Key Points
Question
What are the benefits of applying gene panel sequencing as a first-tier newborn screening test at a population scale?
Findings
In this cohort study including 29 601 newborns, gene panel sequencing of 128 diseases as first-tier screening test identified 59 more patients (1 in 500 newborns). Of these, 20 patients were affected by disorders screened by both biochemical and gene panel sequencing and 39 patients were affected by disorders screened solely by gene panel sequencing.
Meaning
These results suggest that gene panel sequencing is a promising approach to screen newborns for monogenic disorders and would provide a better outcome when incorporated into the current newborn screening program.
This cohort study of Chinese newborns examines the outcomes of gene panel sequencing as a first-tier newborn screening test supplementing biochemical screening for monogenic disorders.
Abstract
Importance
Newborn screening via biochemical tests is in use worldwide. The availability of genetic sequencing has allowed rapid screening for a substantial number of monogenic disorders. However, the outcomes of this strategy have not been evaluated in a general newborn population.
Objective
To evaluate the outcomes of applying gene panel sequencing as a first-tier newborn screening test.
Design, Setting, and Participants
This cohort study included newborns who were prospectively recruited from 8 screening centers in China between February 21 and December 31, 2021. Neonates with positive results were followed up before July 5, 2022.
Exposures
All participants were concurrently screened using dried blood spots. The screen consisted of biochemical screening tests and a targeted gene panel sequencing test for 128 conditions. The biochemical and genomic tests could both detect 43 of the conditions, whereas the other 85 conditions were screened solely by the gene panel.
Main Outcomes and Measures
The primary outcomes were the number of patients detected by gene panel sequencing but undetected by the biochemical test.
Results
This study prospectively recruited 29 601 newborns (15 357 [51.2%] male). The mean (SD) gestational age was 39.0 (1.5) weeks, and the mean (SD) birth weight was 3273 (457) g. The gene panel sequencing screened 813 infants (2.7%; 95% CI, 2.6%-2.9%) as positive. By the date of follow-up, 402 infants (1.4%; 95% CI, 1.2%-1.5%) had been diagnosed, indicating the positive predictive value was 50.4% (95% CI, 50.0%-53.9%). The gene panel sequencing identified 59 patients undetected by biochemical tests, including 20 patients affected by biochemically and genetically screened disorders and 39 patients affected by solely genetically screened disorders, which translates into 1 out of every 500 newborns (95% CI, 1/385-1/625) benefiting from the implementation of gene panels as a first-tier screening test.
Conclusions and Relevance
In this cohort study, the use of gene panel sequencing in a general newborn population as a first-tier screening test improved the detection capability of traditional screening, providing an evidence-based suggestion that it could be considered as a crucial method for first-tier screening.
Introduction
Newborn screening (NBS) to identify severe but treatable disorders in newborns has been extremely successful worldwide in the past few decases.1 Newborn screening using dried blood spot (DBS) specimens was initially designed to detect a single disorder (phenylketonuria) in the 1960s.2 Technological advances, such as the use of tandem mass spectrometry, have played a significant role in the expansion of NBS to a number of disorders simultaneously.3 These DBS-based biochemical NBS methods rely on the measurement of metabolic substances, which resulted in an inherent limitation of screening disorders with specific indicators. Furthermore, biochemical NBS was criticized by its high false-positive rate and the need for differentiate diagnosis.4
Genomic sequencing identifies nucleotide changes that are the underlying etiology of monogenic disorders.5 It is thus regarded as a promising approach to minimize the false positive rate of NBS and facilitate timely case resolution.6 In addition, sequencing-based screening tests could theoretically allow for the identification of disorders for which a NBS assay is not currently available. Meanwhile, genomic sequencing is criticized for its cost.7
A 2020 study6 reported that genomic screening could not replace biochemical NBS because of the insufficient sensitivity. However, the benefits of sequencing-based screening as a first-tier test to be incorporated into the current NBS program remain ambiguous. Although several research groups have investigated the utility,6,8,9,10,11,12 these studies had 2 major limitations. First, genomic screening was always implemented as a second-tier test for high-risk infants who had initially been screened positive by biochemical NBS. The impact of genomic testing as a first-tier screening test for the general population was not determined. Second, these studies were limited by having a small sample size. Given the rarity of monogenic disorders, an adequately large sample is necessary, but few population-scale studies were reported. It also worth noting that most of these studies used whole exome or genome sequencing, which cannot be universally applied in clinical settings because of its high cost currently.7
In this study, we used gene panel sequencing NBS, which may be a promising alternative for whole exome or genome sequencing,11 and designed a targeted gene panel for universal NBS. We prospectively recruited 29 989 newborns from 8 NBS centers across China and performed DBS-based biochemical and genetic NBS in parallel. To our knowledge, this is the first study to incorporate gene panel sequencing into NBS as a concurrent first-tier screening strategy in a large population scale. Our study presents the outcomes of genetic NBS as a first-tier screening test in clinical settings, which is of great importance to clinicians, geneticists, and families for better health care and counseling.
Methods
Study Design and Participants
Newborns were prospectively recruited from 8 NBS centers in China from February 21, 2021, to December 30, 2021. The 8 NBS centers were selected to represent different regions, as they were located in the east, west, south, and north of China (Table 1). The participants were randomly recruited along with a routine NBS program. The inclusion criteria were (1) DBS specimens were available within 3 to 7 days after birth; (2) the parents signed written informed consent; and (3) at least four 8 × 8 mm DBS samples were collected. The exclusion criterion was receiving blood transfusion. DBS specimens were tested for biochemical and genetic NBS simultaneously. Biochemical NBS was performed in NBS centers; genetic NBS was performed in Beijing Genomics Institute. The screening results of each were mutually masked at the 2 laboratories and sent to clinicians for counseling. Patients with positive results were followed up by recalling for confirmatory tests or phone calls. To make medical diagnoses, the medical records were reviewed and discussed by a group of clinical experts, including neonatologists, pediatricians, and geneticists from the 8 NBS centers. This study followed the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) reporting guideline. Approval was obtained from the institutional review board of each center.
Table 1. Characteristics of the Newborns in Study Cohort.
| Characteristic | No. (%) (N = 29 989) |
|---|---|
| NBS center location | |
| Eastern Chinaa | 9978 (33.3) |
| Northern Chinab | 8148 (27.2) |
| Southern Chinac | 6055 (20.2) |
| Western Chinad | 5808 (19.4) |
| Sex | |
| Male | 15 357 (51.2) |
| Female | 14 226 (47.4) |
| Unknown | 406 (1.4) |
| Gestational age | |
| Preterm (<37 wk) | 1309 (4.4) |
| Early term (37-38 wk) | 9166 (30.6) |
| Full term (39-40 wk) | 16 476 (54.9) |
| Late term (41 wk) | 1756 (5.9) |
| Post term (≥42 wk) | 40 (0.1) |
| Unknown | 1242 (4.1) |
| Birth weight | |
| ≤2499 g | 974 (3.2) |
| 2500-4000 g | 26 361 (87.9) |
| >4000 g | 1429 (4.8) |
| Unknown | 1225 (4.1) |
Xinhua Hospital affiliated to Shanghai Jiaotong University School of Medicine; Jinan Maternity and Child Care Hospital.
Shijiazhuang Maternal and Child Health Care Hospital; Inner Mongolia Maternity and Child Health Care Hospital.
Guangzhou Woman and Children’s Medical Center; Hainan Woman and Children’s Medical Center.
Chongqing Health Center for Women and Children; the First People’s Hospital of Yunnan Province.
DBS-Based Biochemical Newborn Screening
The DBS-based biochemical NBS included 3 tests: a fluorometric assay for glucose-6-phosphate dehydrogenase (G6PD) deficiency, a thyroid-stimulating hormone (TSH) test for congenital hypothyroidism and isolated hyperthyrotropinemia (HTT), and a tandem mass spectrometry test for 38 amino acid disorders, organic acid disorders, and fatty acid oxidation disorders (PerkinElmer).
Selection of Diseases and Genes
Following the principles laid out by Wilson and Jungner,13 diseases were selected if they met 1 of the following criteria: (1) diseases in the current NBS program in China, (2) diseases on the first national list of rare diseases in China,14 (3) diseases on the list of major rare diseases in Shanghai (2016 Edition),15 (4) diseases in the NBS program recommended by the American College of Medical Genetics and Genomics,16 or (5) treatable inherited disorders with a high carrier frequency in Southeast Asia.17 Genes linked with these diseases were retrieved from the Online Mendelian Inheritance in Man database.18 In total, 128 diseases and 142 related genes were selected. Of note, 85 diseases involving metabolic, endocrine, hematological, immune, neuromuscular, urologic, and ophthalmologic systems were included only in the gene panel, while 43 diseases were included in both the biochemical panel and the gene panel (eTable 1 in Supplement 1).
Gene Panel–Based Newborn Screening
Genomic DNA was extracted from the DBS using a tissue DNA extraction kit (Magen). DNA fragmentation, end repair, and 3′-end tailing were performed using a fragmentation module (Vazyme), followed by enrichment using a custom-designed panel targeting the coding regions of the 142 genes (Integrated DNA Technologies). The library was sequenced with a minimum depth of 100× coverage on a genetic sequencer (MGI). An in-house verified variant calling pipeline was used to analyze single nucleotide variants, small insertions and deletions, and copy number variants (CNV) including CNVs involving 2 or more continuous exons in DMD, exon 7 deletion of SMN1 and common CNVs involving HBA1/HBA2/HBB.19,20,21
Variants meeting 1 of the following criteria were prioritized: (1) pathogenic or likely pathogenic in ClinVar,22 (2) disease-causing or probably disease-causing in the Human Gene Mutation Database,23 or (3) nonsense, frameshift, or canonical of 1 or 2 splice sites or initiation codon variants in genes with a known mechanism of loss-of-function and with an allele frequency of 1% or less in population databases (GnomAD, ESP6500, and 1000 Genomes).24 Prioritized variants with a matching inheritance pattern (ie, heterozygous variants in genes associated with dominant conditions, bi-allelic variants in genes associated with recessive conditions) were further reviewed manually based on the variant interpretation guidelines recommended by American College of Medical Genetics and Genomics.25 Only curated pathogenic or likely pathogenic variants were reported. A genetically positive result was defined as pathogenic or likely pathogenic variants with a matching inheritance pattern.
Follow-Up and Medical Diagnoses
Infants who were screened positive by either biochemical or genetic NBS were followed up by recalling back or phone calls until diagnosis or censoring on July 5, 2022. For infants with genetic NBS positive results, diagnosis decisions were made if they were with corresponding specific confirmatory test results. The confirmatory tests varied for different diseases and included imaging examinations or biochemical tests such as complete blood count, creatine kinase, kidney function, liver function, ammonia, enzymatic activities, and urine organic acids. For infants with positive biochemical NBS results, diagnosis decisions were made if they were consistent with genetic NBS results or routine confirmatory test results.
Statistical Analysis
For categorical data, summary data were reported as frequencies and percentages (eAppendix 1 in Supplement 1). The Fisher exact test or χ2 was performed for comparative analysis. A 2-sided P value of less than .05 was considered statistically significant. Statistical analyses were performed with SPSS version 24 (IBM Corp), and 95% CIs were calculated using a binomial distribution.
Results
Study Population
A total of 29 989 infants were recruited from 8 NBS centers throughout China (15 357 male [51.2%]). The mean (SD) gestational week was 39.0 (1.5) weeks, and the mean (SD) birth weight was 3273 (457) g (Table 1). Among them, 29 601 (98.7%) had biochemical and genetic NBS screening results, while the other 388 (1.3%) failed the quality control of genetic NBS.
Overview of the Genetic NBS and Follow-Ups
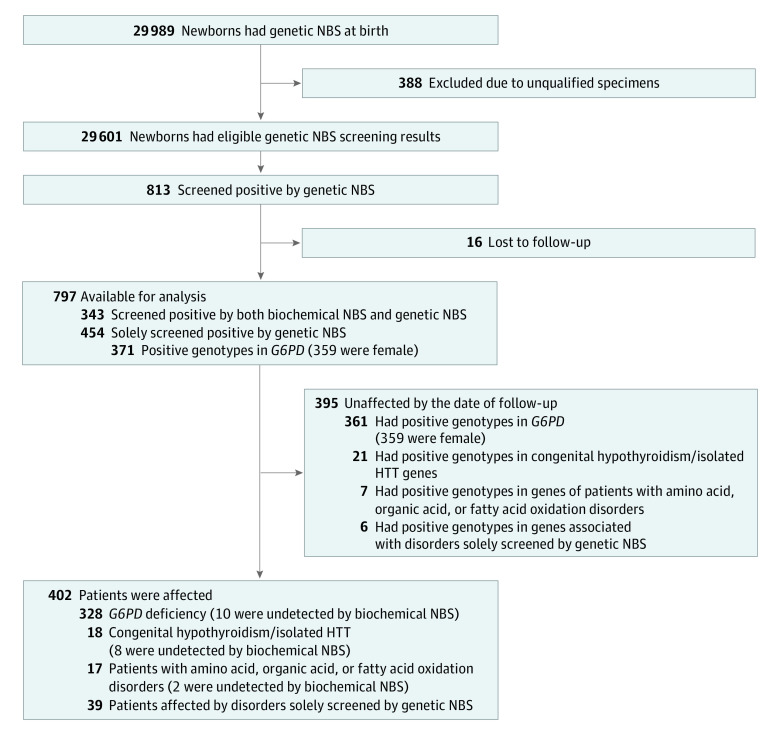
Of the 29 601 newborns with screening results, 820 initially screened positive. To confirm the origin of the variants and eliminate carriers with variants in cis, pedigree analysis was conducted. Among the newborns who had 2 pathogenic variants in recessive genes, samples were successfully collected from parents of 55 newborns. Out of these, 7 newborns were identified as carriers of variants in cis. Finally, 813 (2.7%; 95% CI, 2.6%-2.9%) were screened positive by genetic NBS (Figure). Excluding 16 newborns (2.0%) who were lost to follow-up, 797 newborns (98.0%) were available for further analysis. Upon follow-up at a median age of 1.2 years, 402 infants (1.4%; 95% CI, 1.2%-1.5%) were diagnosed with 26 different diseases. Of note, genetic NBS detected 59 affected patients who were undetected by biochemical NBS. Specifically, 20 patients were affected by disorders screened by both biochemical and genetic NBS (10 G6PD deficiency; 8 congenital hypothyroidism or isolated HTT; 2 amino acid, organic acid, or fatty acid oxidation disorders), whereas 39 patients were affected by disorders screened solely by genetic NBS (Figure). These patients received timely and specific clinical management, including dietary plans, drug therapy, vitamin injections, and systemic chemotherapy (eTables 2-4 and eTable 6 in Supplement 1).
Figure. Study Flowchart.
G6PD indicates glucose-6-phosphate dehydrogenase; HTT, hyperthyrotropinemia; NBS, newborn screening.
Patients Diagnosed With Disorders Screened by Both Genetic and Biochemical NBS
A total of 445 newborns were diagnosed with 13 disorders screened by both genetic and biochemical NBS (Table 2), which included G6PD deficiency (328 newborns), congenital hypothyroidism and isolated HTT (94 newborns), and amino acid, organic acid, or fatty acid oxidation disorders (23 newborns). While 82 of these patients were undetected by genetic NBS (and detected by biochemical NBS), it was notable that 20 of these patients were detected by genetic NBS (while undetected by biochemical NBS).
Table 2. Patients With G6PD Deficiency, Congenital Hypothyroidism, Isolated HTT and Amino Acid, Organic Acid, and Fatty Acid Oxidation Disorders.
| Diseases | Cases detected by both (n = 343) | Undetected cases | All (N = 445) | |
|---|---|---|---|---|
| Biochemical NBS (n = 20) | Genetic NBS (n = 82) | |||
| G6PD deficiency | 318 | 10 | 0 | 328 |
| Male | 272 | 10 | 0 | 282 |
| Female | 46 | 0 | 0 | 46 |
| Congenital hypothyroidism and isolated HTT | 10 | 8 | 76 | 94 |
| Congenital hypothyroidism | 4 | 2 | 24 | 30 |
| Isolated HTT | 6 | 6 | 52 | 64 |
| Amino acids, organic acids, and fatty acid oxidation disorders | 15 | 2 | 6 | 23 |
| Methylmalonic acidemia, cbIC Type | 4 | 1 | 0 | 5 |
| Primary carnitine deficiency | 4 | 0 | 1 | 5 |
| Phenylketonuria | 3 | 0 | 0 | 3 |
| Isobutyryl-CoA dehydrogenase deficiency | 1 | 0 | 1 | 2 |
| Short chain acyl-CoA dehydrogenase deficiency | 1 | 0 | 1 | 2 |
| Medium-chain acyl-coenzyme A dehydrogenase deficiency | 1 | 0 | 0 | 1 |
| Methylmalonic acidemia, cbIA type | 1 | 0 | 0 | 1 |
| Carnitine palmitoyltransferase II deficiency | 0 | 0 | 1 | 1 |
| Maple syrup urine disease, type II | 0 | 0 | 1 | 1 |
| Multiple acyl-CoA dehydrogenase deficiency | 0 | 0 | 1 | 1 |
| Citrin deficiency | 0 | 1 | 0 | 1 |
Abbreviations: HTT, hyperthyrotropinemia; NBS, newborn screening.
G6PD Deficiency
A total of 328 patients with G6PD deficiency (282 male and 46 female) were identified (Table 2). All were detected by genetic NBS, but 10 male patients were not detected by biochemical NBS; instead, they were confirmed in the confirmatory tests (Figure; eTable 2 in Supplement 1). Overall, the fluorometric assay had a sensitivity of 96.95% (95% CI, 94.48%-98.33%) and a specificity of 99.71% (95% CI, 99.64%-99.77%), whereas genetic NBS had a sensitivity of 100% (95% CI, 98.84%-100%) and a specificity of 98.61% (95% CI, 98.46%-98.75%) for G6PD deficiency (Table 3). It is worth noting that the overall positive predictive value (PPV) of genetic NBS was 47.61% for G6PD deficiency (99.30% for male, and 11.36% for female), as the majority of female newborns were unaffected (Figure).
Table 3. Analytic Performance of Biochemical and Genetic NBS for Disorders in Newborns.
| Parameters | Biochemical NBS, newborns, No. (%) | Genetic NBS, newborns, No. (%) | ||||||
|---|---|---|---|---|---|---|---|---|
| G6PD deficiency | Amino acid, organic acid, and fatty acid oxidation disorders | G6PD deficiency | Amino acid, organic acid, and fatty acid oxidation disorders | |||||
| Malea | Femalea | Alla,b | Malea | Femalea | Alla,b | |||
| Newborns with inconclusive resultsc | 0 | 2 | 2 | 146 | 0 | 0 | 0 | 1 |
| Conclusive results | ||||||||
| True negative | 13 343 | 12 601 | 25 962 | 29 055 | 13 381 | 12 278 | 25 677 | 29 424 |
| True positive | 272 | 46 | 318 | 21 | 282 | 46 | 328 | 17 |
| False positive | 40 | 36 | 76 | 376 | 2 | 359 | 361 | 7 |
| False negative | 10 | 0 | 10 | 2 | 0 | 0 | 0 | 6 |
| Sensitivity, % (95% CI) | 96.45 (93.59-98.06) | 100 (92.29-100) | 96.95 (94.48-98.33) | 91.30 (73.20-97.58) | 100 (98.66-100) | 100 (92.29-100) | 100 (98.84-100) | 73.91 (53.53-87.45) |
| Specificity, % (95% CI) | 99.70 (99.59-99.78) | 99.72 (99.61-99.80) | 99.71 (99.64-99.77) | 98.72 (98.59-98.84) | 99.99 (99.95-100.00) | 97.16 (96.86-97.44) | 98.61 (98.46-98.75) | 99.98 (99.96-99.99) |
| PPV, % (95% CI) | 87.18 (83.01-90.44) | 56.10 (45.33-66.33) | 80.71 (76.52-84.30) | 5.29 (3.49-7.95) | 99.30 (97.48-99.81) | 11.36 (8.63-14.82) | 47.61 (43.90-51.34) | 70.83 (50.83-85.08) |
| FNR, % (95% CI) | 3.55 (1.94-6.41) | 0 (0-7.71) | 3.05 (1.67-5.52) | 8.70 (2.42-26.80) | 0 (0-1.34) | 0 (0-7.71) | 0 (0-1.16) | 26.09 (12.55-46.47) |
Abbreviations: FNR, false-negative rate; G6PD, glucose-6-phosphate dehydrogenase; NBS, newborn screening; PPV, positive predictive value.
Biochemical NBS (G6PD screening) was not performed in 3233 newborns from Inner Mongolia Maternity and Child Health Care Hospital consisting of 1692 male and 1541 female.
18 newborns were with unknown sex and were true negative for G6PD deficiency.
Includes newborns with positive results who were lost to follow-up.
Congenital Hypothyroidism and Isolated HTT
The targeted gene panel included 4 genes related to congenital hypothyroidism (DUOXA2, DUOX2, PROP1, TSHR). Gene panel sequencing detected 18 patients (Table 2), and all had DUOX2 variants. They were clinically diagnosed with congenital hypothyroidism (6 patients) and isolated HTT (12 patients). Of note, 8 out of the 18 patients were screened negative by TSH tests (eTable 3 in Supplement 1). In addition, TSH tests identified 76 patients (24 congenital hypothyroidism and 52 isolated HTT) who were undetected by genetic NBS. Overall, screening for the 4 genes identified 19.1% (18 of 94 patients) with congenital hypothyroidism and isolated HTT.
Amino Acid, Organic Acid, or Fatty Acid Oxidation Disorders
A total of 23 newborns were diagnosed with amino acid, organic acid, or fatty acid oxidation disorders by either tandem mass spectrometry or genetic NBS. Of these, 2 were undetected by tandem mass spectrometry while 6 were undetected by genetic NBS (Table 2). The sensitivity and specificity of tandem mass spectrometry for amino acid, organic acid, or fatty acid oxidation disorders were 91.30% (95% CI, 73.20%-97.58%) and 98.72% (95% CI, 98.59%-98.84%), respectively, whereas the sensitivity and specificity of genetic NBS were 73.91% (95% CI, 53.53%-87.45%) and 99.98% (95% CI, 99.96%-99.99%), respectively. The PPV of genetic NBS was 70.83% (95% CI, 50.83%-85.08%), significantly higher than that of tandem mass spectrometry (5.29%; 95% CI, 3.49%-7.95%) (P < .001) (Table 3). The 2 patients not detected by tandem mass spectrometry but revealed by genetic NBS were associated with cbIC-type methylmalonic acidemia and citrin deficiency (eTable 4 in Supplement 1). Among the 6 patients not detected by genetic NBS but revealed by tandem mass spectrometry, indication-based analysis revealed candidate variants in 5 patients (eTable 5 in Supplement 1).
Patients Diagnosed With Disorders Screened Solely by Genetic NBS
A total of 55 genetically positive infants were identified with disorders solely screened by genetic NBS. Of 45 infants who had successful follow-ups, 39 (87%; 95% CI, 73%-94%) had presented clinical symptoms of 13 disorders at a median age of 4.8 months (eTable 6 in Supplement 1). The disorders could be grouped into 7 types: blood disorder (thalassemia, 15 patients), neuromuscular disorder (7 patients), lysosomal storage disorder (10 patients), glycolipid metabolic disorder (1 patient), other metabolic disorder (4 patients), eye disorder (1 patient), and multisystem disorder (1 patient) (Table 4). Five of the infants received timely medical treatment, 4 were scheduled for further treatment with specialists, 27 were counseled to have active surveillance, and 3 were lost to follow-up after clinical diagnosis.
Table 4. Characteristics of Patients Affected by Disorders Screened Solely by Genetic Newborn Screening.
| Group | Total patients, No. | Lost to follow-up | Asymptomatic | Symptomatic | Symptomatic rate, % (95% CI) |
|---|---|---|---|---|---|
| Total | 55 | 10 | 6 | 39 | 87 (73-94) |
| Blood disorder | |||||
| α-Thalassemia | 13 | 1 | 0 | 12 | 100 (70-100) |
| β-Thalassemia | 3 | 0 | 0 | 3 | 100 (31-100) |
| Neuromuscular disorder | |||||
| Duchenne/Becker muscular dystrophy | 9 | 3 | 0 | 6 | 100 (52-100) |
| Early infantile epileptic encephalopathy 6 | 1 | 0 | 0 | 1 | 100 (6-100) |
| Spinal muscular atrophy | 1 | 0 | 1 | 0 | 0 |
| Lysosomal storage disorder | |||||
| Niemann-Pick disease, type A/B | 6 | 1 | 0 | 5 | 100 (46-100) |
| Krabbe disease | 4 | 1 | 1 | 2 | 67 (13-98) |
| Fabry disease | 3 | 0 | 0 | 3 | 100 (31-100) |
| Glycolipid metabolic disorder | |||||
| Glycogen storage disease type IXa | 2 | 1 | 1 | 0 | 0 |
| Glycogen storage disease type IXb | 1 | 0 | 1 | 0 | 0 |
| Glycogen storage disease type Ib | 1 | 0 | 0 | 1 | 100 (6-100) |
| Other metabolic disorder | |||||
| Wilson disease | 3 | 1 | 0 | 2 | 100 (20-100) |
| Hypophosphatasia | 1 | 0 | 1 | 0 | 0 (0-95) |
| X-linked hypophosphatemia | 1 | 0 | 0 | 1 | 100 (6-100) |
| Cerebrotendinous xanthomatosis | 1 | 0 | 0 | 1 | 100 (6-100) |
| Eye disorder | |||||
| Retinoblastoma | 1 | 0 | 0 | 1 | 100 (6-100) |
| Urologic disease | |||||
| Gitelman syndrome | 1 | 0 | 1 | 0 | 0 |
| Alport syndrome 1, X-linked | 1 | 1 | 0 | 0 | NA |
| Multisystem disorder | |||||
| Tuberous sclerosis 1 | 2 | 1 | 0 | 1 | 100 (6-100) |
Abbreviation: NA, not applicable.
Newborns With Suspect Diseases
By the follow-up date, a total of 395 infants (1.3%) who had been screened positive by genetic NBS were unaffected (Figure). Excluding 359 females with G6PD deficiency, the remaining 36 infants were positive for thyroid dyshormonogenesis 6 (21 patients), phenylketonuria (4 patients), G6PD deficiency (2 male patients), glycogen storage disease (2 patients), 2-methylbutyryl glycinuria (1 patient), ornithine transcarbamylase deficiency (1 patient), hyperprolinemia type I (1 patient), spinal muscular atrophy (1 patient), Krabbe disease (1 patient), hypophosphatasia (1 patient), and Gitelman syndrome (1 patient) (eTable 7 in Supplement 1). Further investigation revealed genetic explanations for the unaffected status of three cases. Although pathogenic variants in G6PD (X-linked inheritance) were identified, 1 male infant was confirmed to have an XXY karyotype, and a second male infant harbored a variant c.1386G>T located on exon 12 which was duplicated. The extra copy may play a key role in normal G6PD function. The third case was a homozygous deletion of exon 7 in SMN1, associated with spinal muscular atrophy. Follow-up tests revealed 3 copies of the SMN2 gene, which is a genetic modifier that can decrease the severity.26 The remaining 33 cases may be attributed to childhood onset or mild phenotypes.
Discussion
To investigate the clinical impact of including genomic sequencing as a first-tier screen test in existing NBS programs, we recruited nearly 30 000 newborns in a prospective multicenter design to perform genetic and biochemical NBS concurrently. To our knowledge, this is the largest prospective study to assess the potential benefits of a genomic test as a first-tier screening approach at a population scale.
Our prospective cohort study revealed that gene panel sequencing detected 402 patients out of 29 601 newborns. Of these, 59 patients were undetected by biochemical NBS, including 20 patients affected by disorders screened by both biochemical and genetic NBS and 39 patients affected by disorders screened solely by genetic NBS. This translates into 1 in every 500 newborns who would benefit from the application of genetic NBS, thus receiving timely clinical intervention and management. Therefore, genetic NBS as a first-tier screening test is a promising method to improve the current NBS program.
Another benefit of incorporating genetic with biochemical NBS is to enrich the number of cases screened positive. Our data demonstrate that the PPV of genetic NBS is significantly higher than that of biochemical NBS, except for the fluorometric assay for G6PD in female newborns. The difference was highly apparent for amino acid, organic acid, and fatty acid oxidation disorders (70.83% by genetic NBS vs 5.29% by tandem mass spectrometry; P < .001), which is in line with previous studies.27,28 Reducing the number of false positives would, in turn, relieve the stress and anxiety of parents and decrease the number of follow-ups.29,30
Previous studies primarily focused on the performance of genetic sequencing of inborn errors of metabolism disorders that had been screened by tandem mass spectrometry.6,8,31 There was little investigation of the clinical impact of genetic sequencing for G6PD deficiency, which is important in neonates and prevalent in some regions.1 Our results provide valuable insights on the performance of genetic screening for these common disorders. Specifically, genetic NBS demonstrated a sensitivity of 100% in screening G6PD deficiency, whereas the sensitivity of the fluorometric assay is only 96.95%. Notably, the PPV of genetic NBS was 47.61% and was significantly affected by sex (99.30% in male and 11.36% in female patients). This was mainly attributable to the fact that most female newborns carrying a heterozygous variant in G6PD exhibit normal G6PD activity due to skewed X-chromosome inactivation.32 Together, these data indicate that genetic screening had the potential to supplement biochemical NBS in detection capability but cannot replace it as a universal NBS program, especially for disorders with variable penetrance.
One insight we wish to emphasize is that follow-up tests are crucial in the implementation of genetic NBS in clinical practice. Benefitting from the prospective study design, we carried out pedigree analysis when applicable. Out of 55 cases harboring 2 pathogenic variants in recessive genes, we found that variants in 12.7% (7 of 55) of the cases were in cis configuration. These were ruled out as genetically positive cases. Knowing the phase and origin of pathogenic variants facilitates clinical counseling and is of value for family members for future reproductive planning.
Whole exome or genome sequencing are promising candidates for genomic sequencing tests.6,31 However, their use as a universal screening test in clinical applications is hindered by their high cost. Gene panels targeting genomic regions of interest offer a compromise between cost and the number of genes, thus serving as an alternative solution.30 In this study, we used a gene panel as a first-tier screening test. It demonstrated a sensitivity of 73.91% and a specificity of 99.98% for screening amino acid, organic acid, and fatty acid oxidation disorders in a general newborn population. Although the screening disorders varied between different studies, the performance of the gene panel was comparable with that of whole exome sequencing, which had an overall sensitivity of 88% and specificity of 98.4%.6
Limitations
There are limitations to note about this study. First, newborns who were screened negative were not proactively followed, which may weaken the calculation of the sensitivity and specificity. However, we argue that this is a minor limitation because all the participants were independently and concurrently screened by the biochemical and genetic tests. Second, it was difficult to recruit a representative cohort for the Chinese population as the vast population size and significant heterogeneity across regions. However, we have recruited newborns from local NBS centers in different regions to decrease the influence. The third limitation is that the cost-effectiveness of a gene panel as a first-tier screening test was not evaluated, although we note that gene panels have an advantage in cost compared with whole exome or whole genome sequencing. Cost-effectiveness analysis is warranted in future. Nevertheless, such an analysis is crucial before these tests could be used as a universal screening test in the general newborn population.
Conclusions
In this cohort study, 1 in 500 newborns could additionally benefit from incorporating gene panel sequencing of 142 genes into a traditional, population-scale NBS program as a first-tier screening test, demonstrating that genetic NBS supplemented biochemical NBS by improving detection capability and screening more patients. This provides an evidence-based suggestion that genomic screening could be considered as a crucial method for first-tier screening.
eAppendix. Statistical Methods Used for Calculating Sensitivity, Specificity, PPV and FNR
eTable 1. Diseases and Genes in Genomic Newborn Screening
eTable 2. G6PD-Deficient Patients Undetected by the Initial G6PD Screening but Confirmed by Confirmatory Tests
eTable 3. TSH-Related Patients Undetected by the Initial TSH Screening but Confirmed by Confirmatory Tests
eTable 4. Amino Acid, Organic Acid, or Fatty Acid Oxidation Disorders Patients Undetected by Tandem Mass Spectrometry but Confirmed by Confirmatory Tests
eTable 5. Indication-Based Analysis of Six Amino Acid, Organic Acid, or Fatty Acid Oxidation Disorders Patients Undetected by the Gene Panel
eTable 6. Characteristics of 39 Patients Affected Disorders Screened Solely by Genetic NBS
eTable 7. Characteristics of 36 Unaffected Cases
Data Sharing Statement
References
- 1.Therrell BL, Padilla CD, Loeber JG, et al. Current status of newborn screening worldwide: 2015. Semin Perinatol. 2015;39(3):171-187. doi: 10.1053/j.semperi.2015.03.002 [DOI] [PubMed] [Google Scholar]
- 2.van Spronsen FJ, Blau N, Harding C, Burlina A, Longo N, Bosch AM. Phenylketonuria. Nat Rev Dis Primers. 2021;7(1):36. doi: 10.1038/s41572-021-00267-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Chace DH, Kalas TA, Naylor EW. The application of tandem mass spectrometry to neonatal screening for inherited disorders of intermediary metabolism. Annu Rev Genomics Hum Genet. 2002;3:17-45. doi: 10.1146/annurev.genom.3.022502.103213 [DOI] [PubMed] [Google Scholar]
- 4.Kwon C, Farrell PM. The magnitude and challenge of false-positive newborn screening test results. Arch Pediatr Adolesc Med. 2000;154(7):714-718. doi: 10.1001/archpedi.154.7.714 [DOI] [PubMed] [Google Scholar]
- 5.Biesecker LG, Green RC. Diagnostic clinical genome and exome sequencing. N Engl J Med. 2014;370(25):2418-2425. doi: 10.1056/NEJMra1312543 [DOI] [PubMed] [Google Scholar]
- 6.Adhikari AN, Gallagher RC, Wang Y, et al. The role of exome sequencing in newborn screening for inborn errors of metabolism. Nat Med. 2020;26(9):1392-1397. doi: 10.1038/s41591-020-0966-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Schwarze K, Buchanan J, Taylor JC, Wordsworth S. Are whole-exome and whole-genome sequencing approaches cost-effective? A systematic review of the literature. Genet Med. 2018;20(10):1122-1130. doi: 10.1038/gim.2017.247 [DOI] [PubMed] [Google Scholar]
- 8.Bodian DL, Klein E, Iyer RK, et al. Utility of whole-genome sequencing for detection of newborn screening disorders in a population cohort of 1,696 neonates. Genet Med. 2016;18(3):221-230. doi: 10.1038/gim.2015.111 [DOI] [PubMed] [Google Scholar]
- 9.Pavey AR, Bodian DL, Vilboux T, et al. Utilization of genomic sequencing for population screening of immunodeficiencies in the newborn. Genet Med. 2017;19(12):1367-1375. doi: 10.1038/gim.2017.57 [DOI] [PubMed] [Google Scholar]
- 10.Ceyhan-Birsoy O, Murry JB, Machini K, et al. ; BabySeq Project Team . Interpretation of genomic sequencing results in healthy and ill newborns: results from the BabySeq Project. Am J Hum Genet. 2019;104(1):76-93. doi: 10.1016/j.ajhg.2018.11.016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Huang X, Wu D, Zhu L, et al. Application of a next-generation sequencing (NGS) panel in newborn screening efficiently identifies inborn disorders of neonates. Orphanet J Rare Dis. 2022;17(1):66. doi: 10.1186/s13023-022-02231-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Jian M, Wang X, Sui Y, et al. A pilot study of assessing whole genome sequencing in newborn screening in unselected children in China. Clin Transl Med. 2022;12(6):e843. doi: 10.1002/ctm2.843 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Andermann A, Blancquaert I, Beauchamp S, Déry V. Revisiting Wilson and Jungner in the genomic age: a review of screening criteria over the past 40 years. Bull World Health Organ. 2008;86(4):317-319. doi: 10.2471/BLT.07.050112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.He J, Kang Q, Hu J, Song P, Jin C. China has officially released its first national list of rare diseases. Intractable Rare Dis Res. 2018;7(2):145-147. doi: 10.5582/irdr.2018.01056 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Yang Y, Kang Q, Hu J, et al. Accessibility of drugs for rare diseases in China: policies and current situation. Intractable Rare Dis Res. 2019;8(2):80-88. doi: 10.5582/irdr.2019.01068 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Fabie NAV, Pappas KB, Feldman GL. The current state of newborn screening in the United States. Pediatr Clin North Am. 2019;66(2):369-386. doi: 10.1016/j.pcl.2018.12.007 [DOI] [PubMed] [Google Scholar]
- 17.Bylstra Y, Kuan JL, Lim WK, et al. ; SinGapore Incidental Finding (SGIF) study group . Population genomics in South East Asia captures unexpectedly high carrier frequency for treatable inherited disorders. Genet Med. 2019;21(1):207-212. doi: 10.1038/s41436-018-0008-6 [DOI] [PubMed] [Google Scholar]
- 18.Hamosh A, Scott AF, Amberger JS, Bocchini CA, McKusick VA. Online Mendelian Inheritance in Man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res. 2005;33(Database issue):D514-D517. doi: 10.1093/nar/gki033 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Wei X, Dai Y, Yu P, et al. Targeted next-generation sequencing as a comprehensive test for patients with and female carriers of DMD/BMD: a multi-population diagnostic study. Eur J Hum Genet. 2014;22(1):110-118. doi: 10.1038/ejhg.2013.82 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Shang X, Peng Z, Ye Y, et al. Rapid targeted next-generation sequencing platform for molecular screening and clinical genotyping in subjects with hemoglobinopathies. EBioMedicine. 2017;23:150-159. doi: 10.1016/j.ebiom.2017.08.015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Zhao S, Wang W, Wang Y, et al. NGS-based spinal muscular atrophy carrier screening of 10,585 diverse couples in China: a pan-ethnic study. Eur J Hum Genet. 2021;29(1):194-204. doi: 10.1038/s41431-020-00714-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Landrum MJ, Chitipiralla S, Brown GR, et al. ClinVar: improvements to accessing data. Nucleic Acids Res. 2020;48(D1):D835-D844. doi: 10.1093/nar/gkz972 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Stenson PD, Mort M, Ball EV, et al. The Human Gene Mutation Database: towards a comprehensive repository of inherited mutation data for medical research, genetic diagnosis and next-generation sequencing studies. Hum Genet. 2017;136(6):665-677. doi: 10.1007/s00439-017-1779-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Karczewski KJ, Francioli LC, Tiao G, et al. ; Genome Aggregation Database Consortium . The mutational constraint spectrum quantified from variation in 141,456 humans. Nature. 2020;581(7809):434-443. doi: 10.1038/s41586-020-2308-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Richards S, Aziz N, Bale S, et al. ; ACMG Laboratory Quality Assurance Committee . Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17(5):405-424. doi: 10.1038/gim.2015.30 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Prior TW, Krainer AR, Hua Y, et al. A positive modifier of spinal muscular atrophy in the SMN2 gene. Am J Hum Genet. 2009;85(3):408-413. doi: 10.1016/j.ajhg.2009.08.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Hall PL, Marquardt G, McHugh DM, et al. Postanalytical tools improve performance of newborn screening by tandem mass spectrometry. Genet Med. 2014;16(12):889-895. doi: 10.1038/gim.2014.62 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Messina M, Meli C, Raudino F, et al. Expanded newborn screening using tandem mass spectrometry: seven years of experience in Eastern Sicily. Int J Neonatal Screen. 2018;4(2):12. doi: 10.3390/ijns4020012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Tu WJ, He J, Chen H, Shi XD, Li Y. Psychological effects of false-positive results in expanded newborn screening in China. PLoS One. 2012;7(4):e36235. doi: 10.1371/journal.pone.0036235 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Woerner AC, Gallagher RC, Vockley J, Adhikari AN. The use of whole genome and exome sequencing for newborn screening: challenges and opportunities for population health. Front Pediatr. 2021;9:663752. doi: 10.3389/fped.2021.663752 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Roman TS, Crowley SB, Roche MI, et al. Genomic sequencing for newborn screening: results of the NC NEXUS Project. Am J Hum Genet. 2020;107(4):596-611. doi: 10.1016/j.ajhg.2020.08.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Sudsumrit S, Chamchoy K, Songdej D, et al. Genotype-phenotype association and biochemical analyses of glucose-6-phosphate dehydrogenase variants: implications for the hemolytic risk of using 8-aminoquinolines for radical cure. Front Pharmacol. 2022;13:1032938. doi: 10.3389/fphar.2022.1032938 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
eAppendix. Statistical Methods Used for Calculating Sensitivity, Specificity, PPV and FNR
eTable 1. Diseases and Genes in Genomic Newborn Screening
eTable 2. G6PD-Deficient Patients Undetected by the Initial G6PD Screening but Confirmed by Confirmatory Tests
eTable 3. TSH-Related Patients Undetected by the Initial TSH Screening but Confirmed by Confirmatory Tests
eTable 4. Amino Acid, Organic Acid, or Fatty Acid Oxidation Disorders Patients Undetected by Tandem Mass Spectrometry but Confirmed by Confirmatory Tests
eTable 5. Indication-Based Analysis of Six Amino Acid, Organic Acid, or Fatty Acid Oxidation Disorders Patients Undetected by the Gene Panel
eTable 6. Characteristics of 39 Patients Affected Disorders Screened Solely by Genetic NBS
eTable 7. Characteristics of 36 Unaffected Cases
Data Sharing Statement