Abstract
The roles of DNA methylation in invertebrates are poorly characterized, and critical data are missing for the phylum Annelida. We fill this knowledge gap by conducting the first genome-wide survey of DNA methylation in the deep-sea polychaetes dominant in deep-sea vents and seeps: Paraescarpia echinospica, Ridgeia piscesae, and Paralvinella palmiformis. DNA methylation calls were inferred from Oxford Nanopore sequencing after assembling high-quality genomes of these animals. The genomes of these worms encode all the key enzymes of the DNA methylation metabolism and possess a mosaic methylome similar to that of other invertebrates. Transcriptomic data of these polychaetes support the hypotheses that gene body methylation strengthens the expression of housekeeping genes and that promoter methylation acts as a silencing mechanism but not the hypothesis that DNA methylation suppresses the activity of transposable elements. The conserved epigenetic profiles of genes responsible for maintaining homeostasis under extreme hydrostatic pressure suggest DNA methylation plays an important adaptive role in these worms.
Keywords: epigenomics, DNA methylation, deep-sea, Polychaeta, adaptation, Nanopore
Introduction
DNA methylation is a form of epigenetic control, which integrates genetic, environmental, and stochastic cues (Angers et al. 2020). At the cellular and organismal levels, the variation in DNA methylation patterns upon a given genome facilitates cell differentiation and acclimation. At the individual and population levels, differences in epigenetic makeup have ecological and evolutionary repercussions. Among the known forms of DNA methylation, the addition of a methyl group to the fifth cytosine's carbon (5mC) is common in animals and the best studied, particularly in vertebrate models. In vertebrates, cytosine methylation in the CpG context (i.e., adjacent cytosine and guanine in the 5′–3′ direction) leads to gene silencing by preventing the binding of the transcriptional machinery to the DNA (Lyko 2018), whereas the roles of 5mCpG methylation in invertebrates remain poorly understood. The possible roles for DNA methylation in invertebrates include gene expression regulation, alternative splicing, or silencing of transposable elements (TEs) (Flores et al. 2012; Gavery and Roberts 2014). Furthermore, the genomes of vertebrates are fully methylated with only a few hypomethylated regions generally located in gene promoters (Suzuki and Bird 2008), whereas the genomes of invertebrates display wide variations among species in terms of how much and where DNA methylation occur. Such contrasts in methylation patterns between vertebrates and invertebrates suggest different functions and evolutionary histories (de Mendoza et al. 2020).
The functions and origin of DNA methylation can be better understood by examining more species from different invertebrate phyla. Comparative epigenomic analyses at the species and population levels can provide valuable insights into their adaptive strategies and physiological resilience (McCaw et al. 2020). Phylum Annelida is an invertebrate taxon for which epigenetics data are remarkably scarce. These segmented worms are amongst the most diverse and abundant animals in marine ecosystems, occupy a wide variety of niches from soil and marine sediments and from deposit feeders to pelagic and parasitic lifestyles, and provide important ecological services such as bioturbation (Kvist and Oceguera-Figueroa 2021). So far, there has been no complete genome-wide survey on annelids methylomes. DNA methylation that has been detected in a few annelid species (Newbold et al. 2019; Bicho et al. 2020; Ogunlaja et al. 2020; Planques et al. 2021) responded to environmental shifts, indicating its possible role in acclimation and resilience (Marsh and Pasqualone 2014). Furthermore, DNA methylation of their distant cousins in the phyla Nematoda and Arthropoda presents striking interspecific variation. For instance, insect genomes display much lower levels of DNA methylation (<10%) than those of Crustaceans and Arachnids (∼30%) (Provataris et al. 2018). Amongst nematodes, some taxa have completely lost DNA methylation and its associated machinery (e.g., Caenorhabditis elegans) (de Mendoza et al. 2020). An epigenetic diversity similar to that of Arthropods and Nematodes may exist amongst annelid species.
Deep-sea chemosynthetic ecosystems (e.g., hydrothermal vents, hydrocarbon seeps, and organic falls) are characteristically unstable and typified by extreme gradients of the temperature, concentration of chemicals, and food resources. In hydrothermal vents, the environmental conditions can drastically change in a span of a few centimeters and within minutes (Cuvelier et al. 2011; Lee et al. 2015), whereas in hydrocarbon seeps and organic falls, the conditions are relatively stable but are characterized by strong spatial zonation (Zhao et al. 2020). Endemic polychaetes that inhabit these ecosystems belong to three main families, namely, Siboglinidae, Alvinellidae, and Polynoidae. Siboglinids are sessile gutless tubeworms that rely on intracellular chemolithotrophic bacteria for their nutrition, whereas alvinellids and polynoids are mobile and occupy higher trophic levels. The diversity of their phylogenetic and life history traits and the acute variability of conditions in their habitats make these segmented worms exceptional models for the investigation of how species use different epigenetic traits to cope with variable and unpredictable environments. Accordingly, solid baselines for the epigenomes of these animals need to be established.
Breaking ground on the study of deep-sea adaptive epigenomics, the present study provides the first genome-wide methylome survey of three deep-sea polychaete worms, namely, the siboglinids P. echinospica and Ridgeia piscesae and the alvinellid Paralvinella palmiformis (fig. 1). New genomes for R. piscesae and P. palmiformis were assembled. These two species inhabit hydrothermal vents in the northeastern Pacific Ocean and occupy a wide range of temperature conditions (2–60 °C) (Dilly et al. 2012; Tunnicliffe et al. 2014). Whereas the former is mobile, the latter displays a great degree of phenotypic plasticity associated with the different environments it settles, suggesting a unique epigenetically driven polyphenism in this species (Tunnicliffe et al. 2014). In contrast, P. echinospica, which is closely related to R. piscesae, lives in hydrocarbon seeps in the western Pacific Ocean, does not display much morphotypic variation, and occupies a relatively narrower range of environmental conditions (Sun et al. 2021).
Fig. 1.
Species investigated in this study. Left panel: Colony of P. echinospica at the Haima colds seep in the South China Sea (adopted from Sun et al. (2021)). Right panel: Colocalizing colonies of R. piscesae and P. palmiformis at Endeavour Hydrothermal Vents in the northeastern Pacific Ocean (photo courtesy of Canadian Scientific Submersible Facility/Remotely Operated Platform for Ocean Sciences).
Our epigenomic surveys were conducted using data derived from Oxford Nanopore Technologies (ONT). The putative DNA methylation metabolism and genome-wide somatic 5mCpG methylation landscapes of P. echinospica, R. piscesae, and P. palmiformis were characterized within the context of invertebrate methylomics. The three following hypothesis on the putative roles of DNA methylation were tested: 1) the methylation of TEs represses their activity, 2) DNA methylation located within gene bodies (i.e., the transcriptional region which includes both introns and exons) positively affects their expression, and 3) promoter methylation acts as a gene silencing mechanism. Lastly, we compared the epigenetic profiles of orthologous genes across the three species to identify putative epigenetic adaptation their deep-sea environment. Our results suggest that acclimation and adaptation to the deep-sea environment could arise or take effect at the epigenetic level.
Results
DNA Methylation Metabolism
The methylome of P. echinospica, R. piscesae, and P. palmiformis was characterized by reconstructing high-quality genomes for these species. Based on the previously published genome of P. echinospica (Sun et al. 2021), the genomes of R. piscesae and P. palmiformis were sequenced and assembled using a combination of short (Illumina) and long (Nanopore) reads derived from the same DNA extractions (see detailed genome information in table 1). The contig-level assembly of R. piscesae from this study is the most contiguous to date (Wang et al. 2023) and that of P. palmiformis is the first representative genome for the family Alvinellidae. Our conservative genome annotation protocol, in which only transcriptomic data from their respective species and that of species from the same family were used, resulted in the identification of 22,642 and 31,703 high-quality protein-coding gene models (complete and single-copy BUSCO score > 85%) for R. piscesae and P. palmiformis, respectively (table 1), whereas 24,682 protein-coding genes were obtained for P. echinospica.
Table 1.
Summary of Contig-Level Assembly Statistics and Gene Model Annotations for All Available Deep-Sea Polychaete Genomes.
| P. echinospica a,b | R. piscesae a | P. palmiformis a | L. luymesi c | R. pachyptila d | R. piscesae e | Lamellibrachia satsuma f | R. pachyptila g | Oasisia alvinae g | Osedax frankpressi g | |
|---|---|---|---|---|---|---|---|---|---|---|
| NCBI's WGS accession | JAHLWY01 | JAODUO01 | JAODUP01 | SDWI01 | n.a.h | JALOCR01 | JAHXPS01 | n.a.i | n.a.i | n.a.i |
| Contig-level assembly summary | ||||||||||
| Tot. length (Mbp) | 1,095 | 658 | 601 | 602 | 561 | 530 | 665 | 554 | 808 | 285 |
| # of contigs | 12,702 | 9,128 | 3,622 | 55,092 | 447 | 113,548 | 6,320 | 926 | 785 | 1,194 |
| Max. contig length (Kbp) | 1,903 | 2,546 | 3,777 | 274 | 13,119 | 174 | 1,857 | 8,302 | 10,682 | 2,139 |
| Contig N50 (Kbp) | 243 | 365 | 557 | 24 | 2,870 | 10 | 267 | 1.422 | 2,901 | 426 |
| GC content | 41% | 41% | 36% | 40% | 41% | 41% | 40% | 41% | 41% | 29% |
| # of CpGs | 35,178,402 | 20,589,468 | 14,611,299 | 17,810,943 | 17,346,573 | 16,435,603 | 19,045,649 | 16,783,278 | 28,935,539 | 4,591,061 |
| Annotation summary | ||||||||||
| Coding genome tot. length (Mbp) | 26 | 33 | 32 | 42 | 59 | n.a | 39 | 55 | 59 | 29 |
| # of protein-coding genes (without isoforms) | 22,642 | 31,703 | 24,682 | 40,316 | 25,983 | 24,096 | 33,184 | 37,037 | 37,777 | 18,657 |
| Tot. BUSCO score | 88% | 94% | 96% | 98% | 99% | 93% | 91% | 96% | 97% | 91% |
| Complete and single BUSCO score | 79% | 86% | 87% | 91% | 43% | n.a | 90% | 91% | 90% | 87% |
Note.—BUSCO scores are based on gene models protein searches against the metazoan database (odb10). n.a, not available.
This study.
Patra et al. (in prep).
Data available at https://phaidra.univie.ac.at/detail/o:1220865.
Data available at https://github.com/ChemaMD/OsedaxGenome.
Enzymes essential to the DNA methylation metabolism were detected in all three species (fig. 2 and supplementary tables S1–S8, Supplementary Material online). The transcripts for almost all these enzymes were found in the de novo transcriptomes of worms, with some exceptions in R. piscesae whose transcriptome was of the lowest quality among the three worm species (see supplementary methods, Supplementary Material online section 2.6). Sequences were in some cases fragmented or truncated in either the transcriptomic assemblies (supplementary fig. S1, Supplementary Material online) or the gene models (supplementary fig. S2, Supplementary Material online) but not in both, which suggests these cases were resulted from assembly and annotation issues. We were not able to resolve these issues with our current data.
Fig. 2.
Simplified schematic representation of the DNA methylation metabolism that shows the occurrence and expression level of essential enzymes in the studied worms’ genomes and de novo transcriptomes. 5-MTHF, 5-methyltetrahydrofolate; AHCY, S-adenosyl-homocysteine hydrolase; AHCYL, S-adenosyl-homocysteine hydrolase like; BHMT, betaine homocysteine methyltransferase; DNMT, DNA methyltransferase; Hcy, homocysteine; MAT, methionine adenosyltransferase; Met, methionine; MTR, B12-dependent methionine synthase; SAH, S-adenosyl-homocysteine; SAM, S-adenosyl methionine; TET, methylcytosine dioxygenase; THF, tetrahydrofolate.
The genomes of P. echinospica, R. piscesae, and P. palmiformis encode genes for DNMT1 (domain architecture: DMAP-RFD-zfCXXZ-BAH-DNA methylase), which is the DNA methyltransferase enzyme that is principally responsible for propagating cytosine methylation marks in daughter cells after mitosis; DNMT3 (domain architecture: PWWP-ADD-DNA methylase), which is the de novo cytosine DNA methyltransferase gene; and the methylcytosine dioxygenase TET (domain architecture: CXXC-cysteine rich region-catalytic domain), which is responsible for DNA demethylation. Partial sequences for these enzymes were found in the species de novo transcriptomes (supplementary tables S1–S3, Supplementary Material online), but their expression levels were low. DNMT1, DNMT3, and TET transcripts were also detected in the RNA-Seq data of closely related species (tubeworms Riftia pachyptila and Lamellibrachia luymesi and alvinellid Paralvinella grasslei).
Our gene survey of the worm genomes further highlighted putative differences in the DNA methylation metabolism between the siboglinid and alvinellid worm taxa. Tubeworms possess two enzymes for the remethylation of homocysteine to methionine, the B-12-dependent methionine synthase (MTR) and the betaine homocysteine methyltransferase (BHMT), whereas only MTR was detected in P. palmiformis (genome and transcriptome) and P. grasslei (transcriptomes).
Lastly, two genes for the methionine adenosyltransferases (MATa and MATb) were found in both the tubeworms and alvinellid species (supplementary table S4, Supplementary Material online). Recent genetic and transcriptomic data revealed two versions of MAT also exist in the Alvinellidae's sister group Amphraetidae (Stiller et al. 2020), Osedax (Moggioli et al. 2023), and some species of the Terebellidae family (supplementary fig. S3, Supplementary Material online). Mat paralogs were found in tandem in the vestimentiferans but not in Osedax or P. palmiformis's genomes. Evidence of relaxed selection was only found in the Mat-b paralogs of vestimentiferans (supplementary table S9, Supplementary Material online). No evident episodic diversifying selection was detected across the worm's phylogeny. Taken together, these results suggest the Mat gene may have duplicated multiple times in polychaetes, including one recent duplication in vestimentiferans, but in the absence of a fully resolved phylogeny, the evolutive history of this gene remains uncertain.
Genome-Wide Methylation Profile
Considering the high accuracy of nanopore sequencing data (see supplementary methods, Supplementary Material online section 2.4), they were used to detect 5mC methylation in CpG context (5mCpG) in P. echinospica, R. piscesae, and P. palmiformis. Methylation calls on CpG motifs covered by <10 reads were only considered for P. palmiformis because of its low sequencing depth. After coverage-based filtering, the remaining CpGs called for methylation represented 86%, 87%, and 97% of all CpGs present in the genome assemblies of P. echinospica (35,178,402 CpGs), R. piscesae (20,589,468 CpGs), and P. palmiformis (14,611,299 CpGs), respectively.
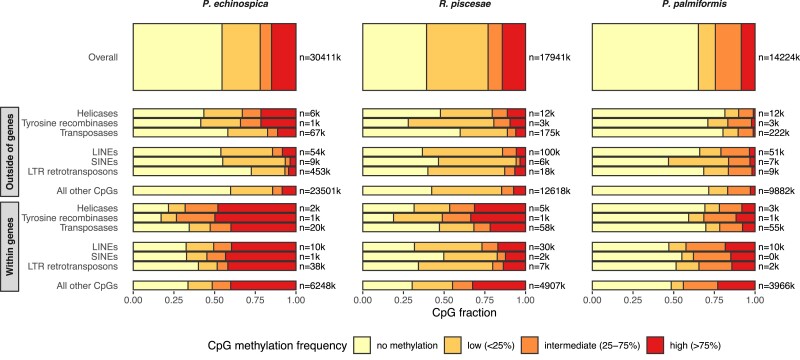
The genomes of R. piscesae, P. echinospica, and P. palmiformis showed similar methylation profiles which are characterized by relatively low levels of CpG methylation (fig. 3). Global CpG methylation in these worms was higher than that of insects but well within the range reported for other arthropod and nematode species. Methylation was present on 35–61% of CpGs. A greater proportion of highly methylated CpGs was found in both tubeworms (∼15%) than in P. palmiformis (8%). In all three polychaetes, CpGs within genes were more methylated than those in intergenic regions (fig. 3). Within genes, methylation was more polarized on exons (supplementary fig. S4, Supplementary Material online) than on introns with the first exon of genes being typically unmethylated (supplementary fig. S5, Supplementary Material online).
Fig. 3.
Genome-wide methylation frequency spectra in three polychaete species and CpG methylation on genetic sequences identified as mobile elements by RepeatMasker outside and within genes. The number of CpGs assessed is indicated on the right. CpGs with <10× coverage were only considered in P. palmiformis because of its low sequencing depth. DNA transposons (helicases, tyrosine recombinases, and transposases) and retrotransposons (LINEs, SINEs, and LTR retrotransposons) were grouped separately.
TEs and Long Terminal Repeat Retrotransposon Methylation
The analysis of the mobilome of P. echinospica, R. piscesae, and P. palmiformis with RepeatMasker (Chen 2004) identified genetic sequences that belong to DNA transposons (helicases, tyrosine recombinases, and transposases), SINE and LINE retrotransposon elements. A more stringent bioinformatic pipeline annotated intact and well-defined long terminal repeat (LTR) retrotransposon (Aroh and Halanych 2021). LTR retrotransposons characteristically possess two identical sets of LTRs, which independently accumulate mutations after their integration in the genome. Thus, insertion times were estimated using the sequence divergence between these paralogs.
CpG methylation levels on mobile elements matched those of their flanking regions (i.e., genomic context) with higher methylation on intragenic elements than intergenic ones (fig. 3). A total of 2,019, 146, and 84 intact LTR retrotransposons were detected in P. echinospica, R. piscesae, and P. palmiformis, respectively (supplementary fig. S6, Supplementary Material online). DNA methylation level (i.e., mean methylation frequency) and insertion time (in Mys) were estimated for most of these LTR retrotransposons (1984, 141, and 84 in P. echinospica, R. piscesae, and P. palmiformis, respectively). Contrary to our expectations, younger LTR retrotransposons were not more methylated than older ones (fig. 4). In fact, when controlling for genomic context, insertion time and methylation load were positively correlated in P. echinospica (supplementary table S10, Supplementary Material online). However, a strong association was observed between insertion time and CpG depletion, indicating that in P. echinospica, the mutation rate on methylated LTRs was considerably higher than on unmethylated ones and may have biased our insertion time estimates (supplementary fig. S7, Supplementary Material online). Still, highly methylated LTR retrotransposons were hypermethylated compared with their flanking regions and inversely for lowly methylated LTR retrotransposons (fig. 4 and supplementary figs. S8 and S9 and supplementary table S11, Supplementary Material online). The epigenetic state of lowly methylated LTR retrotransposons was less dependent on the surrounding epigenetic context than highly methylated ones (supplementary figs. S7–S9, Supplementary Material online) but could not be consistently explained by the other factors tested (LTR retrotransposon size, insertion time, and location).
Fig. 4.
LTR retrotransposon methylation and their genomic context. The average methylation frequency was computed for each LTR retrotransposon and their 4 Kbp upstream and downstream flanking regions (1 Kbp sliding window with a 200 bp step). The step-lines represent the dataset aggregated averages, and the ribbons represent the 95% confidence interval from a 1,000 bootstrap resampling. LTR retrotransposon sizes were normalized. High LTR methylation = mean coverage > 75% and mean depth > 50% and low methylation = mean coverage ≤ 75% and mean depth ≤ 50%. Insertion times for each species are grouped in quantiles.
Gene Body and Promoter Methylation
We found supporting evidence for the role of gene body methylation in the regulation of gene expression in the surveyed worms. The mean methylation frequency (methylation level) of genes was the highest in functional categories associated with housekeeping functions, such as translation and RNA processing, nuclear and ribosomal structure, and cell cycle (supplementary figs. S10 and S11, Supplementary Material online), and was positively correlated to expression (fig. 5 and supplementary fig. S12, Supplementary Material online) even after controlling for the confounding effects of gene length and GC content (supplementary fig. S13 and supplementary table S12, Supplementary Material online for results of partial Spearman correlation analyses).
Fig. 5.
Methylation level around the TSS of genes classified into four groups according to their 1 Kbp upstream and gene methylation state. Genes with missing methylation data were not included. The bottom row represents the normalized gene expression (transcripts per million; TPM) for the gene groups (the whiskers extend to 95% of the data). The “high U low G” group likely contains genes misclassified because of natural stochasticity or noisy methylation data. Moreover, R. piscesae and P. palmiformis transcriptomes include data from different individuals and different tissues than the ones surveyed for methylation, whereas P. echinospica transcriptome was obtained from the same individual and tissues as the epigenome. Differences in gene expression across groups were tested using pairwise KS tests (P value threshold = 0.05) and are presented above the boxes. Gene group description: low UG, low-intermediate upstream and gene methylation (mean coverage ≤ 75% and mean depth ≤ 50%); high U low G, high upstream methylation (mean coverage > 75% and mean depth > 50%) and low-intermediate gene methylation; high UG, high upstream and gene methylation; and low U high G, low-intermediate upstream and high gene methylation. The methylation coverage and depth thresholds were selected to match density clusters in the data (supplementary fig. S38, Supplementary Material online); n = group gene count.
Gene expression was also affected by promoter methylation. Genes characterized by low gene body methylation had higher expression when their 4 Kbp upstream region was heavily methylated (high U low G > low UG in fig. 5). The reverse relation was true amongst genes with high gene body methylation (high UG < low U high G in fig. 5). These correlations held whether the methylomics and transcriptomic data were generated from the same individual and tissues (P. echinospica) or not (R. piscesae and P. palmiformis). Among genes with high gene body methylation, the negative correlation between expression and upstream methylation broke down with increasing distance to the transcription start site (TSS) and mostly held when only the best genes models (i.e., the highly conserved metazoan genes identified by BUSCO (Simão et al. 2015) as complete in our genomes) were included in the analysis (supplementary fig. S14 and supplementary tables S13 and S14, Supplementary Material online). This indicates that our results were not overly confounded by misidentified promoters due to inaccurate gene models. Genes with unmethylated promoters were preferentially associated with protein folding, intracellular trafficking, and energy metabolism, whereas genes with methylated promoters were particularly enriched in signal transduction (in the tubeworms), replication, and repair (in P. palmiformis) functions (supplementary figs. S15 and S16, Supplementary Material online).
Methylation Profiles of Orthologous Genes
A total of 15,559 gene orthologs were identified across the three worm genomes, including 5,132 core orthologs represented by a single gene model in all three species. Gene body methylation could be characterized in all species for 4,942 of these reliable core orthologs (supplementary fig. S17, Supplementary Material online). Among them, 1,185 had contrasting levels of gene body methylation across species and 3,036 had high levels of DNA methylation in all species. The methylation state of gene promoters across species was also characterized for 2,906 of the 3,036 orthologs with conserved hypermethylated gene bodies (supplementary fig. S17, Supplementary Material online). Based on the comparison of the relative expression of orthologs in P. echinospica and P. palmiformis, the association of gene body hypermethylation and promoter demethylation with higher expression was confirmed (supplementary figs. S18 and S19, Supplementary Material online). Moreover, the methylation levels of promoters and gene bodies were tightly matched by their respective methylation-driven mutational biases (CpG observed/expected) but not the gene length and GC content (supplementary figs. S20–S23, Supplementary Material online). The epigenetic and genetic signatures of gene body and promoter methylation were more strongly coupled for genes with conserved methylation across species compared with genes having contrasting methylation profiles (fig. 6 and supplementary fig. S24 and S25, Supplementary Material online). In addition, the coupling strength was correlated with the phylogeny, and it was stronger across worm families than amongst the two siboglinid tubeworms. These results show that the methylation states of a large number of genes and, to a lesser extent, their promoters are stable and persistent over evolutionary time. Genes with conserved gene body hypermethylation did not represent a random subsample of all core orthologs (hypergeometric test P value <0.01) and were most enriched in genes associated with translation and protein folding and degradation functions, particularly in ribosomal proteins, chaperones, and proteins associated with the ubiquitin–proteasome pathway (fig. 7). Together with the genes of the energy metabolism (ATPase and NADH dehydrogenase subunits), orthologs associated with protein homeostasis were also overrepresented amongst orthologs with conserved unmethylated promoters (fig. 7).
Fig. 6.
Stronger genetic signatures are associated with persistent methylation across taxa. For each gene group, the correlation strength was estimated by the goodness of fit of the logistic regression of CpG observed/expected on methylation level. The mean (bar height) and standard deviation (error bar) of 1,000 bootstrap samples are represented.
Fig. 7.
Most enriched functional categories amongst genes with conserved hypermethylated gene bodies or conserved promoter methylation. The top three functional categories (based on enrichment factor) for each of the gene body and promoter gene sets are shown for the KEGG (top) and KOG (bottom) functional annotations. The enrichment factor is defined as the normalized differences between expected and observed relative abundances of functional categories. The significance of the enrichment for each functional category was tested with Fisher tests, followed by Holms correction. Nonsignificant enrichments (Fisher test P value ≥ 0.05) are not shown.
Discussion
The methylome of three polychaete species (P. echinospica, R. piscesae, and P. palmiformis) from deep-sea ecosystems was characterized for the first time. Based on the identification of the key genes of the DNA methylation metabolism in their genomes (two of which were newly assembled in the present study) and available transcriptomes, these worms possessed and expressed the machinery for DNA methylation, but interesting differences were observed across taxa. Considering that the DNA methylation estimates obtained from Nanopore data were as accurate as those recovered from WGBS at the genomic resolution of genes, this third-generation sequencing technology was used to describe the epigenomic landscape across the whole worm genomes.
Mobile genetic elements were not ubiquitously targeted by methylation, although a close examination of well-defined LTR retrotransposons revealed differences in methylation load in comparison with their genomic context. Furthermore, our results strongly support the hypotheses that gene body methylation strengthens and promoter methylation hinders gene expression in these worms. Indeed, the DNA methylation of gene bodies and promoters were positively and negatively correlated with expression. Lastly, we found evidence for the presence of conserved gene and promoter methylation profiles across the species which could reflect epigenetic adaptations to the deep-sea environment.
Siboglinids and Alvinellids Possess the Complete and Active Machinery for Setting Up and Removing DNA Methylation
All key genes for the DNA methylation metabolism were present in the worms’ genomes, although they were not ubiquitously expressed in the somatic tissues of adult worms. DNMT1, DNMT3, and TET were fragmented and poorly represented in the transcriptomic data of muscle and dermal tissues. In both vertebrates (e.g., cattle and Medaka fish) and invertebrates (e.g., oyster Crassostrea gigas and the model polychaete species Platynereis dumerilii), these enzymes were predominantly expressed in oocytes and early embryos (Wang et al. 2014; Duan et al. 2019; Wang and Bhandari 2019; Planques et al. 2021). Our results are consistent with the observations made in other animals, in which most of the DNA methylation marks were removed during the gametogenesis and reset during the development (Smallwood and Kelsey 2012). Epigenetic marks set up during the development are suspected to contribute to the thermal acclimation of R. piscesae and Paralvinella species (Dilly et al. 2012; Tunnicliffe et al. 2014). Future studies on the DNA methylation dynamics in gametogenesis and early development may provide more information on the polychaetes epigenetic landscapes of their ontogenic and adaptive roles.
Putative metabolic differences in DNA metabolism also exist across the two families. Unlike the alvinellids, tubeworms possess two enzymes for the remethylation of homocysteine to methionine (BHMT and MTR). BHMT homologs can be found in other invertebrates, including polychaetes, suggesting a secondary loss in alvinellids. Putative independent duplications of methionine adenosyltransferases (MAT) genes were also observed in tubeworms and alvinellids, respectively. This finding is remarkable, because MAT, which is responsible for synthesizing the main methyl donor S-adenosylmethionine, is a highly conserved enzyme and only a few paralogs have been discovered so far (Sanchez-Perez et al. 2008; Riesgo et al. 2012). The products of BHMT, MTR, and MAT are involved in several metabolic functions beyond the epigenetic metabolism (Finkelstein 1990; Fontecave et al. 2004; Stipanuk 2020), and in polychaetes, their specific roles still remain unclear.
Siboglinids and Alvinellids Possess Intermediate Levels of DNA Methylation
The methylome surveys of the three deep-sea polychaetes revealed that they possessed intermediate levels of DNA methylation (35–61% of methylated CpGs) similar to that of early-branching arthropods (de Mendoza et al. 2020). DNA methylation in the arthropod species Stegodyphus dumicola, Strigamia maritima, and Procambarus virginalis ranges from 17% to 76% of methylated CpGs (de Mendoza et al. 2020), whereas it is typically present in <30% of CpGs in insects (<10% in holometabola) (Provataris et al. 2018).
TEs Carry Weak DNA Methylation Signatures
DNA methylation may act in silencing TE activity via RNA-directed targeting mechanisms (Deniz et al. 2019). TEs are categorized into two clades according to their mode of propagation. DNA transposons move from one genomic location to another as DNA segments, whereas retrotransposons propagate through RNA intermediates, which are reverse-transcribed into DNA (Kojima 2019). In invertebrates, the methylation state of TEs varies widely across element types and species (Lyko et al. 2010; Wang et al. 2014; Gatzmann et al. 2018; de Mendoza, Hatleberg, et al. 2019; Lewis et al. 2020; Ying et al. 2022), suggesting that the silencing of TEs by DNA methylation is a modular function acquired multiple times in their evolutionary histories (de Mendoza et al. 2020). The analyses of mobile DNA methylation did not provide strong support for the hypothesis that TE proliferation is broadly repressed by DNA methylation in the polychaetes. The methylation levels of mobile elements were generally lower (>85% hypomethylated CpGs TEs in the worms) than those of vertebrates (<3% hypomethylated CpGs in humans (Pehrsson et al. 2019)) or plants (∼25% of hypomethylated TEs in Arabidopsis (Ahmed et al. 2011)) and matched that of their genomic context.
Well-defined intact LTR retrotransposons were detected in the worms’ genomes. Their abundance falls within the same range as that in other protostomes (Kim et al. 1994; de la Chaux and Wagner 2011; Thomas-Bulle et al. 2018; Aroh and Halanych 2021). Most LTR retrotransposons (>90%) belonged to the Gypsy superfamily, as previously reported for vestimentiferan tubeworms (Aroh and Halanych 2021; de Oliveira et al. 2022; Wang et al. 2023), mollusks (Thomas-Bulle et al. 2018), arthropods (Kim et al. 1994; Kaminker et al. 2002; Pelisson et al. 2002; Xu et al. 2005; Piednoël et al. 2013; de Mendoza, Pflueger, et al. 2019), and nematodes (de la Chaux and Wagner 2011). Nevertheless, a higher abundance of LTR retrotransposons was observed in P. echinospica than in the other worms, which appears to be correlated with its larger genome size (almost double that of the close-relative siboglinids R. piscesae (Wang et al. 2023), L. luymesi (Li et al. 2019), and R. pachyptila (de Oliveira et al. 2022; Moggioli et al. 2023)). The high abundance of LTR retrotransposons in P. echinospica and their insertion time distribution suggests a recent wave of invasion that contributes to the unique genome expansion observed in this species (Sun et al. 2021).
Younger and presumably more mobile LTR retrotransposons did not display elevated methylation as observed in mammals (Barau et al. 2016) and Cnidaria (Ying et al. 2022), but strong variation of DNA methylation load was observed at LTR retrotransposon boundaries. Such contrasting methylation densities between TEs and their immediate surroundings have also been observed in other species (Lyko et al. 2010; Lewis et al. 2020; Ying et al. 2022) but are hard to interpret without a comprehensive data set of invertebrate methylomes. This finding was obtained possibly because methylation density variation along the DNA sequence rather than absolute methylation level allows the recognition of certain TEs, but this hypothesis needs to be further tested.
Gene Body Methylation Is Associated with Increased Gene Expression
Gene body methylation was observed in these three species and was positively correlated to transcript abundance, thus supporting that it strengthens gene expression. In rat embryos, Neri et al. (2017) demonstrated that the high density of intragenic methylation marks induced by the de novo DNA methyltransferase 3b (DNMT3b) reduces spurious transcription initiation, thus increasing transcription yield, by preventing the aberrant intragenic binding of the RNA polymerase. Although a similar mechanism of transcription stabilization in invertebrates has not been determined, the presence of DNMT3 (a homolog to DNMT3b) in the worms’ genomes supports the hypothesis that gene bodies are specifically targeted by the DNA methylation machinery. DNMT gene knockout experiments have shown that somatic gene expression in plants and insects was not necessarily affected by gene body methylation (Zhang et al. 2006; Bewick et al. 2019). Similar experimental approaches in model polychaete species are needed to assess if DNA methylation exerts direct epigenetic control on gene expression in this group. Since deep-sea worms have not been successfully reared in the laboratory, it will be very hard to conduct in vitro experiments, although the animals can be fixed in situ (Yan et al. 2022).
Promoter Methylation Is Associated with Decreased Gene Expression
In vertebrates, gene promoter methylation silences expression by impeding the binding of transcription factors (Suzuki and Bird 2008). A similar gene silencing mechanism was observed in the mud crab Scylla paramamosain (Jiang et al. 2020). In the deep-sea worms investigated in this study, a negative correlation between the promoter methylation and expression amongst highly methylated genes was observed. This association could reflect a gene silencing mechanism similar to that of vertebrates, although the relative weakness of the association in our samples suggests that promoter methylation may play a supporting role to other gene silencing mechanisms (Keller et al. 2016). Marsh and Pasqualone (2014) showed that DNA methylation in the Antarctic polychaete species Spiophanes tcherniai was linked to thermal acclimation. Future study of the physiological role of DNA methylation in deep-sea worms exposed to strong spatial and temporal thermal gradients, such as those inhabiting hydrothermal vents, could bring new insights into the mechanisms of animal's adaptation to environmental disturbances in the deep-sea habitats.
Conserved Patterns of DNA Methylation in the Deep-Sea Worms May Reflect Adaptations to Hydrostatic Pressure
Methylation profiles, which were conserved across species (for both gene bodies and promoter regions), were matched by a strong genetic signal, indicating their persistence over evolutionary times (methylated cytosines have a higher C→A mutation rate (Coulondre et al. 1978; Ehrlich et al. 1986)) and pervasiveness (the mutations must affect the germ cells to be passed on to future generations). The epigenetic makeup of an individual is partly encoded in its genome and is therefore under the influence of natural selection (Angers et al. 2020). Thus, polychaetes species that are endemic to deep-sea chemosynthetic ecosystems are expected to have acquired adaptive epigenetic buffering strategies aside from behavioral coping methods and molecular alterations to extreme habitats.
In the three species studied, genes with a conserved pattern of promoter hypomethylation and gene body hypermethylation (conducive to upregulation) displayed strong methylation-driven mutational biases and were disproportionately associated with the maintenance of protein homeostasis. Chaperones, which are important for protein folding stability in the deep sea (Cario et al. 2016; Ritchie et al. 2018; Weber et al. 2020), were highly methylated in the worms. Likewise, the ubiquitin–proteasome pathway, which is responsible for degrading misfolded or damaged proteins and plays an important role in cold water–adapted species (Todgham et al. 2017), was overrepresented amongst genes with hypomethylated promoters in all three worms. Our samples were not fixed in situ, and thus, we cannot exclude that the methylation profiles of the worms partly reflect the decompression stress they suffered during their recovery to the ship (Yan et al. 2022). However, methylomes tend to be much more stable in time than transcriptomes (Strader et al. 2020), and thus, environmental stress during the short timeframe of samples recovery (a few hours at most) is unlikely to have had substantial consequences on worms’ methylomes. Further comparative studies on deep-sea invertebrates and their shallow-water relatives could combine transcriptomic and epigenomic data to shed new light onto the mechanisms of adaptation to deep-sea environment.
Conclusion
Deep-sea polychaetes possess a fully functional DNA methylation metabolism, and their genomes are moderately methylated. In P. echinospica, R. piscesae, and P. palmiformis, DNA methylation does not specifically target TEs but is highly concentrated in genes. Among the three worms, gene expression is positively correlated to gene body methylation and, to a lesser extent, negatively correlated to promoter methylation. These chemical modifications of the DNA, which appear to exert control on gene expression, could play roles in metabolic acclimation at the individual level. Finally, the persistent and conserved epigenetic profiles of genes responsible for maintaining homeostasis suggest DNA methylation plays an important adaptive role at the population and species levels. The complete genomes and epigenomes that were assembled for three endemic polychaetes would serve as a valuable reference for future investigations of the ecological and evolutionary roles of DNA methylation in deep-sea animals.
Methods
Sample Collection, Nucleic Acid Extraction, and Sequencing
R. piscesae and P. palmiformis specimens were collected together from the same vent field on the Juan de Fuca Ridge (47°57.00′ N, 129°5.82′ W, 2,196 m) with the ROV ROPOS in June 2016 during a cruise of the Canadian Healthy Oceans Network on board the CCGS John P. Tully. Paraescarpia echinospica was collected from the Haima cold seep in the South China Sea (16°43.80′ N, 110°28.50′ E, 1,390 m) (Sun et al. 2021). Ridgeia piscesae and P. echinospica were frozen at −80 °C immediately after reaching the vessel deck, whereas P. palmiformis was kept in 95% ethanol.
DNA from the vestimentum of R. piscesae and the body wall of P. palmiformis was extracted via phenol–chloroform and ethanol precipitation (Sambrook et al. 1989) and purified using the Genomic DNA Clean & Concentrator-10 kit (ZYMO Research, CA, USA) according to the manufacturer's protocol. DNA from P. echinospica was extracted using the MagAttract HMW DNA kit (Qiagen, Hilden, Germany). RNA was extracted with TRIzol reagent from four tissues of R. piscesae (gills, vestimentum, epiderm, and trophosome) and five tissues of P. echinospica (gills, collar, vestimentum, trophosome, and opistosome). For P. palmiformis, the publicly available whole-body transcriptome of another individual (National Center for Biotechnology Information [NCBI] BioSample: SAMN14351933) was used.
DNA and RNA samples were sequenced by Novogene (Beijing, China) on the Illumina NovaSeq 6000 platform. Paired-end libraries with insert size of 350 bp were constructed using the NEBNext DNA Library Prep Kit. DNA samples were also sequenced on the ONT PromethION platform. All genomic libraries reached a nuclear genome coverage of more than 85×, except for the Nanopore library of P. palmiformis, which reached ∼44× (supplementary fig. S26 and supplementary table S15, Supplementary Material online).
CpG Methylation Calling
Nanopolish v0.13.3 (Simpson et al. 2017) pipeline was used under default parameters to detect CpG methylation in Oxford ONT data, because it offers the best compromise between accuracy and sensitivity of methylation detection and computational efficiency in several benchmarking studies (Liu et al. 2021; Yuen et al. 2021). Nanopolish uses a pretrained hidden Markov model to assign methylation log-likelihood ratios (LLRs) to all CpGs within a 10 bp window. Briefly, nanopore reads were indexed and mapped into the reference genomes. Then, the raw electric signatures (or squiggles) were fetched and aligned back onto their respective mapped reads (supplementary fig. S27, Supplementary Material online). The likelihoods of each CpG-containing 6-mer given a five-base alphabet (with A, T, G, C, and M for methylated cytosines) were then calculated, and CpG call groups (sequences containing one or multiple CpG motifs within 10 bp of each other) were considered methylated if their log-likelihood of being methylated was twice that of being unmethylated (LLR > 2). CpG call groups containing more than one CpG motif represented 26–32% of all calls. The mean group coverage and methylation frequency were attributed to all CpGs within these groups. Accordingly, a bed file of all CpGs for each genome was generated and mapped to the respective methylation annotations (call coverage and frequency) with the bedtools map function (Quinlan and Hall 2010).
The precision of DNA methylation calls derived from Oxford Nanopore sequencing was assessed by comparing the whole-genome DNA methylation estimates of the same DNA extraction of the vestimentum of one R. piscesae individual that was obtained through Nanopore sequencing (sequencing depth >100×) with those obtained via whole-genome bisulfite sequencing (WGBS; sequencing depth >85×). The WGBS method description is available in the Supplementary material. The effect of sequencing effort was tested by artificially reducing the depth of coverage of the Nanopore dataset to 20× and 10× (supplementary fig. S28, Supplementary Material online). Nanopore-derived methylation calls closely matched those of WGBS even at a low sequencing depth (supplementary figs. S29–S31 and supplementary tables S16 and S17, Supplementary Material online). The increasing affordability of DNA sequencing has led to numerous genome sequencing projects, which make use of third-generation sequencing technology to achieve high assembly contiguity (supplementary fig. S32, Supplementary Material online). Our analyses show these data can be reinvestigated for DNA methylation. In the species investigated here, optimal accuracy and sensitivity of DNA methylation estimates was ensured by applying thresholds on the methylation call coverage (10× for P. echinospica and R. piscesae and 1× to P. palmiformis because of its low sequencing depth), and the window size used for averaging methylation calls over genomic regions was set to 1,000 bp.
Genome Assemblies and Annotations
The annotated genome assembly for P. echinospica was based on the study of Sun et al. (2021). For R. piscesae and P. palmiformis, new genome and transcriptome assemblies were constructed and annotated following a similar protocol to that of Sun et al. (2021). The detailed methods for these genome reconstructions are presented as Supplementary material and summarized below.
Contig-level assemblies were produced for the three genomes by using a combination of short (Illumina) and long (ONT) reads. For each genome, multiple assembly pipelines were used, and the assembly with the best completeness and contiguity statistics was retained (supplementary fig. S33 and supplementary table S18, Supplementary Material online). Putative misassemblies were avoided by removing reads from extranuclear compartments from the libraries of R. piscesae and P. palmiformis by mapping them against the reference mitochondrial (KJ872501 (Jun et al. 2016) and OL802212 (Perez et al. 2022)) and endosymbiont (LDXT01 (Perez and Juniper 2016)) genomes.
Prior annotations, repeat regions, and putative TEs were masked using a custom repeat database composed of known repeat motifs and sequences issued from the universal databases RepBase27.02 (Bao et al. 2015) and dfam3.3 (Storer et al. 2021), and species-specific repeats were detected using RepeatModeler (v2.0.1) (Chen 2004). LTR retrotransposons were detected following Aroh and Halanych (Aroh and Halanych 2021) (see supplementary methods, Supplementary Material online). Gene model inferences were obtained using Maker2 (Holt and Yandell 2011) by integrating 1) ab initio gene predictions, 2) species-specific transcriptomes, and 3) additional proteomic evidence from closely related species. The ab initio gene predictions were performed using Augustus, and species-specific sets of modeling parameters were obtained by training the algorithm with a set of high-confidence gene models based on transcriptomic and proteomic evidence. The transcriptomic evidence included transcriptomes assembled from the same individuals that produced the reference genomes (P. echinospica and R. piscesae) and additional assemblies from publicly available databases (R. piscesae and P. palmiformis). These transcriptomes were assembled with Trinity (v2.11.0) (Grabherr et al. 2011) in de novo and genome-guided (when reference genome was available) modes (supplementary fig. S34, Supplementary Material online). Proteomic evidence was obtained from the siboglinid species R. pachyptila (de Oliveira et al. 2022), L. luymesi (Li et al. 2017; Li et al. 2019), and Escarpia spicata (Li et al. 2017) for R. piscesae and the alvinellid species Alvinella pompejana (Gagnière et al. 2010), Paralvinella hessleri (Wang et al., unpublished), and P. grasslei (Stiller et al. 2020) for P. palmiformis (supplementary table S19, Supplementary Material online). The exon-level annotation edit distance (eAED) was superior to 0.5 for more than 85% of the final gene models (supplementary fig. S35, Supplementary Material online) with complete and single-copy BUSCO (Simão et al. 2015) scores >85% against the metazoan database (table 1 and supplementary table S20, Supplementary Material online). The final gene models were Blast-searched against NCBI's nr database and screened for functional annotation against the Gene Ontology (GO), Protein families (Pfam), EuKaryotic Orthologous Groups (KOG), Kyoto Encyclopedia of Genes and Genomes (KEGG), Enzyme Commission (EC), and The Institute for Genomic Research curated protein families (TIGRFAM) databases (supplementary fig. S36, Supplementary Material online). Protein Blast searches against the gene models of P. echinospica and R. pachyptila were further conducted for R. piscesae (supplementary fig. S37, Supplementary Material online). Finally, genes associated with the methylation metabolism were further identified amongst the gene models and de novo transcriptomes through their specific functional domains and homology to protein sequences in UniProt's Swiss-Prot and TrEMBL databases (BlastP searches).
Methylome Annotations
The per-CpG methylation information was averaged over the genomic annotations (and vice versa) by using the bedtools map function (Quinlan and Hall 2010). A detailed description of these steps is available in supplementary methods, Supplementary Material online. For each feature, methylation coverage was defined as the fraction of called CpGs for which methylation was detected, and methylation depth represents the fraction of reads for which methylation was detected amongst methylated CpGs. The parameters are related by the following expression: mean methylation frequency = methylation coverage × methylation depth. Features were also annotated as highly methylated (or hypermethylated) at methylation coverage > 75% and methylation depth > 50% according to the observed density clustering (supplementary fig. S38, Supplementary Material online).
Gene Expression Quantification and Protein Ortholog Identification
The expression of gene model mRNA sequences and de novo transcriptome contigs were quantified with Salmon (Patro et al. 2017) by using their corresponding transcriptomic libraries. Protein orthologs were identified with Proteinortho (Lechner et al. 2014) by using DIAMOND (Buchfink et al. 2021) (parameters: −e = 1e−06 −sim = 1 −identity = 40 −cov = 50) and the “–synteny” option.
Phylogenetic and Statistical Analyses
Phylogenetic trees for the Mat gene were reconstructed using Bayesian inference (BI) and maximum likelihood (ML) methods from codon alignments of polychaetes using the general time-reversible (GTR) substitution model. These alignments included additional Siboglinidae (Moggioli et al. 2023), Sabellida (Tilic et al. 2020), and Terebellida (Stiller et al. 2020) species. Relaxed selection in Mat-b compared with Mat-a paralogs was tested with RELAX (Wertheim et al. 2015), whereas episodic diversifying selection was detected with aBSRel (Smith et al. 2015). Detailed methods for the phylogenetic and selection analyses are presented in the Supplementary material. The multiple sequence alignment files are available on our GitHub page.
Statistical analyses were performed in R (Ihaka and Gentleman 1996). The significance of gene expression differences between methylation group pairs was assessed using Kolmogorov–Smirnov (KS) tests (function ks.test from the stats package v4.1.1) at a P value threshold of 0.05. Hypergeometric tests against 100 random subsamples were used to test whether the distribution of functional categories in a specific gene subset was representative of the whole. Fisher tests followed by Holms correction were used to detect the overrepresentation of certain functional categories within gene subsets. Partial correlations (function pcor.test from the package ppcor v1.1) were used to control for the confounding factors of gene length and GC content for the comparison of promoter methylation and gene expression and epigenomic context for the comparison of LTR retrotransposon methylation level and estimated insertion time. Pairwise Pearson correlations for LTR retrotransposon variables were computed using the function rcorr from the package Hmisc v4.6-0 (Harrell 2019). The strength of the epigenetic/genetic signature coupling was quantified by computing the goodness of fit of the logistic regression of the methylation-driven mutational bias (CpG observed/expected) across two observed methylation categories (high- and low-intermediate); confidence intervals were estimated through 1,000 bootstrap resampling.
Supplementary Material
Acknowledgments
We thank Verena Tunnicliffe, Catherine Stevens, Rachel Boschen-Rose, and the crew members of the CCGS John P. Tully and ROV ROPOS for providing the R. piscesae and P. palmiformis specimen. We acknowledge Jack (Yick Hang) Kwan and Yi Yang for their help in nucleic acid extraction. We are also grateful to Alex de Mendoza for his comments on this study's preliminary results; Chema Martin Duran, Ekin Tillic, and Josefin Stiller for providing additional polychaete genomic and transcriptomic assemblies; and Sandra Ann Binning, Sophie Breton, Marie-Anne Cambon-Bonavita, and three anonymous reviewers for their helpful comments on the manuscript. This study was supported by grants awarded to P.-Y.Q. from the Southern Marine Science and Engineering Guangdong Laboratory (Guangzhou) (2021HJ01 and SMSEGL20SC01), the Major Basic and Applied Research Projects of Guangdong Province (2019B030302004-04), and a GRF and CRF of HKSAR government (16101822 and C2013-22G) and grants awarded to B.A. by the Natural Science and Engineering Research Council of Canada (N238600) and the Digital Research Alliance of Canada (RRG #4257). M.P. received support from the Natural Science and Engineering Research Council's Alexander Graham Bell fellowship and the Fonds de recherche du Québec—Nature et Technologie and Quebec Center for Biodiversity Science's international internship award.
Contributor Information
Maeva Perez, Southern Marine Science and Engineering Guangdong Laboratory (Guangzhou), Guangzhou, China; Department of Ocean Science, The Hong Kong University of Science and Technology, Kowloon, China; Department of Biological Sciences, Université de Montréal, Montréal, Canada.
Oluchi Aroh, Department of Biological Sciences, Auburn University, Auburn, AL, USA.
Yanan Sun, Laboratory of Marine Organism Taxonomy and Phylogeny, Chinese Academy of Sciences, Institute of Oceanology, Qingdao, China.
Yi Lan, Southern Marine Science and Engineering Guangdong Laboratory (Guangzhou), Guangzhou, China; Department of Ocean Science, The Hong Kong University of Science and Technology, Kowloon, China.
Stanley Kim Juniper, School of Earth and Ocean Sciences, University of Victoria, Victoria, Canada.
Curtis Robert Young, National Oceanography Center, Southampton, UK.
Bernard Angers, Department of Biological Sciences, Université de Montréal, Montréal, Canada.
Pei-Yuan Qian, Southern Marine Science and Engineering Guangdong Laboratory (Guangzhou), Guangzhou, China; Department of Ocean Science, The Hong Kong University of Science and Technology, Kowloon, China.
Supplementary Material
Supplementary data are available at Molecular Biology and Evolution online.
Author Contributions
M.P. conceived the study, performed most of the upstream data processing and all downstream analyses, and wrote the manuscript. O.A. performed upstream data processing (LTR retrotransposon detection) and provided advice on the manuscript. Y.S. performed upstream data processing (genome sequencing, assembly, and annotations of P. echinospica genome) and provided advice on the manuscript. Y.L. provided computational support for the upstream data processing and provided advice on the manuscript. S.K.J. and C.R.Y. advised on the manuscript. B.A. and P.-Y.Q. conceived the study and wrote the manuscript.
Data Availability
Raw sequence reads are available in NCBI's SRA database under the accession numbers SRR21707426–SRR21707428 (P. palmiformis Illumina), SRR217033–SRR217038 (Nanopore reads of R. piscesae and P. palmiformis), and SRR21707446–SRR21707448 (R. piscesae Illumina). The whole-genome shotgun projects for P. echinospica, R. piscesae, and P. palmiformis were deposited in GenBank under accession numbers JAHLWY000000000, JAODUO000000000, and JAODUP000000000. The genome versions described in this paper are JAHLWY010000000, JAODUO010000000, and JAODUP010000000. The methylomes are available in NCBI's GEO database under the SuperSeries accession GSE217309. Additional files can be accessed at https://github.com/maepz/deepsea_worms_epigenomes/tree/main/data. All scripts used for the upstream data processing steps and downstream analyses can be accessed at https://github.com/maepz/deepsea_worms_epigenomes.
Conflict of interest statement. The authors declare no competing interests.
References
- Ahmed I, Sarazin A, Bowler C, Colot V, Quesneville H. 2011. Genome-wide evidence for local DNA methylation spreading from small RNA-targeted sequences in Arabidopsis. Nucleic Acids Res. 39:6919–6931. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Angers B, Perez M, Menicucci T, Leung C. 2020. Sources of epigenetic variation and their applications in natural populations. Evol Appl. 13:1262–1278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aroh O, Halanych KM. 2021. Genome-wide characterization of LTR retrotransposons in the non-model deep-sea annelid Lamellibrachia luymesi. BMC Genomics. 22:466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bao W, Kojima KK, Kohany O. 2015. Repbase update, a database of repetitive elements in eukaryotic genomes. Mob DNA. 6:11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barau J, Teissandier A, Zamudio N, Roy S, Nalesso V, Hérault Y, Guillou F, Bourc’his D. 2016. The DNA methyltransferase DNMT3C protects male germ cells from transposon activity. Science 354:909–912. [DOI] [PubMed] [Google Scholar]
- Bewick AJ, Sanchez Z, Mckinney EC, Moore AJ, Moore PJ, Schmitz RJ. 2019. Dnmt1 is essential for egg production and embryo viability in the large milkweed bug, Oncopeltus fasciatus. Epigenetics Chromatin. 12:6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bicho RC, Scott-Fordsmand JJ, Amorim MJB. 2020. Developing an epigenetics model species—from blastula to mature adult, life cycle methylation profile of Enchytraeus crypticus (Oligochaete). Sci Total Environ. 732:139079. [DOI] [PubMed] [Google Scholar]
- Buchfink B, Reuter K, Drost H-G. 2021. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat Methods. 18:366–368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cario A, Jebbar M, Thiel A, Kervarec N, Oger PM. 2016. Molecular chaperone accumulation as a function of stress evidences adaptation to high hydrostatic pressure in the piezophilic archaeon Thermococcus barophilus. Sci Rep. 6:29483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen N. 2004. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr Protoc Bioinformatics. Chapter 4:Unit 4.10. [DOI] [PubMed] [Google Scholar]
- Coulondre C, Miller JH, Farabaugh PJ, Gilbert W. 1978. Molecular basis of base substitution hotspots in Escherichia coli. Nature 274:775–780. [DOI] [PubMed] [Google Scholar]
- Cuvelier D, Sarradin P-M, Sarrazin J, Colaço A, Copley JT, Desbruyères D, Glover AG, Santos RS, Tyler PA. 2011. Hydrothermal faunal assemblages and habitat characterisation at the Eiffel Tower edifice (Lucky Strike, Mid-Atlantic Ridge). Mar Ecol. 32:243–255. [Google Scholar]
- de la Chaux N, Wagner A. 2011. BEL/Pao retrotransposons in metazoan genomes. BMC Evol Biol. 11:154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Mendoza A, Hatleberg WL, Pang K, Leininger S, Bogdanovic O, Pflueger J, Buckberry S, Technau U, Hejnol A, Adamska M, et al. 2019. Convergent evolution of a vertebrate-like methylome in a marine sponge. Nat Ecol Evol. 3:1464–1473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Mendoza A, Lister R, Bogdanovic O. 2020. Evolution of DNA methylome diversity in eukaryotes. J Mol Biol. 432:1687–1705. [DOI] [PubMed] [Google Scholar]
- de Mendoza A, Pflueger J, Lister R. 2019. Capture of a functionally active methyl-CpG binding domain by an arthropod retrotransposon family. Genome Res. 29:1277–1286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deniz Ö, Frost JM, Branco MR. 2019. Regulation of transposable elements by DNA modifications. Nat Rev Genet. 20:417–431. [DOI] [PubMed] [Google Scholar]
- de Oliveira AL, Mitchell J, Girguis P, Bright M. 2022. Novel insights on obligate symbiont lifestyle and adaptation to chemosynthetic environment as revealed by the giant tubeworm genome. Mol Biol Evol. 39, msab347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dilly GF, Young CR, Lane WS, Pangilinan J, Girguis PR. 2012. Exploring the limit of metazoan thermal tolerance via comparative proteomics: thermally induced changes in protein abundance by two hydrothermal vent polychaetes. Proc Biol Sci. 279:3347–3356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duan JE, Jiang ZC, Alqahtani F, Mandoiu I, Dong H, Zheng X, Marjani SL, Chen J, Tian XC. 2019. Methylome dynamics of bovine gametes and in vivo early embryos. Front Genet. 10:512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ehrlich M, Norris KF, Wang RY, Kuo KC, Gehrke CW. 1986. DNA cytosine methylation and heat-induced deamination. Biosci Rep. 6:387–393. [DOI] [PubMed] [Google Scholar]
- Finkelstein JD. 1990. Methionine metabolism in mammals. J Nutr Biochem. 1:228–237. [DOI] [PubMed] [Google Scholar]
- Flores K, Wolschin F, Corneveaux JJ, Allen AN, Huentelman MJ, Amdam GV. 2012. Genome-wide association between DNA methylation and alternative splicing in an invertebrate. BMC Genomics. 13:480. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fontecave M, Atta M, Mulliez E. 2004. S-adenosylmethionine: nothing goes to waste. Trends Biochem Sci. 29:243–249. [DOI] [PubMed] [Google Scholar]
- Gagnière N, Jollivet D, Boutet I, Brélivet Y, Busso D, Da Silva C, Gaill F, Higuet D, Hourdez S, Knoops B, et al. 2010. Insights into metazoan evolution from Alvinella pompejana cDNAs. BMC Genomics. 11:634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gatzmann F, Falckenhayn C, Gutekunst J, Hanna K, Raddatz G, Carneiro VC, Lyko F. 2018. The methylome of the marbled crayfish links gene body methylation to stable expression of poorly accessible genes. Epigenet Chromatin. 11:57. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gavery MR, Roberts SB. 2014. A context dependent role for DNA methylation in bivalves. Briefings Funct Genomics. 13:217–222. [DOI] [PubMed] [Google Scholar]
- Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, Adiconis X, Fan L, Raychowdhury R, Zeng Q, et al. 2011. Trinity: reconstructing a full-length transcriptome without a genome from RNA-Seq data. Nat Biotechnol. 29:644–652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harrell FEJ. 2019. Package ‘hmisc.’ CRAN2018 2019:235–236.
- Holt C, Yandell M. 2011. MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinformatics. 12:491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ihaka R, Gentleman R. 1996. R: a language for data analysis and graphics. J Comput Graphical Stat. 5:299–314. [Google Scholar]
- Jiang Q, Lin D, Huang H, Wang G, Ye H. 2020. DNA methylation inhibits the expression of CFSH in mud crab. Front Endocrinol (Lausanne). 11:263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jun J, Won Y-J, Vrijenhoek RC. 2016. Complete mitochondrial genome of the hydrothermal vent tubeworm, Ridgeia piscesae (Polychaeta, Siboglinidae). Mitochondrial DNA A DNA Mapp Seq Anal. 27:1123–1124. [DOI] [PubMed] [Google Scholar]
- Kaminker JS, Bergman CM, Kronmiller B, Carlson J, Svirskas R, Patel S, Frise E, Wheeler DA, Lewis SE, Rubin GM, et al. 2002. The transposable elements of the Drosophila melanogaster euchromatin: a genomics perspective. Genome Biol. 3:RESEARCH0084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keller TE, Han P, Yi SV. 2016. Evolutionary transition of promoter and gene body DNA methylation across invertebrate–vertebrate boundary. Mol Biol Evol. 33:1019–1028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim A, Terzian C, Santamaria P, Pélisson A, Purd’homme N, Bucheton A. 1994. Retroviruses in invertebrates: the gypsy retrotransposon is apparently an infectious retrovirus of Drosophila melanogaster. Proc Natl Acad Sci U S A. 91:1285–1289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kojima KK. 2019. Structural and sequence diversity of eukaryotic transposable elements. Genes Genet Syst. 94:233–252. [DOI] [PubMed] [Google Scholar]
- Kvist S, Oceguera-Figueroa A. 2021. Phylum Annelida. In: DeSalle R, Schierwater B, editors. Invertebrate zoology. Boca Raton (FL): CRC Press. p. 311–328. [Google Scholar]
- Lechner M, Hernandez-Rosales M, Doerr D, Wieseke N, Thévenin A, Stoye J, Hartmann RK, Prohaska SJ, Stadler PF. 2014. Orthology detection combining clustering and synteny for very large datasets. PLoS One 9:e105015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee RW, Robert K, Matabos M, Bates AE, Juniper SK. 2015. Temporal and spatial variation in temperature experienced by macrofauna at Main Endeavour hydrothermal vent field. Deep Sea Res 1 Oceanogr Res Pap. 106:154–166. [Google Scholar]
- Lewis SH, Ross L, Bain SA, Pahita E, Smith SA, Cordaux R, Miska EA, Lenhard B, Jiggins FM, Sarkies P. 2020. Widespread conservation and lineage-specific diversification of genome-wide DNA methylation patterns across arthropods. PLoS Genet. 16:e1008864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Y, Kocot KM, Whelan NV, Santos SR, Waits DS, Thornhill DJ, Halanych KM. 2017. Phylogenomics of tubeworms (Siboglinidae, Annelida) and comparative performance of different reconstruction methods. Zool Scr. 46:200–213. [Google Scholar]
- Li Y, Tassia MG, Waits DS, Bogantes VE, David KT, Halanych KM. 2019. Genomic adaptations to chemosymbiosis in the deep-sea seep-dwelling tubeworm Lamellibrachia luymesi. BMC Biol. 17:91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y, Rosikiewicz W, Pan Z, Jillette N, Wang P, Taghbalout A, Foox J, Mason C, Carroll M, Cheng A, et al. 2021. DNA methylation-calling tools for Oxford Nanopore sequencing: a survey and human epigenome-wide evaluation. Genome Biol. 22:295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lyko F. 2018. The DNA methyltransferase family: a versatile toolkit for epigenetic regulation. Nat Rev Genet. 19:81–92. [DOI] [PubMed] [Google Scholar]
- Lyko F, Foret S, Kucharski R, Wolf S, Falckenhayn C, Maleszka R. 2010. The honey bee epigenomes: differential methylation of brain DNA in queens and workers. PLoS Biol. 8:e1000506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marsh AG, Pasqualone AA. 2014. DNA methylation and temperature stress in an Antarctic polychaete, Spiophanes tcherniai. Front Physiol. 5:173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McCaw BA, Stevenson TJ, Lancaster LT. 2020. Epigenetic responses to temperature and climate. Integr Comp Biol. 60:1469–1480. [DOI] [PubMed] [Google Scholar]
- Moggioli G, Panossian B, Sun Y, Thiel D, Martín-Zamora FM, Tran M, Clifford AM, Goffredi SK, Rimskaya-Korsakova N, Jékely G, et al. 2023. Distinct genomic routes underlie transitions to specialised symbiotic lifestyles in deep-sea annelid worms. Nat Commun. 14:2814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Neri F, Rapelli S, Krepelova A, Incarnato D, Parlato C, Basile G, Maldotti M, Anselmi F, Oliviero S. 2017. Intragenic DNA methylation prevents spurious transcription initiation. Nature 543:72–77. [DOI] [PubMed] [Google Scholar]
- Newbold LK, Robinson A, Rasnaca I, Lahive E, Soon GH, Lapied E, Oughton D, Gashchak S, Beresford NA, Spurgeon DJ. 2019. Genetic, epigenetic and microbiome characterisation of an earthworm species (Octolasion lacteum) along a radiation exposure gradient at Chernobyl. Environ Pollut. 255:113238. [DOI] [PubMed] [Google Scholar]
- Ogunlaja A, Sharma V, Ghai M, Lin J. 2020. Molecular characterization and DNA methylation profile of Libyodrilus violaceous from oil polluted soil. Mol Biol Res Commun. 9:45–53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patro R, Duggal G, Love MI, Irizarry RA, Kingsford C. 2017. Salmon: fast and bias-aware quantification of transcript expression using dual-phase inference. Nat Methods. 14:417–419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pehrsson EC, Choudhary MNK, Sundaram V, Wang T. 2019. The epigenomic landscape of transposable elements across normal human development and anatomy. Nat Commun. 10:5640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pelisson A, Mejlumian L, Robert V, Terzian C, Bucheton A. 2002. Drosophila germline invasion by the endogenous retrovirus gypsy: involvement of the viral env gene. Insect Biochem Mol Biol. 32:1249–1256. [DOI] [PubMed] [Google Scholar]
- Perez M, Juniper SK. 2016. Insights into symbiont population structure among three vestimentiferan tubeworm host species at eastern pacific spreading centers. Appl Environ Microbiol. 82:5197–5205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perez M, Wang H, Angers B, Qian P-Y. 2022. Complete mitochondrial genome of Paralvinella palmiformis (Polychaeta: Alvinellidae). Mitochondrial DNA B Resour. 7:786–788. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piednoël M, Donnart T, Esnault C, Graça P, Higuet D, Bonnivard E. 2013. LTR-retrotransposons in R. exoculata and other crustaceans: the outstanding success of GalEa-like copia elements. PLoS One 8:e57675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Planques A, Kerner P, Ferry L, Grunau C, Gazave E, Vervoort M. 2021. DNA methylation atlas and machinery in the developing and regenerating annelid Platynereis dumerilii. BMC Biol. 19:148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Provataris P, Meusemann K, Niehuis O, Grath S, Misof B. 2018. Signatures of DNA methylation across insects suggest reduced DNA methylation levels in Holometabola. Genome Biol Evol. 10:1185–1197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Quinlan AR, Hall IM. 2010. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26:841–842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Riesgo A, Andrade SC, Sharma PP, Novo M, Pérez-Porro AR, Vahtera V, González VL, Kawauchi GY, Giribet G. 2012. Comparative description of ten transcriptomes of newly sequenced invertebrates and efficiency estimation of genomic sampling in non-model taxa. Front Zool. 9:33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ritchie H, Jamieson AJ, Piertney SB. 2018. Heat-shock protein adaptation in abyssal and hadal amphipods. Deep Sea Res 2 Top Stud Oceanogr. 155:61–69. [Google Scholar]
- Sambrook J, Fritsch EF, Maniatis T. 1989. Molecular cloning: a laboratory manual. New York: Cold spring Harbor Laboratory Press. [Google Scholar]
- Sanchez-Perez GF, Hampl V, Simpson AG, Roger AJ. 2008. A new divergent type of eukaryotic methionine adenosyltransferase is present in multiple distantly related secondary algal lineages. J Eukaryotic Microbiol. 55:374–381. [DOI] [PubMed] [Google Scholar]
- Simão FA, Waterhouse RM, Ioannidis P, Kriventseva EV, Zdobnov EM. 2015. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31:3210–3212. [DOI] [PubMed] [Google Scholar]
- Simpson JT, Workman RE, Zuzarte PC, David M, Dursi LJ, Timp W. 2017. Detecting DNA cytosine methylation using nanopore sequencing. Nat Methods. 14:407–410. [DOI] [PubMed] [Google Scholar]
- Smallwood SA, Kelsey G. 2012. De novo DNA methylation: a germ cell perspective. Trends Genet. 28:33–42. [DOI] [PubMed] [Google Scholar]
- Smith MD, Wertheim JO, Weaver S, Murrell B, Scheffler K, Kosakovsky Pond SL. 2015. Less is more: an adaptive branch-site random effects model for efficient detection of episodic diversifying selection. Mol Biol Evol. 32:1342–1353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stiller J, Tilic E, Rousset V, Pleijel F, Rouse GW. 2020. Spaghetti to a tree: a robust phylogeny for Terebelliformia (Annelida) based on transcriptomes, molecular and morphological data. Biology (Basel). 9:73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stipanuk MH. 2020. Metabolism of sulfur-containing amino acids: how the body copes with excess methionine, cysteine, and sulfide. J Nutr. 150:2494S–2505S. [DOI] [PubMed] [Google Scholar]
- Storer J, Hubley R, Rosen J, Wheeler TJ, Smit AF. 2021. The Dfam community resource of transposable element families, sequence models, and genome annotations. Mob DNA. 12:2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strader ME, Kozal LC, Leach TS, Wong JM, Chamorro JD, Housh MJ, Hofmann GE. 2020. Examining the role of DNA methylation in transcriptomic plasticity of early stage sea urchins: developmental and maternal effects in a kelp forest herbivore. Front Mar Sci. 7:205. [Google Scholar]
- Sun Y, Sun J, Yang Y, Lan Y, Ip JC-H, Wong WC, Kwan YH, Zhang Y, Han Z, Qiu J-W, et al. 2021. Genomic signatures supporting the symbiosis and formation of chitinous tube in the deep-sea tubeworm Paraescarpia echinospica. Mol Biol Evol. 38:4116–4134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suzuki MM, Bird A. 2008. DNA methylation landscapes: provocative insights from epigenomics. Nat Rev Genet. 9:465–476. [DOI] [PubMed] [Google Scholar]
- Thomas-Bulle C, Piednoël M, Donnart T, Filée J, Jollivet D, Bonnivard É. 2018. Mollusc genomes reveal variability in patterns of LTR-retrotransposons dynamics. BMC Genomics. 19:821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tilic E, Sayyari E, Stiller J, Mirarab S, Rouse GW. 2020. More is needed—thousands of loci are required to elucidate the relationships of the ‘flowers of the sea’ (Sabellida, Annelida). Mol Phylogenet Evol. 151:106892. [DOI] [PubMed] [Google Scholar]
- Todgham AE, Crombie TA, Hofmann GE. 2017. The effect of temperature adaptation on the ubiquitin–proteasome pathway in notothenioid fishes. J Exp Biol. 220:369–378. [DOI] [PubMed] [Google Scholar]
- Tunnicliffe V, Germain CS, Hilário A. 2014. Phenotypic variation and fitness in a metapopulation of tubeworms (Ridgeia piscesae, Jones) at hydrothermal vents. PLoS One. 9:e110578. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang X, Bhandari RK. 2019. DNA methylation dynamics during epigenetic reprogramming of medaka embryo. Epigenetics 14:611–622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang X, Li Q, Lian J, Li L, Jin L, Cai H, Xu F, Qi H, Zhang L, Wu F, et al. 2014. Genome-wide and single-base resolution DNA methylomes of the Pacific oyster Crassostrea gigas provide insight into the evolution of invertebrate CpG methylation. BMC Genomics. 15:1119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang M, Ruan L, Liu M, Liu Z, He J, Zhang L, Wang Y, Shi H, Chen M, Yang F, et al. 2023. The genome of a vestimentiferan tubeworm (Ridgeia piscesae) provides insights into its adaptation to a deep-sea environment. BMC Genomics. 24:72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weber AA-T, Hugall AF, O’Hara TD. 2020. Convergent evolution and structural adaptation to the deep ocean in the protein-folding chaperonin CCTα. Genome Biol Evol. 12:1929–1942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wertheim JO, Murrell B, Smith MD, Kosakovsky Pond SL, Scheffler K. 2015. RELAX: detecting relaxed selection in a phylogenetic framework. Mol Biol Evol. 32:820–832. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu J-S, Xia Q-Y, Li J, Pan G-Q, Zhou Z-Y. 2005. Survey of long terminal repeat retrotransposons of domesticated silkworm (Bombyx mori). Insect Biochem Mol Biol. 35:921–929. [DOI] [PubMed] [Google Scholar]
- Yan G, Lan Y, Sun J, Xu T, Wei T, Qian P-Y. 2022. Comparative transcriptomic analysis of in situ and onboard fixed deep-sea limpets reveals sample preparation-related differences. iScience 25:104092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ying H, Hayward DC, Klimovich A, Bosch TCG, Baldassarre L, Neeman T, Forêt S, Huttley G, Reitzel AM, Fraune S, et al. 2022. The role of DNA methylation in genome defense in Cnidaria and other invertebrates. Mol Biol Evol. 39:msac018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yuen ZW-S, Srivastava A, Daniel R, McNevin D, Jack C, Eyras E. 2021. Systematic benchmarking of tools for CpG methylation detection from nanopore sequencing. Nat Commun. 12:3438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang X, Yazaki J, Sundaresan A, Cokus S, Chan SW, Chen H, Henderson IR, Shinn P, Pellegrini M, Jacobsen SE, et al. 2006. Genome-wide high-resolution mapping and functional analysis of DNA methylation in Arabidopsis. Cell. 126:1189–1201. [DOI] [PubMed] [Google Scholar]
- Zhao Y, Xu T, Law YS, Feng D, Li N, Xin R, Wang H, Ji F, Zhou H, Qiu J-W. 2020. Ecological characterization of cold-seep epifauna in the South China Sea. Deep Sea Res 1 Oceanogr Res Pap. 163:103361. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Raw sequence reads are available in NCBI's SRA database under the accession numbers SRR21707426–SRR21707428 (P. palmiformis Illumina), SRR217033–SRR217038 (Nanopore reads of R. piscesae and P. palmiformis), and SRR21707446–SRR21707448 (R. piscesae Illumina). The whole-genome shotgun projects for P. echinospica, R. piscesae, and P. palmiformis were deposited in GenBank under accession numbers JAHLWY000000000, JAODUO000000000, and JAODUP000000000. The genome versions described in this paper are JAHLWY010000000, JAODUO010000000, and JAODUP010000000. The methylomes are available in NCBI's GEO database under the SuperSeries accession GSE217309. Additional files can be accessed at https://github.com/maepz/deepsea_worms_epigenomes/tree/main/data. All scripts used for the upstream data processing steps and downstream analyses can be accessed at https://github.com/maepz/deepsea_worms_epigenomes.
Conflict of interest statement. The authors declare no competing interests.