SUMMARY
Even-chain acylcarnitine (AC) metabolites, most of which are generated as byproducts of incomplete fatty acid oxidation (FAO), are viewed as biomarkers of mitochondrial lipid stress attributable to one or more metabolic bottlenecks in the beta-oxidation pathway. The origins and functional implications of FAO bottlenecks remain poorly understood. Here, we combined a sophisticated mitochondrial phenotyping platform with state-of-the-art molecular profiling tools and multiple two-state mouse models of respiratory function to uncover a mechanism that connects AC accumulation to lipid intolerance, metabolic inflexibility, and respiratory inefficiency in skeletal muscle mitochondria. These studies also identified a short chain carbon circuit at the C4 node of FAO wherein reverse flux of glucose-derived acetyl-CoA through medium chain ketothiolase enhances lipid tolerance and redox stability in heart mitochondria by regenerating free CoA and NAD+. The findings help to explain why diminished FAO capacity, AC accumulation, and metabolic inflexibility are tightly linked to poor health outcomes.
eTOC Summary
Researchers identify a short chain carbon circuit that enables reverse flux of pyruvate-derived acetyl CoA through medium chain ketothiolase (MKT) to regenerate critical co-factors necessary for beta-oxidation of long chain fatty acids, thereby promoting mitochondrial lipid tolerance. Insufficient MKT flux leads to lipid-induced energy instability and metabolic inflexibility.
Graphical Abstract

INTRODUCTION
In most cells, oxidative ATP generation by mitochondria accounts for a substantial majority of total ATP requirements. This process, known as oxidative phosphorylation (OXPHOS), involves the transfer of energy captured in carbon fuels, such as fatty acids and glucose, to the free energy of ATP hydrolysis (ΔGATP)–the energy currency that powers normal cell and organ function. Catabolism of carbon substrates by dehydrogenase enzymes generates electron energy potential (ΔGredox) in the form of NADH/NAD+ and FADH2/FAD. In turn, these redox pairs supply the electron transport chain (ETC), which couples electron transfer and reduction of molecular oxygen at complex IV to proton pumping at complexes I, III and IV to generate the proton motive force (PMF) across the mitochondrial inner membrane. The PMF provides the ultimate driving force for ATP resynthesis via complex V (ATP synthase). To this point, it is important to underscore that in striated muscles, mitochondria serve two primary functions: i) ATP resynthesis, and ii) displacement of the ATP:ADP ratio far from equilibrium (by ~10 orders of magnitude) to establish and maintain ΔGATP. Thus, mitochondria are typically synthesizing and pumping ATP against backpressure, into a sea of ATP, “up an energetic hill” 1,2.
Not surprisingly, perturbations in oxidative metabolism represent a common feature of aging and age-related metabolic disorders such as obesity, type 2 diabetes, and heart failure 3,4. Mitochondrial dysfunction can manifest in many forms. Among the hallmarks of cardiometabolic diseases is an apparent rigidity in mitochondrial substrate selection, such that various organs and cells fail to appropriately adjust fuel choice in response to nutritional and other physiological cues. This phenomenon, dubbed “metabolic inflexibility” 5, is often accompanied by diminished capacity for fatty acid oxidation (FAO) and the onset of whole-body glucose intolerance and/or insulin resistance 3,6. By contrast, habitual physical activity increases FAO capacity and facilitates robust fuel switching during metabolic transitions 7–9. In aggregate, a large body of evidence has established strong links between diminished FAO capacity, metabolic inflexibility and poor health outcomes 3. Still uncertain, however, are the bioenergetic and molecular mechanisms that underlie this connection.
Whereas fatty acids are known to serve as critical respiratory substrates during starvation, prolonged carbohydrate restriction, and/or endurance exercise, the question of why FAO capacity matters in the context of overnutrition and physical inactivity remains enigmatic. Previous reports have proposed that a persistent oversupply of fatty acid fuels leads to a state of mitochondrial lipid overload, which can give rise to inefficient and/or incomplete FAO along with compromised energy stability 10–14. Among several convincing lines of evidence linking mitochondrial overload to metabolic disease are numerous reports showing that circulating and/or tissue levels of fatty acid-derived acylcarnitine metabolites are elevated in circumstances of lipid stress (e.g. starvation, obesity, diabetes, dyslipidemia, and heart failure) 7,15–25. Considering that most even-chain acylcarnitine metabolites are generated as byproducts of incomplete fatty acid catabolism, these molecules are commonly viewed as biomarkers of mitochondrial lipid stress attributable to one or more metabolic bottlenecks in the beta-oxidation pathway 3. Nonetheless, the origins and functional implications of these FAO bottlenecks are poorly understood.
The overarching goal of this study was to gain insight into the molecular mechanisms that connect metabolic inflexibility and acylcarnitine accumulation to age-related cardiometabolic diseases. While several previous investigations have amassed clues through the application of state-of-the-art molecular profiling technologies, these tools are limited in that they do not provide direct metrics of mitochondrial performance. Herein, we sought to understand how mitochondria resident in specific muscle tissues respond to a heavy influx of lipid fuel, aiming to address two fundamentally important questions: 1) Does mitochondrial lipid overload and/or acylcarnitine accumulation compromise and/or impact OXPHOS thermodynamics? 2) If so, what are the underlying mechanisms? To this end, we combined mass spectrometry-based metabolomics, proteomics and stable isotope tracers with a sophisticated mitochondrial diagnostics platform to assess the interplay between metabolic flexibility, carbon flux, and bioenergetics using multiple two-state models of enhanced or diminished FAO capacity. This approach led us to identify a short chain carbon circuit that enables reverse flux of pyruvate-derived acetyl CoA through medium chain ketothiolase (MKT/Acaa2) to regenerate critical co-factors that support FAO as well as downstream flux through the tricarboxylic acid cycle (TCAC). In turn, this circuit promotes mitochondrial lipid tolerance and metabolic plasticity. By uncovering the important role of pyruvate-supported MKT flux in conferring metabolic resilience in the face of lipid surplus, the study reveals a heretofore unappreciated mechanism linking FAO overload and pyruvate insufficiency to energy instability and metabolic inflexibility.
RESULTS
Mitochondria from mixed SkM are metabolically inflexible and intolerant to long chain lipid fuel.
Mitochondrial diagnostics.
This study leveraged a mitochondrial phenotyping platform that permits multiplexed assessments of energy transduction using freshly isolated mitochondria exposed to a variety of distinct substrate and energetic conditions 1. The platform features a modified version of the creatine kinase (CK) energetic clamp technique (Figure 1A and S. Fig 1A) wherein the extramitochondrial ratio of ATP:ADP, and thus the ΔGATP, is precisely controlled by addition of a large surplus of CK and defined concentrations of creatine, phosphocreatine (PCr) and adenylates, which together confer excess capacitance to compensate for deficits in rates of mitochondrial ATP regeneration 2,26,27. The relationships between the primary CK clamp components (ATP, ADP, Cr, PCr) across a span of physiological ATP free energies are depicted in Supplemental Figure 1A. As mitochondria are exposed to more PCr, the ATP:ADP ratio increases, resulting in diminished respiratory flux. In simple terms, this technique serves as an in vitro “stress test” that gauges mitochondrial respiratory responses to a graded energetic challenge. Transitions from a high to low ATP:ADP ratio mimic an increase in energy demand, akin to a transition between rest and exercise (~VO2peak test). Analysis of the linear relationship between energy demand (ATP:ADP, ΔGATP) and steady-state oxygen flux (JO2) allows for an estimation of respiratory “conductance” (reciprocal of resistance), such that a steeper slope indicates greater sensitivity and improved kinetics (S. Figure 1B). Importantly, the energetic demands and thermodynamic constraints (backpressure imposed by ΔGATP) modeled by the CK clamp are comparable to those observed in vivo.
Figure 1. Mitochondria from mixed SkM are metabolically inflexible and intolerant to long chain lipid fuel.
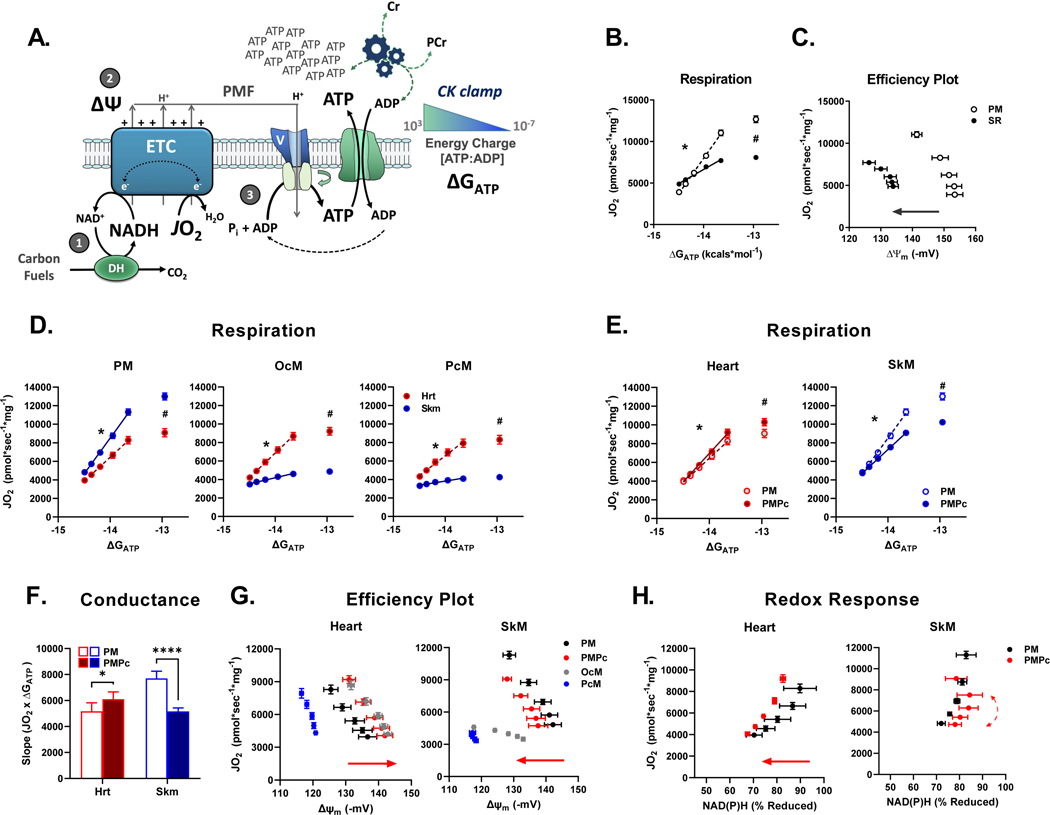
A. The CK clamp technique permits assessment of energy transduction in the context of near-physiological energy demands. Measures of (1) NAD(P)H/NAD(P)+ redox potential, (2) membrane potential ()–the major component of the proton motive force (PMF), and (3) overall respiratory flux (JO2) provide metrics of energy transfer and ATP turnover as a function of the ATP free energy charge (ΔGATP) established by the clamp.
B. Respiration plot of PM (5mM/2.5mM) versus SR (5mM/5μM) as a function of ΔGATP.
C. Respiratory efficiency plot in SkM mitochondria fueled by PM versus SR.
D. Comparison of heart and SkM mitochondria respiratory responses to PM, OcM or PcM.
E. Respiratory responses by heart and SkM mitochondria when Pc is added to PM.
F. Addition of Pc to PM augments respiratory conductance (slope) in heart but is inhibitory in SkM.
G. Addition of Pc to PM inhibits respiratory efficiency in SkM but not heart mitochondria (red arrows).
H. Redox response plot illustrating the relationship between NAD(P)H/NAD(P)+ redox potential and JO2 in heart and SkM mitochondria fueled by PM or PMPc. Curved arrow highlights redox instability with PMPc. See also Figure S1.
CK clamp assay conditions: Mitochondria (0.05 mg) from mouse Heart (Hrt), Mixed Skeletal Muscle (SkM), Red Gastrocnemius (RG) or White Gastrocnemius (WG) were added to 2 ml Buffer Z containing 5mM Creatine, 20U/ml Creatine Kinase, and 1.5mM phosphocreatine (PCr) at 37°C. The CK clamp was engaged upon addition of 5mM ATP. PCr was added to final concentrations of 3, 6, 9, 12, and 15 mM. Mitochondrial membrane potential () and redox status (NAD(P)H) were assessed in parallel in 0.2 ml Buffer Z containing 0.02 mg mitochondria and 0.2μM TMRM. Pyruvate/Malate (5mM/2.5mM; PM), Succinate/Rotenone (5mM/5μM; SR), Octanoyl-L-Carnitine/Malate (0.2mM/2.5mM; OcM), Palmitoyl-L-Carnitine/Malate (20μM/2.5mM; PcM), Pyruvate/Malate/Palmitoyl-L-Carnitine (5mM/2.5mM/20μM;PMPc). All substrates were provided at concentrations that are saturating for JO2.
Respiratory efficiency is defined as the relationship between oxygen flux (JO2) and mitochondrial membrane potential (). Arrows denote a shift in efficiency. Redox response refers to changes in (NAD(P)H/NAD(P)+ redox potential in relation to changes in JO2.
Statistics: n=5 biological replicates for PM/SR study. Data are presented as means ± SEM. Conductance slopes and maximal JO2 responses were tested with unpaired t-tests with Welch’s correction and Holms-Šídák correction for multiple comparisons.
* denotes differences in conductance slope, P<0.05, # = Maximal JO2 differences P<0.05.
Both the absolute rates of oxygen consumption and respiratory conductance (slope) depend on energy gradients and fluxes controlled by three principal regulatory nodes: 1) the dehydrogenase (DH) enzymes, 2) the ETC, and 3) ATP synthesis and transport, which together mediate the transfer of energy from that available in carbon substrates to ΔGATP (Figure 1A). To gain insight into the free energies that drive the transduction process, the foregoing dynamic JO2 assays are combined with parallel assessments of membrane potential (, the primary contributor to the proton motive force, PMF), and NAD(P)H/NAD(P)+ redox state. The example in Figure 1B depicts the respiratory phenotype of skeletal muscle mitochondria fueled by pyruvate/malate (PM) as compared to succinate/rotenone (SR) (S. Table 1, List of Abbreviations). Due to rotenone inhibition of complex I (CI) and heavy electron flux through complex II, the SR condition is characterized by a comparatively lower conductance (shallow slope) and less polarized . The resulting leftward shift in the relationship between JO2 and (Figure 1C) indicates that mitochondria are maintaining a lower (less polarized) for any given rate of oxygen consumption (Fisher-Wellman et al., 2018). In vivo, the energy harnessed in the determines the extent to which complex V (CV) can displace the ATP:ADP ratio from equilibrium. Thus, the ability of mitochondria to maintain and the relationship between and JO2 provide key metrics of respiratory efficiency and mitochondrial readiness to respond to an energy challenge.
Modeling metabolic flexibility in vitro.
Assessments of mitochondrial respiratory function typically involve experiments in which pyruvate and FA substrates are assayed independently. Notably however, one previous study aimed at comparing fuel selection in mitochondria from mammalian hindlimb versus avian flight muscle found that substrate competition and selection can occur at the level of the mitochondrial matrix 28. Here, we sought to investigate the interplay between pyruvate and FA substrates as an approach to modeling metabolic flexibility and mitochondrial lipid tolerance in vitro. As shown earlier (Fisher-Wellman et al., 2018)28, rodent SkM mitochondria have a strong preference for PM as compared with lipid substrates, octanoylcarnitine/malate (OcM) and palmitoylcarnitine/malate (PcM) (Figure 1D). By contrast, in heart mitochondria, JO2 and respiratory conductance are similarly robust when comparing across these three substrate conditions (Figure 1D). Mitochondrial lipid tolerance was tested by examining respiratory responses upon addition of Pc to PM. Notably, adding Pc to PM (PMPc) augmented both respiratory conductance (Figures 1E and 1F) and respiratory efficiency (Figure 1G) in heart mitochondria. Thus, as compared with PM alone, the PMPc respiratory efficiency plot shifted rightward (Figure 1G). Conversely, in SkM mitochondria, addition of Pc to PM reduced both JO2(peak) and conductance while also causing a leftward shift in respiratory efficiency, indicative of a lower (less polarized) for any given rate of oxygen consumption (Figure 1G). These phenomena are defined herein as Pc tolerance (Heart) and Pc intolerance (SkM). Interestingly, the Pc tolerant phenotype of mouse heart mitochondria resembles the profile observed in those from highly oxidative avian flight muscle 28.
Also notable, the long chain fatty acid (LCFA) substrate, PcM, was uniquely inefficient in heart mitochondria when provided without pyruvate. Importantly, in heart mitochondria, the dramatic leftward shift in response to PcM (Figure 1G), reflective of high respiration rates but a low (less polarized) , did not occur in the context of the medium chain FA substrate, OcM. Figure 1H highlights changes in NAD(P)H/NAD(P)+ redox potential in relation to JO2 in the context of PM as compared with PMPc. In heart mitochondria, this redox response plot shifted leftward upon addition of Pc to PM (PMPc), indicating that these mitochondria maintained a more oxidized redox state for any give rate of oxygen consumption when Pc and PM were added together. By contrast, when SkM mitochondria were provided with PMPc the relationship between redox and JO2 trended rightward (more reduced) at lower flux rates but shifted leftward as demand increased (Figure 1H). The curvilinear nature of this plot shows that respiratory inefficiency in SkM mitochondria fueled by PMPc is associated with redox instability.
In sum, we developed a mitochondrial lipid tolerance test that revealed striking tissue-dependent bioenergetic responses to a physiologic mixture of pyruvate and fatty acid fuels. Despite provision of saturating levels of pyruvate (PM), addition of LCFA (Pc) inhibited overall respiratory flux and efficiency in mitochondria derived from SkM, but not those from heart. This “Pc intolerant” phenotype coincided with redox instability and provides evidence that inefficient and/or incomplete FAO (i.e. FAO bottlenecking) might have deleterious thermodynamic consequences in the context of mixed substrate availability, particularly in SkM. Also remarkable was the discordance between JO2 and when heart mitochondria were fueled by PcM in the absence of pyruvate, which in theory could also stem from a bottleneck in LC FAO that destabilizes redox and compromises respiratory efficiency.
Mitochondrial flexibility and lipid tolerance are regulated by fiber type and Pgc1a.
Mitochondria from red fiber types are more lipid tolerant.
We next proceeded to determine if differences in Pc tolerance were evident in isolated mitochondria derived from SkM comprised of disparate fiber types (Figure 2A). As anticipated, PM-supported JO2 and conductance were similar between mitochondria from red gastrocnemius (RG) and white gastrocnemius (WG) (Figure 2B). Notably however, addition of Pc to PM caused much more dramatic reductions in JO2(peak) and conductance in mitochondria from WG as compared with those from RG (Figures 2B and 2C). Moreover, addition of Pc did not affect the respiratory efficiency plot in RG but caused a pronounced leftward and downward shift in WG (Figure 2D), indicating that mitochondria from RG maintained as JO2 increased, whereas those from WG did not. In a subsequent experiment we found that lipid-induced inhibition of respiratory conductance and respiratory efficiency in SkM mitochondria fueled by pyruvate is specific for Pc, as it does not occur with Oc (Figures 2E–2G). This observation aligns with the notion that LCFAs are more vulnerable to metabolic bottlenecks.
Figure 2. Mitochondrial flexibility and lipid tolerance are regulated by fiber type and Pgc1a.
A. Mitochondria were isolated from red (RG) and white (WG) gastrocnemius SkMs.
B. Respiratory responses to PM and PMPc in mitochondria from red gastrocnemius (RG) as compared with white gastrocnemius (WG).
C. Pc-induced inhibition of respiratory conductance is more pronounced in mitochondria from WG as compared with RG.
D. PMPc-supported respiratory efficiency is diminished in mitochondria from WG as compared with RG.
E-G. Addition of Pc but not Oc to PM inhibits respiratory conductance and efficiency in SkM mitochondria.
H. Mitochondria were isolated from SkM of non-transgenic (NT) mice and Mck-Pgc1a transgenic (Tg) littermates.
I. Mitochondrial input was normalized based on mitochondrial protein content. Expression of several proteins belonging to OXPHOS complexes were similar between groups.
J. Respiratory responses were measured in SkM mitochondria exposed to PM, PMPc and PMOc.
K-L. Genotype-specific responses in respiratory conductance (K) and/or respiratory efficiency (L) were observed in SkM mitochondria provided with PMPc or PMOc.
See also Figure S2.
CK clamp assay conditions: Mitochondria (0.05 mg) mitochondria from non-transgenic mouse mixed skeletal muscle (NT) or the same SkM from mice with muscle-specific Pgc1a overexpression (Tg) were added to 2 ml Buffer Z containing 5mM Creatine, 20U/ml Creatine Kinase, and 1.5mM phosphocreatine at 37°C. The CK clamp was engaged upon addition of 5mM ATP. PCr was added to final concentrations of 3, 6, 9, 12, and 15mM. Mitochondrial membrane potential () and redox status (NAD(P)H) were assessed in parallel in 0.2ml Buffer Z containing 0.02mg mitochondria and 0.2μM TMRM. Pyruvate/Malate (5mM/2.5mM; PM), Pyruvate/Malate/Palmitoyl-L-Carnitine (5mM/2.5mM/20μM;PMPc), Pyruvate/Malate/Octanoyl-L-Carnitine (5mM/2.5mM/0.2mM;PMPc). All substrates were provided at concentrations that are saturating for JO2.
Respiratory efficiency is defined as the relationship between oxygen flux (JO2) and mitochondrial membrane potential (). Arrows denote a shift in efficiency.
Statistics: n=6–7 biological replicates with data presented as means ± SEM. Conductance slopes and maximal JO2 responses were tested with unpaired t-tests with Welch’s correction and Holms-Šídák correction for multiple comparisons. * = conductance slopes differences P<0.05, # = Maximal JO2 differences P<0.05.
PGC1a promotes mitochondrial lipid tolerance.
To determine whether the Pc intolerant phenotype of white SkM mitochondria might be adaptable and responsive to physiological and molecular stimuli, we proceeded to evaluate mitochondria derived from transgenic (Tg) mice engineered to overexpress Pgc1a specifically in skeletal muscles (Figure 2H) 29. Not only is this prominent transcriptional co-activator and master regulator of mitochondrial biogenesis expressed abundantly in heart and red SkM, but its activation upon physical activity contributes to improvements in both FAO capacity and exercise endurance 12,30–32. Accordingly, we predicted that mitochondria from mixed SkM of Tg mice might shift to a “red-like” phenotype. Notably, mitochondria from Tg and non-transgenic (NT) SkM had similar amounts OXPHOS subunits belonging to complexes III, IV and V (Figure 2I) and only marginal increases in JO2 and/or respiratory conductance when supported by either PM (Figure 2J), PcM, or OcM (S. Figure 2A–2B). By contrast, clear phenotypes emerged in the context of PMPc (Figure 2J). Again, addition of Pc (but not Oc) to PM inhibited respiratory conductance in NT SkM mitochondria, whereas overexpression of Pgc1a in Tg mice enhanced Pc tolerance (Figure 2K) while also inducing a rightward shift in respiratory efficiency (Figure 2L). In aggregate, these experiments showed that mitochondria from red SkM are more lipid tolerant than those from white SkM, and that muscle-specific overexpression of PGC1a fully prevented Pc intolerance in a heterogeneous population of SkM mitochondria. Moreover, enhanced Pc tolerance in Pgc1a Tg SkM mitochondria was accompanied by only a marginal increase in PcM-supported FAO flux capacity and conductance (S. Figure 2A and 2B), suggesting that the mechanism at play was not related to robust upregulation of LC FAO machinery.
Surplus L-carnitine rescues lipid tolerance in SkM mitochondria independent of CrAT.
Surplus L-carnitine combats mitochondrial lipid intolerance.
Fatty acid oxidation is regulated in part by a family of carnitine acyltransferase enzymes that convert acyl CoA molecules of various chain lengths to their corresponding acylcarnitine metabolites. A member of this family with preference for short chain acyl CoAs, carnitine acetyltransferase (CrAT), has been shown to regulate metabolic flexibility in SkM 33–35. CrAT resides in the mitochondrial matrix and interconverts short chain acyl CoAs and acylcarnitine esters to buffer both the acetyl CoA and free CoA pools (Figure 3A). To test its role in Pc tolerance, bioenergetic responses to surplus L-carnitine were studied in SkM mitochondria from control mice and those lacking CrAT specifically in skeletal and cardiac muscles (Figure 3B). Aligned with the prediction, provision of surplus L-carnitine to SkM mitochondria had no effect when respiration was supported by PM (Figure 3C) but led to robust improvements in respiratory conductance (Figure 3C and 3D), respiratory efficiency (Figure 3E) and redox stability (Figure 3F) when added to PMPc. Although collectively the responses to L-carnitine were blunted in SkM mitochondria derived from muscle-specific CrAT knockout (mKO) mice (Figures 3G–3J), clear effects of L-carnitine remained evident. Thus, contrary to the expected result, carnitine-mediated improvements in SkM mitochondrial bioenergetics fueled by PMPc did not fully depend on CrAT enzyme activity, and thus likely involved CrAT-independent conversion of long and medium chain acyl CoAs to their cognate AC counterparts.
Figure 3. Surplus L-carnitine rescues lipid-induced respiratory inefficiency in SkM mitochondria.
A. CrAT is a carnitine-dependent acyltransferase enzyme with specificity for short chain carbon substrates.
B. Mixed SkM mitochondria isolated from CrATfl/fl control mice (Cont) or muscle-specific CrAT knockout (KO) littermates were provided with PM or PMPc ± surplus L-carnitine and respiratory responses were assayed using the CK clamp technique.
C-F. Respiratory response (C), respiratory conductance (D), respiratory efficiency (E), and redox stability (F) were enhanced in control mitochondria when surplus L-carnitine was added to PMPc but not PM.
G-J. Respiratory response (G), respiratory conductance (H), respiratory efficiency (I), and redox stability (J) were enhanced in control mitochondria when surplus L-carnitine was added to PMPc but not PM.
CK clamp assay conditions: Mitochondria (0.05 mg) from Cont or KO mouse mixed SkM were added to 2 ml Buffer Z containing 5mM Creatine, 20U/ml Creatine Kinase, and 1.5mM phosphocreatine at 37°C. CK clamp was engaged by the addition of 5mM ATP. PCr was added to final concentrations of 3, 6, 9, 12, and 15mM. Mitochondrial membrane potential () and redox status (NAD(P)H) were assessed in parallel in 0.2ml Buffer Z containing 0.02mg mitochondria and 0.2μM TMRM. Pyruvate/Malate (5mM/2.5mM; PM), Pyruvate/Malate/Palmitoyl-L-Carnitine (5mM/2.5mM/20μM; PMPc) ± 2mM L-Carnitine (Carn). All substrates were provided at concentrations that are saturating for JO2.
Respiratory efficiency is defined as the relationship between oxygen flux (JO2) and mitochondrial membrane potential (). Arrows denote a shift in efficiency. Redox response refers to changes in (NAD(P)H/NAD(P)+ redox potential in relation to changes in JO2.
Statistics: n=6 Cont and 5 KO biological replicates per group with data presented as means ± SEM. Conductance slopes and maximal JO2 responses were tested with unpaired t-tests with Welch’s correction and Holms-Šídák correction for multiple comparisons. * = conductance slopes differences P<0.05, # = Maximal JO2 differences P<0.05. For conductance comparisons in (D) and (H), all substrate conditions were tested for significance versus the PM condition and using Dunnett post-hoc testing with correction for multiple comparisons.
Mitochondrial metabolomics link lipid tolerance to free CoA and reverse short chain carbon flux.
Mitochondrial metabolomics and MFA.
In sum, results detailed in Figures 1–3 show that Pc inhibits pyruvate-supported respiratory fluxes in SkM but not heart mitochondria, and that overexpression of Pgc1a or addition of surplus free carnitine in vitro rescued Pc tolerance in the former. Although these findings are reminiscent of the well-known “Randle cycle” 36, wherein acetyl-CoA and NADH generated via FAO inhibits PDH activity, this mechanism appeared unlikely as the root cause of the SkM-specific impingement based on three key observations. First, we and others have shown that the classic Randle effect is robust in the heart. Secondly, PMOc did not mimic the respiratory profile of PMPc. Thirdly, the carnitine-mediated rescue of respiratory conductance and efficiency in SkM mitochondrial did not require CrAT, an enzyme that relieves negative feedback caused by acetyl CoA accumulation 35. We therefore hypothesized that the negative impact of adding Pc to PM in SkM mitochondria was a direct result of LC FAO bottlenecking. To further elucidate the site(s) of the presumed FAO bottleneck and the cause of Pc intolerance, we combined the CK clamp technique fixed at an intermediate energy demand with mass spectrometry-based metabolomics and metabolic flux analysis (MFA). Here, we used uniformly labeled [U-13C]pyruvate as the tracer, added in the absence or presence of Pc with and without surplus L-carnitine. Metabolites with two heavy carbons (M+2) originate from flux through pyruvate dehydrogenase (PDH), which generates M+2 acetyl units (Figure 4A). These substrates were provided to mitochondria isolated from NT hearts, NT SkM, and Pgc1a Tg SkM to assess three distinct mitochondrial pools characterized by disparate responses to PMPc.
Figure 4. Mitochondrial metabolomics and MFA link lipid tolerance to free CoA and reverse SC carbon flux.
A. Schematic of 13C pyruvate tracer experiment performed with isolated heart and SkM mitochondria from non-transgenic (NT) and Mck-Pgc1a transgenic (Tg) mice. Mitochondria were incubated with 5 mM [UL-13C]pyruvate and 2.5mM malate (PM) ± 20μM Pc ± 2mM L-carnitine in the context of 6 mM PCr to mimic moderate intensity exercise. The CK clamp was engaged upon addition of 5mM ATP. Five minutes after ATP addition, mitochondrial pellets were collected by centrifugation. Metabolites were extracted and measured by mass spectrometry.
B. Heat map representing log2 transformed row normalized Z-scores of log2 normalized metabolite abundances.
C-D. Free CoA (C) and Succinyl CoA (D) measured in pelleted heart and SkM mitochondria.
E-H. Acyl CoA metabolites measured in mitochondrial pellets.
I-L. Acylcarnitines measured in mitochondrial pellets.
M-O. Label incorporation from [UL-13C]pyruvate (M+2) into short chain metabolites.
P. Reverse short chain carbon flux estimated by measuring label incorporation into C4OH AC relative to C2. See also Figure S3.
Statistics: Data are presented as means ± SEM and were analyzed by two-way ANOVA (Tissue x Substrate) with no assumptions for equal variance and Dunnet’s correction applied for multiple comparisons. a = P<0.05 for Heart versus NT SkM. b = P<0.05 for Tg SkM versus NT Skm. $ = P<0.05 when PM+Pc condition was different from all other substrate combinations.
This experiment produced a rich (albeit complex) data set summarized as a heat map in Figure 4B. In aggregate, the metabolite profile showed that bottlenecking within the beta-oxidation spiral leads to accumulation of several acyl CoA intermediates and a corresponding reduction in mitochondrial concentrations of free CoA (i.e. CoA trapping). The Pc-induced decline in free CoA was most severe in SkM mitochondria (Figure 4C), which in turn appeared to limit conversion of alpha-ketoglutarate (aKG) to succinyl CoA in NT SkM but not Tg mitochondria (4B and 4D). Conversely, in heart mitochondria, the combination of PM+Pc expanded the succinyl CoA pool. Several enlightening observations are highlighted below.
First, Figure 4E shows that acetyl CoA availability was not a limiting factor in SkM mitochondria. Secondly, pronounced accumulation of C4 and C4OH acyl CoA upon addition of Pc (Figure 4F and 4G), combined with the absolute quantities of C4OH CoA as compared with other short (Figure 4F) and medium chain (Figure 4H) species, strongly implicated the C4 node of LC FAO as a major site of bottlenecking in both SkM and heart mitochondria (Figure 4A). As anticipated, addition of surplus L-carnitine lowered acyl CoAs, increased acylcarnitine metabolites (Figure 4I–4L) and restored levels of free CoA (Figure 4C). Interestingly however, C4OH (Figure 4K) was the only acylcarnitine species that decreased upon addition of L-carnitine (Figures 4B and 4K). The explanation for this odd result emerged from the mass isotopomer profiles (Figure 4M–4P). Consistent with the Randle cycle, addition of Pc lowered [U-13C]pyruvate flux into the acetyl pool, which was most dramatic in mitochondria with the highest FAO capacity (i.e. heart) (Figure 4M). Labeling of acetylcarnitine (C2) (Figure 4N) closely resembled M+2 acetyl CoA. Surprisingly, a substantial M+2 signal was detected in C4OH acylcarnitine (Figure 4O); indicative of flux from pyruvate-derived acetyl CoA to the C4OH-CoA pool via reverse flux through a ketothiolase (KT) enzyme and short chain hydroxy-acyl CoA dehydrogenase (SCHAD) (Figure 4A). M2 C4OH acylcarnitine was measurable only when Pc was present, and the signal declined upon addition of L-carnitine, mirroring the conversion of total acetyl CoA to total acetylcarnitine (Figures 4E and 4I). Thus, addition of surplus L-carnitine appeared to divert the large pool of acetyl CoA away from KT/SCHAD flux and towards the CrAT reaction (Figure 4N). These findings indicated that reverse short chain carbon flux was substantial when free carnitine was limiting. Relative levels of this flux in SkM and heart mitochondria exposed to PMPc were estimated by calculating the ratio of M2 C4OH:M2 acetylcarnitine (Figure 4P); approximately 0.6, 0.2 and 0.4 in heart, SkM and Tg mitochondria, respectively. This ratio correlated strongly with the total mitochondrial pool of free CoA (Figure 4C), which proved to be the best metabolite predictor of changes in respiratory efficiency in response to Pc exposure. In sum, these results showed that C4OH AC can be produced as a byproduct of either forward or reverse short chain carbon flux; and identified an intriguing connection between mitochondrial lipid tolerance and reverse acetyl CoA flux at the C4 node of FAO. We surmised that reverse flux of acetyl CoA to C4OH might support enhanced bioenergetics by regenerating both free CoA and NAD+, thereby diminishing dependence on surplus L-carnitine (Figure 4A).
To determine whether the metabolite profile generated by isolated mitochondria clamped at a heightened energy demand bears resemblance to that occurring in response to an acute bout of exercise, we assayed AC metabolites in heart and SkM obtained from mice that remained sedentary as compared to those that performed an acute graded treadmill test. Tissues were obtained from non-exercised control mice and two additional groups that were sacrificed either 5 min or 60 min following treadmill exercise. The heat map provided in S. Figure 3 shows that the exercise challenge led to: 1) a modest but significant decrease in free carnitine (C0) in SkM but not heart; 2) increased C4OH AC in both tissues; 3) a more dramatic rise in medium and long chain AC in SkM versus heart, and 4) a greater rise in levels of C4OH relative to other AC species in heart as compared with SkM. These profiles align with the prediction that increased contribution of reverse short chain carbon flux to the C4OH pool in heart mitochondria defends against LC FAO bottlenecking and L-carnitine depletion. The similarities between the tissue AC profiles and the mitochondrial assays implied that the in vitro model can indeed provide insights into fluxes and mechanisms of flux control that occur in vivo.
Mass spectrometry-based proteomics identifies MKT as a common feature of lipid-tolerance.
Mitochondrial proteomics.
To further elucidate the proteins and pathways that underlie the distinct metabolic and bioenergetic features of mitochondria from cardiac and skeletal muscles, we performed mass spectrometry-based proteomics on semi-purified mitochondria isolated from heart and mixed SkM. This analysis quantified 580 mitochondrial (Mitocarta 2.0 database) proteins that were detected in both tissues (Figure 5A, S. Table 2A and 2B, and Data S1). At a false discovery rate of 5%, 213 proteins were found to be more abundant in heart as compared with SkM mitochondria. Among the top 100 proteins on this list were 22 with established roles in FAO and/or CoA buffering, including Hadh (SCHAD) and Acaa2 (MKT) (Figure 5A and 5B, S. Table 2A, and Data S1). We then proceeded to perform a similar analysis on SkM mitochondria isolated from Mck-Pgc1a transgenic mice as compared with NT controls (Figure 5C and 5D, S. Table 3A and 3B, and Data S2). This analysis quantified 322 Mitocarta proteins and identified 64 proteins that were more abundant in Pgc1 Tg mitochondria as compared with NT controls when applying the same stringent cutoff of q<0.05 (Figure 5C). Surprisingly, only 8 of the proteins on this list were known to participate in FAO and/or CoA buffering. Top among those were Acaa2 (MKT), Acot13, Acadl and Acadvl (Figure 5C and 5D, S. Table 3A, and Data S2), which together might explain why mitochondria resident in Tg SkM had only slightly higher rates of PcM-supported respiration (Figure 2E) but a pronounced improvement in Pc tolerance. Also noteworthy, Pgc1a overexpression strongly upregulated mt-Co3 (mitochondrially encoded cytochrome C oxidase 3), possibly contributing to enhanced electron transfer capacity in Tg SkM mitochondria. When combining results of the two experiments, we identified only two proteins that were both more abundant in heart versus SkM as well at Tg versus NT controls–Acaa2 (MKT) and Acot13.
Figure 5. MS-based proteomics identifies MKT as a common feature of lipid-tolerant mitochondria.
Mass spectrometry (MS)-based proteomics was performed on semi-purified mitochondria isolated from heart as compared to SkM (n=5 biological replicates per group) and SkM of Mck-Pgc1a transgenic (Tg; n=6) mice compared to Non-transgenic (NT; n=5) littermate controls.
A. Volcano plot (FDR Adjusted P-Values as a function of Log2 fold change) showing proteins that are differentially abundant in mitochondria from heart vs. SkM. Fatty acid oxidation (FAO) proteins are highlighted in red and yellow denotes FAO proteins of high interest.
B. List of the top 15 FAO proteins that were more abundant in mitochondria from heart vs. SkM, ranked by Padj.
C. Volcano plot showing proteins that are differentially abundant in mitochondria from SkM of NT vs. Pgc1a Tg mice. FAO proteins are highlighted in red and yellow denotes FAO proteins of interest.
D. List of the top 15 proteins that were more abundant in SkM mitochondria from Tg vs. NT mice, ranked by Padj.
E. Relationship between proteins differentially abundant in Heart:SkM mitochondria and those changing in response to chronic transaortic constriction relative to sham controls (TAC:Sham) in mouse hearts. The dataset shows 110 proteins that were commonly identified in both sets and differentially abundant at FDR<1% and Log2 fold-change cutoff of at least ±0.5 in heart/SkM.
F. Quantitative results from western blots performed to measure abundance of MKT, SCHAD, CrAT, and CV subunit ATP5A1 in semi-purified mitochondria isolated from heart, RG and WG. semi-purified mitochondria isolated from heart, RG and WG.
G. Quantitative results from western blots performed to measure abundance of MKT, SCHAD, and CrAT in semi-purified SkM mitochondria isolated from NT compared Tg mice.
H. mRNA expression of Acaa2 (MKT) measured in mouse TA muscles harvested at rest or 5 min, 3 h or 24 h after a 90 min bout of acute treadmill exercise.
See also Figure S4.
Previous studies have shown that both Pcg1a and expression of FAO genes are diminished in mouse models of heart failure, in concert with increased accumulation of FA-derived acylcarnitines 32,37,38. We therefore questioned if mitochondria resident in failing hearts might remodel in a manner that shifts towards a SkM phenotype. To test this hypothesis, we compared the proteomics analysis of heart versus SkM (Figure 5A) to a similar analysis performed on mouse hearts exposed to 16 weeks of transaortic constriction (TAC) versus sham controls, as reported in 39. This analysis identified 110 proteins that were common to both sets and differentially abundant at an FDR<1% and Log2 fold-change cutoff of at least ±0.5 in heart/SkM. The resulting scatter plot (R=0.375, P<0.0001) in Figure 5E supports the prediction. Thus, we identified 67 proteins (many involved in FAO) that were both more abundant in heart:SkM mitochondria and less abundant in hearts of TAC:Sham (upper-left quadrant). Interestingly, the top down-regulated protein in TAC:Sham hearts was MKT (Acaa2).
Western blot analyses confirmed the fiber type-dependent expression of MKT and SCHAD, both of which were substantially more abundant in mitochondria isolated from heart and RG than those from WG (Figures 5F and S. 4A and 4B). By contrast, CrAT was only modestly enriched in heart mitochondria (Figure 5F and S. 4C) whereas levels of the complex I subunit, NDUFA9, complex II, and the complex V alpha subunit were similar among all mitochondrial populations, regardless of the tissue source (S. Figure 4D and 4E). Likewise, while OXPHOS proteins were similarly abundant in SkM mitochondria isolated from Tg mice as compared with NT controls (S. Figure 4F), levels of MKT were increased by ~2-fold in the former (Figures 5G and S. 4G). Notably however, mitochondrial SCHAD abundance (Figures 5G and S. 4H) was not affected by Pgc1a overexpression and CrAT was increased by only 33% (Figures 5G and S. 4I); both consistent with the notion that among all FAO-related proteins, MKT is uniquely induced in this model. Because both Pgc1a and mitochondrial FAO capacity are upregulated in response to habitual physical activity 40–42, we suspected that the Acaa2 gene might be induced during acute treadmill running. Indeed, a single bout of exercise led to a robust 3-fold increase in Acaa2 mRNA levels measured in mixed SkM (Figure 5H).
Reverse MKT flux is substantial in cardiac and skeletal muscle tissues and contributes to CrAT-dependent production of 3OH-butyrylcarnitine.
Reverse MKT flux measured in perfused hearts and incubated skeletal muscles.
In aggregate, our findings revealed a strong connection between MKT abundance and mitochondrial responses to LC FA substrate. Moreover, experiments in isolated mitochondria suggested that reverse MKT flux enhances lipid tolerance (Figure 4). To determine whether this same connection was evident in functional tissues, we performed MFA in perfused hearts and explanted soleus and EDL muscles incubated with various U-13C substrates (Figure 6A). Notably, rates of reverse MKT flux supported by [U-13C]glucose-derived acetyl CoA were even more robust in mouse hearts than those detected in purified heart mitochondria. Thus, labeling of both M+2 and M+4 C4OH AC (Figure 6B) and C4OH CoA metabolites (Figure 6C) approximated an enrichment of 20%. Interestingly, the same experiments performed in hearts from CrAT mKO mice suggested that this enzyme might act on C4OH-CoA. For example, CrAT null hearts generated 13C-labeled C4OH-CoA but not its acylcarnitine counterpart, despite similar tissue concentrations of both metabolites (Figure 6D). Likewise, hearts from control mice perfused with [U-13C] acetate showed robust labeling of both C4OH metabolites (Figures 6E and 6F), whereas label incorporation into C4OH acylcarnitine was reduced 50% in CrAT mKO hearts (Figure 6E) and the acetate-induced accumulation of this metabolite was largely abolished (Figure 6G). Similar experiments were performed in soleus (red) and EDL (mixed) muscles from wildtype mice incubated with [U-13C]glucose ±insulin (Figures 6H–6J). Again, results aligned with the in vitro assays; reverse flux of glucose-derived acetyl CoA was greater in soleus than EDL, and insulin responsive only in soleus (Figure 6J).
Figure 6. Reverse short chain carbon flux is substantial in cardiac and skeletal muscle tissues and contributes to CrAT-dependent production of 3OH-butyrylcarnitine.
A. Tracer strategy used for metabolic flux analysis (MFA) performed in perfused hearts harvested from MckCrAT KO mice and fl/fl littermate controls (B-G); or incubated soleus and EDL muscles (H-J).
B-C. [U-13C]glucose incorporation into 3-OH-butyrylcarnitine (B) and 3OH-butyryl-CoA (C) measured in perfused hearts isolated from Mck-CrAT KO mice and fl/fl controls.
D. Heart concentrations of total 3OH-butyrlcarnitine after perfusion with 11 mM [U-13C]glucose.
E-F. [U-13C]acetate incorporation into 3-OH butyrylcarnitine (E) and 3-OH butyryl CoA (F) measured in perfused hearts isolated from Mck-CrAT KO mice and fl/fl controls.
G. Heart concentrations of total 3-OH-butyrlcarnitine after perfusion with 1 mM [U-13C]acetate.
H-I. [U-13C]glucose incorporation into short chain acylcarnitines, including 3-OH-butyrlcarnitine (C4OH), measured in soleus (H) and EDL (I) muscles isolated from wildtype mice and incubated ex vivo in the absence and presence of 100 nM insulin.
J-K. Reverse short chain carbon flux estimated by measuring label incorporation into short chain acylcarnitines (AC) relative to acetylcarnitine (C2) in soleus (J) and EDL (K) muscles.
Statistics: Heart perfusions were performed with n=4 biological replicates per group for 13C-glucose and 6 biological replicates for [U-13C]acetate tracer experiments. Means were tested using Unpaired t-tests with Welch’s correction and Holms-Šídák correction for multiple comparisons. Small muscle incubations were performed with n=6 biological replicates for soleus and EDL. ‡ = main effect of insulin across the reported species.
Insufficient MKT flux results in lipid-induced respiratory inefficiency in heart mitochondria.
Reverse MKT flux confers mitochondrial lipid tolerance.
To gain further insights into the interplay between lipid tolerance and pyruvate flux through MKT, heart mitochondria were exposed to PcM in combination with titrated concentrations of pyruvate ranging from 120 μM to 5 mM (S. Figure 5A.). Two important observations emerged. First, a concentration of 0.6 mM pyruvate was sufficient to fully restore respiratory efficiency (S. Figure 5B). Secondly, the progressive rightward shift in respiratory efficiency as pyruvate concentration increased from 120 μM to 5 mM (S. Figure 5B) coincided perfectly with a more stabilized NADH/NAD+ redox potential (S. Figure 5C). This observation aligns with the premise that reverse flux through MKT and SCHAD establishes a circuit that buffers both the free CoA pool as well as NAD(P)H/NAD+ electron energy potential, which in turn optimizes energy transfer, and respiratory efficiency. In isolated mitochondria fueled by PM±Pc, this circuit was engaged only when mitochondria were provided with a mixture of pyruvate and Pc (PMPc) (Figure 4K), presumably because the FAO bottleneck and CoA trapping caused by addition of Pc establishes the driving forces necessary to push reverse MKT flux.
Assuming that reverse MKT flux is essential for mediating Pc tolerance, replacement of pyruvate with a respiratory substrate that does not expand the acetyl CoA pool should render heart mitochondria more susceptible to Pc-induced respiratory inefficiency. To test this hypothesis, heart and SkM mitochondria fueled by aKG were assayed in the absence and presence of Pc (Figure 7A). As predicted, in this context, addition of Pc dampened conductance (Figure 7B–7C) and caused a pronounced leftward shift in respiratory efficiency, regardless of the tissue source of the mitochondria (Figure 7B). Thus, when pyruvate was replaced by aKG, heart and SkM mitochondria were similarly vulnerable to Pc-induced respiratory inefficiency.
Figure 7. Insufficient MKT flux results in lipid-induced respiratory inefficiency in heart mitochondria.
A-C. Isolated heart and skeletal muscle (SkM) mitochondria were exposed to aKG±Pc followed by assessments of respiration (A), respiratory efficiency (B), and conductance (C). In the absence of PM, addition of Pc inhibits respiratory conductance and disrupts efficiency and redox stability in both heart and SkM mitochondria.
CK clamp assay conditions: Mitochondria (0.05 mg) isolated from mouse Heart (Hrt) or Mixed Skeletal Muscle (SkM) were added to 2 ml Buffer Z containing 5mM Creatine, 20U/ml Creatine Kinase, and 1.5mM phosphocreatine at 37C. CK clamp was engaged upon addition of 5mM ATP. PCr was added to final concentrations of 3, 6, 9, 12, and 15mM. Mitochondrial membrane potential () and redox status (NAD(P)H) were assessed in parallel in 0.2 ml Buffer Z containing 0.02mg mitochondria and 0.2μM TMRM. Alpha-Ketoglutarate (5mM; aKG), alpha-Ketoglutarate/Palmitoyl-L-Carnitine (5mM/20μM; aKG+Pc). All substrates were provided at concentrations that are saturating for JO2.
Respiratory efficiency is defined as the relationship between oxygen flux (JO2) and mitochondrial membrane potential (). Redox response refers to changes in (NAD(P)H/NAD(P)+ redox potential in relation to changes in JO2. Redox transfer refers to the transfer of electron energy potential (NAD(P)H/NAD(P)+ redox) to proton energy potential, which is informed by the relationship between NAD(P)H/NAD(P)+ redox potential and mitochondrial membrane potential ().
Statistics: n=6 biological replicates per condition. Data are presented as means ± SEM. Conductance slopes and maximal JO2 responses were tested with unpaired t-tests with Welch’s correction and Holms-Šídák correction for multiple comparisons. * = conductance slopes differences P<0.05, # = Maximal JO2 differences P<0.05.
D-G. Lipid tolerance was assessed in differentiated primary human skeletal myocytes (HSkMC) using a CK clamp adapted for use with the Seahorse Flux Analyzer. Western blot analysis (15 μg protein per lane) of MKT protein abundance in HSkMC is comparable to heart tissue (D). Addition of 25 uM Pc to PM does not affect respiratory conductance whereas 50 Pc is inhibitory, while 100 uM Oc has no effect (E). MKT-targeted siRNAs diminished protein abundance in HSkMC as compared with cells treated with non-targeting control siRNA (NTC) (F). Knockdown (KD) of MKT protein in HSkMC diminishes respiratory conductance measured with PM+25 uM Pc (G).
Statistics: Conductance measures are from n=4 (E) and n=7 (G) independent Seahorse flux experiments, each with at least 4 technical replicates per condition (i.e. 2 siRNA treatments x 2 substrate mixtures). Data are presented as means ± SEM of n=4 and n=7, respectively. Conductance slopes calculated from protein normalized JO2 measures were analyzed by paired t-tests; * = P<0.05. Quantification of MKT western analysis (F) represents n=7 biological replicates per treatment, each representative of 15–30 technical replicates pooled from 96-well Seahorse plates after permeabilization and JO2 assessment shown in (G).
H-N. Heart mitochondria were isolated from mice that were fed ad libitum or fasted for 6, 12 or 24 hours (H). Blood metabolites (I) were measured by glucometer and ketometer. Mitochondrial assays were performed in parallel (i.e. 4 mice per day, one from each of the 4 timepoints) upon exposure to 0.55 mM aKG + 2 mM beta-hydroxy butyrate (3OHB) followed by assessments of respiratory efficiency (J) and redox response (K).
L-M. Western blot (L) and the corresponding relative quantitation (M) of pAMPK relative to total AMPK in hearts of mice that were fed ad libitum or fasted for 6, 12 or 24 hours.
N. Cardiac function assessed by echocardiography performed in mice that were fed or starved overnight (OS).
O. Working model of a short chain carbon circuit that confers lipid tolerance and metabolic flexibility. AC, acylcarnitine; CoQ, Coenzyme Q; CrAT, carnitine acetyltransferase; ETC, electron transport chain; ETF; electron-transfer flavoprotein system; LC, long chain, MC, medium chain; MKT, medium chain ketothiolase; SC, short chain; SCHAD, short chain hydroxyacylCoA dehydrogenase.
See also Figure S5.
Fasting time course: Mitochondria (0.05 mg) isolated from mouse heart mitochondria were added to 2 ml Buffer Z containing 5mM Creatine, 20U/ml Creatine Kinase, and 1.5mM phosphocreatine at 37°C. CK clamp was engaged upon addition of 5mM ATP. PCr was added to final concentrations of 3, 6, 9, 12, and 15mM. Mitochondrial membrane potential () and redox status (NAD(P)H) were assessed in parallel in 0.2ml Buffer
Z containing 0.02mg mitochondria and 0.2μM TMRM. N=6 biological replicates per timepoint. alpha-Ketoglutarate/3OH-butyrate (0.55mM/1mM; aKG/3OHB).
Statistics: n=6 biological replicates per condition. Data are presented as means ± SEM. Differences in blood metabolites were tested with a one-way ANOVA for differences versus 0 h using Dunnett’s correction for multiple comparisons. Conductance slopes and maximal JO2 responses were tested with unpaired t-tests with Welch’s correction and Holms-Šídák correction for multiple comparisons. * = conductance slopes differences P<0.05, # = Maximal JO2 differences P<0.05.
To further vet the hypothesis that MKT plays a key role in the proposed short chain carbon circuit, we performed loss-of-function experiments in fully differentiated primary human skeletal myocytes (HSkMC). Interestingly, baseline expression of the MKT protein in HSkMC was comparable to that measured in mouse heart tissue (Figure 7D). After optimizing the CK clamp technique for use with permeabilized HSkMC and the Seahorse Flux Analyzer (S. Figure 5D–F), we found that robust expression of MKT was accompanied by a high level of Pc tolerance in these cells. Thus, addition of 25 uM Pc to PM was tolerated without affecting respiratory conductance, whereas 50 uM Pc was inhibitory while 100 uM Oc had no effect (Figure 7E). Next, treatment of HSkMC with MKT-targeted siRNA led to ~50% knockdown of the protein as compared with the control group treated with non-targeting siRNA sequences (Figure 7F and S. Figure 5G). As predicted, knockdown of MKT lowered respiratory conductance in the context of PM+25 uM Pc (Figure 5G), supporting the notion that MKT plays a key role in conferring mitochondrial lipid tolerance.
The findings generated using the foregoing in vitro and ex-vivo systems raised the question of whether or not lipid-induced respiratory inefficiency can occur in vivo when hearts and/or SkMs are deprived of sufficient glucose and pyruvate. To address this question, we performed a time course study to evaluate the impact of prolonged fasting on heart mitochondrial bioenergetics. Mice were housed individually and food was removed at 8 am, 8 pm or 2 am to compare 24, 12 and 6 h fasting to ad libitum fed (0 h) mice. Due to the nature of the parallel study design (Figure 7H), these experiments were limited to three fuel conditions. Those selected featured substrates known to be especially sensitive to changes in redox and CoA buffering capacity (PcM, aKG and aKG+3OHB). As anticipated, fasting led to a time-dependent decline in blood glucose levels and a reciprocal rise in serum NEFAs and ketones (Figure 7I). Notably, the transition to heavy systemic reliance on lipid fuels corresponded with a leftward shift in mitochondrial respiratory efficiency (Figure 7J and S. Figure 7I), a progressive rightward shift in the NADH/NAD+ redox response (Figure 7K), a heightened rate of electron leak (S. Figure 5I), and induction of pAMPK (Figure 7L and 7M), a molecular marker of energy stress. By contrast, mitochondrial abundance of several OXPHOS proteins remained unchanged (S. Figure 5J).
Lastly, we sought to determine if a prolonged fast and the accompanying leftward shift in respiratory efficiency assessed in isolated heart mitochondria coincided with detectable changes in cardiac function. To this end, echocardiography was performed in the fed and overnight starved state using a cohort of C57BL6 mice available in our colony. The results revealed a modest decline in measures of systolic function (Figure 7N). Taken together with Figures 4B–D, these findings point to a Pc-induced bottleneck in the LC FAO pathway that can push free CoA levels below a critical threshold when reverse MKT flux is limited, due either to low abundance of the enzyme and/or insufficient substrate (i.e., driving forces). The results align with the premise that glucose insufficiency constrains the SC carbon circuit (Figure 7O) and can thereby compromise free CoA availability, redox buffering, and energy stability in vitro and in vivo.
DISCUSSION
This investigation combined a sophisticated mitochondrial phenotyping platform with state-of-the-art molecular profiling tools and multiple two-state mouse models of enhanced or diminished mitochondrial respiratory function to uncover a mechanism that connects reverse MKT flux to metabolic flexibility, lipid tolerance and energy stability in SkM and heart mitochondria. The findings align with the widely reported yet imperfect association between systemic acylcarnitine accumulation and cardiometabolic diseases; and help to explain why reduced FAO capacity, high rates of incomplete beta-oxidation, and metabolic inflexibility are tightly linked to poor health outcomes. The discovery that pyruvate plays a prominent role in supporting reverse MKT flux gives expanded meaning to the axiom, “fatty acids burn in a carbohydrate flame.”
The experiments described and the ensuing interpretations were guided by four seminal observations that revealed a heretofore unknown mechanism controlling the interplay between lipid and glucose-derived fuels and its impact on mitochondrial bioenergetics. First, in SkM mitochondria, provision of LCAC impaired pyruvate-supported respiratory efficiency. This “lipid intolerance” phenomenon was less pronounced in mitochondria from red SkM and absent in those from heart. Mitochondrial intolerance to LCFA was subsequently linked to a bottleneck in the LC FAO spiral that results in CoA trapping and an attendant carbon traffic jam that appeared to coincide with reduced flux at aKG dehydrogenase. This unfavorable respiratory phenotype emerged when the matrix pools of both free carnitine and CoA were limiting. The results show that incomplete oxidation of LCFA can impact flux of non-lipid substrates, thereby compromising energy transfer and respiratory efficiency. Also noteworthy, we found that acylcarnitine accumulation and/or a bottleneck in FAO is not necessarily detrimental, provided that the matrix pools of free CoA and free carnitine are maintained above a critical threshold. We therefore surmise that acylcarnitine metabolites serve as biomarkers of mitochondrial congestion but do not act as direct culprits of mitochondrial dysfunction. Instead, these molecules act as vehicles that permit mitochondrial carbon efflux and oppose CoA trapping to defend metabolic stability.
Secondly, we discovered that mitochondrial tolerance to a LCFA substrate correlates with relative abundance of MKT and rates of reverse flux through this enzyme. Although the bidirectional nature of the MKT reaction has been described previously 43, the crucial importance of glucose-derived acetyl CoA as an MKT substrate and the role of reverse flux in preserving mitochondrial respiratory efficiency had not been recognized. Importantly, pyruvate/glucose-supported reverse MKT flux was substantial in isolated mitochondria as well as perfused hearts and excised intact muscles. Moreover, proteomics analysis of “red” versus “white” mitochondria showed that among all FAO enzymes quantified, MKT was uniquely abundant in mitochondria with a high tolerance to LCFA substrate. Additionally, MKT (Acaa2) and Acot13, a thioesterase with strong preference for medium and long chain acyl CoAs 44, were found to be the most upregulated FAO-related proteins in SkM mitochondria derived from Mck-Pgc1a Tg mice as compared with NT controls. Fitting with these results and resembling the phenotype of heart mitochondria, Tg SkM mitochondria better maintained free CoA levels during a lipid challenge. Additionally, we found that Acaa2 mRNA expression was potently induced in response to a single bout of strenuous exercise. Taken together, these findings imply that exercise-induced upregulation of MKT contributes to enhanced metabolic flexibility and possibly improved muscle performance during high work rates.
Thirdly, this study also showed that high capacity for FAO introduces a thermodynamic vulnerability unique to LCFA. Thus, exposure of heart mitochondria to LCFA substrate without sufficient pyruvate resulted in profoundly diminished despite high rates of oxygen consumption. This remarkable profile of respiratory inefficiency could be due to inefficient electron flow. In this context, regeneration of free CoA appears sufficient to maintain high rates of incomplete FAO; however, compromised NADH/NAD+ redox buffering results in imbalanced flow of electrons through FADH2-linked enzymes (e.g. LCAD and MCAD) relative to flux through NADH-linked complexes (Figure 7O). This imbalance could be detrimental because FADH2 delivers electrons directly to the coenzyme Q pool, bypassing proton pumping at complex I, resulting in less energy transfer to Δp per atom of oxygen consumed at complex IV, and thus a lower P:O ratio (i.e. similar to SR, Figure 1B and 1C). In some settings, increased FAD-linked electron flux could lead to complete reduction of coenzyme Q, more electron leak and/or lower rates of electron transfer at complex I, leading to a rise in matrix NADH/NAD+ and further inhibition of forward dehydrogenase flux. Thus, a sizable shift from NADH to FADH2-linked respiration when glucose and pyruvate become limiting would be predicted to increase ROS generation and diminish work efficiency. A key question is whether this form of respiratory inefficiency occurs in vivo. Supportive evidence comes from experiments wherein an overnight fast, which lowers blood glucose and increases acylcarnitine accumulation in cardiac and skeletal muscle tissues 11,19,45,46, was accompanied by a modest but measurable decline in cardiac performance, assessed by echocardiography. Additionally, MKT was found to be strongly downregulated in failing mouse hearts 39, which are characterized by diminished FAO and PDH flux along with redox imbalance 32,47,48. Likewise, two recent studies showed that heart failure in mice and humans is accompanied by reduced expression of the mitochondrial pyruvate carrier (MPC); and that cardiac-specific disruption of MPC results in contractile dysfunction, tissue accumulation of LC acylcarnitines, and profound reductions in both free carnitine and free CoA 38,49. In aggregate, these results imply that mitochondrial inflexibility due to physical inactivity and/or pathological metabolic remodeling (including reduced expression of MKT and MPC) exacerbates lipid-induced mitochondrial stress and thereby threatens energy stability and thus overall muscle function and cardiometabolic health.
Another notable finding underscored by this work is that PcM and OcM, provided alone or in combination with pyruvate, produced distinct bioenergetic profiles in heart and SkM mitochondria. This is a key observation because Pc and Oc have identical stoichiometry of NADH-linked versus FADH2-linked electron fluxes; thus, the chain length-specific responses appear to stem from inefficiencies introduced by the LC arm of the pathway. Also noteworthy, the findings reported herein align with in silico metabolic control simulations predicting that beta-oxidation of LCFA introduces thermodynamic vulnerabilities distinct from those of MC FAO; and that MKT is a major site of negative flux control in the context of elevated lipid substrate 43,50,51. In this simulation, flux decline at the MKT reaction was due both to enzyme promiscuity and well as the highly unfavorable equilibrium constant of the preceding enzyme, SCHAD, such that a high NADH/NAD+ redox state pushes the reaction in the reverse direction. Whereas results of the model simulation fit well with the current and previous investigations identifying C4OH acylcarnitine as a strong metabolite marker of fuel switching and substrate competition 11,32,46,52–54, the findings reported herein now unveil the evolutionary advantage of reverse MKT/SCHAD flux as a mechanism that stabilizes NADH/NAD+ redox potential while also combating catastrophic CoA depletion caused by bottlenecks in LC FAO.
Lastly, these results add two new elements to the Randle cycle by showing that 1) incomplete/inefficient FAO is particularly detrimental to pyruvate oxidation, and 2) restricted PDH flux can diminish FAO efficiency. Whereas the conventional version of the Randle cycle centers on the concept of substrate competition, the model proposed herein highlights the importance of cooperation between these two principal mitochondrial fuels. Together, the findings could explain why metabolically flexible mitochondria confer a survival advantage at the organismal level. This fundamental conceptual advance opens exciting opportunities for understanding the clinical connections between acylcarnitine accumulation, mitochondrial performance, and metabolic resilience.
LIMITATIONS OF STUDY.
Studies performed in isolated mitochondrial systems have caveats and limitations. For example, it is not only possible but highly probable that substrates other than pyruvate and fatty acids contribute to the SC carbon circuit and/or redox and CoA buffering. Thus, the bioenergetic and thermodynamic conditions established in isolated mitochondria exposed to PM, PcM and PMPc do not fully mimic those that occur in vivo. Additionally, the mitochondrial system does not capture regulation imposed by extra-mitochondrial mechanisms. Conversely, the in vitro “stress tests” provide information and insights that cannot be gained by any other approach. Experimental options for assessing mitochondrial energetics and efficiency in vivo are limited, especially in the context of increased work rates (exercise). Included among the key advantages of the in vitro system is the highly tractable nature of the model. Substrate supply and energy demand can be precisely controlled; and mitochondrial carbon fluxes can be studied and interpreted under steady state kinetics without confounding assumptions inherent to multi-compartment models, including regulation of FAO by the carnitine palmitoyltransferase system. Provocative results can then be fully vetted using MFA applied to whole tissues and/or in vivo. Notably, the phenotypes and fluxes observed in the context of the CK clamp assay strongly resemble those identified in whole tissues under comparable conditions. Another noteworthy limitation of the current mitochondrial diagnostics platform is that and redox potential are measured in parallel with JO2 using open chamber microcuvettes positioned in a PTI QuantaMaster spectrofluorometer. Thus, the volume and oxygen supply during these assays are not identical to those applied to the O2K system. Additionally, we are left to infer shifts from NAD- to FAD-linked electron transfer. Also, the mitochondrial diagnostics platform employed by these studies uses TMRM emission spectra at 590 nm following excitation at both 551 and 552 nm to estimate by converting the 572/551 ratio to millivolts, based on a KCl calibration curve performed in the presence of valinomycin. While the KCl titration experiments in the presence of valinomycin allow for mitochondrial quantification, the resulting absolute voltages should be regarded as approximations. Such calculations are of course only as accurate as the inherent assumptions, which for TMRM based -quantification include: 1) equal mitochondrial protein content between groups, and 2) a matrix potassium concentration of 120mM. Validation of this approach is provided in 1.
Lastly, while the results strongly support the proposed working model of the short chain carbon circuit, an Acaa2 knockout mouse model is currently unavailable and therefore we are unable to determine whether targeted loss of MKT per se in heart and/or red SkM would be sufficient to compromise cardiac function and/or exercise tolerance. Nonetheless, a null physiological phenotype of MKT-deficient mice would not contradict the working model, but instead imply that heart and red SkM harbor redundant pathways and mechanisms for buffering the matrix free pools of free CoA and NAD+, which seems highly likely given the prominent role of these cofactors in maintaining energy homeostasis.
STAR METHODS
Resources Availability
Lead contact
Further information and requests for resources and reagents should be directed to and will be fulfilled by the lead contact, Deborah Muoio, PhD (muoio@duke.edu).
Materials availability
This study did not generate new unique reagents.
Data and code availability
Proteomics data have been deposited at ProteomeXchange and jPOST and are publicly available as of the date of publication. Accession numbers are listed in the key resources table. Original western blot images have been shown in Supplemental data. A file with all data presented in figures has been including in Supplemental data.
This manuscript does not report any original code.
Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| MKT (Acaa2) | Millipore Sigma | WH0010449M1; RRID:AB_2219394 |
| CrAT | ABCAM | ab153750; RRID:AB_2935643 |
| SCHAD | ProteinTech | 19828–1AP; RRID:AB_10667408 |
| ATP5A | ABCAM | ab14748; RRID:AB_301447 |
| OxPHOS Cocktail | ab110413; RRID:AB_2629281 | |
| SDHA | ABCAM | ab14715; RRID:AB_301433 |
| NDUFA9 | ABCAM | ab14713; RRID:AB_301431 |
| T-AMPK | CST | 2532; RRID:AB_330331 |
| P-AMPK | CST | 2531; RRID:AB_330330 |
| HSP60 | CST | 12165; RRID:AB_2636980 |
| Chemicals, peptides, and recombinant proteins | ||
| MOPS Free Acid | Millipore Sigma | Cat# M1254 CAS# 1132–61-2 |
| MES Potassium Salt | Millipore Sigma | Cat# M0895 CAS# 39946–25-3 |
| Bovine Serum Albumin (Fatty Acid Free) | Millipore Sigma | Cat# A3803 CAS# 9048–46-6 |
| EDTA | Millipore Sigma | Cat# E0270 CAS# 65501–24-8 |
| Trypsin from Porcine Pancreas (Mitochondrial Isolation) | Millipore Sigma | Cat# T4799 CAS# 9001–51-8 |
| Potassium Chloride | Millipore Sigma | Cat# P5405 CAS# 7447–40-7 |
| Magnesium Sulfate | ||
| Magnesium Chloride Hexahydrate | Millipore Sigma | Cat# M2670 CAS# 7791–18-6 |
| EGTA | Millipore Sigma | Cat# E4378 CAS# 67–42-5 |
| Potassium Dihydrogen Phosphate | Millipore Sigma | Cat# P9791 CAS# 7778–77-0 |
| Creatine Monohydrate | Millipore Sigma | Cat# C3630 CAS# 6020–87-7 |
| Potassium Salt of Phosphocreatine | Millipore Sigma | |
| Palmitoyl-L-carnitine | Millipore Sigma | Cat# P1645 CAS# 18877–64-0 |
| Octanoyl-L-carnitine | Millipore Sigma | Cat# 50892 CAS# 25243–95-2 |
| L-Carnitine hydrochloride | ||
| Malic Acid (Malate) | Millipore Sigma | Cat# M1000 CAS# 97–67-6 |
| α-ketoglutaric acid (aKG) | MilliporeSigma | Cat# K1750 CAS# 328–50-7 |
| (R)-3-Hydroxybutyric acid | Millipore Sigma | Cat# 54920 CAS# 625–72-9 |
| Succinic Acid (Succinate) | Millipore Sigma | Cat# S3674 CAS# 110–15-6 |
| Potassium Pyruvate | Combi-Blocks | Cat# QA-1116 CAS# 4151–33-1 |
| Creatine Kinase from Rabbit Muscle | Roche | Cat# 10127566001 |
| Tetramethylrhodamine Methyl Ester (TMRM) | ThermoFisher | Cat# T668 |
| Rotenone | Millipore Sigma | Cat# R8875 CAS# 83–79-4 |
| Potassium Cyanide | Millipore Sigma | Cat# 60178 CAS# 151–50-8 |
| Alamethicin | AG Scientific | A-1286 CAS#27061–78-5 |
| Methanol | Millipore Sigma | Cat# 439193 CAS# 67–56-1 |
| Protease Inhibitor Cocktail | Millipore Sigma | Cat# P8340 |
| Phosphatase Inhibitor Cocktail 2 | Millipore Sigma | Cat# P5726 |
| Phosphatase Inhibitor Cocktail 3 | Millipore Sigma | Cat# P0044 |
| Pierce Reversible Protein Stain Kit for Nitrocellulose Membranes (Memcode) |
ThermoFisher Scientific |
Cat# 24580 |
| 4–15% Criterion TGX Stain-Free Protein Gel, 18well | Biorad | Cat# 5678084 |
| 10X Tris Glycine SDS Running Buffer | Biorad | Cat# 1610732 |
| 10X Tris Buffered Saline | Biorad | Cat# 1706435 |
| Fish Gelatin | Millipore Sigma | Cat# G7765 |
| Casein | Millipore Sigma | Cat# C0626 |
| Roche cOmplete ULTRA EDTA-free Protease Inhibitor Mini Tablet | Millipore Sigma | Cat# 05892791001 |
| Roche 1x PhosSTOP Phosphatase Inhibitor Cocktail Tablets | Millipore Sigma | Cat# 04906837001 |
| Lysyl Endopeptidase, Mass Spectrometry Grade | Wako Chemicals | Cat# 125–05061 |
| Sequencing Grade Modified Trypsin | Promega | Cat# V5113 |
| tC18 SEP-PAK Solid Phase Extraction Columns (50 mg) | Waters | Cat# WAT054960 |
| tC18 SEP-PAK Solid Phase Extraction Columns (100 mg) | Waters | Cat# WAT036820 |
| Triethylammonium bicarbonate (TEAB) | ThermoFisher | Cat# 90114 |
| TRIzol Reagent | ThermoFisher | Cat# 15596026 |
| Chloroform | Millipore Sigma | Cat# C2432 CAS# 67–66-3 |
| [U-13C]-Glucose | ||
| [U-13C]-Pyruvate | ||
| CelLyticmtCelLytic™ MT Cell Lysis Reagent | SigmaAldrich | C3228 |
| Pentobarbital (Nembutal) | Oak Pharmaceuticals | |
| Critical commercial assays | ||
| Pierce BCA Protein Assay | ThermoFisher | Cat# 23225 |
| RNeasy mini spin columns | Qiagen | Cat# 74004 |
| iScript cDNA Synthesis Kit | BioRad | Cat# 170–8891 |
| PrimeTime Gene Expression Master Mix | IDT | Cat# 1055770 |
| TaqMan Gene Expression Assay Acaa2 | ThermoFisher | Mm00624282 |
| Lipofectamine RNAiMAX | ThermoFisher | 13778 |
| XF Plasma Membrane Permeabilizer | Agilent Technologies | 102504–100 |
| Deposited data | ||
| Proteomics Raw Data Files | This Publication | PXD040326; JPST002051 |
| Proteomics Raw Data Files | https://pubmed.ncbi.nlm.nih.gov/32660330/ | Experiment 3; Supplemental data; Protein SUMMARY_Exp3 Tab |
| Data S4 (Excel File) | This Publication | |
| Experimental models: Organisms/strains | ||
| C57BL/6NJ mice | The Jackson Laboratory | Stock #005304 |
| MCK-Pgc1a C57BL/6-Tg(Ckm-Ppargc1a)31Brsp/J | Dr. Bruce Speigelman | https://pubmed.ncbi.nlm.nih.gov/12181572 |
| MCK-CrAT C57BL/6J | Dr. Randall Mynatt (Pennington Biomedical Research Center) | https://pubmed.ncbi.nlm.nih.gov/22560225/ |
| Oligonucleotides | ||
| ACAA2/MKT siRNA | IDT | hs.Ri.ACAA2 - 13.1, 13.2, 13.3 |
| Non-targeting control sequence | IDT | 51–01-14–03 |
| Software and algorithms | ||
| Proteome Discoverer 2.2 | ThermoFisher | RRID:SCR_014477 |
| DatLab 7 | Oroboros Instruments | Product# 20700 |
| Other | ||
| Oxygraph-2k | Oroboros Instruments | Product# O2k-Core |
| QuantaMaster Spectrofluorometer | Horiba Scientific | Cat# QM-400 |
| Spectrosil Quartz Cuvettes | Starna | 23–5.45-Q-5 |
| Spectromax M2E Spectrophotometer | Molecular Devices | Part#: M2E |
| Thermo Fisher Scientific Q Exactive Plus Orbitrap Mass Spectrometer | ThermoScientific | Cat#: 0726030 |
| Thermo Fisher Scientific nanoEASY nLC | ThermoScientific | Cat #: LC140 |
| Waters Xevo TQ-S triple quadrupole mass spectrometer coupled to a Waters Acquity UPLC system | Waters | Part#: Xevo TQ-S |
| Bio-Rad Turboblot Transfer System | Biorad | Cat# 1704150EDU |
| TissueLyser II | Qiagen | Cat# 85300 |
| NanoDrop 8000 | ThermoScientific | ND-8000-GL |
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
| Antibodies | ||
| Rabbit monoclonal anti-Snail | Cell Signaling Technology | Cat#3879S; RRID: AB_2255011 |
| Mouse monoclonal anti-Tubulin (clone DM1A) | Sigma-Aldrich | Cat#T9026; RRID: AB_477593 |
| Rabbit polyclonal anti-BMAL1 | This paper | N/A |
| Bacterial and virus strains | ||
| pAAV-hSyn-DIO-hM3D(Gq)-mCherry | Krashes et al.1 | Addgene AAV5; 44361-AAV5 |
| AAV5-EF1a-DIO-hChR2(H134R)-EYFP | Hope Center Viral Vectors Core | N/A |
| Cowpox virus Brighton Red | BEI Resources | NR-88 |
| Zika-SMGC-1, GENBANK: KX266255 | Isolated from patient (Wang et al.2) | N/A |
| Staphylococcus aureus | ATCC | ATCC 29213 |
| Streptococcus pyogenes: M1 serotype strain: strain SF370; M1 GAS | ATCC 700294 | |
| Biological samples | ||
| Healthy adult BA9 brain tissue | University of Maryland Brain & Tissue Bank; http://medschool.umaryland.edu/btbank/ | Cat#UMB1455 |
| Human hippocampal brain blocks | New York Brain Bank | http://nybb.hs.columbia.edu/ |
| Patient-derived xenografts (PDX) | Children’s Oncology Group Cell Culture and Xenograft Repository |
http://cogcell.org/ |
| Chemicals, peptides, and recombinant proteins | ||
| MK-2206 AKT inhibitor | Selleck Chemicals | S1078; CAS: 1032350–13-2 |
| SB-505124 | Sigma-Aldrich | S4696; CAS: 694433–59-5 (free base) |
| Picrotoxin | Sigma-Aldrich | P1675; CAS: 12487–8 |
| Human TGF-β | R&D | 240-B; GenPept: P01137 |
| Activated S6K1 | Millipore | Cat#14–486 |
| GST-BMAL1 | Novus | Cat#H00000406P01 |
| Critical commercial assays | ||
| EasyTag EXPRESS 35S Protein Labeling Kit | PerkinElmer | NEG772014MC |
| CaspaseGlo 3/7 | Promega | G8090 |
| TruSeq ChIP Sample Prep Kit | Illumina | IP-202–1012 |
| Deposited data | ||
| Raw and analyzed data | This paper | GEO: GSE63473 |
| B-RAF RBD (apo) structure | This paper | PDB: 5J17 |
| Human reference genome NCBI build 37, GRCh37 | Genome Reference Consortium | http://www.ncbi.nlm.nih.gov/projects/gen ome/assembly/grc/h uman/ |
| Nanog STILT inference | This paper; Mendeley Data | http://dx.doi.org/10.17632/wx6s4mj7s8.2 |
| Affinity-based mass spectrometry performed with 57 genes | This paper; Mendeley Data | Table S8; http://dx.doi.org/10.17632/5hvpvspw82.1 |
| Experimental models: Cell lines | ||
| Hamster: CHO cells | ATCC | CRL-11268 |
| D. melanogaster: Cell line S2: S2-DRSC | Laboratory of Norbert Perrimon | FlyBase: FBtc0000181 |
| Human: Passage 40 H9 ES cells | MSKCC stem cell core facility | N/A |
| Human: HUES 8 hESC line (NIH approval number NIHhESC-09–0021) | HSCI iPS Core | hES Cell Line: HUES-8 |
| Experimental models: Organisms/strains | ||
| C. elegans: Strain BC4011: srl-1(s2500) II; dpy18(e364) III; unc-46(e177)rol-3(s1040) V. | Caenorhabditis Genetics Center | WB Strain: BC4011; WormBase: WBVar00241916 |
| D. melanogaster: RNAi of Sxl: y[1] sc[*] v[1]; P{TRiP.HMS00609}attP2 | Bloomington Drosophila Stock Center | BDSC:34393; FlyBase: FBtp0064874 |
| S. cerevisiae: Strain background: W303 | ATCC | ATTC: 208353 |
| Mouse: R6/2: B6CBA-Tg(HDexon1)62Gpb/3J | The Jackson Laboratory | JAX: 006494 |
| Mouse: OXTRfl/fl: B6.129(SJL)-Oxtrtm1.1Wsy/J | The Jackson Laboratory | RRID: IMSR_JAX:008471 |
| Zebrafish: Tg(Shha:GFP)t10: t10Tg | Neumann and NuessleinVolhard3 | ZFIN: ZDB-GENO060207–1 |
| Arabidopsis: 35S::PIF4-YFP, BZR1-CFP | Wang et al.4 | N/A |
| Arabidopsis: JYB1021.2: pS24(AT5G58010)::cS24:GFP(-G):NOS #1 | NASC | NASC ID: N70450 |
| Oligonucleotides | ||
| siRNA targeting sequence: PIP5K I alpha #1: ACACAGUACUCAGUUGAUA | This paper | N/A |
| Primers for XX, see Table SX | This paper | N/A |
| Primer: GFP/YFP/CFP Forward: GCACGACTTCTTCAAGTCCGCCATGCC | This paper | N/A |
| Morpholino: MO-pax2a GGTCTGCTTTGCAGTGAATATCCAT |
Gene Tools | ZFIN: ZDB-MRPHLNO-061106-5 |
| ACTB (hs01060665_g1) | Life Technologies | Cat#4331182 |
| RNA sequence: hnRNPA1_ligand: UAGGGACUUAGGGUUCUCUCUAGGGACUUAG GGUUCUCUCUAGGGA | This paper | N/A |
| Recombinant DNA | ||
| pLVX-Tight-Puro (TetOn) | Clonetech | Cat#632162 |
| Plasmid: GFP-Nito | This paper | N/A |
| cDNA GH111110 | Drosophila Genomics Resource Center | DGRC:5666; FlyBase:FBcl0130415 |
| AAV2/1-hsyn-GCaMP6-WPRE | Chen et al.5 | N/A |
| Mouse raptor: pLKO mouse shRNA 1 raptor | Thoreen et al.6 | Addgene Plasmid #21339 |
| Software and algorithms | ||
| ImageJ | Schneider et al.7 | https://imagej.nih.gov/ij/ |
| Bowtie2 | Langmead and Salzberg8 | http://bowtiebio.sourceforge.net/bowtie2/index.shtml |
| Samtools | Li et al.9 | http://samtools.sourceforge.net/ |
| Weighted Maximal Information Component Analysis v0.9 | Rau et al.10 | https://github.com/ChristophRau/wMICA |
| ICS algorithm | This paper; Mendeley Data | http://dx.doi.org/10.17632/5hvpvspw82.1 |
| Other | ||
| Sequence data, analyses, and resources related to the ultra-deep sequencing of the AML31 tumor, relapse, and matched normal | This paper | http://aml31.genome.wustl.edu |
| Resource website for the AML31 publication | This paper | https://github.com/chrisamiller/aml31SuppSite |
| Chemicals, peptides, and recombinant proteins | ||
| QD605 streptavidin conjugated quantum dot | Thermo Fisher Scientific | Cat#Q10101MP |
| Platinum black | Sigma-Aldrich | Cat#205915 |
| Sodium formate BioUltra, ≥99.0% (NT) | Sigma-Aldrich | Cat#71359 |
| Chloramphenicol | Sigma-Aldrich | Cat#C0378 |
| Carbon dioxide (13C, 99%) (<2% 18O) | Cambridge Isotope Laboratories | CLM-185–5 |
| Poly(vinylidene fluoride-co-hexafluoropropylene) | Sigma-Aldrich | 427179 |
| PTFE Hydrophilic Membrane Filters, 0.22 μm, 90 mm | Scientificfilters.com/TischScientific | SF13842 |
| Critical commercial assays | ||
| Folic Acid (FA) ELISA kit | Alpha Diagnostic International | Cat# 0365–0B9 |
| TMT10plex Isobaric Label Reagent Set | Thermo Fisher | A37725 |
| Surface Plasmon Resonance CM5 kit | GE Healthcare | Cat#29104988 |
| NanoBRET Target Engagement K-5 kit | Promega | Cat#N2500 |
| Deposited data | ||
| B-RAF RBD (apo) structure | This paper | PDB: 5J17 |
| Structure of compound 5 | This paper; Cambridge Crystallographic Data Center | CCDC: 2016466 |
| Code for constraints-based modeling and analysis of autotrophic E. coli | This paper | https://gitlab.com/elad.noor/sloppy/tree/master/rubisco |
| Software and algorithms | ||
| Gaussian09 | Frish et al.1 | https://gaussian.com |
| Python version 2.7 | Python Software Foundation | https://www.python.org |
| ChemDraw Professional 18.0 | PerkinElmer | https://www.perkinelmer.com/category/chemdraw |
| Weighted Maximal Information Component Analysis v0.9 | Rau et al.2 | https://github.com/ChristophRau/wMICA |
| Other | ||
| DASGIP MX4/4 Gas Mixing Module for 4 Vessels with a Mass Flow Controller | Eppendorf | Cat#76DGMX44 |
| Agilent 1200 series HPLC | Agilent Technologies | https://www.agilent.com/en/products/liquid-chromatography |
| PHI Quantera II XPS | ULVAC-PHI, Inc. | https://www.ulvacphi.com/en/products/xps/phi-quantera-ii/ |
EXPERIMENTAL MODEL AND SUBJECT DETAILS
Animal Studies:
Procedures involving animals were performed in accordance with National Institutes of Health guidelines and approved by the Institutional Animal Care and Use Committee at Duke University School of Medicine. Adult male mice (12–24 weeks of age) were used for all studies and were group housed in a light (12h light/12h dark) and temperature (22°C) controlled room with ad libitum access to standard rodent chow (Lab Diets PicoLab rodent diet 5053) and water throughout studies unless noted otherwise. C57BL/6NJ mice (JAX Labs; Stock No. 000664) were used for experiments comparing heart and skeletal muscle mitochondria and red and white gastrocnemius mitochondria, acute exercise, and the fasting time course. Studies involving transaortic constriction (TAC) were conducted by the Duke Cardiovascular Research Center (CVRC). Methods and results of the TAC studies were described previously39 and the corresponding proteomics data set (https://doi.org/10.1161/CIRCRESAHA.120.31729) was re-analyzed to integrate with new proteomics data generated by current the study. MCK-Pgc1a transgenic mice and non-transgenic controls were bred in-house from the original strain generated and generously gifted by Dr. Bruce Spiegelman 55 and sacrificed after 2 h food removal. Mice with a skeletal muscle and heart-specific deletion of CrAT (CrATM−/−) and control animals (CrATfl/fl) were utilized for heart perfusion studies 35. After 1–2 h food removal starting at 0700 (except where noted) mice were anesthetized with Nembutal (intraperitoneal injection; 100 mg/kgBW) prior to organ and tissue extraction.
METHOD DETAILS
Chemicals and Reagents:
All chemicals were purchased from Sigma-Aldrich unless otherwise stated. Tetramethylrhodamine methyl ester (TMRM) and Amplex Ultra Red were purchased from Thermo Fisher Scientific. Potassium pyruvate was purchased from Combi-Blocks. Creatine kinase was purchased from Roche. Alamethicin was purchased from Enzo Life Sciences. [U-13C]-Glucose was purchased from Cambridge Isotope Laboratories. [U-13C]-Pyruvate for mitochondrial tracer experiments was purchased from Cambridge Isotope Laboratories. Protease (Complete mini EDTA-free) and phosphatase (PhosStop) inhibitor tablets (Roche). 11-plex Tandem Mass Tags (TMT) were purchased from Thermo Fisher Scientific. Sequencing Grade Modified. Sequencing grade trypsin was purchased from Promega. Lysyl Endopeptidase (LysC) was purchased from Wako Chemicals. tC18 SEP-PAK SPE columns were purchased from Waters.
Acute Exercise:
Adult male mice (C57BL/6NJ mice bred in house) 21–24 weeks of age underwent 3 days of treadmill habituation for 15 min each day (Columbus Instruments; Exer3/6). On the day of the study, mice were randomized to non-exercised or exercised groups as well as 5’ and 60’ post-exercise groups. The treadmill protocol for the exercise group was conducted at a fixed 10 degree grade as follows: 20 min at 6m/min, 20 min at 10m/min, 20 min at 12m/min, 20 min at 14m/min, 20 min at 16m/min. Running was encouraged using gentle prodding with a laboratory test tube brush as needed. At 100 min, the treadmill was stopped and mice in the 5’ post-exercise group were injected with 100 mg/kgBW Nembutal and tissue collection commenced 4 min post-injection when areflexic. The heart was removed first followed by skeletal muscles (gastrocnemius and quadriceps). Tissues were quickly rinsed in ice-cold PBS, blotted, freeze clamped using liquid nitrogen chilled clamps and stored at −80C until pulverization and metabolite extraction. The procedure was repeated on mice who underwent 60’ of recovery post-exercise.
Mitochondrial isolation:
Food was removed 1–2 h before experiments and mice were anesthetized with Nembutal. When areflexic, heart and skeletal muscles (one quadricep and two gastrocnemius/plantaris) were extracted and immediately placed in ice-cold PBS containing 10 mM EDTA (pH 7.4). Mitochondria were isolated by differential centrifugation 56 with modifications using the following buffers: Buffer A – 100 mM KCl, 50 mM MOPS, 1 mM EGTA, 5 mM MgSO4 (pH 7.1); and Buffer B – Buffer A supplemented with 2 g/L fatty-acid free bovine serum albumin (BSA). Tissues were minced with surgical scissors in 1.5 ml Buffer A, transferred to a 50 ml conical tube, and incubated in 10 ml of Buffer A containing 0.05% trypsin on ice for 5min with gentle vortexing every 30sec. The minced suspensions were centrifuged at 200g for 5 min at 4°C and the supernatant discarded. The tissue pellet was suspended in 10 ml Buffer B and homogenized with an ice-chilled Teflon pestle and borosilicate glass vessel and centrifuged at 800g for 5 min at 4°C. The supernatant containing mitochondria was filtered through two layers of surgical gauze and centrifuged at 9,000g for 10 min at 4°C. Mitochondrial pellets were suspended in 1.2 ml Buffer B using an ice-chilled Teflon pestle and centrifuged at 9,000g for 3 min at 4°C. Pellets were suspended in 1 ml Buffer A to remove BSA and centrifuged at 9,000g for 3 min at 4°C. Buffer A was aspirated from each tube and mitochondrial pellets were suspended in 50–100 μl Buffer A. Protein concentration was determined using the Pierce BCA protein assay and mitochondria were resuspended at a final concentration of 10 mg/ml for all functional assays. For proteomic and Western blot studies, 0.1 mg aliquots of mitochondria were added to tubes containing Buffer A and protease and phosphatase inhibitors and mixed gently. Samples were pelleted at 9,000g for 3 min at 4°C, the supernatant was aspirated, and mitochondrial pellets were stored at −80°C before lysis.
Mitochondrial Respiratory Control:
High-resolution respirometry was performed using OROBOROS Oxygraph-2k (O2K) using a modified version of the creatine kinase (CK) bioenergetic clamp 57,58 in the presence of excess CK, and known amounts of ATP, creatine (Cr), and phosphocreatine (PCr) as described previously 1. The CK reaction couples the interconversion of ATP and ADP to PCr and Cr. Assays were conducted in 2 ml of Buffer Z (105 mM MES potassium salt, 30 mM KCl, 10 mM KH2PO4, 5 mM MgCl2) supplemented with Cr (5 mM), PCr (1.5 mM), CK (20 U/mL), fatty acid free BSA (2.5 mg/mL), and EGTA (1 mM; pH 7.2) at 37⁰C with constant stirring. At the start of each assay, isolated mitochondria (0.025 mg/mL) were added to the assay buffer followed by the addition of respiratory substrates, then ATP (5 mM). Sequential additions of PCr were added to achieve the final concentrations of 3, 6, 9, 12, and 15 mM allowing tuning of the free energy of ATP hydrolysis (ΔGATP) 1. Substrates and combinations are as described in figure legends for a given study. All substrates were provided at concentrations that are saturating for JO2.
Mitochondrial membrane potential and NAD(P)H:
Simultaneous fluorescent assessment of the mitochondrial membrane potential () and NAD(P)H/NAD(P)+ were performed in parallel with JO2 assays using a PTI QuantaMaster Spectrofluorometer with Quantum Northwest 4-cuvette revolving turret and magnetic stir bars to permit continuous and uniform mixing. Cuvettes were Spectrosil Quartz with a 5mm path length and kept open to air during the assays (Starna; 23–5.45-Q-5). The was determined using the ratiometric fluorescent dye tetramethylrhodamine methyl ester (TMRM) with excitation/emission (λex/λem) parameters of 572/590nm and 551/590nm, respectively. The λex/λem parameters for NAD(P)H were 340/450nm. All assays were conducted in 0.2 mL Buffer Z supplemented with Cr (5mM), PCr (1.5 mM), CK (20 U/mL), and TMRM (0.2 μM) at 37°C with constant stirring. Mitochondria (0.1 mg/mL) were added to the assay buffer followed by addition of respiratory substrates, ATP (5 mM), and then sequential additions of PCr to achieve the final concentrations of 3, 6, 9, 12, and 15 mM. After the last PCr addition, potassium cyanide (4 mM) was added to induce a 100% reduction of the NAD(P)H/NAD(P)+ redox couple followed by alamethicin (12.5 μg/mL) to permeabilize the mitochondria and create a state of 0% reduction. NAD(P)H/NAD(P)+ throughout the experiment was expressed as percent reduction based on the following equation: (F1-F0%)/(F100%-F0%). Mitochondrial was estimated by converting the fluorescent 572/551 nm ratio to millivolts using a KCl standard curve generated in the presence of valinomycin 59. Assuming a matrix potassium concentration of 120 mM, can be reasonably estimated by applying the Nernst equation and buffer ion concentrations resulting from sequential additions of KCl 60.
Mitochondrial H2O2 emission:
Rates of mitochondrial H2O2 emissions (JH2O2) were determined fluorometrically (QuantaMaster Spectrofluorometer; λex/λem 565/600 nm) using Amplex Ultra Red (AUR)/horseradish peroxidase (HRP) detection system. All assays were conducted in 0.2 mL Buffer Z supplemented with Cr (5 mM), PCr (1.5 mM), CK (20 U/mL), AUR (10 μM), HRP (1 U/mL) and superoxide dismutase (20 U/mL) at 37°C with constant stirring. At the start of each assay, isolated mitochondria (0.1 mg/mL) were added to the assay buffer, followed by respiratory substrates, ATP (5 mM), auranofin (AF; 0.1 μM), and then sequential additions of PCr to achieve the final concentrations of 6 and 15 mM. After the final PCr addition, 1-chloro-2,4-dinitrobenzene (CDNB) was added to approximate maximal JH2O2.
HSkMC Cell Culture:
Muscle satellite cells were derived from biopsies of the vastus lateralis under Duke University IRB approved protocol Pro00007394, using previously described methods 61. Cryopreserved primary human myoblasts from five healthy female donors (mean age=25y, BMI=23.5kg/m2) were used to create pooled lots of human myoblasts. Cells were amplified in growth media (DMEM low glucose medium supplemented with 10% FBS, 0.05% BSA, 0.5mg/mL fetuin, 0.01μg/ml human epidermal growth factor, 0.04μg/ml dexamethasone, 0.025mg/mL gentamicin, 0.125μg/ml amphotericin B) in a 5% CO2 and 37°C humidified atmosphere. Upon reaching confluency, myoblasts were switched to differentiation media for 7–8 days (customized low amino acid MEM supplemented with 2% horse serum, 0.05% BSA, 0.5mg/mL fetuin, 100μM L-carnitine, 0.5mM L-glutamine, 5mM galactose, 0.2mM sodium pyruvate, 0.2mM 1:1 oleate: palmitate, 0.025mg/mL gentamicin, 0.125μg/ml amphotericin B) to induce the formation of multinucleated, mature myotubes.
HSkMC siRNA Transfection:
On differentiation day 4, myotubes were transfected with 0.3 nM of siRNA oligonucleotide sequences targeting human ACAA2/MKT (Integrated DNA Technologies, hs.Ri.ACAA2 – 13.1, 13.2, 13.3 (0.1nM each)) or a non-targeting control sequence (IDT, 51–01-14–03) using Lipofectamine RNAiMAX (ThermoFisher Scientific, 13778) in reduced serum Opti-MEM (ThermoFisher, 31985062). Myotubes were switched back to differentiation media on day 5 (24 h after transfection) and assays were performed on day 8.
Permeabilized HSkMC Respiration:
Primary human myoblasts seeded at a density of 10K cells per well were fully differentiated to myotubes. Respiration rates were measured in permeabilized HSkMC using the Seahorse Flux Analyzer (Seahorse Bioscience) and a modified version of the CK bioenergetic clamp. Immediately prior to the assay, differentiation media was replaced with Buffer Z supplemented with Cr (5 mM), PCr (1.5 mM), CK (20U/mL) and respiratory substrates. At the start of the assay, HSkMCs were permeabilized with 1.0 nM XF Plasma Membrane Permeabilizer (PMP; Agilent Technologies, 102504–100), followed by port addition of 5 mM ATP. Sequential port additions of PCr were added to achieve final concentrations of 3 and 6 mM. The mix, wait, measure” phase of each measurement cycle consisted of a 30s mixing step, 30s wait step, and 2 min measurement step. The average oxygen consumption across three measurement periods was used to derive JO2 values for the ATP and PCr addition stages. Following the assay, cells were lysed in 0.5% SDS with protease inhibitors. JO2 measures were normalized to total protein content determined after permeabilization via the BCA assay and used to calculate respiratory conductance. Permeabilized HSkMC pooled from 15–30 wells per condition, per experiment, were used for western blot analysis to confirm MKT KD in each of 7 independent experiments.
Western Blotting:
Mitochondrial pellets were lysed with 2x sample buffer (20 mM Tris-HCl, 4% SDS, 10 mM NaF, 1 mM EDTA, 20% glycerol, pH 6.8) with protease and phosphatase inhibitors. Following lysis, 20–40 μg of mitochondrial protein or 6–15 μg HSkMC protein was loaded onto a 4–15% gradient gel (BioRad) and subjected to electrophoresis to separate proteins based on size. Proteins were transferred from the gradient gel to a nitrocellulose membrane using a Trans TurboBlot system (BioRad). Following transfer, membranes were stained with reversible total protein stain (Thermofisher Scientific, 1858784). Membranes were then imaged to assess total protein levels. Total protein stain was removed with stain eraser (Thermofisher Scientific, 1858786), and membranes were then blocked in a fish gelatin blocking solution for 1 h at room temperature. After blocking, membranes were incubated in primary antibodies overnight (MKT, Sigma WH0010449M1; CrAT, ABCAM ab153750, SCHAD ProteinTech, 19828–1AP; ATP5A, ABCAM ab14748; SDHA, ABCAM, ab14715; Total OxPHOS ABCAM, ab110413; NDUFA9, ABCAM, ab14713; T-AMPK, CST, 2532; P-AMPK, CST, 2531; HSP60, CST, 12165). Following primary incubation, membranes were washed 5 X 5 minutes in TBST and then incubated with appropriate secondary antibodies. Membranes were then washed 5 X 5 min in TBST, and 1 X 5 min in TBS. Protein bands on the membranes were then imaged using an Odyessy Clx system (Licor). Band densitometry was quantified using Image Studio (V5.2; Li-Cor), and then normalized to the quantification of total protein stain within a region encompassing the MW of the protein of interest.
Acute exercise with variable recovery:
Acute exercise test was conducted as previously described 62. After 3 days of habituation to the open-air Exer-3/6 treadmill equipped with a shock grid (Columbus Instruments) the exercise bout started at 5 m/min for 10 min to warm up, speed was increased to 10 m/min for 20 min, and then by 1 m/min every 2 minutes to a final rate of 15 m/min, which was maintained for 1 h. Control mice were habituated but not exercised. Tibialis anterior muscles were collected immediately post (5min) exercise or after 3 or 24 h of recovery and immediately flash frozen.
Gene expression:
Total RNA was extracted from frozen tibialis anterior powdered under liquid nitrogen using Trizol/chloroform extraction method followed by cleanup using RNeasy Mini spin columns (Qiagen) and on column DNase treatment. RNA concentrations were determined using a NanoDrop (ThermoFisher Scientific). First-strand cDNA was synthesized using the iScript cDNA synthesis kit (BioRad). Real-time quantitative PCR for Acaa2 was performed using PrimeTime Gene Expression Master Mix (IDT) and TaqMan#Mm_006224282) on a QuantStudio 6 real-time PCR system (Applied Biosystems). Threshold CT values were normalized to 18S rRNA (18S assay Applied Biosystems #4319413E) to calculate fold change relative to Non-Exercised Controls.
Heart perfusion:
Mice were anesthetized with 5% isoflurane, and the isolated hearts were perfused in the Langendorff mode at 37°C with nonrecirculating perfusate of Krebs-Ringer bicarbonate buffer containing 119 mM NaCl, 4.8 mM KCl, 2.6 mM CaCl2, 1.2 mM KH2PO4, 1.2 mM MgSO4, 25 mM NaHCO3, 11 mM glucose, 0.4 mM palmitate, 3% BSA, 100 μU/ml insulin, and 0.05 mM l-carnitine at a constant flow rate of 1.2 ml/min 63. The hearts were allowed to beat spontaneously throughout the perfusion and were maintained at 37 °C with a water jacketed chamber. The hearts were perfused with Krebs-Ringer bicarbonate buffer for 15 min and then switched to perfusate containing the labeled substrates for 25 min. At the end of each perfusion, hearts were freeze-clamped in liquid nitrogen and stored at −80°C until analysis. The effluent of perfusate sample (~1 ml) was collected during the final minute of perfusion and quickly frozen in liquid nitrogen. In [U-13C]glucose perfusion group, 11 mM glucose in perfusate was replaced by 11 mM [U-13C]glucose. For hearts perfused with [U-13C]acetate, [U-13C]acetate was added into perfusate to reach final concentration at 1 mM.
Mitochondrial and tissue acyl-CoA analysis by LC-MS/MS:
Acyl-CoA analysis was modified from our previous reports 64,65. Briefly, frozen mitochondrial pellets (0.25mg) were sonicated in 1.2 ml methanol for 1 min. After homogenization, 100 μl homogenate spiked with 20 μl 0.01 mM D9 carnitine as internal standard was used for acylcarnitine profile and 1 ml homogenate was spiked with 20 μl 0.01 mM D9 pentanoyl-CoA as internal standard and mixed with 1 ml LC-grade water with 5% acetic acid for acyl-CoA analysis. For tissue samples, ~50 mg tissue sample were homogenized in 1 ml extraction buffer (1:1 methanol/water containing 5% acetic acid) followed by 15-minute centrifugation at 18,000g. For solid phase extraction, the cartridge (1-ml ion exchange cartridge packed with 100 mg of 2–2(pyridyl)ethyl silica gel (Sigma)) was pre-activated with 1 ml of methanol and then with 1 ml of extraction buffer (1:1 methanol/water containing 5% acetic acid). The acyl-CoAs trapped on the silica gel cartridge were released with (i) 1 ml of a 1:1 mixture of 50 mm ammonium formate, pH 6.3, and methanol and then (ii) 1 ml of a 1:3 mixture of 50 mm ammonium formate, pH 6.3 and (iii) 1 ml of methanol. The combined effluent was dried with nitrogen gas and stored at −80 °C until LC-MS analysis. The detailed LC-MS/MS conditions can refer to our previous reports 64,65.
Mitochondrial and tissue acylcarnitine profile by LC-MS/MS:
Acylcarnitines in the mitochondrial pellets or tissues were methylated and profiled by the modified liquid chromatography with tandem mass spectrometry (LC-MS/MS) method. Briefly, for heart or skeletal muscle tissues, ~20 mg frozen and pulverized tissue was spiked with 20 μl 0.01 mM D9 carnitine as internal standard and homogenized with a Folch-extraction approach 63. A 300 μl tissue extract or 100 μl mitochondrial homogenate from the above mitochondrial sample preparation (see Mitochondrial acyl-CoA analysis by LC-MS/MS) was completely dried by nitrogen gas. The dried residue was derivatized with 100 μl 3 M HCl methanol solution at 50 °C for 25 minutes. The derivatized sample was dried again by nitrogen gas and the dried sample was re-dissolved into 20 μl methanol followed by 60 μl LC water. The derivatized samples were analyzed by LC-QTRAP 6500+-MS/MS (Sciex, Concord, Ontario).
A 5 μl sample was injected on a Pursuit XRs 5 C18 column (150 × 2.0 mm, 5μm), protected by a guard column (Pursuit XRs 5 C18 5 μm, 10 × 2.0 mm, 5μm) with temperature controlled at 25 °C, in an ExionLC AD liquid chromatograph system. The chromatogram was developed at 0.4 ml/min (i) for 2 min with 100% buffer A (98% H2O and 2% acetonitrile with 0.1% formic acid), (ii) from 2 to 13 min with a gradient from 0 to 80% buffer B (98% acetonitrile and 2% H2O with 0.1% formic acid), (iii) from 14 to 18.5 min with 10% buffer A and 90% buffer B, (iv) from 18.5 to 19 min with a gradient from 90% to 0 buffer B, and (v) for 10 min of stabilization with 100% buffer A before the next injection. The liquid chromatograph was coupled to a 6500+-MS/MS operated under positive ionization mode with the following source settings: turbo-ion-spray source at 650 °C under N2 nebulization at 65 PSI, N2 heater gas at 65 PSI, curtain gas at 35 PSI, collision-activated dissociation gas pressure held at high, turbo ion-spray voltage at 5,500 V, declustering potential at 33 V, entrance potential at 10 V, collision energy ranged from 24 to 46 V optimized for each acylcarnitine and collision cell exit potential at 14 V. The Analyst software (version 1.6) was used for data record and processing.
Stable isotope tracing data correction
M+0, M+1, …, M+n refer to the isotopologues containing n heavy atoms in a molecule. The isotopologue enrichment or labeling in this work referred to the corrected isotope distribution. Briefly, the labeling of all measured metabolites in samples from 13C treated samples was corrected based on the natural isotope distribution of each metabolite. The natural isotope distribution of each metabolite was to build a matrix for correcting isotope labeling of metabolite in the tracer-treated samples. Details of these methods have been published previously 66,67.
Isolated Skeletal Muscle Incubations:
All skeletal muscle incubations used a base buffer of oxygenated (95% O2, 5% CO2), low calcium Krebs-Henseleit Buffer (KHB) containing 118 mM NaCl, 4.73 mM KCl, 1.18 mM MgSO4·7H2O, 0.47 mM CaCl2·2H2O, 1.17 mM KH2PO4, 25 mM NaHCO3, 12.5 mM HEPES, and 0.5 mM L-carnitine; pH = 7.4 68. Mouse soleus (SOL) and extensor digitorum longus (EDL) were excised from 5h-fasted, 14 week old, male C57BL/6NJ mice and each muscle was individually placed in one well of a 24-well plate. Each well contained 1 mL of KHB supplemented with 5 mM glucose and 0.5% BSA (~22°C). After all tissues were excised, SOL and EDL were transferred to a new plate containing 1mL of KHB + 5 mM glucose + 0.5% BSA, placed into a shaking water bath at 120rpm, and incubated for 15 min at 29°C before increasing the temperature to 37°C for another 15 min. Subsequently, SOL and EDL were transferred to a plate containing KHB + 10 mM [U-13C]glucose + 0.2 mM palmitate + 0.5% BSA. Each contralateral muscle pair interrogated the impact of insulin treatment (0 nM for basal and 100 nM for insulin stimulus). After the 90 min tracer incubation, SOL and EDL were rinsed in ice-cold KHB + 0.5% BSA, trimmed, dabbed dry, and flash frozen in liquid nitrogen. Samples were stored at −80°C until processed.
Tissue Homogenization and Metabolite Extraction:
Frozen SOL and EDL tissues were massed in dry ice chilled 2mL Eppendorf tubes. Tissues were then minced for 1 minute in 400μL of methanol (−20°C) with surgical scissors and homogenized with hand-held rotary device (VWR, VDI12) for 1 minute with a setting of “5.5”. Afterwards, 400 μL of 5 μM norvaline in water (4°C) was added, and the sample was homogenized for another minute. Lastly, 400 μL of chloroform (4°C) was added, followed by a minute of homogenization. Tissue extracts were then centrifuged at 4°C and 14,400g for 20 min and 300 μL of clarified aqueous phase was dried under nitrogen at 37°C via a forced-flow evaporator (Organomation). Samples were stored at −80°C until preparation for acyl-carnitine assay.
Isotopomer Terminology:
A mass isotopomer is the collection of isotope isomers that have the same molecular mass, i.e. same number of 13C-atoms, however, they can be positionally different. The M+0 isotopomer is the simplest as has ‘zero’ 13C-atom incorporation above natural abundance. The isotopomer is a collection of all labeling patterns that have a single 13C-atom incorporated. The mass isotopomer distribution (MID) provides information regarding the distribution of 13C-label in a compound. A metabolite with ‘n’ carbons will have an MID that encompasses isotopomers from up to. The sum of all mass isotopomers equals 1. The MID data can be used to compute an ‘average’ representation of 13C-enrichment in each metabolite. We will refer to this as average 13C-labeling (APE) and is calculated as:
where the number of carbons, and represents the ith mass isotopomer.
Proteomic Sample Preparation:
Two proteomics experiments used similar methods, with technical differences noted. Experiment 1 compared mitochondria from mouse heart (n=5) and SKM (n=5) tissue, while Experiment 2 compared SKM mitochondria from PGC1a transgenic (n=6) and control (n=5) mice. For both experiments, pelleted mitochondria were resuspended in ice-cold 8M Urea Lysis Buffer (8 M urea in 50 mM Tris, pH 8.0, 40 mM NaCl, 2 mM MgCl2, 1x Roche cOmplete ULTRA EDTA-free protease inhibitor cocktail tablet). Samples were disrupted by a combination of vortexing, freeze/thaw (3 cycles switching between dry ice and 32°C), and sonication with a probe sonicator (three 5s bursts, power setting of 3). Samples were clarified by centrifugation at 10,000g for 10 min at 4°C. Protein content was determined via the BCA assay and 50 μg of protein was normalized with Urea Lysis Buffer to an equal concentration. Samples were reduced with 5 mM DTT at 32°C for 30 min, cooled to room temperature (RT), alkylated with 15 mM iodoacetamide for 30 min in the dark, and unreacted iodoacetamide was quenched by the addition of DTT to a final concentration of 15 mM. Samples were digested with Endoproteinase LysC (Wako Chemicals; 1:100 w:w; 0.5 μg enzyme per 50 μg protein) for 4 h at 32°C. Urea was diluted to 1.5 M (using a solution of 50 mM Tris pH 8.0, 5 mM CaCl2) and digested with trypsin (Promega Seq Grade Modified Trypsin, 1:50 w:w; 1 μg enzyme per 50 μg protein) overnight at 32°C. Samples were acidified to 0.5% v/v trifluoroacetic acid (TFA) and centrifuged at 10,000g for 10 min at RT to pellet any undigested material. The supernatant containing soluble peptides was desalted on a 50 mg tC18 SEP-PAK Solid Phase Extraction (SPE) column (Waters) and peptides were eluted once with 500 μL of 25% acetonitrile (ACN)/0.1% TFA and twice with 500 μL 50% ACN/0.1% TFA. The eluate was frozen on dry ice and dried overnight in a speed vac.
Quantitative LC/MS/MS Analysis:
Label-free quantitative nanoLC-MS/MS was performed on 1 μg of digested peptide from all experimental samples and QC pools (made from combining samples). Both experiments used data-dependent acquisition (DDA) on an Orbitrap mass spectrometer (ThermoFisher Scientific) coupled to a UHPLC system via a nanoelectrospray ionization source. Experiment 1 utilized an Orbitrap Fusion Lumos and a nanoAcquity UPLC (Waters Corp). Samples were injected and trapped at 5 μL/min on a Symmetry C18 trapping column (20 mm × 180 μm, Waters) and separated on an Acquity HSS T3 C18 analytical column (1.8 μm 75 μm × 250 mm, Waters) over a 90-min linear gradient (5 to 30% acetonitrile with 0.1% formic acid (FA)) at a flow rate of 400 nL/min and a column temperature of 55°C. MS1 spectra (precursor ions) were collected at a resolution (r) of 120,000, a target AGC value of 2e5 ions, and a maximum injection time (IT) of 50 ms. Using a DDA cycle time of 2 s, precursor ions were selected using a 1.2 m/z isolation window and fragmented using stepped collision energy, with dynamic exclusion (DE) enabled for 20 s. MS2 spectra (product ions) were collected using a Rapid ion trap scan (AGC=5e3, max IT=100 ms). Experiment 2 utilized a Q Exactive Plus Orbitrap and an EASY-nLC UPLC system (Thermo) with Acclaim PepMap (Thermo) trapping (3 um, 75 μm × 20 mm) and analytical (2 um 100 C18, 75 μm × 500 mm column) columns, separating peptides over a 105 min gradient (2 to 40% solvent B (90% ACN/0.1% FA)) at 300 nL/min. MS1 (r=70,000, AGC=3×106, max IT=100 ms) and MS2 (r=17,500 AGC=1×105, max IT=100 ms) spectra were collected by DDA using a loop count of 20 (isolation window=1.2 m/z, DE=30 s, normalized collision energy=27). Newly generated proteomics raw data are available through ProteomeXchange (PXD040326) and jPOST (JPST002051) using the indicated accession numbers.
Proteomic Data Analysis:
Raw data for all experimental samples and QC pools were processed in Proteome Discoverer 2.2 (ThermoFisher). The Minora Feature Detector node was used to align precursor ion features across all runs based on mass and retention time. Relative peptide abundance was calculated using precursor intensity, considering MS1 extracted ion chromatograms (XIC) of the aligned features. Data were searched against a mouse complete proteome database downloaded from UniProt and corresponding “decoy” reversed protein sequences. Considered modifications included oxidation (15.995 Da on M) as variable and carbamidomethyl (57.021 Da on C) as fixed, with up to 2 missed cleavages (full trypsin specificity). FDR for peptide spectral matches (PSMs) was estimated using Percolator 69, after which Peptide Validator was used to collapse PSMs to unique peptides at 1% FDR. Quantitation was normalized for total peptide signal within each sample, providing Normalized Abundance values to adjust for subtle differences in sample loading and LC-MS performance. Peptides were grouped to proteins (strict parsimony), which were filtered to 1% FDR with Protein FDR Validator. “Master Proteins”, or representatives of a group of proteins containing the same peptides, were used for quantitative comparison. Experiment 1 controlled for differences in mitochondrial purity between tissues by the following approach: After searching heart and SKM data separately, quantitative values for MitoCarta 70 proteins confidently identified (1% FDR) in both tissues were summed. Quantitative values for each MitoCarta protein identified in both tissues was divided by the MitoCarta protein sum for a given sample. These “Mitochondrial-normalized Abundance” values represent the fraction of all mitochondrial proteins for which each protein accounts. As Experiment 2 only evaluated one tissue, providing more uniform mitochondrial enrichments, all raw files were searched together and Normalized Abundance values were used directly. Values were converted to log2 space and fold changes (Log2 FC) were calculated across each comparison (Heart/SKM or PGC1a/Control). For each comparison, p-values were calculated using two-tailed Student’s t-test (assuming equal variance) and adjusted p-value was calculated using the Benjamini-Hochburg FDR correction (Padjusted).
QUANTIFICATION AND STATISTICAL ANALYSIS
Statistical analysis:
All data are presented as mean±SEM. Statistical analysis was performed in GraphPad Prism 9.4 (GraphPad Software, San Diego, CA) using 2-tailed unpaired t-tests with Welch’s correction and Holm-Šídák multiple testing correction and one-way ANOVA with Dunnet’s post-hoc testing. Specific tests are indicated in figure legends. Figures were generated using GraphPad Prism 9.4. The level of significance was set at p ≤ 0.05 for all experiments. Unless otherwise noted in figure legends, ns = P>0.05, *=P<0.05, **=P<0.01, ***=P<0.001, ****=P<0.0001.
Supplementary Material
Data S3. Uncropped images of all western blots, related to Figures 2 and 7, and Supplemental Figures S4 and S5.
Data S4. Data used to generate all figures.
Data S1. Mitochondrial Proteomics, Heart versus SkM, related to Figure 5.
Data S2. Mitochondrial Proteomics, PGC1a versus NT SkM, related to Figure 5.
S. Table 2. Mitochondrial Proteomics, Heart versus SkM, related to Figure 5.
S. Table 3. Mitochondrial Proteomics, PGC1a versus NT SkM, related to Figure 5.
HIGHLIGHTS.
The pathway of long chain fatty acid oxidation is prone to metabolic bottlenecks
Bottlenecks in fat oxidation can lead to CoA trapping and respiratory inefficiency
Reverse flux of pyruvate-derived acetyl CoA through MKT regenerates free CoA
High levels of MKT in heart and red muscles promote mitochondrial lipid tolerance
ACKNOWLEDGEMENTS
We thank the DMPI Metabolomics and Biomarker Core Laboratory for assistance with tissue acylcarnitine analyses; Duke Proteomics and Metabolomics Shared Resource for assistance with the label-free proteomics analysis; Jonathan Kasper for his contributions to Figure 7 Supplemental data; Rebecca Wilson for assistance with the fasting time course experiment; and Ashley Williams for assistance with muscle incubation experiments. We recognize Jeffrey Messer, Brian Glancy and Wayne Willis for developing and applying the first prototype of the CK clamp technique, as well as the bioenergetics concepts eloquently conveyed in (Willis et al., 2016) 28. The graphical abstract was created with BioRender.com. This work was supported by National Institutes of Health grants 3R01DK089312 (DMM) and 2R01HL128349 (DMM). The DMPI Metabolomics Core Laboratory is supported by Diabetes and Endocrine Research Center grant P30 DK124723.
Footnotes
Declaration of Interests. None
INCLUSION AND DIVERSITY
We support inclusive, diverse and equitable conduct of research.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
REFERENCES
- 1.Fisher-Wellman KH, Davidson MT, Narowski TM, Lin CT, Koves TR, and Muoio DM (2018). Mitochondrial Diagnostics: A Multiplexed Assay Platform for Comprehensive Assessment of Mitochondrial Energy Fluxes. Cell Rep 24, 3593–3606 e3510. 10.1016/j.celrep.2018.08.091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Willis WT, Jackman MR, Messer JI, Kuzmiak-Glancy S, and Glancy B. (2016). A Simple Hydraulic Analog Model of Oxidative Phosphorylation. Med Sci Sports Exerc 48, 990–1000. 10.1249/MSS.0000000000000884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Muoio DM (2014). Metabolic inflexibility: when mitochondrial indecision leads to metabolic gridlock. Cell 159, 1253–1262. 10.1016/j.cell.2014.11.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Zhou B, and Tian R. (2018). Mitochondrial dysfunction in pathophysiology of heart failure. J Clin Invest 128, 3716–3726. 10.1172/JCI120849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kelley DE, Goodpaster B, Wing RR, and Simoneau JA (1999). Skeletal muscle fatty acid metabolism in association with insulin resistance, obesity, and weight loss. Am J Physiol 277, E1130–1141. 10.1152/ajpendo.1999.277.6.E1130. [DOI] [PubMed] [Google Scholar]
- 6.Muoio DM., Noland RC., Kovalik J-P., Seile SE., Davie MN., DeBalsi KL., Ilkayeva OR., Stevens RD., Kheterpal I., and Zhang J. (2012). Muscle-specific deletion of carnitine acetyltransferase compromises glucose tolerance and metabolic flexibility. Cell metabolism 15, 764–777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Huffman KM, Koves TR, Hubal MJ, Abouassi H, Beri N, Bateman LA, Stevens RD, Ilkayeva OR, Hoffman EP, Muoio DM, and Kraus WE (2014). Metabolite signatures of exercise training in human skeletal muscle relate to mitochondrial remodelling and cardiometabolic fitness. Diabetologia 57, 2282–2295. 10.1007/s00125-014-3343-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Holloszy JO (1967). Biochemical adaptations in muscle. Effects of exercise on mitochondrial oxygen uptake and respiratory enzyme activity in skeletal muscle. J Biol Chem 242, 2278–2282. [PubMed] [Google Scholar]
- 9.Neufer PD, Bamman MM, Muoio DM, Bouchard C, Cooper DM, Goodpaster BH, Booth FW, Kohrt WM, Gerszten RE, Mattson MP, et al. (2015). Understanding the Cellular and Molecular Mechanisms of Physical Activity-Induced Health Benefits. Cell Metab 22, 4–11. 10.1016/j.cmet.2015.05.011. [DOI] [PubMed] [Google Scholar]
- 10.Noland RC, Koves TR, Seiler SE, Lum H, Lust RM, Ilkayeva O, Stevens RD, Hegardt FG, and Muoio DM (2009). Carnitine insufficiency caused by aging and overnutrition compromises mitochondrial performance and metabolic control. J Biol Chem 284, 22840–22852. 10.1074/jbc.M109.032888. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Koves TR, Ussher JR, Noland RC, Slentz D, Mosedale M, Ilkayeva O, Bain J, Stevens R, Dyck JR, Newgard CB, et al. (2008). Mitochondrial overload and incomplete fatty acid oxidation contribute to skeletal muscle insulin resistance. Cell Metab 7, 45–56. 10.1016/j.cmet.2007.10.013. [DOI] [PubMed] [Google Scholar]
- 12.Koves TR, Li P, An J, Akimoto T, Slentz D, Ilkayeva O, Dohm GL, Yan Z, Newgard CB, and Muoio DM (2005). Peroxisome proliferator-activated receptor-gamma co-activator 1alphamediated metabolic remodeling of skeletal myocytes mimics exercise training and reverses lipid-induced mitochondrial inefficiency. J Biol Chem 280, 33588–33598. 10.1074/jbc.M507621200. [DOI] [PubMed] [Google Scholar]
- 13.Muoio DM, and Koves TR (2007). Lipid-induced metabolic dysfunction in skeletal muscle. Novartis Found Symp 286, 24–38; discussion 38–46, 162–163, 196–203. [DOI] [PubMed] [Google Scholar]
- 14.Muoio DM, and Koves TR (2007). Skeletal muscle adaptation to fatty acid depends on coordinated actions of the PPARs and PGC1 alpha: implications for metabolic disease. Appl Physiol Nutr Metab 32, 874–883. 10.1139/H07-083. [DOI] [PubMed] [Google Scholar]
- 15.Ahmad T, Kelly JP, McGarrah RW, Hellkamp AS, Fiuzat M, Testani JM, Wang TS, Verma A, Samsky MD, Donahue MP, et al. (2016). Prognostic Implications of Long-Chain Acylcarnitines in Heart Failure and Reversibility With Mechanical Circulatory Support. Journal of the American College of Cardiology 67, 291–299. 10.1016/j.jacc.2015.10.079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Baker PR 2nd, Boyle KE, Koves TR, Ilkayeva OR, Muoio DM, Houmard JA, and Friedman JE (2015). Metabolomic analysis reveals altered skeletal muscle amino acid and fatty acid handling in obese humans. Obesity (Silver Spring) 23, 981–988. 10.1002/oby.21046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Huffman KM, Redman LM, Landerman LR, Pieper CF, Stevens RD, Muehlbauer MJ, Wenner BR, Bain JR, Kraus VB, Newgard CB, et al. (2012). Caloric restriction alters the metabolic response to a mixed-meal: results from a randomized, controlled trial. PLoS One 7, e28190. 10.1371/journal.pone.0028190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Huffman KM, Shah SH, Stevens RD, Bain JR, Muehlbauer M, Slentz CA, Tanner CJ, Kuchibhatla M, Houmard JA, Newgard CB, and Kraus WE (2009). Relationships between circulating metabolic intermediates and insulin action in overweight to obese, inactive men and women. Diabetes care 32, 1678–1683. 10.2337/dc08-2075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Kien CL, Bunn JY, Poynter ME, Stevens R, Bain J, Ikayeva O, Fukagawa NK, Champagne CM, Crain KI, Koves TR, and Muoio DM (2013). A lipidomics analysis of the relationship between dietary fatty acid composition and insulin sensitivity in young adults. Diabetes 62, 1054–1063. 10.2337/db12-0363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kraus WE, Muoio DM, Stevens R, Craig D, Bain JR, Grass E, Haynes C, Kwee L, Qin X, Slentz DH, et al. (2015). Metabolomic Quantitative Trait Loci (mQTL) Mapping Implicates the Ubiquitin Proteasome System in Cardiovascular Disease Pathogenesis. PLoS Genet 11, e1005553. 10.1371/journal.pgen.1005553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lum H., Sloane R., Huffman KM., Kraus VB., Thompson DK., Kraus WE., Bain JR., Stevens R., Pieper CF., Taylor GA., et al. (2011). Plasma acylcarnitines are associated with physical performance in elderly men. The journals of gerontology. Series A, Biological sciences and medical sciences 66, 548–553. 10.1093/gerona/glr006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Shah AA, Craig DM, Sebek JK, Haynes C, Stevens RC, Muehlbauer MJ, Granger CB, Hauser ER, Newby LK, Newgard CB, et al. (2012). Metabolic profiles predict adverse events after coronary artery bypass grafting. J Thorac Cardiovasc Surg 143, 873–878. 10.1016/j.jtcvs.2011.09.070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Shah SH, Hauser ER, Bain JR, Muehlbauer MJ, Haynes C, Stevens RD, Wenner BR, Dowdy ZE, Granger CB, Ginsburg GS, et al. (2009). High heritability of metabolomic profiles in families burdened with premature cardiovascular disease. Mol Syst Biol 5, 258. 10.1038/msb.2009.11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Koh AS, Gao F, Liu J, Fridianto KT, Ching J, Tan RS, Wong JI, Chua SJ, Leng S, Zhong L, et al. (2018). Metabolomic profile of arterial stiffness in aged adults. Diab Vasc Dis Res 15, 74–80. 10.1177/1479164117733627. [DOI] [PubMed] [Google Scholar]
- 25.Hunter WG, Kelly JP, McGarrah RW 3rd, Khouri MG, Craig D, Haynes C, Ilkayeva O, Stevens RD, Bain JR, Muehlbauer MJ, et al. (2016). Metabolomic Profiling Identifies Novel Circulating Biomarkers of Mitochondrial Dysfunction Differentially Elevated in Heart Failure With Preserved Versus Reduced Ejection Fraction: Evidence for Shared Metabolic Impairments in Clinical Heart Failure. J Am Heart Assoc 5. 10.1161/JAHA.115.003190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Glancy B, Barstow T, and Willis WT (2008). Linear relation between time constant of oxygen uptake kinetics, total creatine, and mitochondrial content in vitro. Am J Physiol Cell Physiol 294, C79–87. 10.1152/ajpcell.00138.2007. [DOI] [PubMed] [Google Scholar]
- 27.Messer JI, Jackman MR, and Willis WT (2004). Pyruvate and citric acid cycle carbon requirements in isolated skeletal muscle mitochondria. Am J Physiol Cell Physiol 286, C565–572. 10.1152/ajpcell.00146.2003. [DOI] [PubMed] [Google Scholar]
- 28.Kuzmiak-Glancy S, and Willis WT (2014). Skeletal muscle fuel selection occurs at the mitochondrial level. J Exp Biol 217, 1993–2003. 10.1242/jeb.098863. [DOI] [PubMed] [Google Scholar]
- 29.Calvo JA, Daniels TG, Wang X, Paul A, Lin J, Spiegelman BM, Stevenson SC, and Rangwala SM (2008). Muscle-specific expression of PPARgamma coactivator-1alpha improves exercise performance and increases peak oxygen uptake. J Appl Physiol (1985) 104, 1304–1312. 10.1152/japplphysiol.01231.2007. [DOI] [PubMed] [Google Scholar]
- 30.Wong KE., Mikus CR., Slentz DH., Seiler SE., DeBalsi KL., Ilkayeva OR., Crai KI., Kinter MT., Kien CL., Stevens RD., and Muoio DM. (2015). Muscle-Specific Overexpression of PGC-1alpha Does Not Augment Metabolic Improvements in Response to Exercise and Caloric Restriction. Diabetes 64, 1532–1543. 10.2337/db14-0827. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Martin OJ, Lai L, Soundarapandian MM, Leone TC, Zorzano A, Keller MP, Attie AD, Muoio DM, and Kelly DP (2014). A role for peroxisome proliferator-activated receptor gamma coactivator-1 in the control of mitochondrial dynamics during postnatal cardiac growth. Circ Res 114, 626–636. 10.1161/CIRCRESAHA.114.302562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Lai L, Leone TC, Keller MP, Martin OJ, Broman AT, Nigro J, Kapoor K, Koves TR, Stevens R, Ilkayeva OR, et al. (2014). Energy metabolic reprogramming in the hypertrophied and early stage failing heart: a multisystems approach. Circ Heart Fail 7, 1022–1031. 10.1161/CIRCHEARTFAILURE.114.001469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Davies MN, Kjalarsdottir L, Thompson JW, Dubois LG, Stevens RD, Ilkayeva OR, Brosnan MJ, Rolph TP, Grimsrud PA, and Muoio DM (2016). The Acetyl Group Buffering Action of Carnitine Acetyltransferase Offsets Macronutrient-Induced Lysine Acetylation of Mitochondrial Proteins. Cell Rep 14, 243–254. 10.1016/j.celrep.2015.12.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Seiler SE, Koves TR, Gooding JR, Wong KE, Stevens RD, Ilkayeva OR, Wittmann AH, DeBalsi KL, Davies MN, Lindeboom L, et al. (2015). Carnitine Acetyltransferase Mitigates Metabolic Inertia and Muscle Fatigue during Exercise. Cell Metab 22, 65–76. 10.1016/j.cmet.2015.06.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Muoio DM, Noland RC, Kovalik JP, Seiler SE, Davies MN, DeBalsi KL, Ilkayeva OR, Stevens RD, Kheterpal I, Zhang J, et al. (2012). Muscle-specific deletion of carnitine acetyltransferase compromises glucose tolerance and metabolic flexibility. Cell metabolism 15, 764–777. 10.1016/j.cmet.2012.04.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Randle PJ (1998). Regulatory interactions between lipids and carbohydrates: the glucose fatty acid cycle after 35 years. Diabetes Metab Rev 14, 263–283. . [DOI] [PubMed] [Google Scholar]
- 37.Aubert G, Martin OJ, Horton JL, Lai L, Vega RB, Leone TC, Koves T, Gardell SJ, Kruger M, Hoppel CL, et al. (2016). The Failing Heart Relies on Ketone Bodies as a Fuel. Circulation 133, 698–705. 10.1161/CIRCULATIONAHA.115.017355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.McCommis KS, Kovacs A, Weinheimer CJ, Shew TM, Koves TR, Ilkayeva OR, Kamm DR, Pyles KD, King MT, Veech RL, et al. (2020). Nutritional modulation of heart failure in mitochondrial pyruvate carrier-deficient mice. Nat Metab 2, 1232–1247. 10.1038/s42255-020-00296-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Davidson MT, Grimsrud PA, Lai L, Draper JA, Fisher-Wellman KH, Narowski TM, Abraham DM, Koves TR, Kelly DP, and Muoio DM (2020). Extreme Acetylation of the Cardiac Mitochondrial Proteome Does Not Promote Heart Failure. Circ Res 127, 1094–1108. 10.1161/CIRCRESAHA.120.317293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Wende AR, Schaeffer PJ, Parker GJ, Zechner C, Han DH, Chen MM, Hancock CR, Lehman JJ, Huss JM, McClain DA, et al. (2007). A role for the transcriptional coactivator PGC-1alpha in muscle refueling. J Biol Chem 282, 36642–36651. 10.1074/jbc.M707006200. [DOI] [PubMed] [Google Scholar]
- 41.Baar K, Wende AR, Jones TE, Marison M, Nolte LA, Chen M, Kelly DP, and Holloszy JO (2002). Adaptations of skeletal muscle to exercise: rapid increase in the transcriptional coactivator PGC-1. FASEB J 16, 1879–1886. 10.1096/fj.02-0367com. [DOI] [PubMed] [Google Scholar]
- 42.Holloszy JO (1975). Adaptation of skeletal muscle to endurance exercise. Med Sci Sports 7, 155–164. [PubMed] [Google Scholar]
- 43.Martines AMF, van Eunen K, Reijngoud DJ, and Bakker BM (2017). The promiscuous enzyme medium-chain 3-keto-acyl-CoA thiolase triggers a vicious cycle in fatty-acid beta-oxidation. PLoS Comput Biol 13, e1005461. 10.1371/journal.pcbi.1005461. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Wei J, Kang HW, and Cohen DE (2009). Thioesterase superfamily member 2 (Them2)/acyl-CoA thioesterase 13 (Acot13): a homotetrameric hotdog fold thioesterase with selectivity for long-chain fatty acyl-CoAs. Biochem J 421, 311–322. 10.1042/BJ20090039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Ellis JM, Mentock SM, Depetrillo MA, Koves TR, Sen S, Watkins SM, Muoio DM, Cline GW, Taegtmeyer H, Shulman GI, et al. (2011). Mouse cardiac acyl coenzyme a synthetase 1 deficiency impairs Fatty Acid oxidation and induces cardiac hypertrophy. Mol Cell Biol 31, 1252–1262. 10.1128/MCB.01085-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Makowski L., Noland RC., Koves TR., Xing W., Ilkayeva OR., Muehlbauer MJ., Stevens RD., and Muoio DM. (2009). Metabolic profiling of PPARalpha−/− mice reveals defects in carnitine and amino acid homeostasis that are partially reversed by oral carnitine supplementation. FASEB J 23, 586–604. 10.1096/fj.08-119420. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Lopaschuk GD, Karwi QG, Tian R, Wende AR, and Abel ED (2021). Cardiac Energy Metabolism in Heart Failure. Circ Res 128, 1487–1513. 10.1161/CIRCRESAHA.121.318241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Chiao YA, Chakraborty AD, Light CM, Tian R, Sadoshima J, Shi X, Gu H, and Lee CF (2021). NAD(+) Redox Imbalance in the Heart Exacerbates Diabetic Cardiomyopathy. Circ Heart Fail 14, e008170. 10.1161/CIRCHEARTFAILURE.120.008170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Zhang Y, Taufalele PV, Cochran JD, Robillard-Frayne I, Marx JM, Soto J, Rauckhorst AJ, Tayyari F, Pewa AD, Gray LR, et al. (2020). Mitochondrial pyruvate carriers are required for myocardial stress adaptation. Nat Metab 2, 1248–1264. 10.1038/s42255-020-00288-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.van Eunen K, Volker-Touw CM, Gerding A, Bleeker A, Wolters JC, van Rijt WJ, Martines AM, Niezen-Koning KE, Heiner RM, Permentier H, et al. (2016). Living on the edge: substrate competition explains loss of robustness in mitochondrial fatty-acid oxidation disorders. BMC Biol 14, 107. 10.1186/s12915-016-0327-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.van Eunen K, Simons SM, Gerding A, Bleeker A, den Besten G, Touw CM, Houten SM, Groen BK, Krab K, Reijngoud DJ, and Bakker BM (2013). Biochemical competition makes fatty-acid beta-oxidation vulnerable to substrate overload. PLoS Comput Biol 9, e1003186. 10.1371/journal.pcbi.1003186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.An J, Muoio DM, Shiota M, Fujimoto Y, Cline GW, Shulman GI, Koves TR, Stevens R, Millington D, and Newgard CB (2004). Hepatic expression of malonyl-CoA decarboxylase reverses muscle, liver and whole-animal insulin resistance. Nature medicine 10, 268–274. [DOI] [PubMed] [Google Scholar]
- 53.Koves TR, Li P, An J, Akimoto T, Slentz D, Ilkayeva O, Dohm GL, Yan Z, Newgard CB, and Muoio DM (2005). Peroxisome proliferator-activated receptor-γ co-activator 1α-mediated metabolic remodeling of skeletal myocytes mimics exercise training and reverses lipid-induced mitochondrial inefficiency. Journal of Biological Chemistry 280, 33588–33598. [DOI] [PubMed] [Google Scholar]
- 54.Muoio DM, and Neufer PD (2012). Lipid-induced mitochondrial stress and insulin action in muscle. Cell Metab 15, 595–605. 10.1016/j.cmet.2012.04.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Lin J, Wu H, Tarr PT, Zhang CY, Wu Z, Boss O, Michael LF, Puigserver P, Isotani E, Olson EN, et al. (2002). Transcriptional co-activator PGC-1 alpha drives the formation of slow-twitch muscle fibres. Nature 418, 797–801. 10.1038/nature00904. [DOI] [PubMed] [Google Scholar]
- 56.Frezza C, Cipolat S, and Scorrano L. (2007). Organelle isolation: functional mitochondria from mouse liver, muscle and cultured fibroblasts. Nature protocols 2, 287–295. 10.1038/nprot.2006.478. [DOI] [PubMed] [Google Scholar]
- 57.Glancy B, Willis WT, Chess DJ, and Balaban RS (2013). Effect of calcium on the oxidative phosphorylation cascade in skeletal muscle mitochondria. Biochemistry 52, 2793–2809. 10.1021/bi3015983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Messer JI, Jackman MR, and Willis WT (2004). Pyruvate and citric acid cycle carbon requirements in isolated skeletal muscle mitochondria. American Journal of Physiology - Cell Physiology 286, C565–C572. 10.1152/ajpcell.00146.2003. [DOI] [PubMed] [Google Scholar]
- 59.Scaduto RC Jr, and Grotyohann LW (1999). Measurement of Mitochondrial Membrane Potential Using Fluorescent Rhodamine Derivatives. Biophysical journal 76, 469–477. 10.1016/S0006-3495(99)77214-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Krumschnabel G., Eigentler A., Fasching M., and Gnaiger E. (2014). Use of safranin for the assessment of mitochondrial membrane potential by high-resolution respirometry and fluorometry. Methods Enzymol 542, 163–181. 10.1016/B978-0-12-416618-9.00009-1. [DOI] [PubMed] [Google Scholar]
- 61.Muoio DM, Way JM, Tanner CJ, Winegar DA, Kliewer SA, Houmard JA, Kraus WE, and Dohm GL (2002). Peroxisome proliferator-activated receptor-alpha regulates fatty acid utilization in primary human skeletal muscle cells. Diabetes 51, 901–909. [DOI] [PubMed] [Google Scholar]
- 62.Koves TR, Sparks LM, Kovalik JP, Mosedale M, Arumugam R, DeBalsi KL, Everingham K, Thorne L, Phielix E, Meex RC, et al. (2013). PPARgamma coactivator-1alpha contributes to exercise-induced regulation of intramuscular lipid droplet programming in mice and humans. Journal of lipid research 54, 522–534. 10.1194/jlr.P028910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Wang Y, Christopher BA, Wilson KA, Muoio D, McGarrah RW, Brunengraber H, and Zhang GF (2018). Propionate-induced changes in cardiac metabolism, notably CoA trapping, are not altered by L-carnitine. American journal of physiology. Endocrinology and metabolism. 10.1152/ajpendo.00081.2018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Li Q, Zhang S, Berthiaume JM, Simons B, and Zhang GF (2014). Novel approach in LC-MS/MS using MRM to generate a full profile of acyl-CoAs: discovery of acyl-dephospho-CoAs. Journal of lipid research 55, 592–602. 10.1194/jlr.D045112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Zhang GF, Kombu RS, Kasumov T, Han Y, Sadhukhan S, Zhang J, Sayre LM, Ray D, Gibson KM, Anderson VA, et al. (2009). Catabolism of 4-hydroxyacids and 4-hydroxynonenal via 4-hydroxy-4-phosphoacyl-CoAs. The Journal of biological chemistry 284, 33521–33534. 10.1074/jbc.M109.055665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Fernandez CA, Des Rosiers C, Previs SF, David F, and Brunengraber H. (1996). Correction of 13C mass isotopomer distributions for natural stable isotope abundance. Journal of mass spectrometry : JMS 31, 255–262. . [DOI] [PubMed] [Google Scholar]
- 67.Tomcik K, Ibarra RA, Sadhukhan S, Han Y, Tochtrop GP, and Zhang GF (2011). Isotopomer enrichment assay for very short chain fatty acids and its metabolic applications. Analytical biochemistry 410, 110–117. 10.1016/j.ab.2010.11.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.DeBalsi KL., Wong KE., Koves TR., Slentz DH., Seiler SE., Wittmann AH., Ilkayeva OR., Stevens RD., Perry CG., Lark DS., et al. (2014). Targeted metabolomics connects thioredoxin-interacting protein (TXNIP) to mitochondrial fuel selection and regulation of specific oxidoreductase enzymes in skeletal muscle. The Journal of biological chemistry 289, 8106–8120. 10.1074/jbc.M113.511535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Käll L, Canterbury JD, Weston J, Noble WS, and MacCoss MJ (2007). Semi-supervised learning for peptide identification from shotgun proteomics datasets. Nature Methods 4, 923–925. 10.1038/nmeth1113. [DOI] [PubMed] [Google Scholar]
- 70.Calvo SE, Clauser KR, and Mootha VK (2015). MitoCarta2.0: an updated inventory of mammalian mitochondrial proteins. Nucleic Acids Research 44, D1251–D1257. 10.1093/nar/gkv1003. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data S3. Uncropped images of all western blots, related to Figures 2 and 7, and Supplemental Figures S4 and S5.
Data S4. Data used to generate all figures.
Data S1. Mitochondrial Proteomics, Heart versus SkM, related to Figure 5.
Data S2. Mitochondrial Proteomics, PGC1a versus NT SkM, related to Figure 5.
S. Table 2. Mitochondrial Proteomics, Heart versus SkM, related to Figure 5.
S. Table 3. Mitochondrial Proteomics, PGC1a versus NT SkM, related to Figure 5.
Data Availability Statement
Proteomics data have been deposited at ProteomeXchange and jPOST and are publicly available as of the date of publication. Accession numbers are listed in the key resources table. Original western blot images have been shown in Supplemental data. A file with all data presented in figures has been including in Supplemental data.
This manuscript does not report any original code.
Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| MKT (Acaa2) | Millipore Sigma | WH0010449M1; RRID:AB_2219394 |
| CrAT | ABCAM | ab153750; RRID:AB_2935643 |
| SCHAD | ProteinTech | 19828–1AP; RRID:AB_10667408 |
| ATP5A | ABCAM | ab14748; RRID:AB_301447 |
| OxPHOS Cocktail | ab110413; RRID:AB_2629281 | |
| SDHA | ABCAM | ab14715; RRID:AB_301433 |
| NDUFA9 | ABCAM | ab14713; RRID:AB_301431 |
| T-AMPK | CST | 2532; RRID:AB_330331 |
| P-AMPK | CST | 2531; RRID:AB_330330 |
| HSP60 | CST | 12165; RRID:AB_2636980 |
| Chemicals, peptides, and recombinant proteins | ||
| MOPS Free Acid | Millipore Sigma | Cat# M1254 CAS# 1132–61-2 |
| MES Potassium Salt | Millipore Sigma | Cat# M0895 CAS# 39946–25-3 |
| Bovine Serum Albumin (Fatty Acid Free) | Millipore Sigma | Cat# A3803 CAS# 9048–46-6 |
| EDTA | Millipore Sigma | Cat# E0270 CAS# 65501–24-8 |
| Trypsin from Porcine Pancreas (Mitochondrial Isolation) | Millipore Sigma | Cat# T4799 CAS# 9001–51-8 |
| Potassium Chloride | Millipore Sigma | Cat# P5405 CAS# 7447–40-7 |
| Magnesium Sulfate | ||
| Magnesium Chloride Hexahydrate | Millipore Sigma | Cat# M2670 CAS# 7791–18-6 |
| EGTA | Millipore Sigma | Cat# E4378 CAS# 67–42-5 |
| Potassium Dihydrogen Phosphate | Millipore Sigma | Cat# P9791 CAS# 7778–77-0 |
| Creatine Monohydrate | Millipore Sigma | Cat# C3630 CAS# 6020–87-7 |
| Potassium Salt of Phosphocreatine | Millipore Sigma | |
| Palmitoyl-L-carnitine | Millipore Sigma | Cat# P1645 CAS# 18877–64-0 |
| Octanoyl-L-carnitine | Millipore Sigma | Cat# 50892 CAS# 25243–95-2 |
| L-Carnitine hydrochloride | ||
| Malic Acid (Malate) | Millipore Sigma | Cat# M1000 CAS# 97–67-6 |
| α-ketoglutaric acid (aKG) | MilliporeSigma | Cat# K1750 CAS# 328–50-7 |
| (R)-3-Hydroxybutyric acid | Millipore Sigma | Cat# 54920 CAS# 625–72-9 |
| Succinic Acid (Succinate) | Millipore Sigma | Cat# S3674 CAS# 110–15-6 |
| Potassium Pyruvate | Combi-Blocks | Cat# QA-1116 CAS# 4151–33-1 |
| Creatine Kinase from Rabbit Muscle | Roche | Cat# 10127566001 |
| Tetramethylrhodamine Methyl Ester (TMRM) | ThermoFisher | Cat# T668 |
| Rotenone | Millipore Sigma | Cat# R8875 CAS# 83–79-4 |
| Potassium Cyanide | Millipore Sigma | Cat# 60178 CAS# 151–50-8 |
| Alamethicin | AG Scientific | A-1286 CAS#27061–78-5 |
| Methanol | Millipore Sigma | Cat# 439193 CAS# 67–56-1 |
| Protease Inhibitor Cocktail | Millipore Sigma | Cat# P8340 |
| Phosphatase Inhibitor Cocktail 2 | Millipore Sigma | Cat# P5726 |
| Phosphatase Inhibitor Cocktail 3 | Millipore Sigma | Cat# P0044 |
| Pierce Reversible Protein Stain Kit for Nitrocellulose Membranes (Memcode) |
ThermoFisher Scientific |
Cat# 24580 |
| 4–15% Criterion TGX Stain-Free Protein Gel, 18well | Biorad | Cat# 5678084 |
| 10X Tris Glycine SDS Running Buffer | Biorad | Cat# 1610732 |
| 10X Tris Buffered Saline | Biorad | Cat# 1706435 |
| Fish Gelatin | Millipore Sigma | Cat# G7765 |
| Casein | Millipore Sigma | Cat# C0626 |
| Roche cOmplete ULTRA EDTA-free Protease Inhibitor Mini Tablet | Millipore Sigma | Cat# 05892791001 |
| Roche 1x PhosSTOP Phosphatase Inhibitor Cocktail Tablets | Millipore Sigma | Cat# 04906837001 |
| Lysyl Endopeptidase, Mass Spectrometry Grade | Wako Chemicals | Cat# 125–05061 |
| Sequencing Grade Modified Trypsin | Promega | Cat# V5113 |
| tC18 SEP-PAK Solid Phase Extraction Columns (50 mg) | Waters | Cat# WAT054960 |
| tC18 SEP-PAK Solid Phase Extraction Columns (100 mg) | Waters | Cat# WAT036820 |
| Triethylammonium bicarbonate (TEAB) | ThermoFisher | Cat# 90114 |
| TRIzol Reagent | ThermoFisher | Cat# 15596026 |
| Chloroform | Millipore Sigma | Cat# C2432 CAS# 67–66-3 |
| [U-13C]-Glucose | ||
| [U-13C]-Pyruvate | ||
| CelLyticmtCelLytic™ MT Cell Lysis Reagent | SigmaAldrich | C3228 |
| Pentobarbital (Nembutal) | Oak Pharmaceuticals | |
| Critical commercial assays | ||
| Pierce BCA Protein Assay | ThermoFisher | Cat# 23225 |
| RNeasy mini spin columns | Qiagen | Cat# 74004 |
| iScript cDNA Synthesis Kit | BioRad | Cat# 170–8891 |
| PrimeTime Gene Expression Master Mix | IDT | Cat# 1055770 |
| TaqMan Gene Expression Assay Acaa2 | ThermoFisher | Mm00624282 |
| Lipofectamine RNAiMAX | ThermoFisher | 13778 |
| XF Plasma Membrane Permeabilizer | Agilent Technologies | 102504–100 |
| Deposited data | ||
| Proteomics Raw Data Files | This Publication | PXD040326; JPST002051 |
| Proteomics Raw Data Files | https://pubmed.ncbi.nlm.nih.gov/32660330/ | Experiment 3; Supplemental data; Protein SUMMARY_Exp3 Tab |
| Data S4 (Excel File) | This Publication | |
| Experimental models: Organisms/strains | ||
| C57BL/6NJ mice | The Jackson Laboratory | Stock #005304 |
| MCK-Pgc1a C57BL/6-Tg(Ckm-Ppargc1a)31Brsp/J | Dr. Bruce Speigelman | https://pubmed.ncbi.nlm.nih.gov/12181572 |
| MCK-CrAT C57BL/6J | Dr. Randall Mynatt (Pennington Biomedical Research Center) | https://pubmed.ncbi.nlm.nih.gov/22560225/ |
| Oligonucleotides | ||
| ACAA2/MKT siRNA | IDT | hs.Ri.ACAA2 - 13.1, 13.2, 13.3 |
| Non-targeting control sequence | IDT | 51–01-14–03 |
| Software and algorithms | ||
| Proteome Discoverer 2.2 | ThermoFisher | RRID:SCR_014477 |
| DatLab 7 | Oroboros Instruments | Product# 20700 |
| Other | ||
| Oxygraph-2k | Oroboros Instruments | Product# O2k-Core |
| QuantaMaster Spectrofluorometer | Horiba Scientific | Cat# QM-400 |
| Spectrosil Quartz Cuvettes | Starna | 23–5.45-Q-5 |
| Spectromax M2E Spectrophotometer | Molecular Devices | Part#: M2E |
| Thermo Fisher Scientific Q Exactive Plus Orbitrap Mass Spectrometer | ThermoScientific | Cat#: 0726030 |
| Thermo Fisher Scientific nanoEASY nLC | ThermoScientific | Cat #: LC140 |
| Waters Xevo TQ-S triple quadrupole mass spectrometer coupled to a Waters Acquity UPLC system | Waters | Part#: Xevo TQ-S |
| Bio-Rad Turboblot Transfer System | Biorad | Cat# 1704150EDU |
| TissueLyser II | Qiagen | Cat# 85300 |
| NanoDrop 8000 | ThermoScientific | ND-8000-GL |
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
| Antibodies | ||
| Rabbit monoclonal anti-Snail | Cell Signaling Technology | Cat#3879S; RRID: AB_2255011 |
| Mouse monoclonal anti-Tubulin (clone DM1A) | Sigma-Aldrich | Cat#T9026; RRID: AB_477593 |
| Rabbit polyclonal anti-BMAL1 | This paper | N/A |
| Bacterial and virus strains | ||
| pAAV-hSyn-DIO-hM3D(Gq)-mCherry | Krashes et al.1 | Addgene AAV5; 44361-AAV5 |
| AAV5-EF1a-DIO-hChR2(H134R)-EYFP | Hope Center Viral Vectors Core | N/A |
| Cowpox virus Brighton Red | BEI Resources | NR-88 |
| Zika-SMGC-1, GENBANK: KX266255 | Isolated from patient (Wang et al.2) | N/A |
| Staphylococcus aureus | ATCC | ATCC 29213 |
| Streptococcus pyogenes: M1 serotype strain: strain SF370; M1 GAS | ATCC 700294 | |
| Biological samples | ||
| Healthy adult BA9 brain tissue | University of Maryland Brain & Tissue Bank; http://medschool.umaryland.edu/btbank/ | Cat#UMB1455 |
| Human hippocampal brain blocks | New York Brain Bank | http://nybb.hs.columbia.edu/ |
| Patient-derived xenografts (PDX) | Children’s Oncology Group Cell Culture and Xenograft Repository |
http://cogcell.org/ |
| Chemicals, peptides, and recombinant proteins | ||
| MK-2206 AKT inhibitor | Selleck Chemicals | S1078; CAS: 1032350–13-2 |
| SB-505124 | Sigma-Aldrich | S4696; CAS: 694433–59-5 (free base) |
| Picrotoxin | Sigma-Aldrich | P1675; CAS: 12487–8 |
| Human TGF-β | R&D | 240-B; GenPept: P01137 |
| Activated S6K1 | Millipore | Cat#14–486 |
| GST-BMAL1 | Novus | Cat#H00000406P01 |
| Critical commercial assays | ||
| EasyTag EXPRESS 35S Protein Labeling Kit | PerkinElmer | NEG772014MC |
| CaspaseGlo 3/7 | Promega | G8090 |
| TruSeq ChIP Sample Prep Kit | Illumina | IP-202–1012 |
| Deposited data | ||
| Raw and analyzed data | This paper | GEO: GSE63473 |
| B-RAF RBD (apo) structure | This paper | PDB: 5J17 |
| Human reference genome NCBI build 37, GRCh37 | Genome Reference Consortium | http://www.ncbi.nlm.nih.gov/projects/gen ome/assembly/grc/h uman/ |
| Nanog STILT inference | This paper; Mendeley Data | http://dx.doi.org/10.17632/wx6s4mj7s8.2 |
| Affinity-based mass spectrometry performed with 57 genes | This paper; Mendeley Data | Table S8; http://dx.doi.org/10.17632/5hvpvspw82.1 |
| Experimental models: Cell lines | ||
| Hamster: CHO cells | ATCC | CRL-11268 |
| D. melanogaster: Cell line S2: S2-DRSC | Laboratory of Norbert Perrimon | FlyBase: FBtc0000181 |
| Human: Passage 40 H9 ES cells | MSKCC stem cell core facility | N/A |
| Human: HUES 8 hESC line (NIH approval number NIHhESC-09–0021) | HSCI iPS Core | hES Cell Line: HUES-8 |
| Experimental models: Organisms/strains | ||
| C. elegans: Strain BC4011: srl-1(s2500) II; dpy18(e364) III; unc-46(e177)rol-3(s1040) V. | Caenorhabditis Genetics Center | WB Strain: BC4011; WormBase: WBVar00241916 |
| D. melanogaster: RNAi of Sxl: y[1] sc[*] v[1]; P{TRiP.HMS00609}attP2 | Bloomington Drosophila Stock Center | BDSC:34393; FlyBase: FBtp0064874 |
| S. cerevisiae: Strain background: W303 | ATCC | ATTC: 208353 |
| Mouse: R6/2: B6CBA-Tg(HDexon1)62Gpb/3J | The Jackson Laboratory | JAX: 006494 |
| Mouse: OXTRfl/fl: B6.129(SJL)-Oxtrtm1.1Wsy/J | The Jackson Laboratory | RRID: IMSR_JAX:008471 |
| Zebrafish: Tg(Shha:GFP)t10: t10Tg | Neumann and NuessleinVolhard3 | ZFIN: ZDB-GENO060207–1 |
| Arabidopsis: 35S::PIF4-YFP, BZR1-CFP | Wang et al.4 | N/A |
| Arabidopsis: JYB1021.2: pS24(AT5G58010)::cS24:GFP(-G):NOS #1 | NASC | NASC ID: N70450 |
| Oligonucleotides | ||
| siRNA targeting sequence: PIP5K I alpha #1: ACACAGUACUCAGUUGAUA | This paper | N/A |
| Primers for XX, see Table SX | This paper | N/A |
| Primer: GFP/YFP/CFP Forward: GCACGACTTCTTCAAGTCCGCCATGCC | This paper | N/A |
| Morpholino: MO-pax2a GGTCTGCTTTGCAGTGAATATCCAT |
Gene Tools | ZFIN: ZDB-MRPHLNO-061106-5 |
| ACTB (hs01060665_g1) | Life Technologies | Cat#4331182 |
| RNA sequence: hnRNPA1_ligand: UAGGGACUUAGGGUUCUCUCUAGGGACUUAG GGUUCUCUCUAGGGA | This paper | N/A |
| Recombinant DNA | ||
| pLVX-Tight-Puro (TetOn) | Clonetech | Cat#632162 |
| Plasmid: GFP-Nito | This paper | N/A |
| cDNA GH111110 | Drosophila Genomics Resource Center | DGRC:5666; FlyBase:FBcl0130415 |
| AAV2/1-hsyn-GCaMP6-WPRE | Chen et al.5 | N/A |
| Mouse raptor: pLKO mouse shRNA 1 raptor | Thoreen et al.6 | Addgene Plasmid #21339 |
| Software and algorithms | ||
| ImageJ | Schneider et al.7 | https://imagej.nih.gov/ij/ |
| Bowtie2 | Langmead and Salzberg8 | http://bowtiebio.sourceforge.net/bowtie2/index.shtml |
| Samtools | Li et al.9 | http://samtools.sourceforge.net/ |
| Weighted Maximal Information Component Analysis v0.9 | Rau et al.10 | https://github.com/ChristophRau/wMICA |
| ICS algorithm | This paper; Mendeley Data | http://dx.doi.org/10.17632/5hvpvspw82.1 |
| Other | ||
| Sequence data, analyses, and resources related to the ultra-deep sequencing of the AML31 tumor, relapse, and matched normal | This paper | http://aml31.genome.wustl.edu |
| Resource website for the AML31 publication | This paper | https://github.com/chrisamiller/aml31SuppSite |
| Chemicals, peptides, and recombinant proteins | ||
| QD605 streptavidin conjugated quantum dot | Thermo Fisher Scientific | Cat#Q10101MP |
| Platinum black | Sigma-Aldrich | Cat#205915 |
| Sodium formate BioUltra, ≥99.0% (NT) | Sigma-Aldrich | Cat#71359 |
| Chloramphenicol | Sigma-Aldrich | Cat#C0378 |
| Carbon dioxide (13C, 99%) (<2% 18O) | Cambridge Isotope Laboratories | CLM-185–5 |
| Poly(vinylidene fluoride-co-hexafluoropropylene) | Sigma-Aldrich | 427179 |
| PTFE Hydrophilic Membrane Filters, 0.22 μm, 90 mm | Scientificfilters.com/TischScientific | SF13842 |
| Critical commercial assays | ||
| Folic Acid (FA) ELISA kit | Alpha Diagnostic International | Cat# 0365–0B9 |
| TMT10plex Isobaric Label Reagent Set | Thermo Fisher | A37725 |
| Surface Plasmon Resonance CM5 kit | GE Healthcare | Cat#29104988 |
| NanoBRET Target Engagement K-5 kit | Promega | Cat#N2500 |
| Deposited data | ||
| B-RAF RBD (apo) structure | This paper | PDB: 5J17 |
| Structure of compound 5 | This paper; Cambridge Crystallographic Data Center | CCDC: 2016466 |
| Code for constraints-based modeling and analysis of autotrophic E. coli | This paper | https://gitlab.com/elad.noor/sloppy/tree/master/rubisco |
| Software and algorithms | ||
| Gaussian09 | Frish et al.1 | https://gaussian.com |
| Python version 2.7 | Python Software Foundation | https://www.python.org |
| ChemDraw Professional 18.0 | PerkinElmer | https://www.perkinelmer.com/category/chemdraw |
| Weighted Maximal Information Component Analysis v0.9 | Rau et al.2 | https://github.com/ChristophRau/wMICA |
| Other | ||
| DASGIP MX4/4 Gas Mixing Module for 4 Vessels with a Mass Flow Controller | Eppendorf | Cat#76DGMX44 |
| Agilent 1200 series HPLC | Agilent Technologies | https://www.agilent.com/en/products/liquid-chromatography |
| PHI Quantera II XPS | ULVAC-PHI, Inc. | https://www.ulvacphi.com/en/products/xps/phi-quantera-ii/ |