Summary
Background
While advancements in imaging techniques have led to major strides in deciphering the human brain, successful interventions are elusive and represent some of the most persistent translational gaps in medicine. Human restricted CHRFAM7A has been associated with neuropsychiatric disorders.
Methods
The physiological role of CHRFAM7A in human brain is explored using multiomics approach on 600 post mortem human brain tissue samples. The emerging pathways and mechanistic hypotheses are tested and validated in an isogenic hiPSC model of CHRFAM7A knock-in medial ganglionic eminence progenitors and neurons.
Findings
CHRFAM7A is identified as a modulator of intracellular calcium dynamics and an upstream regulator of Rac1. Rac1 activation re-designs the actin cytoskeleton leading to dynamic actin driven remodeling of membrane protrusion and a switch from filopodia to lamellipodia. The reinforced cytoskeleton leads to an advantage to tolerate stiffer mechanical properties of the extracellular environment.
Interpretation
CHRFAM7A modifies the actin cytoskeleton to a more dynamic and stiffness resistant state in an α7nAChR dependent manner. CHRFAM7A may facilitate neuronal adaptation to changes in the brain environment in physiological and pathological conditions contributing to risk or recovery. Understanding how CHRFAM7A affects human brain requires human studies in the areas of memory formation and erasure, cognitive reserve, and neuronal plasticity.
Funding
This work is supported in part by the Community Foundation for Greater Buffalo (Kinga Szigeti). Also, in part by the International Society for Neurochemistry (ISN) and The Company of Biologists (Nicolas Rosas). ROSMAP is supported by NIA grants P30AG10161, P30AG72975, R01AG15819, R01AG17917. U01AG46152, and U01AG61356.
Keywords: Human brain, Multiomics analysis, CHRFAM7A, Actin cytoskeleton, iPSC
Research in context.
Evidence before this study
Human restricted genes have been implicated in the evolution of the human brain. CHRFAM7A is one of these human restricted fusion genes but with unique characteristics, such as having two alleles in direct and inverted orientation and population genetics of 75% carriers of the direct, presumed functional allele. CHRFAM7A is implicated in the risk of neuropsychiatric disorders and treatment response to cholinergic therapies. Mechanistic insights to CHRFAM7A function are limited to the direct allele's contribution to the α7nAChR pentamer leading to a hypomorphic receptor.
Added value of this study
Based on omics data we performed an unbiased exploratory analysis to generate testable hypotheses regarding CHRFAM7A function in the human brain. The emerging mechanisms then were validated in an isogenic human iPSC model system. We confirmed that only the direct allele has an effect on α7nAChR receptor function, and for the first time we demonstrated that CHRFAM7A modifies Ca2+ dynamics by decreasing amplitude and prolonging the Ca2+ wave. This change in Ca2+ dynamics is decoded by activation of Rac1 resulting in actin cytoskeleton changes. The actin cytoskeleton switches from filopodia to lamellopodia at all structural levels of the neuron and makes the actin cytoskeleton dynamic. The strengthened actin cytoskeleton facilitates adaptation to changes in tissue stiffness.
Implications of all the available evidence
CHRFAM7A direct allele thus represents an actin cytoskeleton gain of function in 75% of the human population. While overt developmental phenotypes in humans are not seen, these data implicate that the human population has a dichotomy (direct allele carrier versus noncarrier) in brain characteristics that may contribute to risk of neuropsychiatric diseases and to the brain's ability to cope with pathology such as stroke, brain inflammation and brain cancer. Human studies in health and disease are needed to explore the full impact of CHRFAM7A.
Introduction
Human specific fusion genes that have emerged since the human-chimpanzee divergence (e.g. NOTCH2NL, ARHGAP11B, SRGAP2) demonstrated the ability to drive brain development and structure.1,2 CHRFAM7A is one of these human specific fusion genes, where the fusion occurred between the alpha 7 nicotinic acetyl-choline receptor (α7 nAChR) and ULK4, a member of the ULK kinase family.3 Despite its widespread implication in neuropsychiatric diseases, including Alzheimer's disease (AD), schizophrenia, anxiety, and ADHD,3, 4, 5, 6 CHRFAM7A neurobiology remains largely unexplored. The gene underwent strong selective pressure leading to its presence in over 99% of humans.7 CHRFAM7A can be present in 0–4 copies in either direct or inverted orientation, but only the direct allele is translated into the fusion protein.3,7 Understanding the physiological function of CHRFAM7A in the human brain is the first step in deciphering its impact on neuropsychiatric diseases.
α7 nAChR is a homopentamer acetylcholine receptor with a role in fast synaptic transmission. In humans, CHRFAM7A can substitute 1–3 subunits in the α7 nAChR.8,9 CHRFAM7A modifies α7 nAChR to a hypomorphic ionotropic receptor with decreased channel open probability.9, 10, 11, 12, 13 α7 nAChR affects Ca2+-induced Ca2+ release (CICR) through its high Ca2+ conductance from the extracellular space (related to ionotropic function) and IP3-induced Ca2+ release (IICR) via phospholipase C (PLC) activation and IP3 signalling for endoplasmic reticulum (ER) Ca2+ release (related to metabotropic function).14 However, the effect of CHRFAM7A on Ca2+ signalling in neurons is unknown.
The function of CHRFAM7A is best studied in the human context and while live human brain tissue is not accessible, large post mortem datasets could generate testable hypotheses.15 We utilized human transcriptomics15 to uncover CHRFAM7A associated phenotypes and generated human isogenic iPSC derived medial ganglionic eminence (MGE) neuronal progenitors and neurons to decipher the molecular mechanism.5,7,10,16, 17, 18
Methods
Ethics
The Institutional Review Board approved the ROSMAP study.
ROSMAP dataset
The ROS and MAP are longitudinal observational cohort studies of older persons without known dementia at enrollment with deep phenotyping and large-scale omic data generation from the human brain as all participants are organ donors.19 Each study was approved by an Institutional Review Board of Rush University Medical Center and all participants signed an informed consent, an Anatomic Gift Act, and a repository consent. Previous reports described the study design and data collection scheme of each study in detail.15 The autopsy rate in these studies exceeds 86%. DLFC brain tissue undergoes multiomics data generation for each sample, albeit not complete in each case.
Phenotypic and multiomic data are accruing in ROS and MAP and are available for data structure review at the RADC Research Resource Sharing Hub (https://www.radc.rush.edu/). The dataset is shared through Accelerating Medicines Partnership for Alzheimer's disease (AMP-AD) Knowledge Portal (https://www.synapse.org/ampad).
In the ROS and MAP studies, subjects undergo yearly assessments. A final clinical diagnosis is made after death following review of select clinical data while being shielded from pathological data. These diagnoses are based on clinical diagnostic criteria and include Alzheimer's disease (AD), mild cognitive impairment (MCI), or no cognitive impairment (NCI) as previously described.20, 21, 22 Neuropathologic diagnoses are also assigned. Pathologic diagnosis of AD is based on the modified NIA Reagan criteria.
Transciptomics data
Briefly, RNAseq was performed at the Broad Institute's Genomics Platform.19 Detailed method and QC protocol were published and are available in synapse. Sequencing was carried out using the Illumina HiSeq2000 with 101 bp paired end reads for a targeted coverage of 50 M paired reads which were met on average.
Picard metrics (http://broadinstitute.github.io/picard/) from alignment by Tophat31 to the whole human genome reference (hg19) with Bowtie1 as the aligner were collected. The expression levels of genes and transcripts were estimated by RSEM package32. The Gencode V14 annotations were used by RSEM in the quantification process. Fragments Per Kilobase of transcript per Million mapped reads (FPKM) values were the final output of our RNA-Seq pipeline.
SRM proteomics data
Briefly, SRM proteomics were generated at Pacific Northwest National Laboratories.23 On average ∼20 mg of DLPFC brain tissue from each subject was homogenized, alkylated with iodoacetamide followed by digestion with trypsin. The digests were cleaned using C18 solid phase extraction. LC-SRM experiments were performed on a nanoACQUITY UPLC (Waters) coupled to a TSQ Vantage mass spectrometer (ThermoScientific). SRM data were analyzed using the Skyline software package. The peak area ratios of endogenous light peptides and their heavy isotope-labeled internal standards (i.e., L/H peak area ratios) were then automatically calculated by the Skyline software, and the best transition without matrix interference was used for accurate quantification. The signal-to-noise ratio in the quantification of each peptide was calculated as the ratio of variances across the human subject samples versus the technical controls. Peptides with a signal-to-noise ratio of less than 2 were excluded from further analysis. The peptide relative abundances were log2 transformed and centered at the median. The abundance of endogenous peptides was quantified as a ratio to spiked-in synthetic peptides containing stable heavy isotopes. The “light/heavy” ratios were log2 transformed and shifted such that the median log2-ratio was zero. Normalization adjusted for differences in protein amounts among the samples.
Definition of study set
In silico genotyping
In silico genotyping underperformed as the unique sequence breakpoint fragment was only detected in 60% of the samples while 99% of the human population carries CHRFAM7A. This low detection threshold results in ambiguity for genotype calls and introduces a high genotyping error rate. The success of genotyping depends on i) the read depth of the unique breakpoint sequence and ii) the dynamic range of the read depth for exon 6 2bp deletion and wild type alleles. Neither of these were met from WGS data. Challenges in genotyping fusion genes within low copy repeats of the human genome was expected. As CNV effects are associated with dosage, we proceeded to use gene expression level as a surrogate for dosage to uncover CHRFAM7A biology in the human brain.
Data processing and analysis workflow
CHRFAM7A gene expression quantification
As CNV effect is frequently mediated by dosage (number of copies) and associated gene expression level we used normalized RNAseq data to quantify CHRFAM7A gene expression per sample as a predictor in the statistical model. CHRFAM7A gene expression level was extracted from the processed RNAseq data (Synapse).
Aβ and Aβ38 extraction from SRM Proteomics data
The normalized protein quantification data from ROSMAP was extracted from Synapse.
Statistical model
Gene expression values (FPKM) from ROSMAP data were regressed individually against CHRFAM7A expression, Aβ plus their interaction term adjusting for age, sex, APOE genotype and PMI. The interaction coefficients from the linear models were extracted and organized as an input to GSEPreranked24 using C2-CP human collection from msigDB (v7.1). Top normalized enrichment score signatures were examined and the top 20 leading-edge genes’ expression values were plotted in a heatmap. Heatmaps were plotted using variance-stability transformation values implemented in the DESeq225 R package.
Multiple testing correction was performed by the Benjamini-Hochberg approach which controls false discovery rate (FDR).26 Assignment of biological function of the top 100 genes was done by manual curation. Top 100 genes colour coded for Ca2+/small GTPase and cytoskeleton associated genes are depicted in the volcano plot. Quartiles of eight representative genes from the top 100 are depicted as quartiles in violin plots. Gene expression levels per quartile were statistically compared by an independent two-tailed T-test.
iPSC model
Isogenic lines
Isogenic line development was described previously.7 Briefly, we inserted CHRFAM7A coding sequence obtained from Addgene (pcDNA3.1-CHRFAM7A-mCherry, a gift from Henry Lester (Addgene plasmid # 62,635) into the AAVS1 safe harbor site of the constitutively expressed gene PPP1R12C on chr19 (position 19q13.42) of the 0 copy ancestral line (UB068) using TALEN mediated gene editing. The expression vector harbors puromycin resistance gene for selection and fluorescent signal (GFP) for visualization in downstream experiments.
Neuronal differentiation
Neuronal differentiation of iPSC towards Medial Ganglionic Progenitors (MGE) was carried out as described previously.17 Undifferentiated UB068 (null) and UB068_CHRFAM7A (CHRFAM7A KI) cells grown on irradiated mouse embryonic fibroblasts were detached with Dispase at day 0 and continued to float in T-25 flasks with “iPSC Culture Media” without bFGF. When iPSCs formed Embryoid Bodies (EB) at day 5, the medium was replaced with neural induction medium (NIM) [DMEM/F12+Glutamax, N2 (Life Technologies), NEAA (Gibco), Heparin (Stem Cell Technologies) and pen/strep] on day 4. EBs were attached in 6-well plates on day 7, and by day 10 neural rosettes were present, indicating the development of primitive neuroepithelia. Ventralization of primitive neuroepithelia was started on day 10 by adding Purmorphamine 1.5 μM. On day 16, neural tube-like rosettes were detached and transferred into T-25 Flasks in NIM with 2% B27 (Life Technologies) to form neurospheres. The cultures were fed every other day. On day 25, MGE progenitors were dissociated with Accutase (Stem Cell Technologies) and plated onto 6-well plates for further differentiation in neuronal media.
Ca2+ imaging
Image acquisition
Live cell imaging-based measurements of intracellular calcium ion concentration were performed using Fluo-4 fluorescent calcium ion indicator (Sigma, St. Louis MO). The indicator fluorescence was acquired at 2 s intervals before and for up to 30 min Ca2+ dynamic curves were recorded for UB068 and UB068_CHRFAM7A neurons.27,28
MatLab data processing and statistics
Imaging data were analyzed in MATLAB software (Mathsoft, Natick, MA). Individual active regions were automatically identified in the image sequences and fluorescence time courses extracted from them, using the MIN1PIPE package.29 MIN1PIPE is a computer vision algorithm for calcium indicator fluorescence signal extraction from video recordings of neural cells. It is designed to be essentially parameter-free in intended applications as long as the image scale is known. Version v3 alpha (https://github.com/JinghaoLu/MIN1PIPE/releases/tag/v3.0.0) was used with the default settings. The structural element size for the morphological opening operation was set according to the image scale (13 μm).27,28
Software versions and settings
MATLAB R2020b (Mathsoft, Natick MA) was used with VBA v1.9.2 (https://github.com/MBB-team/VBA-toolbox/releases/tag/v1.9.2) and CaBBI distributed as part of the same. Default parameter settings were used.
Descriptive statistics and group comparisons
Posterior mean values of parameters estimated from fluorescence traces were treated in the frequentist framework as parameter sets characterizing the corresponding individual regions. This approach, previously employed in the mentioned Bayesian single-cell study,30 emulates standard handling of direct experimental cell measurements. Summary statistics were compiled across the active regions from the same experimental group. The statistical significance of group differences was evaluated using the Kolmogorov–Smirnov test on the distributions of the given parameter among the active regions of the experimental groups being compared.
Hydrogels
Fibronectin-coated polyacrylamide hydrogels were prepared as previously described.31 The acrylamide concentration remained constant at 7.5% while the bisacrylamide concentrations were 0.03% (2 kPa) and 0.1% (5 kPa) for “soft” and “stiff” hydrogels, respectively.
Atomic force microscopy
Atomic force microscopy (AFM) was used to measure intracellular stiffness.32 iPSC-derived MGE progenitors (null and CHRFAM7A KI) were plated on soft (0.03%) and stiff (0.1%) stiffness polyacrylamide hydrogels for 1 h and were indented using a silicon nitride cantilever (Asylum, BL-AC40TS-C2; spring constant, 0.05–0.12 N/m) with a three-sided pyramidal AFM tip (8 nm in radius). The stiffness of each cell was measured using AFM in contact mode using an NX12 AFM system (Park Systems) mounted on a Nikon ECLIPSE Ti2 inverted microscope. To analyze the stiffness, the first 400 nm of horizontal tip deflection was fit with the Hertz model for a three-sided pyramid and a 35° face angle to determine intracellular stiffness. For each experimental condition, 3 force curves for each cell were acquired for a total of 10 cells. AFM experiments were independently repeated three times (obtaining a total of 90 force curves for each cell type per experimental condition). Using AFM analysis software XEI (Park Systems), the force curves were quantified and converted to Young's modulus (stiffness).
qPCR
Total RNA was isolated using Trizol (Invitrogen) according to manufacturer's protocol. cDNA was synthesized from 500 ng RNA using ImProm-II reverse transcriptase (Promega) and oligo (DT) (Promega) at 42 °C for 1 h. Quantitative gene expression was detected by standard RT-qPCR using specific primers (IDT) listed in Supplementary Table S1 and SYBR green master mix (Biotool) run on Bio-Rad CFX Connect cycler (Bio-Rad). Relative gene expression was quantified by ΔΔCT method normalized to GAPDH expression assayed with 3 technical replicates. Data are presented as the average of the triplicates ± standard error of the mean.
G-LISA
Activity of small GTPases (CDC42, Rac1, and RhoA) in the MGE progenitors derived from the null and direct iPSC lines was measured by G-LISA Activation Assay (Cytoskeleton Inc.) according to the manufacturer's protocol. The MGE progenitors were starved overnight, collected by centrifugation (300 × g, 4 min, RT), plated on 0.03% and/or 0.1% hydrogels, and incubated at 37 °C for 1 h. Afterward, the cells were lysed with an appropriate lysis buffer and centrifugated (10,000 × g, 1 min, 4 °C). Supernatants were immediately frozen and kept at −80 °C until the G-LISA Activity Assay. Protein concentration was measured by Precision Red™ Advanced Protein Assay (Cytoskeleton Inc.).
Actin live imaging
MGE progenitors generated from UB068 and UB068_CHRFAM7A lines and plated on 8-well chambers (30, 000 cells/well) were transfected with 0.5 μg of Lifeact-RFP according to Ma et al. protocol.33 Live imaging was performed 16 h after transfection using Leica DMi8 microscope (objective 63×). The cells were kept at 5% CO2 and 37 °C. The images were acquired every 4 s for 5 min. At least 10 cells/line were imaged in three independent experiments.
Shape index
Images were analyzed by Fiji ImageJ utilizing Trackmate plugin. Regions of interest (ROIs) of 50 × 50 pixels corresponding to an area of 6.63 μm2 were selected to calculate the shape index. The shape index describes the surface topology of the image and was used as a measure of the range of change in the edge of the growth cones.
Synaptic density by western blot
Synaptophysin, a presynaptic marker, and PSD95, a postsynaptic marker, were detected in a crude synaptosome fraction by immunoblotting with the specific antibodies. Using an established protocol,34 crude synaptosomes were isolated from neurons (D30 after plating of MGE) generated from UB068 and UB068_CHRFAM7A lines. Briefly, the cells were collected with accutase, and pelleted by centrifugation (300 × g. 5 min, room temperature). The pellet was resuspended in TE buffer (10 mM TRIS, 1 mM EDTA, pH, 7.5) with protease inhibitors, sonicated, and centrifuged (800 × g, 5 min, 4 °C). After centrifugation, total lysate, the supernatant, was further centrifuged (9200 × g, 20 min, 4 °C). The pellet, a crude synaptosome fraction, was resuspended in an appropriate volume of TE buffer with protease inhibitors. 10 μg of crude synaptosomes were separated on 4–20% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (Bio-Rad), transferred onto polyvinylidene difluoride membrane (Bio-Rad), and incubated overnight at 4 °C with primary antibodies (listed in Supplementary Table S2). Specific immunoreactive bands were detected using ChemiDoc XRS + Imaging Systems (Bio-Rad). Densitometry analysis was performed using Image Lab 6.0.1 Software. β-actin was used as a loading control.
Immunocytochemistry and confocal microscopy
MGE progenitors and/or neurons plated on Matrigel-coated glass coverslips or 8-well glass chambers (Thermo Fisher) were fixed with 4% paraformaldehyde (Mallinckrodt Baker) for 15 min, permeabilized with 0.1% Triton ×100 (Mallinckrodt Baker) for 10 min and blocked with blocking buffer (5% BSA in PBS) for 1 h at room temperature (RT). Cells were incubated overnight at 4 °C with primary antibodies and for 1 h at RT with secondary antibodies (listed in Supplementary Table S2). Both primary and secondary antibodies were diluted in a blocking buffer. Actin was visualized with ActinGreen 488 ReadyProbes (Thermo Fisher Scientific). Slides/coverslips were incubated with NucBlue (DAPI) for 5 min at RT and mounted with a ProLong® Gold Antifade reagent (Life Technologies). Confocal images were captured by using Leica TCS SP8 confocal microscope system (63× and/or 20× objective). Images were acquired using LAS X software.
Image analysis
Dendritic spine quantification
Neurons (D30 after plating MGE progenitors) derived from the null and CHRFAM7A line were double stained with bIII-tubulin and Phalloidin and confocal images were taken as described above. Neuronal processes were classified and quantified in segments of 60 μm of dendrites in at least 10 neurons/line in three independent experiments. Processes were classified based on their morphology as filopodia (>0.2 μm) or dendritic spines (long thin, thin, stubby, mushroom, and branched) as indicated in Fig. 5. The percentage of the population over the total number of processes was then calculated.
Fig. 5.
CHRFAM7A effect on structure of the synapse. a. Schematic depicting dendritogenesis timeline and formation of synapsis (created with BioRender.com). b. Representative Synaptophysin and MAP2 confocal images demonstrating a difference in dendritic spinogenesis in neurons (D 30 after plating MGE progenitors) derived from null and CHRFAM7A KI lines. Quantification of number (c.) and area (d.) of synapsis on neurons derived from both lines. Statistical comparison was performed using the two-sample Kolmogorov–Smirnov test. Representative immunoblot (e.) and densitometric analysis (f.) of presynaptic (synaptophysin) and postsynaptic (PSD95) markers in a crude synaptosome fraction isolated from the null and CHRFAM7A KI iPSC derived neurons (n = 3 independent experiments, independent two-tailed T-test). g. Representative dendritic spine images of null and CHRFAM7A KI iPSC derived neurons. The neurons are stained with phalloidin followed by confocal microscopy and ImageJ processing. h. Dendritic spine length difference between the null UB068 (grey) and CHRFAM7A (orange). (N = 50 from 3 independent experiments, independent two-tailed T-test). i. Schematic depicting dendritic spine types and colour code for quantification (created with BioRender.com). j Quantification of dendritic spines in neurons derived from two iPSC lines. 20 images from 3 independent experiments. Statistical analysis was performed by non-parametric two-tailed Mann–Whitney test.
Synaptic density
Neurons (D30 after plating MGE progenitors) were double stained with Synaptophysin and MAP2 antibodies and confocal images were taken as described above. SynQuant plugin35 (Fiji ImageJ) was employed for synapse characterization and analysis. Number of synapses, their area, and intensity as well as dendritic length and intensity were measured in at least five images/cell line obtained in four independent experiments.
Statistics
Exploratory gene expression analysis was performed on an existing omics dataset (ROSMAP), thus the sample size refers to sample availability with the specified selection criteria: samples required both RNAseq and SRM data to be included in the analysis, resulting in a sample size of 587. A multivariable linear regression model was fitted assuming the response expression (FPKM) values were normally distributed. This model can be interpreted as a GLM assuming normal distribution with identity link function. Kruskal–Wallis test (non-parametric ANOVA) was performed to compare the difference of means between cytoskeleton and Ca2+ signalling associated genes in CHRFAM7A gene expression quartiles.
Single cell data distribution was compared by Kolmogorov–Smirnov statistics. Experimental triplicates were compared by an independent two-tailed T-test. Multiple testing correction was performed by Bonferroni. Dendritic spine morphological distribution was compared by non-parametric two-tailed Mann Whitney test. In box-and-whisker plots, outlying points were plotted for values that were greater than the distance of 1.5∗IQR starting from upper hinge (3rd quartile).
Role of funders
Funders did not have any role in study design, data collection, data analyses, interpretation, or writing of report.
Results
CHRFAM7A associated transcriptomic signature: approach
The ROSMAP post mortem brain dorsolateral frontal cortex (DLFC) dataset includes RNAseq data, amyloid-beta (Aβ) quantification by selective reaction monitoring (SRM) proteomics, and the spectrum of cognitive phenotypes ranging from normal controls (NCI) to Mild Cognitive Impairment (MCI) to Alzheimer's Disease (AD) and Mixed pathology (AD+). Overlap between DLFC RNAseq and SRM proteomics resulted in a sample size of 587 (Fig. 1a and b, Supplementary Fig. S1a). Aβ quantification through SRM proteomics eliminated phenotypic binning (with its inherent misclassification bias). The first assumption is that SRM Aβ level serves as quantified AD biology.
Fig. 1.
RNAseq analysis predicts CHRFAM7A effect on the synapse as a structure. a. Relevant sample and data distribution in the ROSMAP study. b. Demographic characterization of the cohort with RNAseq and SRM proteomics data. c. Kernel distribution of Aβ peptide quantified by SRM in disease groups in the ROSMAP cohort demonstrates disease relevance (created with BioRender.com). d. Aβ peptide and CHRFAM7A gene expression associated pathways (red, upregulated, blue downregulated) using a multivariable regression model. Aβ associated pathways align with AD related neurodegeneration including increased ECM and decreased metabolism. CHRFAM7A top 3 pathways are related to the synapse without neurotransmitter specificity suggesting a functional role in the synapse as a structure. e. Heatmap depicting the leading edges of the top pathway positively associated with CHRFAM7A expression. f. Pie chart depicting the frequency of cytoskeleton (48%) and Ca2+ signalling (19%) associated genes in top 100 positively correlated genes with CHRFAM7A. g. Pie chart depicting the distribution of structural gene categories. h. Violin plots of cytoskeleton (top panel) and Ca2+ signalling associated genes (bottom panel) in CHRFAM7A gene expression quartiles. To compare the difference of means between quartiles, Kruskal–Wallis test (non-parametric ANOVA) was performed.
Gene expression levels typically demonstrate a high correlation with the copy number of structural variants due to gene dosage.36,37 The second assumption is that CHRFAM7A is a dosage sensitive gene. With these two assumptions, we performed co-linear regression using Aβ proteomics and CHRFAM7A gene expression to illuminate CHRFAM7A transcriptomic signature after controlling for Aβ load. The model incorporated age, sex, postmortem interval (PMI), and APOE genotype as covariates (Supplementary Fig. S1a and b).
CHRFAM7A is transcribed and translated in the human brain
CHRFAM7A protein expression in human temporal lobes is confirmed by Western blot (Supplementary Fig. S2). CHRFAM7A and CHRNA7 gene expression (N = 636) in the four binned phenotypes were similar with no significant differences (Supplementary Fig. S1c and d), respectively. Weak correlation (ρ = 0.51) was detected between CHRFAM7A and CHRNA7 gene expression (Supplementary Fig. S1e), dampening the possibility of CHRNA7 shine-through in the RNAseq data. CHRFAM7A to CHRNA7 ratio was used to estimate the number of CHRFAM7A null samples. (Supplementary Fig. S1e). We identified 15 null subjects in 636 samples consistent with the expected 0.7% of nulls in the human population.7 Thus, while CHRFAM7A is expressed in the human brain, it is not related to the disease phenotype and it has no correlation with CHRNA7 gene expression.
Quantification of AD biology using Aβ protein level
SRM proteomics dataset included two amyloid relevant peptides, Aβ and Aβ38. These two peptides demonstrated distinct distribution patterns suggesting that peptide Aβ captures the disease associated Aβ burden (Fig. 1c). There was no correlation found between Aβ (SRM proteomics) and CHRFAM7A (RNAseq), indicating that the two variables are independent (Supplementary Fig. S1b).
CHRFAM7A is associated with neuronal system gain of function in the human brain
To illuminate CHRFAM7A gene dosage associated RNAseq signature we built a statistical model incorporating normalized CHRFAM7A gene expression (continuous variable), Aβ peptide level (binned based on the bimodal distribution) and APOE genotype, age, sex, and PMI as covariates. In this model genes correlating with Aβ highlight Aβ associated effects while CHRFAM7A associated genes uncover CHRFAM7A dependent biology. Pathway analysis of Aβ associated biology demonstrated that the expression levels of genes involved in the extracellular matrix (ECM) and inflammation were increased, while the expression levels of genes involved in glucose metabolism and protein transport were decreased (Fig. 1d). These findings support known AD biology in humans: upregulation of ECM associated pathways is consistent with neuronal loss and greater ECM observed post mortem38; and neuroinflammation is associated with the presence of Aβ plaques.37 Decreases in metabolism correlate with metabolically inactive tissue in AD patients detected by FDG-PET scan in the clinical setting, which is commonly used as a diagnostic tool for AD.39 Interestingly, these transcriptomic hits (of note, after removing the CHRFAM7A effect with the colinear model) are similar to the AD pathology and cognition related proteomic signature of the same cohort.38 These observed Aβ effects indicate that our statistical model is successful in identifying biologically relevant pathways.
We detected a robust signal in C2 canonical pathways from msigDB pathways associated with CHRFAM7A dosage (Fig. 1d). Top three pathways indicate an effect on the neuronal system (Fig. 1e, Supplementary Fig. S1f and g) in general and the chemical synapse in particular, irrespective of neurotransmitters (4 top pathways) followed by NMDA receptors, voltage gated potassium channels and synaptic cell adhesion molecules, neurolexins and neuroligins (Fig. 1d). To understand which genes contribute to the detected pathways with the largest effect size, curation of the top 100 positively correlated genes revealed that 19% (Fig. 1f) are linked to Ca2+ signalling (Supplementary Fig. S1h, depicted in orange on the volcano plot) with three of those genes involving small GTPases RhoA, Rac1, and CDC42. Of these 100 genes, 60% are intrinsic to cellular structure including the cytoskeleton and cell membrane (Supplementary Fig. S1h, depicted in blue on the volcano plot). Furthermore, the majority of structural genes are linked to the cytoskeleton, 65% of which were actin cytoskeleton associated (Fig. 1g). Genes associated with Ca2+ signalling (Fig. 1h bottom panel) and the actin cytoskeleton (Fig. 1h top panel) demonstrated highly significant correlation with CHRFAM7A gene expression quartiles (Fig. 1h, p < 0.001, FDR corrected). These results suggest that CHRFAM7A is related to the synapse through Ca2+ signalling, small GTPases, and the actin cytoskeleton.
The emerging cytoskeletal and membrane biology suggests a structural gain of function through Ca2+ signalling and small-GTPase signal transduction. To mitigate limitations inherent to post mortem human brain RNAseq data based studies we incorporated known epidemiological (age, sex, APOE4) and tissue quality (PMI) covariates. As additional unmeasured and uncontrolled confounding factors are probable we considered the omics results as hypotheses and to increase robustness of the findings we validated the emerging mechanisms experimentally. We proceeded to test these hypotheses in an isogenic iPSC model of null (UB068) and CHRFAM7A knock in (KI) (UB068_CHRFAM7A) lines from AD subjects.7,10,16,17
CHRFAM7A modifies Ca2+ dynamics
CHRFAM7A incorporation into the α7 nAChR pentamer results in a hypomorphic receptor in iPSC derived MGE progenitors with decreased channel open probability.7,40 Intriguingly, fluorescent single cell Ca2+ imaging demonstrated unique Ca2+ peak morphology in the presence of CHRFAM7A (Fig. 2a and b). Distribution of Ca2+ dynamic parameters indicated that amplitude and peak width differ between the null and CHRFAM7A KI lines. The representative montages from the Fluo-4 Ca2+ videos demonstrated the prolonged active state in CHRFAM7A KI cells: in null cells Ca2+ activity persisted in three consecutive frames corresponding to 6 s, and in CHRFAM7A KI - 5 consecutive frames corresponding to 10 s (Fig. 2c). Cumulative density curves of trace characteristics of 50–100 cells demonstrated that CHRFAM7A is associated with lower amplitude and increased width of the Ca2+ tracings (Fig. 2d). The neurophysiology and Ca2+ signal change may suggest a mechanism where the hypomorphic receptor chimera has decreased channel open probability shifting the time ratio spent in CICR to IICR (Fig. 2e and f). The proposed CICR to IICR shift is consistent with the decreased amplitude (mitigated CICR) and increased width (enhanced IICR) in the presence of CHRFAM7A (Fig. 2f). The function of Ca2+ as a signalling molecule is closely linked to its dynamics and propagation within the cell, these changes imply biological consequences.41, 42, 43 Taken together the upregulated genes and pathways from the omics analysis and the distinctly different Ca2+ dynamics in the iPSC model, we hypothesized that CHRFAM7A affects small GTPase activity and consequently actin remodeling.
Fig. 2.
Ca2+dynamics. a. Representative fluorescent Ca2+ time-lapse video recordings of null and CHRFAM7A KI isogenic iPSC derived MGE progenitors. b. Representative fluorescent Ca2+ traces from cells marked in videos in a. c. Representative time-lapse video frames demonstrating the Ca2+ signal change over time in null and CHRFAM7A MGE progenitors (white box active Ca2+ signal time interval in Null, red box active Ca2+ signal time interval in CHRFAM7A KI MGE progenitors). d. Cumulative density curves of Ca2+ peak characteristics in null and CHRFAM7A KI isogenic iPSC derived MGE progenitors. Operator independent analysis of single cell Ca2+ tracings was performed in MatLab. (3 independent experiments, 50–100 peaks per cell line). Statistical comparison was performed using the two-sample Kolmogorov–Smirnov test. Multiple testing correction was performed by the Bonferroni method. (A amplitude, FWHM form width at half maximum amplitude). e. Proposed mechanism of CHRFAM7A modified Ca2+ Flux (created with BioRender.com). f. Ca2+ dynamics curve morphology indicates a shift in time occupancy of CICR to IICR in the presence of CHRFAM7A.
CHRFAM7A is associated with dynamic actin cytoskeleton
Actin cytoskeleton drives fundamental neuronal morphological properties including neurite outgrowth, growth cone, dendritic spine morphology, and axon guidance.37,44,45 Null and CHRFAM7A KI isogenic iPSC derived MGE progenitors demonstrated distinctly different actin cytoskeleton related morphological features (Fig. 3a). Null MGE progenitors harbored filopodia membrane specialization in contrast CHRFAM7A switched the phenotype to lamellipodia. Total neurite branch length was similar between the lines (Fig. 3b).
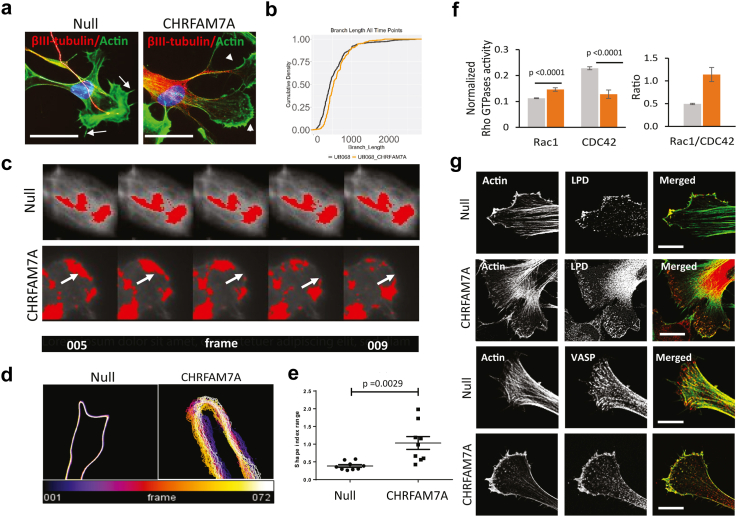
Fig. 3.
Actin cytoskeleton gain of function. a. Representative confocal images of iPSC derived MGE progenitors 48 h after plating on Matrigel. (Red–βIII-tubulin; green—Actin, phalloidin). Filopodia are indicated by arrows and lamellipodia by arrowheads. Scale bar, 10 μm. b. Cumulative density curves of branch length in MGE progenitors (48 h after plating) derived from the null (grey) and CHRFAM7A KI (orange) lines. Statistical comparison was performed using the two-sample Kolmogorov–Smirnov test, not significant. c. Actin lattice change over time depicted in consecutive frames of live actin video recording in MGE progenitors derived from the null and CHRFAM7A KI lines (arrows: area of interest over time). d. Live actin imaging of MGE progenitors: representative image of growth cone surface undulations depicted by overlaying colour coded frames (Supplementary Video 1–6). e. Cell surface topology dynamic changes over time are quantified as variance in shape index in null and CHRFAM7A KI isogenic iPSC derived MGE progenitors (n = 3; independent two-tailed-T-test, the horizontal lines represent median and IQR). f. Activation of small GTPases in response to Matrigel in null (grey) and CHRFAM7A KI isogenic (orange) MGE progenitors detected by G-LISA and Rac1/CDC42 ratio (n = 3; independent two-tailed T-test. Multiple testing correction was performed by Bonferroni). g. Representative confocal images demonstrating differential staining of lamellipodia (by lamellipodin, LPD) and filopodia (by VASP) in MGE progenitors derived from the two lines.
Live-actin imaging of null and CHRFAM7A KI MGE progenitors revealed a strikingly more dynamic actin cytoskeleton potentially leading to dynamic cell shape at the cell body, neurite outgrowth, and growth cone (Supplementary videos). The actin signal demonstrated more dynamic actin polymerization and depolymerization in CHRFAM7A KI MGE progenitors (Fig. 3c). Neurite outgrowth patterns were distinctly different between the null and CHRFAM7A KI progenitors: null cells displayed filopodia tips with low directionality, while the CHRFAM7A KI tips showed dynamic actin architecture leading to lamellipodia with high directionality and retrograde flow (Supplementary videos). Surface topology dynamics in the CHRFAM7A KI growth cone demonstrated a highly dynamic cell membrane compared to null (Fig. 3d and e).46 Dynamic membrane protrusions of the CHRFAM7A KI MGE progenitors in the live videos appeared to be supported by directionally unaligned actin filaments.47 As unique nanoscale architecture of a flexible, signal-dependent actin structure has been associated with Rac1 activation and lamellipodia membrane specialization,47 we performed small GTPase activation assays. The dynamic actin cytoskeleton and the phenotype switch from filopodia to lamellipodia in CHRFAM7A KI MGE progenitors were associated with an increased Rac1 to CDC42 activation ratio (Fig. 3f). Rac1 activation is thought to increase lamellipodin (LPD) localization to the cell periphery in lamellipodia.48 Immunocytochemistry (ICC) showed colocalization of LPD with the membrane-continuous actin signal at the leading edge in CHRFAM7A KI MGE progenitors, in contrast to the small LPD clusters localizing to the filopodia in null cells (Fig. 3g.). Ena/VASP proteins, actin-binding proteins that bind to LPD during migration and localize to focal adhesions at the leading edge,49 demonstrated a similar staining pattern: in CHRFAM7A KI MGE progenitors, VASP was continuous at the leading edge, while in null cells VASP was distributed as small clusters, localizing to the filopodia tips (Fig. 3g).32 Our data suggests that CHRFAM7A can alter Ca2+ dynamics, activate Rac1, and result in a dynamic actin cytoskeleton, leading to a switch in cell shape to lamellipodia.
CHRFAM7A-dependent adaptation to mechanical properties of the extracellular environment
The actin cytoskeleton has been implicated in adaptation to tissue stiffness.50,51 As the mechanical properties of the human and rodent brain are distinctly different, the human brain being stiffer, (Fig. 4a),52, 53, 54 we explored CHRFAM7A function in MGE progenitors in a human relevant context using polyacrylamide hydrogels corresponding to Young's (or elastic) modulus 2 kPa (0.03% bis-acrylamide concentration) and 5 kPa (0.1%), respectively. These stiffnesses represent the shift between the rodent and the human brains (Fig. 4a).52,55 In both cell lines, growth cone (GC) morphology distinctly adapted to tissue stiffness; in high stiffness null GCs developed filopodia, while CHRFAM7A GCs developed lamellipodia (Fig. 4b). Small GTPases activation on 2 kPa hydrogel (baseline) showed that in the presence of CHRFAM7A Rac1/CDC42 ratio favors Rac1, while in null CDC42 is higher. (Fig. 4c). Atomic Force Microscopy (AFM) measuring intracellular elastic modulus demonstrated that MGE progenitor cells derived from the CHRFAM7A KI line respond more efficiently to the stiffness of the environment (hydrogel stiffness) compared to null cells (Fig. 4d). The response is Rac1 depended as the treatment with EHT1864, a specific Rac1 inhibitor, decreased intracellular elastic modulus in both cell lines (Fig. 4d). The increased intracellular stiffness in stiffer environment suggests superior adaptation of the CHRFAM7A KI line.
Fig. 4.
CHRFAM7A-dependent adaptation to mechanical properties of the extracellular environment. a. Schematic depicting Young's modulus of rodent and human brain. Hydrogel concentrations corresponding to the Young's modulus in kPa (gold) is contrasted to the standard procedure of Matrigel coating (blue). b. Changes in growth cone morphology in response to hydrogel stiffness: representative images of MGE progenitors derived from the null and CHRFAM7A KI lines. c. G-LISA demonstrating activation of small GTPases (top panel) on 2 kPa hydrogel; Rac1/CDC42 ratio (bottom panel) in MGE progenitors derived from the null (grey) and CHRFAM7A KI (orange) lines (n = 3; independent two-tailed T-test. Multiple testing correction was performed by Bonferroni). d. Atomic Force microscopy (AFM): intracellular stiffness of MGE progenitors derived from null (grey) and CHRFAM7A KI (orange) iPSC lines and seeded for 2 h on 0.03 and 0.1 stiffness hydrogels in the absence or presence of Rac1 inhibitor, EHT1864. Total of 50 cells were analyzed for each condition from 5 independent experiments. Averages were compared by an independent two-tailed T-test. Multiple testing correction was performed by Bonferroni. e. Representative images of β-III-tubulin immunofluorescent microscopy of null and CHRFAM7A KI isogenic iPSC derived MGE progenitors 48 h after plating on 0.03% and 0.1% hydrogels. f. Arborization of null (grey) and CHRFAM7A KI (orange) isogenic iPSC derived MGE progenitors on 0.03% and 0.1% hydrogels (left panel) (n = 3 independent experiments, 10 images of total 50 MGE progenitors counted; independent two-tailed T-test). g. Percentage of unipolar (Uni), Bipolar (Bi) and Multipolar (Multi) cells in the total population of MGE progenitors plated on the soft and stiff hydrogel (right panel) h. G-LISA of small GTPases CDC42, Rac1 and RhoA in null (grey) and CHRFAM7A KI (orange) isogenic iPSC derived MGE progenitors on 0.03% and 0.1% hydrogels (n = 3 independent experiments; independent two-tailed T-test). i. Representative images showing differences in axon guidance of MGE progenitors plated on 0.1% stiffness hydrogel. j. MMP2 and MMP9 gene expression levels of null (grey) and CHRFAM7A KI (orange) MGE progenitors upon exposure for 1 h to fibronectin coated stiff hydrogel (0.1%) in the absence or presence of Rac1 inhibitor, EHT1864. PCR data from 3 to 5 independent experiments are depicted as bar graphs with SD. Statistical analysis was performed by an independent two-tailed T-test.
Seeding the cells for 48 h on the soft (2 kPa) and stiff (5 kPa) hydrogels resulted in distinct polarization patterns of MGE progenitors derived from the null and CHRFAM7A KI lines (Fig. 4e and f). In the stiffer environment, null cells became predominantly multipolar (>2 branches per cell), while CHRFAM7A_KI cells became mostly bipolar (2 branches with opposite direction) (Fig. 4e and f and g). In mouse models, the switch in the polarization phenotype from bipolar to multipolar has been associated with a change in small Rho GTPase activation, specifically with a decrease in Cdc42 activation.53 The bipolar polarization has been accompanied with an increased Cdc42 activity.56 In addition, Rac1 has been shown to promote the multipolar–bipolar transition.57 Meanwhile, RhoA conveys retraction of the neurites and is regulated in the opposite direction of Rac1 or Cdc42 activity.58,59 In null MGE progenitors, the CDC42 activity was higher than RhoA in the soft environment (Fig. 4h grey), promoting bipolar polarization of the cells in contrast to multipolar morphology in the stiff environment (Fig. 4e–g). In contrast, CHRFAM7A KI cells adapted to higher stiffness by activating both CDC42 and Rac1 and with a limited RhoA activation (Fig. 4h orange), thus promoting bipolar polarization in the stiff environment (Fig. 4e–g). While both cell lines responded to stiffness of the environment with the increase in Rac1 activation, CHRFAM7A KI demonstrated a higher Rac1 activity baseline and response (Fig. 4h). These data are consistent with the notion that polarization of the neuronal lineage is dependent on CDC4256,60 and Rac1,61,62 while RhoA has been linked to the retraction of neurites.63,64
Rac1 has been implicated in axon guidance through an effect on the actin cytoskeleton.65 Directionality of neurite outgrowth of null and CHRFAM7A KI progenitors demonstrated striking differences in the 5 kPa environment. Null MGE progenitors developed pronounced and multidirectional filopodia GCs, while CHRFAM7A KI counterparts had a dense actin tip, indicating the ability to enter the stiff environment (Fig. 4i). Ability of the CHRFAM7A KI MGE progenitors to enter the stiffer environment is associated with increased MMP2 and MMP9 expression (Fig. 4j) suggesting increased ECM degradation as a potential mechanism.66 Inhibition of Rac1 led to a decrease in both MMP2 and MMP9 levels only in the CHRFAM7A KI line (Fig. 4j).
The extracellular matrix (ECM) functions as a scaffold for structural support and forms a reservoir of signalling molecules for the synapse. Matrix metalloproteinases may contribute to the dynamic remodeling of structural and functional plasticity by cleaving ECM components and cell adhesion molecules. In the CNS, MMP9 signalling is required for long-term memory formation that requires synaptic remodeling and MMP2 has been implicated in neurosphere migration.66 Rac1 plays a role in cell invasion across ECM barriers through activation of matrix metalloproteinases (MMP)67,68 and it has been extensively studied in cancer.67,69, 70, 71 MMP2 knockout mice demonstrated increased anxiety underscoring its role in the CNS.66 Our data suggest that both MMP2 and MMP9 are upregulated in stiff environment and both are regulated by Rac1 (Fig. 4j).
CHRFAM7A strengthens the structural foundation of the synapse
Human brain OMICS suggested a synaptic gain of function irrespective of the neurotransmitter (Fig. 1d and e). As actin is known to play a fundamental role in synapse structure and dynamics, we proceeded to study the structure of the synapse.72 MGE progenitors derived from null and CHRFAM7A KI isogenic iPSC were further differentiated into neurons (Fig. 5a). When double stained with dendritic marker MAP2 and presynaptic marker Synaptophysin (Syn), the neurons demonstrated visually different synaptic size and distribution (Fig. 5b). While synaptic density (puncta per dendrite unit) was comparable between null and CHRFAM7A derived neurons (Fig. 5c), the surface area of puncta corresponding to synaptic structure size was higher in CHRFAM7A KI derived neurons compared to null (Fig. 5b and d). This quantitative difference was confirmed by Western blot for both the presynaptic synaptophysin and postsynaptic PSD95 protein expression (Fig. 5e and f). The larger surface area of the synapse as a structure may reflect clustering facilitated by actin cytoskeleton reorganization into lamellipodia in the CHRFAM7A KI neurons.
To test this hypothesis, we characterized dendritic spine morphology (Fig. 5g): dendritic spine length was significantly longer in CHRFAM7A KI neurons compared to null, (Fig. 5h). In addition, there was a distribution shift in dendritic spine type between the neurons; null neurons had a higher percentage of filopodia, while CHRFAM7A KI neurons had a higher percentage of long-thin spines (Fig. 5i and j). Dendritic spine types on the yellow to red scale are various development stages of the “lamellipodia-type” morphology (Fig. 5i and j). In CHRFAM7A neurons, Rac1 activation and actin cytoskeleton reorganization shifts dendritic spine differentiation from filopodia (grey) towards spines with increased head area to stem diameter ratio (orange) which increases synapse clustering (Fig. 5i and j). The clustering, rather than the size of the individual synapse is accountable for the synaptic area and synaptosome increase in the CHRFAM7A KI line (Fig. 5d–f).
Discussion
While the uniquely human CHRFAM7A fusion gene has been linked to neurocognitive disorders, its function in the human brain was poorly understood. Multiomics analysis complemented with human isogenic iPSC model illuminates its impact on the neuronal lineage and deciphers the molecular mechanism. CHRFAM7A utilizes the α7nAChR to modify intracellular Ca2+ dynamics and activate the small GTPase Rac1. This activation leads to a reorganization and energization of the actin cytoskeleton, resulting in a dynamic cell with reinforced actin structure. The actin phenotype affects all neuronal substructures, including the cell body, growth cone, and dendritic spine. The reinforced membrane in CHRFAM7A leads to post-developmental adaptation to changes in stiffness associated with biological and pathological processes. The reinforced neuronal matrix provides the structural basis for more precise connections and enlarged synaptic surface area (“high quality wellcro”) compared to null cells. Whether and how CHRFAM7A affects human brain function is an intriguing question that will require human studies in the areas of memory formation and erasure, cognitive reserve, and neuronal plasticity.
Considering that the presence or absence of CHRFAM7A does not have an overt developmental effect suggests that the strong selective pressure is unlikely to be driven by brain biology. We have previously shown that CHRFAM7A modifies the innate immune response through NFKB in microglia.16 The effect of CHRFAM7A in the innate immune system suggests that it releases the cholinergic anti-inflammatory response and is proinflammatory.73,74 We hypothesize that the strong selective pressure was based on the immune mechanism, and the brain effect is a pleiotropy. One possible interpretation of the data is that CHRFAM7A facilitates neuronal adaptation to changes in the brain environment (inflammation, Amyloid deposition, myelination, aging, glioma, stroke) which may lead to benefit or risk based on the disease context.
Contributors
K.S. conceptualized and designed the study, both the laboratory experiments and OMICS aspects, led the wet lab experimental design, data analysis and data interpretation, and wrote the first draft of the manuscript. I.I. performed the neuronal culture experiments and imaging, contributed to the wet lab experimental design, data analysis and interpretation, and worked extensively on the manuscript. N.R. contributed to live actin imaging and analysis, dendritic spine morphology characterization, and prepared the corresponding figures. R.P.D. contributed to G-LISA experiments and analysis, statistical analysis and figures preparation. Em.N. contributed to OMICS data analysis and figures. E.C.G. performed OMICS data analysis and this section figure. M.H. performed the Min1Pipe analysis of calcium imaging and contributed to this section figure preparation. I.M. conducted calcium imaging on iPSC-derived MGE progenitors and contributed to its analysis. S.P. performed the morphological analyses in ImageJ and statistics. Er.N. performed AFM experiments and data analysis. Y.H. contributed to the mechanosensation experiments and the corresponding figures preparation. B.F. contributed to interpretation of dendritic spine morphology data and critically read the manuscript. D.A.B. contributed to the ROSMAP dataset analysis. W.A.H. contributed to the design of calcium imaging experiments and analysis. A.P. contributed to calcium imaging analysis and critically read the manuscript Y.B. designed the AFM experiments, wrote and edited the corresponding section of the manuscript, contributed to data interpretation for mechanosensation and actin cytoskeleton mechanism, and critically read the manuscript. J.W. designed and managed the OMICS data analysis, wrote and edited this part of the manuscript, and critically read the manuscript.
K.S., Y.B., and J.W. had accessed and verified the underlying data.
All authors read and approved the final version of the manuscript.
Data sharing statement
The data that support the findings of this study are available from the corresponding author (K.S.), upon reasonable request.
Declaration of interests
The authors declare no conflict of interest.
Acknowledgements
We would like to thank Robin Schwartz-Byrne for her assistance in editing the manuscript.
Footnotes
Supplementary data related to this article can be found at https://doi.org/10.1016/j.ebiom.2023.104725.
Appendix A. Supplementary data
References
- 1.Bitar M., Kuiper S., O'Brien E.A., Barry G. Genes with human-specific features are primarily involved withbrain, immune and metabolic evolution. BMC Bioinformatics. 2019;20(9):406. doi: 10.1186/s12859-019-2886-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Florio M., Namba T., Pääbo S., Hiller M., Huttner W.B. A single splice site mutation in human-specific ARHGAP11B causes basal progenitor amplification. Sci Adv. 2016;2(12) doi: 10.1126/sciadv.1601941. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Sinkus M.L., Graw S., Freedman R., Ross R.G., Lester H.A., Leonard S. The human CHRNA7 and CHRFAM7A genes: a review of the genetics, regulation, and function. Neuropharmacology. 2015;96(Pt B):274–288. doi: 10.1016/j.neuropharm.2015.02.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Swaminathan S., Huentelman M.J., Corneveaux J.J., et al. Analysis of copy number variation in Alzheimer's disease in a cohort of clinically characterized and neuropathologically verified individuals. PLoS One. 2012;7(12) doi: 10.1371/journal.pone.0050640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Lew A.R., Kellermayer T.R., Sule B.P., Szigeti K. Copy number variations in adult-onset neuropsychiatric diseases. Curr Genomics. 2018;19(6):420–430. doi: 10.2174/1389202919666180330153842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Zhang Z.G., Li Y., Ng C.T., Song Y.Q. Inflammation in Alzheimer's disease and molecular genetics: recent update. Arch Immunol Ther Exp (Warsz) 2015;63(5):333–344. doi: 10.1007/s00005-015-0351-0. [DOI] [PubMed] [Google Scholar]
- 7.Szigeti K., Ihnatovych I., Birkaya B., et al. CHRFAM7A: a human specific fusion gene, accounts for the translational gap for cholinergic strategies in Alzheimer's disease. eBioMedicine. 2020;59 doi: 10.1016/j.ebiom.2020.102892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Lasala M., Corradi J., Bruzzone A., Esandi M.D.C., Bouzat C. A human-specific, truncated alpha7 nicotinic receptor subunit assembles with full-length alpha7 and forms functional receptors with different stoichiometries. J Biol Chem. 2018;293(27):10707–10717. doi: 10.1074/jbc.RA117.001698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wang Y., Xiao C., Indersmitten T., Freedman R., Leonard S., Lester H.A. The duplicated alpha7 subunits assemble and form functional nicotinic receptors with the full-length alpha7. J Biol Chem. 2014;289(38):26451–26463. doi: 10.1074/jbc.M114.582858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ihnatovych I., Lew A., Lazar E., Sheng A., Kellermayer T., Szigeti K. Timing of wnt inhibition modulates directed differentiation of medial ganglionic eminence progenitors from human pluripotent stem cells. Stem Cells Int. 2018;2018 doi: 10.1155/2018/3983090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.de Lucas-Cerrillo A.M., Maldifassi M.C., Arnalich F., et al. Function of partially duplicated human alpha77 nicotinic receptor subunit CHRFAM7A gene: potential implications for the cholinergic anti-inflammatory response. J Biol Chem. 2011;286(1):594–606. doi: 10.1074/jbc.M110.180067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Araud T., Graw S., Berger R., et al. The chimeric gene CHRFAM7A, a partial duplication of the CHRNA7 gene, is a dominant negative regulator of alpha7∗nAChR function. Biochem Pharmacol. 2011;82(8):904–914. doi: 10.1016/j.bcp.2011.06.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Martín-Sánchez C., Alés E., Balseiro-Gómez S., et al. The human-specific duplicated α7 gene inhibits the ancestral α7, negatively regulating nicotinic acetylcholine receptor-mediated transmitter release. J Biol Chem. 2021;296 doi: 10.1016/j.jbc.2021.100341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.King J.R., Kabbani N. Alpha 7 nicotinic receptors attenuate neurite development through calcium activation of calpain at the growth cone. PLoS One. 2018;13(5) doi: 10.1371/journal.pone.0197247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Bennett D.A., Buchman A.S., Boyle P.A., Barnes L.L., Wilson R.S., Schneider J.A. Religious orders study and Rush memory and aging project. J Alzheimers Dis. 2018;64(s1):S161–S189. doi: 10.3233/JAD-179939. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Ihnatovych I., Birkaya B., Notari E., Szigeti K. iPSC-derived microglia for modeling human-specific DAMP and PAMP responses in the context of Alzheimer's disease. Int J Mol Sci. 2020;21(24):9668. doi: 10.3390/ijms21249668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Ihnatovych I., Nayak T.K., Ouf A., et al. iPSC model of CHRFAM7A effect on alpha7 nicotinic acetylcholine receptor function in the human context. Transl Psychiatry. 2019;9(1):59. doi: 10.1038/s41398-019-0375-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Szigeti K., Kellermayer B., Lentini J.M., et al. Ordered subset analysis of copy number variation association with age at onset of Alzheimer's disease. J Alzheimers Dis. 2014;41(4):1063–1071. doi: 10.3233/JAD-132693. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.De Jager P.L., Ma Y., McCabe C., et al. A multi-omic atlas of the human frontal cortex for aging and Alzheimer's disease research. Sci Data. 2018;5 doi: 10.1038/sdata.2018.142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Bennett D.A., Schneider J.A., Aggarwal N.T., et al. Decision rules guiding the clinical diagnosis of Alzheimer's disease in two community-based cohort studies compared to standard practice in a clinic-based cohort study. Neuroepidemiology. 2006;27(3):169–176. doi: 10.1159/000096129. [DOI] [PubMed] [Google Scholar]
- 21.Bennett D.A., Schneider J.A., Arvanitakis Z., et al. Neuropathology of older persons without cognitive impairment from two community-based studies. Neurology. 2006;66(12):1837–1844. doi: 10.1212/01.wnl.0000219668.47116.e6. [DOI] [PubMed] [Google Scholar]
- 22.Bennett D.A., Wilson R.S., Schneider J.A., et al. Natural history of mild cognitive impairment in older persons. Neurology. 2002;59(2):198–205. doi: 10.1212/wnl.59.2.198. [DOI] [PubMed] [Google Scholar]
- 23.Johnson E.C.B., Dammer E.B., Duong D.M., et al. Large-scale proteomic analysis of Alzheimer's disease brain and cerebrospinal fluid reveals early changes in energy metabolism associated with microglia and astrocyte activation. Nat Med. 2020;26(5):769–780. doi: 10.1038/s41591-020-0815-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Subramanian A., Tamayo P., Mootha V.K., et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci. 2005;102(43):15545–15550. doi: 10.1073/pnas.0506580102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Love M.I., Huber W., Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biology. 2014;15(12):550. doi: 10.1186/s13059-014-0550-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Hochberg Y., Benjamini Y. More powerful procedures for multiple significance testing. Stat Med. 1990;9(7):811–818. doi: 10.1002/sim.4780090710. [DOI] [PubMed] [Google Scholar]
- 27.Maly I.V., Hofmann W.A. Bayesian inference of molecular kinetic parameters from astrocyte calcium imaging data. MethodsX. 2022;9 doi: 10.1016/j.mex.2022.101825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Maly I.V., Hofmann W.A., Pletnikov M.V. Experimental and computational analyses of calcium dynamics in 22q11.2 deletion model astrocytes. Neurosci Lett. 2022;783 doi: 10.1016/j.neulet.2022.136711. [DOI] [PubMed] [Google Scholar]
- 29.Lu J., Li C., Singh-Alvarado J., et al. MIN1PIPE: a miniscope 1-photon-based calcium imaging signal extraction pipeline. Cell Rep. 2018;23(12):3673–3684. doi: 10.1016/j.celrep.2018.05.062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Yao J., Pilko A., Wollman R. Distinct cellular states determine calcium signaling response. Mol Syst Biol. 2016;12(12):894. doi: 10.15252/msb.20167137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Klein E.A., Yung Y., Castagnino P., Kothapalli D., Assoian R.K. Cell adhesion, cellular tension, and cell cycle control. Methods Enzymol. 2007;426:155–175. doi: 10.1016/S0076-6879(07)26008-2. [DOI] [PubMed] [Google Scholar]
- 32.Brazzo J.A., Biber J.C., Nimmer E., et al. Mechanosensitive expression of lamellipodin promotes intracellular stiffness, cyclin expression and cell proliferation. J Cell Sci. 2021;134(12) doi: 10.1242/jcs.257709. [DOI] [PubMed] [Google Scholar]
- 33.Ma Y., Jin J., Dong C., et al. High-efficiency siRNA-based gene knockdown in human embryonic stem cells. RNA. 2010;16(12):2564–2569. doi: 10.1261/rna.2350710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Ve H., Cabana V.C., Gouspillou G., Lussier M.P. Quantitative immunoblotting analyses reveal that the abundance of actin, tubulin, synaptophysin and EEA1 proteins is altered in the brains of aged mice. Neuroscience. 2020;442:100–113. doi: 10.1016/j.neuroscience.2020.06.044. [DOI] [PubMed] [Google Scholar]
- 35.Wang Y., Wang C., Ranefall P., et al. SynQuant: an automatic tool to quantify synapses from microscopy images. Bioinformatics. 2020;36(5):1599–1606. doi: 10.1093/bioinformatics/btz760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Rice A.M., McLysaght A. Dosage-sensitive genes in evolution and disease. BMC Biol. 2017;15(1):78. doi: 10.1186/s12915-017-0418-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Yamasaki M., Makino T., Khor S.-S., et al. Sensitivity to gene dosage and gene expression affects genes with copy number variants observed among neuropsychiatric diseases. BMC Med Genomics. 2020;13(1):55. doi: 10.1186/s12920-020-0699-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Johnson E.C.B., Carter E.K., Dammer E.B., et al. Large-scale deep multi-layer analysis of Alzheimer's disease brain reveals strong proteomic disease-related changes not observed at the RNA level. Nat Neurosci. 2022;25(2):213–225. doi: 10.1038/s41593-021-00999-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Ou Y.-N., Xu W., Li J.-Q., et al. FDG-PET as an independent biomarker for Alzheimer's biological diagnosis: a longitudinal study. Alzheimers Res Ther. 2019;11(1):57. doi: 10.1186/s13195-019-0512-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Lewis A.S., van Schalkwyk G.I., Bloch M.H. Alpha-7 nicotinic agonists for cognitive deficits in neuropsychiatric disorders: a translational meta-analysis of rodent and human studies. Prog Neuropsychopharmacol Biol Psychiatry. 2017;75:45–53. doi: 10.1016/j.pnpbp.2017.01.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Li L., Stefan M.I., Le Novère N. Calcium input frequency, duration and amplitude differentially modulate the relative activation of calcineurin and CaMKII. PLoS One. 2012;7(9) doi: 10.1371/journal.pone.0043810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Qi H., Li X., Jin Z., Simmen T., Shuai J. The oscillation amplitude, not the frequency of cytosolic calcium, regulates apoptosis induction. iScience. 2020;23(11) doi: 10.1016/j.isci.2020.101671. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Sneyd J., Han J.M., Wang L., et al. On the dynamical structure of calcium oscillations. Proc Natl Acad Sci U S A. 2017;114(7):1456–1461. doi: 10.1073/pnas.1614613114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Stern S., Hilton B.J., Burnside E.R., et al. RhoA drives actin compaction to restrict axon regeneration and astrocyte reactivity after CNS injury. Neuron. 2021;109(21):3436–3455.e9. doi: 10.1016/j.neuron.2021.08.014. [DOI] [PubMed] [Google Scholar]
- 45.Zamboni V., Armentano M., Berto G., et al. Hyperactivity of Rac1-GTPase pathway impairs neuritogenesis of cortical neurons by altering actin dynamics. Sci Rep. 2018;8(1):7254. doi: 10.1038/s41598-018-25354-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Leyden F., Uthishtran S., Moorthi U.K., et al. Rac1 activation can generate untemplated, lamellar membrane ruffles. BMC Biol. 2021;19(1):72. doi: 10.1186/s12915-021-00997-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Marston D.J., Anderson K.L., Swift M.F., et al. High Rac1 activity is functionally translated into cytosolic structures with unique nanoscale cytoskeletal architecture. Proc Natl Acad Sci. 2019;116(4):1267–1272. doi: 10.1073/pnas.1808830116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Bae Y.H., Mui K.L., Hsu B.Y., et al. A FAK-Cas-Rac-lamellipodin signaling module transduces extracellular matrix stiffness into mechanosensitive cell cycling. Sci Signal. 2014;7(330):ra57. doi: 10.1126/scisignal.2004838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Krause M., Leslie J.D., Stewart M., et al. Lamellipodin, an Ena/VASP ligand, is implicated in the regulation of lamellipodial dynamics. Dev Cell. 2004;7(4):571–583. doi: 10.1016/j.devcel.2004.07.024. [DOI] [PubMed] [Google Scholar]
- 50.Doss B.L., Pan M., Gupta M., et al. Cell response to substrate rigidity is regulated by active and passive cytoskeletal stress. Proc Natl Acad Sci. 2020;117(23):12817–12825. doi: 10.1073/pnas.1917555117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Tang K., Xin Y., Li K., Chen X., Tan Y. Cell cytoskeleton and stiffness are mechanical indicators of organotropism in breast cancer. Biology (Basel) 2021;10(4):259. doi: 10.3390/biology10040259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Budday S., Ovaert T.C., Holzapfel G.A., Steinmann P., Kuhl E. Fifty shades of brain: a review on the mechanical testing and modeling of brain tissue. Arch Comput Methods Eng. 2020;27(4):1187–1230. [Google Scholar]
- 53.Budday S., Nay R., de Rooij R., et al. Mechanical properties of gray and white matter brain tissue by indentation. J Mech Behav Biomed Mater. 2015;46:318–330. doi: 10.1016/j.jmbbm.2015.02.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Weickenmeier J., de Rooij R., Budday S., Ovaert T.C., Kuhl E. The mechanical importance of myelination in the central nervous system. J Mech Behav Biomed Mater. 2017;76:119–124. doi: 10.1016/j.jmbbm.2017.04.017. [DOI] [PubMed] [Google Scholar]
- 55.Murphy M.C., Jones D.T., Jack C.R., et al. Regional brain stiffness changes across the Alzheimer's disease spectrum. Neuroimage Clin. 2016;10:283–290. doi: 10.1016/j.nicl.2015.12.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Govek E.E., Wu Z., Acehan D., et al. Cdc42 regulates neuronal polarity during cerebellar axon formation and glial-guided migration. iScience. 2018;1:35–48. doi: 10.1016/j.isci.2018.01.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Xu Z., Chen Y., Chen Y. Spatiotemporal regulation of Rho GTPases in neuronal migration. Cells. 2019;8(6):568. doi: 10.3390/cells8060568. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Hall A., Lalli G. Rho and Ras GTPases in axon growth, guidance, and branching. Cold Spring Harb Perspect Biol. 2010;2(2):a001818. doi: 10.1101/cshperspect.a001818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Matsuda S., Fujita T., Kajiya M., et al. Brain-derived neurotrophic factor induces migration of endothelial cells through a TrkB-ERK-integrin alphaVbeta3-FAK cascade. J Cell Physiol. 2012;227(5):2123–2129. doi: 10.1002/jcp.22942. [DOI] [PubMed] [Google Scholar]
- 60.Etienne-Manneville S. Cdc42--the centre of polarity. J Cell Sci. 2004;117(Pt 8):1291–1300. doi: 10.1242/jcs.01115. [DOI] [PubMed] [Google Scholar]
- 61.Burute M., Jansen K.I., Mihajlovic M., Vermonden T., Kapitein L.C. Local changes in microtubule network mobility instruct neuronal polarization and axon specification. Sci Adv. 2022;8(44) doi: 10.1126/sciadv.abo2343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Tahirovic S., Hellal F., Neukirchen D., et al. Rac1 regulates neuronal polarization through the WAVE complex. J Neurosci. 2010;30(20):6930–6943. doi: 10.1523/JNEUROSCI.5395-09.2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Iseppon F., Napolitano L.M., Torre V., Cojoc D. Cdc42 and RhoA reveal different spatio-temporal dynamics upon local stimulation with Semaphorin-3A. Front Cell Neurosci. 2015;9:333. doi: 10.3389/fncel.2015.00333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Govek E.E., Newey S.E., Van Aelst L. The role of the Rho GTPases in neuronal development. Genes Dev. 2005;19(1):1–49. doi: 10.1101/gad.1256405. [DOI] [PubMed] [Google Scholar]
- 65.Hua Z.L., Emiliani F.E., Nathans J. Rac1 plays an essential role in axon growth and guidance and in neuronal survival in the central and peripheral nervous systems. Neural Dev. 2015;10(1):21. doi: 10.1186/s13064-015-0049-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Li Q., Michaud M., Shankar R., Canosa S., Schwartz M., Madri J.A. MMP-2: a modulator of neuronal precursor activity and cognitive and motor behaviors. Behav Brain Res. 2017;333:74–82. doi: 10.1016/j.bbr.2017.06.041. [DOI] [PubMed] [Google Scholar]
- 67.Zhuge Y., Xu J. Rac1 mediates type I collagen-dependent MMP-2 activation. role in cell invasion across collagen barrier. J Biol Chem. 2001;276(19):16248–16256. doi: 10.1074/jbc.m010190200. [DOI] [PubMed] [Google Scholar]
- 68.Shu T., Liu C., Pang M., et al. Effects and mechanisms of matrix metalloproteinase2 on neural differentiation of induced pluripotent stem cells. Brain Res. 2018;1678:407–418. doi: 10.1016/j.brainres.2017.11.006. [DOI] [PubMed] [Google Scholar]
- 69.Shaverdashvili K., Wong P., Ma J., Zhang K., Osman I., Bedogni B. MT1-MMP modulates melanoma cell dissemination and metastasis through activation of MMP2 and RAC1. Pigment Cell Melanoma Res. 2014;27(2):287–296. doi: 10.1111/pcmr.12201. [DOI] [PubMed] [Google Scholar]
- 70.Wang X., Wei Z., Tang Z., et al. IL-37bDelta1-45 suppresses the migration and invasion of endometrial cancer cells by targeting the Rac1/NF-kappaB/MMP2 signal pathway. Lab Invest. 2021;101(6):760–774. doi: 10.1038/s41374-021-00544-2. [DOI] [PubMed] [Google Scholar]
- 71.Xia P., Huang M., Zhang Y., et al. NCK1 promotes the angiogenesis of cervical squamous carcinoma via Rac1/PAK1/MMP2 signal pathway. Gynecol Oncol. 2019;152(2):387–395. doi: 10.1016/j.ygyno.2018.11.013. [DOI] [PubMed] [Google Scholar]
- 72.Konietzny A., Bär J., Mikhaylova M. Dendritic actin cytoskeleton: structure, functions, and regulations. Front Cell Neurosci. 2017;11:147. doi: 10.3389/fncel.2017.00147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Zorbaz T., Madrer N., Soreq H. Cholinergic blockade of neuroinflammation: from tissue to RNA regulators. Neuronal Signal. 2022;6(1) doi: 10.1042/NS20210035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Maroli A., Di Lascio S., Drufuca L., et al. Effect of donepezil on the expression and responsiveness to LPS of CHRNA7 and CHRFAM7A in macrophages: a possible link to the cholinergic anti-inflammatory pathway. J Neuroimmunol. 2019;332:155–166. doi: 10.1016/j.jneuroim.2019.04.012. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.