Abstract
The field of computational chemistry has seen a significant increase in the integration of machine learning concepts and algorithms. In this Perspective, we surveyed 179 open-source software projects, with corresponding peer-reviewed papers published within the last 5 years, to better understand the topics within the field being investigated by machine learning approaches. For each project, we provide a short description, the link to the code, the accompanying license type, and whether the training data and resulting models are made publicly available. Based on those deposited in GitHub repositories, the most popular employed Python libraries are identified. We hope that this survey will serve as a resource to learn about machine learning or specific architectures thereof by identifying accessible codes with accompanying papers on a topic basis. To this end, we also include computational chemistry open-source software for generating training data and fundamental Python libraries for machine learning. Based on our observations and considering the three pillars of collaborative machine learning work, open data, open source (code), and open models, we provide some suggestions to the community.
1. Introduction
Creating models and performing simulations are cornerstones of science. Prior to computers, models were created on paper (e.g., mathematics, diagrams) or physically constructed from material. Modern modeling is done on computers (in silico), allowing one to easily adjust parameters and quickly observe the resulting effects. Today, a plethora of simulation and modeling codes exist, which can be either open or closed to the public. While closed-source software is created by companies for economic reasons, open-source software (OSS) has played an important role in scientific discovery. The OSS philosophy promotes the distribution of code (i.e., tools) and subsequently the natural and computer science knowledge that is embedded within the code. OSS encourages researchers to read the code critically, to understand its mathematical formulations, parameters, and assumptions and the workflow’s logic, and to modify it as desired. Free and open-source software (FOSS), a subcategory of OSS, also demands licensing models that provide a legal framework for the free distribution, use, and development of the code, albeit that commercial usage might still be restricted.
The field of machine learning (ML) has clearly grown, as can be seen by the increased number of research articles published that include it and through the interest shown by the general public. Paraphrasing Sonnenburg et al., OSS benefits the ML field by enabling better reproducibility of scientific results and quicker detection of errors as well as faster, innovative combinations of scientific ideas and their subsequent applications to diverse disciplines.1 To this list are added the benefits of being able to more easily validate the assumptions and approximations made during model building. The very goal and act of making ML algorithms and their trained models open-source has the following three benefits to the field: (1) standardizing interfaces (e.g., adopting specific frameworks), (2) enabling experimentation (e.g., guiding project choices and obtaining alternative perspectives), and (3) community creation (e.g., developer–user interactions and improved educational material).2
The field of computational chemistry has benefited significantly from the advancement of OSS and ML. The development of and access to software tools has enabled nonexperts to apply ML in their chemical and biological research and also enabled ML experts to solve problems in the chemical and biological domains. Informing others about available tools is a 2016 review article written by Pirhadi et al. that covers open-source computational chemistry software that includes some ML approaches.3 The authors also maintain an accompanying GitHub repository software list that includes annotations on open-source molecular modeling software (https://opensourcemolecularmodeling.github.io). The interest in ML by computational chemists is natural given the physics, statistical, algorithmic, and data-based ideas that are already present within the field. At its core, ML is built upon statistics and complex black-box modeling techniques that are implemented, distributed, and made accessible through thoughtful programming. The Python programming language is particularly suited for natural scientists since it is easy to generate readable procedural and object-oriented code that allows for quick and creative exploration of ideas. Python is also the most popular language in the ML community.
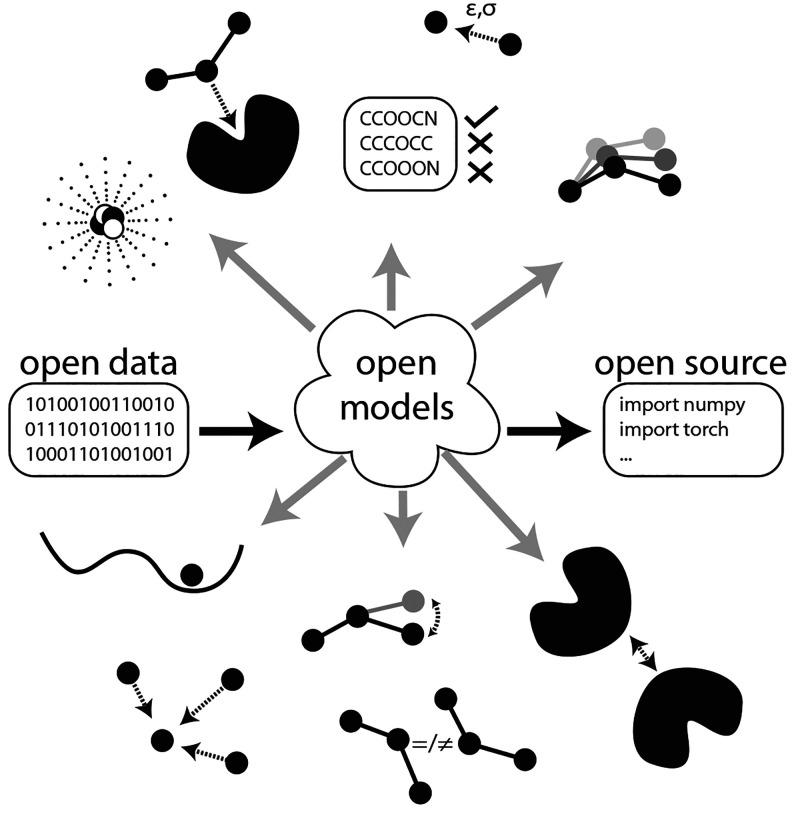
This Perspective’s goal is to provide an introduction concerning open-source Python-based (with a few exceptions) ML tools that are available for computational chemistry researchers who want to start exploring the technology, either in direct usage (i.e., application) of the learned models or in reading the code for purposes of self-education or experimentation (i.e., modification). To this end, four software groups will be covered, starting with general software development tools. Following this, standard scientific Python libraries for data manipulation and visualization are discussed. As the third group, libraries that form the foundation of ML software will be presented. Finally, selected computational-chemistry-specific tools released in the last 5 years will be discussed. While developing and applying ML code is often focused upon, the importance of generating and understanding the data on which ML is trained should not be underestimated. Consequently, we will also include notable computational chemistry software for generating data that are distributed freely through licenses and whose source code is then made available. Our choice for including the application areas (e.g., predicting energies, chemical reactions, partial atomic charges, etc.) is motivated by the ML software that is highly cited (e.g., AlphaFold), recently released research (e.g., via ASAP articles), cited literature within those publications, subsequent literature found from citation searches, and our research interests. A graphical illustration concerning the concept of the openness of machine learning computational chemistry tools is shown in Figure 1. The three building blocks for openness in ML research are considered to be (1) the availability of datasets (open data), (2) the availability of code (open source), and (3) the availability of trained ML models (open models).
Figure 1.
An illustration of how open data, open models, and open source codes are used in the different subdomains, as described herein, of computational chemistry.
In addition to providing a listing of computational-chemistry-focused ML repositories, we also note whether the training data and the resulting model and/or its parameters (i.e., weights) are published. Although this community might often be more interested in the resulting observable predictions, for reproducibility it is very important to ensure that a published model reported in a paper produces the same output as that provided in a repository. In the ML community, it has become standard practice to release training data and model parameters alongside the publication, mostly arising from the model’s size and the amount of training required (i.e., a resource and time issue). In this context, the model parameters refer to the weights that determine its output and should not be confused with the model’s training hyperparameters (e.g., the learning rate). For smaller models that are easily trained on a workstation, the publication of the hyperparameters is sufficient for reproducibility. Nevertheless, publishing the model’s parameters can increase the publication’s robustness against unnoticed changes in the libraries used by the model, outlier random seeds that increase the model’s performance, or mistakes in handling code publication, all of which might lead to differences in the model’s training during experimentation and publication.
As a final comment, we ask the reader and the researchers whose work we included or did not include to be forgiving in their critique of this Perspective. Given the large scope that ML and computational chemistry cover and the speed at which the combined domains are growing, our interpretation and categorization may sometimes be incomplete or imprecisely described. We have limited our focus to OSS that has been released alongside a peer-reviewed paper (i.e., work originating from a preprint manuscript was not included), thereby helping to ensure a higher academic standard of the work. Due to its limited in-depth coverage, the reader is referred to reviews written within the past 3 years that cover the use of ML in specific computational chemistry subdomains for additional information.4−52 Given the numerous review articles and vastness of the topic, we believe that the information herein can serve as an introductory point into the application of ML in computational chemistry.
2. Software and Libraries
2.1. Software Development Tools
The first set of tools to discuss are those forming the software environment that helps one install and organize Python libraries as well as to encode, train, test, and implement an ML workflow. The Conda53 software enables the installation and management of Python and its libraries. Apart from easing library installations, Anaconda54 and Miniconda55 enable the creation of independent environments that isolate projects from one another and from the operating system’s default installed libraries. At an advanced level, this can enable users to test different versions of their code’s imported libraries.
The use of an online code repository, hosted by a service like GitHub56 or GitLab,57 is strongly recommended. Git is a code version control tool that has overtaken older tools such as Apache Subversion (SVN). A git repository (a) serves as a backup to one’s work through a version control system, allowing users to have local and remote instances of the deposited code, (b) enables multiple people to work on the same code (i.e., project) and merge their work back to the centralized source, and (c) enables the creation of branches that allow safe experimentation and implementation that do not immediately affect the main branch. To differentiate the two aforementioned code repository systems, GitHub serves as a public platform for repositories, while GitLab is an entirely independent and self-contained ecosystem that can also be installed on self-managed local servers. A significant number of codes covered in this Perspective are hosted on GitHub.
While computer scientists often use integrated development environment (IDE) software to develop their code (e.g., PyCharm,58 Spyder,59 or Visual Studio Code60), a natural scientist might favor coding in Jupyter notebooks61,62 locally or online using a JupyterHub (private or public server) or Google’s Colaboratory.63 All of these latter tools allow one to mix code (i.e., code cells) with regular textual writing (i.e., markdown cells), thus serving very much like a traditional science lab notebook. Consequently, researchers can transcribe their thoughts while encoding their workflow within a single notebook, which can easily be shared with others (e.g., validation during a manuscript’s peer review; enable reproducibility and transparency). One advantage of the Colaboratory is its free access to reasonably powered GPU computing for executing code and training ML models. A disadvantage of such a third-party hosting solution can be data security and privacy when running code and uploading data. Alternatively, JupyterHub can be used, which enables self-managed collaborative work on multiple shared Jupyter notebooks using centrally organized computational resources and easy sharing of demonstrations with the community. A possible drawback for some researchers is that JupyterHub might require a server and IT expert to install and maintain.
2.2. Standard and Scientific Python Libraries
The Python programming language offers a basic set of common tools in its standard library (https://docs.python.org/3/library), which includes reading and writing using standard data formats, performing numerical and mathematical operations, interacting with the file and operating system, handling data, testing code using unit tests, handling exceptions, and many more things. In addition to the standard library, many libraries have been developed that support more advanced operations and algorithms. Several libraries are considered to be fundamental for programmers who are interested in natural science and machine learning. In the following, we introduce the libraries that are widely used and adapted by researchers within our field. They are maintained by active communities of OSS developers and can be relied upon due to their widespread use.
For mathematical modeling, data analysis, and manipulation, the following libraries are often used: NumPy, Pandas, SciPy, Matplotlib, Seaborn, and SymPy (Table 1). NumPy64 was designed to perform fast numerical calculations on arrays and matrices, which it achieves by interfacing with code that was written in the more low-level C and Fortran programming languages. NumPy calculations are vectorized, which makes its usage inherently parallel. Consequently, many other libraries, such as those mentioned above, use NumPy for their calculations. The Pandas library enables data to be easily imported and exported (e.g., from and to CSV-formatted files) and manipulated (e.g., filtered, sorted).65,66 At a superficial level, Pandas can be considered as Python’s version of a spreadsheet. The focus of the SciPy library is scientific computing, and it includes algorithms for extrapolation, fast Fourier transforms, interpolation, linear algebra, numerical integration, optimization, polynomial fitting, statistics, and signal processing.67 Matplotlib68 and Seaborn69 are libraries for visualizing data through plots. Seaborn is built on top of Matplotlib and simplifies the coding for high-quality, complex visualizations. SymPy is a symbolic mathematics library that includes the ability to compute derivatives, integrals, and the limits of equations, with submodules that focus on matrices, polynomials, series, and quantum mechanics.70 While not a complete list, these libraries are very helpful in ML projects for reading, manipulating, and visualizing data.
Table 1. OSS Available for Scientific Computation.
| software | link | license |
|---|---|---|
| NumPy64 | https://github.com/numpy/numpy | BSD-3-Clause |
| Pandas65,66 | https://github.com/pandas-dev/pandas | BSD-3-Clause |
| SciPy67 | https://github.com/scipy/scipy | BSD-3-Clause |
| Matplotlib68 | https://github.com/matplotlib/matplotlib | BSD-compatible |
| Seaborn69 | https://github.com/mwaskom/seaborn | BSD-3-Clause |
| SymPy70 | https://github.com/sympy/sympy | BSD |
2.3. Machine Learning
ML has become a vast field that can be hard to traverse, even for weathered scientists from the field itself. Development is going at a lightning pace, and sometimes the field even outpaces the scientific review process and suffers from catastrophic forgetting and quality reduction. The lessons we can learn from the fast development of generative, large language, and text-to-image ML models is that it is not only the amount of scientific activity that leads to fast development but rather the active exchange of code, data, and, foremost, trained models. Although large companies with vast computing power are nowadays often the first to increase a model’s size, they do not always share the model’s parameters, partially closing models off for scientific evaluation. These so-called foundation models, which can be used to derive more specifically trained models a posteriori, often end up as closed models whose use requires a fee. Interestingly, a portion of the ML community seems to immediately move to open models as soon as they reach the performance level or outperform the foundation models. These open models are more accessible for scientific evaluation and use and can be more computationally efficient due to the limited resources available outside of large companies.
Increasing the computational efficiency when training ML models enables more groups to do research. It is therefore important to cover classical ML (i.e., “shallow learning”) algorithms that can be more appropriate when dealing with datasets that have certain characteristics (e.g., small number of data points). These models are often much smaller, require less training data, and are trained quicker. Inference, using models for prediction, also tends to require much less computational effort. Some problems might require more rigorous statistics or explainability, for which statistical learning methods such as Bayesian learning are more appropriate. If a lot of training is available, deep learning methods can be used to handle data with an unknown structure. Finally, graph neural networks (GNNs) are of particular use for tasks like molecular modeling due to their inherent graphlike structure that mimics a molecule’s structural formula representation.19,48 Addressing the computational costs of GNNs is a GNN coarsening framework based on functional groups nicknamed FunQG (https://github.com/hhaji/funqg; MIT).
A large amount of ML code has appeared over the last two decades. Herein we focus on the larger Python-focused libraries that are widely used, open-source, and actively maintained.
2.3.1. Classical Frameworks
At a simplistic level, shallow learning can be thought of as having only one or two learning layers in the ML model. Oftentimes, the input data are preprocessed by humans to extract certain known derived data features that help the model learn more easily, reducing the complexity of the models. Table 2 lists the most widely used frameworks.
Table 2. OSS for Classical Machine Learning.
| software | link | license |
|---|---|---|
| scikit-learn71 | https://scikit-learn.org | BSD-3-Clause |
| scikit-cuda72 | https://github.com/lebedov/scikit-cuda | custom |
| ffnet73 | https://ffnet.sourceforge.net | LGPL-3.0 |
| XGBoost74 | https://github.com/dmlc/xgboost | Apache-2.0 |
| LightGBM75 | https://github.com/microsoft/LightGBM | MIT |
Scikit-learn contains a large number of basic shallow learning algorithms for supervised learning (i.e., learning targets from source–target data) and unsupervised learning (i.e., finding structure in unlabeled data).71 The latter include the two major categories of the classical unsupervised ML paradigm: clustering (i.e., grouping data based on similarities) and manifold learning (i.e., resolving a low-dimensional substructure in high-dimensional data). The library provides assistive tools for model selection, inspection, and evaluation as well as data handling and visualization. Scikit-CUDA enables users to easily access GPU-accelerated linear algebra operations.72 The small ffnet package allows the training of shallow feed-forward neural networks (NNs) in Python.73 XGBoost is an optimized, distributed ML library.74 LightGBM offers tree-based learning algorithms.75
2.3.2. Statistical Frameworks
As an alternative to classical ML approaches, statistical learning provides more rigorous models that are mostly based on Gaussian process modeling and regression, collectively called Kriging. This category of ML uses the assumption that a function that we would like to predict, based on input data, is sampled from a normal distribution of functions (just like an input data point is sampled from a normal distribution of points). The stochastic process that produces this distribution is called a Gaussian process. The modeling technique interpolates between data points and follows an underlying assumption that the function is smooth. Other underlying assumptions can be added as well, such as periodicity. We list the most widely used frameworks in Table 3. Representative libraries are GPy,76 GPflow,77 and GPytorch,78 which provide a large array of models and training paradigms. The latter two support fast GPU computations.
Table 3. OSS Available for Statistical ML.
| software | link | license |
|---|---|---|
| GPy76 | http://github.com/SheffieldML/GPy | BSD-3-Clause |
| GPflow77 | https://github.com/GPflow/GPflow | Apache-2.0 |
| GPytorch78 | https://github.com/cornellius-gp/gpytorch | MIT |
2.3.3. Deep Frameworks
Increasing the depth of the model (i.e., the number of layers) opens up the realm of deep learning. These models learn what features are important by themselves based solely on raw data. This is conceptually different from classical ML that uses manually defined features. Deep learning can discover structure and features in much larger datasets, and its concept has had an incredible impact on the usability of ML. The most widely used deep learning frameworks are listed in Table 4. PyTorch79 is a heavily used tensor (i.e., conceptually, a high-dimensional matrix) library for ML, allowing programmers to build vast models and make use of GPUs. Previously, the de facto deep learning framework library was TensorFlow, which is still widely used.80 Together, PyTorch and TensorFlow form the basis of most modern ML codes. Keras81 provides a user-friendly interface to TensorFlow. The PyTorch-based ξ-torch library provides differentiable functions and optimization algorithms for use in deep learning.82 An alternative framework to PyTorch and TensorFlow is Aesara, which grew from Theano,83 but its current use has been limited. Extending beyond Python, Apache MXNet supports deep learning with a wide range of programming languages (e.g., C++, R, Python).84 OpenNMT85 provides an ecosystem for neural machine translation and sequence learning, which can be applied to string-based representation (e.g., SMILES, InChI). It offers model architectures and training procedures for tasks such as string generation and translation. OpenNMT was created for natural language processing and is thus usable with string-based molecular representations.86
Table 4. OSS Available for Deep Learning.
| software | link | license |
|---|---|---|
| PyTorch79 | https://pytorch.org | BSD-style |
| TensorFlow80 | https://www.tensorflow.org | Apache-2.0 |
| Theano/Aesara83 | https://github.com/aesara-devs/aesara | custom |
| Keras81 | https://keras.io | MIT |
| ξ-torch82 | https://github.com/xitorch/xitorch | MIT |
| MXNet84 | https://github.com/apache/incubator-mxnet | Apache-2.0 |
| OpenNMT85 | https://opennmt.net | MIT |
2.3.4. Graph Neural Network Frameworks
A particular type of deep learning model that is specifically useful for computational chemistry is the GNN,19,48 whose most popular frameworks are listed in Table 5. The graph representation conceptually resembles a molecule’s structural formula. Consequently, GNNs are frequently used in ML models that focus on molecule-dependent features (e.g., elemental symbols and bond distances). Exemplifying GNN creations are the PyTorch-based PyG library87 and (currently not peer-reviewed) TensorFlow-based graph_nets library88 for network creation. The Deep Graph Library89 enables the integration of GNNs into the PyTorch, TensorFlow, and Apache MXNet frameworks.
Table 5. OSS Available for GNNs.
| software | link | license |
|---|---|---|
| PyG87 | https://github.com/pyg-team/pytorch_geometric | MIT |
| graph_nets88 | https://github.com/deepmind/graph_nets | Apache-2.0 |
| Deep Graph Library89 | https://www.dgl.ai | Apache-2.0 |
2.4. Automating Machine Learning
ML models must be designed and configured, which is one of the central tasks for ML users. The field of automated machine learning (AutoML) tries to alleviate this task by providing systematic approaches to find the right data features, architecture, and hyperparameters of the model, such as the learning rate, activation function, optimization strategy, and loss function.
2.4.1. Feature Engineering
An important task in classical ML, as opposed to deep learning, is feature engineering, which includes the selection and extraction of high-level features based on the raw data. In deep learning, these features are learned, but oftentimes expert knowledge is used to enable shallow learning, which is much more appropriate when datasets are small. Specific tools and libraries have been developed to assist with selecting, extracting, or calculating the right features from (raw) data. We list the most widely used libraries in Table 6. Featuretools90 is a general library for this task, whereas Feature-engine91 was specifically built for scikit-learn. tsfresh extracts features for time series.92
Table 6. Libraries Used for Automated Feature Extraction.
| software | link | license |
|---|---|---|
| Featuretools90 | https://github.com/alteryx/featuretools | BSD-3-Clause |
| Feature-engine91 | https://github.com/feature-engine/feature_engine | BSD-3-Clause |
| tsfresh92 | https://github.com/blue-yonder/tsfresh | MIT |
2.4.2. Model Selection
Finding a good model architecture (e.g., the number of layers and dimensions) is part of the neural architecture search and the model selection process. The most widely used libraries are provided in Table 7. scikit-learn71 has built-in methods to assist in model selection. Libra is a general library for model selection that supports Keras, TensorFlow, PyTorch, and scikit-learn. PyCaret93 is similar to Libra and supports scikit-learn, XGBoost, LightGBM, Optuna, Hyperopt, and others. Finally, Yellowbrick94 provides visual analysis and diagnostic tools.
Table 7. Libraries Used for Model Selection.
| software | link | license |
|---|---|---|
| scikit-learn71 | https://scikit-learn.org | BSD-3-Clause |
| Yellowbrick94 | https://github.com/DistrictDataLabs/yellowbrick | Apache-2.0 |
| Libra | https://github.com/Palashio/libra | MIT |
| PyCaret93 | https://github.com/pycaret/pycaret | MIT |
2.4.3. Hyperparameter Optimization
Hyperparameters are, in contrast to parameters or weights, not learned by the model but have to be preconfigured. The values of these hyperparameters directly affect the model’s resulting accuracy and should always be reported for reproducibility. Hyperparameter optimization can be done using a variety of libraries (Table 8), including Hyperopt,95 scikit-optimize,96 and Optuna.97 Auto-sklearn98 provides automatic hyperparameter tuning tools and includes visualization. Recent reviews regarding hyperparameter optimization and additional tools for use with Python can be found in the review article by Bischl et al.99 (particularly sections 2, 6.5.4, and 6.5.5) and references within. Syne Tune100 supports many optimization methods and offers distributed computing, multifidelity methods, transfer learning, and multiobjective optimization that can optimize not only model accuracy but latency simultaneously. Using efficient statistical models, GPyOpt101 and SMAC102 provide hyperparametrization using Bayesian optimization via Gaussian process models, with the aim of reducing the number of evaluations needed.
Table 8. Libraries Used for Hyperparameterization.
| software | link | license |
|---|---|---|
| Hyperopt95 | https://github.com/hyperopt/hyperopt | custom |
| scikit-optimize96 | https://scikit-optimize.github.io | BSD-3-Clause |
| Optuna97 | https://optuna.org | MIT |
| auto-sklearn98 | https://github.com/automl/auto-sklearn | BSD-3-Clause |
| Tune and Syne Tune100 | https://github.com/awslabs/syne-tune | Apache-2.0 |
| GPyOpt101 | http://sheffieldml.github.io/GPyOpt | BSD-3-Clause |
| SMAC102 | https://github.com/automl/SMAC3 | BSD-3-Clause |
2.4.4. AutoML for Classical Machine Learning
Fully automating entire ML processes allows users to quickly try and roll out models for specific tasks with less need for in-depth knowledge about ML. However, one should be warned that the more control over the learning process is given to automated frameworks, the more vigilant one should be when analyzing the results. We mention two well-known example libraries that perform AutoML (Table 9). TPOT103 uses genetic programming to create ML pipelines with scikit-learn. AutoGOAL104 uses a framework for program synthesis for AutoML.
Table 9. Libraries Used for AutoML of Classical Machine Learning.
| software | link | license |
|---|---|---|
| TPOT103 | https://github.com/EpistasisLab/tpot | LGPL-3.0 |
| AutoGOAL104 | https://github.com/autogoal/autogoal | MIT |
2.4.5. AutoML for Deep Learning
Although in deep learning we generally do not perform feature engineering, model selection and hyperparameter optimization can still take significant time, effort, and resources. AutoML for deep learning is an active field, whose most widely used libraries are listed in Table 10. Auto-PyTorch105 and AutoKeras106 are two well-known examples that support PyTorch and Keras/TensorFlow models, respectively.
Table 10. Libraries Used for AutoML in Deep Learning.
| software | link | license |
|---|---|---|
| Auto-PyTorch105 | https://github.com/automl/Auto-PyTorch | Apache-2.0 |
| AutoKeras106 | https://github.com/keras-team/autokeras | Apache-2.0 |
3. Computational Chemistry Tools
The usage of ML to improve the primary methodological tools in computational chemistry is uniquely important since they are widely used in researching different problems. In the following, we will focus on the first-principles methods of quantum mechanics (QM) and density functional theory (DFT) and the Newtonian-based methods of molecular mechanics (MM) and molecular dynamics (MD).
3.1. Quantum Mechanics and Density Functional Theory
The fields of QM, DFT, and ML generally overlap in different ways.8,9,14,15,25,34 The largest overlap is the use of QM/DFT calculations to create reliable data for the training and validation of a model. The experimental-level accuracy that can be obtained by QM is offset by the high resource cost of the calculations. However, once these datasets are generated, they provide an exciting opportunity for training significantly faster ML models that compute a variety of wave-function-based observables (e.g., orbital energies, polarizability).8,107−110 Due to their speed, these QM-/DFT-trained ML algorithms are enabling researchers to perform MD simulations in time domains that are generally prohibitive for QM/DFT methods.111
Concerning OSS, there are several tools for performing QM/DFT calculations (Table 11). Psi4 is a sophisticated OSS that is built upon C++ and Python.112,113 Another well-known tool is NorthWest Chemistry (NWChem), which was written in Fortran and C and thus does not natively interface with Python.114,115 OpenMolcas is a modularly developed Fortran code with a Python input parser for control.116,117 Quantum ESPRESSO is a DFT-based software that focuses on material modeling and includes some Python integration in its recent version.118,119 Abacus120 and BigDFT121,122 were designed to perform DFT calculations on very large systems. A software tool that is strongly coupled with Python is the Python-based Simulations of Chemistry Framework (PySCF) library.123,124 Semiempirical software tools for modeling very large systems include MOPAC,125 DivCon,126 and DFTB+.127 Finally, additional electronic-structure-based software that is available for performing specialized modeling (e.g., MD simulations, condensed phase) includes ABINIT128 and Siesta.129
Table 11. OSS Available for Computing QM and DFT Target Data for Training and Validation.
| software | link | license |
|---|---|---|
| Abacus120 | https://github.com/deepmodeling/abacus-develop | LGPL-3.0 |
| ABINIT128 | https://www.abinit.org | GPL-3.0 |
| BigDFT121,122 | https://bigdft.org | multiple |
| DivCon126 | http://www.merzgroup.org/divcon.html | unavailable |
| DFTB+127 | https://dftbplus.org | multiple |
| MOPAC2016125 | http://openmopac.net | unavailable |
| NWChem114,115 | https://www.nwchem-sw.org | Educational Community License-2.0 |
| OpenMolcas116,117 | https://gitlab.com/Molcas/OpenMolcas | LGPL-2.0 |
| Psi4112,113 | https://psicode.org | LGPL-3.0 and GPL-3.0 |
| PySCF123,124 | https://github.com/PySCF/PySCF | Apache-2.0 |
| Quantum ESPRESSO118,119 | https://www.quantum-espresso.org | GPL |
| SIESTA129 | https://siesta-project.org/siesta | GPL-3.0 |
Another overlap between the fields is the use of ML to optimize DFT calculations themselves130 (Table 12). For example, ML can help to optimize or replace the DFT parameters that are embedded within the theory. With the goal of improving functions for DFT theory, several OSS tools have been developed, including PROPerty Prophet (PROPhet; written in C++),131 D3-GP,132 Deep Kohn–Sham (DeepKS),133,134 NeuralXC,135 NNFunctional,136,137 Differentiable Quantum Chemistry (DQC),138 JAX-DFT,139 Compressed scale-Invariant DEnsity Representation (Cider),140 Fourth-order Expansion of the X Hole,141 Symbolic Functional Evolutionary Search (SyFES),142 and CF22D (written in Fortran).143 The D3-GP workflow implements Gaussian process regression and batchwise-variance-based (as opposed to sequential-variance-based) sampling to improve D3-type dispersion corrections in DFT calculations.132 SyFES is unique in that it generates DFT functionals in a symbolic form that are easier to interpret. Including prior knowledge into training exchange functionals was done in JAX-DFT,139 whose GitHub repository links to a Google Colaboratory notebook for demonstration.
Table 12. Computational-Chemistry-Focused ML Tools for Improving DFT Functionals.
| software | link | license | public data | public model |
|---|---|---|---|---|
| CF22D143 | 10.5281/zenodo.7306137 | CC-BY-4.0 | Y | N |
| Cider140 | https://github.com/mir-group/CiderPress | MIT | Y | N |
| D3-GP132 | https://zenodo.org/record/7785794 | unavailable | Y | N |
| DeepKS133,134 | https://github.com/deepmodeling/deepks-kit | LGPL-3.0 | Y | N |
| DQC138 | https://github.com/diffqc/dqc | Apache-2.0 | Y | N |
| Fourth-order Expansion of the X Hole141 | https://gitlab.com/electronic-structure-udem/fourth-order-expansion-of-the-x-hole | unavailable | Y | Y |
| JAX-DFT139 | https://github.com/google-research/google-research/tree/master/jax_dft | Apache-2.0 | Y | Y |
| NeuralXC135 | https://github.com/semodi/neuralxc | BSD-3-Clause | Y | Y |
| NNFunctional136 | https://github.com/ml-electron-project/NNfunctional | MIT | Y | Y |
| PROPhet131 | https://github.com/biklooost/PROPhet | GPL-3.0 | N | N |
| SyFES142 | 10.5281/zenodo.6767222 | CC-BY-4.0 | Y | N |
3.2. Molecular Mechanics Force Fields
Critical to all MM-based methodologies are their employed force fields, whose state-of-the-art parameter optimization is being pursued by using ML. MM potentials are explicitly defined by Newtonian equations, which are generally divided into bonded (e.g., bonds, angles, torsion) and nonbonded (e.g., partial atomic charges and Lennard-Jones) components. Of these, the nonbonded parameters are often the isolated targets of the ML algorithms. Alternatively, the functional form of the force field equation can be replaced entirely by ML that models the potential energy surface (see section 4.3), still enabling MD simulations to be performed.
Table 13 provides representative ML algorithms for determining nonbonded and bonded parameters. GA4AMOEBA (written in Fortran) uses a genetic algorithm and MP2 target data to parametrize polarizable force fields, specifically the electrostatic and van der Waals parameters, for use with AMOEBA.144 The force field precursors for metal–organic frameworks (FFP4MOF) tool was developed for use in materials research and is able to predict nonbonded parameters for metal-containing systems.145 Molecule-specific (i.e., ammonium perchlorate, pentafluoroethane, difluoromethane) examples of optimizing Lennard-Jones parameters through multiobjective surrogate-assisted Gaussian process regression and support vector machine workflows can be found in ref (146) and its two cited GitHub repositories. Similarly, PREMSO uses a presampling-enhanced, surrogate-assisted global evolutionary optimization strategy that allows the use of features at different scales (e.g., single-molecule and bulk-phase observables).147 Also released recently, Thürlemann et al. developed a GNN to predict nonbonded parameters based on QM target data,148 which includes atom typing prediction. Coming out of the Chodera lab, the impressive Extensible Surrogate Potential Optimized by Message-passing Algorithms (espaloma) uses a GNN to perceive chemical environments and then predict bonded and nonbonded parameters.149
Table 13. Computational-Chemistry-Focused ML Tools for Force Field Parameter Determination.
| software | link | license | public data | public model |
|---|---|---|---|---|
| espaloma149 | https://github.com/choderalab/espaloma | MIT | Y | Y |
| GA4AMOEBA144 | https://github.com/AmYingLi/GA4AMOEBA | unavailable | Y | N |
| GNN Parametrized Forcefields148 | https://github.com/rinikerlab/GNNParametrizedFF | MIT | Y | Y |
| FFP4MOF145 | https://github.com/korolewadim/ffp4mof | MIT | Y | Y |
| PREMSO147 | https://github.com/maxm89/PREMSO-2022 | GPL-3.0 | Y | N |
Partial atomic charges (PACs) unto themselves have had several research groups develop ML concepts for their prediction (Table 14). Focusing on small molecules, the Atom-Path-Descriptor (APD) uses a new type of atomic descriptor for training random forest and extreme gradient boosting models for predicting PAC.150 To predict Quantum Theory of Atoms in Molecules’ PACs, the NNAIMQ (an NN model) was created.151 The SuperAtomicCharge model, a feed-forward NN, was written to predict QM-derived RESP, DDEC4, and DDEC78 PACs.152 For metal-containing systems, mpn_charges153 and PAC in Metal–Organic Frameworks (PACMOF)154 were developed using message-passing NN and a random-forest approach, respectively. The PhysNet algorithm predicts both dipole moments and PACs for larger systems such as peptides, as well as their energies and forces.155 To generate PACs for even larger systems (e.g., proteins), the Electron-Passing NN (epnn) was created.156 The drude_electrostatic_dnn algorithm was developed to generate PACs for the Drude polarizable force field.157 Extending the PAC concept, ESP-DNN158 (GNN) predicts the electrostatic potential surface trained on B3LYP/6-311G**//B3LYP/6-31G* data. Thürlemann et al. developed an equivariant GNN approach that predicts multipoles (e.g., dipole and quadrupole) that includes a database of electrostatic potentials and multipoles.159
Table 14. Computational-Chemistry-Focused ML Tools for Partial Atomic Charge Determination.
| software | link | license | public data | public model |
|---|---|---|---|---|
| APD150 | https://github.com/jkwang93/Atom-Path-Descriptor-based-machine-learning | unavailable | Y | N |
| DeepFMPO160 | https://github.com/giovanni-bolcato/deepFMPOv3D | MIT | Y | N |
| drude_electrostatic_dnn157 | https://github.com/mackerell-lab/drude_electrostatic_dnn | BSD-2-Clause | Y | Y |
| epnn156 | https://github.com/derekmetcalf/epnn | unavailable | Y | Y |
| EquivariantMultipoleGNN159 | https://github.com/rinikerlab/EquivariantMultipoleGNN | MIT | Y | Y |
| ESP-DNN158 | https://github.com/AstexUK/ESP_DNN | Apache-2.0 | Y | Y |
| mpn_charges153 | https://github.com/SimonEnsemble/mpn_charges | unavailable | Y | Y |
| NNAIMQ151 | https://github.com/m-gallegos/NNAIMQ | CC-BY-NC-SA-4.0 | Y | Y |
| PACMOF154 | https://github.com/snurr-group/pacmof | BSD-3-Clause New or Revised | Y | N |
| PhysNet155 | https://github.com/MMunibas/PhysNet | MIT | Y | N |
| SuperAtomicCharge152 | https://github.com/zjujdj/SuperAtomicCharge | Apache-2.0 | Y | Y |
3.3. Molecular Dynamics
Classical-physics-based MD simulations can also be used to generate data for use in ML algorithms. OSS available for performing MD simulations includes AmberTools,161 CP2K,162 GROMACS,163 LAMMPS,164 OpenMM,165 and ORAC166 (Table 15). The TorchMD software was created as a framework for MD simulations that can implement a mix of classical and ML potentials.167 PLUMED is another framework for performing and analyzing MD simulations, which interfaces with 10 MD software codes.168
Table 15. OSS Available for Computing Molecular Mechanics and Molecular Dynamics Target Data for Training and Validation.
| software | link | license |
|---|---|---|
| AmberTools161 | https://ambermd.org/AmberTools.php | GPL-3.0 (mostly) |
| CP2K162 | www.cp2k.org | GPL-2.0 |
| GROMACS163 | www.gromacs.org | LGPL-2.1 |
| LAMMPS164 | www.lammps.org | GPL-2.0 |
| OpenMM165 | https://openmm.org | MIT and LGPL |
| ORAC166 | http://www1.chim.unifi.it/orac | GPL |
| TorchMD167 | https://github.com/torchmd | MIT |
| PLUMED168 | www.plumed.org | LGPL-3.0 |
To enhance sampling of MD simulations5,17,18 (Table 16), several groups have developed the following ML approaches: Variational Dynamical Encoder (VDE)169 and Reweighted Autoencoded Variational Bayes for Enhanced Sampling (RAVE).170 As their names suggest, these algorithms focus on how encoders and decoders may be used to generate synthetic data based on known data. Learn the Effective Dynamics (LED) offers a unique approach that uses ML in conjunction with coarse-gaining and atomistic simulations.171 The mapping between the coarse- and fine-grained system is achieved by using an autoencoder, while a recurrent NN advances the latent space dynamics. In a different approach, Deep Generative Markov State Model (DeepGenMSM) predicts new possible configurations for a molecular system.172 GLOW is an algorithmic workflow that combines Gaussian accelerated MD to generate structural maps that are then used in a convolutional NN to identify reaction coordinates of biomolecules.173 To enhance sampling of rare events, Bonati et al. developed the Deep Time-lagged Independent Component Analysis (Deep-TICA).174 In addition to their interest in characterizing a molecular system, collective variables are used to enhance sampling in MD simulations. Identifying collective variables associated with slow or hard-to-model modes in an MD simulation is the focus of Molecular Enhanced Sampling with Autoencoders (MESA),175 FABULOUS (genetic algorithms and NN),176 COVAEM,177 and DeepCV (deep autoencoder NN).178,179 To bypass the use of MD simulations for sampling, Atomistic Adversarial Attacks can generate molecular conformation and nonbonded configurations, which is achieved by combining uncertainty quantification, automatic differentiation, adversarial attacks, and active learning.180
Table 16. Computational-Chemistry-Focused ML Tools for Enhancing MD Simulations.
| software | link | license | public data | public model |
|---|---|---|---|---|
| Atomistic Adversarial Attacks180 | https://github.com/learningmatter-mit/Atomistic-Adversarial-Attacks | MIT | Y | Y |
| COVAEM177 | https://github.com/ai-atoms/covaem | MIT | N | N |
| DeepCV179 | https://lubergroup.pages.uzh.ch/deepcv and https://gitlab.uzh.ch/lubergroup/deepcv | MIT | Y | Y |
| DeepGenMSM172 | https://github.com/markovmodel/deep_gen_msm | unavailable | Y | N |
| Deep-TICA174 | https://github.com/luigibonati/deep-learning-slow-modes | unavailable | Y | N |
| FABULOUS176 | https://github.com/Ensing-Laboratory/FABULOUS | LGPL-3.0 | Y | N |
| GLOW173 | http://miaolab.org/GLOW | MIT | N | N |
| LED171 | https://github.com/cselab/LED | unavailable | N | N |
| MESA175 | https://github.com/weiHelloWorld/accelerated_sampling_with_autoencoder | MIT | Y | N |
| RAVE170 | https://github.com/tiwarylab/RAVE | MIT | Y | N |
| VDE169 | https://github.com/msmbuilder/vde | MIT | Y | N |
Analysis of existing MD-generated data is an additional area where ML meets computational chemistry24,44 (Table 17), primarily involving data dimensionality reduction. A general and popular OSS Python library for analysis is MDAnalysis (https://www.mdanalysis.org; GPL-2),181,182 which is used in various ML projects (e.g., MESA,175 EncoderMap,183,184 MDMachineLearning,185 RTMScore186). EncoderMap combines autoencoders with multidimensional scaling for dimensionality reduction and can generate structures in the reduced space—for example, to visually examine a protein’s conformational changes along a pathway.183,184 Another dimensionality reduction approach to identify metastable states is the Gaussian mixture variational autoencoder (GMVAE)187 and the stateinterpreter.188 DiffNets uses the autoencoder’s dimensionality reduction idea to identify the structural features that are predictive of biochemical differences between protein variants using MD simulations of those variants.189 Additional approaches for using existing MD trajectories to train a machine for predicting protein conformations (e.g., metastable states) are MDMachineLearning185 and Molearn (convolutional NN).190 The State Predictive Information Bottleneck (SPIB) algorithm learns the reaction coordinate within MD trajectories.191 A unique pixel-based approach was developed in the Interpretable Convolutional Neural Network-based deep learning framework for MD (ICNNMD) algorithm.192 ICNNMD represents protein conformations obtained from MD simulations as pixel maps that are subsequently used to perform feature extractions and then classification. Finally, it should be noted that shallow learning concepts (e.g., clustering, principal component analysis) are incorporated into the Markov State Model Python package MSMBuilder (http://msmbuilder.org; LGPL-2.1) for statistical-based predictive modeling that uses MD input data.193
Table 17. Computational-Chemistry-Focused ML Tools for Analyzing MD Simulations.
| software | link | license | public data | public model |
|---|---|---|---|---|
| DiffNets189 | https://github.com/bowman-lab/diffnets | LGPL-3.0 | Y | N |
| EncoderMap183,184 | https://github.com/AG-Peter/EncoderMap | LGPL-3.0 | Y | N |
| GMVAE187 | https://github.com/yabozkurt/gmvae | unavailable | Y | Y |
| ICNNMD192 | https://github.com/Jane-Liu97/ICNNMD | unavailable | Y | N |
| MDMachineLearning185 | https://github.com/Imay-King/MDMachineLearning | MIT | Y | N |
| Molearn190 | https://github.com/Degiacomi-Lab/molearn | GPL-3.0 | Y | N |
| SPIB191 | https://github.com/tiwarylab/State-Predictive-Information-Bottleneck | MIT | N | N |
| Stateinterpreter188 | https://github.com/luigibonati/md-stateinterpreter | MIT | Y | N |
3.4. Docking, Protein–Ligand Interactions, and Virtual Screening
The role that ML has within the docking community was reviewed in refs (16), (30), and (43). OSS for performing small molecule docking to proteins includes Autodock Vina,194 MOLS 2.0,195 rDock,196 SEED,197 and Smina198 (Table 18). GNINA, a recently released tool, docks molecules using an ensemble of trained convolutional NNs as a scoring function.199,200 To facilitate the integration of ML with docking, the DOCKSTRING package (https://dockstring.github.io; Apache 2.0) was developed that enables learned models to be easily benchmarked.201 This package contains Python wrappers, a large dataset of scores and poses, and benchmarking tasks and employs AutoDock Vina as its docking engine. The CABSdock software focuses on the flexible docking of peptides to proteins.202 In addition to peptide–protein docking, the LightDock software can also perform docking between DNA–protein and protein–protein biopolymers.203,204 Very recently, the GNN EDM-Dock was developed that generates protein–ligand poses from distance matrices and implicitly incorporates protein flexibility through coarse-graining of the protein.205 Researchers interested in computer-aided drug design11,27,37,51 should also be aware of the Open Drug Discovery Toolkit (https://github.com/oddt/oddt; BSD-3),206 a Python code that implements the ML scoring functions NNscore207 and RFscore197 (both of which were developed in the early 2010s).
Table 18. OSS Available for Docking Calculations.
| software | link | license |
|---|---|---|
| Autodock Vina194 | https://vina.scripps.edu | Apache 2.0 |
| CABSdock202 | https://bitbucket.org/lcbio/cabsdock/src/master | MIT |
| LightDock203,204 | https://lightdock.org and https://github.com/lightdock/lightdock | GPL-3.0 |
| MOLS 2.0195 | https://sourceforge.net/projects/mols2-0 | LGPL-2.1 |
| Open Drug Discovery Toolkit206 | https://github.com/oddt/oddt | BSD-3-Clause |
| rDock196 | http://rdock.sourceforge.net | LGPL-3.0 |
| SEED197 | https://gitlab.com/CaflischLab/SEED | GPL-3.0 |
| Smina198 | https://sourceforge.net/projects/smina | GPL-2.0 |
| GNINA200 | https://github.com/gnina/gnina | Apache-2.0 and GPL-2.0 |
Concerning scoring functions, significant ML research has focused on improving them for use in docking software and for virtual screening (Table 19). A recent assessment indicated that learned scoring functions can improve predictions, but diligence must be maintained since their performance depends upon the training dataset used (e.g., the degree of protein sequence similarity).208 Such functions include RF-RF-Score-VS,209 Δvina eXreme Gradient Boosting (XGB),210 AEScore (Δ-learning),211 ET-Score (extremely randomized trees),212 ΔLinF9 XGB,213 PharmRF (random forests),214 RTMScore,186 XLPFE,215 DeepRMSD+Vina (multilayer perceptron),216 and GB-Score (Gradient Boosting Trees).217 For docking ligands to RNA, the AnnapuRNA scoring function—a k-nearest neighbors and feed-forward NN scoring function—was developed that uses coarse-grained representations in the modeling.218 Based on KDEEP’s three-dimensional (3D) convolutional NN,219 DeepBSP is an algorithm that predicts the most likely native complex structure from an ensemble of poses generated by a docking software.220
Table 19. Machine-Learned Scoring Functions and Tools for Docking and Virtual Screening.
| software | link | license | public data | public model |
|---|---|---|---|---|
| AEScore211 | https://github.com/bigginlab/aescore | BSD-3-Clause | Y | Y |
| AnnapuRNA218 | https://github.com/filipspl/AnnapuRNA | GPL-3.0 | Y | Y |
| DeepBSP220 | https://github.com/BaoJingxiao/DeepBSP | GPL-3.0 | Y | Y |
| DeepRMSD+Vina216 | https://github.com/zchwang/DeepRMSD-Vina_Optimization | unavailable | Y | Y |
| ΔLinF9XGB213 | https://github.com/cyangNYU/delta_LinF9_XGB | GPL-3.0 | Y | Y |
| ΔvinaXGB210 | https://github.com/jenniening/deltaVinaXGB | GPL-3.0 | Y | Y |
| EDM-Dock205 | https://github.com/MatthewMasters/EDM-Dock | MIT | N | Y |
| ET-Score212 | https://github.com/miladrayka/ET_Score | GPL-3.0 | Y | Y |
| GB-Score217 | https://github.com/miladrayka/GB_Score | AGPL-3.0 | Y | Y |
| NNforDocking221 | https://github.com/mksmd/NNforDocking | MIT | Y | N |
| PharmRF214 | https://github.com/Prasanth-Kumar87/PharmRF | unavailable | Y | Y |
| RF-Score-VS209 | https://github.com/oddt/rfscorevs_binary | BSD-3-Clause | Y | N |
| RTMScore186 | https://github.com/sc8668/RTMScore | MIT | Y | Y |
| XLPFE215 | https://github.com/LinaDongXMU/XLPFE | unavailable | Y | N |
Similar to the goals of docking, several groups have created algorithms for predicting protein–ligand interactions (see Table 20 and the review in ref (20)). DeepDTA,222 DeepConv-DTI,223 DeepCDA,224 and DeepScreen225 make use of a convolutional NN. Using two stacked 3D convolutional NNs—for learning intramolecular and intermolecular interactions, respectively—InteractionGraphNet predicts protein–ligand interactions and binding affinities.226 The ML-ensemble-docking algorithm explored whether ML could aggregate docking scores for better binding predictions.227 GNN_DTI228 and DeepNC229 predict ligand–receptor interactions using GNN. SSnet is built upon a deep learning framework that uses a protein’s secondary structure to predict how a ligand might bind.230 STAMP-DPI uses a protein’s sequence to generate a predicted contact map.231 The NNforDocking code uses an NN to predict possible binding pockets, followed by AutoDock Vina to obtain a possible protein–ligand configuration.221 Of the algorithms listed above, GNN_DTI uses 3D structural information through an adjacency network and a distance-aware graph attention mechanism.228 Protein–Ligand Interaction Fingerprints (ProLIF) is a Python analysis tool for analyzing molecular dynamics trajectories, docking simulations, and experimental structures.232 LUNA is another interaction analysis Python tool that implements Extended Interaction FingerPrint (EIFP), Functional Interaction FingerPrint (FIFP), and Hybrid Interaction FingerPrint (HIFP) for both protein–ligand and protein–protein complexes, outputting a PyMOL session for visualization.233
Table 20. ML Tools for Protein–Ligand Interactions.
| software | link | license | public data | public model |
|---|---|---|---|---|
| DeepCDA224 | https://github.com/LBBSoft/DeepCDA | unavailable | Y | N |
| DeepConv-DTI223 | https://github.com/GIST-CSBL/DeepConv-DTI | GPL-3.0 | Y | N |
| DeepDTA222 | https://github.com/hkmztrk/DeepDTA | unavailable | Y | N |
| DeepNC229 | https://github.com/thntran/DeepNC | unavailable | Y | N |
| DeepScreen225 | https://github.com/cansyl/DEEPscreen | GPL-3.0 | Y | N |
| GNN_DTI228 | https://github.com/jaechanglim/GNN_DTI | unavailable | Y | N |
| InteractionGraphNet226 | https://github.com/zjujdj/InteractionGraphNet/tree/master | unavailable | Y | Y |
| SSnet230 | https://github.com/ekraka/SSnet | MIT | Y | Y |
| STAMP-DPI231 | https://github.com/biomed-AI/STAMP-DPI | GPL-3.0 | Y | Y |
Several recent reviews have been published on virtual screening and ML—for example, see refs (23), (28), and (30). In addition to the scoring functions mentioned above, other goals have been pursued. GATNN is a molecular-graph-focused NN tool that enables scaffold hopping during its virtual screening.234 A fully automated tool for performing virtual screening is PyRMD, which uses a random matrix discriminant to screen and identify potential active ligands based on trained biological activity data.235 This algorithm was designed to be usable by both coding experts and nonexperts. RealVS uses transfer learning and graph attention networks to improve predictions and enable a level of model interpretability.236 An interesting deep-learning-based tool for use in virtual screening is DeepCoy, a GNN that enables researchers to generate property-matched decoy molecules based on a known active ligand input.237 The TocoDecoy tool also creates decoys but uses a conditional recurrent NN.238
4. Selected Research-Focused Topics
4.1. Protein Binding Site Prediction
Very closely related to the ideas just covered but often categorized separately is the goal of predicting small-molecule binding pockets on proteins, whose representative ML algorithms are given in Table 21. One such approach, P2Rank, uses a random forest approach and was developed using Apache’s Groovy and Java programming languages.239,240 The kalasanty algorithm uses image segmentation via a 3D U-net convolutional NN to predict binding sites.241 DeepSurf is a 3D convolutional residual NN that uses a protein surface representation of local 3D voxelized grids to identify binding pockets.242 PUResNet employs a U-net variant residual NN that predicts binding sites based on structure similarity.243 The DeepPocket algorithm employs Fpocket (https://github.com/Discngine/fpocket; MIT)244 and a 3D convolutional NN to identify sites, ranking them using a classification model and mapping the binding sites’ shapes using a segmentation U-net-like model.245 Also using a U-net architecture in a 3D NN, the InDeep algorithm focuses on predicting binding pockets in and near the protein–protein interaction (PPI) interface.246 Using a primary sequence as input, BiRDS is a residual NN that predicts a protein’s amino acids that are most likely to form a binding region.247 PointSite uses an atom-level point cloud segmentation approach in a submanifold sparse convolution NN.248 Mentioned in section 3.4, NNforDocking uses an NN to make a binding pocket prediction as part of its workflow.221
Table 21. ML Tools for Predicting Possible Binding Pockets in Proteins.
| software | link | license | public data | public model |
|---|---|---|---|---|
| BiRDS247 | https://github.com/devalab/BiRDS | unavailable | Y | Y |
| DeepPocket245 | https://github.com/devalab/DeepPocket | MIT | Y | Y |
| DeepSurf242 | https://github.com/stemylonas/DeepSurf | AGPL-3.0 | Y | Y |
| InDeep246 | https://gitlab.pasteur.fr/InDeep/InDeep | unavailable | Y | Y |
| kalasanty241 | https://gitlab.com/cheminfIBB/kalasanty | BSD-3-Clause | Y | Y |
| P2Rank239,240 | https://github.com/rdk/p2rank | MIT | Y | N |
| PointSite248 | https://github.com/PointSite/PointSite | MIT | Y | Y |
| PUResNet243 | https://github.com/jivankandel/PUResNet | unavailable | Y | Y |
4.2. Protein–Protein Interactions and Protein Folding
The prediction of PPIs has seen a lot of ML activity, as reviewed in refs (42), (46), (50), and (249) and given in Table 22. PPI prediction can occur at different resolution levels, for example, sequence-based versus structure-based ML approaches.42 Depending on the focus, the output can range from identification of the primary sequence components (i.e., the interacting amino acids) to 3D “docked” structures. ML-based PPI prediction algorithms include pipgcn,250 masif (convolutional NN),251 GraphPPIS (graph NN),252 Struct2Graph (graph attention network),253 and DeepHomo2 (web server and downloadable package).254 To identify a protein’s possible interfacial binding region, Fout et al. developed pipgcn using graph NN, where each amino acid residue is described by a node.250 Masif employs the unique approach of computing a protein–surface fingerprint, which is then used to predict PPIs (it can also predict protein–ligand interactions). Concerning ML scoring functions for protein–protein docking, DOcking decoy selection with Voxel-based deep neural nEtwork (DOVE) scans PPIs with a 3D voxel, resulting in their ranking.255 iScore provides a scoring function built using a support vector machine and random-walk graph kernels approach.256,257 DeepRank258 and DeepRank-GNN259 were built using a 3D convolutional NN and GNN, respectively. TopNetTree is a novel algorithm that predicts the binding affinity changes for a PPI upon an amino acid mutation.260
Table 22. ML Tools for Exploring Protein–Protein Interactions and Protein Folding.
Data and model are no longer available through the provided link.
Protein structure prediction (i.e., protein folding) is another related topic that is making significant advances due to ML.21,49 MELD uses Bayesian inference to predict protein structure using a limited amount of experimental information and physics-based modeling.261 DeepECA is an end-to-end convolutional NN to predict a protein’s intramolecular contacts and its secondary structure.262 AlphaFold263 and RoseTTAFold264 are well-known, with well-maintained GitHub repositories. A 2022 paper by Bryant et al. describes AlphaFold2, which can predict heterodimeric protein complexes.265 DMPfold2 is a third approach, which uses a multiple sequence alignment as input to generate folded prediction in an ultrafast time frame.266 While not directly involving ML code, ColabFold (https://github.com/sokrypton/ColabFold; MIT) is OSS that couples MMseqs2,267,268 a many-against-many sequence searching and clustering algorithm, with AlphaFold2/RoseTTAFold and can be implemented in Google’s Colaboratory.269
In a closely related topic, DLPacker’s goal is to predict amino acid side-chain conformations using a 3D convolutional U-net architecture.270 For structure validation (e.g., from a prediction using the above method), as one possible application, one can use the Int2Cart algorithm. Int2Cart uses a gated recurrent NN that detects internal coordinate correlation and refines bond distances and bending angles for a given set of torsion rotations.271
4.3. Energies and Forces
In many situations, understanding an experimental observation is significantly aided by elucidating a portion of the system’s potential energy (PE) surface, the corresponding forces, and its free energies. Consequently, this is why much research is devoted to modeling PE using physics-based approaches (i.e., QM and MM) and now by data-driven ML8,12,22,25,29,31,33,38,39 (Table 23). The resulting learned potentials and force fields can be used to predict conformational energies or perform MD simulations. Isayev and co-workers developed Atomic Simulation Environment–Accurate NeurAl networK engINe for Molecular Energies (ASE-ANI),272−274 TorchANI,275 and AIMNet276 as approaches for realizing universal ML interatomic potentials for neutral organic molecules.39 Additional algorithms for modeling PE surfaces include DeePMD-kit,277,278 PES-Learn,279 SchNetPack,280−282 Symmetric Gradient Domain Machine Learning (sGDML),283,284 ml-dft,285 Fast Learning of Atomistic Rare Events (FLARE),286 Module for Ab Initio Structure Evolution (MAISE) (written in C but has a Python wrapper available called MAISE-NET),287 KIM-based learning-integrated fitting framework (KLIFF),288 and AisNet.289 SchNetPack predicts not only PE but also other observables such as atomic forces, formation energies, and dipole moments.282 Building off of SchNet281 and using a trainable encoding module, AisNet can predict the energy and forces for molecules and crystalline materials (e.g., crystalline ceramics and multicomponent alloys).289 Also building off of SchNet,281 the g4mp2-atomization-energy algorithm was developed for predicting atomization energies.290
Table 23. ML Tools for Predicting Molecular Energies, Solvation Energies, and Binding Affinities.
The TorsionNet algorithm enables the prediction of PE curves as a function of torsion angle rotation and was trained using active learning.291 However, a license for the OpenEye Toolkit is needed for its full implementation. The component-based machine-learned intermolecular force field (CLIFF) algorithm can predict intermolecular PE energies by combining physics-based potential forms with ML-based parametrization.292 The concept behind BAND-NN is to predict a molecule’s energy and to enable geometry optimization, which divides the energy into bonds, angles, dihedrals, and nonbonded terms within the NN.293 With a different focus, the D3-GP workflow implements Gaussian process regression and batchwise-variance-based (as opposed to sequential-variance-based) sampling to improve D3-type dispersion corrections in DFT calculations.132 In the domain of materials science, fast_reorg_energy_prediction makes use of the ChiRo ML model to predict the reorganization energy.294
Several groups have worked on using ML to predict binding affinities. Ligand–receptor binding affinities can be computed using Pafnucy,295 DeepAffinity,296 GLXE,297 OctSurf,298 OnionNet-2,299,300 and Hybrid Attention-Based Convolutional Neural Network (HAC-Net).301 Pafnucy’s convolutional NN makes use of a 3D grid input representation with 1 Å resolution for affinity prediction. DeepAffinity was trained to predict binding affinities based on IC50, Ki and Kd.296 The GLXE297 algorithm combines MM/GBSA with shallow and deep ML approaches to predict binding free energies. The OctSurf algorithm computes the 3D surface areas of the protein pocket and ligand to predict a resulting binding affinity.298 The two-dimensional convolutional NN OnionNet-2 generates rotation-free pairwise contacts between protein and ligand atoms to predict binding free energies.299,300 HAC-Net is one of the newest algorithms, which combines the concept of attention with a 3D convolutional NN to compute protein–ligand binding affinity.301
DeepMoleNet is an atomwise NN that predicts a molecule’s internal energy, thermodynamic energies at 298.15 K, HOMO and LUMO energies, and zero-point vibrational energy as well as dipole moment, polarizability, electronic spatial extent, and heat capacity.302 Its model and training data are not directly released but can be requested if used noncommercially. Finally, ML is used to predict solvation free energies, as represented by the following algorithms: Hybrid FEP/ML,303 MLSolvA,304 A3D-PNAConv-FT,305 chemprop_solvation,306 and MolSolv.307
4.4. Molecule Generation
ML has also been used to generate suggestions for new molecules4,13,40 (Table 24). These hypothetical molecules represent new ideas that synthetic chemists can pursue toward various ends (e.g., drug design). Variational autoencoders provide one path for generating new ideas, as implemented in LigDream308 and the Molecular Graph Conditional Variational Autoencoder (MGCVAE).309 LigDream creates structures based on a seed molecule’s volume by coupling a shape autoencoder to convolutional and recurrent NNs.308 MGCVAE includes the use of a GNN to propose molecules that have specific properties (e.g., log P).309 DeepGraphMolGen performs a multiobjective optimization using reinforcement learning based on a graph convolution policy approach for generating molecules with desired properties.310 A unique concept for 3D molecule generation is to include a representation of the receptor’s topology in the model training, as realized by the LiGAN algorithm.311 Boström and co-workers developed Deep Fragment-based Multi-Parameter Optimization (DeepFMPO),160 which generates fragments (i.e., residues) that can be combined to form a new molecule. Extending this idea to biopolymers, Graph-Based Protein Design, an autoregressive language model, was created to generate an amino acid sequence that should fold into a desired 3D structure.312 MOLEcule Generation Using reinforcement Learning with Alternating Reward (MoleGuLAR) is an algorithm that proposes molecules for targeting specific protein binding sites using reinforcement learning.313 Using a ML transformer model that was trained on pairs of similar bioactive molecules, the transform-molecules algorithm can generate new molecules that ideally would have higher potency against a specific protein target.314 Kao et al. developed QuMolGAN and related models to explore the use of quantum generative adversarial networks to create new molecules.315
Table 24. ML Tools for Designing New Molecules.
| software | link | license | public data | public model |
|---|---|---|---|---|
| DeepFMPO160 | https://github.com/giovanni-bolcato/deepFMPOv3D | MIT | Y | N |
| DeepGraphMolGen310 | https://github.com/dbkgroup/prop_gen | unavailable | Y | N |
| DeLinker316 | https://github.com/oxpig/DeLinker | BSD-3-Clause | Y | Y |
| DEVELOP317 | https://github.com/oxpig/DEVELOP | BSD-3-Clause | Y | Y |
| DRLinker319 | https://github.com/biomed-AI/DRlinker | unavailable | Y | Y |
| Graph-Based Protein Design312 | https://github.com/jingraham/neurips19-graph-protein-design | MIT | Y | N |
| LigDream308 | https://github.com/compsciencelab/ligdream | AGPL-3.0 | Y | Y |
| LiGAN311 | https://github.com/mattragoza/liGAN | GPL-2.0 | Y | Y |
| MGCVAE309 | https://github.com/mhlee216/MGCVAE | unavailable | Y | N |
| MoleGuLAR313 | https://github.com/devalab/MoleGuLAR | unavailable | Y | Y |
| QuMolGAN315 | https://github.com/pykao/QuantumMolGAN-PyTorch | MIT | Y | N |
| SyntaLinker318 | https://github.com/YuYaoYang2333/SyntaLinker | MIT | Y | N |
| transform-molecules314 | https://github.com/pfizer-opensource/transform-molecules | Apache-2.0 | Y | N |
With a slightly different end goal, several algorithms approach the creation of new molecules by designing chemical linkers to combine fragments. The graph-based DeLinker NN generates possible linkers for connecting two residues.316 Combining DeLinker with a 3D pharmacophore concept, DEep Vision-Enhanced Lead OPtimisation (DEVELOP) was developed to generate potentially more impactful molecular suggestions.317 SyntaLinker learns and uses the “rules” for linking fragments via syntactic patterns embedded in SMILES strings.318 Using reinforcement learning, DRLinker designs linkers that join two desired residues together, whose resulting molecules are tailored towards specific attributes.319
4.5. Chemical Reactions and Synthesis
Synthetic chemists now have the opportunity to use ML to predict the products from chemical reactions and design synthetic routes to obtain a desired outcome32,47,52 (Table 25). For predicting products, Molecular Transformer uses SMILES strings as input reactants to an autoregressive encoder–decoder.320 Based on a GNN active learning architecture, DeepReac+ was developed to predict reaction products and to help optimize experimental conditions for organic reactions.321 Also using SMILES strings as input and a k-nearest neighbor classifier, the differential reaction fingerprint (DRFP) algorithm was developed for predicting reaction classification and yield prediction.322 The competing-reactions algorithm focuses on predicting the energetic barrier height of possible chemical pathways based on reactants.323 The Open-NMT-py algorithm uses multitask transfer learning to predict the stereoisomeric products of enzyme-catalyzed reactions using a SMILES input.86 Employing a graph-to-graph transformer architecture, G2GT is a retrosynthesis predictor.324 As a pipeline tool, AiZynthTrain can be used to train new synthesis prediction models that are usable by the AiZynthFinder OSS (https://github.com/MolecularAI/aizynthfinder; MIT).325,326
Table 25. ML Tools for Predicting the Products of Reactants and Optimizing Synthesis.
| software | link | license | public data | public model |
|---|---|---|---|---|
| AiZynthTrain326 | https://github.com/MolecularAI/aizynthtrain | Apache-2.0 | N | N |
| competing-reactions323 | https://github.com/ferchault/competing-reactions | unavailable | Y | N |
| DeepReac+321 | https://github.com/bm2-lab/DeepReac | Apache-2.0 | Y | Y |
| DRFP322 | https://github.com/reymond-group/drfp | MIT | Y | Y |
| G2GT324 | https://github.com/Anonnoname/G2GT_2 | unavailable | Y | N |
| Molecular Transformer320 | https://github.com/pschwllr/MolecularTransformer | MIT | Y | Y |
| OpenNMT-py86 | https://github.com/reymond-group/OpenNMT-py | MIT | Y | N |
| ReTReK327 | https://github.com/clinfo/ReTReK | MIT | N | N |
4.6. Conformation Generation
Elucidating and understanding 3D structures and the conformational space of molecules is an important goal in many research fields (e.g., spectroscopy and drug design), whose difficulty increases with a molecule’s number of rotatable bonds. The ML community has generated several algorithms that help predict the structures of conformations (Table 26). The DL4Chem-geometry algorithm uses a conditional variational graph autoencoder to predict conformations by learning the underlying PE surface, which utilizes Cartesian coordinates as part of the input data.328 GraphDG predicts conformations by combining a conditional variational autoencoder with a Euclidean distance geometry algorithm, resulting in an approach that is invariant to rotation and translation.329 Keeping the invariance goal in mind, ConfVAE was developed using bilevel programming to provide an end-to-end generation approach.330 The same group also developed ConfGF, which adds the idea of gradient fields (analogous to force fields) and Langevin dynamics to their ML workflow.331 Auto3D addresses the challenge of sampling configurational stereoisomers when using a SMILES string for generating 3D conformers; it is trained on the author’s atomistic neural network potentials (i.e., AIMNet, ANI-2x, ANI-2xt) and is able to identify the lowest-energy conformer.332 The MolTaut algorithm generates possible tautomer geometries, performs subsequent optimization using the ANI-2x ML model, and ranks the results based on energies (i.e., internal and solvation).307
Table 26. ML Tools for Exploring the Conformational Space of Molecules.
| software | link | license | public data | public model |
|---|---|---|---|---|
| Auto3D332 | https://github.com/isayevlab/Auto3D_pkg | MIT | Y | Y |
| ConfGF331 | https://github.com/DeepGraphLearning/ConfGF | MIT | Y | N |
| ConfVAE330 | https://github.com/MinkaiXu/CGCF-ConfGen | unavailable | Y | Y |
| DL4Chem-geometry328 | https://github.com/nyu-dl/dl4chem-geometry | BSD-3-Clause | Y | Y |
| GraphDG329 | https://github.com/gncs/graphdg | MIT | Y | N |
| MolTaut307 | https://github.com/xundrug/moltaut | GPL-2.0 | Y | Y |
4.7. Spectral Data
A long-standing goal of computational chemistry is the modeling of spectral data, with the recent ML contribution given in Table 27. As one of its goals, SchNarc can predict the UV spectrum by modeling a molecule’s transition dipole moments and excited-state energies using a continuous-filter convolutional NN.333 ML_UVvisModels is an algorithm that extends SchNet,290 SolTranNet,334,335 Chemprop-IR (https://github.com/gfm-collab/chemprop-IR; MIT), and a model developed by Ghosh et al.336 to predict UV–vis spectra.337 The prediction of electronic spectra was explored in the GMM-NEA algorithm, which uses probabilistic machine learning.338 In the GMM-NEA paper, an intriguing application of their model was to identify anomalous QM calculations that could lead to incorrect spectra predictions.
Table 27. ML Tools for Predicting Spectra.
| software | link | license | public data | public model |
|---|---|---|---|---|
| CANDIY-spectrum339 | https://github.com/chopralab/candiy_spectrum | unavailable | Y | N |
| FTIRMachineLearning340 | https://github.com/Ohio-State-Allen-Lab/FTIRMachineLearning | Apache-2.0 | Y | N |
| GMM-NEA338 | https://github.com/lucerlab/GMM-NEA | LGPL-2.1 | Y | N |
| MLforvibspectroscopy341 | https://github.com/elizabeththrall/MLforPChem/tree/main/MLforvibspectroscopy | CC-BY-SA-4.0 | Y | N |
| ML_UVvisModels337 | https://github.com/PNNL-CompBio/ML_UVvisModels | BSD-2-Clause | Y | Y |
| SchNarc333 | https://github.com/schnarc/schnarc | MIT | Y | N |
Not strictly coming from the computational chemistry community but aligning with the goal of assisting experimentalists is the use of ML to help assign functional groups to recorded spectra. For Fourier transform infrared (FTIR) spectroscopy and mass spectrometry, CANDIY-spectrum was developed that uses a multilayer perceptron NN.339 Focusing solely on FTIR spectra, 15 functional group identification models were created using the FTIRMachineLearning algorithms.340 With a focus on educating bachelor students—and thus, a valuable resource for learning—the MLforvibspectroscopy repository contains Jupyter notebooks that demonstrate how one can build a model for identifying functional groups from vibrational frequencies.341
4.8. pKa
For predicting a molecule’s pKa using ML (Table 28), OPERA,342,343 Machine-learning-meets-pKa,344 MolGpKa,345 pKAI,346 pkasolver,347 and DeepKa348 approaches are available as OSS. The pKa models of OPERA consist of a support vector machine, an extreme gradient boosting, and a four-layer fully connected NN. The Machine-learning-meets-pKa model predicts macroscopic pKa values for monoprotic molecules. MolGpKa and pkasolver use a convolutional GNN to make pKa predictions. The pKAI346 model was developed to predict the pKa values of amino acids within a protein. It was trained on the change in pKa values (i.e., ΔpKa) computed when a water-immersed amino acid residue was placed in the protein environment relative to that of its neutral form. To predict the pKa of proteins, one can use DeepKa, which was recently trained and validated using 23817 and 2735 data values, respectively.348,349
Table 28. ML Tools for Predicting pKa.
| software | link | license | public data | public model |
|---|---|---|---|---|
| DeepKa348,349 | https://gitlab.com/yandonghuang/deepka | GPL-3.0 | Y | N |
| Machine-learning-meets-pKa344 | https://github.com/czodrowskilab/Machine-learning-meets-pKa | MIT | Y | N |
| MolGpKa345 | https://github.com/xundrug/molgpka | MIT | Y | Y |
| OPERA343 | https://github.com/NIEHS/OPERA | MIT | Y | Y |
| pKAI346 | https://github.com/bayer-science-for-a-better-life/pKAI | MIT | Y | Y |
| pkasolver347 | https://github.com/mayrf/pkasolver | MIT | Y | Y |
5. A Foray into Additional Topics
In addition to the OSS described above, several additional projects warrant mention (Table 29) but are not easily classified into the above categories. Of particular note is DeepChem, a Python library that was developed to simplify the creation of machine and deep learning models for use in life sciences.350 DeepChem’s deep learning algorithms can be used with Keras, TensorFlow, PyTorch, and Jax frameworks and can include shallow learning libraries like sklearn.
Table 29. ML Tools for Exploring Miscellaneous Topics.
General package.
Molecular fignerprints.
Molecular similarity.
Ionization energy.
Partition coefficient.
Multiple and diverse observables.
Metal–organic framework stability.
Aqueous solubility.
ADMET.
Various interesting and more isolated concepts.
Molecular Fingerprints
There are several existing OSS for generating molecular fingerprints (i.e., representations) that can be used in ML,351,352 most notably OpenBabel353 and RDKit.354 While these tools are not ML algorithms, they are frequently used in ML projects. An ML tool for generating fingerprints is PretrainModels, a self-supervised learning algorithm that uses a bidirectional encoder transformer for reading input SMILES strings.355
Molecular Similarity
Computing molecular similarity is a topic that often arises in pharmaceutical research. As part of a virtual screening project, VS-SVM (implemented in MATLAB) uses support vector machines to predict pairwise similarity.356 Published in 2022, MLKRR is a similarity-based model built using the QM9 dataset and a metric learning approach for kernel ridge regression, and it was used to predict atomization energies.357
ADMET
As part of the drug design process, the prediction of absorption, distribution, metabolism, excretion, and toxicity (ADMET) can be done using ML algorithms.6 Using a molecular fingerprints random forest algorithm written in R and Java, FP-ADMET is a shallow learning approach for predicting ADMET.358 ADMETboost makes use of DeepChem350 and XGBoost359 to generate a predictor; its source code is available on GitHub, with the trained model accessible through a web interface.360
Partition Coefficient
The partition coefficient is a useful observable for environmental chemistry, toxicology, and pharmacology. Using the DeepChem library, log_P_prediction uses convoluted graphs and an NN to predict the octanol–water partition coefficient (i.e., log P).361 Focused on drug lipophilicity, rescoss_logp_ml was developed to predict log P for a given molecule.362 Including liquid chromatography retention time as a molecular descriptor for training, the multilayer perceptron p_chem_prop_CEVR computes log P and the distribution coefficient log D.363
Ionization Energies
The IonEner-Pred GitHub repository contains fourteen different conventional and GNN models that can compute ionization energies.364
Metal–Organic Stability
Specifically for metal–organic molecules, MOFSimplify was trained using graph- and pore-geometry-based representations to predict their thermal stability and their stability upon solvent removal.365
Aqueous Solubility
As mentioned above, aqueous solubility is one of the properties that the chemprop366−368 software can compute. Focusing solely on this observable is SolTranNet,334,335 which uses SMILES strings as the input representation.
Multiple Observables
Many of the above-mentioned projects focus on using ML approaches for investigating a single or a very limited number of observables. However, there are a few projects that are attempting to provide models that can predict diverse observables. One notable example of this is chemprop,366−368 which can predict water solubility, hydrate free energies in water, octanol–water distribution coefficients, protein–ligand binding affinity, toxicity, drug side effects, activation energies, reaction enthalpies, rate constants, yields, and reaction classes to name a few. The Chiral InterRoto-Invariant Neural Network (ChIRo) was designed to enable property predictions that depend upon a molecule’s chirality.369 The hierarchical informative GNN (HiGNN) represents another framework for predicting diverse molecular properties, whose use was demonstrated for a range of physiochemical, biophysics, physiology, and toxicity observables.370
Miscellaneous
To help identify chemical and structural reasons for why molecules predicted by ML models satisfy certain properties, the algorithm exmol was developed;371 it finds counterfactuals and is generalizable to any ML model. The MolScribe algorithm uses an encoder–decoder architecture to create a molecular graph representation from a line drawing (e.g., a skeletal formula, Markush structures).372 To predict glass transition temperatures and melting points for organic molecules, tgBoost uses the random forest and XGBoost frameworks.373
6. Discussion
From our survey of ML algorithms within the computational chemistry domain, some observations can help shape our understanding of the current state. Through hand curation of the 179 repositories listed in Tables 12–14, 16, 17, and 19–29, 94% of the projects reported the availability of the training data, but only 54% reported the availability of a usable model. While the community does well in disclosing training data, there is a clear need to encourage researchers to include an optimized model in their repositories, which includes the code, a list of necessary libraries, and the model parameters (i.e., weights). Alternatively, one can include the necessary hyperparameters to train the model for projects having small training datasets, although this does not guarantee that the model will produce the exact same results.
Building upon the concept of openness, an interesting and helpful development within the ML field is the idea of open ML platforms. Code repositories (e.g., GitHub) are usually used either to publish data, models, and associated scripts for the purpose of reproducibility or to host actively maintained software packages. In contrast, open platforms allow ML experts and users to freely share and organize data, enabling more effective, visible, and collaborative works to be done. The platforms usually collect open data, open models, benchmarks, and applications and enhance these collections through simple search and filter mechanisms. One such example is OpenML (https://www.openml.org; BSD-3), which was created by the Open Machine Learning Foundation.375 Since the existing platforms usually contain an incredibly diverse set of ML problems, it might be worth starting a discussion toward establishing an open ML platform for computational chemistry. Such a platform could house the diverse datasets that computation chemistry researchers are interested in, ranging from QM- to biopolymer-focused data, and enable quick experimentation of ML code across our subdomains.
To tally the number of forks, we conducted a query using GitHub’s application programming interface (API). With a few exceptions (e.g., AlphaFold), many of the ML computational chemistry OSS projects are written, maintained, and expanded upon by small groups of researchers. For a given project, one can assume that these individuals mostly come from a single PI research group. Using the number of forks as a metric, the six most popular projects are AlphaFold (1867 forks), DeepChem (1500), chemprop (455), DeePMD-kit (428), RoseTTAFold (411), and SchNetPack (175). As can be seen in Figure 2, the remaining repositories show only a slight interest by the community, with an average of 13 forks when these top six projects are excluded. Relatively speaking, this is likely due to the highly specific subject matter that the code was developed to investigate (e.g., generating conformers) and thus does not accurately represent its perceived quality or value. To further promote and enhance the reuse of code, we believe that the community would benefit from the creation of an open platform for ML in computational chemistry as well as the other benefits discussed above.
Figure 2.
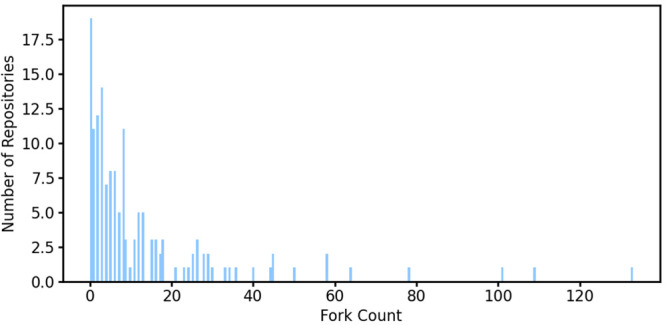
Histogram of the number of forks for projects on Github, leaving out six projects that have ≥175 forks.
Concerning licenses, 78% of the projects surveyed included a license in their release, while 22% did not. Although a large portion of the projects lacked a license, their use of OSS libraries implies that they have an open-source license model, as often dictated through license inheritance. However, it is advised to include a license model when publishing code. As shown in Table 30, the MIT license was predominately favored (50% when unavailable licenses are excluded), followed by GPL-3.0 (14%), Apache-2.0 (11%), and BSD-3-Clause (8%), all of which are FOSS licensing schemes. The MIT license is a BSD-style permissive license that allows for code reuse within proprietary software once proper attribution conditions are met. The GPL license is a copyleft model that requires all subsequent works that use the code to also use a GPL license. For readers who desire additional guidance, we recommend GitHub’s “Choose an open source license’’ website (https://choosealicense.com).
Table 30. License Types and How Often They Are Used for the Reported Computational Chemistry ML Tools, as Provided in Repositories.
| license | count | license | count |
|---|---|---|---|
| MIT | 69 | CC-BY-4.0 | 2 |
| unavailable | 40 | LGPL-2.1 | 2 |
| GPL-3.0 | 20 | BSD-3-Clause New or Revised | 1 |
| Apache-2.0 | 15 | CC-BY-NC-SA-4.0 | 1 |
| BSD-3-Clause | 11 | CC-BY-SA-4.0 | 1 |
| LGPL-3.0 | 5 | MPL-2.0 | 1 |
| GPL-2.0 | 4 | custom | 1 |
| AGPL-3.0 | 3 | multiple | 1 |
| BSD-2-Clause | 2 |
To survey what libraries are used by ML code developers within the computational chemistry domain, a second query using GitHub’s API was done that focused on Python scripts and Jupyter notebooks (i.e., .py and .ipynb files). Specifically, libraries were extracted from the “import” and “from library import” statements and uniquely identified to avoid counting multiple instances within the same repository. For illustration, if two Python scripts called the same library within a given project repository, only one call was counted. The results for the top 40 used libraries are shown in Figure 3, which are classified as being scientific, Python standard, or an additional library. NumPy was the most commonly used library. It offers the benefit of performing fast computations due to its interface with C-encoded algorithms and vectorized computations (i.e., loops are not required to iterate mathematics on data arrays). The other notable scientific software includes Pandas, PyTorch, sklearn, SciPy, Matplotlib, RDKit, TensorFlow, and ASE. The five frequently used third-party libraries were setuptools, tqdm, yaml, utils, and pytest. It is worth noting that PyTorch was used 1.8 times more often than TensorFlow, aligning with the observation that PyTorch has become more popular.376,377 In their comparison of these two libraries, Novac et al. concluded that PyTorch offers a more beginner-friendly experience with faster training and execution, while TensorFlow allows for higher flexibility and better resulting accuracy.378 These top imported libraries represent clear starting points for those who are new to the field and want to start understanding, writing, and using ML algorithms.
Figure 3.
Histogram of the top libraries called (≥24 times) in 167 GitHub Python projects reported herein. Scientific libraries are shown in blue, Python3 standard libraries (https://docs.python.org/3/library) in red, and additional libraries in green.
The value of FOSS to the research community is manyfold, including free access to tools (thus helping to mend global economic inequality), quicker advancement of ideas (both through their application and by providing starting points for extending and modifying algorithms), and increased reproducibility of published data.379,380 Concerning education, having access to code is invaluable, especially when one is learning independently and when coupled with a published paper. The educational interest includes having examples for specific library usage, understanding how the mathematics and physics are encoded, the approximations that are made (e.g., weighting factors, convergence criteria), and the improved elucidation of a specific published workflow. With regard to the last point, while clear and informative writing within a paper’s methodology section is imperative, important details can be omitted, and misunderstanding can occur. When coding is an important component of the research, processing a paper’s information and internalizing its knowledge can be greatly enhanced by accessing the algorithms. Consequently, it is important to release code that is concisely written, logically constructed (e.g., isolation of ideas), and appropriately commented (e.g., docstrings)—essentially, to follow good scientific practices.381−383
7. Conclusions
The prevalence of ML in the computational chemistry domain is growing rapidly. The above survey of ML algorithms indicates exciting endeavors toward improving theoretical modeling and predicting chemical and biological observables. There is a clear push by many journals, reviewers, and principal investigators to publish the raw data used in a paper as well as the code, training data, and parameters (or the hyperparameters at a minimum) of the trained ML models. Assuming that our surveyed projects reasonably represent the computational chemistry community, it is encouraging to see the high prevalence of providing the code and data in an open manner. However, we encourage researchers to also publish their optimized models. Doing so would enable non-ML experts to more easily use the models in their research endeavors and would improve the opportunity to reproduce the published data within the model’s accompanying article. Finally, we believe that the community would benefit from a centralized online platform dedicated to the development of computational-chemistry-focused ML algorithms and datasets.
Acknowledgments
The authors thank Abanoub Abdelmalak, Thomas Gerlach, Nico Piel, and Lena Riem, who contributed to the initial literature searches and discussions on what would be an informative article concept.
The authors declare no competing financial interest.
References
- Sonnenburg S.; Braun M. L.; Ong C. S.; Bengio S.; Bottou L.; Holmes G.; LeCun Y.; Müller K.-R.; Pereira F.; Rasmussen C. E.; Rätsch G.; Schölkopf B.; Smola A.; Vincent P.; Weston J.; Williamson R. The Need for Open Source Software in Machine Learning. J. Mach. Learn. Res. 2007, 8, 2443–2466. [Google Scholar]
- Langenkamp M.; Yue D. N.. How Open Source Machine Learning Software Shapes AI. In Proceedings of the 2022 AAAI/ACM Conference on AI, Ethics, and Society; ACM: New York, 2022; pp 385–395
- Pirhadi S.; Sunseri J.; Koes D. R. Open Source Molecular Modeling. J. Mol. Graphics Modell. 2016, 69, 127–143. 10.1016/j.jmgm.2016.07.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elton D. C.; Boukouvalas Z.; Fuge M. D.; Chung P. W. Deep Learning for Molecular Design – a Review of the State of the Art. Mol. Syst. Des. Eng. 2019, 4, 828–849. 10.1039/C9ME00039A. [DOI] [Google Scholar]
- Bernetti M.; Bertazzo M.; Masetti M. Data-Driven Molecular Dynamics: A Multifaceted Challenge. Pharmaceuticals 2020, 13, 253. 10.3390/ph13090253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cãceres E. L.; Tudor M.; Cheng A. C. Deep Learning Approaches in Predicting ADMET Properties. Future Med. Chem. 2020, 12, 1995–1999. 10.4155/fmc-2020-0259. [DOI] [PubMed] [Google Scholar]
- Cartwright H.Machine Learning in Chemistry: The Impact of Artificial Intelligence; Royal Society of Chemistry, 2020 [Google Scholar]
- Dral P. O. Quantum Chemistry in the Age of Machine Learning. J. Phys. Chem. Lett. 2020, 11, 2336–2347. 10.1021/acs.jpclett.9b03664. [DOI] [PubMed] [Google Scholar]
- Gastegger M.; Marquetand P. In Machine Learning Meets Quantum Physics; Schütt K. T., Chmiela S., von Lilienfeld O. A., Tkatchenko A., Tsuda K., Müller K.-R., Eds.; Lecture Notes in Physics, Vol. 968; Springer International Publishing, 2020; pp 233–252. [Google Scholar]
- Janet J.; Kulik H.. Machine Learning in Chemistry; ACS In Focus; American Chemical Society, 2020 [Google Scholar]
- Jiménez-Luna J.; Grisoni F.; Schneider G. Drug Discovery with Explainable Artificial Intelligence. Nat. Mach. Intell. 2020, 2, 573–584. 10.1038/s42256-020-00236-4. [DOI] [Google Scholar]
- Kang P.-L.; Shang C.; Liu Z.-P. Large-Scale Atomic Simulation via Machine Learning Potentials Constructed by Global Potential Energy Surface Exploration. Acc. Chem. Res. 2020, 53, 2119–2129. 10.1021/acs.accounts.0c00472. [DOI] [PubMed] [Google Scholar]
- von Lilienfeld O. A.; Müller K.-R.; Tkatchenko A. Exploring Chemical Compound Space with Quantum-based Machine Learning. Nat. Rev. Chem. 2020, 4, 347–358. 10.1038/s41570-020-0189-9. [DOI] [PubMed] [Google Scholar]
- Noé F.; Tkatchenko A.; Müller K.-R.; Clementi C. Machine Learning for Molecular Simulation. Annu. Rev. Phys. Chem. 2020, 71, 361–390. 10.1146/annurev-physchem-042018-052331. [DOI] [PubMed] [Google Scholar]
- Machine Learning Meets Quantum Physics; Schütt K., Chmiela S., von Lilienfeld O. A., Tkatchenko A., Tsuda K., Müller K., Eds.;. Lecture Notes in Physics, Vol. 968; Springer International Publishing, 2020 [Google Scholar]
- Shen C.; Ding J.; Wang Z.; Cao D.; Ding X.; Hou T. From Machine Learning to Deep Learning: Advances in Scoring Functions for Protein–ligand Docking. Wiley Interdiscip. Rev.: Comput. Mol. Sci. 2020, 10, e1429. 10.1002/wcms.1429. [DOI] [Google Scholar]
- Sidky H.; Chen W.; Ferguson A. L. Machine Learning for Collective Variable Discovery and Enhanced Sampling in Biomolecular Simulation. Mol. Phys. 2020, 118, e1737742. 10.1080/00268976.2020.1737742. [DOI] [Google Scholar]
- Wang Y.; Lamim Ribeiro J. M.; Tiwary P. Machine Learning Approaches for Analyzing and Enhancing Molecular Dynamics Simulations. Curr. Opin. Struct. Biol. 2020, 61, 139–145. 10.1016/j.sbi.2019.12.016. [DOI] [PubMed] [Google Scholar]
- Wieder O.; Kohlbacher S.; Kuenemann M.; Garon A.; Ducrot P.; Seidel T.; Langer T. A Compact Review of Molecular Property Prediction with Graph Neural Networks. Drug Discovery Today Technol. 2020, 37, 1–12. 10.1016/j.ddtec.2020.11.009. [DOI] [PubMed] [Google Scholar]
- Abbasi K.; Razzaghi P.; Poso A.; Ghanbari-Ara S.; Masoudi-Nejad A. Deep Learning in Drug Target Interaction Prediction: Current and Future Perspectives. Curr. Med. Chem. 2021, 28, 2100–2113. 10.2174/0929867327666200907141016. [DOI] [PubMed] [Google Scholar]
- AlQuraishi M. Machine Learning in Protein Structure Prediction. Curr. Opin. Chem. Biol. 2021, 65, 1–8. 10.1016/j.cbpa.2021.04.005. [DOI] [PubMed] [Google Scholar]
- Behler J. Four Generations of High-Dimensional Neural Network Potentials. Chem. Rev. 2021, 121, 10037–10072. 10.1021/acs.chemrev.0c00868. [DOI] [PubMed] [Google Scholar]
- Ghislat G.; Rahman T.; Ballester P. J. Recent Progress on the Prospective Application of Machine Learning to Structure-based Virtual Screening. Curr. Opin. Chem. Biol. 2021, 65, 28–34. 10.1016/j.cbpa.2021.04.009. [DOI] [PubMed] [Google Scholar]
- Glielmo A.; Husic B. E.; Rodriguez A.; Clementi C.; Noé F.; Laio A. Unsupervised Learning Methods for Molecular Simulation Data. Chem. Rev. 2021, 121, 9722–9758. 10.1021/acs.chemrev.0c01195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang B.; von Lilienfeld O. A. Ab Initio Machine Learning in Chemical Compound Space. Chem. Rev. 2021, 121, 10001–10036. 10.1021/acs.chemrev.0c01303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keith J. A.; Vassilev-Galindo V.; Cheng B.; Chmiela S.; Gastegger M.; Müller K.-R.; Tkatchenko A. Combining Machine Learning and Computational Chemistry for Predictive Insights Into Chemical Systems. Chem. Rev. 2021, 121, 9816–9872. 10.1021/acs.chemrev.1c00107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim J.; Park S.; Min D.; Kim W. Comprehensive Survey of Recent Drug Discovery Using Deep Learning. Int. J. Mol. Sci. 2021, 22, 9983. 10.3390/ijms22189983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kimber T. B.; Chen Y.; Volkamer A. Deep Learning in Virtual Screening: Recent Applications and Developments. Int. J. Mol. Sci. 2021, 22, 4435. 10.3390/ijms22094435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kulichenko M.; Smith J. S.; Nebgen B.; Li Y. W.; Fedik N.; Boldyrev A. I.; Lubbers N.; Barros K.; Tretiak S. The Rise of Neural Networks for Materials and Chemical Dynamics. J. Phys. Chem. Lett. 2021, 12, 6227–6243. 10.1021/acs.jpclett.1c01357. [DOI] [PubMed] [Google Scholar]
- Li H.; Sze K.-H.; Lu G.; Ballester P. J. Machine-learning scoring functions for structure-based virtual screening. Wiley Interdiscip. Rev.: Comput. Mol. Sci. 2021, 11, e1478. 10.1002/wcms.1478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manzhos S.; Carrington T. J. Neural Network Potential Energy Surfaces for Small Molecules and Reactions. Chem. Rev. 2021, 121, 10187–10217. 10.1021/acs.chemrev.0c00665. [DOI] [PubMed] [Google Scholar]
- Meuwly M. Machine Learning for Chemical Reactions. Chem. Rev. 2021, 121, 10218–10239. 10.1021/acs.chemrev.1c00033. [DOI] [PubMed] [Google Scholar]
- Mishin Y. Machine-learning Interatomic Potentials for Materials Science. Acta Mater. 2021, 214, 116980. 10.1016/j.actamat.2021.116980. [DOI] [Google Scholar]
- Morawietz T.; Artrith N. Machine Learning-accelerated Quantum Mechanics-based Atomistic Simulations for Industrial Applications. J. Comput. Aided Mol. Des. 2021, 35, 557–586. 10.1007/s10822-020-00346-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Musil F.; Grisafi A.; Bartók A. P.; Ortner C.; Csãnyi G.; Ceriotti M. Physics-Inspired Structural Representations for Molecules and Materials. Chem. Rev. 2021, 121, 9759–9815. 10.1021/acs.chemrev.1c00021. [DOI] [PubMed] [Google Scholar]
- Nandy A.; Duan C.; Taylor M. G.; Liu F.; Steeves A. H.; Kulik H. J. Computational Discovery of Transition-metal Complexes: From High-throughput Screening to Machine Learning. Chem. Rev. 2021, 121, 9927–10000. 10.1021/acs.chemrev.1c00347. [DOI] [PubMed] [Google Scholar]
- Peña-Guerrero J.; Nguewa P. A.; García-Sosa A. T. Machine Learning, Artificial Intelligence, and Data Science Breaking into Drug Design and Neglected Diseases. Wiley Interdiscip. Rev.: Comput. Mol. Sci. 2021, 11, e1513. 10.1002/wcms.1513. [DOI] [Google Scholar]
- Westermayr J.; Marquetand P. Machine Learning for Electronically Excited States of Molecules. Chem. Rev. 2021, 121, 9873–9926. 10.1021/acs.chemrev.0c00749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zubatiuk T.; Isayev O. Development of Multimodal Machine Learning Potentials: Toward a Physics-Aware Artificial Intelligence. Acc. Chem. Res. 2021, 54, 1575–1585. 10.1021/acs.accounts.0c00868. [DOI] [PubMed] [Google Scholar]
- Bilodeau C.; Jin W.; Jaakkola T.; Barzilay R.; Jensen K. F. Generative Models for Molecular Discovery: Recent Advances and Challenges. Wiley Interdiscip. Rev.: Comput. Mol. Sci. 2022, 12, e1608. 10.1002/wcms.1608. [DOI] [Google Scholar]
- Bojar D.; Lisacek F. Glycoinformatics in the Artificial Intelligence Era. Chem. Rev. 2022, 122, 15971–15988. 10.1021/acs.chemrev.2c00110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Casadio R.; Martelli P. L.; Savojardo C. Machine Learning Solutions for Predicting Protein-protein Interactions. Wiley Interdiscip. Rev.: Comput. Mol. Sci. 2022, 12, e1618. 10.1002/wcms.1618. [DOI] [Google Scholar]
- Crampon K.; Giorkallos A.; Deldossi M.; Baud S.; Steffenel L. A. Machine-learning Methods for Ligand-protein Molecular Docking. Drug Discovery Today 2022, 27, 151–164. 10.1016/j.drudis.2021.09.007. [DOI] [PubMed] [Google Scholar]
- Kaptan S.; Vattulainen I. Machine Learning in the Analysis of Biomolecular Simulations. Adv. Phys.: X 2022, 7, 2006080. 10.1080/23746149.2021.2006080. [DOI] [Google Scholar]
- Kuntz D.; Wilson A. K. Machine Learning, Artificial Intelligence, and Chemistry: How Smart Algorithms Are Reshaping Simulation and the Laboratory. Pure Appl. Chem. 2022, 94, 1019–1054. 10.1515/pac-2022-0202. [DOI] [Google Scholar]
- Li S.; Wu S.; Wang L.; Li F.; Jiang H.; Bai F. Recent advances in predicting protein–protein interactions with the aid of artificial intelligence algorithms. Curr. Opin. Struct. Biol. 2022, 73, 102344. 10.1016/j.sbi.2022.102344. [DOI] [PubMed] [Google Scholar]
- Park S.; Han H.; Kim H.; Choi S. Machine Learning Applications for Chemical Reactions. Chem. - Asian J. 2022, 17, e202200203. 10.1002/asia.202200203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reiser P.; Neubert M.; Eberhard A.; Torresi L.; Zhou C.; Shao C.; Metni H.; van Hoesel C.; Schopmans H.; Sommer T.; Friederich P. Graph Neural Networks for Materials Science and Chemistry. Commun. Mater. 2022, 3, 93. 10.1038/s43246-022-00315-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schauperl M.; Denny R. A. AI-based Protein Structure Prediction in Drug Discovery: Impacts and Challenges. J. Chem. Inf. Model. 2022, 62, 3142–3156. 10.1021/acs.jcim.2c00026. [DOI] [PubMed] [Google Scholar]
- Soleymani F.; Paquet E.; Viktor H.; Michalowski W.; Spinello D. Protein–protein Interaction Prediction with Deep Learning: A Comprehensive Review. Comput. Struct. Biotechnol. J. 2022, 20, 5316–5341. 10.1016/j.csbj.2022.08.070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Staszak M.; Staszak K.; Wieszczycka K.; Bajek A.; Roszkowski K.; Tylkowski B. Machine Learning in Drug Design: Use of Artificial Intelligence to Explore the Chemical Structure–biological Activity Relationship. Wiley Interdiscip. Rev.: Comput. Mol. Sci. 2022, 12, e1568. 10.1002/wcms.1568. [DOI] [Google Scholar]
- Tu Z.; Stuyver T.; Coley C. W. Predictive Chemistry: Machine Learning for Reaction Deployment, Reaction Development, and Reaction Discovery. Chem. Sci. 2023, 14, 226–244. 10.1039/D2SC05089G. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Conda, 2022. https://github.com/conda/conda (accessed 2023-06-21).
- Anaconda Documentation, 2020. https://docs.anaconda.com/ (accessed 2023-06-21).
- Continuum Analytics, Inc. Miniconda, 2017. https://docs.conda.io/en/latest/miniconda.html (accessed 2023-06-21).
- GitHub, Inc., GitHub. https://github.com (accessed 2022-08-28).
- GitLab, Inc. GitLab. https://gitlab.com (accessed 2022-08-28).
- JetBrains s.r.o. PyCharm. https://www.jetbrains.com/pycharm (accessed 2023-06-07).
- Raybaut P.; Córdoba C.; International Community of Volunteers . Spyder: The Scientific Python Development Environment. https://www.spyder-ide.org (accessed 2022-08-28).
- Microsoft . Visual Studio Code. https://code.visualstudio.com (accessed 2022-08-28).
- Kluyver T.; Ragan-Kelley B.; Pérez F.; Granger B. E.; Bussonnier M.; Frederic J.; Kelley K.; Hamrick J.; Grout J.; Corlay S.; Ivanov P.; Avila D.; Abdalla S.; Willing C.; Jupyter Development Team ; Jupyter Notebooks—A Publishing Format for Reproducible Computational Workflows In Positioning and Power in Academic Publishing: Players, Agents and Agendas; Loizides F., Schmidt B., Eds.; IOS Press, 2016; pp 87–90. 10.3233/978-1-61499-649-1-87. [DOI] [Google Scholar]
- Jupyter. https://jupyter.org (accessed 2022-08-28).
- Welcome to Colaboratory. https://colab.research.google.com (accessed 2022-08-28).
- Harris C. R.; Millman K. J.; van der Walt S. J.; Gommers R.; Virtanen P.; Cournapeau D.; Wieser E.; Taylor J.; Berg S.; Smith N. J.; Kern R.; Picus M.; Hoyer S.; van Kerkwijk M. H.; Brett M.; Haldane A.; del Río J. F.; Wiebe M.; Peterson P.; Gérard-Marchant P.; Sheppard K.; Reddy T.; Weckesser W.; Abbasi H.; Gohlke C.; Oliphant T. E. Array Programming with NumPy. Nature 2020, 585, 357–362. 10.1038/s41586-020-2649-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McKinney W.Data Structures for Statistical Computing in Python. In Proceedings of the 9th Python in Science Conference, 2010; pp 56–61.
- The Pandas Development Team . pandas-dev/pandas: Pandas, 2020. https://doi.org/10.5281/zenodo.3509134 (accessed 2023-06-21). 10.5281/zenodo.3509134. [DOI]
- Virtanen P.; Gommers R.; Oliphant T. E.; Haberland M.; Reddy T.; Cournapeau D.; Burovski E.; Peterson P.; Weckesser W.; Bright J.; van der Walt S. J.; Brett M.; Wilson J.; Millman K. J.; Mayorov N.; Nelson A. R. J.; Jones E.; Kern R.; Larson E.; Carey C. J.; Polat d.; Feng Y.; Moore E. W.; VanderPlas J.; Laxalde D.; Perktold J.; Cimrman R.; Henriksen I.; Quintero E. A.; Harris C. R.; Archibald A. M.; Ribeiro A. H.; Pedregosa F.; van Mulbregt P.; SciPy 1.0: Fundamental Algorithms for Scientific Computing in Python. Nat. Methods 2020, 17, 261–272. 10.1038/s41592-019-0686-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hunter J. D. Matplotlib: A 2D Graphics Environment. Comput. Sci. Eng. 2007, 9, 90–95. 10.1109/MCSE.2007.55. [DOI] [Google Scholar]
- Waskom M. L. Seaborn: Statistical Data Visualization. J. Open Source Software 2021, 6, 3021. 10.21105/joss.03021. [DOI] [Google Scholar]; https://seaborn.pydata.org (accessed 2023-06-07).
- Meurer A.; Smith C. P.; Paprocki M.; Čertík O.; Kirpichev S. B.; Rocklin M.; Kumar A.; Ivanov S.; Moore J. K.; Singh S.; Rathnayake T.; Vig S.; Granger B. E.; Muller R. P.; Bonazzi F.; Gupta H.; Vats S.; Johansson F.; Pedregosa F.; Curry M. J.; Terrel A. R.; Roučka u.; Saboo A.; Fernando I.; Kulal S.; Cimrman R.; Scopatz A. SymPy: Symbolic Computing in Python. PeerJ Comput. Sci. 2017, 3, e103. 10.7717/peerj-cs.103. [DOI] [Google Scholar]
- Pedregosa F.; Varoquaux G.; Gramfort A.; Michel V.; Thirion B.; Grisel O.; Blondel M.; Prettenhofer P.; Weiss R.; Dubourg V.; Vanderplas J.; Passos A.; Cournapeau D.; Brucher M.; Perrot M.; Duchesnay E. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Givon L. E.; Unterthiner T.; Erichson N. B.; Chiang D. W.; Larson E.; Pfister L.; Dieleman S.; Lee G. R.; van der Walt S.; Menn B.; Moldovan T. M.; Bastien F.; Shi X.; Schlüter J.; Thomas B.; Capdevila C.; Rubinsteyn A.; Forbes M. M.; Frelinger J.; Klein T.; Merry B.; Merill N.; Pastewka L.; Liu L. Y.; Clarkson S.; Rader M.; Taylor S.; Bergeron A.; Ukani N. H.; Wang F.; Lee W.-K.; Zhou Y.. scikit-cuda 0.5.3: A Python Interface to GPU-Powered Libraries, 2019. 10.5281/zenodo.3229433. [DOI]
- Wojciechowski M.Solving Differential Equations by Means of Feed-Forward Artificial Neural Networks. In Artificial Intelligence and Soft Computing; Lecture Notes in Computer Science, Vol. 7267; Springer: Berlin, 2012; pp 187–195.
- Chen T.; Guestrin C.. XGBoost: A Scalable Tree Boosting System. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery & Data Mining; ACM: New York, 2016; pp 785–794.
- Ke G.; Meng Q.; Finley T.; Wang T.; Chen W.; Ma W.; Ye Q.; Liu T.-Y.. LightGBM: A Highly Efficient Gradient Boosting Decision Tree. In Advances in Neural Information Processing Systems 30 (NIPS 2017); Curran Associates, 2017; pp 3149–3157. [Google Scholar]
- GPy: A Gaussian Process Framework in Python, 2012. http://github.com/SheffieldML/GPy (accessed 2023-06-20).
- Matthews A. G. d. G.; van der Wilk M.; Nickson T.; Fujii K.; Boukouvalas A.; León-Villagrá P.; Ghahramani Z.; Hensman J. GPflow: A Gaussian Process Library using TensorFlow. J. Mach. Learn. Res. 2017, 18, 1–6. [Google Scholar]
- Gardner J.; Pleiss G.; Weinberger K. Q.; Bindel D.; Wilson A. G.. GPyTorch: Blackbox Matrix-Matrix Gaussian Process Inference with GPU Acceleration. In Proceedings of the 32nd International Conference on Neural Information Processing Systems (NIPS 2018); Curran Associates, 2018; 7576–7586.
- Paszke A.; Gross S.; Chintala S.; Chanan G.; Yang E.; DeVito Z.; Lin Z.; Desmaison A.; Antiga L.; Lerer A.. Automatic Differentiation in PyTorch. In 31st Conference on Neural Information Processing Systems (NIPS 2017), 2017; https://openreview.net/forum?id=BJJsrmfCZ
- Abadi M.; Agarwal A.; Barham P.; Brevdo E.; Chen Z.; Citro C.; Corrado G. S.; Davis A.; Dean J.; Devin M.; Ghemawat S.; Goodfellow I.; Harp A.; Irving G.; Isard M.; Jia Y.; Jozefowicz R.; Kaiser L.; Kudlur M.; Levenberg J.; Mane D.; Monga R.; Moore S.; Murray D.; Olah C.; Schuster M.; Shlens J.; Steiner B.; Sutskever I.; Talwar K.; Tucker P.; Vanhoucke V.; Vasudevan V.; Viegas F.; Vinyals O.; Warden P.; Wattenberg M.; Wicke M.; Yu Y.; Zheng X.. TensorFlow: Large-scale Machine Learning on Heterogeneous Distributed Systems. arXiv (SSSS), March 16, 2016, 1603.04467, ver. 2. https://arxiv.org/abs/1603.04467 (accessed 2023-05-04).
- Gulli A.; Pal S.. Deep Learning with Keras; Packt Publishing Ltd., 2017 [Google Scholar]
- Kasim M. F.; Vinko S. M. ξ-torch: Differentiable Scientific Computing Library. arXiv (Computer Science.Machine Learning), October 5, 2020, 2010.01921, ver. 1. https://arxiv.org/abs/2010.01921 (accessed 2023-05-04).
- The Theano Development Team . Theano: A Python framework for fast computation of mathematical expressions. arXiv (Computer Science.Symbolic Computation), May 9, 2016, 1605.02688, ver. 1. https://arxiv.org/abs/1605.02688 (accessed 2023-05-04).
- Chen T.; Li M.; Li Y.; Lin M.; Wang N.; Wang M.; Xiao T.; Xu B.; Zhang C.; Zhang Z.. MXNet: A Flexible and Efficient Machine Learning Library for Heterogeneous Distributed Systems. arXiv (Computer Science.Distributed, Parallel, and Cluster Computing), December 3, 2015, 1512.01274, ver. 1. https://arxiv.org/abs/1512.01274 (accessed 2023-05-04).
- Klein G.; Kim Y.; Deng Y.; Senellart J.; Rush A.. OpenNMT: Open-Source Toolkit for Neural Machine Translation. In Proceedings of ACL 2017, System Demonstrations, Vancouver, Canada, 2017; pp 67–72.
- Kreutter D.; Schwaller P.; Reymond J.-L. Predicting Enzymatic Reactions with a Molecular Transformer. Chem. Sci. 2021, 12, 8648–8659. 10.1039/D1SC02362D. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fey M.; Lenssen J. E.. Fast Graph Representation Learning with PyTorch Geometric. arXiv (Computer Science.Machine Learning), April 25, 2019, 1903.02428, ver. 3. https://arxiv.org/abs/1903.02428 (accessed 2023-05-04).
- Battaglia P. W.; Hamrick J. B.; Bapst V.; Sanchez-Gonzalez A.; Zambaldi V.; Malinowski M.; Tacchetti A.; Raposo D.; Santoro A.; Faulkner R.; Gulcehre C.; Song F.; Ballard A.; Gilmer J.; Dahl G.; Vaswani A.; Allen K.; Nash C.; Langston V.; Dyer C.; Heess N.; Wierstra D.; Kohli P.; Botvinick M.; Vinyals O.; Li Y.; Pascanu R.. Relational Inductive Biases, Deep Learning, and Graph Networks. arXiv (Computer Science.Machine Learning), October 17, 2018, 1806.01261, ver. 3. https://arxiv.org/abs/1806.01261 (accessed 2023-05-04).
- Wang M.; Zheng D.; Ye Z.; Gan Q.; Li M.; Song X.; Zhou J.; Ma C.; Yu L.; Gai Y.; Xiao T.; He T.; Karypis G.; Li J.; Zhang Z.. Deep Graph Library: A Graph-Centric, Highly-Performant Package for Graph Neural Networks. arXiv (Computer Science.Machine Learning), August 25, 2020, 1909.01315, ver. 2. https://arxiv.org/abs/1909.01315 (accessed 2023-05-04).
- Kanter J. M.; Veeramachaneni K.. Deep Feature Synthesis: Towards Automating Data Science Endeavors. In 2015 IEEE International Conference on Data Science and Advanced Analytics (DSAA 2015), Paris, France, October 19–21, 2015; IEEE, 2015; pp 717–726. 10.1109/DSAA.2015.7344858. [DOI]
- Galli S.Python Feature Engineering Cookbook: Over 70 Recipes for Creating, Engineering, and Transforming Features to Build Machine Learning Models; Packt Publishing Ltd., 2022. [Google Scholar]
- Christ M.; Braun N.; Neuffer J.; Kempa-Liehr A. W. Time Series Feature Extraction on Basis of Scalable Hypothesis Tests (tsfresh – a Python Package). Neurocomputing 2018, 307, 72–77. 10.1016/j.neucom.2018.03.067. [DOI] [Google Scholar]
- Ali M.PyCaret: An Open Source, Low-Code Machine Learning Library in Python, version 1.0.0, 2020.
- Bengfort B.; Bilbro R. Yellowbrick: Visualizing the Scikit-Learn Model Selection Process. J. Open Source Software 2019, 4, 1075–1079. 10.21105/joss.01075. [DOI] [Google Scholar]
- Bergstra J.; Yamins D.; Cox D. D.. Hyperopt: A Python Library for Optimizing the Hyperparameters of Machine Learning Algorithms. In Proceedings of the 12th Python in Science Conference, 2013; p 20.
- Head T.; et al. scikit-optimize/scikit-optimize: v0.5.2, 2018. 10.5281/zenodo.1207017. [DOI]
- Akiba T.; Sano S.; Yanase T.; Ohta T.; Koyama M.. Optuna: A Next-Generation Hyperparameter Optimization Framework. In Proceedings of the 25th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining; ACM: New York, 2019; pp 2623–2631.
- Feurer M.; Klein A.; Eggensperger K.; Springenberg J.; Blum M.; Hutter F.. Efficient and Robust Automated Machine Learning. In Advances in Neural Information Processing Systems 28 (NIPS 2015); Curran Associates, 2015; pp 2962–2970.
- Bischl B.; Binder M.; Lang M.; Pielok T.; Richter J.; Coors S.; Thomas J.; Ullmann T.; Becker M.; Boulesteix A.-L.; Deng D.; Lindauer M. Hyperparameter Optimization: Foundations, Algorithms, Best Practices, and Open Challenges. Wiley Interdiscip. Rev.: Data Min. Knowl. Discovery 2023, 13, e1484. 10.1002/widm.1484. [DOI] [Google Scholar]
- Salinas D.; Seeger M.; Klein A.; Perrone V.; Wistuba M.; Archambeau C. Syne Tune: A Library for Large Scale Hyperparameter Tuning and Reproducible Research. Proc. Mach. Learn. Res. 2022, 188, 16. [Google Scholar]
- GPyOpt: A Bayesian Optimization Framework in Python, 2016. http://github.com/SheffieldML/GPyOpt (accessed 2023-06-07).
- Lindauer M.; Eggensperger K.; Feurer M.; Biedenkapp A.; Deng D.; Benjamins C.; Ruhkopf T.; Sass R.; Hutter F. SMAC3: A Versatile Bayesian Optimization Package for Hyperparameter Optimization. J. Mach. Learn. Res. 2022, 23, 1–9. [Google Scholar]
- Olson R. S.; Bartley N.; Urbanowicz R. J.; Moore J. H.. Evaluation of a Tree-Based Pipeline Optimization Tool for Automating Data Science. In Proceedings of the Genetic and Evolutionary Computation Conference 2016, New York, 2016; pp 485–492.
- Estevez-Velarde S.; Gutiérrez Y.; Almeida-Cruz Y.; Montoyo A. General-purpose hierarchical optimization of machine learning pipelines with grammatical evolution. Inf. Sci. 2021, 543, 58–71. 10.1016/j.ins.2020.07.035. [DOI] [Google Scholar]
- Mendoza H.; Klein A.; Feurer M.; Springenberg J. T.; Urban M.; Burkart M.; Dippel M.; Lindauer M.; Hutter F.. Towards Automatically-Tuned Deep Neural Networks. In Automated Machine Learning: Methods, Systems, Challenges; Hutter F., Kotthoff L., Vanschoren J., Eds.; Springer, 2019; Chapter 7, pp 135–149. [Google Scholar]
- Jin H.; Chollet F.; Song Q.; Hu X. AutoKeras: An AutoML Library for Deep Learning. J. Mach. Learn. Res. 2023, 24, 1–6. [Google Scholar]
- Schütt O.; VandeVondele J. Machine Learning Adaptive Basis Sets for Efficient Large Scale Density Functional Theory Simulation. J. Chem. Theory Comput. 2018, 14, 4168–4175. 10.1021/acs.jctc.8b00378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coe J. P. Machine Learning Configuration Interaction for ab Initio Potential Energy Curves. J. Chem. Theory Comput. 2019, 15, 6179–6189. 10.1021/acs.jctc.9b00828. [DOI] [PubMed] [Google Scholar]
- Manzhos S. Machine Learning for the Solution of the Schrödinger Equation. Mach. Learn.: Sci. Technol. 2020, 1, 013002. 10.1088/2632-2153/ab7d30. [DOI] [Google Scholar]
- Townsend J.; Vogiatzis K. D. Transferable MP2-Based Machine Learning for Accurate Coupled-Cluster Energies. J. Chem. Theory Comput. 2020, 16, 7453–7461. 10.1021/acs.jctc.0c00927. [DOI] [PubMed] [Google Scholar]
- Brockherde F.; Vogt L.; Li L.; Tuckerman M. E.; Burke K.; Müller K.-R. Bypassing the Kohn-Sham Equations with Machine Learning. Nat. Commun. 2017, 8, 872. 10.1038/s41467-017-00839-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parrish R. M.; Burns L. A.; Smith D. G. A.; Simmonett A. C.; DePrince A. E.; Hohenstein E. G.; Bozkaya U.; Sokolov A. Y.; Di Remigio R.; Richard R. M.; Gonthier J. F.; James A. M.; McAlexander H. R.; Kumar A.; Saitow M.; Wang X.; Pritchard B. P.; Verma P.; Schaefer H. F.; Patkowski K.; King R. A.; Valeev E. F.; Evangelista F. A.; Turney J. M.; Crawford T. D.; Sherrill C. D. Psi4 1.1: An Open-Source Electronic Structure Program Emphasizing Automation, Advanced Libraries, and Interoperability. J. Chem. Theory Comput. 2017, 13, 3185–3197. 10.1021/acs.jctc.7b00174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Turney J. M.; Simmonett A. C.; Parrish R. M.; Hohenstein E. G.; Evangelista F. A.; Fermann J. T.; Mintz B. J.; Burns L. A.; Wilke J. J.; Abrams M. L.; Russ N. J.; Leininger M. L.; Janssen C. L.; Seidl E. T.; Allen W. D.; Schaefer H. F.; King R. A.; Valeev E. F.; Sherrill C. D.; Crawford T. D. Psi4: An Open-source Ab Initio Electronic Structure Program. Wiley Interdiscip. Rev.: Comput. Mol. Sci. 2012, 2, 556–565. 10.1002/wcms.93. [DOI] [Google Scholar]
- Aprá E.; Bylaska E. J.; de Jong W. A.; Govind N.; Kowalski K.; Straatsma T. P.; Valiev M.; van Dam H. J. J.; Alexeev Y.; Anchell J.; Anisimov V.; Aquino F. W.; Atta-Fynn R.; Autschbach J.; Bauman N. P.; Becca J. C.; Bernholdt D. E.; Bhaskaran-Nair K.; Bogatko S.; Borowski P.; Boschen J.; Brabec J.; Bruner A.; Cauët E.; Chen Y.; Chuev G. N.; Cramer C. J.; Daily J.; Deegan M. J. O.; Dunning T. H.; Dupuis M.; Dyall K. G.; Fann G. I.; Fischer S. A.; Fonari A.; Früchtl H.; Gagliardi L.; Garza J.; Gawande N.; Ghosh S.; Glaesemann K.; Götz A. W.; Hammond J.; Helms V.; Hermes E. D.; Hirao K.; Hirata S.; Jacquelin M.; Jensen L.; Johnson B. G.; Jónsson H.; Kendall R. A.; Klemm M.; Kobayashi R.; Konkov V.; Krishnamoorthy S.; Krishnan M.; Lin Z.; Lins R. D.; Littlefield R. J.; Logsdail A. J.; Lopata K.; Ma W.; Marenich A. V.; Martin del Campo J.; Mejia-Rodriguez D.; Moore J. E.; Mullin J. M.; Nakajima T.; Nascimento D. R.; Nichols J. A.; Nichols P. J.; Nieplocha J.; Otero-de-la Roza A.; Palmer B.; Panyala A.; Pirojsirikul T.; Peng B.; Peverati R.; Pittner J.; Pollack L.; Richard R. M.; Sadayappan P.; Schatz G. C.; Shelton W. A.; Silverstein D. W.; Smith D. M. A.; Soares T. A.; Song D.; Swart M.; Taylor H. L.; Thomas G. S.; Tipparaju V.; Truhlar D. G.; Tsemekhman K.; Van Voorhis T.; Vãzquez-Mayagoitia Å.; Verma P.; Villa O.; Vishnu A.; Vogiatzis K. D.; Wang D.; Weare J. H.; Williamson M. J.; Windus T. L.; Woliński K.; Wong A. T.; Wu Q.; Yang C.; Yu Q.; Zacharias M.; Zhang Z.; Zhao Y.; Harrison R. J. NWChem: Past, Present, and Future. J. Chem. Phys. 2020, 152, 184102. 10.1063/5.0004997. [DOI] [PubMed] [Google Scholar]
- Valiev M.; Bylaska E.; Govind N.; Kowalski K.; Straatsma T.; Van Dam H.; Wang D.; Nieplocha J.; Apra E.; Windus T.; de Jong W. NWChem: A Comprehensive and Scalable Open-source Solution for Large Scale Molecular Simulations. Comput. Phys. Commun. 2010, 181, 1477–1489. 10.1016/j.cpc.2010.04.018. [DOI] [Google Scholar]
- Fdez. Galván I.; Vacher M.; Alavi A.; Angeli C.; Aquilante F.; Autschbach J.; Bao J. J.; Bokarev S. I.; Bogdanov N. A.; Carlson R. K.; Chibotaru L. F.; Creutzberg J.; Dattani N.; Delcey M. G.; Dong S. S.; Dreuw A.; Freitag L.; Frutos L. M.; Gagliardi L.; Gendron F.; Giussani A.; Gonzãlez L.; Grell G.; Guo M.; Hoyer C. E.; Johansson M.; Keller S.; Knecht S.; Kovačević G.; Källman E.; Li Manni G.; Lundberg M.; Ma Y.; Mai S.; Malhado J. a. P.; Malmqvist P. Å.; Marquetand P.; Mewes S. A.; Norell J.; Olivucci M.; Oppel M.; Phung Q. M.; Pierloot K.; Plasser F.; Reiher M.; Sand A. M.; Schapiro I.; Sharma P.; Stein C. J.; Sørensen L. K.; Truhlar D. G.; Ugandi M.; Ungur L.; Valentini A.; Vancoillie S.; Veryazov V.; Weser O.; Wesołowski T. A.; Widmark P.-O.; Wouters S.; Zech A.; Zobel J. P.; Lindh R. OpenMolcas: From Source Code to Insight. J. Chem. Theory Comput 2019, 15, 5925–5964. 10.1021/acs.jctc.9b00532. [DOI] [PubMed] [Google Scholar]
- Aquilante F.; Autschbach J.; Baiardi A.; Battaglia S.; Borin V. A.; Chibotaru L. F.; Conti I.; De Vico L.; Delcey M.; Fdez. Galvãn I.; Ferré N.; Freitag L.; Garavelli M.; Gong X.; Knecht S.; Larsson E. D.; Lindh R.; Lundberg M.; Malmqvist P. Å.; Nenov A.; Norell J.; Odelius M.; Olivucci M.; Pedersen T. B.; Pedraza-Gonzãlez L.; Phung Q. M.; Pierloot K.; Reiher M.; Schapiro I.; Segarra-Martí J.; Segatta F.; Seijo L.; Sen S.; Sergentu D.-C.; Stein C. J.; Ungur L.; Vacher M.; Valentini A.; Veryazov V. Modern Quantum Chemistry with [Open] Molcas. J. Chem. Phys. 2020, 152, 214117. 10.1063/5.0004835. [DOI] [PubMed] [Google Scholar]
- Giannozzi P.; Baroni S.; Bonini N.; Calandra M.; Car R.; Cavazzoni C.; Ceresoli D.; Chiarotti G. L.; Cococcioni M.; Dabo I.; Dal Corso A.; de Gironcoli S.; Fabris S.; Fratesi G.; Gebauer R.; Gerstmann U.; Gougoussis C.; Kokalj A.; Lazzeri M.; Martin-Samos L.; Marzari N.; Mauri F.; Mazzarello R.; Paolini S.; Pasquarello A.; Paulatto L.; Sbraccia C.; Scandolo S.; Sclauzero G.; Seitsonen A. P.; Smogunov A.; Umari P.; Wentzcovitch R. M. QUANTUM ESPRESSO: A Modular and Open-source Software Project for Quantum Simulations of Materials. J. Phys.: Condens. Matter 2009, 21, 395502. 10.1088/0953-8984/21/39/395502. [DOI] [PubMed] [Google Scholar]
- Giannozzi P.; Andreussi O.; Brumme T.; Bunau O.; Buongiorno Nardelli M.; Calandra M.; Car R.; Cavazzoni C.; Ceresoli D.; Cococcioni M.; Colonna N.; Carnimeo I.; Dal Corso A.; de Gironcoli S.; Delugas P.; DiStasio R. A.; Ferretti A.; Floris A.; Fratesi G.; Fugallo G.; Gebauer R.; Gerstmann U.; Giustino F.; Gorni T.; Jia J.; Kawamura M.; Ko H.-Y.; Kokalj A.; Küçükbenli E.; Lazzeri M.; Marsili M.; Marzari N.; Mauri F.; Nguyen N. L.; Nguyen H.-V.; Otero-de-la-Roza A.; Paulatto L.; Poncé S.; Rocca D.; Sabatini R.; Santra B.; Schlipf M.; Seitsonen A. P.; Smogunov A.; Timrov I.; Thonhauser T.; Umari P.; Vast N.; Wu X.; Baroni S. Advanced Capabilities for Materials Modelling with Quantum ESPRESSO. J. Phys.: Condens. Matter 2017, 29, 465901. 10.1088/1361-648X/aa8f79. [DOI] [PubMed] [Google Scholar]
- Li P.; Liu X.; Chen M.; Lin P.; Ren X.; Lin L.; Yang C.; He L. Large-scale Ab Initio Simulations Based on Systematically Improvable Atomic Basis. Comput. Mater. Sci. 2016, 112, 503–517. 10.1016/j.commatsci.2015.07.004. [DOI] [Google Scholar]
- Mohr S.; Ratcliff L. E.; Genovese L.; Caliste D.; Boulanger P.; Goedecker S.; Deutsch T. Accurate and Efficient Linear Scaling DFT Calculations with Universal Applicability. Phys. Chem. Chem. Phys. 2015, 17, 31360–31370. 10.1039/C5CP00437C. [DOI] [PubMed] [Google Scholar]
- Ratcliff L. E.; Dawson W.; Fisicaro G.; Caliste D.; Mohr S.; Degomme A.; Videau B.; Cristiglio V.; Stella M.; D’Alessandro M.; Goedecker S.; Nakajima T.; Deutsch T.; Genovese L. Flexibilities of Wavelets As a Computational Basis Set for Large-scale Electronic Structure Calculations. J. Chem. Phys. 2020, 152, 194110. 10.1063/5.0004792. [DOI] [PubMed] [Google Scholar]
- Sun Q.; Berkelbach T. C.; Blunt N. S.; Booth G. H.; Guo S.; Li Z.; Liu J.; McClain J. D.; Sayfutyarova E. R.; Sharma S.; Wouters S.; Chan G. K.-L. PySCF: The Python-based Simulations of Chemistry Framework. Wiley Interdiscip. Rev.: Comput. Mol. Sci. 2018, 8, e1340. 10.1002/wcms.1340. [DOI] [Google Scholar]
- Sun Q.; Zhang X.; Banerjee S.; Bao P.; Barbry M.; Blunt N. S.; Bogdanov N. A.; Booth G. H.; Chen J.; Cui Z.-H.; Eriksen J. J.; Gao Y.; Guo S.; Hermann J.; Hermes M. R.; Koh K.; Koval P.; Lehtola S.; Li Z.; Liu J.; Mardirossian N.; McClain J. D.; Motta M.; Mussard B.; Pham H. Q.; Pulkin A.; Purwanto W.; Robinson P. J.; Ronca E.; Sayfutyarova E. R.; Scheurer M.; Schurkus H. F.; Smith J. E. T.; Sun C.; Sun S.-N.; Upadhyay S.; Wagner L. K.; Wang X.; White A.; Whitfield J. D.; Williamson M. J.; Wouters S.; Yang J.; Yu J. M.; Zhu T.; Berkelbach T. C.; Sharma S.; Sokolov A. Y.; Chan G. K.-L. Recent Developments in the PySCF Program Package. J. Chem. Phys. 2020, 153, 024109. 10.1063/5.0006074. [DOI] [PubMed] [Google Scholar]
- Stewart J. J. P.MOPAC2016; Stewart Computational Chemistry: Colorado Springs, CO, 2016; http://OpenMOPAC.net (accessed 2023-06-21).
- Vincent J. J.; Dixon S. L.; Merz K. M. Jr. Parallel Implementation of a Divide and Conquer Semiempirical Algorithm. Theor. Chem. Acc. 1998, 99, 220–223. 10.1007/s002140050329. [DOI] [Google Scholar]
- Hourahine B.; Aradi B.; Blum V.; Bonafé F.; Buccheri A.; Camacho C.; Cevallos C.; Deshaye M. Y.; Dumitrică T.; Dominguez A.; Ehlert S.; Elstner M.; van der Heide T.; Hermann J.; Irle S.; Kranz J. J.; Köhler C.; Kowalczyk T.; Kubař T.; Lee I. S.; Lutsker V.; Maurer R. J.; Min S. K.; Mitchell I.; Negre C.; Niehaus T. A.; Niklasson A. M. N.; Page A. J.; Pecchia A.; Penazzi G.; Persson M. P.; Řezáč J.; Sãnchez C. G.; Sternberg M.; Stöhr M.; Stuckenberg F.; Tkatchenko A.; Yu V. W.-z.; Frauenheim T. DFTB+, a Software Package for Efficient Approximate Density Functional Theory Based Atomistic Simulations. J. Chem. Phys. 2020, 152, 124101. 10.1063/1.5143190. [DOI] [PubMed] [Google Scholar]
- Gonze X.; Amadon B.; Anglade P.-M.; Beuken J.-M.; Bottin F.; Boulanger P.; Bruneval F.; Caliste D.; Caracas R.; Côté M.; Deutsch T.; Genovese L.; Ghosez P.; Giantomassi M.; Goedecker S.; Hamann D.; Hermet P.; Jollet F.; Jomard G.; Leroux S.; Mancini M.; Mazevet S.; Oliveira M.; Onida G.; Pouillon Y.; Rangel T.; Rignanese G.-M.; Sangalli D.; Shaltaf R.; Torrent M.; Verstraete M.; Zerah G.; Zwanziger J. ABINIT: First-principles Approach to Material and Nanosystem Properties. Comput. Phys. Commun. 2009, 180, 2582–2615. 10.1016/j.cpc.2009.07.007. [DOI] [Google Scholar]
- García A.; Papior N.; Akhtar A.; Artacho E.; Blum V.; Bosoni E.; Brandimarte P.; Brandbyge M.; Cerdã J. I.; Corsetti F.; Cuadrado R.; Dikan V.; Ferrer J.; Gale J.; García-Fernãndez P.; García-Suãrez V. M.; García S.; Huhs G.; Illera S.; Korytãr R.; Koval P.; Lebedeva I.; Lin L.; López-Tarifa P.; Mayo S. G.; Mohr S.; Ordejón P.; Postnikov A.; Pouillon Y.; Pruneda M.; Robles R.; Sãnchez-Portal D.; Soler J. M.; Ullah R.; Yu V. W.-z.; Junquera J. Siesta: Recent Developments and Applications. J. Chem. Phys. 2020, 152, 204108. 10.1063/5.0005077. [DOI] [PubMed] [Google Scholar]
- Kalita B.; Li L.; McCarty R. J.; Burke K. Learning to Approximate Density Functionals. Acc. Chem. Res. 2021, 54, 818–826. 10.1021/acs.accounts.0c00742. [DOI] [PubMed] [Google Scholar]
- Kolb B.; Lentz L. C.; Kolpak A. M. Discovering Charge Density Functionals and Structure-property Relationships with PROPhet: A General Framework for Coupling Machine Learning and First-principles Methods. Sci. Rep. 2017, 7, 1192. 10.1038/s41598-017-01251-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Proppe J.; Gugler S.; Reiher M. Gaussian Process-Based Refinement of Dispersion Corrections. J. Chem. Theory Comput. 2019, 15, 6046–6060. 10.1021/acs.jctc.9b00627. [DOI] [PubMed] [Google Scholar]
- Chen Y.; Zhang L.; Wang H.; E W. Ground State Energy Functional with Hartree–Fock Efficiency and Chemical Accuracy. J. Phys. Chem. A 2020, 124, 7155–7165. 10.1021/acs.jpca.0c03886. [DOI] [PubMed] [Google Scholar]
- Chen Y.; Zhang L.; Wang H.; E W. DeePKS: A Comprehensive Data-driven Approach toward Chemically Accurate Density Functional Theory. J. Chem. Theory Comput. 2021, 17, 170–181. 10.1021/acs.jctc.0c00872. [DOI] [PubMed] [Google Scholar]
- Dick S.; Fernandez-Serra M. Machine Learning Accurate Exchange and Correlation Functionals of the Electronic Density. Nat. Commun. 2020, 11, 3509. 10.1038/s41467-020-17265-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagai R.; Akashi R.; Sugino O. Completing Density Functional Theory by Machine Learning Hidden Messages from Molecules. npj Comput. Mater. 2020, 6, 43. 10.1038/s41524-020-0310-0. [DOI] [Google Scholar]
- Nagai R.; Akashi R.; Sugino O. Machine-learning-based exchange correlation functional with physical asymptotic constraints. Phys. Rev. Res. 2022, 4, 013106. 10.1103/PhysRevResearch.4.013106. [DOI] [Google Scholar]
- Kasim M. F.; Vinko S. M. Learning the Exchange-Correlation Functional from Nature with Fully Differentiable Density Functional Theory. Phys. Rev. Lett. 2021, 127, 126403. 10.1103/PhysRevLett.127.126403. [DOI] [PubMed] [Google Scholar]
- Li L.; Hoyer S.; Pederson R.; Sun R.; Cubuk E. D.; Riley P.; Burke K. Kohn-Sham Equations as Regularizer: Building Prior Knowledge into Machine-Learned Physics. Phys. Rev. Lett. 2021, 126, 036401. 10.1103/PhysRevLett.126.036401. [DOI] [PubMed] [Google Scholar]
- Bystrom K.; Kozinsky B. CIDER: An Expressive, Nonlocal Feature Set for Machine Learning Density Functionals with Exact Constraints. J. Chem. Theory Comput. 2022, 18, 2180–2192. 10.1021/acs.jctc.1c00904. [DOI] [PubMed] [Google Scholar]
- Cuierrier E.; Roy P.-O.; Wang R.; Ernzerhof M. The Fourth-order Expansion of the Exchange Hole and Neural Networks to Construct Exchange–Correlation Functionals. J. Chem. Phys. 2022, 157, 171103. 10.1063/5.0122761. [DOI] [PubMed] [Google Scholar]
- Ma H.; Narayanaswamy A.; Riley P.; Li L. Evolving Symbolic Density Functionals. Sci. Adv. 2022, 8, eabq0279. 10.1126/sciadv.abq0279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y.; Zhang C.; Liu Z.; Truhlar D. G.; Wang Y.; He X. Supervised Learning of a Chemistry Functional with Damped Dispersion. Nat. Comput. Sci. 2023, 3, 48–58. 10.1038/s43588-022-00371-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Y.; Li H.; Pickard F. C.; Narayanan B.; Sen F. G.; Chan M. K. Y.; Sankaranarayanan S. K. R. S.; Brooks B. R.; Roux B. Machine Learning Force Field Parameters from Ab Initio Data. J. Chem. Theory Comput. 2017, 13, 4492–4503. 10.1021/acs.jctc.7b00521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Korolev V. V.; Nevolin Y. M.; Manz T. A.; Protsenko P. V. Parametrization of Nonbonded Force Field Terms for Metal–Organic Frameworks Using Machine Learning Approach. J. Chem. Inf. Model. 2021, 61, 5774–5784. 10.1021/acs.jcim.1c01124. [DOI] [PubMed] [Google Scholar]
- Befort B. J.; DeFever R. S.; Tow G. M.; Dowling A. W.; Maginn E. J. Machine Learning Directed Optimization of Classical Molecular Modeling Force Fields. J. Chem. Inf. Model. 2021, 61, 4400–4414. 10.1021/acs.jcim.1c00448. [DOI] [PubMed] [Google Scholar]
- Müller M.; Hagg A.; Strickstrock R.; Hülsmann M.; Asteroth A.; Kirschner K. N.; Reith D. Determining Lennard-Jones Parameters Using Multiscale Target Data through Presampling-Enhanced, Surrogate-Assisted Global Optimization. J. Chem. Inf. Model. 2023, 63, 1872–1881. 10.1021/acs.jcim.2c01231. [DOI] [PubMed] [Google Scholar]
- Thürlemann M.; Böselt L.; Riniker S. Regularized by Physics: Graph Neural Network Parametrized Potentials for the Description of Intermolecular Interactions. J. Chem. Theory Comput. 2023, 19, 562–579. 10.1021/acs.jctc.2c00661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y.; Fass J.; Kaminow B.; Herr J. E.; Rufa D.; Zhang I.; Pulido I.; Henry M.; Bruce Macdonald H. E.; Takaba K.; Chodera J. D. End-to-end Differentiable Construction of Molecular Mechanics Force Fields. Chem. Sci. 2022, 13, 12016–12033. 10.1039/D2SC02739A. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang J.; Cao D.; Tang C.; Chen X.; Sun H.; Hou T. Fast and Accurate Prediction of Partial Charges Using Atom-path-descriptor-based Machine Learning. Bioinformatics 2020, 36, 4721–4728. 10.1093/bioinformatics/btaa566. [DOI] [PubMed] [Google Scholar]
- Gallegos M.; Guevara-Vela J. M.; Pendás A. M. NNAIMQ: A Neural Network Model for Predicting QTAIM Charges. J. Chem. Phys. 2022, 156, 014112. 10.1063/5.0076896. [DOI] [PubMed] [Google Scholar]
- Jiang D.; Sun H.; Wang J.; Hsieh C.-Y.; Li Y.; Wu Z.; Cao D.; Wu J.; Hou T. Out-of-the-box Deep Learning Prediction of Quantum-mechanical Partial Charges by Graph Representation and Transfer Learning. Briefings Bioinf. 2022, 23, bbab597. 10.1093/bib/bbab597. [DOI] [PubMed] [Google Scholar]
- Raza A.; Sturluson A.; Simon C. M.; Fern X. Message Passing Neural Networks for Partial Charge Assignment to Metal–Organic Frameworks. J. Phys. Chem. C 2020, 124, 19070–19082. 10.1021/acs.jpcc.0c04903. [DOI] [Google Scholar]
- Kancharlapalli S.; Gopalan A.; Haranczyk M.; Snurr R. Q. Fast and Accurate Machine Learning Strategy for Calculating Partial Atomic Charges in Metal–Organic Frameworks. J. Chem. Theory Comput. 2021, 17, 3052–3064. 10.1021/acs.jctc.0c01229. [DOI] [PubMed] [Google Scholar]
- Unke O. T.; Meuwly M. PhysNet: A Neural Network for Predicting Energies, Forces, Dipole Moments, and Partial Charges. J. Chem. Theory Comput. 2019, 15, 3678–3693. 10.1021/acs.jctc.9b00181. [DOI] [PubMed] [Google Scholar]
- Metcalf D. P.; Jiang A.; Spronk S. A.; Cheney D. L.; Sherrill C. D. Electron-Passing Neural Networks for Atomic Charge Prediction in Systems with Arbitrary Molecular Charge. J. Chem. Inf. Model. 2021, 61, 115–122. 10.1021/acs.jcim.0c01071. [DOI] [PubMed] [Google Scholar]
- Kumar A.; Pandey P.; Chatterjee P.; MacKerell A. D. J. Deep Neural Network Model to Predict the Electrostatic Parameters in the Polarizable Classical Drude Oscillator Force Field. J. Chem. Theory Comput. 2022, 18, 1711–1725. 10.1021/acs.jctc.1c01166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rathi P. C.; Ludlow R. F.; Verdonk M. L. Practical High-Quality Electrostatic Potential Surfaces for Drug Discovery Using a Graph-Convolutional Deep Neural Network. J. Med. Chem. 2020, 63, 8778–8790. 10.1021/acs.jmedchem.9b01129. [DOI] [PubMed] [Google Scholar]
- Thürlemann M.; Böselt L.; Riniker S. Learning Atomic Multipoles: Prediction of the Electrostatic Potential with Equivariant Graph Neural Networks. J. Chem. Theory Comput. 2022, 18, 1701–1710. 10.1021/acs.jctc.1c01021. [DOI] [PubMed] [Google Scholar]
- Bolcato G.; Heid E.; Boström J. On the Value of Using 3D Shape and Electrostatic Similarities in Deep Generative Methods. J. Chem. Inf. Model. 2022, 62, 1388–1398. 10.1021/acs.jcim.1c01535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Case D.; Aktulga H.; Belfon K.; Ben-Shalom I.; Brozell S.; Cerutti D.; Cheatham T. E. III; Cruzeiro V.; Darden T.; Duke R.; Giambasu G.; Gilson M.; Gohlke H.; Goetz A.; Harris R.; Izadi S.; Izmailov S.; Jin C.; Kasavajhala K.; Kaymak M.; King E.; Kovalenko A.; Kurtzman T.; Lee T.; LeGrand S.; Li P.; Lin C.; Liu J.; Luchko T.; Luo R.; Machado M.; Man V.; Manathunga M.; Merz K. M.; Miao Y.; Mikhailovskii O.; Monard G.; Nguyen C.; Nguyen H.; O’Hearn K.; Onufriev A.; Pan F.; Pantano S.; Qi R.; Rahnamoun A.; Roe D.; Roitberg A.; Sagui C.; Schott-Verdugo S.; Shen J.; Simmerling C.; Skrynnikov N.; Smith J.; Swails J.; Walker R.; Wang J.; Wei H.; Wolf R.; Wu X.; Xue Y.; York D.; Zhao S.; Kollman P.. AMBER20 and AmberTools21; University of California, San Francisco, 2021; http://ambermd.org.
- Kühne T. D.; Iannuzzi M.; Del Ben M.; Rybkin V. V.; Seewald P.; Stein F.; Laino T.; Khaliullin R. Z.; Schütt O.; Schiffmann F.; Golze D.; Wilhelm J.; Chulkov S.; Bani-Hashemian M. H.; Weber V.; Borštnik U.; Taillefumier M.; Jakobovits A. S.; Lazzaro A.; Pabst H.; Müller T.; Schade R.; Guidon M.; Andermatt S.; Holmberg N.; Schenter G. K.; Hehn A.; Bussy A.; Belleflamme F.; Tabacchi G.; Glöß A.; Lass M.; Bethune I.; Mundy C. J.; Plessl C.; Watkins M.; VandeVondele J.; Krack M.; Hutter J. CP2K: An Electronic Structure and Molecular Dynamics Software Package - Quickstep: Efficient and Accurate Electronic Structure Calculations. J. Chem. Phys. 2020, 152, 194103. 10.1063/5.0007045. [DOI] [PubMed] [Google Scholar]
- GROMACS Development Team . GROMACS. https://www.gromacs.org/ (accessed 2023-06-21).
- Thompson A. P.; Aktulga H. M.; Berger R.; Bolintineanu D. S.; Brown W. M.; Crozier P. S.; in ’t Veld P. J.; Kohlmeyer A.; Moore S. G.; Nguyen T. D.; Shan R.; Stevens M. J.; Tranchida J.; Trott C.; Plimpton S. J. LAMMPS - a Flexible Simulation Tool for Particle-based Materials Modeling at the Atomic, Meso, and Continuum Scales. Comput. Phys. Commun. 2022, 271, 108171. 10.1016/j.cpc.2021.108171. [DOI] [Google Scholar]
- Eastman P.; Swails J.; Chodera J. D.; McGibbon R. T.; Zhao Y.; Beauchamp K. A.; Wang L.-P.; Simmonett A. C.; Harrigan M. P.; Stern C. D.; Wiewiora R. P.; Brooks B. R.; Pande V. S. OpenMM 7: Rapid Development of High Performance Algorithms for Molecular Dynamics. PLoS Comput. Biol. 2017, 13, e1005659. 10.1371/journal.pcbi.1005659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Procacci P. Hybrid MPI/OpenMP Implementation of the ORAC Molecular Dynamics Program for Generalized Ensemble and Fast Switching Alchemical Simulations. J. Chem. Inf. Model. 2016, 56, 1117–1121. 10.1021/acs.jcim.6b00151. [DOI] [PubMed] [Google Scholar]
- Doerr S.; Majewski M.; Pérez A.; Krämer A.; Clementi C.; Noe F.; Giorgino T.; De Fabritiis G. TorchMD: A Deep Learning Framework for Molecular Simulations. J. Chem. Theory Comput. 2021, 17, 2355–2363. 10.1021/acs.jctc.0c01343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tribello G. A.; Bonomi M.; Branduardi D.; Camilloni C.; Bussi G. PLUMED 2: New Feathers for an Old Bird. Comput. Phys. Commun. 2014, 185, 604–613. 10.1016/j.cpc.2013.09.018. [DOI] [Google Scholar]
- Hernández C. X.; Wayment-Steele H. K.; Sultan M. M.; Husic B. E.; Pande V. S. Variational Encoding of Complex Dynamics. Phys. Rev. E 2018, 97, 062412. 10.1103/PhysRevE.97.062412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ribeiro J. M. L.; Bravo P.; Wang Y.; Tiwary P. Reweighted Autoencoded Variational Bayes for Enhanced Sampling (RAVE). J. Chem. Phys. 2018, 149, 072301. 10.1063/1.5025487. [DOI] [PubMed] [Google Scholar]
- Vlachas P. R.; Arampatzis G.; Uhler C.; Koumoutsakos P. Multiscale Simulations of Complex Systems by Learning Their Effective Dynamics. Nat. Mach. Intell. 2022, 4, 359–366. 10.1038/s42256-022-00464-w. [DOI] [Google Scholar]
- Wu H.; Mardt A.; Pasquali L.; Noe F.. Deep Generative Markov State Models. In Proceedings of the 32nd International Conference on Neural Information Processing Systems (NIPS 2018); Curran Associates, 2018; pp 3979–3988.
- Do H. N.; Wang J.; Bhattarai A.; Miao Y. GLOW: A Workflow Integrating Gaussian-Accelerated Molecular Dynamics and Deep Learning for Free Energy Profiling. J. Chem. Theory Comput. 2022, 18, 1423–1436. 10.1021/acs.jctc.1c01055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bonati L.; Piccini G.; Parrinello M. Deep Learning the Slow Modes for Rare Events Sampling. Proc. Natl. Acad. Sci. U.S.A. 2021, 118, e2113533118. 10.1073/pnas.2113533118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen W.; Ferguson A. L. Molecular Enhanced Sampling with Autoencoders: On-the-fly Collective Variable Discovery and Accelerated Free Energy Landscape Exploration. J. Comput. Chem. 2018, 39, 2079–2102. 10.1002/jcc.25520. [DOI] [PubMed] [Google Scholar]
- Hooft F.; Pérez de Alba Ortíz A.; Ensing B. Discovering Collective Variables of Molecular Transitions via Genetic Algorithms and Neural Networks. J. Chem. Theory Comput. 2021, 17, 2294–2306. 10.1021/acs.jctc.0c00981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baima J.; Goryaeva A. M.; Swinburne T. D.; Maillet J.-B.; Nastar M.; Marinica M.-C. Capabilities and Limits of Autoencoders for Extracting Collective Variables in Atomistic Materials Science. Phys. Chem. Chem. Phys. 2022, 24, 23152–23163. 10.1039/D2CP01917E. [DOI] [PubMed] [Google Scholar]
- Ketkaew R.; Creazzo F.; Luber S. Machine Learning-Assisted Discovery of Hidden States in Expanded Free Energy Space. J. Phys. Chem. Lett. 2022, 13, 1797–1805. 10.1021/acs.jpclett.1c04004. [DOI] [PubMed] [Google Scholar]
- Ketkaew R.; Luber S. DeepCV: A Deep Learning Framework for Blind Search of Collective Variables in Expanded Configurational Space. J. Chem. Inf. Model. 2022, 62, 6352–6364. 10.1021/acs.jcim.2c00883. [DOI] [PubMed] [Google Scholar]
- Schwalbe-Koda D.; Tan A. R.; Gómez-Bombarelli R. Differentiable Sampling of Molecular Geometries with Uncertainty-based Adversarial Attacks. Nat. Commun. 2021, 12, 5104. 10.1038/s41467-021-25342-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Michaud-Agrawal N.; Denning E. J.; Woolf T. B.; Beckstein O. MDAnalysis: A Toolkit for the Analysis of Molecular Dynamics Simulations. J. Comput. Chem. 2011, 32, 2319–2327. 10.1002/jcc.21787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gowers R. J.; Linke M.; Barnoud J.; Reddy T. J. E.; Melo M. N.; Seyler S. L.; Domański J.; Dotson D. L.; Buchoux S.; Kenney I. M.; Oliver B.. MDAnalysis: A Python Package for the Rapid Analysis of Molecular Dynamics Simulations. In Proceedings of the 15th Python in Science Conference, 2016; pp 98–105.
- Lemke T.; Peter C. EncoderMap: Dimensionality Reduction and Generation of Molecule Conformations. J. Chem. Theory Comput. 2019, 15, 1209–1215. 10.1021/acs.jctc.8b00975. [DOI] [PubMed] [Google Scholar]
- Lemke T.; Berg A.; Jain A.; Peter C. EncoderMap(II): Visualizing Important Molecular Motions with Improved Generation of Protein Conformations. J. Chem. Inf. Model. 2019, 59, 4550–4560. 10.1021/acs.jcim.9b00675. [DOI] [PubMed] [Google Scholar]
- Jin Y.; Johannissen L. O.; Hay S. Predicting New Protein Conformations from Molecular Dynamics Simulation Conformational Landscapes and Machine Learning. Proteins: Struct., Funct., Bioinf. 2021, 89, 915–921. 10.1002/prot.26068. [DOI] [PubMed] [Google Scholar]
- Shen C.; Zhang X.; Deng Y.; Gao J.; Wang D.; Xu L.; Pan P.; Hou T.; Kang Y. Boosting Protein–Ligand Binding Pose Prediction and Virtual Screening Based on Residue–Atom Distance Likelihood Potential and Graph Transformer. J. Med. Chem. 2022, 65, 10691–10706. 10.1021/acs.jmedchem.2c00991. [DOI] [PubMed] [Google Scholar]
- Bozkurt Varolgüneş Y.; Bereau T.; Rudzinski J. F. Interpretable Embeddings from Molecular Simulations Using Gaussian Mixture Variational Autoencoders. Mach. Learn.: Sci. Technol. 2020, 1, 015012. 10.1088/2632-2153/ab80b7. [DOI] [Google Scholar]
- Novelli P.; Bonati L.; Pontil M.; Parrinello M. Characterizing Metastable States with the Help of Machine Learning. J. Chem. Theory Comput. 2022, 18, 5195–5202. 10.1021/acs.jctc.2c00393. [DOI] [PubMed] [Google Scholar]
- Ward M. D.; Zimmerman M. I.; Meller A.; Chung M.; Swamidass S. J.; Bowman G. R. Deep Learning the Structural Determinants of Protein Biochemical Properties by Comparing Structural Ensembles with Diffnets. Nat. Commun. 2021, 12, 3023. 10.1038/s41467-021-23246-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramaswamy V. K.; Musson S. C.; Willcocks C. G.; Degiacomi M. T. Deep Learning Protein Conformational Space with Convolutions and Latent Interpolations. Phys. Rev. X 2021, 11, 011052. 10.1103/PhysRevX.11.011052. [DOI] [Google Scholar]
- Wang D.; Tiwary P. State Predictive Information Bottleneck. J. Chem. Phys. 2021, 154, 134111. 10.1063/5.0038198. [DOI] [PubMed] [Google Scholar]
- Li C.; Liu J.; Chen J.; Yuan Y.; Yu J.; Gou Q.; Guo Y.; Pu X. An Interpretable Convolutional Neural Network Framework for Analyzing Molecular Dynamics Trajectories: a Case Study on Functional States for G-Protein-Coupled Receptors. J. Chem. Inf. Model. 2022, 62, 1399–1410. 10.1021/acs.jcim.2c00085. [DOI] [PubMed] [Google Scholar]
- Harrigan M. P.; Sultan M. M.; Hernãndez C. X.; Husic B. E.; Eastman P.; Schwantes C. R.; Beauchamp K. A.; McGibbon R. T.; Pande V. S. MSMBuilder: Statistical Models for Biomolecular Dynamics. Biophys. J. 2017, 112, 10–15. 10.1016/j.bpj.2016.10.042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trott O.; Olson A. J. AutoDock Vina: Improving the Speed and Accuracy of Docking with a New Scoring Function, Efficient Optimization, and Multithreading. J. Comput. Chem. 2009, 31, 455–461. 10.1002/jcc.21334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paul D. S.; Gautham N. MOLS 2.0: Software Package for Peptide Modeling and Protein-Ligand Docking. J. Mol. Model. 2016, 22, 239. 10.1007/s00894-016-3106-x. [DOI] [PubMed] [Google Scholar]
- Ruiz-Carmona S.; Alvarez-Garcia D.; Foloppe N.; Garmendia-Doval A. B.; Juhos S.; Schmidtke P.; Barril X.; Hubbard R. E.; Morley S. D. rDock: A Fast, Versatile and Open Source Program for Docking Ligands to Proteins and Nucleic Acids. PLoS Comput. Biol. 2014, 10, e1003571. 10.1371/journal.pcbi.1003571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ballester P. J.; Mitchell J. B. O. A Machine Learning Approach to Predicting Protein–ligand Binding Affinity with Applications to Molecular Docking. Bioinformatics 2010, 26, 1169–1175. 10.1093/bioinformatics/btq112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koes D. R.; Baumgartner M. P.; Camacho C. J. Lessons Learned in Empirical Scoring with smina from the CSAR 2011 Benchmarking Exercise. J. Chem. Inf. Model. 2013, 53, 1893–1904. 10.1021/ci300604z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ragoza M.; Hochuli J.; Idrobo E.; Sunseri J.; Koes D. R. Protein–Ligand Scoring with Convolutional Neural Networks. J. Chem. Inf. Model. 2017, 57, 942–957. 10.1021/acs.jcim.6b00740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McNutt A. T.; Francoeur P.; Aggarwal R.; Masuda T.; Meli R.; Ragoza M.; Sunseri J.; Koes D. R. GNINA 1.0: Molecular Docking with Deep Learning. J. Cheminf. 2021, 13, 43. 10.1186/s13321-021-00522-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- García-Ortegón M.; Simm G. N. C.; Tripp A. J.; Hernãndez-Lobato J. M.; Bender A.; Bacallado S. DOCKSTRING: Easy Molecular Docking Yields Better Benchmarks for Ligand Design. J. Chem. Inf. Model. 2022, 62, 3486–3502. 10.1021/acs.jcim.1c01334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kurcinski M.; Pawel Ciemny M.; Oleniecki T.; Kuriata A.; Badaczewska-Dawid A. E.; Kolinski A.; Kmiecik S. CABS-dock Standalone: A Toolbox for Flexible Protein-peptide Docking. Bioinformatics 2019, 35, 4170–4172. 10.1093/bioinformatics/btz185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiménez-García B.; Roel-Touris J.; Romero-Durana M.; Vidal M.; Jiménez-Gonzãlez D.; Fernãndez-Recio J. LightDock: A New Multi-scale Approach to Protein-Protein Docking. Bioinformatics 2018, 34, 49–55. 10.1093/bioinformatics/btx555. [DOI] [PubMed] [Google Scholar]
- Roel-Touris J.; Bonvin A. M. J. J.; Jiménez-García B. Lightdock Goes Information-driven. Bioinformatics 2020, 36, 950–952. 10.1093/bioinformatics/btz642. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Masters M. R.; Mahmoud A. H.; Wei Y.; Lill M. A. Deep Learning Model for Efficient Protein-Ligand Docking with Implicit Side-Chain Flexibility. J. Chem. Inf. Model. 2023, 63, 1695–1707. 10.1021/acs.jcim.2c01436. [DOI] [PubMed] [Google Scholar]
- Wójcikowski M.; Zielenkiewicz P.; Siedlecki P. Open Drug Discovery Toolkit (ODDT): A New Open-source Player in the Drug Discovery Field. J. Cheminf. 2015, 7, 26. 10.1186/s13321-015-0078-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Durrant J. D.; McCammon J. A. NNScore 2.0: A Neural-Network Receptor–Ligand Scoring Function. J. Chem. Inf. Model. 2011, 51, 2897–2903. 10.1021/ci2003889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shen C.; Hu Y.; Wang Z.; Zhang X.; Zhong H.; Wang G.; Yao X.; Xu L.; Cao D.; Hou T. Can Machine Learning Consistently Improve the Scoring Power of Classical Scoring Functions? Insights into the Role of Machine Learning in Scoring Functions. Briefings Bioinf. 2021, 22, 497–514. 10.1093/bib/bbz173. [DOI] [PubMed] [Google Scholar]
- Wójcikowski M.; Ballester P. J.; Siedlecki P. Performance of Machine-learning Scoring Functions in Structure-based Virtual Screening. Sci. Rep. 2017, 7, 46710. 10.1038/srep46710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu J.; Hou X.; Wang C.; Zhang Y. Incorporating Explicit Water Molecules and Ligand Conformation Stability in Machine-Learning Scoring Functions. J. Chem. Inf. Model. 2019, 59, 4540–4549. 10.1021/acs.jcim.9b00645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meli R.; Anighoro A.; Bodkin M. J.; Morris G. M.; Biggin P. C. Learning Protein-Ligand Binding Affinity with Atomic Environment Vectors. J. Cheminf. 2021, 13, 59. 10.1186/s13321-021-00536-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rayka M.; Karimi-Jafari M. H.; Firouzi R. ET-score: Improving Protein-ligand Binding Affinity Prediction Based on Distance-weighted Interatomic Contact Features Using Extremely Randomized Trees Algorithm. Mol. Inform. 2021, 40, 2060084. 10.1002/minf.202060084. [DOI] [PubMed] [Google Scholar]
- Yang C.; Zhang Y. Delta Machine Learning to Improve Scoring-Ranking-Screening Performances of Protein–Ligand Scoring Functions. J. Chem. Inf. Model. 2022, 62, 2696–2712. 10.1021/acs.jcim.2c00485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kumar S. P.; Dixit N. Y.; Patel C. N.; Rawal R. M.; Pandya H. A. PharmRF: A Machine-learning Scoring Function to Identify the Best Protein-ligand Complexes for Structure-based Pharmacophore Screening with High Enrichments. J. Comput. Chem. 2022, 43, 847–863. 10.1002/jcc.26840. [DOI] [PubMed] [Google Scholar]
- Dong L.; Qu X.; Wang B. XLPFE: A Simple and Effective Machine Learning Scoring Function for Protein–Ligand Scoring and Ranking. ACS Omega 2022, 7, 21727–21735. 10.1021/acsomega.2c01723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Z.; Zheng L.; Wang S.; Lin M.; Wang Z.; Kong A. W.-K.; Mu Y.; Wei Y.; Li W. A Fully Differentiable Ligand Pose Optimization Framework Guided by Deep Learning and a Traditional Scoring Function. Briefings Bioinf. 2023, 24, bbac520. 10.1093/bib/bbac520. [DOI] [PubMed] [Google Scholar]
- Rayka M.; Firouzi R. GB-score: Minimally designed machine learning scoring function based on distance-weighted interatomic contact features. Mol. Inf. 2023, 42, 2200135. 10.1002/minf.202200135. [DOI] [PubMed] [Google Scholar]
- Stefaniak F.; Bujnicki J. M. AnnapuRNA: A Scoring Function for Predicting RNA-small Molecule Binding Poses. PLoS Comput. Biol. 2021, 17, e1008309. 10.1371/journal.pcbi.1008309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiménez J.; Škalič M.; Martínez-Rosell G.; De Fabritiis G. KDEEP: Protein–Ligand Absolute Binding Affinity Prediction via 3D-Convolutional Neural Networks. J. Chem. Inf. Model. 2018, 58, 287–296. 10.1021/acs.jcim.7b00650. [DOI] [PubMed] [Google Scholar]
- Bao J.; He X.; Zhang J. Z. H. DeepBSP—a Machine Learning Method for Accurate Prediction of Protein–Ligand Docking Structures. J. Chem. Inf. Model. 2021, 61, 2231–2240. 10.1021/acs.jcim.1c00334. [DOI] [PubMed] [Google Scholar]
- Simončič M.; Lukšič M.; Druchok M. Machine Learning Assessment of the Binding Region as a Tool for More Efficient Computational Receptor-Ligand Docking. J. Mol. Liq. 2022, 353, 118759. 10.1016/j.molliq.2022.118759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Öztürk H.; Özgür A.; Ozkirimli E. DeepDTA: Deep Drug-Target Binding Affinity Prediction. Bioinformatics 2018, 34, i821–i829. 10.1093/bioinformatics/bty593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee I.; Keum J.; Nam H. DeepConv-DTI: Prediction of Drug–Target Interactions Via Deep Learning with Convolution on Protein Sequences. PLoS Comput. Biol. 2019, 15, e1007129. 10.1371/journal.pcbi.1007129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Abbasi K.; Razzaghi P.; Poso A.; Amanlou M.; Ghasemi J. B.; Masoudi-Nejad A. DeepCDA: Deep Cross-domain Compound–Protein Affinity Prediction through LSTM and Convolutional Neural Networks. Bioinformatics 2020, 36, 4633–4642. 10.1093/bioinformatics/btaa544. [DOI] [PubMed] [Google Scholar]
- Rifaioglu A. S.; Nalbat E.; Atalay V.; Martin M. J.; Cetin-Atalay R.; Doğan T. DEEPScreen: High Performance Drug–target Interaction Prediction with Convolutional Neural Networks Using 2-D Structural Compound Representations. Chem. Sci. 2020, 11, 2531–2557. 10.1039/C9SC03414E. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang D.; Hsieh C.-Y.; Wu Z.; Kang Y.; Wang J.; Wang E.; Liao B.; Shen C.; Xu L.; Wu J.; Cao D.; Hou T. InteractionGraphNet: A Novel and Efficient Deep Graph Representation Learning Framework for Accurate Protein–Ligand Interaction Predictions. J. Med. Chem. 2021, 64, 18209–18232. 10.1021/acs.jmedchem.1c01830. [DOI] [PubMed] [Google Scholar]
- Ricci-Lopez J.; Aguila S. A.; Gilson M. K.; Brizuela C. A. Improving Structure-Based Virtual Screening with Ensemble Docking and Machine Learning. J. Chem. Inf. Model. 2021, 61, 5362–5376. 10.1021/acs.jcim.1c00511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lim J.; Ryu S.; Park K.; Choe Y. J.; Ham J.; Kim W. Y. Predicting Drug–Target Interaction Using a Novel Graph Neural Network with 3D Structure-Embedded Graph Representation. J. Chem. Inf. Model. 2019, 59, 3981–3988. 10.1021/acs.jcim.9b00387. [DOI] [PubMed] [Google Scholar]
- Tran H. N. T.; Thomas J. J.; Ahamed Hassain Malim N. H. DeepNC: A Framework for Drug-target Interaction Prediction with Graph Neural Networks. PeerJ 2022, 10, e13163. 10.7717/peerj.13163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Verma N.; Qu X.; Trozzi F.; Elsaied M.; Karki N.; Tao Y.; Zoltowski B.; Larson E. C.; Kraka E. SSnet: A. Deep Learning Approach for Protein-Ligand Interaction Prediction. Int. J. Mol. Sci. 2021, 22, 1392. 10.3390/ijms22031392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang P.; Zheng S.; Jiang Y.; Li C.; Liu J.; Wen C.; Patronov A.; Qian D.; Chen H.; Yang Y. Structure-Aware Multimodal Deep Learning for Drug–Protein Interaction Prediction. J. Chem. Inf. Model. 2022, 62, 1308–1317. 10.1021/acs.jcim.2c00060. [DOI] [PubMed] [Google Scholar]
- Bouysset C.; Fiorucci S. ProLIF: A Library to Encode Molecular Interactions As Fingerprints. J. Cheminf. 2021, 13, 72. 10.1186/s13321-021-00548-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fassio A. V.; Shub L.; Ponzoni L.; McKinley J.; O’Meara M. J.; Ferreira R. S.; Keiser M. J.; de Melo Minardi R. C. Prioritizing Virtual Screening with Interpretable Interaction Fingerprints. J. Chem. Inf. Model. 2022, 62, 4300–4318. 10.1021/acs.jcim.2c00695. [DOI] [PubMed] [Google Scholar]
- Stojanović L.; Popović M.; Tijanić N.; Rakočević G.; Kalinić M. Improved Scaffold Hopping in Ligand-Based Virtual Screening Using Neural Representation Learning. J. Chem. Inf. Model. 2020, 60, 4629–4639. 10.1021/acs.jcim.0c00622. [DOI] [PubMed] [Google Scholar]
- Amendola G.; Cosconati S. PyRMD: A New Fully Automated AI-Powered Ligand-Based Virtual Screening Tool. J. Chem. Inf. Model. 2021, 61, 3835–3845. 10.1021/acs.jcim.1c00653. [DOI] [PubMed] [Google Scholar]
- Yin Y.; Hu H.; Yang Z.; Xu H.; Wu J. RealVS: Toward Enhancing the Precision of Top Hits in Ligand-Based Virtual Screening of Drug Leads from Large Compound Databases. J. Chem. Inf. Model. 2021, 61, 4924–4939. 10.1021/acs.jcim.1c01021. [DOI] [PubMed] [Google Scholar]
- Imrie F.; Bradley A. R.; Deane C. M. Generating Property-matched Decoy Molecules Using Deep Learning. Bioinformatics 2021, 37, 2134–2141. 10.1093/bioinformatics/btab080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang X.; Shen C.; Liao B.; Jiang D.; Wang J.; Wu Z.; Du H.; Wang T.; Huo W.; Xu L.; Cao D.; Hsieh C.-Y.; Hou T. TocoDecoy: A New Approach to Design Unbiased Datasets for Training and Benchmarking Machine-Learning Scoring Functions. J. Med. Chem. 2022, 65, 7918–7932. 10.1021/acs.jmedchem.2c00460. [DOI] [PubMed] [Google Scholar]
- Krivák R.; Hoksza D. Improving Protein-Ligand Binding Site Prediction Accuracy by Classification of Inner Pocket Points Using Local Features. J. Cheminf. 2015, 7, 12. 10.1186/s13321-015-0059-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krivák R.; Hoksza D. P2Rank: Machine Learning Based Tool for Rapid and Accurate Prediction of Ligand Binding Sites from Protein Structure. J. Cheminf. 2018, 10, 39. 10.1186/s13321-018-0285-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stepniewska-Dziubinska M. M.; Zielenkiewicz P.; Siedlecki P. Improving Detection of Protein-ligand Binding Sites with 3D Segmentation. Sci. Rep. 2020, 10, 5035. 10.1038/s41598-020-61860-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mylonas S. K.; Axenopoulos A.; Daras P. DeepSurf: A Surface-based Deep Learning Approach for the Prediction of Ligand Binding Sites on Proteins. Bioinformatics 2021, 37, 1681–1690. 10.1093/bioinformatics/btab009. [DOI] [PubMed] [Google Scholar]
- Kandel J.; Tayara H.; Chong K. T. PUResNet: Prediction of Protein–Ligand Binding Sites Using Deep Residual Neural Network. J. Cheminf. 2021, 13, 65. 10.1186/s13321-021-00547-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Le Guilloux V.; Schmidtke P.; Tuffery P. Fpocket: An Open Source Platform for Ligand Pocket Detection. BMC Bioinf. 2009, 10, 168. 10.1186/1471-2105-10-168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aggarwal R.; Gupta A.; Chelur V.; Jawahar C. V.; Priyakumar U. D. DeepPocket: Ligand Binding Site Detection and Segmentation using 3D Convolutional Neural Networks. J. Chem. Inf. Model. 2022, 62, 5069–5079. 10.1021/acs.jcim.1c00799. [DOI] [PubMed] [Google Scholar]
- Mallet V.; Checa Ruano L.; Moine Franel A.; Nilges M.; Druart K.; Bouvier G.; Sperandio O. InDeep: 3D Fully Convolutional Neural Networks to Assist In Silico Drug Design on Protein–Protein Interactions. Bioinformatics 2022, 38, 1261–1268. 10.1093/bioinformatics/btab849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chelur V. R.; Priyakumar U. D. BiRDS - Binding Residue Detection from Protein Sequences Using Deep ResNets. J. Chem. Inf. Model. 2022, 62, 1809–1818. 10.1021/acs.jcim.1c00972. [DOI] [PubMed] [Google Scholar]
- Yan X.; Lu Y.; Li Z.; Wei Q.; Gao X.; Wang S.; Wu S.; Cui S. PointSite: A Point Cloud Segmentation Tool for Identification of Protein Ligand Binding Atoms. J. Chem. Inf. Model. 2022, 62, 2835–2845. 10.1021/acs.jcim.1c01512. [DOI] [PubMed] [Google Scholar]
- Liu S.; Liu C.; Deng L. Machine Learning Approaches for Protein–Protein Interaction Hot Spot Prediction: Progress and Comparative Assessment. Molecules 2018, 23, 2535. 10.3390/molecules23102535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fout A.; Byrd J.; Shariat B.; Ben-Hur A.. Protein Interface Prediction using Graph Convolutional Networks. In Proceedings of the 31st International Conference on Neural Information Processing Systems (NIPS 2017); Curran Associates, 2017; pp 6533–6542.
- Gainza P.; Sverrisson F.; Monti F.; Rodolá E.; Boscaini D.; Bronstein M. M.; Correia B. E. Deciphering Interaction Fingerprints from Protein Molecular Surfaces Using Geometric Deep Learning. Nat. Methods 2020, 17, 184–192. 10.1038/s41592-019-0666-6. [DOI] [PubMed] [Google Scholar]
- Yuan Q.; Chen J.; Zhao H.; Zhou Y.; Yang Y. Structure-aware Protein-protein Interaction Site Prediction Using Deep Graph Convolutional Network. Bioinformatics 2021, 38, 125–132. 10.1093/bioinformatics/btab643. [DOI] [PubMed] [Google Scholar]
- Baranwal M.; Magner A.; Saldinger J.; Turali-Emre E. S.; Elvati P.; Kozarekar S.; VanEpps J. S.; Kotov N. A.; Violi A.; Hero A. O. Struct2Graph: A Graph Attention Network for Structure Based Predictions of Protein-protein Interactions. BMC Bioinf. 2022, 23, 370. 10.1186/s12859-022-04910-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin P.; Yan Y.; Huang S.-Y. DeepHomo2.0: Improved Protein–Protein Contact Prediction of Homodimers by Transformer-enhanced Deep Learning. Briefings Bioinf. 2023, 24, bbac499. 10.1093/bib/bbac499. [DOI] [PubMed] [Google Scholar]
- Wang X.; Terashi G.; Christoffer C. W.; Zhu M.; Kihara D. Protein Docking Model Evaluation by 3D Deep Convolutional Neural Networks. Bioinformatics 2020, 36, 2113–2118. 10.1093/bioinformatics/btz870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geng C.; Jung Y.; Renaud N.; Honavar V.; Bonvin A. M. J. J.; Xue L. C. iScore: A Novel Graph Kernel-based Function for Scoring Protein-protein Docking Models. Bioinformatics 2020, 36, 112–121. 10.1093/bioinformatics/btz496. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Renaud N.; Jung Y.; Honavar V.; Geng C.; Bonvin A. M. J. J.; Xue L. C. iScore: An MPI Supported Software for Ranking Protein–Protein Docking Models Based on a Random Walk Graph Kernel and Support Vector Machines. SoftwareX 2020, 11, 100462. 10.1016/j.softx.2020.100462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Renaud N.; Geng C.; Georgievska S.; Ambrosetti F.; Ridder L.; Marzella D. F.; Réau M. F.; Bonvin A. M. J. J.; Xue L. C. DeepRank: A Deep Learning Framework for Data Mining 3D Protein–Protein Interfaces. Nat. Commun. 2021, 12, 7068. 10.1038/s41467-021-27396-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Réau M.; Renaud N.; Xue L. C.; Bonvin A. M. J. J. DeepRank-GNN: A Graph Neural Network Framework to Learn Patterns in Protein-protein Interfaces. Bioinformatics 2023, 39, btac759. 10.1093/bioinformatics/btac759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang M.; Cang Z.; Wei G.-W. A Topology-based Network Tree for the Prediction of Protein-protein Binding Affinity Changes Following Mutation. Nat. Mach. Intell. 2020, 2, 116–123. 10.1038/s42256-020-0149-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- MacCallum J. L.; Perez A.; Dill K. A. Determining Protein Structures by Combining Semireliable Data with Atomistic Physical Models by Bayesian Inference. Proc. Natl. Acad. Sci. U. S. A. 2015, 112, 6985–6990. 10.1073/pnas.1506788112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fukuda H.; Tomii K. DeepECA: An End-to-end Learning Framework for Protein Contact Prediction from a Multiple Sequence Alignment. BMC Bioinf. 2020, 21, 10. 10.1186/s12859-019-3190-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jumper J.; Evans R.; Pritzel A.; Green T.; Figurnov M.; Ronneberger O.; Tunyasuvunakool K.; Bates R.; Žídek A.; Potapenko A.; Bridgland A.; Meyer C.; Kohl S. A. A.; Ballard A. J.; Cowie A.; Romera-Paredes B.; Nikolov S.; Jain R.; Adler J.; Back T.; Petersen S.; Reiman D.; Clancy E.; Zielinski M.; Steinegger M.; Pacholska M.; Berghammer T.; Bodenstein S.; Silver D.; Vinyals O.; Senior A. W.; Kavukcuoglu K.; Kohli P.; Hassabis D. Highly Accurate Protein Structure Prediction with AlphaFold. Nature 2021, 596, 583–589. 10.1038/s41586-021-03819-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baek M.; DiMaio F.; Anishchenko I.; Dauparas J.; Ovchinnikov S.; Lee G. R.; Wang J.; Cong Q.; Kinch L. N.; Schaeffer R. D.; Millãn C.; Park H.; Adams C.; Glassman C. R.; DeGiovanni A.; Pereira J. H.; Rodrigues A. V.; van Dijk A. A.; Ebrecht A. C.; Opperman D. J.; Sagmeister T.; Buhlheller C.; Pavkov-Keller T.; Rathinaswamy M. K.; Dalwadi U.; Yip C. K.; Burke J. E.; Garcia K. C.; Grishin N. V.; Adams P. D.; Read R. J.; Baker D. Accurate Prediction of Protein Structures and Interactions Using a Three-track Neural Network. Science 2021, 373, 871–876. 10.1126/science.abj8754. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bryant P.; Pozzati G.; Elofsson A. Improved Prediction of Protein-protein Interactions Using AlphaFold2. Nat. Commun. 2022, 13, 1265. 10.1038/s41467-022-28865-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kandathil S. M.; Greener J. G.; Lau A. M.; Jones D. T. Ultrafast End-to-end Protein Structure Prediction Enables High-throughput Exploration of Uncharacterized Proteins. Proc. Natl. Acad. Sci. U. S. A. 2022, 119, e2113348119. 10.1073/pnas.2113348119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steinegger M.; Söding J. MMseqs2 Enables Sensitive Protein Sequence Searching for the Analysis of Massive Data Sets. Nat. Biotechnol. 2017, 35, 1026–1028. 10.1038/nbt.3988. [DOI] [PubMed] [Google Scholar]
- Mirdita M.; Steinegger M.; Söding J. MMseqs2 Desktop and Local Web Server App for Fast, Interactive Sequence Searches. Bioinformatics 2019, 35, 2856–2858. 10.1093/bioinformatics/bty1057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mirdita M.; Schütze K.; Moriwaki Y.; Heo L.; Ovchinnikov S.; Steinegger M. ColabFold: Making Protein Folding Accessible to All. Nat. Methods 2022, 19, 679–682. 10.1038/s41592-022-01488-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Misiura M.; Shroff R.; Thyer R.; Kolomeisky A. B. DLPacker: Deep Learning for Prediction of Amino Acid Side Chain Conformations in Proteins. Proteins: Struct., Funct., Bioinf. 2022, 90, 1278–1290. 10.1002/prot.26311. [DOI] [PubMed] [Google Scholar]
- Li J.; Zhang O.; Lee S.; Namini A.; Liu Z. H.; Teixeira J. M. C.; Forman-Kay J. D.; Head-Gordon T. Learning Correlations between Internal Coordinates to Improve 3D Cartesian Coordinates for Proteins. J. Chem. Theory Comput. 2023, 10.1021/acs.jctc.2c01270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith J. S.; Isayev O.; Roitberg A. E. ANI-1: An Extensible Neural Network Potential with DFT Accuracy at Force Field Computational Cost. Chem. Sci. 2017, 8, 3192–3203. 10.1039/C6SC05720A. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith J. S.; Isayev O.; Roitberg A. E. ANI-1, a Data Set of 20 Million Calculated Off-equilibrium Conformations for Organic Molecules. Sci. Data 2017, 4, 170193. 10.1038/sdata.2017.193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith J. S.; Nebgen B.; Lubbers N.; Isayev O.; Roitberg A. E. Less Is More: Sampling Chemical Space with Active Learning. J. Chem. Phys. 2018, 148, 241733. 10.1063/1.5023802. [DOI] [PubMed] [Google Scholar]
- Gao X.; Ramezanghorbani F.; Isayev O.; Smith J. S.; Roitberg A. E. TorchANI: A Free and Open Source PyTorch-Based Deep Learning Implementation of the ANI Neural Network Potentials. J. Chem. Inf. Model. 2020, 60, 3408–3415. 10.1021/acs.jcim.0c00451. [DOI] [PubMed] [Google Scholar]
- Zubatyuk R.; Smith J. S.; Leszczynski J.; Isayev O. Accurate and Transferable Multitask Prediction of Chemical Properties with an Atoms-in-molecules Neural Network. Sci. Adv. 2019, 5, eaav6490. 10.1126/sciadv.aav6490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang L.; Han J.; Wang H.; Car R.; E W. Deep Potential Molecular Dynamics: A Scalable Model with the Accuracy of Quantum Mechanics. Phys. Rev. Lett. 2018, 120, 143001. 10.1103/PhysRevLett.120.143001. [DOI] [PubMed] [Google Scholar]
- Wang H.; Zhang L.; Han J.; E W. DeePMD-kit: A Deep Learning Package for Many-body Potential Energy Representation and Molecular Dynamics. Comput. Phys. Commun. 2018, 228, 178–184. 10.1016/j.cpc.2018.03.016. [DOI] [Google Scholar]
- Abbott A. S.; Turney J. M.; Zhang B.; Smith D. G. A.; Altarawy D.; Schaefer H. F. PES-Learn: An Open-Source Software Package for the Automated Generation of Machine Learning Models of Molecular Potential Energy Surfaces. J. Chem. Theory Comput. 2019, 15, 4386–4398. 10.1021/acs.jctc.9b00312. [DOI] [PubMed] [Google Scholar]
- Schütt K. T.; Kindermans P.-J.; Sauceda H. E.; Chmiela S.; Tkatchenko A.; Müller K.-R.. SchNet: A Continuous-Filter Convolutional Neural Network for Modeling Quantum Interactions. In Proceedings of the 31st International Conference on Neural Information Processing Systems, 2017; pp 992–1002.
- Schütt K. T.; Sauceda H. E.; Kindermans P.-J.; Tkatchenko A.; Müller K.-R. SchNet – a Deep Learning Architecture for Molecules and Materials. J. Chem. Phys. 2018, 148, 241722. 10.1063/1.5019779. [DOI] [PubMed] [Google Scholar]
- Schütt K. T.; Kessel P.; Gastegger M.; Nicoli K. A.; Tkatchenko A.; Müller K.-R. SchNetPack: A Deep Learning Toolbox For Atomistic Systems. J. Chem. Theory Comput. 2019, 15, 448–455. 10.1021/acs.jctc.8b00908. [DOI] [PubMed] [Google Scholar]
- Chmiela S.; Sauceda H. E.; Müller K.-R.; Tkatchenko A. Towards Exact Molecular Dynamics Simulations with Machine-Learned Force Fields. Nat. Commun. 2018, 9, 3887. 10.1038/s41467-018-06169-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chmiela S.; Sauceda H. E.; Poltavsky I.; Müller K.-R.; Tkatchenko A. sGDML: Constructing Accurate and Data Efficient Molecular Force Fields Using Machine Learning. Comput. Phys. Commun. 2019, 240, 38–45. 10.1016/j.cpc.2019.02.007. [DOI] [Google Scholar]
- Bogojeski M.; Vogt-Maranto L.; Tuckerman M. E.; Müller K.-R.; Burke K. Quantum Chemical Accuracy from Density Functional Approximations via Machine Learning. Nat. Commun. 2020, 11, 5223. 10.1038/s41467-020-19093-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vandermause J.; Torrisi S. B.; Batzner S.; Xie Y.; Sun L.; Kolpak A. M.; Kozinsky B. On-the-fly Active Learning of Interpretable Bayesian Force Fields for Atomistic Rare Events. npj Comput. Mater. 2020, 6, 20. 10.1038/s41524-020-0283-z. [DOI] [Google Scholar]
- Hajinazar S.; Thorn A.; Sandoval E. D.; Kharabadze S.; Kolmogorov A. N. MAISE: Construction of Neural Network Interatomic Models and Evolutionary Structure Optimization. Comput. Phys. Commun. 2021, 259, 107679. 10.1016/j.cpc.2020.107679. [DOI] [Google Scholar]
- Wen M.; Afshar Y.; Elliott R. S.; Tadmor E. B. KLIFF: A Framework to Develop Physics-based and Machine Learning Interatomic Potentials. Comput. Phys. Commun. 2022, 272, 108218. 10.1016/j.cpc.2021.108218. [DOI] [Google Scholar]
- Hu Z.; Guo Y.; Liu Z.; Shi D.; Li Y.; Hu Y.; Bu M.; Luo K.; He J.; Wang C.; Du S. AisNet: A Universal Interatomic Potential Neural Network with Encoded Local Environment Features. J. Chem. Inf. Model. 2023, 63, 1756–1765. 10.1021/acs.jcim.3c00077. [DOI] [PubMed] [Google Scholar]
- Ward L.; Blaiszik B.; Foster I.; Assary R. S.; Narayanan B.; Curtiss L. Machine Learning Prediction of Accurate Atomization Energies of Organic Molecules from Low-fidelity Quantum Chemical Calculations. MRS Commun. 2019, 9, 891–899. 10.1557/mrc.2019.107. [DOI] [Google Scholar]
- Rai B. K.; Sresht V.; Yang Q.; Unwalla R.; Tu M.; Mathiowetz A. M.; Bakken G. A. TorsionNet: A Deep Neural Network to Rapidly Predict Small-Molecule Torsional Energy Profiles with the Accuracy of Quantum Mechanics. J. Chem. Inf. Model. 2022, 62, 785–800. 10.1021/acs.jcim.1c01346. [DOI] [PubMed] [Google Scholar]
- Schriber J. B.; Nascimento D. R.; Koutsoukas A.; Spronk S. A.; Cheney D. L.; Sherrill C. D. CLIFF: A Component-based, Machine-learned, Intermolecular Force Field. J. Chem. Phys. 2021, 154, 184110. 10.1063/5.0042989. [DOI] [PubMed] [Google Scholar]
- Laghuvarapu S.; Pathak Y.; Priyakumar U. D. BAND NN: A Deep Learning Framework for Energy Prediction and Geometry Optimization of Organic Small Molecules. J. Comput. Chem. 2020, 41, 790–799. 10.1002/jcc.26128. [DOI] [PubMed] [Google Scholar]
- Li C.-H.; Tabor D. P. Reorganization Energy Predictions with Graph Neural Networks Informed by Low-Cost Conformers. J. Phys. Chem. A 2023, 127, 3484–3489. 10.1021/acs.jpca.2c09030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stepniewska-Dziubinska M. M.; Zielenkiewicz P.; Siedlecki P. Development and Evaluation of a Deep Learning Model for Protein-ligand Binding Affinity Prediction. Bioinformatics 2018, 34, 3666–3674. 10.1093/bioinformatics/bty374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karimi M.; Wu D.; Wang Z.; Shen Y. DeepAffinity: interpretable deep learning of compound-protein affinity through unified recurrent and convolutional neural networks. Bioinformatics 2019, 35, 3329–3338. 10.1093/bioinformatics/btz111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dong L.; Qu X.; Zhao Y.; Wang B. Prediction of Binding Free Energy of Protein-Ligand Complexes with a Hybrid Molecular Mechanics/Generalized Born Surface Area and Machine Learning Method. ACS Omega 2021, 6, 32938–32947. 10.1021/acsomega.1c04996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Q.; Wang P.-S.; Zhu C.; Gaines B. B.; Zhu T.; Bi J.; Song M. OctSurf: Efficient Hierarchical Voxel-based Molecular Surface Representation for Protein-Ligand Affinity Prediction. J. Mol. Graphics Modell. 2021, 105, 107865. 10.1016/j.jmgm.2021.107865. [DOI] [PubMed] [Google Scholar]
- Zheng L.; Fan J.; Mu Y. OnionNet: a Multiple-Layer Intermolecular-Contact-Based Convolutional Neural Network for Protein–Ligand Binding Affinity Prediction. ACS Omega 2019, 4, 15956–15965. 10.1021/acsomega.9b01997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Z.; Zheng L.; Liu Y.; Qu Y.; Li Y.-Q.; Zhao M.; Mu Y.; Li W. OnionNet-2: A Convolutional Neural Network Model for Predicting Protein-Ligand Binding Affinity Based on Residue-Atom Contacting Shells. Front. Chem. 2021, 9, 753002. 10.3389/fchem.2021.753002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kyro G. W.; Brent R. I.; Batista V. S. HAC-Net: A Hybrid Attention-Based Convolutional Neural Network for Highly Accurate Protein-Ligand Binding Affinity Prediction. J. Chem. Inf. Model. 2023, 63, 1947–1960. 10.1021/acs.jcim.3c00251. [DOI] [PubMed] [Google Scholar]
- Liu Z.; Lin L.; Jia Q.; Cheng Z.; Jiang Y.; Guo Y.; Ma J. Transferable Multilevel Attention Neural Network for Accurate Prediction of Quantum Chemistry Properties via Multitask Learning. J. Chem. Inf. Model. 2021, 61, 1066–1082. 10.1021/acs.jcim.0c01224. [DOI] [PubMed] [Google Scholar]
- Scheen J.; Wu W.; Mey A. S. J. S.; Tosco P.; Mackey M.; Michel J. Hybrid Alchemical Free Energy/Machine-Learning Methodology for the Computation of Hydration Free Energies. J. Chem. Inf. Model. 2020, 60, 5331–5339. 10.1021/acs.jcim.0c00600. [DOI] [PubMed] [Google Scholar]
- Lim H.; Jung Y. MLSolvA: Solvation Free Energy Prediction from Pairwise Atomistic Interactions by Machine Learning. J. Cheminf. 2021, 13, 56. 10.1186/s13321-021-00533-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang D.; Xia S.; Zhang Y. Accurate Prediction of Aqueous Free Solvation Energies Using 3D Atomic Feature-Based Graph Neural Network with Transfer Learning. J. Chem. Inf. Model. 2022, 62, 1840–1848. 10.1021/acs.jcim.2c00260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chung Y.; Vermeire F. H.; Wu H.; Walker P. J.; Abraham M. H.; Green W. H. Group Contribution and Machine Learning Approaches to Predict Abraham Solute Parameters, Solvation Free Energy, and Solvation Enthalpy. J. Chem. Inf. Model. 2022, 62, 433–446. 10.1021/acs.jcim.1c01103. [DOI] [PubMed] [Google Scholar]
- Pan X.; Zhao F.; Zhang Y.; Wang X.; Xiao X.; Zhang J. Z. H.; Ji C. MolTaut: A Tool for the Rapid Generation of Favorable Tautomer in Aqueous Solution. J. Chem. Inf. Model. 2023, 63, 1833–1840. 10.1021/acs.jcim.2c01393. [DOI] [PubMed] [Google Scholar]
- Skalic M.; Jiménez J.; Sabbadin D.; De Fabritiis G. Shape-Based Generative Modeling for de Novo Drug Design. J. Chem. Inf. Model. 2019, 59, 1205–1214. 10.1021/acs.jcim.8b00706. [DOI] [PubMed] [Google Scholar]
- Lee M.; Min K. MGCVAE: Multi-Objective Inverse Design via Molecular Graph Conditional Variational Autoencoder. J. Chem. Inf. Model. 2022, 62, 2943–2950. 10.1021/acs.jcim.2c00487. [DOI] [PubMed] [Google Scholar]
- Khemchandani Y.; O’Hagan S.; Samanta S.; Swainston N.; Roberts T. J.; Bollegala D.; Kell D. B. DeepGraphMolGen, a multi-objective, computational strategy for generating molecules with desirable properties: a graph convolution and reinforcement learning approach. J. Cheminf. 2020, 12, 53. 10.1186/s13321-020-00454-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ragoza M.; Masuda T.; Koes D. R. Generating 3D Molecules Conditional on Receptor Binding Sites with Deep Generative Models. Chem. Sci. 2022, 13, 2701–2713. 10.1039/D1SC05976A. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ingraham J.; Garg V.; Barzilay R.; Jaakkola T.. Generative Models for Graph-Based Protein Design. In Advances in Neural Information Processing Systems 32 (NeurIPS 2019); Curran Associates, 2019; pp 15820–15831.
- Goel M.; Raghunathan S.; Laghuvarapu S.; Priyakumar U. D. MoleGuLAR: Molecule Generation Using Reinforcement Learning with Alternating Rewards. J. Chem. Inf. Model. 2021, 61, 5815–5826. 10.1021/acs.jcim.1c01341. [DOI] [PubMed] [Google Scholar]
- Tysinger E. P.; Rai B. K.; Sinitskiy A. V. Can We Quickly Learn to “Translate” Bioactive Molecules with Transformer Models?. J. Chem. Inf. Model. 2023, 63, 1734–1744. 10.1021/acs.jcim.2c01618. [DOI] [PubMed] [Google Scholar]
- Kao P.-Y.; Yang Y.-C.; Chiang W.-Y.; Hsiao J.-Y.; Cao Y.; Aliper A.; Ren F.; Aspuru-Guzik A.; Zhavoronkov A.; Hsieh M.-H.; Lin Y.-C. Exploring the Advantages of Quantum Generative Adversarial Networks in Generative Chemistry. J. Chem. Inf. Model. 2023, 63, 3307–3318. 10.1021/acs.jcim.3c00562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Imrie F.; Bradley A. R.; van der Schaar M.; Deane C. M. Deep Generative Models for 3D Linker Design. J. Chem. Inf. Model. 2020, 60, 1983–1995. 10.1021/acs.jcim.9b01120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Imrie F.; Hadfield T. E.; Bradley A. R.; Deane C. M. Deep Generative Design with 3D Pharmacophoric Constraints. Chem. Sci. 2021, 12, 14577–14589. 10.1039/D1SC02436A. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang Y.; Zheng S.; Su S.; Zhao C.; Xu J.; Chen H. SyntaLinker: Automatic Fragment Linking with Deep Conditional Transformer Neural Networks. Chem. Sci. 2020, 11, 8312–8322. 10.1039/D0SC03126G. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tan Y.; Dai L.; Huang W.; Guo Y.; Zheng S.; Lei J.; Chen H.; Yang Y. DRlinker: Deep Reinforcement Learning for Optimization in Fragment Linking Design. J. Chem. Inf. Model. 2022, 62, 5907–5917. 10.1021/acs.jcim.2c00982. [DOI] [PubMed] [Google Scholar]
- Schwaller P.; Laino T.; Gaudin T.; Bolgar P.; Hunter C. A.; Bekas C.; Lee A. A. Molecular Transformer: A Model for Uncertainty-Calibrated Chemical Reaction Prediction. ACS Cent. Sci. 2019, 5, 1572–1583. 10.1021/acscentsci.9b00576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gong Y.; Xue D.; Chuai G.; Yu J.; Liu Q. DeepReac+: Deep Active Learning for Quantitative Modeling of Organic Chemical Reactions. Chem. Sci. 2021, 12, 14459–14472. 10.1039/D1SC02087K. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Probst D.; Schwaller P.; Reymond J.-L. Reaction Classification and Yield Prediction Using the Differential Reaction Fingerprint DRFP. Digital Discovery 2022, 1, 91–97. 10.1039/D1DD00006C. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heinen S.; von Rudorff G. F.; von Lilienfeld O. A. Toward the Design of Chemical Reactions: Machine Learning Barriers of Competing Mechanisms in Reactant Space. J. Chem. Phys. 2021, 155, 064105. 10.1063/5.0059742. [DOI] [PubMed] [Google Scholar]
- Lin Z.; Yin S.; Shi L.; Zhou W.; Zhang Y. J. G2GT: Retrosynthesis Prediction with Graph-to-Graph Attention Neural Network and Self-Training. J. Chem. Inf. Model. 2023, 63, 1894–1905. 10.1021/acs.jcim.2c01302. [DOI] [PubMed] [Google Scholar]
- Genheden S.; Thakkar A.; Chadimovã V.; Reymond J.-L.; Engkvist O.; Bjerrum E. AiZynthFinder: A Fast, Robust and Flexible Open-source Software for Retrosynthetic Planning. J. Cheminf. 2020, 12, 70. 10.1186/s13321-020-00472-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Genheden S.; Norrby P.-O.; Engkvist O. AiZynthTrain: Robust, Reproducible, and Extensible Pipelines for Training Synthesis Prediction Models. J. Chem. Inf. Model. 2023, 63, 1841–1846. 10.1021/acs.jcim.2c01486. [DOI] [PubMed] [Google Scholar]
- Ishida S.; Terayama K.; Kojima R.; Takasu K.; Okuno Y. AI-Driven Synthetic Route Design Incorporated with Retrosynthesis Knowledge. J. Chem. Inf. Model. 2022, 62, 1357–1367. 10.1021/acs.jcim.1c01074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mansimov E.; Mahmood O.; Kang S.; Cho K. Molecular Geometry Prediction using a Deep Generative Graph Neural Network. Sci. Rep. 2019, 9, 20381. 10.1038/s41598-019-56773-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simm G. N. C.; Hernández-Lobato J. M.. A Generative Model for Molecular Distance Geometry. In Proceedings of the 37th International Conference on Machine Learning, 2020; pp 8949–8958.
- Xu M.; Wang W.; Luo S.; Shi C.; Bengio Y.; Gomez-Bombarelli R.; Tang J.. An End-to-End Framework for Molecular Conformation Generation via Bilevel Programming. In Proceedings of the 38th International Conference on Machine Learning, 2021; pp 11537–11547.
- Shi C.; Luo S.; Xu M.; Tang J.. Learning Gradient Fields for Molecular Conformation Generation. In Proceedings of the 38th International Conference on Machine Learning, 2021; pp 9558–9568.
- Liu Z.; Zubatiuk T.; Roitberg A.; Isayev O. Auto3D: Automatic Generation of the Low-Energy 3D Structures with ANI Neural Network Potentials. J. Chem. Inf. Model. 2022, 62, 5373–5382. 10.1021/acs.jcim.2c00817. [DOI] [PubMed] [Google Scholar]
- Westermayr J.; Marquetand P. Deep Learning for UV Absorption Spectra with Schnarc: First Steps toward Transferability in Chemical Compound Space. J. Chem. Phys. 2020, 153, 154112. 10.1063/5.0021915. [DOI] [PubMed] [Google Scholar]
- Francoeur P. G.; Koes D. R. SolTranNet-A Machine Learning Tool for Fast Aqueous Solubility Prediction. J. Chem. Inf. Model. 2021, 61, 2530–2536. 10.1021/acs.jcim.1c00331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Francoeur P. G.; Koes D. R. Correction to “SolTranNet–A Machine Learning Tool for Fast Aqueous Solubility Prediction”. J. Chem. Inf. Model. 2021, 61, 4120–4123. 10.1021/acs.jcim.1c00806. [DOI] [PubMed] [Google Scholar]
- Ghosh K.; Stuke A.; Todorović M.; Jørgensen P. B.; Schmidt M. N.; Vehtari A.; Rinke P. Deep Learning Spectroscopy: Neural Networks for Molecular Excitation Spectra. Adv. Sci. 2019, 6, 1801367. 10.1002/advs.201801367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McNaughton A. D.; Joshi R. P.; Knutson C. R.; Fnu A.; Luebke K. J.; Malerich J. P.; Madrid P. B.; Kumar N. Machine Learning Models for Predicting Molecular UV-Vis Spectra with Quantum Mechanical Properties. J. Chem. Inf. Model. 2023, 63, 1462–1471. 10.1021/acs.jcim.2c01662. [DOI] [PubMed] [Google Scholar]
- Cerdãn L.; Roca-Sanjuán D. Reconstruction of Nuclear Ensemble Approach Electronic Spectra Using Probabilistic Machine Learning. J. Chem. Theory Comput. 2022, 18, 3052–3064. 10.1021/acs.jctc.2c00004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fine J. A.; Rajasekar A. A.; Jethava K. P.; Chopra G. Spectral Deep Learning for Prediction and Prospective Validation of Functional Groups. Chem. Sci. 2020, 11, 4618–4630. 10.1039/C9SC06240H. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Enders A. A.; North N. M.; Fensore C. M.; Velez-Alvarez J.; Allen H. C. Functional Group Identification for FTIR Spectra Using Image-Based Machine Learning Models. Anal. Chem. 2021, 93, 9711–9718. 10.1021/acs.analchem.1c00867. [DOI] [PubMed] [Google Scholar]
- Thrall E. S.; Lee S. E.; Schrier J.; Zhao Y. Machine Learning for Functional Group Identification in Vibrational Spectroscopy: A Pedagogical Lab for Undergraduate Chemistry Students. J. Chem. Educ. 2021, 98, 3269–3276. 10.1021/acs.jchemed.1c00693. [DOI] [Google Scholar]
- Mansouri K.; Grulke C. M.; Judson R. S.; Williams A. J. OPERA Models for Predicting Physicochemical Properties and Environmental Fate Endpoints. J. Cheminf. 2018, 10, 10. 10.1186/s13321-018-0263-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mansouri K.; Cariello N. F.; Korotcov A.; Tkachenko V.; Grulke C. M.; Sprankle C. S.; Allen D.; Casey W. M.; Kleinstreuer N. C.; Williams A. J. Open-source QSAR Models for pKa Prediction Using Multiple Machine Learning Approaches. J. Cheminf. 2019, 11, 60. 10.1186/s13321-019-0384-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baltruschat M.; Czodrowski P. Machine Learning Meets pKa. F1000Research 2020, 9, 113. 10.12688/f1000research.22090.2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pan X.; Wang H.; Li C.; Zhang J. Z. H.; Ji C. MolGpka: A Web Server for Small Molecule pKa Prediction Using a Graph-Convolutional Neural Network. J. Chem. Inf. Model. 2021, 61, 3159–3165. 10.1021/acs.jcim.1c00075. [DOI] [PubMed] [Google Scholar]
- Reis P. B. P. S.; Bertolini M.; Montanari F.; Rocchia W.; Machuqueiro M.; Clevert D.-A. A Fast and Interpretable Deep Learning Approach for Accurate Electrostatics-driven pKa Predictions in Proteins. J. Chem. Theory Comput. 2022, 18, 5068–5078. 10.1021/acs.jctc.2c00308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mayr F.; Wieder M.; Wieder O.; Langer T. Improving Small Molecule pKa Prediction Using Transfer Learning With Graph Neural Networks. Front. Chem. 2022, 10, 866585. 10.3389/fchem.2022.866585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai Z.; Liu T.; Lin Q.; He J.; Lei X.; Luo F.; Huang Y. Basis for Accurate Protein pKa Prediction with Machine Learning. J. Chem. Inf. Model. 2023, 63, 2936–2947. 10.1021/acs.jcim.3c00254. [DOI] [PubMed] [Google Scholar]
- Cai Z.; Luo F.; Wang Y.; Li E.; Huang Y. Protein pKa Prediction with Machine Learning. ACS Omega 2021, 6, 34823–34831. 10.1021/acsomega.1c05440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramsundar B.; Eastman P.; Walters P.; Pande V.; Leswing K.; Wu Z.. Deep Learning for the Life Sciences; O’Reilly Media, 2019 [Google Scholar]
- Raghunathan S.; Priyakumar U. D. Molecular Representations for Machine Learning Applications in Chemistry. Int. J. Quantum Chem. 2022, 122, e26870. 10.1002/qua.26870. [DOI] [Google Scholar]
- Wigh D. S.; Goodman J. M.; Lapkin A. A. A Review of Molecular Representation in the Age of Machine Learning. Wiley Interdiscip. Rev.: Comput. Mol. Sci. 2022, 12, e1603. 10.1002/wcms.1603. [DOI] [Google Scholar]
- O’Boyle N.; Banck M.; James C.; Morley C.; Vandermeersch T.; Hutchison G. Open Babel: An Open Chemical Toolbox. J. Cheminf. 2011, 3, 33. 10.1186/1758-2946-3-33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- RDKit: Open-Source Cheminformatics Software. https://www.rdkit.org/ (accessed 2023-06-20).
- Chen D.; Zheng J.; Wei G.-W.; Pan F. Extracting Predictive Representations from Hundreds of Millions of Molecules. J. Phys. Chem. Lett. 2021, 12, 10793–10801. 10.1021/acs.jpclett.1c03058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Safizadeh H.; Simpkins S. W.; Nelson J.; Li S. C.; Piotrowski J. S.; Yoshimura M.; Yashiroda Y.; Hirano H.; Osada H.; Yoshida M.; Boone C.; Myers C. L. Improving Measures of Chemical Structural Similarity Using Machine Learning on Chemical-Genetic Interactions. J. Chem. Inf. Model. 2021, 61, 4156–4172. 10.1021/acs.jcim.0c00993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fabregat R.; van Gerwen P.; Haeberle M.; Eisenbrand F.; Corminboeuf C. Metric Learning for Kernel Ridge Regression: Assessment of Molecular Similarity. Mach. Learn.: Sci. Technol 2022, 3, 035015. 10.1088/2632-2153/ac8e4f. [DOI] [Google Scholar]
- Venkatraman V. FP-ADMET: A Compendium of Fingerprint-based ADMET Prediction Models. J. Cheminf. 2021, 13, 75. 10.1186/s13321-021-00557-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen T.; Guestrin C.. XGBoost: A Scalable Tree Boosting System. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery & Data Mining, New York, 2016; pp 785–794.
- Tian H.; Ketkar R.; Tao P. ADMETboost: A Web Server for Accurate ADMET Prediction. J. Mol. Model. 2022, 28, 408. 10.1007/s00894-022-05373-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ulrich N.; Goss K.-U.; Ebert A. Exploring the Octanol-water Partition Coefficient Dataset Using Deep Learning Techniques and Data Augmentation. Commun. Chem. 2021, 4, 90. 10.1038/s42004-021-00528-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Isert C.; Kromann J. C.; Stiefl N.; Schneider G.; Lewis R. A. Machine Learning for Fast, Quantum Mechanics-Based Approximation of Drug Lipophilicity. ACS Omega 2023, 8, 2046–2056. 10.1021/acsomega.2c05607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Win Z.-M.; Cheong A. M. Y.; Hopkins W. S. Using Machine Learning To Predict Partition Coefficient (Log P) and Distribution Coefficient (Log D) with Molecular Descriptors and Liquid Chromatography Retention Time. J. Chem. Inf. Model. 2023, 63, 1906–1913. 10.1021/acs.jcim.2c01373. [DOI] [PubMed] [Google Scholar]
- Liu Y.; Li Z. Predict Ionization Energy of Molecules Using Conventional and Graph-Based Machine Learning Models. J. Chem. Inf. Model. 2023, 63, 806–814. 10.1021/acs.jcim.2c01321. [DOI] [PubMed] [Google Scholar]
- Nandy A.; Terrones G.; Arunachalam N.; Duan C.; Kastner D. W.; Kulik H. J. MOFSimplify, Machine Learning Models with Extracted Stability Data of Three Thousand Metal-Organic Frameworks. Sci. Data 2022, 9, 74. 10.1038/s41597-022-01181-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang K.; Swanson K.; Jin W.; Coley C.; Eiden P.; Gao H.; Guzman-Perez A.; Hopper T.; Kelley B.; Mathea M.; Palmer A.; Settels V.; Jaakkola T.; Jensen K.; Barzilay R. Analyzing Learned Molecular Representations for Property Prediction. J. Chem. Inf. Model. 2019, 59, 3370–3388. 10.1021/acs.jcim.9b00237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang K.; Swanson K.; Jin W.; Coley C.; Eiden P.; Gao H.; Guzman-Perez A.; Hopper T.; Kelley B.; Mathea M.; Palmer A.; Settels V.; Jaakkola T.; Jensen K.; Barzilay R. Correction to Analyzing Learned Molecular Representations for Property Prediction. J. Chem. Inf. Model. 2019, 59, 5304–5305. 10.1021/acs.jcim.9b01076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heid E.; Green W. H. Machine Learning of Reaction Properties via Learned Representations of the Condensed Graph of Reaction. J. Chem. Inf. Model. 2022, 62, 2101–2110. 10.1021/acs.jcim.1c00975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Adams K.; Pattanaik L.; Coley C. W.. Learning 3D Representations of Molecular Chirality with Invariance to Bond Rotations. Presented at the International Conference on Learning Representations, 2022.
- Zhu W.; Zhang Y.; Zhao D.; Xu J.; Wang L. HiGNN: A Hierarchical Informative Graph Neural Network for Molecular Property Prediction Equipped with Feature-Wise Attention. J. Chem. Inf. Model. 2023, 63, 43–55. 10.1021/acs.jcim.2c01099. [DOI] [PubMed] [Google Scholar]
- Wellawatte G. P.; Seshadri A.; White A. D. Model Agnostic Generation of Counterfactual Explanations for Molecules. Chem. Sci. 2022, 13, 3697–3705. 10.1039/D1SC05259D. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qian Y.; Guo J.; Tu Z.; Li Z.; Coley C. W.; Barzilay R. MolScribe: Robust Molecular Structure Recognition with Image-to-Graph Generation. J. Chem. Inf. Model. 2023, 63, 1925–1934. 10.1021/acs.jcim.2c01480. [DOI] [PubMed] [Google Scholar]
- Galeazzo T.; Shiraiwa M. Predicting Glass Transition Temperature and Melting Point of Organic Compounds Via Machine Learning and Molecular Embeddings. Environ. Sci.: Atmos. 2022, 2, 362–374. 10.1039/D1EA00090J. [DOI] [Google Scholar]
- Hormazabal R. S.; Kang J. W.; Park K.; Yang D. R. Not from Scratch: Predicting Thermophysical Properties through Model-Based Transfer Learning Using Graph Convolutional Networks. J. Chem. Inf. Model. 2022, 62, 5411–5424. 10.1021/acs.jcim.2c00846. [DOI] [PubMed] [Google Scholar]
- Vanschoren J.; van Rijn J. N.; Bischl B.; Torgo L. OpenML: Networked Science in Machine Learning. SIGKDD Explor. Newsl. 2014, 15, 49–60. 10.1145/2641190.2641198. [DOI] [Google Scholar]
- He H. The State of Machine Learning Frameworks in 2019. The Gradient, October 10, 2019. https://thegradient.pub/state-of-ml-frameworks-2019-pytorch-dominates-research-tensorflow-dominates-industry (accessed 2023-06-20).
- Kezmann J. M.Pytorch vs Tensorflow in 2022—Pros & Cons and Which Framework Is Best for You. Medium, September 28, 2022. https://medium.com/mlearning-ai/pytorch-vs-tensorflow-in-2022-9a7106b1f606 (accessed 2023-06-20).
- Novac O.-C.; Chirodea M. C.; Novac C. M.; Bizon N.; Oproescu M.; Stan O. P.; Gordan C. E. Analysis of the Application Efficiency of TensorFlow and PyTorch in Convolutional Neural Network. Sensors 2022, 22, 8872. 10.3390/s22228872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lowndes J. S. S.; Best B. D.; Scarborough C.; Afflerbach J. C.; Frazier M. R.; O’Hara C. C.; Jiang N.; Halpern B. S. Our Path to Better Science in Less Time Using Open Data Science Tools. Nat. Ecol. Evol. 2017, 1, 0160. 10.1038/s41559-017-0160. [DOI] [PubMed] [Google Scholar]
- Ivie P.; Thain D. Reproducibility in Scientific Computing. ACM Comput. Surv. 2019, 51, 63. 10.1145/3186266. [DOI] [Google Scholar]
- Wilson G.; Aruliah D. A.; Brown C. T.; Chue Hong N. P.; Davis M.; Guy R. T.; Haddock S. H. D.; Huff K. D.; Mitchell I. M.; Plumbley M. D.; Waugh B.; White E. P.; Wilson P. Best Practices for Scientific Computing. PLoS Biol. 2014, 12, e1001745. 10.1371/journal.pbio.1001745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson G.; Bryan J.; Cranston K.; Kitzes J.; Nederbragt L.; Teal T. K. Good Enough Practices in Scientific Computing. PLoS Comput. Biol. 2017, 13, e1005510. 10.1371/journal.pcbi.1005510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hunter-Zinck H.; de Siqueira A. F.; Vásquez V. N.; Barnes R.; Martinez C. C. Ten Simple Rules on Writing Clean and Reliable Open-source Scientific Software. PLoS Comput. Biol. 2021, 17, e1009481. 10.1371/journal.pcbi.1009481. [DOI] [PMC free article] [PubMed] [Google Scholar]