Abstract
We have sequenced the long unique region (LUR) and characterized the terminal repeats of the genome of a rhesus rhadinovirus (RRV), strain 17577. The LUR as sequenced is 131,364 bp in length, with a G+C content of 52.2% and a CpG ratio of 1.11. The genome codes for 79 open reading frames (ORFs), with 67 of these ORFs similar to genes found in both Kaposi’s sarcoma-associated herpesvirus (KSHV) (formal name, human herpesvirus 8) and herpesvirus saimiri. Eight of the 12 unique genes show similarity to genes found in KSHV, including genes for viral interleukin-6, viral macrophage inflammatory protein, and a family of viral interferon regulatory factors (vIRFs). Genomic organization is essentially colinear with KSHV, the primary differences being the number of cytokine and IRF genes and the location of the gene for dihydrofolate reductase. Highly repetitive sequences are located in positions corresponding to repetitive sequences found in KSHV. Phylogenetic analysis of several ORFs supports the similarity between RRV and KSHV. Overall, the sequence, structural, and phylogenetic data combine to provide strong evidence that RRV 17577 is the rhesus macaque homolog of KSHV.
Kaposi’s sarcoma (KS) is a vascular disorder frequently occurring in patients infected with the human immunodeficiency virus (HIV). The nature of KS is unclear, with evidence for both neoplasia (47) and hyperplasia (9, 22). This debate notwithstanding, the occurrence of non-HIV-associated KS in regions of Africa (24), in elderly Mediterranean men (12), and as an occasional iatrogenic complication of organ transplants (58) is indicative of a KS pathogen independent of HIV. A putative rhadinovirus agent for KS, Kaposi’s sarcoma-associated herpesvirus (KSHV), was recently characterized (7, 38, 51) and determined to possess a number of unique genes which distinguish it from other herpesviruses, including genes for viral interleukin-6 (vIL-6), viral macrophage inflammatory proteins (vMIPs), and viral interferon regulatory factors (vIRFs). KSHV, known formally as human herpesvirus 8, has been the subject of intense scrutiny since its discovery, but the mechanism of its involvement with KS has not yet been determined.
Despite disagreement over a causative role for KSHV in KS (16, 17), the preponderance of evidence indicates that the virus is critical for KS development. KSHV has been linked epidemiologically to KS in a number of studies. Chang et al. (7) identified DNA indicative of the virus in 90% of KS tissue and in 15% of non-KS tissue from AIDS patients but not in tissue from non-HIV-infected individuals. Large-scale PCR screening, in situ hybridization of KS tissue, and serological testing has associated KSHV with all forms of KS, regardless of HIV status (2, 12, 25, 36, 52). Serological studies indicate that KSHV infection precedes KS and is predictive of KS development (31, 60).
In addition to its involvement with KS, KSHV is associated with a type of B-cell lymphoma referred to as primary effusion lymphoma (PEL) (40). It is also associated with nonneoplastic lymphadenopathy (29) and with a subset of multicentric Castleman’s disease (53). The determination of a role for KSHV in PEL development is complicated by ubiquitous coinfection with Epstein-Barr virus (EBV).
KSHV possesses a number of potential transforming genes, including K1 (27), an IL-8 receptor-like G protein-coupled receptor (3, 4, 6, 20), kaposin (39), and several vIRF genes (18, 28, 63). In addition, IL-6 and, by implication its viral homolog, promote continued development of KS and PEL (34, 41, 43). This large number of potential transforming genes suggests that the virus’s pathogenic potential is complex. Unraveling the function of these genes would be aided by the use of recombinant virus in an animal model. KSHV does not induce KS in monkeys, but close relatives of KSHV exist in various macaque species and may be involved in a variety of diseases similar to KSHV-associated disorders. Desrosiers and colleagues (10) recently reported a 10,595-bp segment of a rhesus macaque rhadinovirus (RRV) (isolate H26-95; Genbank accession no. AF029302) that included, from left to right, a partial gene for open reading frame 7 (ORF 7); intact genes for glycoprotein B (ORF 8), DNA polymerase (ORF 9), ORFs 10 and 11, and vIL-6; and a partial gene for thymidylate synthase (TS) (ORF 70). This arrangement of genes is very similar to the arrangement of KSHV, but no pathology has been reported for H26-95. PCR-derived DNA polymerase sequences similar to KSHV polymerase have also been recovered from tissues isolated from KS-like retroperitoneal fibromatosis (RF) in pigtail macaques (Macaca nemestrina) and rhesus macaques (Macaca mulatta) (49).
We have isolated an RRV (isolate 17577, referred to in this paper as RRV) from a simian immunodeficiency virus (SIV)-infected macaque with a lymphoproliferative disorder (LPD) (61). A preliminary characterization of the virus indicated a nucleotide structure almost identical to that of RRV H26-95, and PCR analysis showed that the virus was present in diseased tissue but not in normal tissue. Moreover, experimental RRV infection of SIV-infected macaques resulted in the induction of an LPD similar to that observed in multicentric Castleman’s disease patients (61). In this study we characterize the genome of this rhesus macaque rhadinovirus. Analysis of the primary structure reveals that RRV is highly similar to KSHV, possesses a number of KSHV-specific genes, and is likely to be the rhesus macaque equivalent to KSHV.
MATERIALS AND METHODS
Preparation of viral DNA.
Primary rhesus fibroblasts grown in two 850-cm2 roller bottles were infected with RRV at a multiplicity of infection of 0.1, and the virus was harvested from the culture supernatant and infected monolayers at 10 to 12 days postinfection. Cellular debris was removed from the culture supernatant by centrifugation at 1,000 × g for 10 min. Intracellular virus particles were released by sonication followed by centrifugation to pellet debris.
The two clarified supernatants were then combined, and the virus was pelleted by centrifugation at 12,500 × g for 1 h at 4°C and further purified through a six-step sorbitol gradient ranging from 20 to 70%. Gradients were spun in a Beckman SW41 rotor for 2 h at 18,000 rpm at 4°C. The interface containing the virus was collected and diluted with cold buffered saline solution. The virus was then pelleted by centrifugation in the SW41 rotor for 50 min at 18,000 rpm. The virus pellet was resuspended in 9.2 ml of 10 mM Tris (pH 8.0)–1 mM EDTA (TE) before the addition of 0.6 ml of 10% sodium dodecyl sulfate (SDS) and 0.2 ml of proteinase K (10 mg/ml) to release the viral DNA. Viral DNA was isolated by CsCl2 gradient centrifugation in a Beckman Ti75 rotor at 38,400 rpm for 72 h, collected, and dialyzed against TE.
To ensure that the isolated DNA contained all of the sequences required for RRV replication, DNA was transfected, in duplicate, into primary rhesus fibroblasts by the calcium phosphate method without dimethyl sulfoxide shock, and cells were observed for cytopathic effects. Control transfections, lacking viral DNA or calcium phosphate, did not develop cytopathic effects.
Construction and analysis of the cosmid library.
Approximately 100 μg of purified viral DNA was partially digested with Sau3AI. Aliquots taken at various time points were run on a 0.5% agarose gel and examined for the fraction which gave the desired range of fragments (30 to 42 kb). The selected fraction was dephosphorylated with calf intestinal alkaline phosphatase, and 1 μg of the dephosphorylated DNA was ligated into the cosmid vector SuperCos 1, prepared as described by the manufacturer (Stratagene). The resulting ligation product was packaged by using GigaPack II Gold packaging extract (Stratagene) and grown for the isolation of recombinant cosmids.
Individual cosmids were selected for sequencing by Southern blot analysis. Recombinant cosmids were grown in 3-ml cultures, and cosmid DNA was isolated by alkaline lysis. Cosmid DNA was digested with EcoRI, and the DNA fragments were separated on a 0.8% agarose gel. The separated fragments were transferred to a duralon membrane and probed with various PCR amplification products corresponding to specific KSHV ORFs (ORFs 25, 56, and 69) and to RRV TS and DNA polymerase (61). The KSHV probes were generated by PCR with DNA isolated from the BCBL-1 cell line (48). Hybridization of the probes to the transferred recombinant cosmids was done under conditions of moderate stringency (2× SSC [1× SSC is 0.15 M NaCl plus 0.015 M sodium citrate]–0.1% SDS at 55°C) for each of the KSHV-specific probes and at high stringency (0.2× SSC–0.1% SDS at 60°C) for the RRV-specific probes. Cosmids which hybridized to the probes were grown in 50-ml cultures and purified by cesium chloride gradient centrifugation. Purified cosmids were end sequenced with T3 and T7 primers.
Cloning and sequencing.
Ten micrograms of each purified recombinant cosmid was digested with EcoRI. The digested products were isolated from a 0.8% agarose gel by using the QiaQuick gel extraction protocol (Qiagen) and ligated into pSP73 (Promega). Sequencing templates were prepared by alkaline lysis, followed by precipitation with 6.5% polyethylene glycol–0.8 M NaCl. Templates were resuspended at a concentration of 0.1 μg/μl, and end sequences were determined with primers corresponding to the SP6 and T7 promoters of pSP73. Internal sequences were determined by a combination of subcloning with convenient restriction sites and custom primers. DNA sequencing reactions were performed with Applied Biosystems (ABI) PRISM Dye Terminator Cycle Sequencing Ready Reaction kits with Amplitaq DNA polymerase per the manufacturer’s instructions. Sequence data were acquired with an ABI 373A sequencer in the Molecular Biology Core at the Oregon Regional Primate Research Center. The primary EcoRI products were sequentially arranged by sequencing across the EcoRI sites in the intact cosmids with custom primers. Corresponding EcoRI fragments from overlapping cosmids were verified by end sequencing. Except for those regions containing long, high G+C repeat units, the entire viral DNA sequence was determined with a redundancy of three- to fourfold.
Sequences not accessible to custom primers or restriction subcloning were determined by deletion subcloning with the Exo Size Deletion kit (New England Biolabs). Minor modifications were made to the manufacturer’s recommended incubation times and concentrations to optimize for the use of polyethylene glycol-precipitated DNA as a template for the deletions. Deletion products were size selected by restriction digests of DNA recovered from 3-ml cultures, and selected plasmids were prepared for sequencing as described.
Assembly of sequence, assignment of ORFs, and nomenclature.
Factura (ABI) and Autoassembler (ABI) were used to assemble the final sequence from individual sequencing runs. ORFs in the RRV sequence were determined with the program MacVector, version 6.01 (Oxford Molecular Group), by using a target size of 100 or more amino acids. Putative ORFs were then translated and compared to a database of KSHV ORFs. RRV ORFs which matched KSHV ORFs were then compared to GenBank by using BLASTP to verify similarity, followed by a Gap analysis (Wisconsin GCG analysis package, version 9.1; Oxford Molecular Group) to determine the levels of similarity and identity between the RRV and KSHV proteins. When a gap in the genome of RRV corresponded to the location of a KSHV ORF with less than 100 amino acids, MacVector was reset to a lower limit. RRV ORFs were assigned the names of herpesvirus saimiri (HVS) ORFs when they showed similarity to KSHV ORFs with the same name. In this fashion, some RRV ORFs, such as ORFs 28 and 45, were named as such despite having no similarity to HVS ORFs.
Sequence alignment and phylogenetic analysis.
Phylogenetic analysis of specific RRV ORFs was performed similarly to that described for the alphaherpesviruses (32, 33). Sequence alignment was performed with ClustalW, version 1.4 (55), as implemented by MacVector. Blosum 30 was used for pairwise alignment, and the Blosum series was used for multiple sequence alignment. Pairwise and multiple sequence alignments both used a gap introduction penalty of 10 and a gap extension penalty of 0.1. Bootstrap analysis was performed with the programs Seqboot, Protpars, and Consense from the Phylip package, version 3.572c (copyrighted and distributed without charge by Joseph Felsenstein and the University of Washington). The Consense treefile was displayed with TreeView (45).
Viral sequences for phylogenetic analysis were acquired from GenBank. The viruses and their accession numbers are as follows: HVS, X64346; KSHV, U756998; alcelaphine herpesvirus (AHV), AF005370; murine herpesvirus 68 (MHV), U97553; EBV, V01555, J02070, K01729, K01730, V01554, X00498, X00499, and X00784; cytomegalovirus (CMV), X17403. The AHV submission is not annotated, so ORFs were derived from the published table (13).
The proteins used for phylogenetic analysis are single-stranded DNA binding protein (ssDBP), glycoprotein B (gB), DNA polymerase (Pol), major capsid protein (MCP), helicase (Hel), and uracyl-DNA glycosylase (UDG). The respective ORFs for AHV, HVS, KSHV, MHV, and RRV are ORFs 6, 8, 9, 25, 44, and 46; those for EBV are BALF2, BALF4, BALF5, BCLF1, BBLF4, and BKRF3; and those for CMV are UL57, UL55, UL54, UL86, UL105, and UL114.
Nucleotide sequence accession number.
RRV 17577 nucleotide sequence data have been deposited in the GenBank, EMBL, and DDBJ nucleotide sequence databases under accession no. AF083501.
RESULTS
Primary structure of the genome of RRV.
The nucleotide sequence of the genome of RRV was determined by using 29 EcoRI fragments from seven overlapping isolates of a partial Sau3AI cosmid library. Cosmids were selected by hybridization with PCR products from KSHV ORFs 25, 56, and 69 (51) and RRV ORFs 9 and 70 (61). EcoRI fragments from each cosmid were subcloned into pSP73 and sequenced. The EcoRI fragments were arranged in the proper order by sequencing across the EcoRI junctions in the parent cosmids with custom primers. Greater than 98% of the viral long unique region (LUR) was determined on both strands. The average sequencing redundancy was between 3 and 4, but three regions were sequenced on only one strand. One of these regions is a 106-bp segment of ORF 61 that was blocked on one side by an apparent hairpin. This segment was sequenced multiple times in one direction with templates derived from independent overlapping cosmids. The other two regions are long, high G+C, repetitive sequences. These segments, which are discussed in more detail below, were sequenced on one strand with a combination of custom primers and exonuclease III deletions. Terminal repeats have been identified at each end of the LUR. The terminal repeat structure is discussed below.
The sequence between the left and right terminal repeats was designated the LUR of the genome. The first base to the right of the left terminal repeat was designated base one. The LUR, as sequenced, is 131,364 bp long, with evidence of endogenous variability in one repetitive element. The G+C content of RRV is 52.2%, which is comparable to the 53.5% G+C content of KSHV but considerably higher than the 34.5% G+C content of the HVS genome. The CpG ratio is 1.11, which is substantially higher than the ratio found for other gammaherpesviruses (21).
Potential ORFs were identified by MacVector and compared to a database containing the full complement of known KSHV ORFs. To ensure that the use of the KSHV database did not bias our evaluation of potential ORFs, all deduced RRV proteins were compared to GenBank with BLASTP (default parameters). These searches resulted in KSHV or HVS proteins as the primary match in all but 11 cases. ORF 2 (dihydrofolate reductase [DHFR]), R3 (vMIP), ORF 70 (TS), and R15 (NCAM Ox-2) scored most highly with their cellular homologs. ORFs 48 and 52 aligned best with the corresponding proteins from AHV and MHV, respectively; the corresponding KSHV protein was the second highest score for each. R8, R12, and R13 were most similar to a variety of human IRFs. R1 aligned with a number of Fc receptors but did not align with any viral proteins. R2 (vIL-6) returned no significant matches.
Seventy-nine RRV ORFs were identified, with 67 of these corresponding to ORFs found in both KSHV and HVS. In accordance with the standard nomenclature for rhadinoviruses, these ORFs were labeled according to the HVS designation. The 12 ORFs not found in HVS were assigned labels beginning with R (for rhesus), indicating their presence in RRV but not HVS. Most of these genes have counterparts in KSHV. The organization of the ORFs is highly similar to the arrangement of KSHV. The organization also conforms to the structural pattern developed for gammaherpesviruses, in which blocks of conserved sequences (the HVS-like ORFs) are interrupted by blocks of acquired cellular genes referred to as divergent loci, as has been diagrammed elsewhere (44).
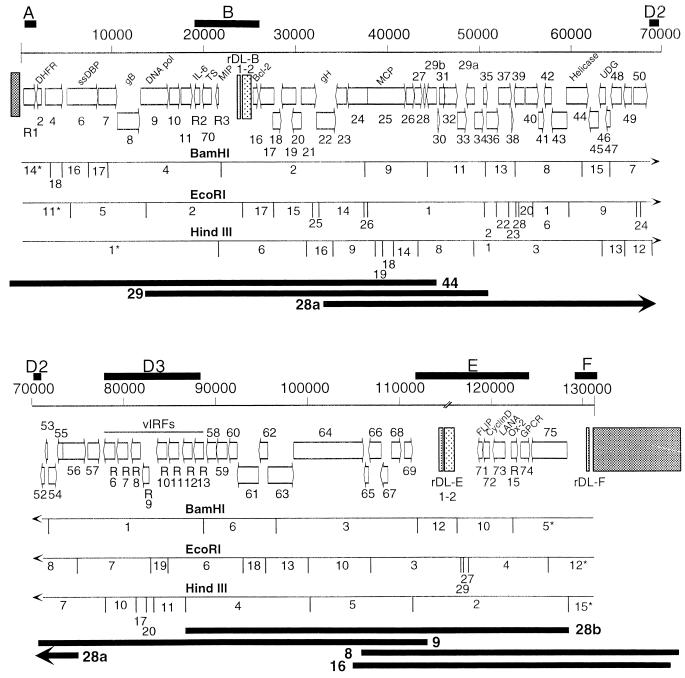
A map of the genome of RRV 17577 is presented in Fig. 1, with all identified ORFs and their orientations. Cosmids used for sequencing of the genome are indicated on the figure. There is a substantial overlap among the cosmids, which reduces the possibility that the organization of the genome as determined by the sequence data is compromised by rearrangements in the cosmid library. Three restriction maps are presented with the genome map: BamHI, EcoRI, and HindIII. Fragment sizes for each restriction map are presented in Table 1. The BamHI and HindIII maps were generated from the final compiled sequence. The EcoRI map was also generated from the final compiled sequence, but it was further characterized by sequencing across the EcoRI junctions in the parent cosmids. All BamHI and HindIII sites identified by the compiled sequence were verified by restriction mapping of appropriate cosmid subclones.
FIG. 1.
Map of the genome of RRV 17577. The ORFs of RRV 17577 are represented by arrows indicating the direction of transcription. Divergent loci are indicated by black bars above the scale; their assignment is based on the work of Nicholas et al. (44). ORF numbers are presented above or below the appropriate ORF. ORFs resident in the divergent loci are named, as are the ORFs used for bootstrap analysis. Terminal repeat sequences are represented by the dark hatched boxes at either end of the LUR. Textured boxes within the LUR represent repeat sequences, which have been named for the appropriate divergent loci. BamHI, EcoRI, and HindIII deduced restriction maps are presented. Digest products are numbered based on the sizing presented in Table 1. Fragments marked with asterisks contain terminal repeat sequences; the sizes listed for these fragments represent the portion of the fragment in the LUR. The hash marks on the map above the repeat unit rDL-E indicate that there is some ambiguity about the length of the repeats. ORFs found in DL-D2 and DL-F are not shown on the map, pending verification of transcription from these regions.
TABLE 1.
Restriction fragments of the RRV 17577 genomea
|
BamHI
|
EcoRI
|
HindIII
|
|||
|---|---|---|---|---|---|
| Fragment no. | Fragment size (bp) | Fragment no. | Fragment size (bp) | Fragment no. | Fragment size (bp) |
| 1 | 17,189 | 1 | 12,476 | 1∗ | 22,006 |
| 2 | 15,598 | 2 | 10,341 | 2 | 17,108 |
| 3 | 15,441 | 3 | 9,565 | 3 | 14,134 |
| 4 | 12,360 | 4 | 9,213 | 4 | 13,511 |
| 5∗ | 8,943 | 5 | 8,465 | 5 | 11,516 |
| 6 | 7,747 | 6 | 8,031 | 6 | 10,743 |
| 7 | 7,718 | 7 | 7,969 | 7 | 8,452 |
| 8 | 7,142 | 8 | 7,416 | 8 | 5,995 |
| 9 | 6,667 | 9 | 7,278 | 9 | 4,679 |
| 10 | 6,474 | 10 | 7,002 | 10 | 3,374 |
| 11 | 6,333 | 11∗ | 5,400 | 11 | 3,030 |
| 12 | 3,978 | 12∗ | 5,054 | 12 | 2,963 |
| 13 | 3,411 | 13 | 4,907 | 13 | 2,891 |
| 14∗ | 3,157 | 14 | 4,771 | 14 | 2,849 |
| 15 | 3,008 | 15 | 4,272 | 15∗ | 2,832 |
| 16 | 2,916 | 16 | 4,099 | 16 | 1,599 |
| 17 | 2,210 | 17 | 3,516 | 17 | 1,272 |
| 18 | 1,343 | 18 | 2,102 | 18 | 1,016 |
| 19 | 1,868 | 19 | 853 | ||
| 20 | 1,603 | 20 | 811 | ||
| 21 | 1,512 | ||||
| 22 | 1,221 | ||||
| 23 | 910 | ||||
| 24 | 624 | ||||
| 25 | 609 | ||||
| 26 | 592 | ||||
| 27 | 584 | ||||
| 28 | 122 | ||||
| 29 | 107 | ||||
∗, fragment size excludes terminal repeat sequences.
To further verify that the sequence derived from the overlapping cosmids was colinear with the sequence of the viral genome, RRV genomic DNA was digested with the restriction enzymes BamHI, EcoRI, and HindIII (Fig. 2). The results of the digests were compared to the predicted digests generated from the completed sequence of the genome. The BamHI digest produced a number of large, similarly sized fragments which were more difficult to interpret than the patterns for EcoRI and HindIII. The composite nature of several of the BamHI bands was determined by electrophoresis of substantially smaller amounts of genomic DNA (data not shown), which allowed visualization of only the largest bands. The BamHI digest provides important evidence for the added complexity of the repeat structure of the RRV genome. The 3.9-kb BamHI digest product (BamHI fragment 12) that is predicted by the sequence data, and which contains one of the high G+C repeat elements, is missing from the BamHI digest. The absence of this band appears to reflect natural variability within the virus and is discussed below in the section describing RRV repetitive sequences.
FIG. 2.
Restriction digests of RRV 17577 genomic DNA. Genomic DNA (15 μg) was precipitated and then resuspended and distributed for three restriction digests. DNA was digested overnight with 10 units of either BamHI, EcoRI, or HindIII at 37°C overnight. The digests were supplemented with 10 units of the appropriate enzyme and incubated for a further 4 h before the digest products were separated on a 0.7% agarose gel. Numbering of fragments is based on the data in Table 1. The numbers 5 + 14, 11 + 12, and 1 + 15 indicate restriction fragments that are attached to the terminal repeats. The band in the BamHI digest at 11 kb is a residual partial digestion product. The faint bands seen at 1 kb in the BamHI and EcoRI lanes are artifacts of reproduction; only the band in the HindIII lane is visible on the original photograph. Bands smaller than 1 kb were too faint to see. The absence of band 12 in the BamHI digest lane is discussed in the text. The standards used are 1-kb standards (Life Technologies); the largest band is 12 kb, and the smallest band visible in the figure is 1 kb.
Correspondence between predicted restriction digest patterns and actual patterns is not necessarily confirmation that the structure of the virus matches that predicted by the sequence data. This is particularly important regarding the BamHI digest. The use of three different enzymes to compare the predicted and actual structures, though, strengthens the argument significantly. The combination of the restriction digest data with the use of independent overlapping clones strongly supports the organization presented in this paper as the correct structure for RRV.
Comparison of RRV 17577 to RRV H26-95.
The segment of the genome of RRV H26-95 available in Genbank (10) was compared to the corresponding region of RRV 17577. Of the 10,595 bases compared, there are 84 differences. All of these occur within ORFs, and most code for silent mutations. ORFs 9, 10, and 11 are identical at the protein level. Only partial sequences for H26-95 ORFs 7 and 70 are available, but these are also identical at the protein level to the corresponding ORFs of 17577. ORF 8, gB, has 19 amino acid differences, all occurring between residues 276 and 539, inclusively (Fig. 3). Sixteen of these alterations occur in the smaller region from residue 346 to residue 440. Seven of the changes in ORF 8 result in a change in the charge of the side group, which may have significance in gB function. None of the variations between H26-95 and 17577 occur at residues common to all gB sequences as aligned by ClustalW (data not shown). Strain variations in the sequence of gB have been noted previously for CMV (8, 15, 50), and it is likely that the sequences noted here indicate that the two sequences for RRV represent unique strains of the virus.
FIG. 3.
Comparison of gB from RRV 17577 and RRV H26-95. The deduced sequences for gB from the two strains of RRV were aligned by ClustalW. No variations occurred between the two except in the regions between residues 274 and 540. These variations in gB account for all of the variations at the protein level between the two strains.
Positioning of RRV, HVS, and KSHV DHFR and TS.
The structure of RRV is highly similar to that of KSHV and, to a much lesser extent, to that of HVS. Notably, ORFs found in all three of the primate rhadinoviruses are colocalized, with two exceptions. The genes for KSHV DHFR and TS (ORFs 2 and 70, respectively) are found in the cytokine-containing segment of the KSHV genome and not in locations corresponding to HVS ORFs 2 and 70 (51). In contrast to this, the location of the RRV DHFR gene corresponds with that of the HVS DHFR gene. RRV, though, is similar to KSHV in having its TS gene located near the cytokine genes.
Comparison of HVS-like ORFs identified in RRV and KSHV.
Data for RRV ORFs are presented in Table 2, along with the results of the Gap analysis of ORFs shared by RRV, KSHV, and HVS. All HVS-like ORFs found in KSHV are found in RRV. Similarities between RRV and KSHV proteins range from a low of 23.6% for ORF 73, the latent nuclear antigen and a potential leucine zipper protein, to a high of 79.9% for ORF 25, the major capsid protein. ORF 73 also displays the only significant size difference found among the ORFs of the two viruses: RRV ORF 73 is 447 residues long, while KSHV ORF 73 is 1,162 residues long. Of the 67 HVS-like ORFs shared by RRV and KSHV, 48 show greater than 50% similarity when RRV and KSHV ORFs are compared by Gap analysis, and 27 show greater than 60% similarity. There appears to be no general clustering of more- or less-conserved ORFs in particular regions of the genome.
TABLE 2.
Comparison of RRV, KSHV, and HVS ORFsa
| ORFb | RRV
|
KSHV
|
HVS
|
Putative functionc | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Start | Stop | Strand | Size (aa) | Size (aa) | % Sim | % Id | Size (aa) | % Sim | % Id | ||
| R11 | 513 | 1784 | + | 423 | |||||||
| ORF 2 | 2418 | 1852 | − | 188 | 210 | 55.1 | 46.0 | 187 | 65.6 | 54.8 | DHFR |
| ORF 42 | 2836 | 4773 | + | 645 | 550 | 40.9 | 35.7 | 360 | 42.0 | 35.3 | Complement binding protein |
| 287 | 44.0 | 38.6 | |||||||||
| ORF 6 | 5205 | 8603 | + | 1,132 | 1,133 | 71.3 | 63.3 | 1,128 | 65.2 | 53.5 | ssDBP |
| ORF 7 | 8628 | 10688 | + | 686 | 695 | 60.1 | 51.5 | 679 | 58.1 | 47.7 | Transport protein |
| ORF 8 | 10675 | 13164 | + | 829 | 845 | 73.3 | 65.5 | 808 | 62.4 | 53.1 | gB |
| ORF 9 | 13282 | 16326 | + | 1,014 | 1,312 | 75.0 | 67.0 | 1,009 | 71.0 | 62.5 | Pol |
| ORF 10 | 16421 | 17575 | + | 384 | 418 | 43.5 | 34.8 | 407 | 33.6 | 23.3 | |
| ORF 11 | 17681 | 18910 | + | 409 | 407 | 41.3 | 31.7 | 405 | 46.3 | 32.4 | |
| R21 | 19705 | 19082 | − | 207 | vIL-6 | ||||||
| ORF 70 | 20939 | 19938 | − | 333 | 337 | 72.1 | 66.1 | 294 | 72.1 | 64.6 | TS |
| R33 | 21753 | 21406 | − | 115 | 95 | 41.9 | 32.3 | vMIP | |||
| ORF 16 | 26007 | 26570 | + | 187 | 175 | 58.0 | 46.0 | 160 | 31.4 | 21.4 | Bcl-2 homolog |
| ORF 17 | 28286 | 26676 | − | 536 | 553 | 50.6 | 44.3 | 475 | 49.0 | 42.2 | Capsid protein |
| ORF 18 | 28159 | 29058 | + | 299 | 257 | 68.1 | 58.0 | 256 | 60.2 | 48.8 | |
| ORF 19 | 30709 | 29066 | − | 547 | 549 | 61.1 | 52.8 | 543 | 55.5 | 46.9 | Tegument protein |
| ORF 20 | 31256 | 30204 | − | 350 | 320 | 51.8 | 44.7 | 303 | 43.2 | 35.6 | |
| ORF 21 | 31255 | 32928 | + | 557 | 580 | 54.0 | 44.6 | 527 | 39.0 | 31.7 | Thymidine kinase |
| ORF 22 | 32915 | 35029 | + | 704 | 730 | 50.1 | 40.7 | 717 | 42.3 | 31.5 | gH |
| ORF 23 | 36234 | 35026 | − | 402 | 404 | 56.8 | 48.5 | 253 | 40.5 | 29.8 | |
| ORF 24 | 38482 | 36284 | − | 732 | 752 | 66.3 | 58.7 | 731 | 56.3 | 46.8 | |
| ORF 25 | 38484 | 42620 | + | 1,378 | 1,376 | 79.9 | 72.5 | 1,371 | 76.7 | 67.5 | MCP |
| ORF 26 | 42652 | 43575 | + | 307 | 305 | 71.8 | 64.3 | 304 | 69.1 | 58.2 | Capsid protein |
| ORF 27 | 43600 | 44409 | + | 269 | 290 | 33.6 | 25.3 | 280 | 35.0 | 27.1 | |
| ORF 28 | 44575 | 44850 | + | 91 | 102 | 30.1 | 26.5 | 93 | |||
| ORF 29b | 45946 | 44900 | − | 348 | 351 | 77.6 | 66.4 | 387 | 74.4 | 62.9 | Packaging protein |
| ORF 30 | 46072 | 46302 | + | 76 | 77 | 51.3 | 38.2 | 75 | 40.3 | 29.2 | |
| ORF 31 | 46260 | 46913 | + | 217 | 224 | 56.0 | 45.4 | 208 | 50.5 | 39.9 | |
| ORF 32 | 46850 | 48244 | + | 464 | 454 | 49.9 | 41.8 | 441 | 43.2 | 34.1 | |
| ORF 33 | 48216 | 49226 | + | 336 | 312 | 52.1 | 42.1 | 330 | 49.1 | 39.1 | |
| ORF 29a | 50127 | 49147 | − | 327 | 312 | 66.7 | 61.2 | 303 | 57.8 | 49.8 | Packaging protein |
| ORF 34 | 50126 | 51109 | + | 327 | 327 | 58.9 | 48.5 | 316 | 53.7 | 40.6 | |
| ORF 35 | 51090 | 51539 | + | 149 | 151 | 47.7 | 35.6 | 150 | 51.0 | 37.4 | |
| ORF 36 | 51445 | 52752 | + | 435 | 444 | 56.0 | 46.1 | 431 | 38.4 | 28.7 | Kinase |
| ORF 37 | 52733 | 54175 | + | 480 | 486 | 72.4 | 63.5 | 483 | 63.0 | 53.2 | Alkaline exonuclease |
| ORF 38 | 54130 | 54339 | + | 69 | 61 | 56.7 | 45.0 | 66 | 39.4 | 34.8 | |
| ORF 39 | 55558 | 54422 | − | 378 | 399 | 73.0 | 59.3 | 366 | 67.1 | 57.0 | gM |
| ORF 40 | 55693 | 57099 | + | 468 | 457 | 42.2 | 32.7 | 450 | 39.1 | 28.1 | Helicase-primase |
| ORF 41 | 57084 | 57695 | + | 203 | 205 | 33.5 | 26.0 | 161 | 37.3 | 29.1 | Helicase-primase |
| ORF 42 | 58510 | 57692 | − | 272 | 278 | 56.8 | 46.1 | 265 | 51.2 | 38.1 | |
| ORF 43 | 60194 | 58464 | − | 576 | 605 | 69.7 | 61.6 | 563 | 66.4 | 56.6 | Capsid protein |
| ORF 44 | 60133 | 62505 | + | 790 | 788 | 73.9 | 66.0 | 781 | 71.1 | 62.6 | Helicase-primase |
| ORF 45 | 63604 | 62546 | − | 352 | 407 | 31.2 | 24.9 | 257 | |||
| ORF 46 | 64413 | 63646 | − | 255 | 255 | 71.9 | 60.1 | 252 | 67.5 | 59.1 | UDG |
| ORF 47 | 64898 | 64389 | − | 169 | 167 | 31.9 | 27.7 | 141 | 33.3 | 23.9 | gL |
| ORF 48 | 66335 | 65166 | − | 389 | 402 | 36.2 | 29.2 | 797 | 34.1 | 25.8 | |
| ORF 49 | 67470 | 66565 | − | 301 | 302 | 66.1 | 54.2 | 303 | 35.1 | 23.3 | |
| ORF 50 | 67661 | 69205 | + | 514 | 631 | 46.6 | 37.8 | 535 | 29.7 | 21.6 | Transactivator |
| ORF 52 | 71990 | 71571 | − | 139 | 131 | 58.5 | 45.4 | 115 | 41.7 | 30.4 | |
| ORF 53 | 72368 | 72054 | − | 104 | 110 | 51.0 | 46.2 | 90 | 43.3 | 28.9 | |
| ORF 54 | 72444 | 73316 | + | 290 | 318 | 48.6 | 41.0 | 287 | 46.5 | 36.4 | dUTPase |
| ORF 55 | 74009 | 73377 | − | 210 | 227 | 62.9 | 55.2 | 200 | 52.5 | 44.4 | |
| ORF 56 | 74021 | 76507 | + | 828 | 843 | 61.2 | 52.5 | 835 | 54.0 | 43.6 | DNA replication protein |
| ORF 57 | 76748 | 78076 | + | 442 | 275 | 60.6 | 47.1 | 416 | 40.3 | 31.5 | Immediate-early protein |
| R64 | 79683 | 78436 | − | 415 | 449 | 26.0 | 21.1 | vIRF | |||
| R74 | 81103 | 79856 | − | 415 | 449 | 28.3 | 20.7 | vIRF | |||
| R84 | 82487 | 81432 | − | 351 | 449 | 28.9 | 19.4 | vIRF | |||
| R94 | 83422 | 82661 | − | 253 | vIRF | ||||||
| R104 | 85379 | 84222 | − | 385 | 449 | 33.7 | 26.2 | vIRF | |||
| R114 | 86697 | 85525 | − | 390 | 449 | 30.0 | 21.8 | vIRF | |||
| R124 | 88131 | 87064 | − | 355 | vIRF | ||||||
| R134 | 89386 | 88292 | − | 364 | vIRF | ||||||
| ORF 58 | 90714 | 89632 | − | 360 | 357 | 45.2 | 38.2 | 357 | 39.9 | 29.5 | |
| ORF 59 | 91909 | 90725 | − | 394 | 396 | 60.3 | 51.8 | 368 | 40.7 | 32.7 | DNA replication protein |
| ORF 60 | 92982 | 92038 | − | 314 | 305 | 78.2 | 70.0 | 305 | 71.0 | 62.4 | Ribonucleotide reductase, small |
| ORF 61 | 95330 | 92964 | − | 788 | 792 | 69.3 | 61.7 | 767 | 64.4 | 53.3 | Ribonucleotide reductase, large |
| ORF 62 | 96328 | 95333 | − | 331 | 331 | 64.4 | 56.5 | 330 | 53.8 | 41.9 | Assembly/DNA maturation |
| ORF 63 | 96327 | 99146 | + | 939 | 927 | 51.8 | 42.6 | 899 | 43.4 | 34.6 | Tegument protein |
| ORF 64 | 99150 | 106796 | + | 2,548 | 2,635 | 49.6 | 40.2 | 2,469 | 39.2 | 29.4 | Tegument protein |
| ORF 65 | 107316 | 106807 | − | 169 | 170 | 48.2 | 38.6 | 139 | 41.0 | 33.1 | Capsid protein |
| ORF 66 | 108668 | 107322 | − | 448 | 429 | 51.9 | 46.4 | 435 | 43.6 | 32.3 | |
| ORF 67 | 109368 | 108694 | − | 224 | 271 | 69.6 | 64.7 | 253 | 58.6 | 51.4 | Tegument protein |
| ORF 68 | 109779 | 111152 | + | 457 | 545 | 53.2 | 44.8 | 436 | 53.5 | 44.3 | Glycoprotein |
| ORF 69 | 111174 | 112067 | + | 297 | 225 | 73.1 | 65.5 | 261 | 57.5 | 49.0 | |
| ORF 71 | 118905 | 118381 | − | 174 | 139 | 38.8 | 30.9 | 167 | 25.3 | 15.1 | FLIP homolog |
| ORF 72 | 119728 | 118964 | − | 254 | 257 | 49.8 | 38.6 | 254 | 37.5 | 29.2 | Cyclin D homolog |
| ORF 73 | 121379 | 120036 | − | 447 | 1,162 | 23.6 | 16.8 | 407 | 29.0 | 20.8 | Latency-associated nuclear antigen |
| R155 | 122033 | 122794 | + | 253 | 348 | 35.2 | 31.2 | N-CAM Ox-2 homolog | |||
| ORF 74 | 123091 | 124119 | + | 342 | 342 | 51.6 | 41.1 | 321 | 41.1 | 32.1 | G-protein coupled receptor |
| ORF 75 | 128120 | 124224 | − | 1,298 | 1,296 | 52.2 | 44.0 | 1,299 | 43.2 | 34.4 | Tegument protein/FGARAT |
Sim, similar; Id, identical; aa, amino acids.
1, no similarity found; 2, compared to HVS ORF 4a and 4b; 3, compared to KSHV R4; 4, compared to KSHV K9; 5, compared to KSHV K14.
FGARAT, N-formalglycinamide ribotide amidotransferase.
Comparison of HVS-like ORFs identified in RRV and HVS.
In general, RRV and HVS ORFs are highly similar when the corresponding RRV and KSHV ORFs are highly similar, although the Gap values are generally lower. Of the 67 ORFs shared by RRV and HVS, only 30 have a similarity greater than 50% and only 16 have a similarity greater than 60%. The range of similarity based on Gap analysis between RRV and HVS extends from nonexistent for ORFs 28 and 45 to 76.7% for ORF 25. Note that RRV ORFs 28 and 45, which have no similarity to HVS ORFs 28 and 45, are named for their similarity to the corresponding KSHV proteins. Note also that HVS ORF 73, which is closer in size to RRV ORF 73 than is ORF 73 of KSHV, also has very low similarity to RRV ORF 73, with a Gap value of 29.0%. A small number of RRV ORFs are more similar to HVS ORFs than to their KSHV counterparts, but the differences in similarity are small, with the exception of ORF 2, DHFR. RRV DHFR is 55.1% similar to the KSHV DHFR gene but 65.6% similar to the HVS DHFR gene. As noted above, RRV DHFR is also located similarly to the same gene in HVS but not KSHV.
Comparison of ORFs unique to RRV and KSHV.
RRV codes for a number of genes not found in HVS but present in KSHV. The shared presence of these unique genes in RRV and KSHV is evidence of the close similarity of these two viruses.
In DL-A, R1 colocalizes with, but has no similarity to, K1, a KSHV gene that has been demonstrated to have in vivo transforming ability (27). K1 and R1 both colocalize with HVS ORF1, or STP (saimiri transforming protein) (11), although both K1 and R1 are in opposite orientations compared to STP and have no sequence similarity to STP. A BLASTP search of GenBank with R1 reveals a limited amino-terminal similarity to a series of Fc receptors, including a potential transmembrane domain. These data suggest that R1, like K1 and STP, has transforming potential via transmembrane signaling.
In the region between ORFs 11 and 16, DL-B, both viruses possess ORFs for vIL-6, TS, and vMIP, although the number of MIP genes varies between the viruses. As mentioned above, TS is also found in HVS, but the location of TS in HVS is different than in KSHV and RRV. R2 has functional homology to K2, the vIL-6 gene of KSHV. Gap analysis of the vIL-6 genes from KSHV and RRV shows no notable similarity, but both possess four conserved cysteines found in cellular IL-6 (61). In addition, RRV vIL-6 has IL-6-like activity in cell culture (23). R3 has a low, but clear, similarity to KSHV K4, a vMIP1β gene. R3 is the only vMIP gene in RRV, compared to the three vMIP genes found in KSHV. In addition to these shared genes and the above-mentioned DHFR gene, the cytokine region of KSHV also contains two zinc finger proteins which are absent from the genome of RRV.
vIRFs, unique features for RRV and KSHV and among the most significant similarities between the two viruses, exist between ORFs 57 and 58 in DL-D3. The numbers of vIRFs differ between the two viruses. KSHV has four vIRFs, one of which, K10.5, is much smaller than the others. RRV has eight vIRFs, one of which, R9, is also notably smaller than the others. The RRV vIRFs are discussed in greater detail below.
In the region between ORFs 69 and 75, DL-E, RRV and KSHV code for an NCAM Ox-2 homolog (R15 and K14, respectively) and all three of the primate rhadinoviruses have genes for vFLIP (ORF 71), cyclin (ORF 72), and an IL-8 receptor-like G protein-coupled receptor (ORF 74). Although KSHV vFLIP was designated a unique KSHV gene, K13 (37), recent analysis has indicated that the corresponding HVS ORF, ORF 71, is also related to the FLIP family (42). Because of the HVS similarity, we have chosen to refer to the RRV FLIP homolog as ORF 71. Other groups have used the ORF 71 designation for KSHV (42).
ORFs are also found in RRV in regions corresponding to the locations of K8 and K8.1 (46), in DL-D2, and K12, in DL-E. However, none of the proteins deduced from these potential ORFs are similar to the KSHV gene products, and we have no evidence of transcription products derived from these regions. An analysis of potential promoter and polyadenylation sites is inconclusive. Although we were inclined to designate these potential RRV ORFs unique genes with R numbers, the lack of supporting evidence weighed against this. In anticipation of further evidence, though, we have reserved the corresponding R assignments (R4, R5, and R14) from our tabulation for possible use following transcriptional analysis of these regions. The lack of an RRV product corresponding to K12, kaposin, is of particular note, since kaposin is ubiquitously expressed in KSHV-infected cells (54, 62).
Multiple sequence analysis.
A phylogenetic analysis of the relationship of KSHV to other herpesviruses has previously been done (38). To extend this analysis and to more rigorously determine the relationship between RRV and KSHV, several ORFs found in six sequenced gammaherpesviruses were examined by bootstrap analysis by the maximum parsimony method. The corresponding ORFs from human CMV were used as outgroups.
Alignments were performed with ClustalW (55). All alignments were visually examined to ensure adequacy (i.e., no sequences were offset from the primary alignment, and there were significant numbers of identities in columns throughout the alignment). The alignment for UDG is presented as an example (Fig. 4). For each aligned ORF, columns containing gaps in any row were manually deleted, as were the unaligned N-terminal and C-terminal ends, to allow tree construction based on substitution rates alone. The sizes of the aligned sequences following gap removal and the number of identities in each alignment are listed in the legend to Fig. 5.
FIG. 4.
ClustalW alignment of UDG. The deduced sequences of UDG from seven viruses (Table 3) were aligned with ClustalW, version 1.4, as implemented by MacVector, version 6.01. Blosum 30 was used as the scoring matrix for the pairwise alignment and for the multiple sequence alignment. The gap introduction penalty was 10, and the gap extension penalty was 0.1. Identities, shown in white on black, are defined as columns containing the same residue in all rows.
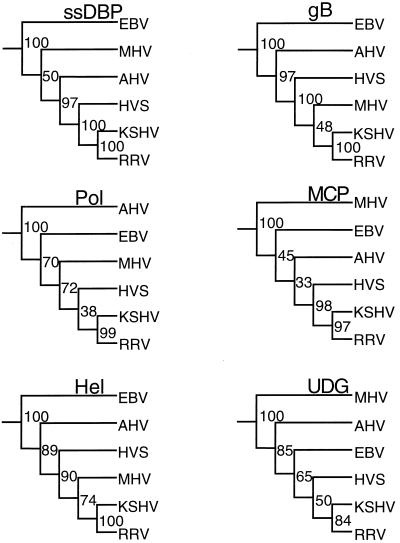
FIG. 5.
Phylogenetic trees for gammaherpesvirus proteins. Six proteins were selected for phylogenetic analysis with the Phylip programs. Proteins were aligned by ClustalW. Columns containing any gaps were removed. Unaligned N- and C-terminal residues were removed. The adjusted alignments were then used to generate 100 subsets with the program Seqboot. Trees were generated from each of these subsets with the program Protpars. A final consensus tree was generated with the program Consense. The numbers represent the number of Protpars trees, out of 100, which contained the same sequences to the right of the branch point as are found in the consensus tree. Proteins used, residues remaining after gaps were removed, and number of identities in the alignment are as follows: ssDBP, 1,072 residues, 117 identities; gB, 788 residues, 123 identities; Pol, 972 residues, 253 identities; MCP, 1,318 residues, 223 identities; Hel, 759 residues, 166 identities; UDG, 240 residues, 69 identities.
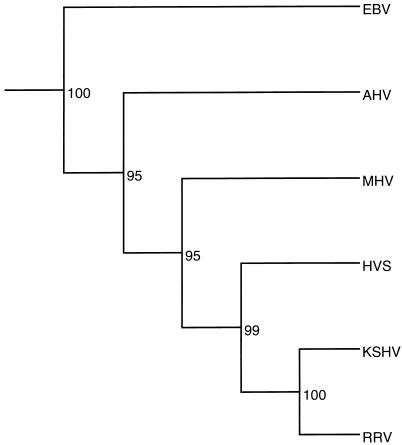
One hundred subsets were generated from each alignment by Seqboot, individual trees for each subset were generated by Protpars, and a final consensus tree was generated from the Protpars trees by using the program Consense. Trees for each of the ORFs are presented in Fig. 5. Of the six viruses examined, only RRV and KSHV always segregated on the same branch when individual proteins were examined. In addition to the individual analyses, the protein files were combined into a single concatenated file with 5,149 residues per virus, similar to the analysis of KSHV with a nine-gene set (38). This file was then analyzed by the bootstrapping procedure described above. The final tree shows that RRV and KSHV are closely related, with HVS as their nearest neighbor (Fig. 6). The close relationship of RRV and KSHV based on this phylogenetic analysis supports the similarity based on the organization of the two genomes and on the cumulative Gap analysis data.
FIG. 6.
Phylogenetic tree of the gammaherpesviruses. A concatenated file was made by merging all of the sequences used to generate the individual trees in Fig. 5. The concatenated file was then processed in the same fashion as described in the legend to Fig. 5. The outgroup CMV branch has been removed from the figure.
Analysis of RRV vIRFs.
KSHV and RRV possess a number of vIRF genes, all clustered in DL-D3. K9, the most studied of the KSHV vIRFs, does not have a DNA binding domain but has been demonstrated to inhibit the endogenous cellular interferon response pathways (18). Based on Gap analysis (Table 3), five of the RRV vIRFs (R6, R7, R8, R10, and R11) are similar to K9, though only R10 has a similarity greater than 30%. The remaining similarities fall between 26 and 30%. There is no measurable similarity between any RRV vIRF and any KSHV vIRF other than K9. There is, however, a pattern of higher similarity between members of the RRV vIRF family. R6, R7, R8, and R9 are most similar to R10, R11, R12, and R13, respectively, with the similarities falling between 50 and 62%, suggesting a gene amplification event in which an ancestral four-vIRF block was duplicated in toto to generate the eight vIRFs now found in RRV.
TABLE 3.
Comparison of vIRFs encoded by RRV 17577 and KSHV
| vIRF | % Similarity/% identitya:
|
||||||||
|---|---|---|---|---|---|---|---|---|---|
| KSHV K9 | RRV R6 | RRV R7 | RRV R8 | RRV R9 | RRV R10 | RRV R11 | RRV R12 | RRV R13 | |
| KSHV K9 | 100.000 | 26.044 | 28.291 | 28.857 | NA | 33.705 | 29.972 | NA | NA |
| 100.000 | 21.130 | 20.728 | 19.427 | 26.184 | 21.849 | ||||
| RRV R6 | 100.000 | NA | 26.393 | 29.918 | 54.427 | NA | NA | NA | |
| 100.000 | 19.062 | 22.131 | 47.917 | ||||||
| RRV R7 | 100.000 | 34.513 | NA | NA | 50.773 | 33.038 | NA | ||
| 100.000 | 26.254 | 41.495 | 24.484 | ||||||
| RRV R8 | 100.000 | NA | 31.412 | 35.693 | 61.254 | 28.818 | |||
| 100.000 | 24.207 | 23.849 | 50.997 | 21.037 | |||||
| RRV R9 | 100.000 | 28.980 | 28.216 | 30.364 | 58.103 | ||||
| 100.000 | 18.367 | 21.577 | 18.623 | 52.964 | |||||
| RRV R10 | 100.000 | 32.951 | 33.526 | NA | |||||
| 100.000 | 23.496 | 25.723 | |||||||
| RRV R11 | 100.000 | 33.923 | 31.124 | ||||||
| 100.000 | 23.849 | 25.072 | |||||||
| RRV R12 | 100.000 | NA | |||||||
| 100.000 | |||||||||
| RRV R13 | 100.000 | ||||||||
| 100.000 | |||||||||
NA, not applicable.
Similarity between KSHV vIRF K9 and two human interferon response factors, hICSBP and hISGF3γ, was demonstrated by ClustalW alignment of the three sequences (35). RRV vIRFs were individually aligned with hICSBP and hISGF3γ to examine their potential relationship to these proteins. Of the RRV vIRFs, R10 showed the greatest similarity to hICSBP and hISGF3γ (Fig. 7). That R10, of the RRV vIRFs, showed the greatest similarity to human IRFs is consistent with the Gap data, which showed greater similarity between K9 and R10 than between K9 and any of the remaining RRV vIRFs.
FIG. 7.
Comparison of RRV vIRF R10 to two human IRFs. RRV vIRF R10 was aligned to human ICSBP (59) and ISGF3γ (57). Identities between R10 and either of the human sequences are highlighted in white on black. The conserved tryptophans found in the DNA binding domain of mammalian IRFs are marked by asterisks.
The relationships among the RRV and KSHV vIRFs were further examined by multiple sequence alignment. KSHV K10 and K11 could not be accommodated by the ClustalW alignment; the use of either of these KSHV vIRFs significantly reduced the number of identities and similarities aligned by the program. However, K9 aligned well with the RRV vIRFs, although R9, which is significantly shorter than the remaining vIRFs, did not align with the amino-terminal segments of the other vIRFs (Fig. 8). The similarities among the vIRFs are generally clustered on either side of the extensively gapped central segment of the alignment. Bootstrap analysis with this alignment supports the Gap data in showing the relatedness of R6, R7, R8, and R9 to R10, R11, R12, and R13, respectively, and in showing that K9 is most closely related to R6 and R10 (data not shown).
FIG. 8.
Alignment of RRV vIRFs and KSHV K9. All eight RRV vIRFs and KSHV K9 were aligned by ClustalW. Identities and similarities are highlighted in white on black. Similarities are ranked as having the same residue in greater than half of the rows in a column. Tryptophans found in R8 and R12 that correspond to conserved tryptophans in mammalian IRFs are marked with asterisks.
The vIRF alignment shows that there is no overall conservation of the five tryptophan residues found in the DNA binding domain of mammalian IRFs (14). This lack of conserved tryptophans suggests that, like K9 (18), most RRV vIRFs possess no DNA binding activity. However, R8 and R12 each possess three tryptophans which align with the first three conserved tryptophans in mammalian IRFs and a fourth which is slightly out of alignment with the fifth conserved tryptophan of mammalian IRFs (Fig. 7). Whether this degree of conservation is sufficient to allow R8 and R12 to bind DNA similarly to mammalian IRFs has not been determined.
Colocalization of repeat units in RRV and KSHV.
The RRV genome contains three highly repetitive regions, which correspond to three of the repetitive regions of KSHV (Table 4). All three of the RRV repetitive regions occur in divergent loci and have been named for the locus in which they appear. The two tandem repeats, rDL-B 1 and 2 (for “repeat in divergent locus B” 1 and 2) and rDL-E 1 and 2, correspond to KSHV tandem repeats frnk and zppa, respectively. The RRV short single repeat rDL-F corresponds to the longer, but similarly structured, mdsk repeat of KSHV. The rDL-B 2 repeat is a long (∼1 kb), high G+C (>79.9%) segment that could not be sequenced in a single reaction. Instead, rDL-B 2 was sequenced from either end with custom primers. The sum of the bases derived from these reactions was greater than the length of the repeat element based upon rigorous restriction mapping. Thus, we were able to sequence the complete element but only on a single strand. Because this element was deleted when isolated from its surrounding DNA, we were also unable to reliably use exonuclease III deletion to determine the sequence of the second strand.
TABLE 4.
Comparison of corresponding repeats in RRV and KSHV
| Virus | Insert namea | Length (bp)
|
G+C content (%) | G+A content (%)b | |
|---|---|---|---|---|---|
| Total | Repeat unit | ||||
| KSHV | frnk | 332 | 20 | 80.1 | |
| 292 | 30 | 84.9 | |||
| RRV | rDL-B 1 | 304 | 26 | 53.3 | |
| rDL-B 2 | 1,008 | 25 | 79.9 | ||
| KSHV | zppa | 308 | 23 | 74.0 | |
| 244 | 23 | 77.9 | |||
| RRV | rDL-E 1 | 405 | 19 | 74.6 | |
| rDL-E 2 | 1,029 | 32 | 84.4 | ||
| KSHV | mdsk | 409 | —c | 75.4 | |
| RRV | rDL-F | 196 | 13 | 81.6 | |
KSHV frnk and zppa and RRV rDL-B and rDL-E are tandem repeats.
G+A content refers to the unusually high number of G’s and A’s in the viral strand which reads left to right.
—, KSHV mdsk is a complex repeat with no defined unit length.
The second high G+C repeat, rDL-E 1 and 2, is part of the 3.9-kb BamHI fragment (BamHI fragment 12) predicted by the sequence data but not present in the genomic digest. To determine the reason for the differences between the predicted and the actual patterns, we digested three independent cosmids that span this region with BamHI. Two, cosmids 8 and 16, produce the 3.9-kb fragment. A third cosmid, cosmid 28b, originally exhibited a 5.4-kb product, but upon repeat culturing in bacteria it produced a faint 3.9-kb product and a much more visible 3.4-kb product, with smearing that suggested a heterologous mix of sizes within this region (data not shown). Southern blot analysis of RRV genomic DNA identifies the BamHI fragment bearing rDL-E 1 and 2 as a smear between 6.0 and 3.4 kb (data not shown). This corresponds to the smear between 6 and 3.4 kb observed by ethidium bromide in the BamHI genomic digest (Fig. 2). Therefore, it is likely that the heterogeneity observed in the cosmids reflects heterogeneity in the virus as well. Sequencing of the various cosmids shows that the DNA flanking rDL-E 1 and 2 is the same in all cosmids examined, so the likely source of the variability found in the BamHI fragment is rDL-E 1 and 2. This heterogeneity is not apparent in the EcoRI and HindIII digests, possibly because the EcoRI and HindIII fragments containing rDL-E 1 and 2 are significantly larger than BamHI fragment 12, thereby obscuring the size differences.
At the right end of the genome, in DL-F, KSHV has the mdsk repeat, the plus strand of which has a high G+A content. A similar though shorter repeat, rDL-F, is found in the RRV genome. It also occurs to the right of all identified ORFs and close to the right terminal repeat.
Not all repeat elements found in KSHV have corresponding repeats in RRV. This includes the KSHV vnct, waka/jwka, and moi repeats. The moi repeat is located in the center of the KSHV ORF 73 and is responsible for the divergent lengths of RRV and KSHV ORF 73. moi is described in the annotations to the KSHV GenBank entry as having 15 different 11- to 16-bp repeats. The result of this repeat element is the presence in ORF 73 of a highly acidic central domain, with a large number of glutamate residues encoded by a repeating GAG codon. KSHV ORF 73 is a potential leucine zipper protein, with a number of leucine zipper sites in the repeat region. RRV lacks the moi repeat and its concomitant acidic domain. It also lacks any evidence for a leucine zipper, suggesting that the biology of ORF 73 in RRV is substantially different than the biology of ORF 73 in KSHV.
Terminal repeats.
We have examined the structure of the terminal repeats and have included references to them in Fig. 1, although we have only sequenced segments of them. We have determined that each intact terminal repeat unit contains a single BglII site. BglII restriction digests of cosmid inserts containing the right terminal repeat demonstrate that the size of the terminal repeat unit is about 2,100 bp. We have isolated two independent 2.1-kb sequences. One possesses intact flanking BglII sites. The second possess a single BglII site on the left side and a corrupted BglII site on the right side. This corrupted BglII site corresponds to the Sau3AI site used to create the parent cosmid.
An 800-bp ApaI/BglII fragment has been identified and sequenced on the right end of each of these fragments. The same 800-bp sequence has been identified at the end of a 5-kb BglII fragment containing the right end of the LUR. It has also been identified as flanking the left end of the LUR, although it terminates 30 bases to the left of the expected BglII site and segues into the unique region of the genome. A Sau3AI site at the left end of this left terminal repeat sequence was used to generate the only cosmid we have isolated containing the left end of the genome. The sequence to the left of the ApaI/BglII fragment has a high G+C content and appears to have a complex repeat structure, which, like the elements rDL-B and rDL-E, has been resistant to sequencing. To date we have not isolated any cosmids which contain the terminal repeats flanked on both sides by sequence from the LUR, as was done with KSHV (26), so we cannot define the full length of the terminal repeat structure of RRV.
DISCUSSION
We have sequenced the entire LUR of the genome of an RRV, strain 17577. Comparisons of genomic organization, ORF structure, phylogenetic analysis, and the positioning of repetitive elements strongly support our conclusion that RRV is a rhesus macaque equivalent to KSHV. In addition to these criteria, the presence of genes for vIL-6, vMIP, and multiple vIRFs, genes found in no other herpesviruses, also supports this conclusion.
The criteria for classification of KSHV and RRV as rhadinoviruses are their similarity to HVS, the prototypical primate rhadinovirus, and their lymphotropism. KSHV and RRV, however, have characteristics unlike other rhadinoviruses. KSHV and RRV do not have the low G+C content characteristic of rhadinoviruses. Also, unlike other rhadinoviruses, RRV does not display CpG suppression, whereas KSHV does, although to a lesser extent than other rhadinoviruses. Rhadinoviruses in general have a reduced CpG content, with increased amounts of TpG and CpA (21). This is the result of methylation of the C in the dinucleotide, followed by mutation to T (CpA is the opposite-strand complement of TpG). Suppression of CpG is measured as a ratio of observed to expected. For human EBV, this ratio is 0.60; for HVS, the ratio is 0.33; for MHV, the ratio is 0.43; and for KSHV, the ratio is 0.81. The CpG ratio of RRV is 1.11, which indicates that CpG suppression does not alter the base composition of this virus. CpG suppression is believed to be an indicator of latent state methylation in lymphoid cells, so the lack of CpG suppression in RRV may indicate that its latent state is somewhat different than that of KSHV and other rhadinoviruses.
A rhesus herpesvirus with similarity to KSHV, RRV H26-95, was previously reported (10). This previous report described 10,595 bases of the virus. This segment of H26-95 is greater than 99% identical at the DNA level to the corresponding region of RRV 17577. Based upon this similarity, we conclude that H26-95 and 17577 are the same virus. However, there is evidence, in the form of variations in the structure of gB, that they represent different strains. This strain variation may have important implications. RRV strains may be preferentially associated with certain types of illness, much as KSHV strain A appears to be more closely associated with AIDS-associated body cavity-based lymphoma, while strains B and C tend to be associated with AIDS- and non-AIDS-associated LPD (30).
No disease association has been reported for H26-95. In contrast, RRV 17577 is linked to SIV-associated LPD (61). These potential differences in pathogenesis may be associated with the variations in RRV gB. This would be consistent with studies on the variability of gB in CMV. Variations in CMV gB alter its immunological characteristics and may alter the virus’s ability to infect cells (15, 50). Major loci for CMV gB variation occur between codons 27 and 67 and between codons 448 and 480 (8). Minor loci of variation occur at codons 181 to 195, 312 to 316, and 387 to 408. In the multiple sequence analysis of gB (data not shown), the extended region of RRV gB variability (residues 274 to 534) aligns with CMV sequence from 305 to 572. The most variable region of RRV gB (residues 346 to 440) aligns with CMV codons 381 to 477, thereby straddling one minor locus and one major locus of CMV gB variability. This similarity between RRV and CMV gB variability suggests that RRV gB variability also alters its immunogenicity and possibly its ability to target cells. It is quite possible that the difference in pathogenic potential between the two strains is a factor of currently unknown differences, since the segment of H26-95 that has been reported accounts for less than 10% of the complete virus. However, it is interesting that the only known differences between H26-95 and 17577 are differences that may alter host cell targeting and, therefore, alter the virus’s ability to induce disease.
Although RRV 17577 and H26-95 ORFs are extremely similar, the range of similarity between RRV and KSHV ORFs is broad. The most notable difference between KSHV and RRV ORFs, however, is not in the range of similarity but, instead, in the variation in the size of ORF 73, the latency associated nuclear antigen. The low similarity for ORF 73 (23.6%) may be misleading because the Gap analysis overlooks the difference in the sizes of the two proteins. ORF 73 from KSHV contains 1,162 residues, while ORF 73 from RRV contains only 447. The difference in size between RRV and KSHV ORF 73 is not unique; ORF 73 is variable in size across the range of gammaherpesviruses. ORF 73 from HVS (1), MHV (44), and AHV (11) contains 407, 314, and 1,300 residues, respectively. Both KSHV and AHV have centrally located, highly repetitive, acidic domains that are not present in the smaller proteins. These acidic domains also dominate the comparisons of the proteins. If the carboxyterminal 300 residues of KSHV ORF 73, which do not include any portion of the acidic domain, are compared to RRV ORF 73, the similarity is 44.3%, indicating a moderately conserved region outside the KSHV acidic domain.
We have been unable to find RRV ORFs corresponding to K8, K8.1 (46), or K12. K8.1 is a highly immunogenic glycoprotein that has been demonstrated to be a useful marker for epidemological studies of KSHV. Although there are ORFs in this region of RRV, they have no similarity to K8 or K8.1 and they lack any of the necessary transcriptional control elements to suggest that they are transcribed. K12 is a gene for kaposin, a transcript ubiquitous in KSHV-infected cells (54, 62). In addition to being ubiquitous, kaposin is a transforming gene in vitro (38). RRV codes for no protein with similarity to K12. We have examined all possible translations of the region between ORF 69 and the nearby tandem repeats and have found no similarity to K12. ORFs similar to K12 have also not been found elsewhere in the RRV genome. The ubiquitous expression of K12 and its transforming potential suggests that it is important in the involvement of KSHV in disease, and so the lack of an RRV counterpart for this gene has important implications for the comparability of the two viruses. It is possible that RRV encodes a protein of similar function but with no notable similarity to kaposin, much as the RRV vIL-6 lacks significant sequence similarity to cellular or KSHV vIL-6. Transcriptional analysis of RRV will be necessary to determine if it codes for genes corresponding to KSHV K8, K8.1, and K12.
Kaposin is one of several KSHV ORFs that have been implicated as transforming genes. However, the transforming capability of most of them has been determined only in vitro. The exception to this is K1, which colocalizes with HVS ORF 1, which is also called saimiri transforming protein (STP). Both K1 (27) and STP (11) are demonstrated transforming genes in vivo. K1 inserted into the genome of an STP-negative strain of HVS restores the virus’s ability to transform T cells in marmosets. RRV ORF R1 colocalizes with K1 and STP. By Gap analysis, R1 is not similar to K1 or STP. BLASTP searches of GenBank with R1 and K1 show that the N terminus of each is similar to Fc receptors, suggesting that R1 is a transmembrane receptor and therefore may be involved in signal transduction and transformation. If R1 is a transforming gene, then the identification of a third rhadinovirus with a transforming gene as its first ORF would clearly designate this region an important locus for viral pathogenesis.
The presence of potential transforming genes does not in itself prove transforming potential for the virus. Direct evidence of virus-mediated transformation is necessary for this. Experimental evidence demonstrates such an association between RRV 17577 and SIV-associated LPD (61), suggesting that this is a good model for AIDS-related Castleman’s disease. If RRV is responsible for LPD in SIV-infected rhesus macaques, then RRV clearly will be important in understanding the role of primate rhadinoviruses in lymphoproliferative disease in general. However, RRV may not be adequate as a model for KSHV involvement in KS. The ideal model system would be an RRV-associated rhesus KS. Such a model does not currently exist, since KS has never been identified in macaques. However, type D simian retrovirus type 2-infected pigtail and rhesus macaques have been shown to develop RF, a disease affecting different tissues than KS but with similar morphology (19, 56). DNA sequences extremely similar to KSHV DNA polymerase have been obtained by nested PCR from RF tissue (5, 49), suggesting that there is a rhadinovirus involved in RF development. Only a careful analysis of the biology of RF development and manner of RRV involvement in RF will determine if this disease can serve as a functional nonhuman model for the virology behind KS.
The presence of a virus in nonhuman primates that is similar to KSHV and that causes an illness similar to an LPD associated with KSHV (i.e., Castleman’s disease) suggests that RRV will be a useful model for some aspects of KSHV pathogenesis. The study of RRV LPD associations is promoted by the ease with which RRV can be propagated in cultured cells. This allows molecular dissection of genes postulated to be related to virus-mediated pathogenesis and, therefore, allows us to identify viral determinants of pathogenesis. We are currently using these features of RRV to determine the role of the cytokine-like proteins and other viral ORFs in RRV-induced LPD.
ACKNOWLEDGMENTS
This work was supported by NIH grants RR00163 (M.K.A. and S.W.W.) and CA75922 (S.W.W.).
We thank Gary S. Hayward, Diane LoPiccolo, and Johnan Kaleeba for helpful discussions; Felix Lee and Caroline Hettrick for technical assistance; Yibing Jia and Kalama Taylor of the Oregon Regional Primate Research Center Molecular Biology Core for performing sequencing reactions; and Lori Boshears for expert editorial assistance.
REFERENCES
- 1.Albrecht J C, Nicholas J, Biller D, Cameron K R, Biesinger B, Newman C, Wittmann S, Craxton M A, Coleman H, Fleckenstein B, Honess R W. Primary structure of the herpesvirus saimiri genome. J Virol. 1992;66:5047–5058. doi: 10.1128/jvi.66.8.5047-5058.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ambroziak J A, Blackbourn D J, Herndier B G, Glogau R G, Gullett J H, McDonald A R, Lennette E T, Levy J A. Herpes-like sequences in HIV-infected and uninfected Kaposi’s sarcoma patients. Science. 1995;268:582–583. doi: 10.1126/science.7725108. [DOI] [PubMed] [Google Scholar]
- 3.Arvanitakis L, Geras-Raaka E, Varma A, Gershengorn M C, Cesarman E. Human herpesvirus KSHV encodes a constitutively active G-protein-coupled receptor linked to cell proliferation. Nature. 1997;385:347–350. doi: 10.1038/385347a0. [DOI] [PubMed] [Google Scholar]
- 4.Bais C, Santomasso B, Coso O, Arvanitakis L, Raaka E G, Gutkind J S, Asch A S, Cesarman E, Gershengorn M C, Mesri E A, Gerhengorn M C. G-protein-coupled receptor of Kaposi’s sarcoma-associated herpesvirus is a viral oncogene and angiogenesis activator. Nature. 1998;391:86–89. doi: 10.1038/34193. [DOI] [PubMed] [Google Scholar]
- 5.Bosch M L, Strand K B, Rose T M. Gammaherpesvirus sequence comparisons. J Virol. 1998;72:8458–8459. doi: 10.1128/jvi.72.10.8458-8459.1998. . (Letter.) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Cesarman E, Nador R G, Bai F, Bohenzky R A, Russo J J, Moore P S, Chang Y, Knowles D M. Kaposi’s sarcoma-associated herpesvirus contains G protein-coupled receptor and cyclin D homologs which are expressed in Kaposi’s sarcoma and malignant lymphoma. J Virol. 1996;70:8218–8223. doi: 10.1128/jvi.70.11.8218-8223.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Chang Y, Cesarman E, Pessin M S, Lee F, Culpepper J, Knowles D M, Moore P S. Identification of herpesvirus-like DNA sequences in AIDS-associated Kaposi’s sarcoma. Science. 1994;266:1865–1869. doi: 10.1126/science.7997879. [DOI] [PubMed] [Google Scholar]
- 8.Chou S. Comparative analysis of sequence variation in gp116 and gp55 components of glycoprotein B of human cytomegalovirus. Virology. 1992;188:388–390. doi: 10.1016/0042-6822(92)90771-g. [DOI] [PubMed] [Google Scholar]
- 9.Delabesse E, Oksenhendler E, Lebbe C, Verola O, Varet B, Turhan A G. Molecular analysis of clonality in Kaposi’s sarcoma. J Clin Pathol. 1997;50:664–668. doi: 10.1136/jcp.50.8.664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Desrosiers R C, Sasseville V G, Czajak S C, Zhang X, Mansfield K G, Kaur A, Johnson R P, Lackner A A, Jung J U. A herpesvirus of rhesus monkeys related to the human Kaposi’s sarcoma-associated herpesvirus. J Virol. 1997;71:9764–9769. doi: 10.1128/jvi.71.12.9764-9769.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Duboise S M, Guo J, Czajak S, Desrosiers R C, Jung J U. STP and Tip are essential for herpesvirus saimiri oncogenicity. J Virol. 1998;72:1308–1313. doi: 10.1128/jvi.72.2.1308-1313.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Dupin N, Grandadam M, Calvez V, Gorin I, Aubin J T, Havard S, Lamy F, Leibowitch M, Huraux J M, Escande J P, Agut H. Herpesvirus-like DNA sequences in patients with Mediterranean Kaposi’s sarcoma. Lancet. 1995;345:761–762. doi: 10.1016/s0140-6736(95)90642-8. [DOI] [PubMed] [Google Scholar]
- 13.Ensser A, Pflanz R, Fleckenstein B. Primary structure of the alcelaphine herpesvirus 1 genome. J Virol. 1997;71:6517–6525. doi: 10.1128/jvi.71.9.6517-6525.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Escalante C R, Yie J, Thanos D, Aggarwal A K. Structure of IRF-1 with bound DNA reveals determinants of interferon regulation. Nature. 1998;391:103–106. doi: 10.1038/34224. [DOI] [PubMed] [Google Scholar]
- 15.Fries B C, Chou S, Boeckh M, Torok-Storb B. Frequency distribution of cytomegalovirus envelope glycoprotein genotypes in bone marrow transplant recipients. J Infect Dis. 1994;169:769–774. doi: 10.1093/infdis/169.4.769. [DOI] [PubMed] [Google Scholar]
- 16.Gallo R C. The enigmas of Kaposi’s sarcoma. Science. 1998;282:1837–1839. doi: 10.1126/science.282.5395.1837. [DOI] [PubMed] [Google Scholar]
- 17.Gallo R C. Some aspects of the pathogenesis of HIV-1-associated Kaposi’s sarcoma. Monogr Natl Cancer Inst. 1998;23:55–57. doi: 10.1093/oxfordjournals.jncimonographs.a024174. [DOI] [PubMed] [Google Scholar]
- 18.Gao S J, Boshoff C, Jayachandra S, Weiss R A, Chang Y, Moore P S. KSHV ORF K9 (vIRF) is an oncogene which inhibits the interferon signaling pathway. Oncogene. 1997;15:1979–1985. doi: 10.1038/sj.onc.1201571. [DOI] [PubMed] [Google Scholar]
- 19.Giddens W E, Jr, Tsai C C, Morton W R, Ochs H D, Knitter G H, Blakley G A. Retroperitoneal fibromatosis and acquired immunodeficiency syndrome in macaques. Pathologic observations and transmission studies. Am J Pathol. 1985;119:253–263. [PMC free article] [PubMed] [Google Scholar]
- 20.Guo H G, Browning P, Nicholas J, Hayward G S, Tschachler E, Jiang Y W, Sadowska M, Raffeld M, Colombini S, Gallo R C, Reitz M S., Jr Characterization of a chemokine receptor-related gene in human herpesvirus 8 and its expression in Kaposi’s sarcoma. Virology. 1997;228:371–378. doi: 10.1006/viro.1996.8386. [DOI] [PubMed] [Google Scholar]
- 21.Honess R W, Gompels U A, Barrell B G, Craxton M, Cameron K R, Staden R, Chang Y N, Hayward G S. Deviations from expected frequencies of CpG dinucleotides in herpesvirus DNAs may be diagnostic of differences in the states of their latent genomes. J Gen Virol. 1989;70:837–855. doi: 10.1099/0022-1317-70-4-837. [DOI] [PubMed] [Google Scholar]
- 22.Kaaya E E, Parravicini C, Ordonez C, Gendelman R, Berti E, Gallo R C, Biberfeld P. Heterogeneity of spindle cells in Kaposi’s sarcoma: comparison of cells in lesions and in culture. J Acquired Immune Defic Syndr Hum Retrovirol. 1995;10:295–305. [PubMed] [Google Scholar]
- 23.Kaleeba, J. A., E. P. Bergquam, and S. W. Wong. A rhesus macaque rhadinovirus related to Kaposi’s sarcoma-associated herpesvirus (KSHV) encodes a functional homologue of interleukin-6. Unpublished data. [DOI] [PMC free article] [PubMed]
- 24.Kasolo F C, Mpabalwani E, Gompels U A. Infection with AIDS-related herpesviruses in human immunodeficiency virus-negative infants and endemic childhood Kaposi’s sarcoma in Africa. J Gen Virol. 1997;78:847–855. doi: 10.1099/0022-1317-78-4-847. [DOI] [PubMed] [Google Scholar]
- 25.Kedes D H, Operskalski E, Busch M, Kohn R, Flood J, Ganem D. The seroepidemiology of human herpesvirus 8 (Kaposi’s sarcoma-associated herpesvirus): distribution of infection in KS risk groups and evidence for sexual transmission. Nat Med. 1996;2:918–924. doi: 10.1038/nm0896-918. [DOI] [PubMed] [Google Scholar]
- 26.Lagunoff M, Ganem D. The structure and coding organization of the genomic termini of Kaposi’s sarcoma-associated herpesvirus. Virology. 1997;236:147–154. doi: 10.1006/viro.1997.8713. [DOI] [PubMed] [Google Scholar]
- 27.Lee H, Veazey R, Williams K, Li M, Guo J, Neipel F, Fleckenstein B, Lackner A, Desrosiers R C, Jung J U. Deregulation of cell growth by the K1 gene of Kaposi’s sarcoma-associated herpesvirus. Nat Med. 1998;4:435–440. doi: 10.1038/nm0498-435. [DOI] [PubMed] [Google Scholar]
- 28.Li M, Lee H, Guo J, Neipel F, Fleckenstein B, Ozato K, Jung J U. Kaposi’s sarcoma-associated herpesvirus viral interferon regulatory factor. J Virol. 1998;72:5433–5440. doi: 10.1128/jvi.72.7.5433-5440.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Luppi M, Barozzi P, Maiorana A, Artusi T, Trovato R, Marasca R, Savarino M, Ceccherini-Nelli L, Torelli G. Human herpesvirus-8 DNA sequences in human immunodeficiency virus-negative angioimmunoblastic lymphadenopathy and benign lymphadenopathy with giant germinal center hyperplasia and increased vascularity. Blood. 1996;87:3903–3909. [PubMed] [Google Scholar]
- 30.Luppi M, Barozzi P, Marasca R, Ferrari M G, Torelli G. Human herpesvirus 8 strain variability in clinical conditions other than Kaposi’s sarcoma. J Virol. 1997;71:8082–8083. doi: 10.1128/jvi.71.10.8082-8083.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Martin J N, Ganem D E, Osmond D H, Page-Shafer K A, Macrae D, Kedes D H. Sexual transmission and the natural history of human herpesvirus 8 infection. N Engl J Med. 1998;338:948–954. doi: 10.1056/NEJM199804023381403. [DOI] [PubMed] [Google Scholar]
- 32.McGeoch D J, Cook S. Molecular phylogeny of the alphaherpesvirinae subfamily and a proposed evolutionary timescale. J Mol Biol. 1994;238:9–22. doi: 10.1006/jmbi.1994.1264. [DOI] [PubMed] [Google Scholar]
- 33.McGeoch D J, Cook S, Dolan A, Jamieson F E, Telford E A. Molecular phylogeny and evolutionary timescale for the family of mammalian herpesviruses. J Mol Biol. 1995;247:443–458. doi: 10.1006/jmbi.1995.0152. [DOI] [PubMed] [Google Scholar]
- 34.Molden J, Chang Y, You Y, Moore P S, Goldsmith M A. A Kaposi’s sarcoma-associated herpesvirus-encoded cytokine homolog (vIL-6) activates signaling through the shared gp130 receptor subunit. J Biol Chem. 1997;272:19625–19631. doi: 10.1074/jbc.272.31.19625. [DOI] [PubMed] [Google Scholar]
- 35.Moore P S, Boshoff C, Weiss R A, Chang Y. Molecular mimicry of human cytokine and cytokine response pathway genes by KSHV. Science. 1996;274:1739–1744. doi: 10.1126/science.274.5293.1739. [DOI] [PubMed] [Google Scholar]
- 36.Moore P S, Chang Y. Detection of herpesvirus-like DNA sequences in Kaposi’s sarcoma in patients with and without HIV infection. N Engl J Med. 1995;332:1181–1185. doi: 10.1056/NEJM199505043321801. [DOI] [PubMed] [Google Scholar]
- 37.Moore P S, Chang Y. Kaposi’s sarcoma-associated herpesvirus-encoded oncogenes and oncogenesis. Monogr Natl Cancer Inst. 1998;23:65–71. doi: 10.1093/oxfordjournals.jncimonographs.a024176. [DOI] [PubMed] [Google Scholar]
- 38.Moore P S, Gao S J, Dominguez G, Cesarman E, Lungu O, Knowles D M, Garber R, Pellett P E, McGeoch D J, Chang Y. Primary characterization of a herpesvirus agent associated with Kaposi’s sarcoma. J Virol. 1996;70:549–558. doi: 10.1128/jvi.70.1.549-558.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Muralidhar S, Pumfery A M, Hassani M, Sadale M R, Azumi N, Kishishita M, Brady J N, Doniger J, Medveczky P, Rosenthal L J. Identification of kaposin (open reading frame K12) as a human herpesvirus 8 (Kaposi’s sarcoma-associated herpesvirus) transforming gene. J Virol. 1998;72:4980–4988. doi: 10.1128/jvi.72.6.4980-4988.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Nador R G, Cesarman E, Chadburn A, Dawson D B, Ansari M Q, Sald J, Knowles D M. Primary effusion lymphoma: a distinct clinicopathologic entity associated with the Kaposi’s sarcoma-associated herpes virus. Blood. 1996;88:645–656. [PubMed] [Google Scholar]
- 41.Neipel F, Albrecht J C, Ensser A, Huang Y Q, Li J J, Friedman-Kien A E, Fleckenstein B. Human herpesvirus 8 encodes a homolog of interleukin-6. J Virol. 1997;71:839–842. doi: 10.1128/jvi.71.1.839-842.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Neipel F, Albrecht J-C, Fleckenstein B. Human herpesvirus 8—the first human rhadinovirus. Monogr Natl Cancer Inst. 1998;23:73–77. doi: 10.1093/oxfordjournals.jncimonographs.a024178. [DOI] [PubMed] [Google Scholar]
- 43.Nicholas J, Ruvolo V R, Burns W H, Sandford G, Wan X, Ciufo D, Hendrickson S B, Guo H G, Hayward G S, Reitz M S. Kaposi’s sarcoma-associated human herpesvirus-8 encodes homologues of macrophage inflammatory protein-1 and interleukin-6. Nat Med. 1997;3:287–292. doi: 10.1038/nm0397-287. [DOI] [PubMed] [Google Scholar]
- 44.Nicholas J, Zong J-C, Alcendor D J, Ciufo D M, Poole L J, Sarisky R T, Chiou C-J, Zhang X, Wan X, Guo H-G, Reitz M S, Hayward G S. Novel organizational features, captured cellular genes, and strain variability within the genome of KSHV/HHV8. Monogr Natl Cancer Inst. 1998;23:79–88. doi: 10.1093/oxfordjournals.jncimonographs.a024179. [DOI] [PubMed] [Google Scholar]
- 45.Page R D. TreeView: an application to display phylogenetic trees on personal computers. Comput Appl Biosci. 1996;12:357–358. doi: 10.1093/bioinformatics/12.4.357. [DOI] [PubMed] [Google Scholar]
- 46.Raab M-S, Albrecht J-C, Birkmann A, Yaguboglu S, Lang D, Fleckenstein B, Neipel F. The immunogenic glycoprotein gp35-37 of human herpesvirus 8 is encoded by open reading frame K8.1. J Virol. 1998;72:6725–6731. doi: 10.1128/jvi.72.8.6725-6731.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Rabkin C S, Janz S, Lash A, Coleman A E, Musaba E, Liotta L, Biggar R J, Zhuang Z. Monoclonal origin of multicentric Kaposi’s sarcoma lesions. N Engl J Med. 1997;336:988–993. doi: 10.1056/NEJM199704033361403. [DOI] [PubMed] [Google Scholar]
- 48.Renne R, Zhong W, Herndier B, McGrath M, Abbey N, Kedes D, Ganem D. Lytic growth of Kaposi’s sarcoma-associated herpesvirus (human herpesvirus 8) in culture. Nat Med. 1996;2:342–346. doi: 10.1038/nm0396-342. [DOI] [PubMed] [Google Scholar]
- 49.Rose T M, Strand K B, Schultz E R, Schaefer G, Rankin G W, Jr, Thouless M E, Tsai C C, Bosch M L. Identification of two homologs of the Kaposi’s sarcoma-associated herpesvirus (human herpesvirus 8) in retroperitoneal fibromatosis of different macaque species. J Virol. 1997;71:4138–4144. doi: 10.1128/jvi.71.5.4138-4144.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Roy D M, Grundy J E, Emery V C. Sequence variation within neutralizing epitopes of the envelope glycoprotein B of human cytomegalovirus: comparison of isolates from renal transplant recipients and AIDS patients. J Gen Virol. 1993;74:2499–2505. doi: 10.1099/0022-1317-74-11-2499. [DOI] [PubMed] [Google Scholar]
- 51.Russo J J, Bohenzky R A, Chien M C, Chen J, Yan M, Maddalena D, Parry J P, Peruzzi D, Edelman I S, Chang Y, Moore P S. Nucleotide sequence of the Kaposi sarcoma-associated herpesvirus (HHV8) Proc Natl Acad Sci USA. 1996;93:14862–14867. doi: 10.1073/pnas.93.25.14862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Schalling M, Ekman M, Kaaya E E, Linde A, Biberfeld P. A role for a new herpes virus (KSHV) in different forms of Kaposi’s sarcoma. Nat Med. 1995;1:707–708. doi: 10.1038/nm0795-707. [DOI] [PubMed] [Google Scholar]
- 53.Soulier J, Grollet L, Oksenhendler E, Cacoub P, Cazals-Hatem D, Babinet P, d’Agay M F, Clauvel J P, Raphael M, Degos L, et al. Kaposi’s sarcoma-associated herpesvirus-like DNA sequences in multicentric Castleman’s disease. Blood. 1995;86:1276–1280. [PubMed] [Google Scholar]
- 54.Staskus K A, Zhong W, Gebhard K, Herndier B, Wang H, Renne R, Beneke J, Pudney J, Anderson D J, Ganem D, Haase A T. Kaposi’s sarcoma-associated herpesvirus gene expression in endothelial (spindle) tumor cells. J Virol. 1997;71:715–719. doi: 10.1128/jvi.71.1.715-719.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Thompson J D, Higgins D G, Gibson T J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994;22:4673–4680. doi: 10.1093/nar/22.22.4673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Tsai C C, Roodman S T, Woon M D. Mesenchymoproliferative disorders (MPD) in simian AIDS associated with SRV-2 infection. J Med Primatol. 1990;19:189–202. [PubMed] [Google Scholar]
- 57.Veals S A, Schindler C, Leonard D, Fu X Y, Aebersold R, Darnell J E, Jr, Levy D E. Subunit of an alpha-interferon-responsive transcription factor is related to interferon regulatory factor and Myb families of DNA-binding proteins. Mol Cell Biol. 1992;12:3315–3324. doi: 10.1128/mcb.12.8.3315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Vella J P, Mosher R, Sayegh M H. Kaposi’s sarcoma after renal transplantation. N Engl J Med. 1997;336:1761. doi: 10.1056/nejm199706123362416. [DOI] [PubMed] [Google Scholar]
- 59.Weisz A, Marx P, Sharf R, Appella E, Driggers P H, Ozato K, Levi B Z. Human interferon consensus sequence binding protein is a negative regulator of enhancer elements common to interferon-inducible genes. J Biol Chem. 1992;267:25589–25596. [PubMed] [Google Scholar]
- 60.Whitby D, Howard M R, Tenant-Flowers M, Brink N S, Copas A, Boshoff C, Hatzioannou T, Suggett F E, Aldam D M, Denton A S, Miller R F, Weller I V D, Weiss R A, Tedder R S, Schulz T F. Detection of Kaposi sarcoma associated herpesvirus in peripheral blood of HIV-infected individuals and progression to Kaposi’s sarcoma. Lancet. 1995;346:799–802. doi: 10.1016/s0140-6736(95)91619-9. [DOI] [PubMed] [Google Scholar]
- 61.Wong, S. W., E. P. Bergquam, R. M. Swanson, F. W. Lee, S. M. Shiigi, N. A. Avery, and M. K. Axthelm. Induction of B-cell hyperplasia in SIV-infected monkeys with the simian homologue of Kaposi’s sarcoma-associated herpesvirus. Submitted for publication. [DOI] [PMC free article] [PubMed]
- 62.Zhong W, Wang H, Herndier B, Ganem D. Restricted expression of Kaposi sarcoma-associated herpesvirus (human herpesvirus 8) genes in Kaposi sarcoma. Proc Natl Acad Sci USA. 1996;93:6641–6646. doi: 10.1073/pnas.93.13.6641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Zimring J C, Goodbourn S, Offermann M K. Human herpesvirus 8 encodes an interferon regulatory factor (IRF) homolog that represses IRF-1-mediated transcription. J Virol. 1998;72:701–707. doi: 10.1128/jvi.72.1.701-707.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]