Abstract
The battle against emerging viral infections has been uneven, as there is currently no broad-spectrum drug available to contain the spread of novel pathogens throughout the population. Consequently, the pandemic outbreak that occurred in early 2020 laid bare the almost empty state of the pandemic box. Therefore, the development of novel treatments with broad specificity has become a paramount concern in this post-pandemic era.
Here, we propose copolymers of poly (sodium 2-(acrylamido)-2-methyl-1-propanesulfonate) (PAMPS) and poly (sodium 11-(acrylamido)undecanoate (AaU), both block (PAMPS75-b-PAaUn) and random (P(AMPSm-co-AaUn)) that show efficacy against a broad range of alpha and betacoronaviruses.
Owing to their intricate architecture, these polymers exhibit a highly distinctive mode of action, modulating nano-mechanical properties of cells and thereby influencing viral replication. Through the employment of confocal and atomic force microscopy techniques, we discerned perturbations in actin and vimentin filaments, which correlated with modification of cellular elasticity and reduction of glycocalyx layer. Intriguingly, this process was reversible upon polymer removal from the cells. To ascertain the applicability of our findings, we assessed the efficacy and underlying mechanism of the inhibitors using fully differentiated human airway epithelial cultures, wherein near-complete abrogation of viral replication was documented.
Given their mode of action, these polymers can be classified as biologically active nanomaterials that exploit a highly conserved molecular target—cellular plasticity—proffering the potential for truly broad-spectrum activity while concurrently for drug resistance development is minimal.
Keywords: Polymers, Nanomaterials, Coronaviruses, Antivirals, Broad-spectrum
Graphical abstract
1. Introduction
The Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) is the third highly pathogenic zoonotic coronavirus that emerged in the human population in the 21st century. Although the Severe Acute Respiratory Syndrome Coronavirus (SARS-CoV) and the Middle East Respiratory Syndrome Coronavirus (MERS-CoV) heightened our awareness of the potential biological hazards posed by this viral family, SARS-CoV-2 has had a far more devastating impact on global health, social structures, and economic stability, unparalleled since the Spanish influenza pandemic [[1], [2], [3]]. As per the data accessible via the World Health Organization (WHO) website, the SARS-CoV-2 pandemic has thus far resulted in over 6.9 million fatalities, notwithstanding the availability of vaccines and targeted therapeutics (https://covid19.who.int/, accessed May 18, 2023). However, it is crucial to note that the actual death toll is estimated to be approximately three to four times higher than the reported figures [4].
Coronaviruses comprise positive-sense, single-stranded RNA viruses that encompass a diverse array of pathogens known to infect numerous mammalian and avian species, including livestock and companion animals. These viruses are responsible for a broad spectrum of diseases, ranging from relatively benign to consistently fatal outcomes [5]. The long-known human coronaviruses, HCoV-229E and HCoV-OC43, as well as the more recently identified HCoV-NL63 and HCoV-HKU1, circulate among humans and typically cause seasonal, comparatively mild respiratory tract infections, often characterized by symptoms of the common cold. In stark contrast, the highly pathogenic SARS-CoV, MERS-CoV, and SARS-CoV-2 cause severe illnesses that have proven fatal for a proportion of infected individuals [6]. Consequently, coronaviruses pose a threat to public health but also raise veterinary and economic concerns [7,8].
Polymers were attributed with antiviral properties ∼80 years ago [9]. Initial investigations centered on natural polysaccharides, predominantly dextran sulfate, heparin, and agar [[10], [11], [12], [13]]. The multimeric character of polymers allows for the organized multiplication of the active moieties. Consequently, the sum of the weak interactions can result in a surprisingly strong antiviral effect that can be compared to a zipper. During the HIV epidemic of the early 1980s, interest in antiviral polymers surged, culminating in several promising compounds advancing to clinical trial phases; regrettably, these efforts were met with limited success [[14], [15], [16]]. Although polymers typically act as early-stage blockers of a viral infection, interfering with the virus attachment and entry, other mechanisms of action have been also reported as well as their activity toward different viral species [[17], [18], [19], [20], [21], [22], [23]].
While natural polymers undeniably offer certain advantages, their limited clinical success has resulted in diminished confidence in their therapeutic potential. Nevertheless, advancements in chemical synthesis have facilitated the efficient generation of synthetic polymers, the properties, efficacy, and toxicity of which can be readily modulated and optimized to better suit clinical applications.
In our recent study, we documented the potent antiviral properties of a diblock copolymer, PAMPS-b-PAaU, comprising poly (sodium 2-(acrylamido)-2-methylpropanesulfonate) (PAMPS) and poly (sodium 11-(acrylamido) undecanoate) (PAaU), against the Zika virus [24]. This diblock copolymer demonstrates pH-induced self-association, leading to the formation of core-shell-type, negatively charged nanoparticles (with a zeta potential of −25 mV and hydrodynamic radius of 4–12 nm) [25]. Our investigations revealed that the copolymers impeded infection by exerting a protective influence on host cells; however, the precise molecular mechanism remains to be elucidated [24].
Here, we evaluated a series of PAMPS-b-PAaU block copolymers against both high (HPHCV) and low-pathogenic human coronaviruses (LPHCV). The PAMPS-b-PAaU copolymers demonstrated inhibitory effects on a wide array of coronaviruses, including SARS-CoV-2, exhibiting IC50 values within the picomolar range and exceptionally high selectivity. These characteristics were further corroborated in fully differentiated human airway epithelium (HAE) cultures, wherein the copolymers effectively abrogated SARS-CoV-2 replication. Mechanistic analyses, encompassing atomic force and confocal microscopies, revealed a substantial influence of PAMPS-b-PAaU copolymers on cellular plasticity and membrane rigidity. We speculate that this may affect the production of infectious virus particles and cell-to-cell spread. Notably, this impact was reversible. Given their mode of action and activity, these polymers can be classified as biologically active nanomaterials targeting a highly conserved molecular feature—cellular plasticity—thereby offering the potential for truly broad-spectrum antiviral activity and significantly reduced likelihood of resistance emergence.
2. Materials and methods
2.1. Compounds synthesis
PAMPS homopolymers and diblock copolymers of poly (sodium 2-acrylamido-2-methyl-1-propanesulfonate) (PAMPS) with poly (sodium 11-acrylamido undecanoate) (PAaU) were synthesized via reversible addition-fragmentation chain transfer (RAFT) radical polymerization following the procedure described before [25]. Their characteristics are provided in our previous paper [19].
2.2. Cells and viruses
Vero (Cercopithecus aethiops; kidney epithelial; ATCC CCL-81); HCT-8 (ATCC CCL-244), a derivative of HRT-18 (Homo sapiens; male; ileocecal colorectal adenocarcinoma; ATCC CCL-244); MRC-5 (Homo sapiens; male; lung fibroblast; ATCC: CCL-171) cells were maintained in Dulbecco-modified Eagle's medium (DMEM, high glucose, ThermoFisher Scientific, Poland) supplemented with 5% heat-inactivated fetal bovine serum (FBS, ThermoFisher Scientific, Poland). The medium was supplemented with penicillin (100 U/mL, ThermoFisher Scientific, Poland) and streptomycin (100 μg/ml, ThermoFisher Scientific, Poland). Cells were cultured at 37 °C in an atmosphere containing 5% CO2 and humidity. Every two weeks, cells were tested for mycoplasma contamination.
The Human Airway Epithelium culture (HAE) derived from primary human bronchial epithelial cells was purchased (Epithelix Sarl, Switzerland) and expanded in the bronchial epithelial growth medium (BEGM) in-house. When confluent, cells were detached using trypsin and seeded onto permeable Thincert™ culture inserts (Greiner Bio-One, cat. no. 662641, Austria). Cells were cultured submerged in BEGM medium on the apical and basolateral side until confluent; next, the apical medium was discarded, while the basolateral medium was changed to an air-liquid interface (ALI) medium. Cells were cultured for 4 weeks to form fully-differentiated, polarized cultures that manifested in vivo pseudostratified mucociliary epithelium phenotype. All cells were maintained at 37 °C under 5% CO2.
Reference SARS-CoV-2 strain was isolated in-house in the spring of 2020 in Poland. The variant (B.1.13) is designated hCoV-19/Poland/PL_P7/2020 (GISAID accession code: EPI_ISL_428930). The SARS-CoV-2 stock was generated by infecting monolayers of Vero cells. The cells were incubated for 2 days at 37 °C under 5% CO2. The virus-containing medium was collected, aliquoted, and stored at −80 °C. Control samples from mock-infected cells were prepared in the same manner.
The HCoV-OC43 (ATCC: VR-1558) and HCoV-229E (NCPV: 0310051v) stocks were generated by infecting monolayers of HCT-8 and MRC-5 cells, respectively. The cells were incubated at 32 °C under 5% CO2 and then lysed by two freeze-thaw cycles at 5 or 6 days post-infection (p.i.), respectively. Control samples from mock-infected cells were prepared in the same manner. The virus- and mock-containing liquids were aliquoted and stored at −80 °C.
Virus yields were assessed by titration on fully confluent Vero, HCT-8, and MRC-5 cells in 96-well plates (TPP, Trasadingen, Switzerland) according to the method of Reed and Muench. Plates were incubated at 37 °C or 32 °C, and the cytopathic effect (CPE) was scored by observation under an inverted microscope.
2.3. Cell viability assay
Cell viability was evaluated using a commercially available Cell Proliferation Kit (XTT-based) (Biological Industries, USA). The kit was used according to the manufacturer's protocol. Fully confluent MRC-5 and HCT-8 cells were cultured on 96-well plates and incubated with compounds for 2 h at 37 °C, the medium was discarded, and fresh compounds were added. Cells with compounds were incubated for 3 days at 37 °C. After incubation, the medium was discarded, and 100 μl of fresh medium was overlaid on the cells. Next, 50 μl of activated 2,3-bis-(2-methoxy-4-nitro-5-sulfenyl)-(2H)-tetrazolium-5-carboxanilide (XTT) solution was added to each well, and plates were incubated for 2 h at 37 °C. The absorbance (λ = 490 nm) was measured using SpectraMax iD5 Multi-Mode Microplate Reader (Molecular Devices, San Jose, CA, USA). Data are shown as results normalized to the control samples (untreated cells), where cell viability was set to 100%.
2.4. Virus inhibition assay
Vero, HCT-8 or MRC-5 cells were seeded in a culture medium on a 96-well plate 1 day before infection. Fully confluent cells were infected with the SARS-CoV-2, HCoV-OC43, or HCoV-229E virus at 1600 (SARS-CoV-2 and HCoV-OC43) or 4000 (HCoV-229E) 50% tissue culture infectious dose (TCID50)/ml in the presence or absence of studied polymers. After 2 h of incubation at 37 °C (SARS-CoV-2) or 32 °C (HCoV-OC43 and HCoV-229E), cells were washed twice with PBS, and a fresh medium with polymers was added. The infection was carried out for 2 (SARS-CoV-2) or 5 (HCoV-OC43 and HCoV-229E) days, and the cytopathic effect (CPE) was assessed. Supernatants were diluted 10 × before collection, and 10 μl of each sample was added to the isolation buffer for viral RNA isolation. The number of SARS-CoV-2 RNA copies was assessed using RT-qPCR.
Virus replication inhibition ex vivo was evaluated by infecting HAE cultures with SARS-CoV-2 virus at 5000 TCID50/ml in the presence of PAMPS75-b-PAaU39, P(AMPS50-co-AaU50), or PBS. The compounds in concentrations of 100 μg/ml and the control were added to the apical side of the inserts, followed by the addition of the virus diluted in PBS for 2 h. Next, the apical side of the HAEs was washed thrice with PBS, and each compound was re-applied and incubated for 30 min at 37 °C. Samples were collected and diluted 10 ×; the HAEs were maintained in air-liquid interphase. Every 24 h until 96 h, the HAEs were incubated for 30 min with the compounds or controls, and 10 μl of apical washes were collected. Virus yield was assessed using the RT-qPCR, as described below.
2.5. Mechanism of action
Four different assays were performed to discover the mechanism of action of the copolymers. The Vero cell line and PAMPS75-b-PAaU39 were used in each experiment. In detail, cells were seeded in a culture medium on a 12-well plate 1 day before infection. Fully confluent cells were inoculated with the SARS-CoV-2 virus in the presence or absence of PAMPS75-b-PAaU39 polymer [26]. A schematic illustration of the time of addition experiment is shown in Supplementary Fig. 1.
2.5.1. Assay I – virus inactivation assay
The virus at 500,000 TCID50/ml was preincubated with 500 μg/ml of the tested copolymer for 1 h at RT. The mixture was then diluted to reach the final virus titer of 1600 TCID50/ml and polymer below active concentration and applied to the cells. After 2 h of infection, the cells were washed twice, and a fresh portion of the medium was overlaid. The replication was conducted for 24 h, and cells were washed and fixed with 3.7% w/v paraformaldehyde (PFA).
2.5.2. Assay II – virus attachment assay
Cells were chilled for 30 min at 4 °C, and the ice-cold virus at 1600 TCID50/ml was applied in the presence or absence of 50 μg/ml PAMPS75-b-PAaU39. After 2 h, cells were washed twice with cold PBS and fixed with 3.7% w/v PFA.
2.5.3. Assay III – virus entry assay
Cells were chilled for 30 min at 4 °C, and the ice-cold virus was applied at 1600 TCID50/ml. After 2 h of incubation, the cells were washed twice with cold PBS, and the 50 μg/ml of ice-cold polymer was added and incubated for 10 min at 4 °C. Next, cells were incubated for 1 h at 37 °C, then washed twice with PBS and fixed with 3.7% w/v PFA.
2.5.4. Assay IV – virus replication, assembly, and egress assay
Cells were infected at 1600 TCID50/ml and infection was carried out for 2 h at 37 °C. Next, cells were washed twice with PBS to remove any residual virus, and fresh medium, with or without the polymer, was overlaid on the cells. The replication was conducted for 24 h, and cells were washed and fixed with 3.7% w/v PFA.
After fixation, the cells were stained to visualize SARS-CoV-2.
2.6. Isolation of nucleic acids and quantitative PCR
A viral DNA/RNA kit (A&A Biotechnology, Gdansk, Poland) was used for nucleic acid isolation from cell culture supernatants and cells. RNA was isolated according to the manufacturer's instructions. Viral RNA was quantified using quantitative PCR coupled with reverse transcription (RT-qPCR) (GoTaq Probe 1-Step RT-qPCR System, Promega, Poland) using a CFX96 Touch real-time PCR detection system (Bio-Rad, Munich, Germany). The reaction was carried out in the presence of the probes and primers used before [27,28]. The heating scheme was as follows: 15 min at 45 °C and 2 min at 95 °C, followed by 40 cycles of 15 s at 95 °C and 1 min at 56 °C. To assess the copy number of the N gene, standards were prepared. The PCR product was amplified and cloned into pTZ57 R/T plasmids using an InsTAclone PCR cloning kit (Thermo Fisher Scientific, Poland). The resulting plasmid was linearized using the HindIII restriction enzyme. The linear product was purified with the GeneJet PCR purification kit (Thermo Fisher Scientific), and its concentration was assessed using a NanoDrop™ 2000 spectrophotometer (Thermo Fisher Scientific, Poland). The number of DNA copies per milliliter was calculated using the approximate molecular weight of deoxyribonucleotide (320 g/mol) and Avogadro's constant. Eight 10-fold serial dilutions were used as a qPCR template to develop a standard curve.
2.7. PAMPS75-b-PAaU28 visualization
To examine compounds entry into the cells, fluorescein and 50 μg/ml fluorescein-conjugated PAMPS75-b-PAaU28 were overlaid on Vero cells seeded on glass bottom dishes (HBSt-5040, Willco Wells) 24 h earlier. Nuclear DNA was stained with NucBlue Live ReadyProbes Reagent (Hoechst 33342, 2 drops/ml, 10 min, RT, R37605, Invitrogen) 10 min before the start of imaging, and PAMPS75-b-PAaU28-FL or an equal number of moles of free fluorescein were added on cells 2 min before imaging. Stacks of photos were collected every 2 min for 48 min using LSM 880 confocal microscope with ZEN 2.1 SP3 FP3 software. Obtained data were processed using ImageJ Fiji [29].
2.8. Staining
2.8.1. SARS-CoV-2 visualization
After fixation, cells were permeabilized using 0.5% Triton X-100 (13 min, RT). Unspecific binding sites were blocked with 5% bovine serum albumin (BSA) in PBS (overnight, 4 °C) before staining. For SARS-CoV-2 virions visualization, mouse anti-SARS-CoV-2 nucleocapsid protein antibody (1:200, 2 h, RT, MA5-29981, Invitrogen) followed by donkey anti-mouse Alexa Fluor 488 antibody (1:400, 1 h, RT) were used. After incubating with each antibody, cells were washed thrice with 0.5% Tween-20. Then, the actin cytoskeleton was visualized using Alexa Fluor 546 conjugated phalloidin (4 U/mL, 1 h, RT, A22283, Invitrogen), and nuclear DNA was stained with 4′,6-diamidino-2-phenylindole dihydrochloride (DAPI, 0.1 mg/ml, 20 min, RT, D9542, Sigma-Aldrich). Stained coverslips were mounted on glass slides with Prolong Diamond antifade mountant (P36961, Invitrogen).
2.8.2. Cytoskeleton visualization
For cytoskeleton visualization, Vero cells were seeded on coverslips in 12-well plates 1 day before infection. The PAMPS75-b-PAaU39 at a concentration of 50 μg/ml was administered onto cells and incubated for 15 min, 1 h, 2 h, 3 h, 4 h, and 6 h; H2O was used as a control. After incubation, cells were fixed using 3.7% w/v PFA (15 min, RT) and permeabilized using 0.5% Trition-X100 (13 min, RT), and unspecific binding sites were blocked with 5% bovine serum albumin (BSA) in PBS (overnight, 4 °C) prior to staining. For cytoskeleton visualization, mouse anti-vinculin antibody (1:300, 2 h, RT, V9131, Sigma-Aldrich) followed by donkey anti-mouse Alexa Fluor 488 antibody (1:400, 1 h, RT, A21202, Invitrogen), and rabbit anti-vimentin antibody conjugated with Alexa Fluor 647 (1:400, O/N, 4 °C, 9856, Cell Signaling Technology) were used. After incubating with each antibody, cells were washed thrice with 0.5% Tween-20. Then, the actin cytoskeleton was visualized using Alexa Fluor 546 conjugated phalloidin (4 U/ml, 1 h, RT, A22283, Invitrogen), and nuclear DNA was stained with 4′,6-diamidino-2-phenylindole dihydrochloride (DAPI, 0.1 mg/ml, 20 min, RT, D9542, Sigma-Aldrich). Stained coverslips were mounted on glass slides with Prolong Diamond antifade mountant (P36961, Invitrogen).
Images were acquired under a Zeiss LSM 710 confocal microscope (Carl Zeiss Microscopy GmbH; release version 8.1) using ZEN 2012 SP1 software (Carl Zeiss Microscopy GmbH; black edition, version 8.1.0.484) and collected as z-stacks (step size 0.20 μm (Vero cells), frame size 2048 × 2048 pixels, pixel size 0.12 μm). Within a single biological repeat of the experiment, images were processed equally using the ImageJ FIJI version and are presented as the maximal projections. Virus particles and nuclei were counted using the “analyze particles” software tool.
2.9. Virus particles counting
Calculation of viral particles from confocal images was performed using the 3D Object Counter v2.0 tool incorporated into the ImageJ Fiji image processing package. The number of viral particles calculated for each stack was divided by the number of cells on the same image to estimate the number of particles per cell. Threshold and size filters were selected experimentally using pictures from control and non-infected cells; cell nuclei areas were excluded from calculations due to excessive non-specific signals. Particles internalized into cells were counted manually from orthogonal views of images with actin visualizing the boundary of the cell.
2.10. AFM imaging
2.10.1. Elasticity measurement
Vero cells were seeded on a glass coverslip for 24 h to reach a confluence of about 70%. Next, the sample with cells was gently mounted into AFM liquid cell (BioCell, JPK Instruments, Billerica, USA) filled with Hanks' Balanced Salt Solution (HBSS, Cat. No. 55037C, Sigma-Aldrich, St. Louis, USA) supplemented with 1% FBS (ATCC). The sample was kept at a stable temperature of 37 °C throughout the measurement. PAMPS75-b-PAaU39 at a concentration of 50 μg/ml was added directly to the sample mounted in a liquid cell and the elasticity measurements were performed after subsequent incubation times: 15 min, 1 h, 2 h, 3 h, 4 h, and 6 h. Non-treated cells were measured as the negative control (control) throughout the experiment. The elasticity measurements were performed using a NanoWizard 3 NanoScience AFM (JPK Instruments, Billerica, USA) working in force mapping mode. For each cell, a spatial map of force vs. distance (FD) curves at a grid of 5 × 5 points and a square surface of 20 μm × 20 μm were measured. The position of scan areas was controlled by inverted optical microscopy (IX-71, Olympus, Tokyo, Japan). Force-distance curves were measured at a speed of 1 μm/s with the maximal applied force of 1 nN. To evaluate changes in the elastic modulus of cells and glycocalyx detection, the non-covered spherical polystyrene probe with a radius of 2.2 μm (Novascan, Chicago, USA) mounted on the triangular cantilever with the spring constant 0.03 N/m was used. Before each measurement, the cantilever spring constant was calibrated using specialistic software (SPM Software, JPK Instruments, Billerica, USA). In order to calculate the elastic modulus of cell [E of the cell], the elastic modulus of the cellular cortex [E of the cortex], and the thickness of glycocalyx layer [T of Glycocalyx layer], a method of FD analysis described in previous works [30,31] was used. The analysis was performed by using software written in the Matlab environment.
2.10.2. AFM quantitative imaging
Cells were prepared according to the protocol described in the previous section and incubated with PAMPS75-b-PAaU39 (50 μg/ml) for 15 min, 1 h, 2 h, 3 h, 4 h, and 6 h. After incubation, cells were fixed at the appropriate time points. For this purpose, the slides with cells were first gently washed with Hanks' Balanced Salt Solution (HBSS, Cat. No. 55037C, Sigma-Aldrich, St. Louis, USA) and then 0.5% glutaraldehyde solution (Cat. No. G7651, Sigma-Aldrich, St. Louis, MO, USA) was added for 2 min at room temperature. After this time, cells were washed and placed in Hanks' Balanced Salt Solution (HBSS, Cat. No. 55037C, Sigma-Aldrich, St. Louis, USA) and imaged sequentially. All experiments were repeated two times.
2.10.3. QI-AFM imaging: for fixed cells
AFM imaging was performed using V-shaped reflective gold-coated cantilevers (MLCT-BIO, Bruker, Billerica, USA) with a nominal spring constant of 0.03 N/m. All experiments were performed in a drop of Hanks' Balanced Salt Solution (HBSS, Cat. No. 55037C, Sigma-Aldrich, St. Louis, USA). Images (64 × 64 pixels) were obtained at a scan size of 20 × 20 μm2. Topographical images were performed using force-distance (FD)–based imaging mode (QI; JPK Instruments, Billerica, USA), allowing for high-resolution imaging of cells. In this method, a single FD-curve measurement is performed at every pixel of the image and then translated from the selected trigger force into images of cell topography. The loading force varied from 1.0 to 1.2 nN and was adjusted to obtain a clear contrast of the cell surface. The obtained images of topography were analyzed using JPK Data Processing Software.
2.11. Statistics
The experiments were carried out in at least three replicates. The data are shown as means ± standard deviations (S.D.). The statistical significance of the data presented in the manuscript was assessed with the non-parametric Kruskal–Wallis test, and P values below 0.05 were considered significant unless stated otherwise. Statistical analysis was performed using GraphPad Prism 9 (GraphPad Software Inc., San Diego, CA). IC50 doses were determined using a non-linear regression curve fit.
The AFM nanoindentation data were presented in the form of box plots. Each point in box-plot graphs represents the mean value of elasticity modulus (E) or glycocalyx thickness (T) calculated for a single cell. Statistical significance was tested using a one-way analysis of variance ANOVA (p < 0.05) followed by post-hoc Tukey's range tests. All statistical analyses and graphs were prepared in OriginPro 2022 (Academic).
3. Results
3.1. PAMPS75-b-PAaUn and P(AMPS50-co-AaU50) copolymers inhibit SARS-CoV-2 replication in vitro
The antiviral activity of the PAMPS75-b-PAaUn and P(AMPS50-co-AaU50) copolymers was verified against highly pathogenic SARS-CoV-2 in vitro using the Vero cell line. First, the compounds' cytotoxic effect was assessed. The results are presented in Table 1. Second, the initial evaluation of the antiviral activity was performed using the CPE-reduction assay, and the results are presented in Fig. 1. Most of the compounds tested showed antiviral activity. In the case of block copolymers, the efficacy seemed to correlate with the PAaU block length.
Table 1.
Antiviral efficacy of tested copolymers. The table shows CC50, IC50, and SI obtained for the SARS-CoV-2 and Vero cells. *data adopted from the previous study [24].
| Polymers | CC50* μg/ml (μM) |
IC50 μg/ml (μM) |
SI |
|---|---|---|---|
| PAMPS75-b-PAaU39 | 2.4 × 104 (8.9 × 101) | 1.9 × 10−2 (7.1 × 10−4) | 1.3 × 105 |
| PAMPS75-b-PAaU28 | 2.4 × 104 (1 × 102) | 5.4 × 10−2 (2.3 × 10−3) | 4.5 × 104 |
| PAMPS75-b-PAaU12 | 2.9 × 104 (1.5 × 102) | 2.1 × 10−1 (1.1 × 10−2) | 1.4 × 104 |
| PAMPS75-b-PAaU3 | 4.1 × 104 (2.5 × 102) | 1.1 (6.8 × 10−2) | 3.6 × 103 |
| P(AMPS50-co-AaU50) | 3.2 × 104 (1.3 × 102) | 1.8 × 10−1 (7.4 × 10−3) | 1.8 × 104 |
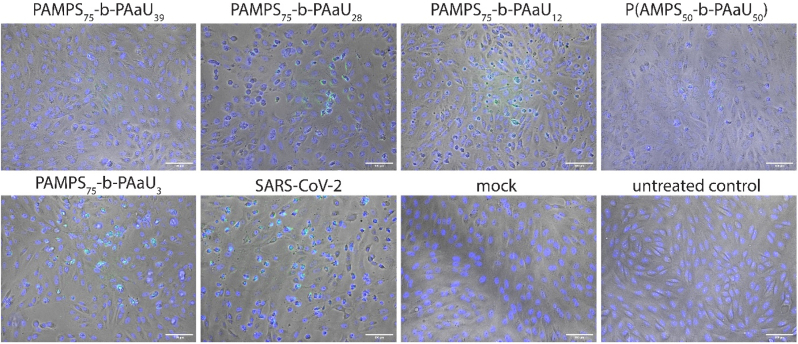
Fig. 1.
Polymers inhibit the cytopathic effect. CPE effect in Vero cells infected with SARS-CoV-2 at 1600 TCID50/ml 48 h post-infection in the presence or absence of 25 μg/ml of polymers. The live cell staining was conducted with Blue/Green Cell Viability Imaging Kit. The blue color denotes the nuclei of all cells, while the green color denotes only the nuclei of cells with compromised plasma membrane integrity. Light microscope images were collected on the EVOS Floid Imaging system using 20× objective. Scale bar 100 μm. (For interpretation of the references to color in this figure legend, the reader is referred to the Web version of this article.)
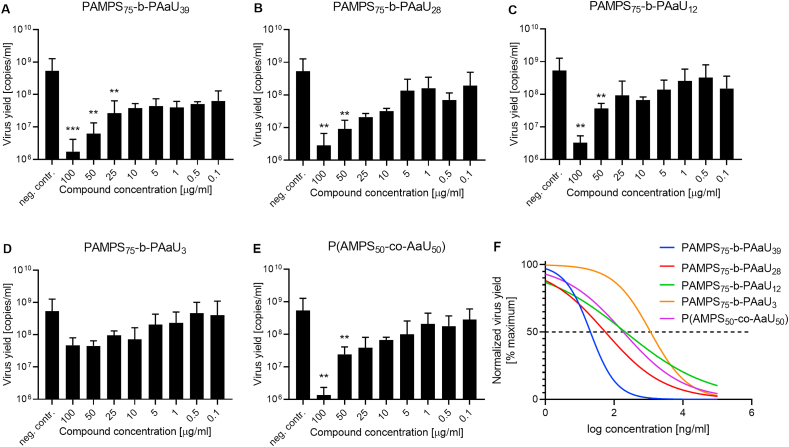
To validate the results, the virus replication was evaluated by means of quantitative PCR coupled with reverse transcription (RT-qPCR) analysis of the cell culture supernatants (Fig. 2). All selected copolymers hampered coronavirus replication, and the antiviral effect correlated with the length of the PAaU chain. No inhibition was recorded in the tested concentration range for the copolymer with the shortest PAaU chain, i.e., PAMPS75-b-PAaU3 (Fig. 2D). The other polymers showed statistically significant inhibitory properties at 50 and 100 μg/ml, with PAMPS75-b-PAaU39 additionally at 25 μg/ml. For the copolymer with the longest chain, PAMPS75-b-PAaU39, sub-nanomolar IC50 was recorded (1.9 × 10−2 μg/ml, 7.1 × 10−4 μM); Table 1). Based on the CC50 and IC50, the selectivity index (SI) values were calculated. The SI of all tested compounds was high, particularly for PMAPS75-b-PAaU39, for which the SI exceeded 1.3 × 105. The random polymer (in which monomers are not organized in blocks, but instead stochastically distributed) P(AMPS50-co-AaU50) showed intermediate antiviral properties with IC50 of 1.8 × 10−1 μg/ml and SI of 1.8 × 104 (Table 1). This suggests that the block structure enhances the antiviral properties of the copolymers.
Fig. 2.
The copolymers are potent SARS-CoV-2 inhibitors. The figure shows RT-qPCR analysis of supernatants collected from cells infected with SARS-CoV-2 at 1600 TCID50/ml 48 h post-infection in the absence (neg.contr.) or presence of compounds in the range of concentrations 0.1–100 μg/ml (A–E) and the dose-response curves of normalized data (F). All experiments were performed in at least two biological repetitions, each in triplicate. The results are presented as average values with standard deviations (error bars). Values statistically significant are indicated by asterisks: **p < 0.05, ***p < 0.005.
3.2. PAMPS-b-PAaU and P(AMPS50-co-AaU50) effectively inhibit the replication of low pathogenic coronaviruses
The most effective block copolymer PAMPS75-b-PAaU39 and random copolymer P(AMPS50-co-AaU50) were selected for further study. Two other coronaviruses representing the alphacoronavirus (HCoV-229E) and betacoronavirus (HCoV-OC43) evolutionary branches were used. In vitro assays were carried out using the MRC-5 cells for the HCoV-229E virus and HCT-8 cell line for the HCoV-OC43 virus. Copolymers showed marked toxicity at 2 mg/ml (Fig. 3). In the replication reduction tests, both compounds displayed properties similar to those recorded for SARS-CoV-2. The RT-qPCR results showed the effect of PAMPS75-b-PAaU39 and P(AMPS50-co-AaU50) at submicromolar concentrations (Fig. 4, Table 2). In both cases, the block copolymer PAMPS75-b-PAaU39 performed better than the random copolymer P(AMPS50-co-AaU50) (Fig. 4A and B). The IC50 values of block copolymer were 698 ng/ml and 77 ng/ml against HCoV-OC43 and HCoV-229E, respectively (Table 2).
Fig. 3.
PAMPS-PAaU copolymers are not toxic to MRC-5 and HCT-8 cells in the low μg/ml concentration range. Results of the XTT assay for PAMPS75-b-PAaU39 and P(AMPS50-co-AaU50) copolymers at concentrations of 4000, 3000, 2000, 1000, 500, and 100, 50, and 10 μg/ml on MRC-5 and HCT-8 cells. All experiments were performed in duplicate. Average values with standard deviation (error bars) are shown.
Fig. 4.
The PAMPS75-b-PAaU39block and P(AMPS50-co-AaU50) random copolymers as broad-spectrum antivirals. The figure shows RT-qPCR analysis of cell culture supernatants collected from cells infected with HCoV-OC43 at 1600 TCID50/ml 72 h post-infection (A, C) and HCoV-229E at 4000 TCID50/ml 72 h post-infection (B, D), neg. contr. denotes cells infected without polymer. All experiments were performed in at least three biological repetitions, each in triplicate. The results are presented as average values with standard deviations (error bars). *Significantly different from the control (p < 0.05).
Table 2.
Antiviral efficacy of tested compounds for LPHCV.
| Polymer | HCT-8 |
HCoV-OC43 |
SI | MRC-5 |
HCoV-229E |
SI |
|---|---|---|---|---|---|---|
| CC50 |
IC50 |
CC50 |
IC50 |
|||
| μg/ml (μM) | μg/ml (μM) | μg/ml (μM) | μg/ml (μM) | |||
| PAMPS75-b-PAaU39 | 3.1 × 103 | 7.0 × 10−1 | 4.4 × 103 | 1.8 × 103 | 7.7 × 10−2 | 2.3 × 104 |
| (1.2 × 102) | (2.6 × 10−2) | (6.6 × 101) | (3 × 10−3) | |||
| P(AMPS50-co-AaU50) | 3.5 × 103 | 7.8 × 10−1 | 4.4 × 103 | 2.8 × 103 | 1.1 × 10−1 | 2.7 × 104 |
| (1.4 × 102) | (3.2 × 10−2) | (1.2 × 102) | (4 × 10−3) |
3.3. Copolymers inhibit SARS-CoV-2 infection in HAE cultures
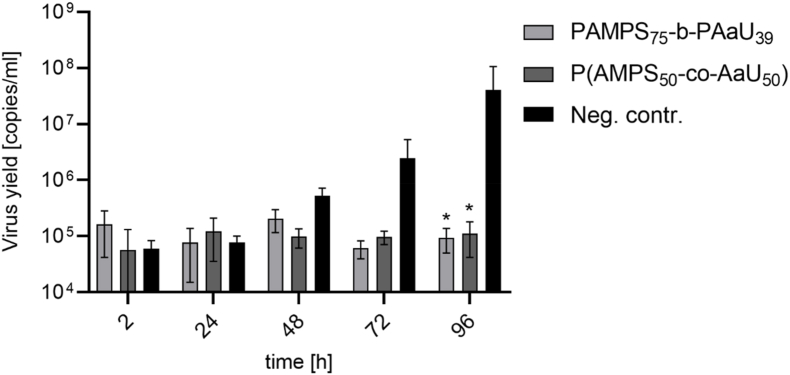
The antiviral activity of copolymers was further validated using fully differentiated 3D cultures of HAE infected with SARS-CoV-2. Cultures were infected from the apical side in the presence of PAMPS75-b-PAaU39 or P(AMPS50-co-AaU50), at 100 μg/ml; H2O was used as a control. Both compounds were used at higher concentrations, exceeding their IC90 concentration, and showed similar antiviral properties (Fig. 5).
Fig. 5.
Copolymers block SARS-CoV-2 replication in ex vivo model. The graph presents the number of viral RNA copies collected apically after the indicated time. HAE culture was infected with SARS-CoV-2 at 5000 TCID50/ml until 96 h post-infection, neg. contr. denotes cells infected without polymer. Each experiment was performed in 2 biological repetitions, each in triplicate. The results are presented as average values with standard deviations (error bars). *Significantly different from the control (p < 0.05).
3.4. PAMPS-b-PAaUs interfere with virus attachment to the host cells and influence the late stage of virus infection
The compound PAMPS75-b-PAaU39 which showed the best antiviral properties was selected for further research. First, a virus inactivation assay was performed to see if the compound could bind to the virus particle, thus blocking its attachment. The virus stock and polymer were incubated at RT for 1 h and then diluted to a concentration of 1 μg/ml, at which the compounds showed no antiviral properties. Twenty-four hours after infection, cells were fixed and labeled with SARS-CoV-2 protein N. No differences were observed between cells infected in the presence of the polymer and untreated control (Fig. 6A). Then, an attachment assay was performed to see whether the compounds affected this stage. To study this, precooled cells were incubated with the virus stock together with the polymer at 4 °C, to see whether polymers are able to block the attachment of virus particles to cellular receptors. Subsequently, cells were fixed and SARS-CoV-2 N protein was visualized. The significantly lower virus adhesion to the cell surface for cells pre-treated with the polymers was observed (Fig. 6B). Next, the effect of inhibitors on virus internalization to permissive cells was tested in the virus entry assay. For this, cells were preincubated with the virus at 4 °C, unattached particles were washed out, and PAMPS75-b-PAaU39 was added to the cells and incubated for another 10 min at 4 °C, followed by 1 h incubation at 37 °C, and then fixation. The internalized virus particles on confocal images were counted, and the quantitative results are shown in Fig. 6C and D. We observed a twofold reduction in the number of viral particles adhered to the cell surface after the PAMPS75-b-PAaU39 treatment. However, this did not translate to the number of internalized particles suggesting that the adhesion inhibition mechanism did not directly affect the virus entry, and its relevance may be limited. Finally, the effect on the late phases of virus replication was tested. To do this, cells were infected with the virus for 2 h at 37 °C, washed, and overlaid with media supplemented with the polymer. Twenty-four hours after infection, the cells were fixed. For cells to which PAMPS75-b-PAaU39 was added after infection, a significant reduction in the number of infected cells and a different distribution of infection were observed (Fig. 6E). In control, infected individual cells were primarily observed in large infection foci of infected cells suggesting a cell-to-cell transmission. In contrast, in polymer-treated samples, only isolated infected cells were noted (Fig. 6E). The schematic representation of the assays described above is shown in Supplementary Fig. 1.
Fig. 6.
PAMPS-b-PAaUs inhibit the SARS-CoV-2 attachment to host cells and the replication cycle. Antiviral activity of copolymers was tested in Vero cells infected with the SARS-CoV-2, as described in the Material and Method section: virus inactivation assay (A), virus attachment assay (B), virus entry assay (C) with the graph showing the number of SARS-CoV-2 virions attached and entered to the cells with line at median with 95% CI (D), and virus replication, assembly and egress assay (E). F-actin – red; SARS-CoV-2 N protein – green; cell nuclei – blue. Scale bar = 25 μm. (For interpretation of the references to color in this figure legend, the reader is referred to the Web version of this article.)
3.5. PAMPS75-b-PAaUn copolymers enter the cell but are rapidly removed
In previous work, we proposed that the copolymers mainly act in the early stages of ZIKV replication by influencing host cells but do not affect the virus itself [24]. Also, in the case of the SARS-CoV-2 virus, we confirmed the impact of compounds on the attachment and late stages of replication (Fig. 6). The activity at late stages requires the ability of the polymer to reach the replication or egress site, and therefore we explored this topic.
The fluorescently labeled copolymer PAMPS75-b-PAaU28-FL was used to study the ability of copolymers to penetrate living cells; fluorescein (FL) was used as a control. In line with previous experiments, we observed that copolymers could quickly (<2 min) and efficiently invade the cell but are also rapidly removed (∼30 min) (Fig. 7 and Supplementary Video 1). In the second minute after the administration of PAMPS75-b-PAaU28-FL, film recording was started when the compound was already inside the cells. Over the next minutes, the compound was gradually removed from the cell's bodies. After about 30 min, the signal in the cells was no longer noticeable, while the background signal in the extracellular space increased. Fluorescein penetrated the cells to a lower extent (Fig. 7). It is worth emphasizing that the observed phenomenon of signal decrease inside the cell is not related to the bleaching of the fluorescent dye but clearly results from the removal of PAMPS75-b-PAaU28-FL from the cell (Supplementary Video 1).
Fig. 7.
The PAMPS75-b-PAaU28quickly penetrates the cells. The figure shows the signal from fluorescein attached to PAMPS75-b-PAaU28 (upper panel) and fluorescein (lower panel) administrated to the cells. The signal distribution was observed for up to 48 min. The results are presented as maximum projections from chosen time points: 2, 16, and 30 min. Scale bar = 20 μm.
3.6. The PAMPS75-b-PAaU copolymers change cell elasticity and reduce glycocalyx layer
Given the ability of copolymers to quickly enter and exit cells, we investigated their effect on the mechanical properties of cells. We assumed that PAMPS75-b-PAaU39 directly affects cellular elasticity by modification of the glycocalyx layer associated with cytoskeleton structure.
In the first step, measurements were made on unfixed cells incubated directly with the copolymer (Fig. 8 A-C). By using the AFM nanoindentation method with the spherical probe, we have proved that PAMPS75-b-PAaU39 copolymer modified the elasticity of the whole cell, and the most significant changes were observed after 3 h incubation (Fig. 8A). If we focus on the cortical part of the cell, then for this layer we observe an increase in stiffness after 15 min of incubation and it is maintained for subsequent incubation times. Interestingly, the increase of the cortical stiffness of cell (Fig. 8B) is accompanied by a reduction in the thickness of the glycocalyx layer (Fig. 8C). In the next step, using the QI-AFM method, spatial maps were measured showing the topography of the cell (height, Fig. 8D - top row) correlated with the spatial distribution of the elastic modulus (elasticity, Fig. 8D - bottom row). All images shown in Fig. 8D depict the central part of cell detailing location of the cell nucleus surrounded by the cytoskeleton fibers. After 2 h and 3 h incubation with copolymer (Fig. 8D - bottom row), the elasticity maps depict the stiff well recognized cytoskeleton fibers. This indicates an increase of cellular stiffness correlated with the cytoskeleton fibers polymerization.
Fig. 8.
Changes of the nano-mechanical parameters of Vero cells induced by PAMPS75-b-PAaU39. Box plots depict the quantitative analysis of elastic moduli counted for the entire range of indentation corresponding to the whole cell (A) and a specific range of nanoindentation corresponding to the elasticity of the cortex of the non-treated cells (control) or cells incubated with PAMPS75-b-PAaU39 for 15 min, 1 h, 2 h, 3 h, 4 h and 6 h (B). Box plots depict the quantitative analysis of thickness of the glycocalyx layer for non-treated cells (control) and cells incubated with PAMPS75-b-PAaU39 for 15 min, 1 h, 2 h, 3 h, 4 h, and 6 h (C). The representative AFM maps of height (topography, top row) and elasticity (bottom row) were taken for Vero cells incubated with PAMPS75-b-PAaU39 for 15 min, 1 h, 2 h, 3 h, 4 h, and 6 h. Non-treated cells were used as a reference (control). All images were taken for 0.5% GA fixed cells (D). Statistics: p values were determined by one-way analysis of variance ANOVA followed by Tukey's post-hoc test. *Significantly different from the control (p < 0.05).
3.7. PAMPS-b-PAaU inhibits viral replication through cellular cytoskeleton modification
The changes in cell elasticity observed with AFM can be explained by cytoskeleton rearrangement. To verify this hypothesis, cells were treated with 100 μg/ml PAMPS75-b-PAaU39 for 15 min, 1 h, 2 h, 3 h, 4 h, or 6 h. Then, cytoskeletal proteins such as actin, vimentin, and vinculin were immunofluorescently labeled and visualized with confocal microscopy (Fig. 9). The change in organization was observed for all immunolabeled proteins. The copolymers induced a marked reduction in the organization of the actin filaments 1–4 h p.i. (Supplementary Fig. 2), which may be the main reason for the rise of stiffness observed in the AFM analysis. In the case of vimentin, a destabilization of its organization was also observed, starting from 15 min (Fig. 9B), which resulted in the diffusion of fluorescence signal from its original location at the nuclear periphery into the cytoplasm after 2 h (Fig. 9C) and back to the nucleus perimeter after (Fig. 9D). In turn, in the case of vinculin, which is responsible for, among others, focal adhesion sites and anchoring F-actin to the membrane, a difference in distribution and reduction in its clusters in the form of green dots was observed at 2 h p.i. (Fig. 9C). The images showed that the cell cytoskeleton returned to the normal state, the same as that of untreated cells, confirming that the effect of the copolymers is reversible (Fig. 9D). Such effects of polymers on the cell cytoskeleton can undoubtedly translate into the inhibition of viral infection in cells. This is similar to the other work showing a correlation between the disruption of actin and other cytoskeletal elements by cytochalasin D or jasplakinolide on viral replication [32,33]. That also confirms the importance of the cell cytoskeleton in viral infection [[34], [35], [36], [37]].
Fig. 9.
PAMPS-b-PAaU reversibly affects cell skeleton. Confocal images of Vero cells non-treated (A), treated with 100 μg/ml PAMPS75-b-PAaU39 at 15 min (B), 2 h (C), and 6 h (D) F-actin – red; Vimentin – white; Vinculin – green; cell nuclei – blue. Scale bar = 10 μm. (For interpretation of the references to color in this figure legend, the reader is referred to the Web version of this article.)
4. Discussion
The COVID-19 pandemic, incited by the SARS-CoV-2 virus, has inflicted extensive damage on global public health, despite the unprecedented expeditious deployment of vaccinations. Effective antivirals were introduced only at the beginning of 2022, a full two years post-pandemic onset. Consequently, there is a pressing need to identify and develop broad-spectrum antiviral agents that can be rapidly implemented and deployed in response to future epidemic threats.
In the present study, we describe our explorative study on PAMPS-b-PAaU block copolymers as potent inhibitors of human coronaviruses and members of the Flaviviridae family. Our in vitro investigations demonstrated that the polymers effectively obstruct the progression of infection, with the most efficacious copolymer exhibiting an IC50 value of 1.9 × 10−2 μg/ml, corresponding to 7.1 × 10−4 μM. Moreover, these copolymers exhibit minimal cytotoxicity, with a CC50 > 2000 μg/ml. Importantly, our findings also reveal that the evaluated compounds impede SARS-CoV-2 replication in human airway epithelial cultures, which accurately represent the human bronchial milieu.
The glycocalyx plays an important role in the non-specific attachment of the virus to cells [38]. The reduction of the glycocalyx caused by the copolymers, proven in AFM measurements, could be the first point of antiviral action of PAMPS75-b-PAaUn. Loss of glycocalyx layer contributes to limited virus attachment and therefore to the protection of the epithelial cells.
Heparan sulfate proteoglycans (HSPGs), integral constituents of the glycocalyx, play an essential role in facilitating interactions with a variety of ligands [39,40]. They enable viral attachment to cellular surfaces, an initial stage of infection that has been reported for coronaviruses such as HCoV-NL63, SARS-CoV, and SARS-CoV-2 [[41], [42], [43], [44], [45], [46], [47], [48]]. Moreover, the glycocalyx molecules are associated with the cytoskeleton through their membrane domains [49]. Therefore, modification of the glycocalyx layer induces the reorganization of the cellular cytoskeleton [50,51]. In this paper, we have shown that the reduction of cellular glycocalyx results in the increase in stiffness of the cortical layer of epithelial cells treated with PAMPS75-b-PAaU39. Interestingly, despite the cell surface modification persisting throughout all periods of incubation, the impact on global cellular elasticity is temporary, up to 3 h, and after this time the cell elasticity returns to the normal level. This effect can be attributed to the removal of the compound from the cell, as shown in Fig. 7 and Supplementary Video 1. However, at the time of administration and penetration into the cell, the compound has a strong effect on the cellular cytoskeleton, causing a significant change in the elasticity of the entire cell. Actin acts as a cellular scaffold but also plays a critical role in numerous cellular processes, including phagocytosis, intercellular communication, organelle distribution, and cellular motility [31]. Notably, it is integral to every phase of viral infection, such as viral entry, replication, and egress [34,52]. Consequently, viral infections remodel cytoskeletal arrangements [53], inducing alterations in cellular mechanical properties, but may also be affected by any alterations [54]. To give an example, the actin cortex beneath the plasma membrane may function as a barrier for viral entry or exit [55]. Vinculin, a pivotal protein for cell adhesion, can bind to actin and is a constituent of focal adhesions and adherens junctions [56,57]. Studies have shown that overexpression of vinculin and talin-1, which interact and localize to focal adhesion, inhibits infection with pseudotyped human immunodeficiency virus type 1 (HIV-1) and Moloney murine leukemia virus. In contrast, the use of siRNA targeting these proteins increases infection [58]. Another important element of the cytoskeleton involved in viral infection is vimentin. Vimentin is a major intermediate filament responsible for many functions, such as signaling and intracellular organization, cell migration, and adhesion [59,60]. During viral infection, vimentin may have several functions and play a role as an important factor [34,61]. Extracellular vimentin can serve as a co-receptor to facilitate entry into the cell; in the downstream stages of infection, vimentin can facilitate the movement of the virus deep into the cell to replication sites or facilitate replication itself by providing the necessary elements for replication and assembly of progeny virions [[62], [63], [64]]. For example, reducing the amount or knockdown of extracellular vimentin decreased the number of Japanese encephalitis viral particles that could bind to the cell surface [65]. In addition, vimentin-NS4A interaction was observed during DENV replication, as well as its reorganization, and its role in replication complex formation was proven. Interestingly, the silencing of vimentin was found to cause significant changes in the distribution of the replication complex in infected cells. This finding strengthens the concept that the intact state of the vimentin scaffold is essential for DENV replication [66]. In summary, all the cell cytoskeleton elements we studied may be affected by viral infection. In the present study, we demonstrated that polymers modify the normal structure of the cytoskeleton and, thus, their function. Notably, the observed changes decreased over time and did not affect cell death rate.
The literature offers an example of another compound, brilacidin, which also has a dual mechanism of action. It affects the interaction of coronaviruses with HSPG and hampers early stages of the infection [67]. Antiviral polymers primarily inhibit virus-cell interactions through direct binding to viral particles or the cell surface receptors, consequently obstructing viral entry [17,18,68,69]. Previous studies on HIV-1 also suggest that PAMPS polymers can efficiently inhibit virus adsorption [70]. This aligns with our previous findings on Zika virus and the current study on SARS-CoV-2 [24]. However, this study describes the copolymers that traverse cellular membranes without inducing cytotoxic effects. Their antiviral mechanism, at first glance, is based on inhibiting viral adherence to the susceptible cells. However, a deeper analysis of this phenomenon shows that it does not significantly alter viral entry or infection. The functional studies showed that the polymers are active at later stages of the infection. The assessment of potential mechanisms shows that while infected cells replicate the virus and produce viral particles, the virus transmission between cells is affected. Interestingly, the inhibition correlated with the altered cellular plasticity and cytoskeleton rearrangements. This may suggest that the polymers may block the production of infectious viral particles and cell-to-cell transfer in a non-specific manner. Having said that, we would like to underline that all the cellular alterations have diminished over time, and we did not observe tissue remodeling in the HAE tissues. While PAMPS-PAaUs may raise controversies regarding safety in their current form, it is an exciting and relatively unexplored direction in antiviral drug development.
Our results show that the best antiviral properties were shown by copolymers containing the longest PAaU block, creating nanoparticles with the largest size. PAMPS75-b-PAaU39 and P(AMPS50-co-AaU50) have a hydrodynamic radius of about 12 and 8 nm, respectively, while the others are below 5 nm [24]. Research on cell-internalizing nanoparticles shows that their size is important in terms of their bioavailability. This suggests that the properties of PAMPS-PAaU copolymers can be improved by increasing their diameter. However, this may result in changes in the exocytosis rate of the copolymers from the cells, as smaller nanoparticles are removed from cells more efficiently, and thus affect their cytotoxicity [[71], [72], [73]].
In evaluating the antiviral potential of polymers, the potential adverse effects of these compounds, particularly their impact on blood clotting, need to be taken into account given their resemblance to HSPGs. It is, however, worth mentioning that the polymers may also exhibit advantageous properties such as anticoagulation, anti-inflammatory effects, modulation of angiogenesis, and anti-tumor metastasis [40,74]. Despite the observed influence of these compounds on cytoskeletal rearrangement, we did not discern any other effects indicative of toxicity. No notable morphological changes of the treated cells were observable and the assays studying the potential toxicity of polymers reveled a very promising safety profile. Notably, the alterations of the cytoskeleton were transient, with cells reverting to their initial state over time.
Antiviral polymers often inhibit multiple viruses simultaneously. Thus, developing broad-spectrum antiviral polymers has excellent potential to combat current viral diseases and as part of a preparedness strategy for future threats from emerging viruses. As the tested materials inhibit the development of infections of both Coronaviruses and Flaviviruses, and earlier reports showed an anti-HIV effect, it seems reasonable to investigate their potential further.
Credit author statement
Agnieszka Dabrowska: Conceptualization, Methodology, Investigation, Validation, Formal analysis, Writing - Original Draft, Visualization; Pawel Botwina: Conceptualization, Methodology, Investigation, Formal analysis, Writing - Original Draft, Visualization; Emilia Barreto-Duran: Methodology, Investigation; Agata Kubisiak: Methodology, Investigation, Visualization; Magdalena Obloza: Methodology, Investigation Aleksandra Synowiec: Methodology, Investigation, Visualization; Artur Szczepanski: Methodology, Investigation, Visualization, Formal analysis; Marta Targosz-Korecka: Methodology, Investigation, Resources, Visualization; Krzysztof Szczubialka: Conceptualization, Resources, Writing - Review & Editing, Supervision, Project administration; Maria Nowakowska: Conceptualization, Resources, Writing - Review & Editing, Supervision, Project administration, Funding acquisition; Krzysztof Pyrc: Conceptualization, Writing - Review & Editing, Supervision, Project administration, Funding acquisition.
Funding
This work was supported by the National Science Centre, Poland, in the form of Grants No. 2017/27/B/ST5/01108 to M.N. and 2016/21/B/NZ6/01307 to K.P. A.S.'s work was supported by the Foundation for Polish Science (FNP).
Declaration of competing interest
The authors declare the following financial interests/personal relationships which may be considered as potential competing interests Krzysztof Pyrc reports financial support was provided by National Science Centre Poland. Maria Nowakowska reports financial support was provided by National Science Centre Poland.
Acknowledgment
We would like to thank Haider Ali and Malgorzata Bzowska for their support in the flow cytometry experiments. Graphical abstract and Supplementary Fig. 1 were created with BioRender.com.
Footnotes
www: http://virogenetics.info/
Supplementary data to this article can be found online at https://doi.org/10.1016/j.mtbio.2023.100763.
Appendix A. Supplementary data
The following are the Supplementary data to this article:
Data availability
No data was used for the research described in the article.
References
- 1.Zhu N., Zhang D., Wang W., Li X., Yang B., Song J., et al. A novel coronavirus from patients with pneumonia in China. N. Engl. J. Med. 2020 doi: 10.1056/nejmoa2001017. 2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Yang J., Chen X., Deng X., Chen Z., Gong H., Yan H., et al. Disease burden and clinical severity of the first pandemic wave of COVID-19 in Wuhan, China. Nat. Commun. 2020 doi: 10.1038/s41467-020-19238-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Brinkworth J.F., Rusen R.M. SARS-CoV-2 is not special, but the pandemic is: the ecology, evolution, policy, and future of the deadliest pandemic in living memory. Annu. Rev. Anthropol. 2022;51:527–548. doi: 10.1146/annurev-anthro-041420-100047. [DOI] [Google Scholar]
- 4.Collaborators E.M. Estimating excess mortality due to the COVID-19 pandemic: a systematic analysis of COVID-19-related mortality, 2020-21. Lancet (London, England) 2022;399:1513–1536. doi: 10.1016/S0140-6736(21)02796-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Fields B.N. Fields virology. Fields Virol. 2013:2664. [Google Scholar]
- 6.Payne S. 2017. Family Coronaviridae. Viruses. [DOI] [Google Scholar]
- 7.Swelum A.A., Shafi M.E., Albaqami N.M., El-Saadony M.T., Elsify A., Abdo M., et al. COVID-19 in human, animal, and environment: a review. Front. Vet. Sci. 2020 doi: 10.3389/fvets.2020.00578. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Compton S.R. Overview of coronaviruses in veterinary medicine. Comp. Med. 2021;71:333–341. doi: 10.30802/AALAS-CM-21-000007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ginsberg H.S., Goebel W.F., Horsfall F.L. The inhibitory effect of polysaccharide on mumps virus multiplication. J. Exp. Med. 1948 doi: 10.1084/jem.87.5.385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Takemoto K.K., Liebhaber H. Virus-polysaccharide interactions. I. An agar polysaccharide determining plaque morphology of EMC virus. Virology. 1961 doi: 10.1016/0042-6822(61)90338-5. [DOI] [PubMed] [Google Scholar]
- 11.Nahmias A.J., Kibrick S. Inhibitory effect of heparin on herpes simplex virus. J. Bacteriol. 1964;87:1060–1066. doi: 10.1128/jb.87.5.1060-1066.1964. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.De Somer P., De Clercq E., Billiau A., Schonne E., Claesen M. Antiviral activity of polyacrylic and polymethacrylic acids. I. Mode of action in vitro. J. Virol. 1968;2:878–885. doi: 10.1128/jvi.2.9.878-885.1968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Liebhaber H., Takemoto K.K. Alteration of plaque morphology of EMC virus with polycations. Virology. 1961 doi: 10.1016/0042-6822(61)90349-X. [DOI] [PubMed] [Google Scholar]
- 14.Bianculli R.H., Mase J.D., Schulz M.D. Antiviral polymers: past approaches and future possibilities. Macromolecules. 2020 doi: 10.1021/acs.macromol.0c01273. [DOI] [Google Scholar]
- 15.Pirrone V., Wigdahl B., Krebs F.C. The rise and fall of polyanionic inhibitors of the human immunodeficiency virus type 1. Antivir. Res. 2011;90:168–182. doi: 10.1016/j.antiviral.2011.03.176. [DOI] [PubMed] [Google Scholar]
- 16.Zmonarski S., Stojanowski J., Zmonarska J. Polymers with antiviral properties: a brief review. Polim. Med. 2020 doi: 10.17219/pim/131643. [DOI] [PubMed] [Google Scholar]
- 17.Pyrć K., Milewska A., Duran E.B., Botwina P., Lopes R., Arenas-Pinto A., et al. SARS-CoV-2 inhibition in human airway epithelial cells using a mucoadhesive, amphiphilic chitosan that may serve as an anti-viral nasal spray. bioRxiv. 2020 doi: 10.1101/2020.12.10.413609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Milewska A., Kaminski K., Ciejka J., Kosowicz K., Zeglen S., Wojarski J., et al. HTCC: broad range inhibitor of coronavirus entry. PLoS One. 2016 doi: 10.1371/journal.pone.0156552. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Ciejka J., Wolski K., Nowakowska M., Pyrc K., Szczubiałka K. Biopolymeric nano/microspheres for selective and reversible adsorption of coronaviruses. Mater. Sci. Eng., C. 2017;76:735–742. doi: 10.1016/j.msec.2017.03.047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ciejka J., Milewska A., Wytrwal M., Wojarski J., Golda A., Ochman M., et al. Novel polyanions inhibiting replication of influenza viruses. Antimicrob. Agents Chemother. 2016 doi: 10.1128/AAC.02183-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Ciejka J., Botwina P., Nowakowska M., Szczubiałka K., Pyrc K. Synthetic sulfonated derivatives of poly(allylamine hydrochloride) as inhibitors of human metapneumovirus. PLoS One. 2019;14 doi: 10.1371/journal.pone.0214646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Synowiec A., Gryniuk I., Pachota M., Strzelec Ł., Roman O., Kłysik-Trzciańska K., et al. Cat flu: broad spectrum polymeric antivirals. Antivir. Res. 2019 doi: 10.1016/j.antiviral.2019.104563. [DOI] [PubMed] [Google Scholar]
- 23.Serrano-Aroca Á., Ferrandis-Montesinos M., Wang R. Antiviral properties of alginate-based biomaterials: promising antiviral agents against SARS-CoV-2. ACS Appl. Bio Mater. 2021;4(8):5897–5907. doi: 10.1021/acsabm.1c00523. [DOI] [PubMed] [Google Scholar]
- 24.Botwina P., Obłoza M., Zatorska-Płachta M., Kamiński K., Mizusaki M., Yusa S.-I., et al. Self-organized nanoparticles of random and block copolymers of sodium 2-(Acrylamido)-2-methyl-1-propanesulfonate and sodium 11-(acrylamido)undecanoate as safe and effective Zika virus inhibitors. Pharmaceutics. 2022;14 doi: 10.3390/pharmaceutics14020309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Mizusaki M., Shimada Y., Morishima Y., Yusa S.I.S.-I. PH-responsive intra- and inter-molecularly micelle formation of anionic diblock copolymer in water. Polymers (Basel) 2016;8:1–11. doi: 10.3390/polym8020056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Pachota M., Grzywa R., Iwanejko J., Synowiec A., Iwan D., Kamińska K., et al. Novel inhibitors of HSV-1 protease effective in vitro and in vivo. Antivir. Res. 2023;213 doi: 10.1016/j.antiviral.2023.105604. [DOI] [PubMed] [Google Scholar]
- 27.Napolitano V., Dabrowska A., Schorpp K., Mourão A., Barreto-Duran E., Benedyk M., et al. Acriflavine, a clinically approved drug, inhibits SARS-CoV-2 and other betacoronaviruses. Cell Chem. Biol. 2022 doi: 10.1016/j.chembiol.2021.11.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Owczarek K., Szczepanski A., Milewska A., Baster Z., Rajfur Z., Sarna M., et al. Early events during human coronavirus OC43 entry to the cell. Sci. Rep. 2018 doi: 10.1038/s41598-018-25640-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Schindelin J., Arganda-Carreras I., Frise E., Kaynig V., Longair M., Pietzsch T., et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods. 2012 doi: 10.1038/nmeth.2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Giergiel M., Malek-Zietek K.E., Konior J., Targosz-Korecka M. Endothelial glycocalyx detection and characterization by means of atomic force spectroscopy: comparison of various data analysis approaches. Micron. 2021 doi: 10.1016/j.micron.2021.103153. [DOI] [PubMed] [Google Scholar]
- 31.Pollard T.D., Cooper J.A. Actin, a central player in cell shape and movement. Science. 2009 doi: 10.1126/science.1175862. 80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Dietzel E., Kolesnikova L., Maisner A. Actin filaments disruption and stabilization affect measles virus maturation by different mechanisms. Virol. J. 2013 doi: 10.1186/1743-422X-10-249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Wang J.L., Zhang J.L., Chen W., Xu X.F., Gao N., Fan D.Y., et al. Roles of small GTPase RAc1 in the regulation of actin cytoskeleton during dengue virus infection. PLoS Neglected Trop. Dis. 2010 doi: 10.1371/journal.pntd.0000809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Wen Z., Zhang Y., Lin Z., Shi K., Jiu Y. Cytoskeleton - a crucial key in host cell for coronavirus infection. J. Mol. Cell Biol. 2020 doi: 10.1093/jmcb/mjaa042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Gudheti M.V., Curthoys N.M., Gould T.J., Kim D., Gunewardene M.S., Gabor K.A., et al. Actin mediates the nanoscale membrane organization of the clustered membrane protein influenza hemagglutinin. Biophys. J. 2013;104:2182–2192. doi: 10.1016/j.bpj.2013.03.054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Döhner K., Sodeik B. The role of the cytoskeleton during viral infection. Curr. Top. Microbiol. Immunol. 2004 doi: 10.1007/3-540-26764-6_3. [DOI] [PubMed] [Google Scholar]
- 37.Zhang Y., Zhao S., Li Y., Feng F., Li M., Xue Y., et al. Host cytoskeletal vimentin serves as a structural organizer and an RNA-binding protein regulator to facilitate Zika viral replication. Proc. Natl. Acad. Sci. U.S.A. 2022 doi: 10.1073/pnas.2113909119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Naskalska A., Dabrowska A., Szczepanski A., Milewska A., Jasik K.P., Pyrc K. Membrane protein of human coronavirus NL63 is responsible for interaction with the adhesion receptor. J. Virol. 2019 doi: 10.1128/jvi.00355-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Sarrazin S., Lamanna W.C., Esko J.D. Heparan sulfate proteoglycans. Cold Spring Harbor Perspect. Biol. 2011 Jul 1;3(7) doi: 10.1101/cshperspect.a004952. PMID: 21690215; PMCID: PMC3119907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Paluck S.J., Nguyen T.H., Maynard H.D. Heparin-mimicking polymers: synthesis and biological applications. Biomacromolecules. 2016 Nov 14;17(11):3417–3440. doi: 10.1021/acs.biomac.6b01147. Epub 2016 Oct 14. PMID: 27739666; PMCID: PMC5111123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Paluck Samantha J., Nguyen Thi H., Maynard Heather D. Heparin-mimicking polymers: synthesis and biological applications. Biomacromolecules. 2016;17(11):3417–3440. doi: 10.1021/acs.biomac.6b01147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Connell B.J., Lortat-Jacob H. Human immunodeficiency virus and heparan sulfate: from attachment to entry inhibition. Front. Immunol. 2013 Nov 20;4:385. doi: 10.3389/fimmu.2013.00385. PMID: 24312095; PMCID: PMC3834540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Cagno V., Tseligka E.D., Jones S.T., Tapparel C. Heparan sulfate proteoglycans and viral attachment: true receptors or adaptation bias? Viruses. 2019 Jul 1;11(7):596. doi: 10.3390/v11070596. PMID: 31266258; PMCID: PMC6669472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Milewska A., Zarebski M., Nowak P., Stozek K., Potempa J., Pyrc K. Human coronavirus NL63 utilizes heparan sulfate proteoglycans for attachment to target cells. J. Virol. 2014 Nov;88(22):13221–13230. doi: 10.1128/JVI.02078-14. Epub 2014 Sep 3. PMID: 25187545; PMCID: PMC4249106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Kearns F.L., Sandoval D.R., Casalino L., Clausen T.M., Rosenfeld M.A., Spliid C.B., Amaro R.E., Esko J.D. Spike-heparan sulfate interactions in SARS-CoV-2 infection. Curr. Opin. Struct. Biol. 2022 Oct;76 doi: 10.1016/j.sbi.2022.102439. Epub 2022 Jul 6. PMID: 35926454; PMCID: PMC9257145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Naskalska A., Dabrowska A., Szczepanski A., Milewska A., Jasik K.P., Pyrc K. Membrane protein of human coronavirus NL63 is responsible for interaction with the adhesion receptor. J. Virol. 2019;93(19) doi: 10.1128/JVI.00355-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Hu Y., Meng X., Zhang F., Xiang Y., Wang J. The in vitro antiviral activity of lactoferrin against common human coronaviruses and SARS-CoV-2 is mediated by targeting the heparan sulfate co-receptor. Emerg. Microb. Infect. 2021 Dec;10(1):317–330. doi: 10.1080/22221751.2021.1888660. PMID: 33560940; PMCID: PMC7919907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Lang J., Yang N., Deng J., Liu K., Yang P., Zhang G., Jiang C. Inhibition of SARS pseudovirus cell entry by lactoferrin binding to heparan sulfate proteoglycans. PLoS One. 2011;6(8) doi: 10.1371/journal.pone.0023710. Epub 2011 Aug 22. PMID: 21887302; PMCID: PMC3161750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Möckl L. The emerging role of the mammalian glycocalyx in functional membrane organization and immune system regulation. Front. Cell Dev. Biol. 2020 doi: 10.3389/fcell.2020.00253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Chighizola M., Dini T., Marcotti S., D'Urso M., Piazzoni C., Borghi F., et al. The glycocalyx affects the mechanotransductive perception of the topographical microenvironment. J. Nanobiotechnol. 2022 doi: 10.1186/s12951-022-01585-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Thi M.M., Tarbell J.M., Weinbaum S., Spray D.C. The role of the glycocalyx in reorganization of the actin cytoskeleton under fluid shear stress: a “bumper-car” model. Proc. Natl. Acad. Sci. U.S.A. 2004 doi: 10.1073/pnas.0407474101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Spear M., Wu Y. Viral exploitation of actin: force-generation and scaffolding functions in viral infection. Virol. Sin. 2014 doi: 10.1007/s12250-014-3476-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Taylor M.P., Koyuncu O.O., Enquist L.W. Subversion of the actin cytoskeleton during viral infection. Nat. Rev. Microbiol. 2011 doi: 10.1038/nrmicro2574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Radtke K., Döhner K., Sodeik B. Viral interactions with the cytoskeleton: a hitchhiker's guide to the cell. Cell Microbiol. 2006 doi: 10.1111/j.1462-5822.2005.00679.x. [DOI] [PubMed] [Google Scholar]
- 55.Marsh M., Bron R. SFV infection in CHO cells: cell-type specific restrictions to productive virus entry at the cell surface. J. Cell Sci. 1997 doi: 10.1242/jcs.110.1.95. [DOI] [PubMed] [Google Scholar]
- 56.Bays J.L., DeMali K.A. Vinculin in cell–cell and cell–matrix adhesions. Cell. Mol. Life Sci. 2017 doi: 10.1007/s00018-017-2511-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Tolbert C.E., Burridge K., Campbell S.L. Vinculin regulation of F-actin bundle formation: what does it mean for the cell? Cell Adhes. Migrat. 2013 doi: 10.4161/cam.23184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Brown C., Morham S.G., Walsh D., Naghavi M.H. Focal adhesion proteins talin-1 and vinculin negatively affect paxillin phosphorylation and limit retroviral infection. J. Mol. Biol. 2011 doi: 10.1016/j.jmb.2011.03.076. [DOI] [PubMed] [Google Scholar]
- 59.Lowery J., Kuczmarski E.R., Herrmann H., Goldma R.D. Intermediate filaments play a pivotal role in regulating cell architecture and function. J. Biol. Chem. 2015 doi: 10.1074/jbc.R115.640359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Ivaska J., Pallari H.M., Nevo J., Eriksson J.E. Novel functions of vimentin in cell adhesion, migration, and signaling. Exp. Cell Res. 2007 doi: 10.1016/j.yexcr.2007.03.040. [DOI] [PubMed] [Google Scholar]
- 61.Ramos I., Stamatakis K., Oeste C.L., Pérez-Sala D. Vimentin as a multifaceted player and potential therapeutic target in viral infections. Int. J. Mol. Sci. 2020 doi: 10.3390/ijms21134675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Miller M.S., Hertel L. Onset of human cytomegalovirus replication in fibroblasts requires the presence of an intact vimentin cytoskeleton. J. Virol. 2009 doi: 10.1128/jvi.00398-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Teo C.S.H., Chu J.J.H. Cellular vimentin regulates construction of dengue virus replication complexes through interaction with NS4A protein. J. Virol. 2014 doi: 10.1128/jvi.01249-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Zhang Y., Wen Z., Shi X., Liu Y.J., Eriksson J.E., Jiu Y. The diverse roles and dynamic rearrangement of vimentin during viral infection. J. Cell Sci. 2021 doi: 10.1242/jcs.250597. [DOI] [PubMed] [Google Scholar]
- 65.Stefanovic S., Windsor M., Nagata K., Inagaki M., Wileman T. Vimentin rearrangement during african swine fever virus infection involves retrograde transport along microtubules and phosphorylation of vimentin by calcium calmodulin kinase II. J. Virol. 2005 doi: 10.1128/jvi.79.18.11766-11775. 2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Liang J.J., Yu C.Y., Liao C.L., Lin Y.L. Vimentin binding is critical for infection by the virulent strain of Japanese encephalitis virus. Cell Microbiol. 2011 doi: 10.1111/j.1462-5822.2011.01624.x. [DOI] [PubMed] [Google Scholar]
- 67.Hu Y., Jo H., DeGrado W.F., Wang J. Brilacidin, a COVID-19 drug candidate, demonstrates broad-spectrum antiviral activity against human coronaviruses OC43, 229E, and NL63 through targeting both the virus and the host cell. J. Med. Virol. 2022;94(5):2188–2200. doi: 10.1002/jmv.27616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Li J., Yu F., Chen Y., Oupický D. Polymeric drugs: advances in the development of pharmacologically active polymers. J. Contr. Release. 2015 doi: 10.1016/j.jconrel.2015.09.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Pyrć K., Milewska A., Duran E.B., Botwina P., Dabrowska A., Jedrysik M., et al. SARS-CoV-2 inhibition using a mucoadhesive, amphiphilic chitosan that may serve as an anti-viral nasal spray. Sci. Rep. 2021 doi: 10.1038/s41598-021-99404-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Mohan P., Schols D., Baba M., De Clercq E. Sulfonic acid polymers as a new class of human immunodeficiency virus inhibitors. Antivir. Res. 1992 doi: 10.1016/0166-3542(92)90034-3. [DOI] [PubMed] [Google Scholar]
- 71.Oh N., Park J.H. Endocytosis and exocytosis of nanoparticles in mammalian cells. Int. J. Nanomed. 2014;9:51–63. doi: 10.2147/IJN.S26592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Ding L., Yao C.J., Yin X.F., Li C.C., Huang Y.N., Wu M., Wang B., Guo X.Y., Wang Y.L., Wu M.H. Small. 2018;14 doi: 10.1002/smll.201801451. [DOI] [PubMed] [Google Scholar]
- 73.Chithrani B.D. Chan WC Elucidating the mechanism of cellular uptake and removal of protein-coated gold nanoparticles of different sizes and shapes. Nano Lett. 2007;7(6):1542–1550. doi: 10.1021/nl070363y. [DOI] [PubMed] [Google Scholar]
- 74.Mohamed S., Coombe D.R. Heparin mimetics: their therapeutic potential. Pharmaceuticals (Basel) 2017 Oct 2;10(4):78. doi: 10.3390/ph10040078. PMID: 28974047; PMCID: PMC5748635. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
No data was used for the research described in the article.