esophageal squamous cell carcinoma. Front Oncol. . 2022;12:893186. doi: 10.3389/fonc.2022.893186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Xue C, Chu Q, Zheng Q, Jiang S, Bao Z, Su Y, Lu J, et al. Role of main RNA modifications in cancer: N6-methyladenosine, 5-methylcytosine, and pseudouridine. Sig Transduct Target Ther. . 2022;7:142. doi: 10.1038/s41392-022-01003-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Yankova E, Blackaby W, Albertella M, Rak J, De Braekeleer E, Tsagkogeorga G, Pilka ES, et al. Small-molecule inhibition of METTL3 as a strategy against myeloid leukaemia. Nature. . 2021;593:597–601. doi: 10.1038/s41586-021-03536-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Li J, Gregory RI. Mining for METTL3 inhibitors to suppress cancer. Nat Struct Mol Biol. . 2021;28:460–462. doi: 10.1038/s41594-021-00606-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Du Y, Yuan Y, Xu L, Zhao F, Wang W, Xu Y, Tian X. Discovery of METTL3 small molecule inhibitors by virtual screening of natural products. Front Pharmacol. . 2022;13:878135. doi: 10.3389/fphar.2022.878135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Xie G, Wu XN, Ling Y, Rui Y, Wu D, Zhou J, Li J, et al. A novel inhibitor of N6-methyladenosine demethylase FTO induces mRNA methylation and shows anti-cancer activities. Acta Pharmaceutica Sin B. . 2022;12:853–866. doi: 10.1016/j.apsb.2021.08.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Cui Q, Shi H, Ye P, Li L, Qu Q, Sun G, Sun G, et al. m6A RNA methylation regulates the self-renewal and tumorigenesis of glioblastoma stem cells. Cell Rep. . 2017;18:2622–2634. doi: 10.1016/j.celrep.2017.02.059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Su R, Dong L, Li C, Nachtergaele S, Wunderlich M, Qing Y, Deng X, et al. R-2HG exhibits anti-tumor activity by targeting FTO/m6A/MYC/CEBPA signaling. Cell. . 2018;172:90–105.e23. doi: 10.1016/j.cell.2017.11.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Huang Y, Su R, Sheng Y, Dong L, Dong Z, Xu H, Ni T, et al. Small-molecule targeting of oncogenic FTO demethylase in acute myeloid leukemia. Cancer Cell. . 2019;35:677–691.e10. doi: 10.1016/j.ccell.2019.03.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Weng H, Huang F, Yu Z, Chen Z, Prince E, Kang Y, Zhou K, et al. The m6A reader IGF2BP2 regulates glutamine metabolism and represents a therapeutic target in acute myeloid leukemia. Cancer Cell. . 2022;40:1566–1582.e10. doi: 10.1016/j.ccell.2022.10.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Li N, Kang Y, Wang L, Huff S, Tang R, Hui H, Agrawal K, et al. ALKBH5 regulates anti–PD-1 therapy response by modulating lactate and suppressive immune cell accumulation in tumor microenvironment. Proc Natl Acad Sci USA. . 2020;117:20159–20170. doi: 10.1073/pnas.1918986117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Fang Z, Mu B, Liu Y, Guo N, Xiong L, Guo Y, Xia A, et al. Discovery of a potent, selective and cell active inhibitor of m6A demethylase ALKBH5. Eur J Medicinal Chem. . 2022;238:114446. doi: 10.1016/j.ejmech.2022.114446. [DOI] [PubMed] [Google Scholar]
- 104.Visvanathan A, Patil V, Arora A, Hegde AS, Arivazhagan A, Santosh V, Somasundaram K. Essential role of METTL3-mediated m6A modification in glioma stem-like cells maintenance and radioresistance. Oncogene. . 2018;37:522–533. doi: 10.1038/onc.2017.351. [DOI] [PubMed] [Google Scholar]
- 105.Lin EW, Karakasheva TA, Hicks PD, Bass AJ, Rustgi AK. The tumor microenvironment in esophageal cancer. Oncogene. . 2016;35:5337–5349. doi: 10.1038/onc.2016.34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Wan W, Ao X, Chen Q, Yu Y, Ao L, Xing W, Guo W, et al. METTL3/IGF2BP3 axis inhibits tumor immune surveillance by upregulating N6-methyladenosine modification of PD-L1 mRNA in breast cancer. Mol Cancer. . 2022;21:60. doi: 10.1186/s12943-021-01447-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Liu Z, Wang T, She Y, Wu K, Gu S, Li L, Dong C, et al. N6-methyladenosine-modified circIGF2BP3 inhibits CD8+ T-cell responses to facilitate tumor immune evasion by promoting the deubiquitination of PD-L1 in non-small cell lung cancer. Mol Cancer. . 2021;20:105. doi: 10.1186/s12943-021-01398-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Ni Z, Sun P, Zheng J, Wu M, Yang C, Cheng M, Yin M, et al. JNK signaling promotes bladder cancer immune escape by regulating METTL3-Mediated m6A modification of PD-L1 mRNA. Cancer Res. . 2022;82:1789–1802. doi: 10.1158/0008-5472.CAN-21-1323. [DOI] [PubMed] [Google Scholar]
- 109.Wei W, Sun J, Zhang H, Xiao X, Huang C, Wang L, Zhong H, et al. Circ0008399 interaction with WTAP promotes assembly and activity of the m6A methyltransferase complex and promotes cisplatin resistance in bladder cancer . Cancer Res. . 2021;81:6142–6156. doi: 10.1158/0008-5472.CAN-21-1518. [DOI] [PubMed] [Google Scholar]
- 110.Li M, Xia M, Zhang Z, Tan Y, Li E, Guo Z, Fang M, et al. METTL3 antagonizes 5‑FU chemotherapy and confers drug resistance in colorectal carcinoma. Int J Oncol. . 2022;61:106. doi: 10.3892/ijo.2022.5396. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Li H, Wang C, Lan L, Yan L, Li W, Evans I, Ruiz EJ, et al. METTL3 promotes oxaliplatin resistance of gastric cancer CD133+ stem cells by promoting PARP1 mRNA stability. Cell Mol Life Sci. . 2022;79:135. doi: 10.1007/s00018-022-04129-0. [DOI] [PMC free article] [PubMed] [Google Scholar]