Abstract
BACKGROUND
Preimplantation genetic testing for aneuploidy has been proven to be effective in determining the embryo's chromosomal or ploidy status. The test requires a biopsy of embryonic cells on day 3, 5, or 6 from which complete information on the chromosomes would be obtained. The main drawbacks of preimplantation genetic testing for aneuploidy include its relatively invasive approach and the lack of research studies on the long-term effects of preimplantation genetic testing for aneuploidy.
OBJECTIVE
Computer-assisted predictive modeling through machine learning and deep learning algorithms has been proposed to minimize the use of invasive preimplantation genetic testing for aneuploidy. The capability to predict morphologic characteristics of embryo ploidy status creates a meaningful support system for decision-making before further treatment.
STUDY DESIGN
Image processing is a component in developing a predictive model specialized in image classification through which a model is able to differentiate images based on unique features. Image processing is obtained through image augmentation to capture segmented embryos and perform feature extraction. Furthermore, multiple machine learning and deep learning algorithms were used to create prediction-based modeling, and all of the prediction models undergo similar model performance assessments to determine the best model prediction algorithm.
RESULTS
An efficient artificial intelligence model that can predict embryo ploidy status was developed using image processing through a histogram of oriented gradient and then followed by principal component analysis. The gradient boosting algorithm showed an advantage against other algorithms and yielded an accuracy of 0.74, an aneuploid precision of 0.83, and an aneuploid predictive value (recall) of 0.84.
CONCLUSION
This research study proved that machine-assisted technology perceives the embryo differently than human observation and determined that further research on in vitro fertilization is needed. The study finding serves as a basis for developing a better computer-assisted prediction model.
Key words: artificial intelligence, image processing, in vitro fertilization, noninvasive embryo assessment, preimplantation genetic testing for aneuploid, ploidy status, prediction model
AJOG MFM at a Glance.
Why was this study conducted?
Because of the lack of studies on preimplantation genetic testing for aneuploid (PGT-A) treatment, an alternative to the invasive PGT-A method is needed.
Key findings
Using computer-assisted technology to assess embryo ploidy status is doable; however, there is a limitation to the quantity of datasets being used. Nevertheless, computer-assisted technology can be adopted for decision support of a noninvasive PGT-A.
What does this add to what is known?
Computer-assisted technology for the assessment of ploidy status has been developed; however, computer-generated prediction is not dependable, and medical personnel is still required to assess the results.
Introduction
Infertility is a disease characterized by the inability of couples to conceive despite 1 year of regular sexual intercourse without contraception. As part of assisted reproduction technology, in vitro fertilization (IVF) has become an effective solution for infertility. During IVF, patients will undergo a single- or multiple-embryo transfer and a preimplantation embryo selection, which is a crucial part of this process.
Embryo selection based on ploidy status has been proven to be effective in reducing the risk of miscarriage while increasing the implantation rate.1,2 Ploidy status is assessed through preimplantation genetic testing for aneuploidy (PGT-A), which accurately specifies the chromosome number in biopsied embryonic cells.3 However, embryo biopsy holds unknown risks on the safety of embryo development before and after implantation,4,5 and studies are limited to support the normalcy of PGT-A–assessed embryos at birth or early childhood.6,7
Noninvasive methods, such as image processing, could be used to differentiate morphologic differences. Machine learning (ML) and deep learning (DL), parts of artificial intelligence (AI), could overcome the limitations of conventional human observation3,8 to predict the ploidy status of embryos based on morphologic differences. ML and DL have been successful in classifying images according to various medical image domains, such as blastocyst morphologic quality,9,10 embryo development stages,11,12 and implantation potential.13
Materials and Methods
Dataset
Data sources were obtained from the Morula IVF Jakarta Clinic, Jakarta, Indonesia, using 2 separate extraction methods: static image extraction from time-lapse videos recorded through a closed incubator system (MIRI time-lapse incubators) (37°C, 6% CO2, and 5% O2) and direct image extraction captured using an inverted microscope (Olympus IX71 or Nikon Eclipse Ti, Japan). By combining 483 couples, we were able to obtain 1123 embryo samples. The ploidy status of all embryo samples was determined through PGT-A, with baseline and clinical characteristics shown in Supplemental Table.
Image reduction was performed to the dataset, consisting of images in which the embryos were not fully captured, images containing additional objects attached to the embryo (eg, holding or biopsy pipette), and unclear images. This ML approach was performed through supervised learning, with known embryo ploidy classifications, such as euploid, aneuploid, or mosaic. This research was split into 2 case studies, with case study 1 employing 3 classifications—euploid, aneuploid, and mosaic—and case study 2 using binary classification. In binary classification, aneuploid and mosaic embryos were both categorized into 1 classification14 against the euploid classification. This research did not use image duplicates or multiplication; hence, each image was unique. Figure 1 shows the data distribution for each class. A total of 865 blastocyst images were used. The dataset was divided into training and testing sets with an 80:20 split. Separate training and testing datasets would produce unbiased results, especially in cases with limited datasets.15,16
Figure 1.
Data distribution
Danardono. Ploidy status classification. Am J Obstet Gynecol Glob Rep 2023.
Preimplantation genetic testing for aneuploid laboratory protocol
Embryo biopsy was performed as previously described in Polim et al's17 study. Briefly, a trophectoderm biopsy was performed on day 5 or 6, depending on the embryologist's assessment regarding the blastocyst quality. Of note, 3 to 5 cells of Trophectoderm (TE) were biopsied and transferred into a sterile microcentrifuge tube containing phosphate-buffered saline solution (Cell Signaling Technologies, Danvers, MA), which was supplemented with 1% polyvinylpyrrolidone (Origio). The samples were stored at −20°C before further use of genetic analysis. Whole-genome amplification was performed using the SurePlex DNA Amplification System (Illumina, San Diego, CA). The Veriseq PGS-MiSeq kit (Illumina, San Diego, CA) was used for the Next-generation sequencing (NGS) procedure, and the MiSeq sequencer (Illumina, San Diego, CA) was used for sequencing. Data interpretation used the BlueFuse Multi Software (version 4.5; Illumina, San Diego, CA; 32178). The threshold for calling mosaic was a 30% to 80% mixture of euploid and aneuploid cells (<30% was euploid, and >80% was aneuploid).
Methods
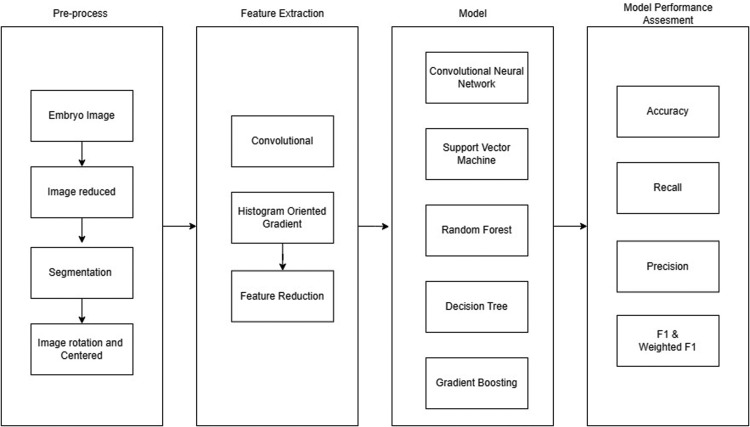
In developing the prediction model, performing feature extraction to obtain a unique differentiator in each classification became a significant step. Figure 2 illustrates the step-by-step approach to developing the AI model.
Figure 2.
Research workflow
Danardono. Ploidy status classification. Am J Obstet Gynecol Glob Rep 2023.
Image preprocessing
Image preprocessing was proposed to be beneficial toward AI model performance, reduce the possibility of any interference from the image background, and capture important information about an image object.18
Image segmentation was performed to minimize noise and dark fields surrounding the embryo and remove any interfering objects. This procedure was proposed to improve morphologic analysis and to reduce the possibility of misclassification because of interfering objects.19 Figure 3 shows a detailed process of image augmentation.
-
1.
Original data: A copy image of an embryo was generated. During the augmentation process, both the original and copy images were used. The copy image was used for the augmentation process and to make changes to complement the following process while still keeping the original image for further use.
-
2.
Strip alpha channel: The alpha channel or transparency density should be normalized for all images. Maximizing the value of the alpha channel would eliminate the pixel transparency differences within the image.
-
3.
Image blurring: Image blurring was used to eliminate any pixel degradation.
-
4.
Color enhancement: The foreground (image object) and background were distinguished through color enhancement to intensify color differences. This approach was achieved through a color balance algorithm,20 in which a high-contrast color balance was applied.
-
5.
Gray scale: The colored image was converted to gray scales to determine the image threshold.
-
6.
Image blurring: The second stage of image blurring was performed to complement pixel degradation after conversion to gray scale.
-
7.
Thresholding: Thresholding was performed to produce binary images and differentiate the image foreground from the image background.21,22
-
8.
Masking and padding: Masking and padding were applied to the original image with the threshold image as the mask parameter, which would be used to segment the embryo images.23
-
9.
Image to center: Before finding the image centroid and extreme vertical and horizontal alignment, moving the embryo object into the center of the canvas or frame is important.
-
10.
Detect centroid and extreme coordinate: Detect centroid and extreme coordinate determine the coordinates of the center-weighted embryo: most outer left, right, top, and bottom of the embryo.
-
11.Calculate embryo orientation: Calculating the embryo orientation uses a slope function to determine the vertical and horizontal gradients, establishing embryo orientation and the rotational degree needed to align the embryo.
(1) -
12.
Rotate image and center image: Rotate image and center image apply the rotational degree obtained from the image orientation calculation with the image centroid as the center point and reapply the image to the center step to designate the embryo at the middle of the frame.
Figure 3.
Image augmentation
Danardono. Ploidy status classification. Am J Obstet Gynecol Glob Rep 2023.
Feature extraction
Convolutional feature extraction
Convolutional feature extraction adopted the convolutional neural network (CNN) model architecture. It has a similar approach to the DL method.24,25 Adjustment was made before the output layer to foresee the possibility of performing the classification with ML algorithms. This method used a convolutional function; therefore, feature transformation holds similar characteristics to CNN images. The advantage of convolution feature extraction is its ability to map spatial and temporal correlations in image data.26 This approach enables us to use pretrained model architecture to produce image features. Here, VGG19,27 DenseNet,28 and ResNet29 pretrained models were used.
Histogram of oriented gradient and feature reduction
Images were converted into dimensional arrays and then became image feature descriptors, with each array cell represented by pixel values. Using the histogram of oriented gradient (HOG) algorithm, the grayscale image was denoted as a 2-dimensional array. The HOG works by subsequently dividing the dimensional array into small spatial regions and then producing pixel orientation toward its surrounding pixels.30,31 The combination of pixel orientation for each region will produce an image that will show features of an image in a gradient field perspective.
The massive number of features extracted from an image could lead to an overwhelming process of information. Feature reduction is a method to merge multiple variables into smaller numbers of variables. Principal component analysis (PCA) is a variable reduction process through dimensionality reduction to extract the most important information from the image or features and compress the size of the image without losing significant information about the features.32,33
Model prediction
This study adopted multiple AI approaches and assessed the performance of each model, ultimately identifying the model with the most optimal prediction results. Although CNN is considered a state-of-the-art prediction model, it has a disadvantage because it requires a massive dataset to achieve appropriate outcomes. In contrast, the earlier generations of ML techniques were more suited to a lower amount of datasets. Thus, the size of the available dataset is crucial in determining the optimal approach. This study deployed 5 ML algorithms and 1 DL approach. Each approach had different conditional input-output requirements yet served similar purposes nonetheless.
Convolutional neural network
A DL approach for a prediction model consists of a multilayer perceptron in which each layer produces mathematical calculations concerning the final classification outcome. The CNN model was specifically developed for image classification. The convolutional layer within the model contributed to image feature extraction for the CNN model.
Decision tree
A decision tree (DT) is a supervised ML approach used to develop a prediction model based on a derivative function. A DT serves as a rule-based predictor that uses a combination of parameters and derivative functions for a linear decision rule.
Random forest
A random forest (RF) is an extension of a DT, which essentially functions by combining multiple smaller trees to create a single giant tree.
Gradient boosting
Gradient boosting (GB) is an extension of a DT with a regression function to create the next iteration of the decision rule.
Support vector machine
A supervised ML approach uses linear or nonlinear relationships to differentiate various classifications. Image classification using a support vector machine (SVM) achieved a higher accuracy and better performance compared with DT and RF algorithms.34
Logistic regression
Logistic regression (LR) is a supervised ML approach that uses a linear relationship among different classes. Generally, LR and SVM operate similarly on classification algorithms. One of the main differences between LR and SVM is the mathematical approach they use. The LR approach is based on statistical properties, whereas the SVM approach is based on geometrical properties.
Model performance assessment
Standardization was imperative for the model performance analysis; accuracy, recall, precision, and F1 from each model were evaluated to measure the model performance. Multiple assessment is necessary to determine the prediction model with the most optimal performance. The implementation of multiple matrix calculation is needed as it holds different advantages and disadvantages in interpreting a model performance, as there is no golden standard for calculating model performance.
Accuracy
| (2) |
Accuracy is a performance matrix that considers all conditional matrices as equal contributors. In the case of a balanced dataset, accuracy would yield a general report on the model performance on prediction tasks. However, in the case of an imbalanced dataset, further performance matrix measurements should be considered. The accuracy matrix is an early indicator to describe a model's performance, yet it cannot be used as a benchmark for the overall model performance.
Recall
| (3) |
An imbalanced dataset affects a model's performance in predicting binary or multiclass classification, and recall would be beneficial for this issue. Recall works on predicting a classification without caring whether the other classification is mislabeled. This means that the recall matrix is a way to ignore false classifications while keeping high chances of catching all targeted classifications.
Precision
| (4) |
The precision matrix measures the model performance in classifying a prediction as a correct positive prediction. Although the recall matrix disregards mislabeled classification, the precision matrix recognizes misclassifications and calculates the reliability of the model in making predictions.
F1 and weighted F1
The F1-score is used to compute a harmonic combination of precision and recall. If both precision and recall values are low, it will produce low F1-scores, and vice versa. The F1 score complies with a specific prediction output, whereas the weighted F1-score observes a proportionate prediction per classification over the total data sample.
| (5) |
| (6) |
Result
Our research study used AI to predict an embryo's ploidy status by using an augmented image while maintaining the original embryo information. Subsequently, we conducted an assessment to identify models with the highest performance as presented in 2 case study comparisons.
Table 1 shows that the euploid-aneuploid classification (case study 2) was distinguished remarkably compared with the euploid-aneuploid-mosaic classification (case study 1). Low accuracy scores in case study 1 indicated that almost all of the models did not achieve the appropriate euploid predictive scores. In contrast, in case study 2, the models had fewer tasks while performing binary classification, which led to a superior model performance as it reduced the classification option and increased the probability of prediction.
Table 1.
Model performance
| No. | Model | Accuracy | Recall |
||
|---|---|---|---|---|---|
| Euploid predicted value | Aneuploid predictive value | Mosaic predicted value | |||
| Case study 1 | |||||
| 1 | DenseNet121 (SVM) | 0.42 | 0.05 | 0.75 | 0.60 |
| 2 | VGG16 (CNN) | 0.43 | 0.07 | 0.78 | 0.46 |
| 3 | DT (HOG-PCA256) | 0.40 | 0.07 | 0.69 | 0.23 |
| 4 | Logistic regression (HOG-PCA256) | 0.39 | 0.40 | 0.35 | 0.44 |
| 5 | Random forest (HOG-PCA256) | 0.41 | 0.02 | 0.55 | 0.35 |
| Case study 2 | |||||
| 6 | DenseNet121 (CNN) | 0.58 | 0.14 | 1.00 | — |
| 7 | DenseNet121 (SVM) | 0.56 | 0.00 | 1.00 | — |
| 8 | DenseNet169 (CNN) | 0.44 | 0.07 | 1.00 | — |
| 9 | DT (HOG-PCA256)a | 0.70a | 0.50a | 0.75a | — |
| 10 | Gradient boosting (HOG-PCA256)a | 0.74a | 0.33a | 0.84a | — |
CNN, convolutional neural network; HOG, histogram of oriented gradient; SVM, support vector machine; PCA, Principal Component Analysis.
Top 2 models from recall & accuracy model assessment.
Danardono. Ploidy status classification. Am J Obstet Gynecol Glob Rep 2023.
The top 2 models from case study 2 achieved significant accuracy scores of 0.70 and 0.74 using DT and GB algorithms, respectively. Table 2 shows that the GB model has a lower precision matrix with a higher F1-score matrix for predicting aneuploid embryos. Figure 4 shows the receiver operating characteristic (ROC) curve and precision-recall graph, with the GB algorithm slightly having a higher area under the curve (AUC) than the DT algorithm. The AUC calculates the region under the curve, with greater value yielding a better general model performance. The ROC curve observes the relationship between the true positive rate (sensitivity) and the false positive rate (1-specificity), and the precision-recall graph observes the relationship between precision and recall.
Table 2.
Model performance
| No. | Model | Prediction | Precision | F1-score | Weighted F1-score |
|---|---|---|---|---|---|
| 1 | Decision tree (HOG-PCA256) | Aneuploid | 0.85 | 0.80 | 0.72 |
| Euploid | 0.35 | 0.41 | |||
| 2 | Gradient boosting (HOG-PCA256)a | Aneuploid | 0.83a | 0.83a | 0.73a |
| Euploid | 0.35a | 0.34a |
HOG, histogram of oriented gradient; PCA, Principal Component Analysis.
Top model performance.
Danardono. Ploidy status classification. Am J Obstet Gynecol Glob Rep 2023.
Figure 4.
ROC and precision—recall graph
ROC, receiver operating characteristic.
Danardono. Ploidy status classification. Am J Obstet Gynecol Glob Rep 2023.
In developing prediction models with class imbalance, multiple matrix analysis was considered. A high confidence level for the prediction is important.
Here, the most robust prediction model, based on recall, precision, and F1-score for predicting aneuploid embryos, was developed using the GB algorithm and featured extraction using the HOG and then followed by the PCA. An optimal model should be able to classify different classes with similar accuracy, precision, and recall. In contrast, the performance of our model was still imbalanced with a bias toward one of the classifications. This model could be used to predict an aneuploid embryo, with a side note on the use of the model. When the model produces an euploid prediction, further tests should be performed to validate the real ploidy outcome. This approach will result in increasing the probability of finding a euploid embryo while reducing the number of embryos going through the PGT-A test.
Discussion
This study demonstrated the use of AI in providing an alternative noninvasive determination of embryo chromosomal ploidy status. Our model could differentiate morphologic differences between euploid and aneuploid-mosaic embryos with an overall accuracy of 0.74, precision of 0.83 for aneuploid precision, and aneuploid prediction value of 0.84. Computer-assisted technology has been around in the medical field to assist healthcare personnel during decision-making to proceed further with treatment. In our research, we observed that a recall value higher than 0.90 was caused by the overfitting of the model. Overfitting can be caused primarily by insufficient data, model complexity, or both, further exacerbating the effects. Although a recall value close to 0.00 was caused by the underfitting of the model because of an imbalanced and low dataset, a low and imbalanced data might cause a biased classification on account of the lesser amount of euploid embryos compared with mosaic and aneuploid embryos.
A previous study by Chavez-Badiola et al35 has demonstrated the possibility of using an AI model to predict embryo ploidy status with an accuracy of 0.70, an euploid (positive) predictive value of 0.79, and an aneuploid (negative) predictive value of 0.66. The datasets used were embryo images with known ploidy and implantation status, split into 2 groups, that is, euploid and/or successful implantation and aneuploid and/or implantation failure.
VerMilyea et al23 proposed an AI model to predict embryo viability based on Gardner scoring. The classification was split between viable and nonviable embryos with a Gardner score of 50% as the benchmark.36 A similar methodology was adopted by Diakiw et al37 to predict the ploidy status with adjustments on VerMilyea's approach for image preprocessing. Of note, 15,192 embryo images with known ploidy status were fed to the CNN to create a prediction model with multiple embryo classifications. Low mosaic embryos were grouped as euploid, and high mosaic embryos were grouped as aneuploid. The AI model did not specifically differentiate embryo ploidy. Diakiw et al's37 AI model had an overall accuracy of 65.4% and a sensitivity of 97.3%.
Barnes et al38 combined the embryo parameters to identify the most optimal model performance. Model parameters of the image, maternal age, morphokinetics, and Blastocysts (BL) score yielded the most ideal model performance with a higher AUC value of 0.7614 and a lower accuracy of 69.68%, compared with a model with similar parameters, which excluded morphokinetics, obtaining a lower AUC of 0.7558 and a higher accuracy of 70.23%.
Currently, Berntsen et al39 used the greatest number of 115,832 embryo datasets with known implantation data or known fetal heartbeat or known ploidy status for AI-based prediction modeling. Eventually, the datasets were classified into 2 categories: fetal heartbeat positive and fetal heartbeat negative (FH−). They achieved an astonishing result, an AUC of 0.95 when multivariables were calculated, and the discarded embryos were pseudolabeled as FH−.
To date, noninvasive AI models are still inferior to invasive genetic testing in predicting an embryo's ploidy status. Further scrutiny on the possibility to use ML prediction models as an alternative to PGT-A is imperative. The end goal is to create a robust model with high confidence and accuracy in predicting the ploidy status of embryos. Although the state-of-the-art technology for prediction models is an ongoing development, existing models could be modified and enhanced to perform specific prediction tasks.
Conclusion
We have developed a computer-assisted ML model that noninvasively predicts ploidy status. Particularly, our model has the ability to predict aneuploid embryos with high confidence levels. In contrast, our model lacks the capacity to classify euploid embryos. Possessing a 1-sided classification comes with the advantage of being able to draw conclusions about a specific class and disregard the other classification. Ultimately, our result has significantly proven the model's ability to predict ploidy status by using day 5 or 6 embryo image features with a minimum dataset. In our study, the embryo dataset, a major key aspect in constructing the ML model, was imbalanced, which influenced our model performance significantly. To date, the efficacy of PGT-A in determining the true complete chromosome status is unmatched by any noninvasive ML approaches. AI in the medical field would gradually advance with the continuous growth of available datasets to train the model. Prospectively, the study authors would continue to gather available datasets, minimize the gap between imbalanced classes, and conduct an external validation process.
Acknowledgments
The authors would like to thank the embryologists’ teams at Morula IVF Jakarta Clinic, Jakarta, Indonesia, for their assistance in collecting retrospective data, time-lapse video, and static embryo images.
Footnotes
The authors report no conflict of interest.
Data are available upon reasonable request.
This study was conducted according to the guidelines of the Declaration of Helsinki of 1975, as amended. The local research ethics committee in the Faculty of Medicine, Universitas Indonesia, Jakarta, Indonesia, has approved the study protocol (identification number: KET-572/UN2.F1/ETIK/PPM.00.02/2021).
Because of the nature of a retrospective study, a waiver of informed consent was granted by the ethics committee of the Faculty of Medicine, University of Indonesia, Jakarta, Indonesia, on March 1, 2021 (identification number: KET-572/UN2.F1/ETIK/PPM.00.02/2021).
This study received no funding.
Cite this article as: Danardono GB, Handayani N, Louis CM, et al. Embryo ploidy status classification through computer-assisted morphology assessment. Am J Obstet Gynecol Glob Rep 2023;XX:x.ex–x.ex.
Supplementary material associated with this article can be found in the online version at doi:10.1016/j.xagr.2023.100209.
Appendix. Supplementary materials
References
- 1.Forman EJ, Tao X, Ferry KM, Taylor D, Treff NR, Scott RT., Jr. Single embryo transfer with comprehensive chromosome screening Results in improved ongoing pregnancy rates and decreased miscarriage rates. Hum Reprod. 2012;27:1217–1222. doi: 10.1093/humrep/des020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Neal SA, Morin SJ, Franasiak JM, et al. Preimplantation genetic testing for aneuploidy is cost-effective, shortens treatment time, and reduces the risk of failed embryo transfer and clinical miscarriage. Fertil Steril. 2018;110:896–904. doi: 10.1016/j.fertnstert.2018.06.021. [DOI] [PubMed] [Google Scholar]
- 3.Pennetta F, Lagalla C, Borini A. Embryo morphokinetic characteristics and euploidy. Curr Opin Obstet Gynecol. 2018;30:185–196. doi: 10.1097/GCO.0000000000000453. [DOI] [PubMed] [Google Scholar]
- 4.Zacchini F, Arena R, Abramik A, Ptak GE. Embryo biopsy and development: the known and the unknown. Reproduction. 2017;154:R143–R148. doi: 10.1530/REP-17-0431. [DOI] [PubMed] [Google Scholar]
- 5.Cimadomo D, Capalbo A, Ubaldi FM, et al. The impact of biopsy on human embryo developmental potential during preimplantation genetic diagnosis. BioMed Res Int. 2016;2016 doi: 10.1155/2016/7193075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Desmyttere S, De Rycke M, Staessen C, et al. Neonatal follow-up of 995 consecutively born children after embryo biopsy for PGD. Hum Reprod. 2012;27:288–293. doi: 10.1093/humrep/der360. [DOI] [PubMed] [Google Scholar]
- 7.Natsuaki MN, Dimler LM. Pregnancy and child developmental outcomes after preimplantation genetic screening: a meta-analytic and systematic review. World J Pediatr. 2018;14:555–569. doi: 10.1007/s12519-018-0172-4. [DOI] [PubMed] [Google Scholar]
- 8.Lagalla C, Tarozzi N, Sciajno R, et al. Embryos with morphokinetic abnormalities may develop into euploid blastocysts. Reprod Biomed Online. 2017;34:137–146. doi: 10.1016/j.rbmo.2016.11.008. [DOI] [PubMed] [Google Scholar]
- 9.Thirumalaraju P, Kanakasabapathy MK, Bormann CL, et al. Evaluation of deep convolutional neural networks in classifying human embryo images based on their morphological quality. Heliyon. 2021;7:e06298. doi: 10.1016/j.heliyon.2021.e06298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Khosravi P, Kazemi E, Zhan Q, et al. Deep learning enables robust assessment and selection of human blastocysts after in vitro fertilization. NPJ Digit Med. 2019;2:21. doi: 10.1038/s41746-019-0096-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Raudonis V, Paulauskaite-Taraseviciene A, Sutiene K, Jonaitis D. Towards the automation of early-stage human embryo development detection. Biomed Eng Online. 2019;18:120. doi: 10.1186/s12938-019-0738-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Dirvanauskas D, Maskeliunas R, Raudonis V, Damasevicius R. Embryo development stage prediction algorithm for automated time lapse incubators. Comput Methods Programs Biomed. 2019;177:161–174. doi: 10.1016/j.cmpb.2019.05.027. [DOI] [PubMed] [Google Scholar]
- 13.Carrasco B, Arroyo G, Gil Y, et al. Selecting embryos with the highest implantation potential using data mining and decision tree based on classical embryo morphology and morphokinetics. J Assist Reprod Genet. 2017;34:983–990. doi: 10.1007/s10815-017-0955-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Viotti M, Victor AR, Barnes FL, et al. Using outcome data from one thousand mosaic embryo transfers to formulate an embryo ranking system for clinical use. Fertil Steril. 2021;115:1212–1224. doi: 10.1016/j.fertnstert.2020.11.041. [DOI] [PubMed] [Google Scholar]
- 15.Vabalas A, Gowen E, Poliakoff E, Casson AJ. Machine learning algorithm validation with a limited sample size. PLoS One. 2019;14 doi: 10.1371/journal.pone.0224365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Gholamy A, Kreinovich V, Kosheleva O. Why 70/30 or 80/20 relation between training and testing sets : a pedagogical explanation. 2018. Available at: https://scholarworks.utep.edu/cs_techrep/1209/. Accessed September 29, 2022.
- 17.Adrainus Polim A, Handayani N, Kesumapramudya Nurputra D, et al. Birth of spinal muscular atrophy unaffected baby from genetically at-risk parents following a pre-implantation genetic screening: a case report. Int J Reprod Biomed. 2022;20:779–786. doi: 10.18502/ijrm.v20i9.12068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Poostchi M, Silamut K, Maude RJ, Jaeger S, Thoma G. Image analysis and machine learning for detecting malaria. Transl Res. 2018;194:36–55. doi: 10.1016/j.trsl.2017.12.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Khan A, Gould S, Salzmann M. Segmentation of developing human embryo in time-lapse microscopy. 2016 IEEE ISBI. 2016:930–934. [Google Scholar]
- 20.Kay DY. Simplest color balance with OpenCV [w/code]: a very simple color balancing algorithm in OpenCV C++. 2015. Available at:https://www.morethantechnical.com/2015/01/14/simplest-color-balance-with-opencv-wcode/. Accessed July 28, 2022.
- 21.Otsu N. A threshold selection method from gray-level histograms. IEEE Trans Syst Man Cybern. 1979;9:62–66. [Google Scholar]
- 22.Bangare SL, Dubal A, Bangare PS, Patil ST. Reviewing Otsu's method for image thresholding. Int J App Eng Res. 2015;10:21777–21783. [Google Scholar]
- 23.VerMilyea M, Hall JMM, Diakiw SM, et al. Development of an artificial intelligence-based assessment model for prediction of embryo viability using static images captured by optical light microscopy during IVF. Hum Reprod. 2020;35:770–784. doi: 10.1093/humrep/deaa013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Chen Y, Jiang H, Li C, Jia X, Ghamisi P. Deep feature extraction and classification of hyperspectral images based on convolutional neural networks. IEEE Trans Geosci Remote Sens. 2016;54:6232–6251. [Google Scholar]
- 25.Yang A, Yang X, Wu W, Liu H, Zhuansun Y. Research on feature extraction of tumor image based on convolutional neural network. IEEE Access. 2019;7:24204–24213. [Google Scholar]
- 26.Khan A, Sohail A, Zahoora U, Qureshi AS. A survey of the recent architectures of deep convolutional neural networks. Artif Intell Rev. 2020;53:5455–5516. [Google Scholar]
- 27.Simonyan K, Zisserman A. Very deep convolutional networks for large-scale image recognition. arXiv. 2015 https://arxiv.org/abs/1409.1556 Available at: Accessed July 14, 2022. [Google Scholar]
- 28.Huang G, Liu Z, Van Der Maaten L, Weinberger KQ. Densely connected convolutional networks. CVPR. 2017:4700–4708. [Google Scholar]
- 29.He K, Zhang X, Ren S, Sun J. Deep residual learning for image recognition. CVPR. 2016:770–778. [Google Scholar]
- 30.Dalal N, Triggs B. Histograms of oriented gradients for human detection. IEEE CVPR. 2005:886–893. [Google Scholar]
- 31.Daniel N, Wasserman T, Adler Z, Czyzewski T, Savir Y. Machine learning reveals the effect of maternal age on the mouse pre-implantation embryo developmental timing. bioRxiv. 2022 https://www.biorxiv.org/content/10.1101/2022.05.17.492244v1 Available at: Accessed June 29, 2022. [Google Scholar]
- 32.Abdi H, Williams LJ. Principal component analysis. Wiley Interdiscip Rev Comput Stat. 2010;2:433–459. [Google Scholar]
- 33.Jolliffe IT, Cadima J. Principal component analysis: a review and recent developments. Philos Trans A Math Phys Eng Sci. 2016;374 doi: 10.1098/rsta.2015.0202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Chavez-Badiola A, Flores-Saiffe Farias A, Mendizabal-Ruiz G, Garcia-Sanchez R, Drakeley AJ, Garcia-Sandoval JP. Predicting pregnancy test results after embryo transfer by image feature extraction and analysis using machine learning. Sci Rep. 2020;10:4394. doi: 10.1038/s41598-020-61357-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Chavez-Badiola A, Flores-Saiffe-Farías A, Mendizabal-Ruiz G, Drakeley AJ, Cohen J. Embryo Ranking Intelligent Classification Algorithm (ERICA): artificial intelligence clinical assistant predicting embryo ploidy and implantation. Reprod Biomed Online. 2020;41:585–593. doi: 10.1016/j.rbmo.2020.07.003. [DOI] [PubMed] [Google Scholar]
- 36.Gardner DK, Sakkas D. Assessment of embryo viability: the ability to select a single embryo for transfer–a review. Placenta. 2003;24(SupplB) doi: 10.1016/s0143-4004(03)00136-x. S5–12. [DOI] [PubMed] [Google Scholar]
- 37.Diakiw SM, Hall JMM, Vermilyea MD, et al. Development of an artificial intelligence model for predicting the likelihood of human embryo euploidy based on blastocyst images from multiple imaging systems during IVF. Hum Reprod. 2022;37:1746–1759. doi: 10.1093/humrep/deac131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Barnes J, Malmsten J, Zhan Q, et al. Noninvasive detection of blastocyst ploidy (euploid vs. aneuploid) using artificial intelligence (AI) with deep learning methods. Fertil Steril. 2020;114:e76. [Google Scholar]
- 39.Berntsen J, Rimestad J, Lassen JT, Tran D, Kragh MF. Robust and generalizable embryo selection based on artificial intelligence and time-lapse image sequences. PLoS One. 2022;17 doi: 10.1371/journal.pone.0262661. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.