Significance
Aminoacyl-tRNA synthetases (aaRSs) ensure both speed and accuracy of messenger RNA (mRNA) translation. Amino acid misincorporation due to loss of editing of cytoplasmic aaRSs is harmful to cellular functions and causes neurodegeneration and cardioproteinopathy. However, mitochondrial translation in mammals is the most simplified, generating only 13 proteins with 3,789 codons. Several mitochondrial aaRSs have lost editing activity during evolution, implying that mammalian mitochondrial protein synthesis is able to tolerate mistranslation. Based on our comprehensive studies using mouse mitochondrial threonyl-tRNA synthetase as a model, we found that the correct amino acid incorporation during mitochondrial gene expression is essential for the structure and function of the organelle. Inaccurate translation triggers profound cellular and tissue disorders due to abnormal cell proliferation induced by oxidative stress.
Keywords: aminoacyl-tRNA synthetase, tRNA, editing, mitochondria, cardiomyopathy
Abstract
Proofreading (editing) of mischarged tRNAs by cytoplasmic aminoacyl-tRNA synthetases (aaRSs), whose impairment causes neurodegeneration and cardiac diseases, is of high significance for protein homeostasis. However, whether mitochondrial translation needs fidelity and the significance of editing by mitochondrial aaRSs have been unclear. Here, we show that mammalian cells critically depended on the editing of mitochondrial threonyl-tRNA synthetase (mtThrRS, encoded by Tars2), disruption of which accumulated Ser-tRNAThr and generated a large abundance of Thr-to-Ser misincorporated peptides in vivo. Such infidelity impaired mitochondrial translation and oxidative phosphorylation, causing oxidative stress and cell cycle arrest in the G0/G1 phase. Notably, reactive oxygen species (ROS) scavenging by N-acetylcysteine attenuated this abnormal cell proliferation. A mouse model of heart-specific defective mtThrRS editing was established. Increased ROS levels, blocked cardiomyocyte proliferation, contractile dysfunction, dilated cardiomyopathy, and cardiac fibrosis were observed. Our results elucidate that mitochondria critically require a high level of translational accuracy at Thr codons and highlight the cellular dysfunctions and imbalance in tissue homeostasis caused by mitochondrial mistranslation.
mRNA translation is a key step in the expression of genetic codes. Aminoacyl-tRNA synthetases (aaRSs) are ubiquitously expressed and essential for mRNA translation, catalyzing the ligation of amino acids (aminoacylation) with tRNAs to generate aminoacyl-tRNAs (1, 2). Protein synthesis shows a very high level of fidelity (approximately 1 error in every 10,000 codons) (3). Achieving such a high degree of fidelity is challenging for some aaRSs, as large amounts of structurally and chemically similar amino acids and metabolites are present in cells. To overcome this difficulty, approximately one-half of all aaRSs have evolved a proofreading (editing) function that enables clearance of mischarged tRNAs (4–7). Disruption or failure of aaRS editing function due to mutations in the editing domain results in translation errors due to amino acid misincorporation, e.g., the generation of pools of statistical and heterogeneous proteins, which could be detrimental to protein folding and function. Numerous studies have shown that editing is critical for optimal cell growth and cell viability in bacteria and eukaryotes (8–13).
Mammalian cells harbor two gene expression systems, one in the cytoplasm and one in mitochondria. Mitochondrial DNA (mtDNA) is a double-stranded circular DNA sequence of 16,569 bp and encodes 37 genes, including 22 tRNAs, 2 rRNAs, and 11 mRNAs, producing only 13 mtDNA-encoded proteins (14). Two sets of translation machinery are based on two different sets of aaRSs, which are encoded by 37 nuclear genes (15).
Given the pivotal role of editing in translational accuracy, approximately one-half of cytoplasmic aaRSs have retained intact editing domains in mammalian cells throughout evolution. Indeed, several cytoplasmic aaRSs, including alanyl-tRNA synthetase (ctAlaRS), leucyl-tRNA synthetase, valyl-tRNA synthetase (ctValRS), threonyl-tRNA synthetase (ctThrRS), and ThrRS-like protein (ctThrRS-L), have been reported to exhibit posttransfer editing activity in vitro (8, 9, 16–18). An apoptotic response was stimulated by editing-deficient mouse ValRS in NIH3T3 cells (8). Similarly, expression of editing-deficient ValRS activated a p53-dependent DNA damage response in cells and shortened the lifespan of zebrafish (19). Notably, mice harboring a single missense mutation in the editing domain of ctAlaRS exhibited severe ataxia accompanied by Purkinje cell degeneration (9). This mutation, A734E, only slightly decreased editing activity of ctAlaRS, by only approximately twofold. Indeed, a more severe reduction in editing function of ctAlaRS caused by the C723A mutation resulted in ubiquitinated protein aggregates and progressive cardiac fibrosis and dysfunction (10). Recently, a vertebrate-specific protein, Ankrd16, was shown to interact with ctAlaRS to accept misactivated Ser and thus to prevent mistranslation due to defect in ctAlaRS editing (20). Moreover, cytoplasmic translation involves several ubiquitously expressed trans-editing factors that further promote and ensure fidelity in mRNA translation, adding another layer of quality control (21–23). All these studies highlight the clear necessity to prevent mistranslation of cytoplasmic mRNAs.
In contrast to the clear requirement for cytoplasmic aaRS editing, the requirement for mammalian mitochondrial aaRS editing is unclear. Mitochondrial phenylalanyl-tRNA synthetase (mtPheRS) has lost an editing domain during evolution (24); in addition, mtLeuRS harbors a truncated editing domain, and the editing catalytic site has degenerated (25, 26). In contrast, yeast mtLeuRS has retained an editing function (27). In addition, yeast mitochondrial prolyl-tRNA synthetase (mtProRS) carries an editing domain that has been lost in mammalian mtProRS, which is likely an editing-deficient aaRS. The rationale for the loss of mtPheRS, mtLeuRS, and mtProRS editing function is unclear but suggests the possibility that mitochondrial translation does not require the same level of accuracy as cytoplasmic mRNA translation (28, 29). However, mammalian mitochondrial ThrRS (mtThrRS), isoleucyl-tRNA synthetase (mtIleRS), mtValRS, and mtAlaRS carry intact editing domains and editing active sites (30). Indeed, our in vitro data have clearly shown that mtThrRS and mtAlaRS hydrolyze Ser-mischarged mitochondrial tRNAThr (Ser-mtRNAThr) and Ser/Gly-mischarged mitochondrial tRNAAla, respectively (11, 31, 32). Moreover, loss of the editing function of mouse mtAlaRS caused embryonic lethality, highlighting the essential role played by mtAlaRS editing in embryonic development (11). However, considering that mtDNA contains only 3,789 codons (excluding stop codons) and generates only 13 polypeptides, it is questionable whether such a simple mitochondrial translation requires a high level of precision from aaRSs (whose editing domains are intact).
In this work, we revealed that mtThrRS possesses editing capability. NIH3T3 cells depend on the mtThrRS editing function to prevent the generation of Thr-to-Ser misincorporation into proteins. Using CRISPR/Cas9 editing technology, we established a mutant NIH3T3 cell line expressing editing-defective mtThrRS, which led to impaired mitochondrial translation and oxidative phosphorylation (OXPHOS), triggering oxidative stress and blocking the cell cycle in the G0/G1 phase. Furthermore, we established a heart tissue–specific mouse model with defective mtThrRS editing, and these mice exhibited obvious cardiomyopathy. Our results reveal a critical role of mtThrRS editing in mitochondrial RNA translation and function.
Results
Mouse mtThrRS Catalyzed Posttransfer Editing In Vitro, Which was Reliant on His138 and His142.
Mammalian mtThrRS carries an N1 domain, an N2 editing domain, an aminoacylation domain, and a C-terminal tRNA-binding domain (33) (SI Appendix, Fig. S1A). Through its N2 editing domain, human mtThrRS (encoded by TARS2, hereafter hmtThrRS) catalyzes posttransfer editing of Ser-mtRNAThr in vitro (31). To determine whether mouse mtThrRS (encoded by Tars2, hereafter mmtThrRS) exhibits the same editing activity as its human ortholog, we purified the N-terminal domain (Leu22-Pro287, designated mmtThrRS-N, which includes the N1 and N2 domains) of mmtThrRS based on the primary sequence alignment between hmtThrRS and mmtThrRS and the cleavage site (between Arg19 and Leu20) in the hmtThrRS precursor after its import into mitochondria (31, 34) (SI Appendix, Fig. S1A). Hydrolysis of preformed Ser-mtRNAThr (generated by editing-deficient hmtThrRS-H133A/H137A) (31) clearly showed that mmtThrRS-N performed robust posttransfer editing activity (SI Appendix, Fig. S1 B and C). The hydrolysis of Ser-mtRNAThr depends on two absolutely conserved His residues (His133 and His137) in hmtThrRS, which have been completely conserved in ThrRSs in all domains of life (31). We mutated the His counterparts in mmtThrRS-N2 (His138 and His142) to generate mmtThrRS-N-H138A/H142A (SI Appendix, Fig. S1A). We found that mmtThrRS-N-H138A/H142A similarly lost posttransfer editing of Ser-mtRNAThr (SI Appendix, Fig. S1 B and C). In summary, the biochemical data showed that mmtThrRS exhibited posttransfer editing activity and that the His138 and His142 residues played essential roles in its editing function.
H138A/H142A Mutations Led to Thr-to-Ser Misincorporation into Proteins In Vivo.
mmtThrRS exhibits editing activity in vitro; however, Ser-mtRNAThr hydrolysis has not been demonstrated in vivo, and it is unclear whether mitochondrial protein synthesis requires this proofreading activity. To address these questions, we constructed a mouse NIH3T3 cell line with the His138 and His142 residues simultaneously replaced with Ala residues via CRISPR/Cas9-mediated genome editing (Fig. 1A). This cell line is referred to as the NIH3T3-MU cell line hereafter. The NIH3T3-MU cell line carried the same amount of mtRNAThr as the wild-type NIH3T3 (NIH3T3-WT) cell line, as determined by northern blot under denaturing conditions (SI Appendix, Fig. S2A). Similarly, no change in mtRNAThr charging levels was observed in the northern blot under acidic conditions, which preserved the amino acid–tRNA conjugates (SI Appendix, Fig. S2B). These results showed that mmtThrRS editing deficiency did not influence the steady-state abundance or charging level of mtRNAThr in vivo. However, mtRNAThr charging level data obtained from northern blot assays did not enable the differentiation of amino acid identities (Thr or Ser) on mtRNAThr. To overcome this limitation, mtRNAThr was purified using a tRNA pulldown method based on a solid-phase biotinylated DNA probe designed to be complementary to mtRNAThr under acidic conditions. Then, amino acids were detached from charged mtRNAThr by incubating the samples under alkaline conditions. The identity of the amino acids was determined by liquid chromatography–tandem mass spectrometry (LC–MS/MS) (API 400 Q-TRAP) (SI Appendix, Fig. S2C), which showed that approximately 3.35% of mtRNAThr was incorrectly charged with noncognate Ser in NIH3T3-MU cells, and the level of Ser-mtRNAThr was undetectable in NIH3T3-WT cells (Fig. 1B). To determine whether this mischarged Ser-mtRNAThr had been efficiently incorporated by the mitochondrial protein synthesis apparatus, we pulled down the OXPHOS supercomplex (35) by immunoprecipitation using an anti-Mt-Atp6 antibody directed against mtDNA-encoded adenosine 5'-triphosphate (ATP) synthase membrane subunit 6. The complexes immunoprecipitated from either NIH3T3-WT or NIH3T3-MU cells were then subjected to LC–MS (Q Exactive Orbitrap) analysis to identify the peptides in which Ser residues had been incorporated in the place of Thr codons on the basis of the mass difference between Thr and Ser residues (SI Appendix, Fig. S2D). In the NIH3T3-WT cell line, no peptides with a Thr-to-Ser misincorporation were found in the OXPHOS complex. In contrast, complexes isolated from NIH3T3-MU cells carried a significant level of misincorporated Ser residues. Among the 67 peptides analyzed from 10 mtDNA-encoded proteins, 15 peptides in 7 of the proteins carried erroneously incorporated Ser residue at the Thr position. In total, 16 cases of Ser misincorporation were identified among 81 Thr codons (Fig. 1C and SI Appendix, Fig. S3). Collectively, these data clearly showed that mmtThrRS editing deficiency led to Ser-mtRNAThr, which was used by the mitochondrial protein synthesis machinery, leading to Ser insertion at Thr codons in vivo. These data demonstrated that the editing activity of mmtThrRS is necessary for the fidelity of mitochondrial mRNA translation.
Fig. 1.
mmtThrRS editing defects lead to Thr-to-Ser misincorporation during mitochondrial translation in vivo. (A) Schematic showing the NIH3T3-MU cell line construction by CRISPR/Cas9-mediated homologous recombination. DNA sequencing of the mmtThrRS gene in NIH3T3-MU cells is shown. Codons His138 and His142 (CAC and CAT) have been changed to GCC and GCT Ala codons, respectively. (B) The amount of each amino acid moiety attached to mtRNAThr in the two cell lines was determined by LC–MS/MS. The results are reported as the averages of two independent trials with the SD indicated. (C) LC–MS analysis showing Thr-to-Ser misincorporation during mitochondrial translation in the NIH3T3-MU cell line.
The Abundance of mmtThrRS-H138A/H142A in NIH3T3-MU Cells was Significantly Reduced due to Protein Degradation.
We then compared the steady-state levels of mmtThrRS between NIH3T3-WT and NIH3T3-MU cells by western blot. The data showed that the level of mmtThrRS was significantly reduced in NIH3T3-MU cells compared to NIH3T3-WT cells. No significant variation in the expression of other mitochondrial aaRSs, including mtIleRS and mtAlaRS, between the two cell lines was observed (SI Appendix, Fig. S4A). The decrease in mmtThrRS protein expression may have been a result of transcriptional and/or translational and/or posttranslational effects because of His138 and His142 mutations in NIH3T3-MU cells. To distinguish among these possibilities, we first measured the mmtThrRS mRNA abundance by real-time quantitative PCR (RT–qPCR) and found that the mutant mmtThrRS mRNA level was increased in NIH3T3-MU cells (SI Appendix, Fig. S4B). To compare the mRNA translation efficiency of mmtThrRS-WT and mmtThrRS-H138A/H142A, NIH3T3 cells were transfected with a plasmid encoding either a C-terminal FLAG-tagged mmtThrRS (mmtThrRS-FLAG) or mmtThrRS-H138A/H142A (mmtThrRS-H138A/H142A-FLAG). Western blot analysis revealed comparable levels of mmtThrRS and mmtThrRS-H138A/H142A proteins (SI Appendix, Fig. S4C), indicating that in the presence of endogenous wild-type mmtThrRS, the translation efficiencies were comparable between the mmtThrRS and mmtThrRS-H138A/H142A mRNAs. Furthermore, similar yields of mmtThrRS-N and mmtThrRS-N-H138A/142A were observed in E. coli expression systems (SI Appendix, Fig. S4D). These data indicated that mRNA translation efficiency was not negatively affected by the two mutations in mammalian and bacterial cells. To determine whether the decrease in mmtThrRS-H138A/H142A in NIH3T3-MU cells was due to posttranslational protein degradation, the cells were treated with MG132, a proteasome inhibitor (36). The results showed that the abundance of mmtThrRS-H138A/H142A increased significantly after MG132 treatment. However, the MG132-induced accumulation of mmtThrRS in NIH3T3-WT cells was less pronounced (SI Appendix, Fig. S4E), suggesting that mmtThrRS-H138A/H142A was degraded via the action of the ubiquitin-proteasome system. To further investigate the accelerated degradation of mmtThrRS-H138A/H142A in NIH3T3-MU cells compared to mmtThrRS degradation in NIH3T3-WT cells, both cell lines were treated with cycloheximide to stop cytoplasmic mRNA translation. The levels of mmtThrRS and mmtThrRS-H138A/H142A remaining in the cells were measured at different times. Western blot analysis revealed that the protein level of mmtThrRS-H138A/H142A decreased more rapidly over time in NIH3T3-MU cells than in the NIH3T3-WT cells (SI Appendix, Fig. S4F). Together, these data collectively showed that the reduced level of mmtThrRS-H138A/H142A in NIH3T3-MU cells was mainly due to ubiquitin-mediated proteasome degradation of the protein.
We hypothesized that the degradation of mmtThrRS-H138A/ H142A in NIH3T3-MU cells might result from its toxicity due to the production of mischarged Ser-mtRNAThr and to the Thr-to-Ser misincorporation in vivo. Alternatively, His138 and His142 mutations may lead to local conformation changes and subsequent instability of the protein. To distinguish between these possibilities, we compared the thermal stabilities of purified mmtThrRS- N and mmtThrRS-N-H138A/142A in vitro, but the results showed little difference (SI Appendix, Fig. S4G). Furthermore, circular dichroism (CD) spectroscopy analysis showed that there was little difference in the secondary structure of the two proteins (SI Appendix, Fig. S4H). In addition, we measured the stability of purified mmtThrRS-N and mmtThrRS-N-H138A/142A using a proteinase K hydrolysis experiment, and the results showed comparable proteinase K sensitivity (SI Appendix, Fig. S4I). These data indicated that at least in vitro, the two mutations did not cause local structural alterations.
Overall, these data suggested that His138 and His142 mutations had no effects on the local architecture or stability of the protein. However, the abundance of mmtThrRS-H138A/H142A in NIH3T3-MU cells was significantly lower due to the degradation of the protein. We proposed that cell toxicity of mmtThrRS-H138A/H142A could be one of the main reasons for its lower abundance in NIH3T3-MU cells, perhaps to avoid the accumulation of large amounts of Ser-mtRNAThr. However, the detailed degradation mechanism of mmtThrRS-H138A/H142A in the mutant cells is unclear at present. Notably, mmtThrRS-H138A/H142A was not completely degraded, likely because it was essential for mitochondrial translation and cell survival. Therefore, the observed abundance of mmtThrRS-H138A/ H142A probably results from a balance between survival mechanisms and the potentially induced toxicity in cells.
Mitochondrial Translation, Biogenesis, and Morphology were Impaired in NIH3T3-MU Cells.
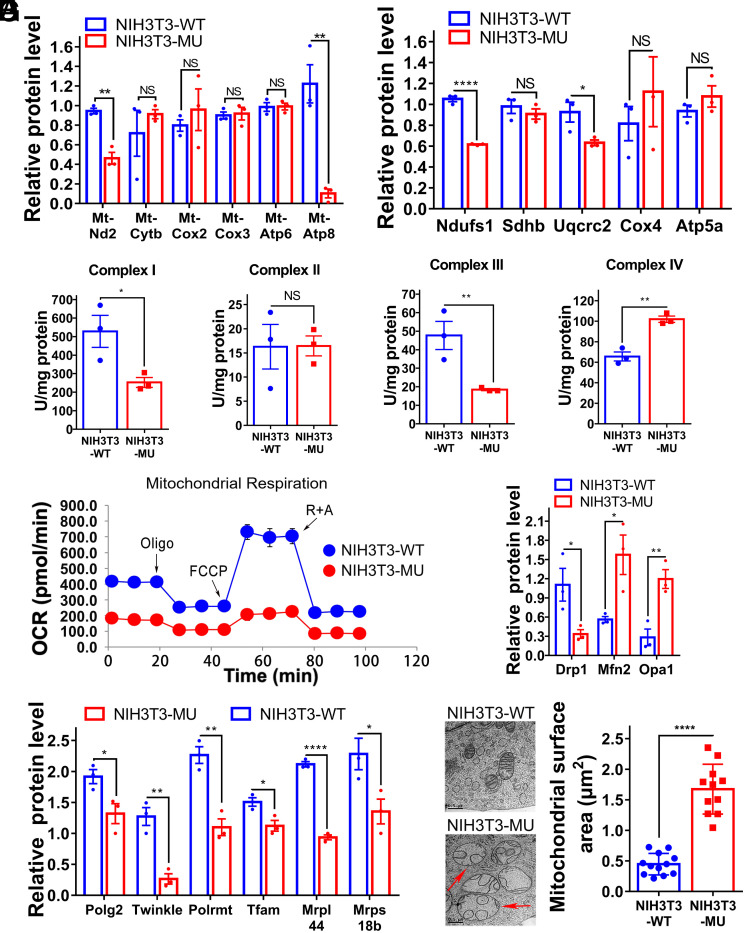
Having established that the editing-defective mmtThrRS-H138A/H142A protein generated Ser-mtRNAThr, which functions in conjunction with the mitochondrial ribosome in vivo, we investigated whether mitochondrial translation is affected by mischarged tRNAThr. To this end, we analyzed the abundance of several mtDNA-encoded proteins in various OXPHOS complexes, including Mt-Nd2 (Complex I subunit), Mt-Cytb (Complex III subunit), Mt-Cox2 and Mt-Cox3 (Complex IV subunit), and Mt-Atp6 and Mt-Atp8 (Complex V subunit). Western blot analyses revealed that mischarged tRNAThr exerted different effects on the levels of these proteins. The amount of some of these proteins, such as Mt-Cytb, Mt-Cox2, Mt-Cox3, and Mt-Atp6, was not altered, while the abundance of other proteins, including Mt-Nd2 and Mt-Atp8, was significantly lower in NIH3T3-MU cells. In particular, Mt-Atp8 was barely detectable (Fig. 2A and SI Appendix, Fig. S5A). In addition, nuclear DNA (nDNA)-encoded proteins in various OXPHOS complexes were also analyzed, and the results showed that the levels of Ndufs1 (Complex I subunit) and Uqcrc2 (Complex III subunit) were significantly lower, while the protein levels of Sdhb (Complex II subunit), Cox4 (Complex IV subunit), and Atp5a (Complex V subunit) were not changed (Fig. 2B and SI Appendix, Fig. S5B). Thus, the disruption of the mmtThrRS editing activity not only affected the specific subunits encoded by mtDNA but also those encoded by nDNA. We then compared the respiratory chain complex activities in NIH3T3-WT and NIH3T3-MU cells. The results showed that the activity levels of Complexes I and III were significantly lower, while the activity level of Complex IV was significantly higher in NIH3T3-MU cells compared to NIH3T3-W cells (Fig. 2C), possibly due to a compensatory mechanism. In addition, the activity level of Complex II was not altered, which was consistent with the fact that subunits of Complex II are all nDNA-encoded. We then compared the mitochondrial OXPHOS capacity of NIH3T3-WT and NIH3T3-MU cells. The oxygen consumption rate (OCR) measurements indicated that both basal and maximal respiration capacities were significantly lower in the NIH3T3-MU cells (Fig. 2D and SI Appendix, Fig. S5 C and D). As expected, the reduction in OCR rates resulted in decreased ATP production in the NIH3T3-MU cells (SI Appendix, Fig. S5E). These data clearly showed that mitochondrial function was disrupted in the NIH3T3-MU cells, likely because of mitochondrial mistranslation. To determine whether mitochondrial translation infidelity affects mitochondrial biogenesis, we first counted the number of mtDNA copies, which we found to be markedly reduced (SI Appendix, Fig. S5F). Furthermore, the steady-state levels of nDNA-encoded mitochondrial proteins, including Polg2 and Twinkle (which function in mtDNA replication), Polrmt and Tfam (which function in mtDNA transcription) (37), and Mrpl44 and Mrps18b (which function in mitochondrial ribosome biogenesis) (38), were significantly lower in the NIH3T3-MU cells (Fig. 2E and SI Appendix, Fig. S5G). These data suggested that mitochondrial biogenesis was influenced by mitochondrial translation infidelity. In addition, the amount of Drp1 (39), which is essential for mitochondrial fission, was lower in the NIH3T3-MU cells. In contrast, the amounts of Mfn2 and Opa1 (39), which are essential for mitochondrial fusion of the outer and inner mitochondrial membranes, were higher in the NIH3T3-MU cells (Fig. 2F and SI Appendix, Fig. S5H), suggesting altered mitochondrial morphology in NIH3T3-MU cells. Transmission electron microscopy (TEM) results indeed showed that mitochondria were swollen in NIH3T3-MU cells, as evidenced by an enlarged mitochondrial surface. In addition, less delineated mitochondrial ridges were observed in the NIH3T3-MU cells (Fig. 2G). Altogether, these results indicated that disruption to mmtThrRS-mediated mitochondrial translation fidelity impaired mitochondrial translation, biogenesis, structure, and function.
Fig. 2.
Mitochondrial translation, biogenesis, and morphology were impaired in the NIH3T3-MU cell line. (A and B) Relative level of several proteins in various OXPHOS complexes. (C) Enzymatic activity levels of mitochondrial oxidative respiratory chain complexes. (D) OCR of NIH3T3-WT (blue) and NIH3T3-MU (red) cells. Oligo, oligomycin; FCCP, carbonyl cyanide-p-trifluoromethoxyphenylhydrazone; R+A, rotenone + antimycin A. (E) Relative level of proteins in mitochondrial replication, transcription, and translation. (F) Relative level of mitochondrial morphology-associated proteins. (G) Representative TEM images (Left) and quantification (Right) of mitochondria in NIH3T3-WT and NIH3T3-MU cells. Swollen mitochondria are marked by red arrowheads. The data are presented as the mean ± SEM (n = 3 to 12, ∗P < 0.05; ∗∗P < 0.01; ∗∗∗∗P < 0.0001; NS, not significant). Student’s t test was performed.
The NIH3T3-MU Cell Proliferation Rate was Lower due to Cell Cycle Arrest.
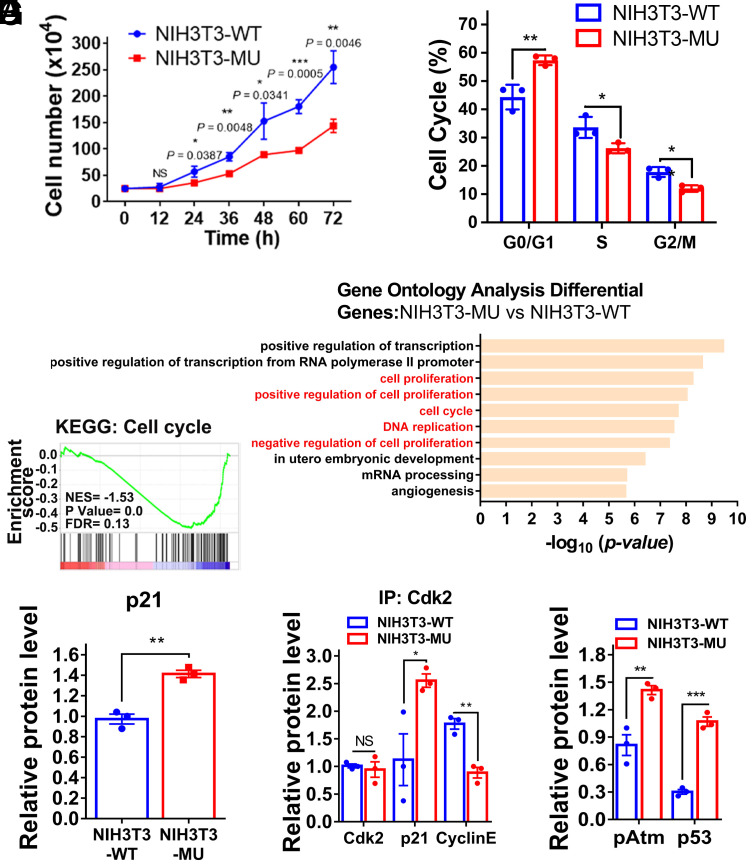
The phenotypes of NIH3T3-MU cells were then characterized. The cell growth rate was clearly lower after the loss of the mmtThrRS editing function (Fig. 3A). To evaluate whether cell proliferation was blocked, we examined cell proliferation capability by 5-ethynyl-2′-deoxyuridine labeling (40). NIH3T3-MU cells showed significantly lower proliferation rates than NIH3T3-WT cells (SI Appendix, Fig. S6A). Flow cytometry analysis revealed that the mutant cell line was more frequently arrested in G0/G1 phase than the wild-type cell line (Fig. 3B). High-throughput sequencing of total RNA (RNA-seq) from NIH3T3-WT and NIH3T3-MU cells identified 3,582 protein-coding genes that were differentially expressed after mmtThrRS was mutated (1,603 up-regulated genes and 1,979 down-regulated genes) (SI Appendix, Fig. S6B). A Gene Ontology (GO) analysis of the down-regulated genes in the biological process subset (BP) revealed a high gene enrichment in terms of cell proliferation and cell cycle (Fig. 3C). An enrichment score (ES) analysis of all genes also revealed that cell cycle signaling pathway was highly enriched (Fig. 3D). In summary, our results showed that the NIH3T3-MU cell line exhibits suppressed cell cycle progression and reduced proliferation.
Fig. 3.
NIH3T3-MU cell lines showed reduced cell proliferation due to cell cycle arrest. (A) Cell growth determination of NIH3T3-WT (blue) and NIH3T3-MU (red) cells over time. (B) Cell cycle analysis performed with flow cytometry. (C) GO enrichment analysis of the subset of dysregulated genes in the RNA-seq data. Cell cycle relative BP subsets were marked with red. (D) The cell cycle pathway was enriched by ES enrichment. (E) Relative level of p21. (F) A higher level of p21-Cdk2 interaction in NIH3T3-MU cells was observed by coimmunoprecipitation with an anti-Cdk2 antibody. (G) Relative level of pAtm and p53. The data are presented as the mean ± SEM (n = 3 to 4, ∗P < 0.05; ∗∗P < 0.01; ∗∗∗P < 0.001; NS, not significant). Student’s t test was performed.
Cyclins and cyclin-dependent kinases are key regulators of the eukaryotic cell cycle (41). Cdk2 forms a complex with Cyclin E to control the G1/S transition, which is inhibited by p21 via its competitive binding to Cdk2 (42–44) (SI Appendix, Fig. S6C). To understand the potential mechanism underlying the observed G0/G1 arrest of the mutant cells, we first measured the transcript levels of the relevant genes encoding Cdk inhibitor family proteins by RT–qPCR. The mRNA level of p21 was higher in the NIH3T3-MU cells than in the NIH3T3-WT cells (SI Appendix, Fig. S6D). Western blot analyses showed that the p21 level was consistently higher in the NIH3T3-MU cells (Fig. 3E and SI Appendix, Fig. S6E). We hypothesized that the Cdk2-Cyclin E interaction was inhibited in NIH3T3-MU cells. Therefore, we enriched endogenous Cdk2 using an anti-Cdk2 antibody and found that the amount of immunoprecipitated Cyclin E was indeed significantly reduced. In addition, the level of p21 pulled down was markedly increased in NIH3T3-MU cells (Fig. 3F and SI Appendix, Fig. S6F).
In a similar experiment, we precipitated endogenous p21 using an anti-p21 antibody and found that Cdk2 was more abundant in the p21-enriched sample of NIH3T3-MU cells (SI Appendix, Fig. S6G). DNA damage–induced activation of the cell-cycle checkpoint pathway is involved in mammalian cell-cycle arrest (45) (SI Appendix, Fig. S6C). Phosphorylation of ataxia telangiectasia mutated (Atm) at Ser1987 (pAtm), a key player in the DNA-damage response (DDR) pathway, was higher in the NIH3T3-MU cells (Fig. 3G and SI Appendix, Fig. S6H). In addition, the protein level of p53 (a transcription factor related to p21 expression) (46, 47), whose transcription is activated by pAtm, was higher in the NIH3T3-MU cells (Fig. 3G and SI Appendix, Fig. S6H). These data collectively suggested that increased p21 prevented Cdk2 from interacting with Cyclin E and thus likely contributed to cell cycle arrest during the G1/S phase transition in NIH3T3-MU cells.
Oxidative Stress Led to Cell Cycle Arrest in NIH3T3-MU Cells.
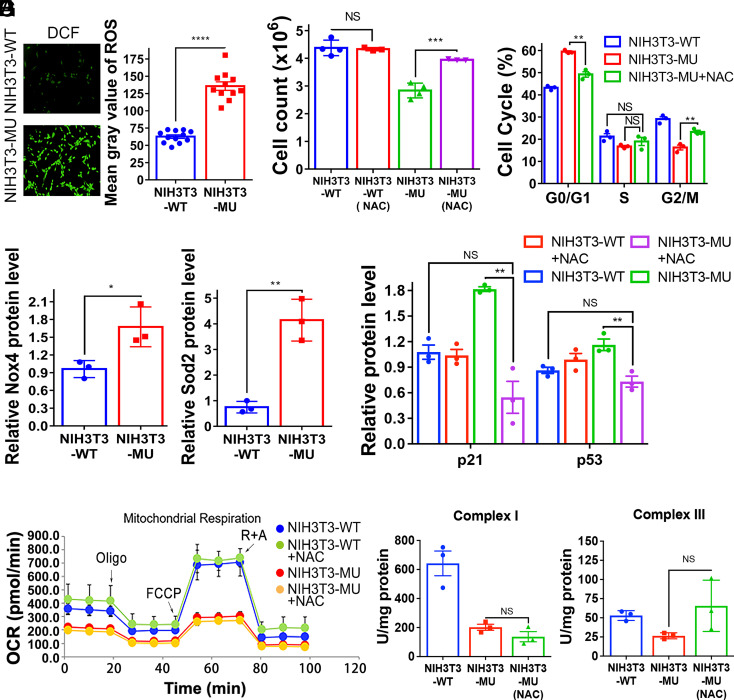
Atm is frequently activated by oxidative stress (48). Indeed, abnormal mitochondrial function frequently leads to excessive ROS (reactive oxygen species) production and cellular oxidative stress (49). Specifically, Complex I is a main source of ROS production (50). To determine whether disruption of mmtThrRS editing leads to oxidative stress, 2′,7′-dichlorofluorescein (DCF), a cell-permeable fluorescent probe, was used to measure global oxidative stress (51). The ROS content in NIH3T3-MU cells was significantly higher than that in NIH3T3-WT cells (Fig. 4A). Oxidative stress is thought to lead to the posttranslational modification of proteins, including carbonylation (52). Protein carbonylation assays showed a higher level of carbonylation in the NIH3T3-MU cells, confirming a higher level of ROS production in the mutant cells (SI Appendix, Fig. S7A). Besides Complex I, Nox4 is a validated source of ROS production (53). Western blot analysis showed that the level of Nox4 was significantly higher in NIH3T3-MU cells (Fig. 4B and SI Appendix, Fig. S7B). In addition, the level of Sod2, a protein involved in ROS scavenging (54), was significantly higher in the mutant cells (Fig. 4B and SI Appendix, Fig. S7B). Taken together, these data revealed that defective editing of mmtThrRS led to high levels of ROS production in NIH3T3-MU cells.
Fig. 4.
Oxidative stress contributed to NIH3T3-MU cell cycle arrest. (A) Fluorescence intensity (Left) and level (Right) of ROS measured using a DCF fluorescent probe in vivo. (B) Relative level of the ROS-related proteins Nox4 (Left) and Sod2 (Right). (C) Number of NIH3T3-WT and NIH3T3-MU cells before and after NAC treatment. (D) Analysis of the NIH3T3-MU cell cycle in the absence or presence of NAC treatment, as determined via flow cytometry. Ratios of cells in various phases to the total number of cells were calculated and compared with those of NIH3T3-WT cells. (E) Relative protein level of p21 and p53 in NIH3T3-WT and NIH3T3-MU cells before and after NAC treatment. (F) OCR in cells before [NIH3T3-WT (blue), NIH3T3-MU (red)) and after (NIH3T3-WT + NAC (green), NIH3T3-MU (yellow)] NAC treatment. (G) Activities of Complexes I and III in NIH3T3-MU cells after NAC treatment. The data are presented as the mean ± SEM (n = 3-12, ∗P < 0.05; ∗∗P < 0.01; ∗∗∗P < 0.001; ∗∗∗∗P < 0.0001; NS, not significant). Student’s t test was performed.
We hypothesized that the higher level of ROS might contribute to the activation of the DDR pathway, as shown by a higher level of pAtm, and subsequently lead to cell cycle arrest. N-acetylcysteine (NAC) is a precursor of cysteine and glutathione synthesis and normally functions as an antioxidant that contributes to cellular redox balance (55, 56). We treated the NIH3T3-MU and NIH3T3-WT cell lines with NAC and measured the protein carbonylation levels, which indeed showed that the NIH3T3-MU cell line exhibited significantly reduced overall protein carbonylation after NAC treatment. A reduction in protein carbonylation in the NIH3T3-WT cells was also evident, but it was not as pronounced as that in the NIH3T3-MU cells (SI Appendix, Fig. S7C). Cell number determination showed that NAC treatment exerted no effect on the number of NIH3T3-WT cells. In contrast, the number of the NIH3T3-MU cells increased significantly after NAC treatment, reaching a number comparable to that of the NIH3T3-WT cells (Fig. 4C). Cell cycle analysis showed that the ratio of NIH3T3-MU cells in the G0/G1 phase after NAC treatment was significantly reduced, whereas it was increased in G2/M (Fig. 4D). No significant difference was observed in S phase (Fig. 4D). Consistently, the abundance of p53 and p21 proteins was markedly reduced in NIH3T3-MU cells after NAC treatment, but not in NIH3T3-WT cells (Fig. 4E and SI Appendix, Fig. S7D). These data indicated that NAC effectively reduced ROS levels in the NIH3T3-MU cells. Additional immunofluorescence assay showed that the level of pAtm in the NIH3T3-MU cells after NAC treatment was reduced and comparable to that in the NIH3T3-WT cells (SI Appendix, Fig. S7E).
We then evaluated whether mitochondrial function was influenced by NAC treatment. The OCR and ATP production were similar before and after NAC treatment in both NIH3T3-WT and NIH3T3-MU cells (Fig. 4F and SI Appendix, Fig. S7F). Furthermore, we measured Complexes I and III activity levels after NAC treatment, which were not significantly increased (Fig. 4G). Indeed, NAC treatment reduced only the levels of ROS and did not rescue the mitochondrial mRNA translation fidelity that had been abolished.
All these data indicated that NIH3T3-MU cells underwent oxidative stress, which contributed to cell cycle arrest. NAC treatment alleviated oxidative stress and thus restored cell cycle progression.
Impaired Mitochondrial Translation, Elevated ROS Levels, and Cell Cycle Arrest were Conserved Effects after mmtThrRS Editing was Rendered Deficient in Other Mammalian Cells.
We further explored whether the impaired mitochondrial translation, elevated ROS levels, and cell cycle arrest were common outcomes of mmtThrRS-editing deficiency in other cell lines. The HL-1 cell line, derived from the AT1 mouse atrial cardiomyocyte tumor cell line (57), is a mouse cardiac muscle cell line (hereafter, the HL-1-WT cell line). H138A/H142A double mutations were established in the HL-1 cell line (hereafter, the HL-1-MU cell line) using CRISPR/Cas9-mediated genome editing (SI Appendix, Fig. S8A). As in the NIH3T3-MU cells, the mmtThrRS-H138A/H142A level was substantially lower in the HL-1-MU cells (SI Appendix, Fig. S8B). The abundance of Mt-Atp6 was significantly lower in the HL-1-MU cell line. The protein level of Mt-Nd2 was also decreased, but the reduction was not significant (SI Appendix, Fig. S8C). These results indicated impaired mitochondrial translation in the HL-1-MU cells. Moreover, with respect to the OXPHOS rate, the activity levels of Complexes I and III were significantly lower while the activity levels of Complexes II and IV were not substantially changed in the HL-1-MU cells (SI Appendix, Fig. S8D). The OCR results showed that ATP production capacity was significantly lower in the HL-1-MU cells (SI Appendix, Fig. S8E). A DCF fluorescent probe assay showed that the ROS content was significantly higher (SI Appendix, Fig. S8F) and was accompanied by an elevated level of global protein carbonylation (SI Appendix, Fig. S8G) in the HL-1-MU cells. Notably, the abundance of p21 and p53 proteins was significantly higher in the HL-1-MU cell line (SI Appendix, Fig. S8B). We then compared the number of HL-1-MU and HL-1-WT cells in each cell cycle phase via flow cytometry, and the results revealed that the HL-1-MU cell line was more frequently arrested in G0/G1 phase than the HL-1-WT cell line (SI Appendix, Fig. S8H). Altogether, these results indicated that disruption of mmtThrRS editing activity induced conserved cellular responses, including impaired mitochondrial translation, elevated ROS levels, and disruption of the cell cycle in different mammalian cells.
Cardiomyocyte-Specific mmtThrRS H138A/H142A Mutations Led to Cardiac Dysfunction.
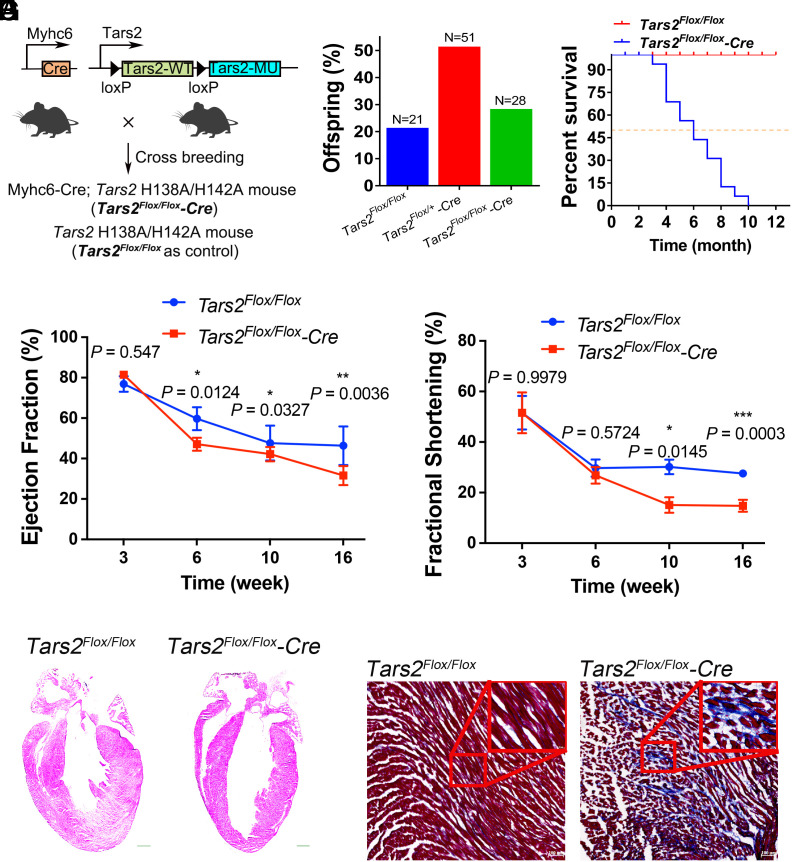
To investigate the role of the mmtThrRS editing function in mice, we constructed a conditional mmtThrRS H138A/H142A-mutant C57BL/6 mouse model using the Cre-loxP recombinase system (Tars2Flox/Flox) (SI Appendix, Fig. S9A). We introduced the mmtThrRS H138A/H142A mutation throughout the whole body of the mice using Cre recombinase controlled by the adenovirus EIIa promoter (58). Only mice with the Tars2Flox/+-EIIa Cre−/− genotype (Tars2-EIIa Cre−/−, only wild-type mmtThrRS expressed in the whole body due to absence of Cre recombinase) were among the offspring, and they showed no obvious abnormalities. However, mice with Tars2Flox/+-EIIa Cre+/− genotype (Tars2-EIIaCre+/−, both mmtThrRS and mmtThrRS-H138A/H142A expressed in the whole body) were not obtained (SI Appendix, Fig. S9B). This outcome suggests that the editing function of mmtThrRS was essential for embryonic development, as its loss resulted in lethality even in heterozygotes, likely due to the manifestation of a gain-of-function mutation that generated a large amount of Ser-mtRNAThr. Western blot analysis revealed that mmtThrRS protein expression was the highest in the heart compared to multiple other mouse tissues (SI Appendix, Fig. S9C). Notably, the heart is sensitive to mitochondrial dysfunction due to its high energy consumption (59). Given the high expression level of mmtThrRS in the heart and to gain an understanding of any potential role for mmtThrRS editing in cardiac function, we crossed Myhc6-Cre mice (in which the cardiac-specific mouse alpha-myosin heavy chain promoter drove the expression of Cre) (60) with Tars2Flox/Flox mice to obtain cardiac-specific mmtThrRS-H138A/H142A mutant mouse offspring (Tars2Flox/Flox-Cre, only mmtThrRS-H138A/H142A expressed in cardiomyocytes, designated the mutant mice) (Fig. 5A). We were able to obtain knock-in mice that followed Mendel's laws of inheritance (mice without Cre (Tars2Flox/Flox, only wild-type mmtThrRS expressed in cardiomyocytes) were the control group, designated the wild-type mice) (Fig. 5B). The mutant mice showed a significantly shorter lifespan than the wild-type mice, with a median survival rate of 26 wk, suggesting that the editing function of mmtThrRS was essential for the maintenance of normal cardiac function (Fig. 5C). We continuously monitored the cardiac function of the mice from week 3 to week 16. A longitudinal echocardiographic analysis of the left ventricle in the mouse heart revealed cardiac dysfunction in the mutant mice (SI Appendix, Fig. S9D). The mutant mice showed a decrease in left ventricular ejection fraction (LVEF) and left ventricular fractional shortening (LVFS) from approximately the sixth week (Fig. 5 D and E). In addition, we found that the left ventricular internal diameter end diastole (d) and the left ventricular cavity in the mutant mice started to dilate in the sixth week, meaning that the mutant mice exhibited systolic heart dysfunction (Fig. 5F and SI Appendix, Fig. S9E). Masson’s trichrome staining revealed cardiac fibrosis in the hypertrophic hearts of the mutant mice (Fig. 5G). Altogether, these results showed that the cardiomyocyte-specific loss of the mmtThrRS editing function led to cardiomyopathy in mice.
Fig. 5.
Cardiomyocyte-restricted mmtThrRS-H138A/ H142A mutations resulted in cardiac dysfunction. (A) mmtThrRS-H138A/H142A was expressed in cardiomyocytes by flipping the floxed mmtThrRS-WT cassette between the mmtThrRS gene promoter and mmtThrRS-MU and crossing the mice carrying this mutation with mice harboring Myhc6 promoter-driven Cre recombinase. Myhc6-Cre; Mice expressing mmtThrRS-H138A/H142A in cardiomyocytes were designated Tars2Flox/Flox-Cre (mutant) mice. Litter-mate mice expressing wild-type mmtThrRS in cardiomyocytes were designated Tars2Flox/Flox (control, wild-type) mice. (B) The genotyping results after crossing Myhc6 promoter-Cre mice with Tars2Flox/Flox mice; the outcome followed Mendelian laws. (C) Survival curves for the wild-type (n = 13) and the mutant mice (n = 16). (D and E) LVEF (Tars2Flox/Flox, n = 7; Tars2Flox/Flox-Cre, n = 6) and LVFS (Tars2Flox/Flox, n = 7; Tars2Flox/Flox-Cre, n = 4) measurements of the wild-type and the mutant mice over time. (F) Representative hematoxylin and eosin (H&E) staining of the wild-type and the mutant mouse hearts. (G) Representative image showing Masson’s trichrome staining of the wild-type and the mutant mouse hearts. The data are presented as the mean ± SEM (∗P < 0.05; ∗∗P < 0.01; ∗∗∗P < 0.001; NS, not significant). Student’s t test was performed.
The Mutant Mice Exhibited Increased ROS Levels and Inhibited Cardiomyocyte Proliferation.
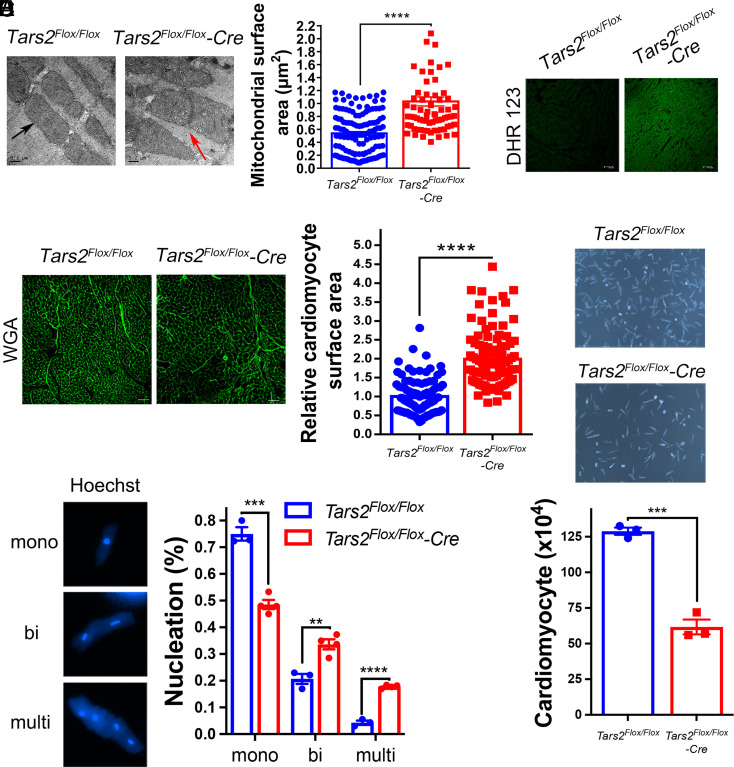
To explore the molecular basis of cardiac dysfunction in the mutant mice, we examined the morphology of mitochondria in cardiomyocytes using TEM (Fig. 6A). Mitochondria with abnormal cristae structures and a large surface area, indicating altered mitochondrial structure, had accumulated in the hearts of the mutant mice (Fig. 6A). Furthermore, we observed a decrease in the number of mitochondria, which was consistent with the TEM results obtained by observation of the NIH3T3-MU cell line (SI Appendix, Fig. S9F). Considering the increased ROS levels in NIH3T3-MU cells, we examined the ROS levels in cardiomyocytes isolated from the wild-type and mutant mice separately and found that the ROS levels were significantly higher in the mutant mice, as indicated by a protein carbonylation assay (SI Appendix, Fig. S9G). An immunofluorescence assay of the ROS probe dihydrorhodamine 123 (DHR 123), a probe commonly used for the measurement of tissue ROS, also showed a higher level of ROS content in the hearts of the mutant mice (Fig. 6B). We then determined the mouse heart weight to body weight ratio (HW/BW) and found that the mutant mice showed an HW/BW comparable to that of control mice (SI Appendix, Fig. S9H). By performing a wheat germ agglutinin (WGA) staining assay, we found that the cardiomyocytes in the mutant mice were significantly larger (Fig. 6C). We then isolated cardiomyocytes and found that the number of cardiomyocytes in the mutant mice was significantly lower than that in the wild-type mice (Fig. 6D), implying that excessive ROS levels reduced cardiomyocyte proliferation, similar to their effect on NIH3T3-MU cell proliferation. To validate the inhibition of cardiomyocyte proliferation in the mutant mice, we analyzed the karyotype of the cardiomyocytes isolated from both the mutant and wild-type mice and found that the number of mononuclear cardiomyocytes was significantly lower and that the number of bi- or multinucleated cardiomyocytes was significantly higher in the mutant mice (Fig. 6E). These results revealed increased ROS levels and inhibited cardiomyocyte proliferation in the mutant mice.
Fig. 6.
Increased ROS levels and inhibited proliferation of cardiomyocytes from the mutant mice. (A) Mitochondria (indicated by a black arrow) were examined by TEM (×30,000). Representative images (Left) showing disorganized mitochondrial cluttering (indicated by a red arrow) in the mutant mice hearts (scale bar: 0.2 µm.) The quantification (Right) of the surface area of mitochondria from the wild-type (n = 128) and the mutant (n = 73) mice. (B) Representative images showing DHR 123 staining of heart tissue sections from the wild-type and the mutant mice. (C) Representative images showing WGA staining (Left) and quantification (Right) of parameters measured in the heart tissue sections prepared from the wild-type (n = 3) and the mutant (n = 3) mice. (D) Bright-field images (Upper) and number (Lower) of isolated mouse cardiomyocytes (the wild-type and the mutant mice, n = 3). (E) The types (Left) and quantification (Right) of the nuclei in the isolated mouse cardiomyocytes from the wild-type (n = 3) and the mutant (n = 4) mice. The data are presented as the mean ± SEM (∗∗P < 0.01; ∗∗∗P < 0.001; ∗∗∗∗P < 0.0001). Student’s t test was performed.
Discussion
AaRSs are generally considered the initial checkpoint of “translation quality” because they ensure accurate tRNA charging via aminoacylation and proofreading functions. In all three domains of life, with few exceptions, Class I aaRSs, namely, LeuRS, IleRS, and ValRS, and Class II aaRSs, namely, AlaRS, ThrRS, ProRS (bacterial type), and PheRS, all carry editing domains (4, 5). This conservation highlights the high selective pressure to maintain the proofreading of mischarged tRNAs, which is necessary to maintain a high level of translational accuracy during mRNA decoding. Indeed, misincorporation of amino acids is likely to generate a statistical proteome with effects of loss or gain of function, including toxicity and misfolding, leading to various outcomes for both bacteria and mammalian cells/tissues. For example, ctValRS editing deficiency leads to mistranslation in the mammalian cytoplasm (8). As a consequence, metabolic dysfunctions and diseases in mice and humans have been observed (9, 10). Mammalian mitochondrial aaRSs are unique in that only some of these enzymes carry intact editing domains: mtIleRS, mtValRS, mtAlaRS, and mtThrRS (30). The yeast mtLeuRS carries an intact CP1 editing domain and can edit mischarged tRNALeu (27). However, in mammalian cells, peptide deletions and loss of catalytic residues in the CP1 domain in mtLeuRS have resulted in the loss of proofreading ability (25, 26). Several studies have suggested that although the amino acids Ile, Val, and norvaline (Nva) share very similar structures and physicochemical properties, all LeuRSs misactivate Nva at significantly higher efficiency than Ile or Val (16, 61–63). Nva seems to be the bona fide substrate for the editing activity of LeuRS in vivo (64, 65). The failure to detect free Nva in the amino acid pool of mammalian mitochondria may explain the loss of editing activity of mtLeuRS (66). On the other hand, the CP1 domain also contributes to the structure of the enzyme and thus to the aminoacylation of tRNALeu, which may explain the reason that the editing domain has been retained in these enzymes, even though they are inactive (26). Notably, there is a clear dichotomy between bacterial ProRSs (with an editing domain) and eukaryotic ctProRSs (without an editing domain) enzymes. The prokaryotic editing domain, or “INS”, is critical for removing Ala-tRNAPro (67). This editing activity is complemented by freestanding trans-editing factors, including ProX (22, 23) and YbaK, which remove Ala-tRNAPro and Cys-tRNAPro, respectively (68). Mitochondrial ProRSs in lower and higher eukaryotes lack an INS homologous domain and are thus thought to be incapable of editing functions (30). Furthermore, no trans-editing factors have been identified, suggesting that no proofreading mechanism is available to correct misacylated tRNAPro in mitochondria. The possibility that mtProRSs efficiently reject noncognate Ala and Cys during the amino acid activation step, thus eliminating the need for editing function, cannot be discounted. Finally, yeast mtPheRS misactivates noncognate Tyr but cannot edit Tyr-tRNAPhe, suggesting that mitochondrial translation is inherently less accurate than cytoplasmic translation at Phe codons (28). Similar to yeast mtPheRS, mammalian mtPheRS does not carry an editing domain (30). The loss of the editing function of mammalian mtPheRS implies efficient Tyr discrimination or toleration of inaccuracy at Phe codons (29).
Overall, the reason that many mitochondrial tRNA synthetases have lost their editing function is unclear, although some hypotheses have been suggested. First, mitochondrial tRNA synthetases may exhibit greater fidelity via improved synthetic active sites that have rendered them to show higher specificity and be less prone to error (29). Second, the proteinogenic or nonproteinogenic amino acid content in mitochondria may be reduced, preventing competition with cognate amino acids (66). Third, mitochondrial translation tolerates a certain degree of amino acid misincorporation at Leu, Pro, and Phe codons. Fourth, the possibility that translation specificity is enhanced by other factors, such as a quality control program orchestrated by mitochondrial elongation factors, similar to counterparts in bacteria that can reject nonhomologous aa-tRNA pairs, cannot be ruled out (69, 70). Finally, the possibility that certain unidentified trans-editing factors exist in mitochondria cannot be absolutely excluded.
The translation of a complete cytoplasmic proteome (approximately 40.6 × 106 codons in a human genome) (https://www.kazusa.or.jp/codon/) is accompanied by thousands of misincorporation events, which may explain the need for proofreading by tRNA synthetases. The high complexity of an organism seems to justify the need for a proofreading function that reduces the error rate (3, 71, 72). The human mitochondrial genome harbors only 3,789 codons (excluding stop codons). Given the small number of codons for the 13 mitochondrial proteins, one may question the need of editing. With an overall translation error rate of 10−3 to 10−5 and considering that the mitochondrial ribosome is similarly faithful (3, 72), we may deduce that the translation of the 3,789 mitochondrial codons can take place almost without any incorporation errors. However, in a previous work, we showed that mtAlaRS misactivated noncognate Ser and Gly and relied on robust tRNA-independent pretransfer editing to correct misactivated Gly and posttransfer editing to correct mischarged Ser-mtRNAAla (32). This proofreading function seems essential since disruption of the mtAlaRS editing function led to early embryonic lethality in mice (11). In the present work, we established an NIH3T3 cell line and a mouse model with mtThrRS editing deficiency. This is a previously undescribed cell line or mouse model with the mitochondrial translation quality control abolished. Our results clearly showed that without mtThrRS editing, noncognate Ser was readily charged to mtRNAThr, and the resulting Ser-mtRNAThr was efficiently utilized by the mitochondrial translation apparatus, as shown by the misincorporated Ser residues identified at Thr positions in seven mtDNA-encoded proteins (Fig. 7). Considering the peptides identified, we roughly estimated a replacement rate of approximately 20% at Thr positions in the mutant cells, which is a significantly high rate, although this ratio should be interpreted with caution due to the small number of peptides sequenced. In contrast to mammalian mtThrRS, yeast mtThrRS lacks an editing domain, retaining only the aminoacylation and tRNA-binding domains (73). The specificity of this minimal ThrRS is ensured by two editing functions: tRNA-independent pretransfer editing and tRNAThr-species-specific pretransfer editing (74, 75). These observations point to species-specific strategies to prevent the replacement of Thr by Ser in mitochondria. Overall, our data showed that mammalian mtThrRS misactivated noncognate Ser and hydrolyzed Ser-mtRNAThr, which were essential for controlling the translational quality of mitochondrial protein synthesis.
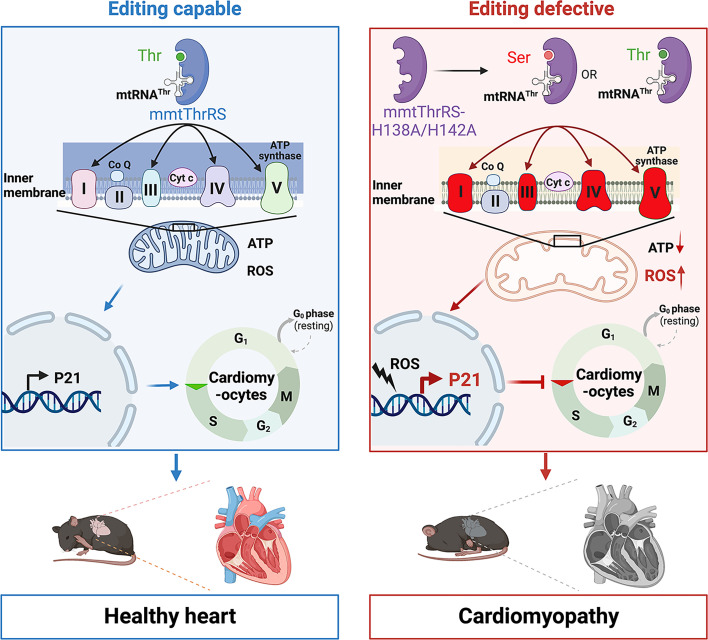
Fig. 7.
Proposed model showing how mmtThrRS editing deficiency leads to cellular and heart dysfunctions. Mutations in His138 and His142 abolished the posttransfer editing function of mmtThrRS, leading to gain-of-function effects that increased the Ser-mtRNAThr mischarging, in addition to Thr-mtRNAThr charging, by mmtThrRS-H138A/H142A. The accumulation of Ser-mtRNAThr led to abundant Thr-to-Ser misincorporation during mitochondrial mRNA translation. This disruption to translation impaired the assembly and function of the mitochondrial respiratory chain complexes and caused oxidative stress, which subsequently led to cell cycle arrest in the G0/G1 phase. This comprehensive mitochondrial and cellular dysfunction likely affected cardiomyocytes after mutations of His138 and His142, which were associated with dilated cardiomyopathy in heart-specific mmtThrRS editing–defective mice. This figure was created with BioRender.
The components of the OXPHOS chain are multiprotein complexes that must be assembled from subunits encoded by both nDNA and mtDNA (except for Complex II) (37). In mtThrRS-mutated cells, altered content and quality of mtDNA-encoded proteins are thought to impair the faithful assembly and functions of OXPHOS complexes. Indeed, we observed a significant reduction in the activity levels of Complexes I and III, which led to the disruption of basic cellular functions. Moreover, the production of ATP by Complex V was significantly reduced, leading to a deficient cellular energy supply. Finally, the abnormality in electron transport generated oxidative stress in the cells (Fig. 7).
In this work, we showed that ROS overload triggered oxidative damage to nuclear genes and activated the DDR (Fig. 7). The damaged nuclear genome induced the phosphorylation of Atm, a key kinase upstream of the DDR signaling pathway. Atm activated the p53 protein, a transcription factor that mediates p21 expression. Ultimately, p21 bound to Cdk2 to suppress the Cdk2-Cyclin E interaction, and therefore, the G1/S transition was arrested. Our results suggested that increased levels of p53 in mutant cell lines were involved in cell cycle arrest, a finding similar to that of previous studies (42, 46, 47). The p53-dependent DDR was also activated in zebrafish models expressing editing-defective cytoplasmic ValRS, suggesting a common connection between cytoplasmic and mitochondrial translation fidelity with p53 (19). A recent study has reported that mtThrRS can interact with RagC to regulate mTORC1 signaling outside of mitochondria. However, it has been shown that the H133A/H137A double mutant in the hmtThrRS editing site does not influence the mtThrRS-RagC interaction (76). It is worth noting that treatment of NIH3T3-MU cells with NAC efficiently reversed cell cycle arrest, indicating that ROS generation is the primary driver of NIH3T3-MU cell proliferation suppression. However, mitochondrial dysfunction itself is directly associated with mTORC1 regulation (77, 78); the potential alteration in mTORC1 activity in NIH3T3-MU cells requires further investigation.
We observed abnormal mitochondrial morphology and elevated ROS in the cardiomyocytes of mutant mice, with these cardiomyocytes undergoing cell cycle arrest and generating excessive ROS, which in turn activated nuclear responses to suppress the expression of electron transport chain subunits and further impair electron transfer capability. This vicious cycle induces high level of oxidative stress in cardiomyocytes, resulting in cardiomyocyte cell cycle arrest (45). In addition, echocardiography results showed that the values of the LVEF and LVFS parameters were reduced in mutant mice, indicating that they had acquired the classic cardiomyopathy phenotype (79). In cardiac dysfunction, metabolic substrates of the heart are converted from fatty acids to glucose, which leads to a decrease in the mitochondrial ATP production capacity of the heart and exacerbates cardiac dysfunction (80–82). Considering these analyses, we suggest that both the reduction in the ATP supply and cell cycle arrest, likely caused by oxidative stress, contributed to the impaired cardiac function of mtThrRS-mutant mice (Fig. 7).
We and others have reported several pathogenic hmtThrRS mutations (including P282L, S258L, F323C, E425G, R452W, etc.) in clinical studies (31, 83, 84). Both S258L and P282L were in the editing domain of hmtThrRS. However, none of the patients showed heart dysfunction. It is likely that the lack of cardiac dysfunction was a result from weak impairment to Ser-mtRNAThr editing induced by these hmtThrRS mutants. We observed robust posttransfer editing of mischarged tRNAThr by the P282L mutant (31) and no accumulation of Ser-mtRNAThr by the S258L, F323C, E425G, and R452W mutants (83). Thus, the clinical effects of these hmtThrRS mutants were rather the result of a deficiency in Thr-mtRNAThr production by enzymes whose levels were also decreased due to their structural instability. These mutants were clearly different from the H138A/H142A double mutant constructed in this study, which resulted in the accumulation and incorporation of Ser-mtRNAThr into mitochondrial protein synthesis. Notably, the genetically induced mutations of H133 and H137 (mouse equivalents of H138 and H142) have never been isolated in clinical studies, which may corroborate the very strong deleterious effect observed in this study. Indeed, if these mutations were introduced, embryonic development would be stopped, and the patient would not give birth. Here, when the mouse H138A/H142A mutations were introduced by a widely expressed Cre recombinase, no offspring were obtained. Consistent with this hypothesis, we observed early embryonic lethality due to whole-body mutations in two critical editing sites in mtAlaRS (11). Similarly, “sticky” mice expressing ctAlaRS with a minor defect (shown an approximately twofold loss of function) in editing activity exhibited neurodegeneration (9). Homozygous embryos expressing ctAlaRS-C723A, with abolished editing activity, died at an earlier stage of development (10). We anticipate that hmtThrRS editing site mutations, with attenuated disruptions of editing, can be found in patients with cardiopathies. In this regard, hmtThrRS may be a valuable marker in patients with heart diseases in future clinical studies.
Materials and Methods
Generation of Knock-in mmtThrRS Mutation in Mice.
Tars2Flox/Flox knock-in mice were generated by homologous recombination using CRISPR/Cas9 by Cyagen (Suzhou) Biotechnology Co., Ltd. In short, a target vector containing the loxP-endogenous SA-Tars2 exon 4~18 CDS-polyA-SDA-Neo-SDA-loxP sequence was inserted into Intron 3, and the H138A (CAC to GCC) and H142A (CAT to GCT) mutations were introduced into Exon 4 in the 3′ homology arm. To express the mmtThrRS mutant in the whole mouse body, Tars2Flox/Flox mice were crossed with EIIa-Cre mice. Similarly, Tars2Flox/Flox mice were crossed with Myh6-Cre mice to obtain Tars2Flox/Flox-Cre mice with the mmtThrRS mutant specifically expressed in cardiomyocytes.
Detailed methods are described in SI Appendix, Materials and Methods.
Supplementary Material
Appendix 01 (PDF)
Acknowledgments
We thank the National Center for Protein Science Shanghai for LC–MS analysis of Thr-to-Ser misincorporation in mtDNA-encoded proteins. We are grateful to Dr. Zi Li (Shanghai Institute of Nutrition and Health, CAS) for LC–MS/MS analysis of amount of amino acid moiety attached to mtRNAThr. We also thank Dr. Huang-Tian Yang (Shanghai Institute of Nutrition and Health, CAS) for valuable suggestions in mouse experiments and Drs. Bao-Jin Wu and Kang Kang (Animal Core Facility, Shanghai Institute of Biochemistry and Cell Biology, CAS) for echocardiographic analysis of mice. This work was supported by the National Key Research and Development Program of China (2021YFA1300800 and 2021YFC2700903 to X.-L.Z.); the Natural Science Foundation of China (32271300 and 81870896 to X.-L.Z.; 91940302 and 31870811 to E.-D.W.); the Committee of Science and Technology in Shanghai (22ZR1481300 and 22JC1400503 to X.-L.Z.); and the CAS Project for Young Scientists in Basic Research (YSBR-075 to X.-L.Z.).
Author contributions
W.-Q.Z. and X.-L.Z. designed research; W.-Q.Z., J.-H.Z., Z.-H.L., X.L., Y.Z., and S.H. performed research; W.-Q.Z., J.L., B.Z., G.E., E.-D.W., and X.-L.Z. analyzed data; and X.-L.Z. wrote the paper.
Competing interests
The authors declare no competing interest.
Footnotes
This article is a PNAS Direct Submission.
Contributor Information
En-Duo Wang, Email: edwang@sibcb.ac.cn.
Xiao-Long Zhou, Email: xlzhou@sibcb.ac.cn.
Data, Materials, and Software Availability
All data presented in this study are available within the Figures, Table, and in SI Appendix. The RNA-seq data have been deposited to the NCBI Gene Expression Omnibus (GEO) under accession number GSE232410 (85).
Supporting Information
References
- 1.Guo M., Yang X. L., Schimmel P., New functions of aminoacyl-tRNA synthetases beyond translation. Nat. Rev. Mol. Cell Biol. 11, 668–674 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Kim S., You S., Hwang D., Aminoacyl-tRNA synthetases and tumorigenesis: More than housekeeping. Nat. Rev. Cancer 11, 708–718 (2011). [DOI] [PubMed] [Google Scholar]
- 3.Ibba M., Soll D., Quality control mechanisms during translation. Science 286, 1893–1897 (1999). [DOI] [PubMed] [Google Scholar]
- 4.Ling J., Reynolds N., Ibba M., Aminoacyl-tRNA synthesis and translational quality control. Annu. Rev. Microbiol. 63, 61–78 (2009). [DOI] [PubMed] [Google Scholar]
- 5.Zhou X., Wang E., Transfer RNA: A dancer between charging and mis-charging for protein biosynthesis. Sci. China Life Sci. 56, 921–932 (2013). [DOI] [PubMed] [Google Scholar]
- 6.Lin L., Hale S. P., Schimmel P., Aminoacylation error correction. Nature 384, 33–34 (1996). [DOI] [PubMed] [Google Scholar]
- 7.Jakubowski H., Quality control in tRNA charging. Wiley Interdiscip. Rev. RNA 3, 295–310 (2012). [DOI] [PubMed] [Google Scholar]
- 8.Nangle L. A., Motta C. M., Schimmel P., Global effects of mistranslation from an editing defect in mammalian cells. Chem. Biol. 13, 1091–1100 (2006). [DOI] [PubMed] [Google Scholar]
- 9.Lee J. W., et al. , Editing-defective tRNA synthetase causes protein misfolding and neurodegeneration. Nature 443, 50–55 (2006). [DOI] [PubMed] [Google Scholar]
- 10.Liu Y., et al. , Deficiencies in tRNA synthetase editing activity cause cardioproteinopathy. Proc. Natl. Acad. Sci. U.S.A. 111, 17570–17575 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Hilander T., et al. , Editing activity for eliminating mischarged tRNAs is essential in mammalian mitochondria. Nucleic Acids Res. 46, 849–860 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Kelly P., et al. , Alanyl-tRNA synthetase quality control prevents global dysregulation of the Escherichia coli proteome. mBio 10, e02921-19 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Zhou X. L., et al. , Role of tRNA amino acid-accepting end in aminoacylation and its quality control. Nucleic Acids Res. 39, 8857–8868 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Anderson S., et al. , Sequence and organization of the human mitochondrial genome. Nature 290, 457–465 (1981). [DOI] [PubMed] [Google Scholar]
- 15.Antonellis A., Green E. D., The role of aminoacyl-tRNA synthetases in genetic diseases. Annu. Rev. Genomics Hum. Genet. 9, 87–107 (2008). [DOI] [PubMed] [Google Scholar]
- 16.Chen X., et al. , Modular pathways for editing non-cognate amino acids by human cytoplasmic leucyl-tRNA synthetase. Nucleic Acids Res. 39, 235–247 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Zhou X. L., Ruan Z. R., Huang Q., Tan M., Wang E. D., Translational fidelity maintenance preventing Ser mis-incorporation at Thr codon in protein from eukaryote. Nucleic Acids Res. 41, 302–314 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Chen Y., et al. , A threonyl-tRNA synthetase-like protein has tRNA aminoacylation and editing activities. Nucleic Acids Res. 46, 3643–3656 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Song Y., et al. , p53-Dependent DNA damage response sensitive to editing-defective tRNA synthetase in zebrafish. Proc. Natl. Acad. Sci. U.S.A. 113, 8460–8465 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Vo M. N., et al. , ANKRD16 prevents neuron loss caused by an editing-defective tRNA synthetase. Nature 557, 510–515 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Kumar P., Bhatnagar A., Sankaranarayanan R., Chiral proofreading during protein biosynthesis and its evolutionary implications. FEBS Lett. 596, 1615–1627 (2022). [DOI] [PubMed] [Google Scholar]
- 22.Ruan L. L., Zhou X. L., Tan M., Wang E. D., Human cytoplasmic ProX edits mischarged tRNAPro with amino acid but not tRNA specificity. Biochem. J. 450, 243–252 (2013). [DOI] [PubMed] [Google Scholar]
- 23.Vargas-Rodriguez O., et al. , Human trans-editing enzyme displays tRNA acceptor-stem specificity and relaxed amino acid selectivity. J. Biol. Chem. 295, 16180–16190 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Klipcan L., et al. , Crystal structure of human mitochondrial PheRS complexed with tRNA(Phe) in the active “open” state. J. Mol. Biol. 415, 527–537 (2012). [DOI] [PubMed] [Google Scholar]
- 25.Lue S. W., Kelley S. O., An aminoacyl-tRNA synthetase with a defunct editing site. Biochemistry 44, 3010–3016 (2005). [DOI] [PubMed] [Google Scholar]
- 26.Ye Q., et al. , Degenerate connective polypeptide 1 (CP1) domain from human mitochondrial leucyl-tRNA synthetase. J. Biol. Chem. 290, 24391–24402 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Karkhanis V. A., Boniecki M. T., Poruri K., Martinis S. A., A viable amino acid editing activity in the leucyl-tRNA synthetase CP1-splicing domain is not required in the yeast mitochondria. J. Biol. Chem. 281, 33217–33225 (2006). [DOI] [PubMed] [Google Scholar]
- 28.Roy H., Ling J., Alfonzo J., Ibba M., Loss of editing activity during the evolution of mitochondrial phenylalanyl-tRNA synthetase. J. Biol. Chem. 280, 38186–38192 (2005). [DOI] [PubMed] [Google Scholar]
- 29.Reynolds N. M., et al. , Cell-specific differences in the requirements for translation quality control. Proc. Natl. Acad. Sci. U.S.A. 107, 4063–4068 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Peng G. X., et al. , RNA granule-clustered mitochondrial aminoacyl-tRNA synthetases form multiple complexes with the potential to fine-tune tRNA aminoacylation. Nucleic Acids Res. 50, 12951–12968 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Wang Y., et al. , A human disease-causing point mutation in mitochondrial threonyl-tRNA synthetase induces both structural and functional defects. J. Biol. Chem. 291, 6507–6520 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Zeng Q. Y., et al. , The G3–U70-independent tRNA recognition by human mitochondrial alanyl-tRNA synthetase. Nucleic Acids Res. 47, 3072–3085 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Sankaranarayanan R., et al. , The structure of threonyl-tRNA synthetase-tRNA(Thr) complex enlightens its repressor activity and reveals an essential zinc ion in the active site. Cell 97, 371–381 (1999). [DOI] [PubMed] [Google Scholar]
- 34.Carapito C., et al. , Two proteomic methodologies for defining N-termini of mature human mitochondrial aminoacyl-tRNA synthetases. Methods 113, 111–119 (2017). [DOI] [PubMed] [Google Scholar]
- 35.Guo R., Zong S., Wu M., Gu J., Yang M., Architecture of human mitochondrial respiratory megacomplex I(2)III(2)IV(2). Cell 170, 1247–1257.e12 (2017). [DOI] [PubMed] [Google Scholar]
- 36.Han Y. H., Moon H. J., You B. R., Park W. H., The effect of MG132, a proteasome inhibitor on HeLa cells in relation to cell growth, reactive oxygen species and GSH. Oncol. Rep. 22, 215–221 (2009). [PubMed] [Google Scholar]
- 37.Gustafsson C. M., Falkenberg M., Larsson N. G., Maintenance and expression of mammalian mitochondrial DNA. Annu. Rev. Biochem. 85, 133–160 (2016). [DOI] [PubMed] [Google Scholar]
- 38.Rath S., et al. , MitoCarta3.0: An updated mitochondrial proteome now with sub-organelle localization and pathway annotations. Nucleic Acids Res. 49, D1541–D1547 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Hoppins S., Lackner L., Nunnari J., The machines that divide and fuse mitochondria. Annu. Rev. Biochem. 76, 751–780 (2007). [DOI] [PubMed] [Google Scholar]
- 40.Salic A., Mitchison T. J., A chemical method for fast and sensitive detection of DNA synthesis in vivo. Proc. Natl. Acad. Sci. U.S.A. 105, 2415–2420 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.King R. W., Jackson P. K., Kirschner M. W., Mitosis in transition. Cell 79, 563–571 (1994). [DOI] [PubMed] [Google Scholar]
- 42.Harper J. W., Adami G. R., Wei N., Keyomarsi K., Elledge S. J., The p21 Cdk-interacting protein Cip1 is a potent inhibitor of G1 Cyclin-dependent kinases. Cell 75, 805–816 (1993). [DOI] [PubMed] [Google Scholar]
- 43.Gu Y., Turck C. W., Morgan D. O., Inhibition of Cdk2 activity in-vivo by an associated 20k regulatory subunit. Nature 366, 707–710 (1993). [DOI] [PubMed] [Google Scholar]
- 44.Xiong Y., et al. , p21 is a universal inhibitor of Cyclin kinases. Nature 366, 701–704 (1993). [DOI] [PubMed] [Google Scholar]
- 45.Puente B. N., et al. , The oxygen-rich postnatal environment induces cardiomyocyte cell-cycle arrest through DNA damage response. Cell 157, 565–579 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Eldeiry W. S., et al. , Waf1, a potential mediator of p53 tumor suppression. Cell 75, 817–825 (1993). [DOI] [PubMed] [Google Scholar]
- 47.Kastan M. B., Onyekwere O., Sidransky D., Vogelstein B., Craig R. W., Participation of p53 protein in the cellular-response to DNA damage. Cancer Res. 51, 6304–6311 (1991). [PubMed] [Google Scholar]
- 48.Guo Z., Kozlov S., Lavin M. F., Person M. D., Paull T. T., ATM activation by oxidative stress. Science 330, 517–521 (2010). [DOI] [PubMed] [Google Scholar]
- 49.Dan Dunn J., Alvarez L. A., Zhang X., Soldati T., Reactive oxygen species and mitochondria: A nexus of cellular homeostasis. Red. Biol. 6, 472–485 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Kussmaul L., Hirst J., The mechanism of superoxide production by NADH: Ubiquinone oxidoreductase (complex I) from bovine heart mitochondria. Proc. Natl. Acad. Sci. U.S.A. 103, 7607–7612 (2006). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Hans C., Saini R., Sachdeva M. U. S., Sharma P., 2’,7’-Dichlorofluorescein (DCF) or 2’,7’-dichlorodihydrofluorescein diacetate (DCFH2-DA) to measure reactive oxygen species in erythrocytes. Biomed. Pharmacother 138, 111512 (2021). [DOI] [PubMed] [Google Scholar]
- 52.Nystrom T., Role of oxidative carbonylation in protein quality control and senescence. EMBO J. 24, 1311–1317 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Kuroda J., et al. , NADPH oxidase 4 (Nox4) is a major source of oxidative stress in the failing heart. Proc. Natl. Acad. Sci. U.S.A. 107, 15565–15570 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Flynn J. M., Melov S., SOD2 in mitochondrial dysfunction and neurodegeneration. Free Radic. Biol. Med. 62, 4–12 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Halasi M., et al. , ROS inhibitor N-acetyl-L-cysteine antagonizes the activity of proteasome inhibitors. Biochem. J. 454, 201–208 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Kalyanaraman B., NAC, NAC, Knockin’ on Heaven’s door: Interpreting the mechanism of action of N-acetylcysteine in tumor and immune cells. Redox. Biol. 57, 102497 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Lanson N. A. Jr., Glembotski C. C., Steinhelper M. E., Field L. J., Claycomb W. C., Gene expression and atrial natriuretic factor processing and secretion in cultured AT-1 cardiac myocytes. Circulation 85, 1835–1841 (1992). [DOI] [PubMed] [Google Scholar]
- 58.Lakso M., et al. , Efficient in vivo manipulation of mouse genomic sequences at the zygote stage. Proc. Natl. Acad. Sci. U.S.A. 93, 5860–5865 (1996). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Bertero E., Maack C., Metabolic remodelling in heart failure. Nat. Rev. Cardiol. 15, 457–470 (2018). [DOI] [PubMed] [Google Scholar]
- 60.Agah R., et al. , Gene recombination in postmitotic cells. Targeted expression of Cre recombinase provokes cardiac-restricted, site-specific rearrangement in adult ventricular muscle in vivo. J. Clin. Invest. 100, 169–179 (1997). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Zhou X. L., et al. , Aminoacylation and translational quality control strategy employed by leucyl-tRNA synthetase from a human pathogen with genetic code ambiguity. Nucleic Acids Res. 41, 9825–9838 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Zhu B., Yao P., Tan M., Eriani G., Wang E. D., tRNA-independent pretransfer editing by class I leucyl-tRNA synthetase. J. Biol. Chem. 284, 3418–3424 (2009). [DOI] [PubMed] [Google Scholar]
- 63.Yao P., et al. , Unique residues crucial for optimal editing in yeast cytoplasmic Leucyl-tRNA synthetase are revealed by using a novel knockout yeast strain. J. Biol. Chem. 283, 22591–22600 (2008). [DOI] [PubMed] [Google Scholar]
- 64.Ji Q. Q., Fang Z. P., Ye Q., Chi C. W., Wang E. D., Self-protective responses to norvaline-induced stress in a leucyl-tRNA synthetase editing-deficient yeast strain. Nucleic Acids Res. 45, 7367–7381 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Cvetesic N., Palencia A., Halasz I., Cusack S., Gruic-Sovulj I., The physiological target for LeuRS translational quality control is norvaline. EMBO J. 33, 1639–1653 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Ross-Inta C., Tsai C. Y., Giulivi C., The mitochondrial pool of free amino acids reflects the composition of mitochondrial DNA-encoded proteins: Indication of a post- translational quality control for protein synthesis. Biosci. Rep. 28, 239–249 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Wong F. C., Beuning P. J., Silvers C., Musier-Forsyth K., An isolated class II aminoacyl-tRNA synthetase insertion domain is functional in amino acid editing. J. Biol. Chem. 278, 52857–52864 (2003). [DOI] [PubMed] [Google Scholar]
- 68.An S., Musier-Forsyth K., Trans-editing of Cys-tRNAPro by Haemophilus influenzae YbaK protein. J. Biol. Chem. 279, 42359–42362 (2004). [DOI] [PubMed] [Google Scholar]
- 69.LaRiviere F. J., Wolfson A. D., Uhlenbeck O. C., Uniform binding of aminoacyl-tRNAs to elongation factor Tu by thermodynamic compensation. Science 294, 165–168 (2001). [DOI] [PubMed] [Google Scholar]
- 70.Schrader J. M., Chapman S. J., Uhlenbeck O. C., Tuning the affinity of aminoacyl-tRNA to elongation factor Tu for optimal decoding. Proc. Natl. Acad. Sci. U.S.A. 108, 5215–5220 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Loftfield R. B., Vanderjagt D., The frequency of errors in protein biosynthesis. Biochem. J. 128, 1353–1356 (1972). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Rodnina M. V., Quality control of mRNA decoding on the bacterial ribosome. Adv Protein Chem. Struct. Biol. 86, 95–128 (2012). [DOI] [PubMed] [Google Scholar]
- 73.Pape L. K., Koerner T. J., Tzagoloff A., Characterization of a yeast nuclear gene (MST1) coding for the mitochondrial threonyl-tRNA1 synthetase. J. Biol. Chem. 260, 15362–15370 (1985). [PubMed] [Google Scholar]
- 74.Ling J., Peterson K. M., Simonovic I., Soll D., Simonovic M., The mechanism of pre-transfer editing in yeast mitochondrial threonyl-tRNA synthetase. J. Biol. Chem. 287, 28518–28525 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Zhou X. L., et al. , A minimalist mitochondrial threonyl-tRNA synthetase exhibits tRNA-isoacceptor specificity during proofreading. Nucleic Acids Res. 42, 13873–13886 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Kim S. H., et al. , Mitochondrial threonyl-tRNA synthetase TARS2 is required for threonine-sensitive mTORC1 activation. Mol. Cell 81, 398–407.e4 (2021). [DOI] [PubMed] [Google Scholar]
- 77.Chung C. Y., Valdebenito G. E., Chacko A. R., Duchen M. R., Rewiring cell signalling pathways in pathogenic mtDNA mutations. Trends Cell Biol. 32, 391–405 (2022). [DOI] [PubMed] [Google Scholar]
- 78.Rackham O., Filipovska A., Organization and expression of the mammalian mitochondrial genome. Nat. Rev. Genet. 23, 606–623 (2022). [DOI] [PubMed] [Google Scholar]
- 79.Tsutsui H., Tsuchihashi M., Takeshita A., Mortality and readmission of hospitalized patients with congestive heart failure and preserved versus depressed systolic function. Am. J. Cardiol. 88, 530–533 (2001). [DOI] [PubMed] [Google Scholar]
- 80.Wende A. R., Brahma M. K., McGinnis G. R., Young M. E., Metabolic origins of heart failure. JACC Basic Transl. Sci. 2, 297–310 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Allard M. F., Schönekess B. O., Henning S. L., English D. R., Lopaschuk G. D., Contribution of oxidative metabolism and glycolysis to ATP production in hypertrophied hearts. Am. J. Physiol. 267, H742–H750 (1994). [DOI] [PubMed] [Google Scholar]
- 82.Lopaschuk G. D., Karwi Q. G., Tian R., Wende A. R., Abel E. D., Cardiac energy metabolism in heart failure. Circ. Res. 128, 1487–1513 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Zheng W. Q., et al. , Elucidating the molecular mechanisms associated with TARS2-related mitochondrial disease. Hum. Mol. Genet. 31, 523–534 (2022). [DOI] [PubMed] [Google Scholar]
- 84.Diodato D., et al. , VARS2 and TARS2 mutations in patients with mitochondrial encephalomyopathies. Hum. Mutat. 35, 983–989 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Zheng W., et al. , Mammalian mitochondrial translation infidelity leads to oxidative stress-induced cell cycle arrest and cardiomyopathy. NCBI Gene Expression Omnibus (GEO). https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=gse232410. Deposited 25 August 2023. [DOI] [PMC free article] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Appendix 01 (PDF)
Data Availability Statement
All data presented in this study are available within the Figures, Table, and in SI Appendix. The RNA-seq data have been deposited to the NCBI Gene Expression Omnibus (GEO) under accession number GSE232410 (85).