Summary
We present a protocol for quantifying the expression of the receptor gp130 using a calibrated flow cytometric approach. We describe pitfalls for receptor quantification such as titration of primary antibodies and standardizing cell culture. Receptors are stained with primary antibodies and fluorophore-coupled secondary antibodies. Beads covered with defined numbers of immunoglobulin G stained with fluorophore-coupled secondary antibodies serve as calibrators. In this way, the fluorescence intensity of cells is converted to the number of receptors on the cell surface.
For complete details on the use and execution of this protocol, please refer to Reeh et al. (2019).1
Subject areas: Single Cell, Flow Cytometry/Mass Cytometry, Signal Transduction
Graphical abstract

Highlights
-
•
Quantification of membrane-bound receptors using calibrated flow cytometry
-
•
IgG-covered beads allow converting fluorescence intensity to the number of receptors
-
•
Optimization steps include titration of antibodies and cell culture conditions
-
•
Optimized for quantification of gp130 but can easily be expanded to other receptors
Publisher’s note: Undertaking any experimental protocol requires adherence to local institutional guidelines for laboratory safety and ethics.
We present a protocol for quantifying the expression of the receptor gp130 using a calibrated flow cytometric approach. We describe pitfalls for receptor quantification such as titration of primary antibodies and standardized cell culture. Receptors are stained with primary antibodies and fluorophore-coupled secondary antibodies. Beads covered with defined numbers of immunoglobulin G stained with fluorophore-coupled secondary antibodies serve as calibrators. In this way, the fluorescence intensity of cells is converted to the number of receptors on the cell surface.
Before you begin
Quantitative experimental data are essential for understanding mechanisms of cellular signaling.2 Absolute quantification of intracellular signaling proteins and membrane-bound receptors is fraught with various challenges such as the requirement to lyse or permeabilize cells for quantification of cytoplasmic and nuclear proteins, and the need to focus on membrane-bound receptors available for ligand binding. Thus, they require specific approaches. Here, we describe a method to quantify membrane-bound receptors.
In the past receptor availability was analyzed by radioactive binding assays using 125I labeled cytokines.3,4 These assays have the disadvantages that quantification of receptor subunits in heteromeric receptor complexes is difficult and that they require a handling permit for working with radioactive materials. As a remedy, we present a flow cytometry-based method that relies on binding of specific antibodies to the receptors.
Several kits to quantify membrane-bound proteins by flow cytometry are commercially available. In principle they all make use of artificial Calibrator beads that are either coupled to a defined number of fluorophores (e.g., Quantibrite Beads, BD Biosciences) or primary antibodies (e.g., QIFIKIT, Agilent). We used both approaches successfully for quantification of cytokine receptors.1,5 However, when using fluorophore-coupled Calibrator beads the number of fluorophores on antibody-stained cells does not equal to the number of receptors in some cases. First, the usage of polyclonal antibodies recognizing an unknown number of epitopes increases the number of fluorophores on the cells in an unpredictable extent. Second, the number of fluorophores coupled to antibodies is often higher than one and is also not predictable. The first problem can be addressed by using monoclonal primary antibodies coupled to fluorophores. In contrast, the second problem is less easy to solve. For accurate quantification it is necessary to purify antibody-fluorophore-conjugates with a defined number of fluorophores e.g., by gel filtration. Otherwise, the mean number of fluorophores bound to an antibody can be determined spectrophotometrically (based on Beerś Law).
Alternatively, as described here in detail, beads coupled to a defined number of primary antibodies can be used. After staining of cells with a monoclonal antibody that specifically recognizes the receptor of interest, cells and beads are stained with the same fluorophore-coupled secondary antibody (here, FITC-conjugate). Thereby, amplification of the signal caused by 1) binding of unknown numbers of secondary antibodies to a single primary monoclonal antibody and 2) an undefined number of fluorophores coupled to a single secondary antibody affects both, beads and cells to the same extent. Consequently, the mean fluorescence intensity of the stained cells can be directly related to the number of primary monoclonal antibodies.
Of note, quantification of receptor expression based on flow cytometric data enables to determine not only the mean expression but also the heterogeneity of receptor expression on individual cells within a cell population.6
In the following, the IL-6 receptor subunit gp130 serves as an example for the quantification of membrane-bound receptors. Information on how to adopt the procedure to the quantification of other receptors is given in the Notes sections.
Before you begin prepare all buffers and media needed according to the materials and equipment section. While it is preferable to have all buffers freshly prepared, for convenience, some may be stored long-term, as indicated in the materials and equipment section.
Institutional permissions
Before starting work, inform yourself about the safe handling of the chemicals and cells described here and about the local regulations regarding protection and disposal.
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| Anti-gp130 (clone B-R3, IgG) (1:50) | Diaclone | Cat# 852.060.000 |
| Mouse IgG Isotype Control (RRID: AB_2801452) (1:25) | BioLegend | Cat# 401401 |
| Chemicals, peptides, and recombinant proteins | ||
| DMEM/F-12 with GlutaMAX | Gibco | Cat# 31331-028 |
| EDTA | Carl Roth | Cat# 8043.1 |
| Fetal calf serum (FCS) | Gibco | Cat# 10270-106 |
| Sodium chloride (NaCl) | ITW Reagents | Cat# 131659.1214 |
| Sodium hydrogen phosphate dihydrate (Na2HPO4 × 2 H2O) | Carl Roth | Cat# 4984 |
| Penicillin (10,000 U/mL)/streptomycin (10,000 μg/mL) | Gibco | Cat# 15140-122 |
| Potassium chloride (KCl) | Carl Roth | Cat# 6781 |
| Potassium dihydrogen phosphate (KH2PO4) | Carl Roth | Cat# 3904 |
| Critical commercial assays | ||
| QIFIKIT containing Set-up beads, Calibrator beads, and FITC-conjugated goat anti-mouse immunoglobulins (FITC-conjugate) | Agilent | Cat# K007811-8 |
| Experimental models: Cell lines | ||
| HepG2 | ATCC | HB-8065 |
| Software and algorithms | ||
| FlowJo | BD Biosciences | v#10.6.1 |
| Excel or another spreadsheet program | Microsoft | v#2019 |
| Other | ||
| 5 mL FACS tube | Fisher Scientific | Cat# 10186360 |
| Cell culture equipment including incubator, sterile hood, centrifuge, vacuum pump | Any company | |
| Neubauer chamber or other equipment to determine cell count | Any company | |
| Flow cytometer | Any company | |
Materials and equipment
All chemicals were used in the quality pro analysis.
PBS
| Reagent | Final concentration | Amount |
|---|---|---|
| NaCl | 200 mM | 11.69 g |
| KCl | 2.5 mM | 0.186 g |
| Na2HPO4 × 2 H2O | 8 mM | 1.42 g |
| KH2PO4 | 1.5 mM | 0.204 g |
| ddH2O | Add up to 1 L | |
| Total | 1 L |
Note: Adjust pH to 7.4 with HCl before adjusting to final volume with ddH2O.
Note: Autoclave PBS directly after preparation and subsequently store the sterile solution at 20°C–22°C for up to one year.
-
•
PBS/EDTA: 2 mM EDTA in PBS.
Note: Store at 4°C and use within one day.
-
•
FACS buffer: 2% FCS in PBS/EDTA.
Note: Store at 4°C and use within one day.
Step-by-step method details
Titration of primary antibodies
Timing: 28 h
Perform this step once before quantification to determine the saturating concentration of the monoclonal primary antibody for your antibody and cell line of choice. The saturating concentration indicates the number of antibodies needed to stain all receptors on the surface of the cells that will be analyzed.
-
1.
Seed 106 cells each on two 10 cm cell culture dishes in standard medium.
-
2.
Incubate the cells at 37°C, 10% CO2 in a humidified atmosphere for 24 h.
CRITICAL: Always use a certified biosafety cabinet and sterilized equipment and solutions. Follow aseptic technique when working with cells.
CRITICAL: Cytokine binding might interfere with antibody binding, so it is best to quantify receptor expression at the cell surface in the absence of cytokines. When isolating primary cells, receptors might be bound by cytokines. Carefully wash cells with PBS several times before receptor quantification to remove these cytokines. Additionally, test whether binding of the primary antibody is affected by cytokine binding by comparing fluorescence intensity of stained unstimulated cells and stained cells stimulated with recombinant cytokines.
Note: For HepG2 cells standard medium refers to DMEM/F-12 with Glutamax plus 10% FCS plus Pen/Strep. For other cell types adjust medium and supplements.
-
3.Detach cells from cell culture dishes.
-
a.Remove cell culture medium by vacuum aspiration and gently wash cells once with sterile PBS.
-
b.Add PBS/EDTA to cover the surface of the cell culture dish (e.g., 2 mL/10 cm dish).
-
c.Incubate cells on ice until they detach from the cell culture dish.
-
d.Resuspend cells with 4 mL ice-cold FACS buffer.
-
e.Transfer the cell suspension to a FACS tube.
-
a.
Note: Avoid using trypsin to detach cells from the cell surface as trypsin cleaves gp130 and other surface proteins. Instead of detaching cells with PBS/EDTA, you can also use any other protocol to detach and singularize cells until it does not cause receptor cleavage.
Note: At this point, you should create a suspension of single cells without cell aggregates. You can easily control this with a microscope. If cells continue to form aggregates, you can use a syringe (0.9 × 40 mm needle size) to mechanically disrupt cell-cell contacts or increase the incubation time with PBS/EDTA. Keep in mind that intensive physical steering stress may reduce receptor expression.
Note: We describe this method for adherent HepG2 cells. You need to adapt some of the steps to the cell line you are analyzing, particularly when detaching cells from tissue culture dishes. To harvest suspension cells, take them directly out of the culture flask and centrifuge them (233 g, 4°C, 5 min). Resolve the cell pellet in PBS and centrifuge again.
-
4.Wash cells.
-
a.Add 4 mL ice-cold FACS buffer to each FACS tube.
-
b.Vortex cells briefly.
-
c.Centrifuge cells (233 g, 4°C, 5 min).
-
d.Remove supernatant by vacuum aspiration.
-
a.
-
5.
Resuspend cells in 2 mL ice-cold FACS buffer.
-
6.
Determine the cell number.
-
7.
Resuspend 200,000 cells each in 100 μL ice-cold FACS buffer with various dilutions of the primary antibody (e.g., dilute the antibody 1:20, 1:50, 1:100, 1:200, 1:500). Leave two aliquots of cells unstained as controls to account for 1) auto-fluorescence of the cells and 2) unspecific binding of the FITC-conjugate, respectively. Resuspend these control cells in 100 μL FACS buffer each.
Note: Only the concentration of the primary antibody needs to be titrated. Saturating concentration of the FITC-conjugate is used according to the QIFIKIT manual.
Note: The choice of the primary antibody depends on your protein of interest. If you want to use QIFIKIT for quantification, as described here, you need a mouse monoclonal IgG primary antibody, as the calibration beads are covered with mouse monoclonal IgG.
Note: Test the specificity of the primary antibody by staining cells with the antibody that are known to not express the receptor or where the gene coding for the receptor has been knocked out. If staining is positive in cells that do not express the receptor, it is necessary to choose another antibody clone.
Note: Complete resuspension of the cell pellet is crucial for homogenous binding of the antibodies. However, the cells are alive during staining, so avoid strong and long vortexing at all steps performed and keep cells on ice.
Note: If you are analyzing immune cells, such as monocytes, macrophages, dendritic cells, and B-cells, that express Fc receptors, include an incubation step with an Fc blocking reagent following manufacturerś description. This ensures that only antigen-specific binding is analyzed. For human cells, e.g., BD Pharmingen Human BD Fc Block (Cat# 564219) can be used.
-
8.
Incubate cell suspensions on ice for 30 min.
-
9.Wash cells twice.
-
a.Add 4 mL ice-cold FACS buffer to each FACS tube.
-
b.Vortex cells briefly.
-
c.Centrifuge cells (233 g, 4°C, 5 min).
-
d.Remove supernatant by vacuum aspiration.
-
a.
-
10.
Resuspend unstained control cells in 100 μL ice-cold FACS buffer.
-
11.
Resuspend all other cells, including cells that serve as control for unspecific binding of the secondary antibody, in 100 μL FACS buffer with FITC-conjugate (1:50 in FACS buffer) each.
-
12.
Incubate cells on ice in the dark for 1 h.
CRITICAL: Protect cells stained with FITC-conjugate from light-induced destruction of the fluorophore by incubating them in the dark.
-
13.
Wash cells as described in step 9.
-
14.
Resuspend cells in 300 μL ice-cold FACS buffer per FACS tube.
-
15.Analyze the fluorescence intensity of the cells using a flow cytometer.
-
a.Perform flow cytometer start-up.
-
b.Separate the cell population of interest from debris and large cell aggregates based on an SSC-A against FSC-A scatter plot (Figure 1A).
-
c.Gate single cells in an FSC-H against FSC-W scatter plot (Figure 1B).
-
d.Record the fluorescence intensity within the FITC channel for 10,000 single cells/sample and calculate the mean fluorescence intensity of these cells.
-
a.
-
16.
Determine the concentration of antibodies needed for saturation of the FITC signal, i.e., identify the concentration of antibodies that does not further increase the fluorescence intensity of the cells compared to the next lower antibody concentration (Figure 1C).
Note: For 200,000 HepG2 cells the saturating amount of the anti-gp130 (BR3) antibody is 1:50 (20 μg/mL). This dilution strongly depends on the concentration of the antibody stock solution (here, 1 μg/μL), the number of cells analyzed, and the number of epitopes present on the cells. Thus, each cell/antibody pair requires determining the dilution of the antibody that results in saturated binding.
Figure 1.
Gating of single cells
(A) SSC-A and FSC-A of HepG2 cells were analyzed by flow cytometry. The depicted gate marks living cells outside of bigger aggregates.
(B) FSC-H and FSC-W of living HepG2 cells from Figure 1A. The depicted gate marks single cells. Figure 1 A) and B) are modified from Reeh et al.1 with permission from Springer Nature.
(C) HepG2 cells were stained with anti-gp130 primary antibody followed by staining with FITC-conjugate. Unstained cells and cells only incubated with FITC-conjugate serve as control.
Flow cytometric quantification of surface receptors
Timing: 28 h
-
17.
Seed 106 cells /10 cm cell culture dish in standard medium.
-
18.
Incubate the cells at 37°C, 10% CO2 in a humidified atmosphere for 24 h.
-
19.Detach cells from cell culture dish.
-
a.Remove cell culture medium by vacuum aspiration.
-
b.Gently wash cells once with PBS.
-
c.Add PBS/EDTA to cover the surface of the cell culture dish (e.g., 2 mL/10 cm dish).
-
d.Incubate cells on ice until they detach from the cell culture dish.
-
e.Resuspend cells with 2 mL ice-cold FACS buffer.
-
f.Transfer 200 μL each of the cell suspension to two FACS tubes on ice.Note: Cells in one FACS tube will be stained with anti-gp130 antibody (IgG) and FITC-conjugate, cells in the other FACS tube will be incubated with a corresponding IgG isotype control and FITC-conjugate as control for unspecific binding of the primary antibody and the FITC-conjugate. Unspecific binding results in increased fluorescence and thus overestimates receptor numbers.Note: Perform all steps with pre-chilled buffers (store in the refrigerator or on ice) and on ice to prevent receptor internalization.Note: Working sterile is not necessary after this step as the incubation periods are short and performed on ice, thus contamination of the sample is highly unlikely.Note: Ensure that the number of stained cells is not too high, so that you do not need too much of the primary antibody to reach saturation of staining. Ideally, use 100,000 to 200,000 cells. Since only 10,000 cells are finally analyzed by flow cytometry, fewer cells are perfectly fine. However, some cells will get lost during washing steps, and others will be out-gated when gating for single-cells. Thus, a moderate excess in cell number is beneficial.
-
g.Add 2 mL ice-cold FACS buffer to wash the cells.
-
h.Centrifuge cells (233 g, 4°C, 5 min).
-
i.Remove FACS buffer by vacuum aspiration.
-
a.
-
20.Staining of cells with primary antibodies.
-
a.Resuspend one aliquot of cells in 100 μL ice-cold FACS buffer with primary antibodies at saturating concentration (anti-gp130, 1:50, final concentration 20 μg/mL).
-
b.Resuspend one aliquot of cells in 100 μL ice-cold FACS buffer with IgG isotype control (1:25, final concentration 20 μg/mL) at the same concentration.
-
c.Incubate cells on ice for 1 h.
-
a.
Note: It is also possible to stain the cells in v-bottom 96-well dishes or 1.5 mL reaction tubes instead of FACS tubes. In this case, wash cells with 200 μL FACS buffer in the next steps.
Note: Complete resuspension of the cell pellet is crucial for homogenous binding of the antibodies to the cells. However, the cells are alive during staining, so avoid strong and long vortexing at all steps performed and keep cells always on ice.
-
21.Staining of cells and beads with FITC-conjugate.
-
a.Vortex Set-up beads and Calibrator beads briefly.
-
b.Transfer 20 μL of Set-up beads and 20 μL of Calibrator beads into separate FACS tubes.
-
c.Wash Set-up beads, Calibrator beads and the cells with 3 mL FACS buffer.Note: Set-up beads contain two populations of beads: blank beads and beads conjugated with a high number of mouse IgG. They are used to pre-adjust the detector gain of the FITC channel of the flow cytometer to ensure linearity of the measurement. Calibrator beads contain five populations of beads coupled to different numbers of mouse IgG.
-
d.Centrifuge Set-up beads, Calibrator beads, and cells (233 g, 4°C, 5 min).
-
e.Remove FACS buffer by vacuum aspiration.
-
f.Resuspend Set-up beads, Calibrator beads, and cells each in 100 μL ice-cold FACS buffer with FITC-conjugate (1:50).
-
g.Incubate Set-up beads, Calibrator beads, and cells on ice in the dark for 1 h.
 CRITICAL: Protect cells and beads stained with FITC-conjugate from light-induced destruction of the fluorophore by incubating them in the dark.Note: Always prepare a master mix containing FACS buffer and FITC-conjugate for the use in parallel samples. This ensures that you add the exact same amount of FITC-conjugate to the cells and beads.Note: Dead cells absorb antibodies and can therefore increase the fluorescence signal. To exclude dead cells from analyses you can use a live-dead assay as described by the manufacturer in addition to staining of the cells with primary antibodies and FITC-conjugate. Live/dead stains are available in different colors and should be chosen depending on the laser and filter equipment of your flow cytometer.
CRITICAL: Protect cells and beads stained with FITC-conjugate from light-induced destruction of the fluorophore by incubating them in the dark.Note: Always prepare a master mix containing FACS buffer and FITC-conjugate for the use in parallel samples. This ensures that you add the exact same amount of FITC-conjugate to the cells and beads.Note: Dead cells absorb antibodies and can therefore increase the fluorescence signal. To exclude dead cells from analyses you can use a live-dead assay as described by the manufacturer in addition to staining of the cells with primary antibodies and FITC-conjugate. Live/dead stains are available in different colors and should be chosen depending on the laser and filter equipment of your flow cytometer. -
h.Wash Set-up beads, Calibrator beads and cells with 3 mL FACS buffer.
-
i.Centrifuge Set-up beads, Calibrator beads, and cells (233 g, 4°C, 5 min).
-
j.Remove FACS buffer by vacuum aspiration.
-
k.Resuspend cells and beads in 300 μL FACS buffer.
-
a.
-
22.Analyze the fluorescence intensity using a flow cytometer.
-
a.Perform flow cytometer start-up.
-
b.Determine optimal gain of FITC detector channel:
-
i.Gate Set-up beads in the SSC-A against FSC-A scatter plot (Figure 2A).
-
ii.Gate living cells in the SSC-A against FSC-A scatter plot and single cells in the FSC-H against FSC-W scatter plot (Figures 1A and 1B).
-
iii.Adjust the detector gain of the FITC channel so that fluorescence of blank Set-up beads, control cells stained with IgG isotype control and FITC-conjugate, and the Set-up bead population with a high number of IgGs are depicted on the same log scale (Figure 2B).Note: This step is crucial to determine the linear range of the measurement. Once you start the measurement, you must not change the detector gain of the FITC channel as this would make it impossible to compare FITC fluorescence intensity between different samples for absolute quantification. However, changing the detector gain of FSC and SSC between the measurements of cells and beads may be needed as they differ greatly in size and granularity.
-
i.
-
c.Record fluorescence intensity within the FITC channel of 10,000 Set-up beads, 10,000 Calibrator beads and 10,000 single cells/sample without changing the detector gain of the FITC channel.
-
d.Export FCS files.
-
e.Calculate amount of gp130 as described in the quantification and statistical analysis section.
-
a.
Figure 2.
Determination of optimal gain of FITC detector channel
(A) SSC-A and FSC-A of Set-up beads were analyzed by flow cytometry. The depicted gate marks single beads.
(B) HepG2 cells were treated with IgG isotype control and FITC-conjugate (blue). Set-up beads were stained with FITC-conjugate (green). Analysis was performed by flow cytometry. Figure 2 is modified from Reeh et al.1 with permission from Springer Nature.
Expected outcomes
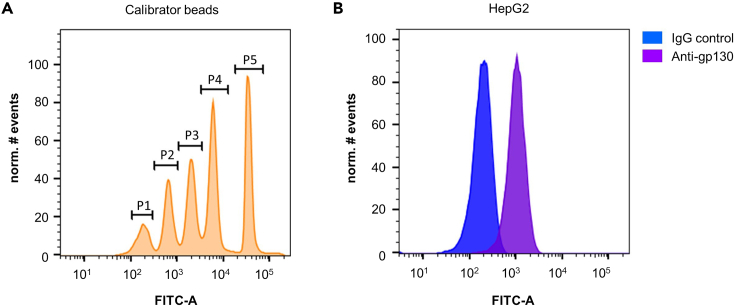
An exemplary result of quantification of gp130 is shown in Figure 3. The QIFIKIT contains five different populations of Calibrator beads that differ in the amount of IgG bound to their surface. Upon staining with FITC-conjugate these five populations can easily be differentiated based on FITC fluorescence intensity (Figure 3A). The five populations are marked with gates p1–p5. HepG2 cells were incubated with an anti-gp130 antibody and FITC-conjugate or with an IgG isotype control and FITC-conjugate. Subsequently, fluorescence intensity of the cells was determined with the same detector gain of the FITC channel as the Calibrator beads. Fluorescence of HepG2 cells incubated with the IgG isotype control and the FITC-conjugate is caused by autofluorescence of the cells and unspecific binding of either the IgG isotype control or the FITC-conjugate (Figure 3B, blue). Thus, the mean fluorescence intensity of cells incubated with the IgG isotype control and FITC-conjugate has to be subtracted from the mean fluorescence intensity of cells stained with anti-gp130 antibody and FITC-conjugate (Figure 3B, violet). Note, that the fluorescence intensity of the cells is within the linear range of the Calibrator beads (Figure 3A).
Figure 3.
Quantification of gp130 surface expression by calibrated flow cytometry
(A) Calibrator beads stained with FITC-conjugate were analyzed by flow cytometry. The five bead populations with different amounts of IgG were marked with gates p1-p5.
(B) HepG2 cells were incubated with IgG isotype control and FITC-conjugate or anti-gp130 antibody and FITC-conjugate, respectively. Analysis was performed by flow cytometry. Figure 3 is modified from Reeh et al.1 with permission from Springer Nature.
Quantification and statistical analysis
-
1.
Import FCS files into an analysis program such as FlowJo.
Note: Export to an analysis program is not strictly necessary and analysis can also be performed using the flow cytometer software. However, graphical presentation of the data is often more illustrative in analysis programs such as FlowJo.
-
2.
Gate single cells and bead singlets as visualized in Figures 1A and 1B and 2A.
-
3.
Plot the fluorescence intensity of the Calibrator beads in a histogram and gate the five populations (Figure 3A).
-
4.
Determine the mean FITC fluorescence intensity for each of the five Calibrator bead populations (Table 1).
- 5.
| (1) |
where a is the slope and b is the y-intercept of the calibration curve.
Note: Perform analysis of the results and linear regression in Excel or any other spreadsheet program.
Note: The lot-specific number of IgG on the different bead populations is provided by the manufacturer.
-
6.
Determine the mean FITC fluorescence intensity of cells stained with anti-gp130 antibody and FITC-conjugate and cells incubated with IgG isotype control and FITC-conjugate (Figure 3B).
-
7.
Subtract the mean background fluorescence, i.e., fluorescence intensity of control cells incubated with IgG isotype control and FITC-conjugate, from the mean fluorescence of cells stained with anti-gp130 antibodies and FITC-conjugate (Corrected mean FITC intensity in Table 2).
-
8.
Calculate the Log10 of the corrected mean FITC intensity of the cells (Log10 (corrected mean FITC intensity) in Table 2).
-
9.
Calculate Log10 (receptor/cell) based on the formula resulting from linear regression as described in Step 5b (Figure 4) and the Log10 (corrected mean FITC intensity) of the cells. Y in the formula represents the Log10 (corrected mean FITC intensity) while x represents Log10 (receptor/cell).
-
10.
Transform the obtained value back to linear by calculating 10 log(receptor/cell). The result corresponds to the mean number of receptors/cell (Table 2).
Table 1.
Exemplary fluorescence intensity of Calibrator beads
| Bead population | Mean FITC intensity | Log10 (mean FITC intensity) | IgG/bead | Log10 (IgG/bead) |
|---|---|---|---|---|
| Calibration p1 | 178 | 2.2 | 2700 | 3.4 |
| Calibration p2 | 646 | 2.8 | 12,000 | 4.1 |
| Calibration p3 | 2006 | 3.3 | 39,000 | 4.6 |
| Calibration p4 | 5906 | 3.8 | 127,000 | 5.1 |
| Calibration p5 | 37,467 | 4.6 | 787,000 | 5.9 |
Calibrator beads (Figure 3A) stained with FITC-conjugate were analyzed by flow cytometry. Log10 of mean fluorescence intensities of the analyzed bead populations and log10 of the lot-specific IgG amount per bead in p1-p5 are presented.
Figure 4.
Calibration curve
Linear dependency of Log10 (corrected mean FITC intensity) and Log10 (IgG/bead) (Table 1).
The equation displayed is the result of a linear regression. Figure 4 is modified from Reeh et al.1 with permission from Springer Nature.
Table 2.
Exemplary quantification of gp130
| Sample | Mean FITC intensity | Corrected mean FITC intensity | Log10 (corrected mean FITC intensity) | Log10 (receptors/cell) | Receptors/cell |
|---|---|---|---|---|---|
| HepG2 IgG control | 192 | ||||
| HepG2 Anti-gp130 | 1038 | 846 | 2.9 | 4.3 | 19,952 |
Limitations
This method only allows to quantify membrane-bound proteins such as cytokine receptors on the cell surface, and not intracellular proteins. Therefore, any receptors that have been internalized or are in the secretory pathway within the cell will not be detected. While this allows to precisely monitor the number of cytokine-available receptors, it underestimates the number of total receptors in a cell. As a remedy, quantification of intracellular signaling proteins can also be achieved through calibrated Western blotting.7
Quantification of intracellular signaling proteins with flow cytometric approaches is not straightforward. Staining of intracellular proteins needs fixation and permeabilization of the cells. Especially for staining of some nuclear transcription factors not only the cell membrane but also the nuclear membrane has to be permeabilized.8 Additionally, intracellular staining with a combination of primary and secondary antibodies is prone to high unspecific binding and thus background noise. Therefore, only fluorophore-coupled primary antibodies should be applied for intracellular flow cytometry. However, the approach presented here depends on staining of primary antibodies bound to cells and Calibrator beads with FITC-conjugates. Alternatively, artificial Calibrator beads that are coupled to a lot-specific well-defined number of fluorophores (e.g., Quantibrite Beads, BD Biosciences) could be used as calibrators when using fluorophore-coupled primary antibodies. However, fluorescence emitted intracellularly might be scattered and as a consequence fluorescence intensity of intracellularly stained cells might not be comparable with fluorescence intensity emitted from the Calibrator beads.
Beside this, quantification of the stoichiometry of receptor complexes is not possible with this protocol, as this protocol only measures the total number of receptors on the cell surface and not the number of receptors within a receptor complex.
Troubleshooting
Problem 1
The fluorescence intensity of cells stained with primary antibody and FITC-conjugate is not stronger than the fluorescence intensity of control cells incubated with IgG isotype control and FITC-conjugate (Step 22).
Potential solution 1
If incubation of the cells with primary antibody does not increase the fluorescence, this indicates that the receptor is not expressed or that the primary antibody does not recognize the receptor. In the latter case, it is necessary to choose another antibody clone or to optimize the staining conditions, such as increasing the incubation time with the antibody or the antibody concentration. Note, that improper handling of cells might cause receptor internalization, so make sure to perform all steps on ice.
Problem 2
High background fluorescence of cells incubated with IgG isotype control and FITC-conjugate (Step 22).
Potential solution 2
This indicates highly unspecific binding of the FITC-conjugate. Try to increase the percentage of BSA in the FACS buffer to block unspecific binding to the cell surface or to reduce the concentration of FITC-conjugate. Alternatively, you can increase the number of washing steps to three or four times per washing step to remove unbound antibodies.
Problem 3
The results of independent experiments differ, i.e., quantification yields in different numbers of membrane-bound receptors (Step 22 e).
Potential solution 3
The cell surface expression of receptors can be affected by cell culture conditions such as passage number, cell density, medium, composition of starvation medium, and length of starvation. To control the influence of these factors on the quantification, set standard conditions for quantitative experiments and strictly adhere to them.
Problem 4
No specific mouse monoclonal IgG against the membrane-bound protein under investigation is available on the market (Step 7).
Potential solution 4
You cannot use QIFIKIT in this case. Alternatively, you can use any primary monoclonal antibody directly coupled to phycoerythrin and use the Quantibrite kit for quantification.
Problem 5
Instead of membrane-bound proteins, intracellular proteins should be quantified.
Potential solution 5
Quantification of intracellular signaling proteins can be performed by calibrated Western blotting as described in ref. 7.
Resource availability
Lead contact
Further information and requests for resources and/or reagents should be directed to and will be fulfilled by the lead contact:
Dr. Anna Dittrich, Institute of Biology, Department of Systems Biology, Otto-von-Guericke University Magdeburg, Universitätsplatz 2, Building 28, 39106 Magdeburg E-Mail: anna.dittrich@ovgu.de.
Materials availability
This study did not generate new unique reagents.
Acknowledgments
This work was funded by the Deutsche Forschungsgemeinschaft (DFG) to A.D. (SCHA 785/15-1). We acknowledge support by the Open Access Publication Fund of Magdeburg University. The graphical abstract was created using BioRender (https://biorender.com).
Author contributions
Conceptualization, A.D.; investigation, A.D., N.M., and N.K.; visualization, A.D. and N.M.; writing – original draft, N.M.; writing – review and editing, A.D., N.M., and N.K.; supervision, A.D.
Declaration of interests
The authors declare no competing interests.
Data and code availability
This paper does not report original code. Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
References
- 1.Reeh H., Rudolph N., Billing U., Christen H., Streif S., Bullinger E., Schliemann-Bullinger M., Findeisen R., Schaper F., Huber H.J., Dittrich A. Response to IL-6 trans- and IL-6 classic signalling is determined by the ratio of the IL-6 receptor alpha to gp130 expression: fusing experimental insights and dynamic modelling. Cell Commun. Signal. 2019;17:46. doi: 10.1186/s12964-019-0356-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Dittrich A., Hessenkemper W., Schaper F. Systems biology of IL-6, IL-12 family cytokines. Cytokine Growth Factor Rev. 2015;26:595–602. doi: 10.1016/j.cytogfr.2015.07.002. [DOI] [PubMed] [Google Scholar]
- 3.Baumann H., Isseroff H., Latimer J.J., Jahreis G.P. Phorbol ester modulates interleukin 6- and interleukin 1-regulated expression of acute phase plasma proteins in hepatoma cells. J. Biol. Chem. 1988;263:17390–17396. doi: 10.1016/S0021-9258(19)77848-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Pietzko D., Zohlnhöfer D., Graeve L., Fleischer D., Stoyan T., Schooltink H., Rose-John S., Heinrich P.C. The hepatic interleukin-6 receptor. Studies on its structure and regulation by phorbol 12-myristate 13-acetate-dexamethasone. J. Biol. Chem. 1993;268:4250–4258. doi: 10.1016/S0021-9258(18)53603-6. [DOI] [PubMed] [Google Scholar]
- 5.Dittrich A., Quaiser T., Khouri C., Görtz D., Mönnigmann M., Schaper F. Model-driven experimental analysis of the function of SHP-2 in IL-6-induced Jak/STAT signaling. Mol. Biosyst. 2012;8:2119–2134. doi: 10.1039/c2mb05488d. [DOI] [PubMed] [Google Scholar]
- 6.Schaper F., Jetka T., Dittrich A. Vol. 24. 2022. Decoding Cellular Communication: An Information Theoretic Perspective on Cytokine and Endocrine Signaling. (Decoding Cellular Communication: An Information Theoretic Perspective on Cytokine and Endocrine Signaling). [DOI] [Google Scholar]
- 7.Köhler N., Miri N., Dittrich A. Quantification of total and phosphorylated STAT3 by calibrated Western blotting. Star Protoc. 2023;4 doi: 10.1016/j.xpro.2023.102508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Billing U., Jetka T., Nortmann L., Wundrack N., Komorowski M., Waldherr S., Schaper F., Dittrich A. Robustness and Information Transfer within IL-6-induced JAK/STAT Signalling. Commun. Biol. 2019;2:27. doi: 10.1038/s42003-018-0259-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
This paper does not report original code. Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.