Abstract
The negative-sense segmented RNA genome of influenza virus is transcribed into capped and polyadenylated mRNAs, as well as full-length replicative intermediates (cRNAs). The mechanism that regulates the two forms of transcription remains unclear, although several lines of evidence imply a role for the viral nucleoprotein (NP). In particular, temperature-shift and biochemical analyses of the temperature-sensitive viruses A/WSN/33 ts56 and A/FPV/Rostock/34/Giessen tsG81 containing point mutations within the NP coding region have indicated specific defects in replicative transcription at the nonpermissive temperature. To identify the functional defect, we introduced the relevant mutations into the NP of influenza virus strain A/PR/8/34. Both mutants were temperature sensitive for influenza virus gene expression in transient-transfection experiments but localized and accumulated normally in transfected cells. Similarly, the mutants retained the ability to self-associate and interact with the virus polymerase complex whether synthesized at the permissive or the nonpermissive temperatures. In contrast, the mutant NPs were defective for RNA binding when expressed at the nonpermissive temperature but not when expressed at 30°C. This suggests that the RNA-binding activity of NP is required for replicative transcription.
The influenza A virus genome consists of eight segments of negative-sense single-stranded RNA, which encode 10 identified polypeptides. These polypeptides are expressed through transcription of the virion-associated genomic RNAs (vRNA) into capped and polyadenylated positive-sense mRNAs. The 5′-cap structure is not synthesized de novo by virus enzymes but is recycled along with 10 to 15 nucleotides from host cell heterogeneous nuclear RNAs by endonucleolytic cleavage and used to prime influenza virus transcription (36). The poly(A) tail is generated by repetitive transcription of a short stretch of uridines located 15 to 22 bases from the 5′ end of the template, beyond which processive transcription does not occur (28). The non-template-directed 5′ and 3′ modifications to viral mRNAs render them ineffective as substrates for the synthesis of new vRNA segments. Progeny vRNAs are instead synthesized via a replicative intermediate plus sense transcript (cRNA) that is not capped, contains no host-cell sequences, and is not polyadenylated (reviewed in reference 24). Four viral proteins are necessary and sufficient to carry out these processes (19); the three subunits of an RNA-dependent RNA polymerase (the P proteins PB1, PB2, and PA) and a single-stranded RNA-binding nucleoprotein (NP). Indeed, the functional form of the v- and cRNA segments are always associated with these polypeptides to form ribonucleoprotein (RNP) structures, which are thought to be composed of one copy of the trimeric polymerase and approximately one NP polypeptide per 20 bases of RNA (24). The mechanisms underlying viral mRNA transcription are comparatively well understood. The polymerase complex binds to the 5′ end of the vRNA template (47) in a step which activates cap recognition by the PB2 subunit (7). Subsequent interaction with the 3′ end of vRNA via a combination of base-pairing between terminal inverted repeats (18) and direct protein-RNA contacts with the PB1 subunit (25) activates endonuclease activity (15), which is possibly mediated by the PB2 subunit (5, 42). PB1 then initiates transcription by using the capped RNA fragment as a primer (Braam et al. [6]). Polyadenylation is thought to occur because the polymerase remains bound to the 5′ sequences immediately upstream of the polyuridine stretch, preventing processive transcription through the latter region (38, 39, 47).
In contrast, the mechanisms which induce the influenza virus polymerase to switch from transcribing mRNAs to making replicative intermediate cRNAs remain relatively ill defined. The form of RNP packaged into virions will only synthesize mRNA in vitro (44), and although the same vRNA templates are transcribed into cRNA after infection of cells, an initial round of mRNA transcription and subsequent protein expression is essential (16). This transcriptional switch is likely to be multifactorial: genetic evidence points to the involvement of the viral polymerase (29), and the direct participation of host cell proteins is a possibility (43). In addition, multiple lines of genetic and biochemical evidence implicate the virus NP as a major factor. Several conditional lethal virus mutants with lesions in the NP gene have been isolated that are specifically defective for replicative transcription (22, 30, 40, 41, 46), and infected cell extracts that synthesize c- and vRNA in vitro depend on a supply of non-RNP-associated NP (3, 41). Moreover, by using a combination of the genetic and biochemical approaches, Shapiro and Krug (41) showed that extracts from cells infected with the mutant A/WSN/33 ts56 virus (containing a lesion in the NP gene) synthesized m-, c- and vRNA in vitro at the permissive temperature but only mRNA at the nonpermissive temperature. Nevertheless, we do not understand the molecular mechanisms by which NP functions in replicative transcription.
NP is a multifunctional protein, capable of binding RNA (37) and a variety of viral and cellular proteins, including itself (37), two of the three subunits of the viral polymerase (4), cellular importin α (35), and filamentous actin (10). Since the function of NP in mRNA transcription is genetically and biochemically distinguishable from its replicative function, we reasoned that the two roles might require different subsets of the individual activities of the polypeptide. Moreover, identification of the NP functions required for genome replication would be informative as to the mechanisms involved. We therefore examined the biochemical activities of mutant NP molecules containing temperature-sensitive (ts) lesions which disrupt replicative transcription. The ts mutations from A/WSN/33 (WSN) ts56 and A/FPV/Rostock/34 (FPV) tsG81 conferred temperature sensitivity on the ability of the A/PR/8/34 NP to support influenza virus gene expression, without affecting the cellular localization or accumulation of the polypeptides. The mutations did not affect the ability of the polypeptides to make NP-NP or NP-P protein-protein contacts, but they did induce ts RNA-binding activity. This suggests that the RNA-binding activity of NP is essential for replicative transcription.
MATERIALS AND METHODS
Plasmids and protein expression.
Plasmids containing the wild-type A/PR/8/34 PB1, PB2 PA and NP genes under the control of a bacteriophage T7 RNA polymerase promoter (pKT1, -2, -3, and -5, respectively) or lacking a heterologous gene (pKT0) have previously been described (5, 10). Plasmids containing the NP gene fused to either Escherichia coli maltose-binding protein (pMAL-NP) or Schistosoma japonicum glutathione S-transferase (GST) (pGEX-NP) have also been reported (10). Point mutations were introduced by using oligonucleotide-directed mutagenesis according to standard procedures (23). Plasmid pPB2CAT9 containing an antisense chloramphenicol acetyltransferase (CAT) gene flanked by the 5′- and 3′-noncoding regions from influenza virus segment 1 was the generous gift of Mark Krystal. Rabbit polyclonal antisera directed against influenza FPV RNPs or amino acids 340 to 498 of A/PR/8/34 NP have already been described (10).
Radiolabelled NP polypeptides were synthesized by using a coupled in vitro transcription-translation system (8). Rabbit reticulocyte lysate (Promega) was supplemented with 0.6 μCi of [35S]methionine per ml, 2 U of T7 RNA polymerase (Gibco-BRL) per ml, 3.3 mM MgCl2, 0.2 mM nucleoside triphosphates, and 20 to 100 μg of plasmid DNA per ml before incubation at 30 or 37°C for 90 min.
Xenopus laevis oocytes were maintained and microinjected with synthetic mRNAs encoding the influenza virus P proteins (transcribed from plasmids pKT 1 to 3) as previously described (5, 9).
Maltose-binding protein (MBP) or MBP fused to NP (MBP-NP) were purified from extracts of E. coli cultures containing plasmids pMAL-c2 (New England Biolabs) or pMAL-NP (or mutant derivatives thereof) by affinity chromatography on amylose resin columns (New England Biolabs) as previously described (10). GST or GST fused to NP (GST-NP) were similarly obtained from E. coli cultures containing plasmid pGEX-4T1 (Pharmacia) or plasmid pGEX-NP as described earlier (10).
Transfection of tissue culture cells, transient replication assays, and indirect immunofluorescence.
CV1 or BHK cells were infected with recombinant vaccinia viruses separately encoding the three subunits of the influenza virus polymerase (PB1-, PB2-, and PA-VAC [45]) and bacteriophage T7 RNA polymerase (VTF7 [14]) at a multiplicity of infection of 5 of each virus for 2 h at 37°C. The cells were washed three times with serum-free medium before transfection with plasmid DNA encoding NP and pPB2CAT9 in vitro-transcribed RNA by using a cationic liposome mixture (Lipofectin [GIBCO-BRL] or Escort [Sigma-Aldrich]) as previously described (10). After 20 to 24 h incubation at 30 to 31°C (the permissive temperature [PT]) or 37°C (the nonpermissive temperature [NPT]), the cells were lysed, and the accumulation of CAT polypeptide was measured by using a commercial enzyme-linked immunosorbent assay (ELISA; Boehringer Mannheim). The NPT was defined as 37°C throughout these experiments because the wild-type (WT) influenza virus components failed to produce detectable quantities of CAT polypeptide in the transfection system at the more usual NPTs of 39.5 or 40.5°C (data not shown).
For immunofluorescence analysis, BHK cells on coverslips were infected with VTF7 only and transfected with plasmid DNA as described above. After 4 h of incubation at 31 or 37°C, the cells were washed with phosphate-buffered saline (PBS) containing 1% newborn calf serum, fixed in PBS containing 4% formaldehyde, and stained for NP by using anti-RNP serum as previously described (10). Fluorescence was viewed on an Olympus IX70 microscope, and images were captured by using an Hamamatsu Colour Chilled 3CCD video camera.
RNA transcription and binding assays.
For in vitro RNA binding assays, a radiolabelled 178-nucleotide synthetic RNA target corresponding to segment 8 but with nucleotides 84 to 795 (inclusive) deleted, and a C-to-A transversion of the penultimate base was generated by in vitro transcription of plasmid pKT8-Δ3′5′ with bacteriophage T7 RNA polymerase in the presence of [α-32P]CTP (100 μM; specific activity, 2 Ci/mmol) as previously described (10). Filter binding assays were performed by incubating protein samples with 20 fmol of RNA (ca. 5,000 cpm) in 25 mM Tris-Cl (pH 7.6)–50 mM NaCl–5 mM MgCl2–0.5 mM dithiothreitol–5% glycerol at room temperature for 20 min. The reaction mixtures were filtered under vacuum through nitrocellulose membrane equilibrated in 20 mM Tris-Cl (pH 7.6)–50 mM NaCl and washed three times with 200 μl of the same buffer. Bound radioactivity was quantified by liquid scintillation counting. For UV cross-linking experiments, reactions were incubated for 20 min at room temperature as described above before irradiation at 254 nm for 5 min at an intensity of 4 mW/cm2 in a Spectronics Corporation XL-1500 UV cross-linker. Free RNA was removed by digestion with 2 μg of RNase A per ml for 30 min at room temperature before analysis by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and autoradiography.
Protein binding assays.
For precipitation reactions, 1 μl of in vitro-translated protein or 2.5 μl of oocyte lysate (corresponding to one-quarter of an oocyte) was incubated with 0.5 μg of fusion protein in 100 μl of IP buffer (100 mM KCl; 50 mM Tris-Cl, pH 7.6; 5 mM MgCl2; 1 mM dithiothreitol; 0.1% Nonidet P-40) for 1 h at room temperature. Then, 50 μl of a 50% (vol/vol) slurry of glutathione-Sepharose (Pharmacia) or amylose resin (New England Biolabs) in PBS was added as appropriate, and incubation continued for a further 30 min with gentle mixing. The solid phase was collected by centrifugation and washed three times with 750 μl of IP buffer. Bound material was eluted by boiling in 40 μl of SDS-PAGE sample buffer and then analyzed by SDS-PAGE and autoradiography.
RESULTS
Induction of temperature sensitivity in the A/PR/8/34 strain NP.
For a functional study of the NPs of WSN ts56 and FPV tsG81, we needed sources of recombinant protein. Although we did not possess cDNA clones of the WT or mutant NP genes from either virus strain, the ts lesions have been identified by comparison of the sequences of mutant and parental viruses (26, 31). We therefore introduced the relevant mutations into a cDNA clone of the influenza A/PR/8/34 (PR8) strain NP (plasmid pKT5 [10]) to create plasmids pKT5 S314-N and pKT5 A332-T. Although the WT WSN and FPV NP amino acid sequences are closely related to that of PR8 (96 and 94% identical, respectively [26, 31, 50]), including identity at residues 314 and 332, we examined the ts phenotype of the mutations in the PR8 strain NP. To test the ability of the mutant genes to support influenza virus gene expression, CV1 cells were infected with recombinant vaccinia viruses expressing the three influenza virus P proteins and bacteriophage T7 RNA polymerase and then were transfected with WT or mutant pKT5 plasmids and an RNA containing an antisense CAT gene flanked by the promoter sequences from influenza virus segment 1 (PB2-CAT). After maintenance at either the PT (31°C) or the NPT (37°C) for 20 h, the cells were lysed and the accumulation of CAT polypeptide was measured (10). The synthetic influenza virus segment was recognized by the WT components of the virus transcriptase and transcribed and replicated to yield CAT polypeptide expression (data not shown). Figure 1 shows the activities of the mutants relative to the WT gene at each temperature: the S314 polypeptide supported the synthesis of essentially WT quantities of CAT at 31°C, while the A332 mutant showed substantial activity, producing about two-thirds the amount made in the presence of WT NP (Fig. 1A). However, the activity of both mutants was much reduced at 37°C, producing on average less than 10% of the amount seen with the WT gene (Fig. 1). Similar results were obtained when the plasmids were titrated, with the mutants showing equal temperature sensitivity at plasmid doses between 1 μg and 10 ng/2 × 105 cells (data not shown). Thus, the mutations confer temperature sensitivity on the transcriptional function of PR8 strain NP. To ascertain whether the failure of the mutants to support CAT synthesis at 37°C resulted from differential stability of the mutants at the NPT, we analyzed parallel transfections by SDS-PAGE and Western blotting with an anti-NP serum. Both mutants expressed NP at levels similar to those of the WT at 31 and 37°C and accumulated to higher levels at 37°C (Fig. 1B). This indicates that the mutations do not significantly affect the intracellular stability of either polypeptide.
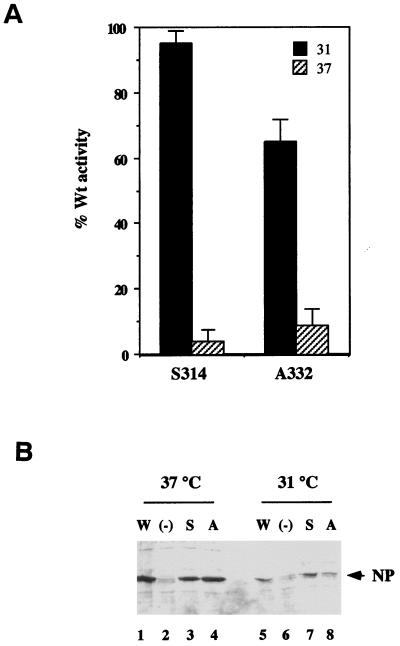
FIG. 1.
Transcriptional activity and cellular accumulation of NP mutants. (A) The ability of the indicated mutants to support virus gene expression at 31 or 37°C was measured by transfecting CV1 cells containing the influenza virus RNA polymerase with plasmids pKT5, pKT5 S314-N, or A332-T and a synthetic influenza RNA segment containing an antisense CAT gene and then measuring the subsequent accumulation of CAT polypeptide. Data are plotted as the percentage (with the standard error) of the value obtained with the WT gene, and result from at least three independent determinations. (B) Cellular accumulation of WT and mutant NPs. CV1 cells were transfected with plasmids encoding WT NP (W), NP S314-N (S), NP A332-T (A), or plasmid vector (pKT0) not containing a heterologous gene (−), maintained at the indicated temperature for 6 h, detergent lysed, and examined for their NP content by SDS-PAGE and Western blotting with an anti-NP serum. The position of NP is indicated by the arrow.
Although NP contains at least two nuclear localization signals (34, 48, 49) and can localize to the nucleus in the absence of other influenza virus proteins (27), more recent work has shown that transiently expressed NP will accumulate in the cytoplasm under certain circumstances (10, 34), similarly to the late stages of authentic virus infection. We have shown that the balance between nuclear import and cytoplasmic accumulation is affected by the amount of NP expressed and its ability to bind filamentous actin (10). We therefore examined the cellular distribution of the mutants at the PT and NPT in cells transfected with various amounts of plasmid by immunofluorescence with anti-NP serum. As with the WT gene (10), transfection of cells with a low dose of plasmid (3 ng/105 cells) resulted in predominantly nuclear accumulation of the mutant polypeptides at the PT (Fig. 2b and f). Similarly, transfection of cells at the PT with a high dose of plasmid (300 ng/105 cells) resulted in substantial cytoplasmic accumulation of the polypeptides (Fig. 2a and e). This pattern of localization was not substantially altered at the NPT, with both mutants still localizing to the nucleus when expressed in low abundance (Fig. 2d and h) and to the cytoplasm when expressed at high level (Fig. 2c and g). Thus, neither ts lesion prevents the polypeptide from localizing normally within the cell.
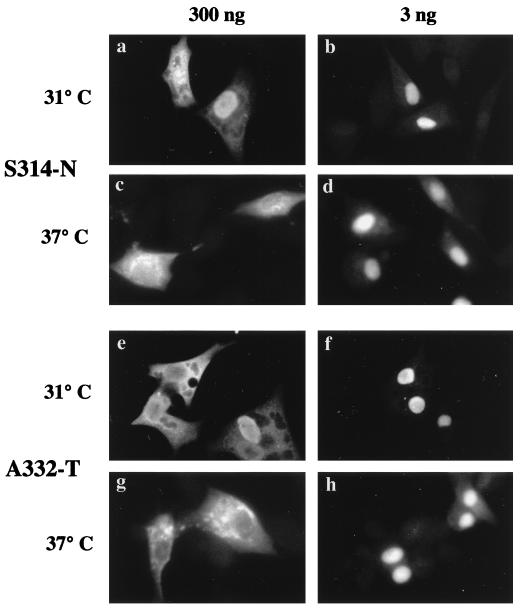
FIG. 2.
Cellular distribution of NP mutants. BHK cells (105) were transfected with 3 or 300 ng of plasmid and maintained for 4 h at 31 or 37°C as labelled. The cells were formaldehyde fixed, detergent permeabilized, and examined for NP content by indirect immunofluorescence with an anti-RNP serum.
Having established that the mutant NPs displayed the expected ts phenotype and having ruled out failure to express or localize to the nucleus as a cause of the defect, we went on to assay individual biochemical functions of the polypeptides. The mutant NP genes were subcloned into plasmid pMAL-NP to allow the expression and affinity purification of MBP-NP fusion proteins from E. coli as previously described (10). Nonfused MBP was prepared from bacteria grown at 37°C, while the WT and mutant fusion proteins were prepared from cultures incubated at 30 or 37°C. By comparison with WT MBP-NP, which expressed well and copurified with only small amounts of a presumably proteolyzed MBP-NP “stub” (Fig. 3, lane 1 [10]), the mutants contained greater quantities of the contaminating shorter polypeptide. However, the ratios of full-length fusion protein to degradation product were not dramatically different between the preparations made at the PT or NPT (Fig. 3, compare lanes 3 and 4 and lanes 5 and 6). In addition, the MBP-NP stub contains only a short NP sequence (compare mobilities with MBP, lane 2) that does not possess detectable RNA binding (see below), oligomerization, or polymerase-binding properties (data not shown) and can therefore be disregarded.
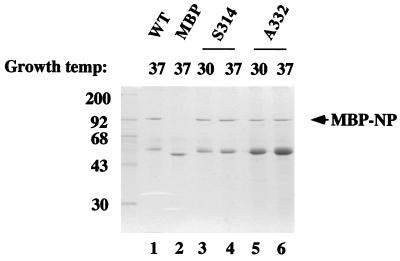
FIG. 3.
Expression and purification of WT and mutant NP fusion proteins. Lysates from E. coli cultures containing plasmids encoding WT MBP-NP (WT), MBP-NP A332-T (A332), MBP-NP S314-N (S314), or MBP were grown at the indicated temperatures and fractionated over amylose resin columns. Bound proteins were eluted by the addition of maltose, dialyzed, and analyzed by SDS-PAGE and stained with Coomassie brilliant blue dye. Also indicated are the sizes of molecular mass markers (in kilodaltons).
Recently, we have determined that in vitro-translated NP binds to NP fusion proteins in a specific and quantitative manner (12). We used this method to test whether the mutant NP polypeptides were defective for NP-NP interactions. Radiolabelled WT NP, NP S314-N, and NP A332-T were prepared by in vitro transcription and translation of the appropriate plasmids in rabbit reticulocyte lysate at the NPT and then tested for their ability to bind immobilized MBP or WT and mutant MBP-NP fusion proteins in a binding assay. Substantial quantities of WT NP translated in vitro bound to WT MBP-NP but not to MBP (Fig. 4, lanes 2 and 3), indicating a specific interaction between the two NP moieties. However, similar quantities of WT NP were bound by the mutant MBP-NP fusion proteins irrespective of whether they were synthesized at the permissive or nonpermissive temperature (lanes 4 to 7). Moreover, the mutant MBP-NP molecules showed little or no temperature dependence in their ability to bind the homologous in vitro-translated mutant (lanes 8 to 15). Thus, the ts lesions have no significant effect on NP oligomerization.
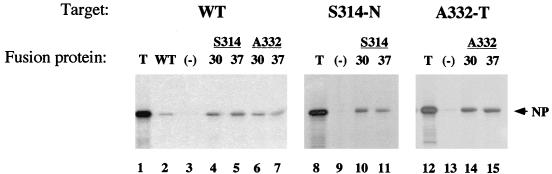
FIG. 4.
Oligomerization of NP mutants. Aliquots of WT, S314-N, or A332-T NP translated in vitro at 37°C were analyzed by SDS-PAGE and autoradiography before (T) or after binding to MBP (−), WT MBP-NP (WT), or mutant MBP-NP fusion protein prepared at the indicated temperature.
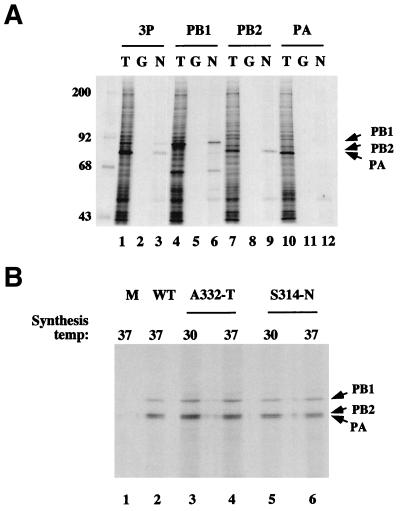
While this study was in progress, Biswas et al. (4) published experiments showing that NP was capable of forming direct protein-protein contacts with the influenza virus polymerase. We also find that NP binds the virus RNA polymerase. Previously, we have shown that the three influenza virus P proteins assemble into a complex in X. laevis oocytes microinjected with the appropriate synthetic mRNAs and that this system provides a convenient source of soluble radiolabelled protein for binding studies (5, 9). When lysates from oocytes microinjected with mRNAs encoding all three P proteins were incubated with GST-NP and glutathione-Sepharose, all three P proteins became associated with the solid phase (Fig. 5A, lane 3). The precipitation was specific, since the majority of the cellular polypeptides did not bind GST-NP (cf. lanes 1 and 3), and no polypeptides were bound by immobilized GST (lane 2). Thus, NP interacts with the polymerase complex. To determine which of the polymerase subunits NP interacted with, we microinjected oocytes with mRNAs encoding the individual P proteins and assayed the resulting lysates for binding to GST or GST-NP. Both PB1 and PB2 were precipitated by GST-NP but not by GST (lanes 5, 6, 8, and 9) but, in contrast, PA failed to bind to either protein (lanes 11 and 12). Therefore, in agreement with the findings of Biswas et al. (4), NP interacts with the two basic P proteins but shows much less affinity for PA. Accordingly, we went on to test the ability of the ts NP mutants to bind the polymerase complex. Oocyte lysates containing the three P proteins were incubated with MBP, WT MBP-NP, or the mutant MBP-NP polypeptides prepared at the PT or NPT and amylose resin; the solid phase was collected, washed, and examined by SDS-PAGE and autoradiography. All three P proteins, including PA (confirmed by Western blot analysis [data not shown]), were precipitated by MBP-NP but not by MBP alone (Fig. 5B, lanes 1 and 2), indicating that NP fused to MBP also interacts with the polymerase complex. Similar quantities of the P proteins were precipitated by the mutant MBP-NP proteins, with no obvious differential activity between polypeptides prepared at 30 or 37°C (lanes 3 to 6). In addition, no differences were found between the abilities of WT and ts mutant MBP-NP fusion proteins to interact with PB1 or PB2 alone (data not shown). Therefore, the ts lesions do not affect the ability of NP to bind the influenza virus polymerase.
FIG. 5.
Ability of WT and mutant NP fusion proteins to bind influenza P proteins. (A) Subunit specificity. Radiolabelled lysates from Xenopus oocytes microinjected with synthetic mRNAs encoding the indicated P proteins (3P corresponds to a mixture of PB1, PB2, and PA mRNAs) were analyzed by SDS-PAGE and autoradiography before (T) or after precipitation with immobilized GST (G) or GST-NP (N). The positions of the P proteins are marked by arrows. The size of molecular mass markers are also indicated (in kilodaltons). (B) Association of WT and mutant MBP-NP fusion proteins with the polymerase complex. Radiolabelled lysates containing all three P proteins were analyzed as described above after precipitation with MBP (M), WT MBP-NP (WT), or mutant MBP-NP fusion proteins prepared at the indicated temperatures.
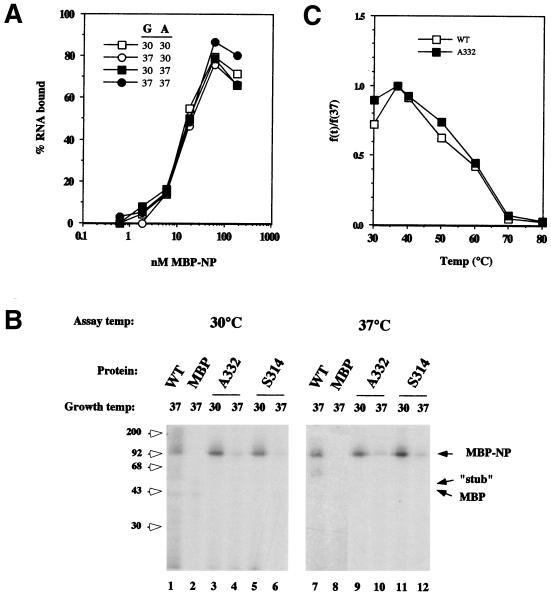
Previously, we have shown that MBP-NP (grown at 37°C) binds RNA with an affinity similar to that of authentic NP purified from virions and that MBP does not possess measurable binding activity (10). We tested the effects of growth and assay temperature on WT MBP-NP by preparing protein at 30 or 37°C and then titrating the polypeptides’ RNA binding activities at the two temperatures. At ca. 20 nM MBP-NP grown at 37°C was required to retain 50% of a 1 nM solution of a radiolabelled RNA on nitrocellulose filters at 37°C (Fig. 6a), which under the assay conditions corresponds to an estimate of the dissociation constant of the interaction (10). The binding curves derived from MBP-NP grown at 37°C and assayed at 30°C or from MBP-NP grown at 30°C and assayed at either temperature were essentially indistinguishable (Fig. 6A). In replicate experiments, the MBP-NP:RNA dissociation constants for protein grown at 30 or 37°C and assayed at 37°C were found to be 14.7 ± 5.2 and 16.7 ± 7.0 nM, respectively (n = 3), which is in good agreement with the values previously measured for MBP-NP at room temperature (10) or for authentic NP at 37°C (2). Thus, the RNA-binding activity of WT MBP-NP is invariant over the PT and NPT ranges used here. Next, we compared the RNA-binding activities of the mutant polypeptides to WT MBP-NP by using a UV cross-linking assay. The protein preparations (containing equal amounts of full-length MBP-NP) were incubated with a radiolabelled RNA at either 30 or 37°C, UV irradiated, incubated with RNase to degrade unbound RNA, and analyzed by SDS-PAGE and autoradiography to detect covalently bound RNA. As expected, the WT polypeptide became radiolabelled at 30 and 37°C, indicating that it bound the RNA at either temperature (Fig. 6B, lanes 1 and 7). No radiolabelled product was seen in reactions containing MBP (lanes 2 and 8), nor was any radiolabel acquired by MBP-NP in the absence of UV irradiation (data not shown). If synthesized at 30°C, the full-length S314 and A332 polypeptides bound RNA similarly to WT MBP-NP when assayed at either 30 or 37°C, while no activity was seen from the smaller MBP-NP “stub” (Fig. 6b, lanes 3, 5, 9, and 11). However, when synthesized at 37°C, the polypeptides displayed greatly reduced binding activity at either assay temperature (lanes 4, 6, 10, and 12). Thus, the S314 and A332 mutations render the RNA-binding activity of NP temperature sensitive. However, if synthesized at the PT, the polypeptides are functional at the NPT. To further examine this aspect, we compared the thermal stability of the RNA-binding activity of the mutant and of the WT MBP-NP polypeptides. Samples containing 1 nM radiolabelled RNA and 80 nM WT or A332 MBP-NP fusion protein (synthesized at 30°C) were incubated at increasing temperatures, and aliquots were examined for bound RNA by using a nitrocellulose filter binding assay. WT MBP-NP showed slightly higher binding activity at 37 than at 30°C, but thereafter the binding decreased, reaching half the original value at ca. 60°C, and it was largely lost by 80°C (Fig. 6c). The A332 mutant produced a binding curve essentially indistinguishable from that of the WT protein, indicating no significant change in the thermal stability of the mutant polypeptide. Similarly, no decrease in the relative stability was seen with the S314 protein (data not shown). Thus, the S314 and A332 polypeptides fail to bind RNA with high affinity if synthesized at the NPT, but they possess thermostable binding activity equal to that of the WT polypeptide if they are made at the permissive temp.
FIG. 6.
RNA-binding activity of WT and mutant MBP-NP polypeptides. (A) Effect of growth and assay temperature on RNA-binding activity of WT MBP-NP. Proteins were grown (G) at the indicated temperatures and titrated for their ability to bind and retain a radiolabelled RNA on nitrocellulose filters at the indicated assay (A) temperature. (B) UV cross-linking assay. Equal amounts of MBP, WT, and mutant MBP-NP polypeptides (prepared at 30 or 37°C as labelled) were incubated with radiolabelled RNA at the indicated temperatures and subjected sequentially to UV irradiation, RNase digestion, SDS-PAGE, and autoradiography. The positions of MBP-NP, MBP, and the MBP-NP “stub” (as determined by staining of the gel with Coomassie brilliant blue [data not shown]) are indicated by arrows. (C) Thermal denaturation of RNA-binding activity. WT or mutant A332-T MBP-NPs (the latter synthesized at 30°C) were incubated with radiolabelled RNA at increasing temperatures, and aliquots were filtered through a nitrocellulose membrane to separate bound and free RNA. The bound values are plotted as a fraction of the amount retained at 37°C.
DISCUSSION
The NP of influenza virus is required for the replicative synthesis of viral RNAs. The ts mutations in WSN ts56 (S314-N) and FPV tsG81 (A332-T) specifically disrupt replicative transcription. If cells are infected with either virus at the permissive temperature for 3 to 5 h and then shifted to the NPT, mRNA transcription continues but vRNA synthesis ceases (22, 40, 41). The synthesis of cRNA also ceases in ts56 (41), but it is not known whether tsG81 behaves similarly. In addition, the ts56 NP is unable to support c- or vRNA synthesis in vitro at the NPT (41).
NP is a multifunctional protein and to identify activities essential for replicative transcription, we have analyzed the biochemical properties of engineered NP mutants which contain the amino acid substitutions found in the ts56 and ts81 NPs. Here we show that the ts mutations S314-N and A332-T render the RNA-binding activity of A/PR/8/34 NP temperature sensitive (Fig. 6). The fact that the mutants showed apparently WT RNA-binding activity and thermostability when synthesized at 30°C but were defective when synthesized at 37°C suggests that the mutations affect the folding of the polypeptide. However, the effect on RNA binding is specific, since the mutations do not substantially alter the cellular accumulation or distribution of the polypeptides (Fig. 1 and 2), suggesting that the polypeptides interact with importin α, F-actin, and the nuclear export machinery similarly to WT NP. Neither do the mutations prevent the NP molecules from self-associating or interacting with the virus polymerase normally (Fig. 4 and 5). Further arguing against gross effects on polypeptide structure, no significant differences were seen between the sensitivities of the mutant and WT MBP-NP polypeptides to partial proteolytic digestion with trypsin or chymotrypsin (data not shown).
Several hypotheses have been put forward to explain the specific role of NP in the switch between mRNA and replicative transcription. A recent suggestion concerns the ability of NP to interact with the polymerase: in this model NP alters the transcriptional function of the polymerase through direct protein-protein contacts (4, 33). In contrast, the encapsidation hypothesis proposes that NP does not have a regulatory function as such and that other elements determine the choice between cap-primed and unprimed transcription initiation but that NP is required to cotranscriptionally coat the nascent c- and vRNA segments (41). Alternatively, the template modification hypothesis holds that the interaction of free NP (that is not already present in the RNP structure) with the promoter element of the template RNA alters the modes of transcription initiation and termination (13, 18, 20). This is plausible, since the terminal sequences of the vRNA template are partially base paired to form a panhandle structure (2, 18), and recognition of this structure by the polymerase is intimately connected with the mechanisms of mRNA transcription initiation (7, 15, 47) and polyadenylation (38, 39). Here we show that two ts mutations which disrupt replicative transcription render the RNA-binding activity of NP ts (Fig. 6), suggesting that NP-RNA contacts are essential for the process. The encapsidation and template modification hypotheses predict NP-RNA interactions during RNA replication. Thus, our data support both hypotheses but do not differentiate between the two. The hypotheses are not necessarily exclusive, but further experimental tests are necessary to determine whether one or both mechanisms occur. In addition, the regulation of vRNA transcription may differ from that of cRNA, and indeed genetic evidence indicates that the roles of NP in c- and vRNA synthesis are mutationally separable (33, 46). The role of NP in vRNA synthesis is perhaps simpler, as cRNA templates do not support cap-primed transcription or contain a polyadenylation signal (7). Certainly, the premature termination of in vitro vRNA transcription in the absence of soluble NP is consistent with the necessity of NP for cotranscriptional encapsidation of the nascent segment (41). The hypothesis that NP-P interactions regulate transcription is not lent support by our results. In addition, the competence of the ts NPs studied here to form protein contacts with the influenza polymerase indicates that NP-P interactions alone are not sufficient to support replicative transcription. This is also supported by the observation that the addition of excess NP to virion RNPs does not induce the synthesis of cRNA (44). Although neither set of experiments exclude the possibility that direct NP-P contacts are involved in regulating the mode of transcription, it is also conceivable that protein-protein contacts between NP and the polymerase play a structural role during the assembly and transcription of RNPs.
Temperature-shift experiments carried out with ts56 and tsG81 showed that mRNA transcription persisted at the NPT (22, 40, 41). However, RNA-binding activity would also be expected to be essential for mRNA synthesis, because NP is a structural component of the RNP required for processive transcription (17). The temperature stability of the RNA-binding activity of mutant NPs made at the PT provides a plausible explanation for the persistence of mRNA transcription, as RNP cores made at 31°C would be predicted to retain full functionality. However, we note that, in the experiments reported by Shapiro and Krug (41), extracts from WSN ts56-infected cells made at 31°C did not support replicative transcription at 39.5°C, a finding which would not be predicted from the temperature stability of our S314-N mutant. This discrepancy might be explained by our using a lower NPT or, alternatively, by the non-ts-associated sequence changes between the PR8 and WSN NPs.
The S314-N and A332-T mutations lie outside of the region of NP previously identified as the minimal RNA-binding domain (approximately residues 1 to 180 [1, 21]). However, we have recently determined that sequences throughout the length of NP are in direct contact with RNA and have identified several other point mutations outside of the N-terminal third of the polypeptide that similarly disrupt the interaction of the protein with RNA (11). Thus, our current model is that high-affinity RNA binding by NP requires the concerted function of multiple regions of the polypeptide.
Two further points arise from the tsG81/A332-T mutation. A partial revertant of FPV tsG81A has been isolated that retains the original ts NP gene, thus implying the extragenic suppression of the ts defect (32). The location of the suppressor mutation was difficult to identify, but after a complex genetic study Mandler and colleagues concluded that the mutation most likely lay in the PB2 gene (32). We find no evidence of any alteration in the ability of NP A332-T to interact with WT PB2, either in the context of the polymerase complex (Fig. 6) or alone (data not shown), but we have not studied NP binding to the mutant PB2 isolated by Mandler et al. (32). Finally, a recent study reported the effects of deliberately introduced NP point mutations on m-, c-, and vRNA synthesis (33) in which the authors mutated residue M331 (among others), adjacent to the ts lesion in NP tsG81 (A332). Alteration to threonine had little effect on NP function, but when it was changed to lysine, the polypeptide became ts for gene expression and specifically defective for cRNA synthesis (33). No further characterization of the polypeptide was reported, but our results would predict that NP M331-K has a ts RNA-binding activity similar to that of NP A332-T.
ACKNOWLEDGMENTS
We thank Ian Brierley and John McCauley for critical appraisal of the manuscript and Laurence Tiley for helpful discussion.
This work was supported by grants from the Royal Society and Wellcome Trust (048911) to P.D. P.D. is a Royal Society University Research Fellow.
REFERENCES
- 1.Albo C, Valencia A, Portela A. Identification of an RNA binding region within the N-terminal third of the influenza A virus nucleoprotein. J Virol. 1995;69:3799–3806. doi: 10.1128/jvi.69.6.3799-3806.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Baudin F, Bach C, Cusak S, Ruigrok R W H. Structure of influenza RNP I. Influenza virus nucleoprotein melts secondary structure in panhandle RNA and exposes the bases to solvent. EMBO J. 1994;13:3158–3165. doi: 10.1002/j.1460-2075.1994.tb06614.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Beaton A R, Krug R M. Transcription antitermination during influenza viral template RNA synthesis requires the nucleocapsid proteins and the absence of a 5′ capped end. Proc Natl Acad Sci USA. 1986;83:6282–6286. doi: 10.1073/pnas.83.17.6282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Biswas S K, Boutz P L, Nayak D P. Influenza virus nucleoprotein interacts with influenza virus polymerase proteins. J Virol. 1998;72:5493–5501. doi: 10.1128/jvi.72.7.5493-5501.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Blok V, Cianci C, Tibbles K W, Inglis S C, Krystal M, Digard P. Inhibition of the influenza virus RNA-dependent RNA polymerase by antisera directed against the carboxy-terminal region of the PB2 subunit. J Gen Virol. 1996;77:1025–1033. doi: 10.1099/0022-1317-77-5-1025. [DOI] [PubMed] [Google Scholar]
- 6.Braam J, Ulmanen I, Krug R M. Molecular model of a eukaryotic transcription complex: functions and movements of influenza P proteins during capped RNA-primed transcription. Cell. 1983;34:609–618. doi: 10.1016/0092-8674(83)90393-8. [DOI] [PubMed] [Google Scholar]
- 7.Cianci C, Tiley L, Krystal M. Differential activation of the influenza virus polymerase by template RNA binding. J Virol. 1995;69:3995–3999. doi: 10.1128/jvi.69.7.3995-3999.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Craig D, Howell M T, Gibbs C L, Hunt T, Jackson R J. Plasmid cDNA-directed protein synthesis in a coupled in vitro transcription-translation system. Nucleic Acids Res. 1992;20:4987–4995. doi: 10.1093/nar/20.19.4987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Digard P, Blok V C, Inglis S C. Complex formation between influenza virus polymerase proteins expressed in Xenopus oocytes. Virology. 1989;171:162–169. doi: 10.1016/0042-6822(89)90523-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Digard P, Elton D, Bishop K, Medcalf E, Weeds A, Pope B. Modulation of nuclear localization of influenza virus nucleoprotein through interaction with actin filaments. J Virol. 1999;73:2222–2231. doi: 10.1128/jvi.73.3.2222-2231.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Elton D, Medcalf E, Bishop K, Harrison D, Digard P. Identification of amino acid residues of influenza virus nucleoprotein essential for RNA binding. J Virol. 1999;73:7357–7367. doi: 10.1128/jvi.73.9.7357-7367.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Elton, D., E. Medcalf, K. Bishop, and P. Digard. Analysis of self-association by the influenza virus nucleoprotein. Virology, in press.
- 13.Fodor E, Pritlove D C, Brownlee G G. The influenza virus panhandle is involved in the initiation of transcription. J Virol. 1994;68:4092–4096. doi: 10.1128/jvi.68.6.4092-4096.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Fuerst T R, Earl P L, Moss B. Use of a hybrid vaccinia virus-T7 RNA polymerase system for expression of target genes. Mol Cell Biol. 1987;7:2538–2544. doi: 10.1128/mcb.7.7.2538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hagen M, Chung T D Y, Butcher J A, Krystal M. Recombinant influenza virus polymerase: requirement of both 5′ and 3′ viral ends for endonuclease activity. J Virol. 1994;68:1509–1515. doi: 10.1128/jvi.68.3.1509-1515.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Hay A J, Lomnizi B, Bellamy A R, Skehel J J. Transcription of the influenza virus genome. Virology. 1977;83:337–355. doi: 10.1016/0042-6822(77)90179-9. [DOI] [PubMed] [Google Scholar]
- 17.Honda A, Ueda K, Nagata K, Ishihama A. RNA polymerase of influenza virus: role of NP on RNA chain elongation. J Biochem. 1988;104:1021–1026. doi: 10.1093/oxfordjournals.jbchem.a122569. [DOI] [PubMed] [Google Scholar]
- 18.Hsu M-T, Parvin J D, Gupta S, Krystal M, Palese P. Genomic RNAs of influenza viruses are held in a circular conformation in virions and in infected cells by a terminal panhandle. Proc Natl Acad Sci USA. 1987;84:8140–8144. doi: 10.1073/pnas.84.22.8140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Huang T-S, Palese P, Krystal M. Determination of influenza virus proteins required for genome replication. J Virol. 1990;64:5669–5673. doi: 10.1128/jvi.64.11.5669-5673.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Klump K, Ruigrok R W H, Baudin F. Roles of the influenza virus polymerase and nucleoprotein in forming a functional RNP structure. EMBO J. 1997;16:1248–1257. doi: 10.1093/emboj/16.6.1248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Kobayashi M, Toyoda T, Adyshev D M, Azuma Y, Ishihama A. Molecular dissection of influenza virus nucleoprotein: deletion mapping of the RNA binding domain. J Virol. 1994;68:8433–8436. doi: 10.1128/jvi.68.12.8433-8436.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Krug R M, Ueda M, Palese P. Temperature-sensitive mutants of influenza WSN virus defective in virus-specific RNA synthesis. J Virol. 1975;16:790–796. doi: 10.1128/jvi.16.4.790-796.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kunkel T A. Rapid and efficient site-specific mutagenesis without phenotypic selection. Proc Natl Acad Sci USA. 1985;82:488–492. doi: 10.1073/pnas.82.2.488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Lamb R A, Krug R M. Orthomyxoviridae: the viruses and their replication. In: Fields B N, Knipe D M, Howley P M, editors. Fields virology. Philadelphia, Pa: Lippincott-Raven; 1996. pp. 1353–1396. [Google Scholar]
- 25.Li M-L, Ramirez B C, Krug R M. RNA-dependent activation of primer RNA production by influenza virus polymerase: different regions of the same protein subunit constitute the two required RNA-binding sites. EMBO J. 1998;17:5844–5852. doi: 10.1093/emboj/17.19.5844. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Li R, Palese P, Krystal M. Complementation and analysis of an NP mutant of influenza virus. Virus Res. 1989;12:97–112. doi: 10.1016/0168-1702(89)90057-9. [DOI] [PubMed] [Google Scholar]
- 27.Lin B-C, Lai C-J. The influenza virus nucleoprotein synthesized from cloned DNA in a simian virus 40 vector is detected in the nucleus. J Virol. 1983;45:434–438. doi: 10.1128/jvi.45.1.434-438.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Luo G, Luytes W, Enami M, Palese P. The polyadenylation signal of influenza virus RNA involves a stretch of uridines followed by the RNA duplex of the panhandle structure. J Virol. 1991;65:2861–2867. doi: 10.1128/jvi.65.6.2861-2867.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Mahy B W J, Barret T, Nichol S T, Penn C R, Wolstenholme A J. Analysis of the functions of influenza virus genome RNA segments by the use of temperature-sensitive mutants of fowl plague virus. In: Bishop D H L, Compans R W, editors. The replication of negative strand viruses. Amsterdam, The Netherlands: Elsevier/North Holland; 1981. pp. 379–387. [Google Scholar]
- 30.Mahy B W J. Mutants of influenza virus. In: Palese P, Kingsbury D W, editors. Genetics of influenza virus. New York, N.Y: Springer-Verlag; 1983. pp. 192–254. [Google Scholar]
- 31.Mandler J, Scholtissek C. Localization of the temperature-sensitive defect in the nucleoprotein of an influenza A/FPV/Rostock/34 virus. Virus Res. 1989;12:113. doi: 10.1016/0168-1702(89)90058-0. [DOI] [PubMed] [Google Scholar]
- 32.Mandler J, Muler K, Scholtissek C. Mutants and revertants of an avian influenza A virus with temperature-sensitive defects in the nucleoprotein and PB2. Virology. 1991;181:512–519. doi: 10.1016/0042-6822(91)90883-d. [DOI] [PubMed] [Google Scholar]
- 33.Mena I, Jambrina E, Albo C, Perales B, Ortin J, Arrese M, Vallejo D, Portela A. Mutational analysis of influenza A virus nucleoprotein: identification of mutations that affect RNA replication. J Virol. 1999;73:1186–1194. doi: 10.1128/jvi.73.2.1186-1194.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Neumann G, Castrucci M R, Kawaoka Y. Nuclear import and export of influenza virus nucleoprotein. J Virol. 1997;71:9690–9700. doi: 10.1128/jvi.71.12.9690-9700.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.O’Neill R E, Palese P. NPI-1, the human homologue of SRP-1, interacts with influenza virus nucleoprotein. Virology. 1995;206:116–125. doi: 10.1016/s0042-6822(95)80026-3. [DOI] [PubMed] [Google Scholar]
- 36.Plotch S J, Bouloy M, Ulmanen I, Krug R M. A unique cap(m7GpppXm)-dependent influenza virion endonuclease cleaves capped RNAs to generate the primers that initiate viral RNA transcription. Cell. 1981;23:847–858. doi: 10.1016/0092-8674(81)90449-9. [DOI] [PubMed] [Google Scholar]
- 37.Pons M W, Schulze I T, Hirst G K, Hauser R. Isolation and characterization of the ribonucleoprotein of influenza virus. Virology. 1969;39:25–259. doi: 10.1016/0042-6822(69)90045-2. [DOI] [PubMed] [Google Scholar]
- 38.Poon L L M, Pritlove D C, Sharps J, Brownlee G G. The RNA polymerase of influenza virus, bound to the 5′ end of virion RNA, acts in cis to polyadenylate mRNA. J Virol. 1998;72:8214–8219. doi: 10.1128/jvi.72.10.8214-8219.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Pritlove D C, Poon L L M, Fodor E, Sharps J, Brownlee G G. Polyadenylation of influenza virus mRNA transcribed in vitro from model virion RNA templates: requirement for 5′ conserved sequences. J Virol. 1998;72:1280–1286. doi: 10.1128/jvi.72.2.1280-1286.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Scholtissek C. The genome of influenza virus. Curr Top Microbiol Immunol. 1978;80:139–169. doi: 10.1007/978-3-642-66956-9_5. [DOI] [PubMed] [Google Scholar]
- 41.Shapiro G I, Krug R M. Influenza virus RNA replication in vitro: synthesis of viral template RNAs and virion RNAs in the absence of an added primer. J Virol. 1988;62:2285–2290. doi: 10.1128/jvi.62.7.2285-2290.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Shi L, Summers D F, Peng Q, Galarza J M. Influenza A virus RNA polymerase subunit PB2 is the endonuclease which cleaves host cell mRNA and functions only as the trimeric enzyme. Virology. 1995;208:38–47. doi: 10.1006/viro.1995.1127. [DOI] [PubMed] [Google Scholar]
- 43.Shimizu K, Handa H, Nakada S, Nagata K. Regulation of influenza virus RNA polymerase activity by cellular and viral factors. Nucleic Acids Res. 1994;22:5047–5053. doi: 10.1093/nar/22.23.5047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Skorko R, Summers D F, Galarza J M. Influenza A virus in vitro transcription: roles of NS1 and NP proteins in regulating RNA synthesis. Virology. 1991;180:668–677. doi: 10.1016/0042-6822(91)90080-u. [DOI] [PubMed] [Google Scholar]
- 45.Smith G L, Levin J Z, Palese P, Moss B. Synthesis and cellular location of the ten influenza polypeptides individually expressed by recombinant vaccinia viruses. Virology. 1987;160:336–345. doi: 10.1016/0042-6822(87)90004-3. [DOI] [PubMed] [Google Scholar]
- 46.Thierry F, Danos O. Use of specific single-stranded DNA probes cloned in M13 to study the RNA synthesis of four temperature-sensitive mutants of HK/68 influenza virus. Nucleic Acids Res. 1982;10:2925–2937. doi: 10.1093/nar/10.9.2925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Tiley L, Hagen M, Matthews J T, Krystal M. Sequence-specific binding of the influenza virus RNA polymerase to sequences located at the 5′ ends of the viral RNAs. J Virol. 1994;68:5108–5116. doi: 10.1128/jvi.68.8.5108-5116.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Wang P, Palese P, O’Neil R E. The NPI-1/NPI-3 (karyopherin α) binding site on the influenza A virus nucleoprotein NP is a nonconventional nuclear localization signal. J Virol. 1997;71:1850–1856. doi: 10.1128/jvi.71.3.1850-1856.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Weber F, Kochs G, Gruber S, Haller O. A classical bipartite nuclear localization signal on Thogoto and influenza A virus nucleoproteins. Virology. 1998;250:9–18. doi: 10.1006/viro.1998.9329. [DOI] [PubMed] [Google Scholar]
- 50.Winter G, Fields S. The structure of the gene encoding the nucleoprotein of human influenza virus A/PR/8/34. Virology. 1981;114:423–428. doi: 10.1016/0042-6822(81)90223-3. [DOI] [PubMed] [Google Scholar]