Summary
To explore mechanisms of response to combined PD-1/CTLA-4 immune checkpoint blockade (ICB) treatment in individual cell types, we generated scRNA-seq using a mouse model of invasive urothelial carcinoma with three conditions: untreated tumor, treated tumor, and tumor treated after CD4+ T cell depletion. After classifying tumor cells based on detection of somatic variants and assigning non-tumor cell types using SingleR, we performed differential expression analysis, overrepresentation analysis, and gene set enrichment analysis (GSEA) within each cell type. GSEA revealed that endothelial cells were enriched for upregulated IFN-g response genes when comparing treated cells to both untreated cells and cells treated after CD4+ T cell depletion. Functional analysis showed that knocking out IFNgR1 in endothelial cells inhibited treatment response. Together, these results indicated that IFN-g signaling in endothelial cells is a key mediator of ICB induced anti-tumor activity.
Subject areas: Oncology, Immunology, Transcriptomics
Graphical abstract

Highlights
-
•
scRNA-seq reveals basal- and luminal-like tumor populations in a bladder cancer model
-
•
scRNA-seq identifies a rare endothelial cell population in the microenvironment
-
•
IFN-g response is upregulated in endothelial cells during ICB treatment
-
•
IFNgR1 knockout in endothelial cells negates effective ICB treatment response
Oncology; Immunology; Transcriptomics
Introduction
Bladder cancer is a common malignancy worldwide (6th most common among men and 17th most common among women) and accounts for over 500,000 new cancer diagnoses and 200,000 cancer-related deaths per year.1 While over 95% of bladder cancer cases are classified as urothelial carcinomas, they encompass a range of molecular subtypes, which are primarily distinguished by differential expression of differentiation markers and may predict for response to specific treatments.2,3 Initial diagnosis stages can be broadly grouped into non-muscle invasive (NMIBC), muscle invasive (MIBC), and metastatic disease. About 75% of cases are initially diagnosed as NMIBC, 20% as MIBC, and the remaining 5% as metastatic. Depending on the initial degree of invasiveness and metastasis, 5-year survival rates can range from 96% to 6%.4
Standard treatment recommendations likewise depend on the initial degree of invasiveness as well as the risk stratification of recurrence and progression. NMIBC is typically treated with a transurethral resection followed by either chemotherapy, Bacillus Calmette-Guerin immunotherapy, or radical cystectomy in high risk cases.2 MIBC is typically treated with neoadjuvant chemotherapy followed by radical cystectomy and, in some cases, adjuvant immunotherapy. Previous research has suggested that response to treatment may differ by subtype. For example, basal/squamous bladder cancers may have better response to neoadjuvant chemotherapy than luminal-infiltrated tumors.3 While neoadjuvant chemotherapy has historically been used most commonly, clinical trials looking at the use of neoadjuvant immune checkpoint blockade (ICB) treatments have shown promise as well.5,6
Currently, there are several ICB treatments approved by the FDA for treatment of bladder cancer, all of which are either PD-1 or PD-L1 inhibitors.7,8 Initially, these treatments were approved specifically for treatment of advanced disease, targeting patients who were ineligible for cisplatin treatment.9 Over time, ICB use has become more widespread and has been applied across the range of bladder cancer stages from NMIBC to metastatic disease.8,10,11 While ICB therapy shows great promise for treatment of bladder cancer, there are still many patients who do not receive benefit from ICB treatment. Thus, there remains a need to improve treatment methods, determine which patients will respond well to treatment, understand mechanisms of response to treatment, and identify potential predictors of response.5
Clinical trials examining the benefit of PD-1/PD-L1 inhibitors in urothelial carcinoma have found that high IFN-g expression is associated with treatment response, suggesting that IFN-g signatures could serve as a predictor of response.12 Additionally, treatment response has been associated with high expression of CXCL9 and CXCL10, two IFN-g induced chemokines that have been associated with increased T cell infiltration in multiple tumor types.13,14 However, these trials did not fully explore how or where IFN-g may be acting to help induce or improve treatment response. Clinical trials have also looked at improving treatment response by combining PD-1/PD-L1 inhibitor treatments with CTLA-4 inhibitor treatments. These trials have shown greater response rates compared to PD-1/PD-L1 monotherapy.15,16,17,18 Nevertheless, the challenges of identifying ideal patients for treatment as well as identifying mechanisms and predictors of response remain.
To study mechanisms of response to combined PD-1/CTLA-4 ICB treatment, we used a murine muscle-invasive urothelial carcinoma cell line generated by exposing mice to 4-hydroxybutyl(butyl)nitrosamine (BBN), which caused them to develop areas of invasive disease. These tumor bearing bladders were then resected and used to propagate an organoid cell line, MCB6C.19 Previous analysis showed that MCB6C is responsive to ICB treatments and achieves the best treatment response with combined PD-1/CTLA-4 ICB treatment. Additionally, this previous work showed that treatment response was dependent on CD4+ T cells and not dependent CD8+ T cells, consistent with research showing that CD4 T cells may be the primary mediators of anti-tumor activity in human bladder cancer.19,20 Analysis of the MCB6C model also showed that ICB treatment led to an increase of IFN-g producing CD4+ T cells with a Th-1 like phenotype. Neutralizing IFN-g in the tumor negated the anti-tumor activity of combined treatment, indicating that IFN-g was a key mediator of response. Surprisingly, this research showed that knocking out IFNgR1 in the tumor cells themselves did not affect treatment response, suggesting that IFN-g was mediating treatment response through non-tumoral cells in the tumor microenvironment (TME).19
To better understand mechanisms of treatment response in this model, we performed single cell RNA sequencing (scRNA-seq) on MCB6C tumors isolated from mice under three conditions: untreated tumor, tumor treated with combined PD-1/CTLA-4 ICB treatment, and tumor treated with combined ICB treatment after CD4+ T cell depletion (Figure 1A). For each condition in each replicate, tumors from three mice were resected and pooled to generate single cell suspensions for 10x Genomics 5′ gene expression sequencing as well as B cell and T cell receptor (TCR) sequencing (Figure 1B, STAR Methods). This sequencing was performed for five biological replicates. In addition to scRNA-seq, whole genome and exome sequencing of the tumor cell line were performed, along with matched normal whole genome and exome sequencing of a tail sample (Figure 1C, STAR Methods).
Figure 1.
Experimental design for single cell RNA and bulk DNA sequencing
(A) Timelines for generating tumor samples for individual mice for each condition.
(B) Workflow for generating single cell suspensions for single cell RNA sequencing for one of five biological replicates sequenced. For each condition in each replicate, tumors from three individual mice were pooled into one suspension and used to create libraries for 10x Genomics 5′ single cell gene expression (GEX), B cell receptor (BCR), and T cell receptor (TCR) sequencing.
(C) Sources for normal and tumor bulk DNA sequencing. DNA was isolated from a normal mouse tail sample and an MCB6C tumor cell line sample for whole genome sequencing (WGS) and whole exome sequencing (WES). ICB = combined PD-1/CTLA-4 immune checkpoint blockade treatment, ICBΔT = combined PD-1/CTLA-4 immune checkpoint blockade treatment received after CD4+ T cell depletion, Isotype control = rat IgG2a and mouse IgG2b.
Results
Bulk DNA sequencing shows that the MCB6C cell line has a high mutation burden, normal ploidy, and a stable genome
Bulk whole genome sequencing (WGS) of the tumor cell line generated over one billion paired reads, 88% of which produced high quality alignments (i.e., had a mapping score of Q20 or greater). Bulk WGS of the normal tail sample produced over 1.1 billion reads, with approximately 91% of reads having high quality alignments. Bulk whole exome sequencing (WES) of the tumor cell line produced over 55 million reads with over 90% of reads having high quality alignments, while WES of the tail sample produced over 77 million reads with over 90% of reads having high quality alignments (Table S1A).
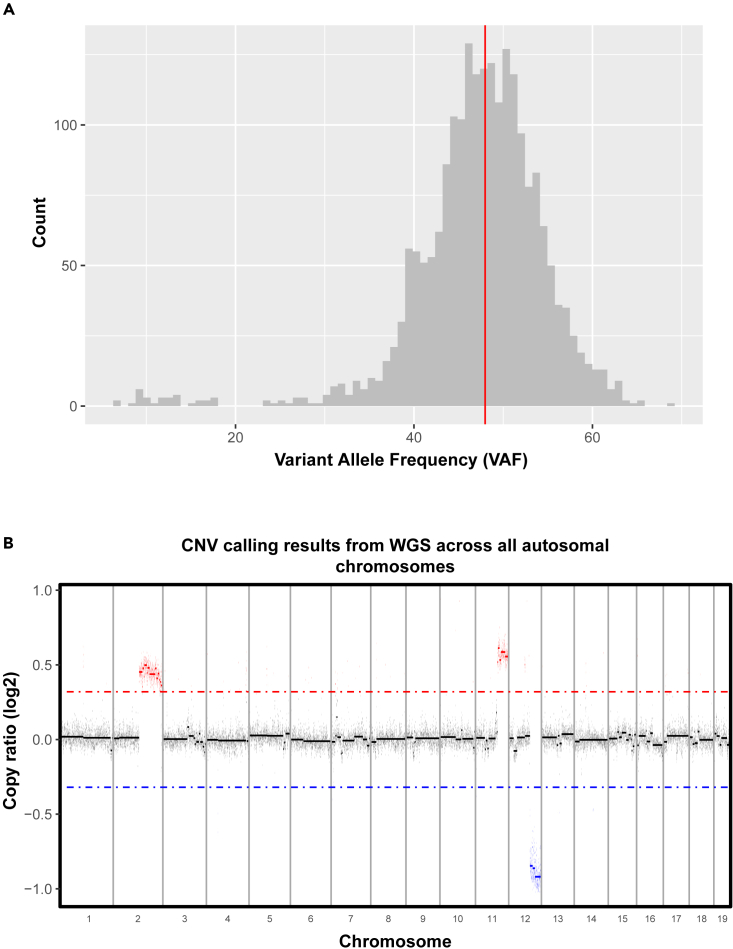
After alignment, we performed somatic variant calling with the WES data and identified 16,449 possible somatic variants, including 16,315 single nucleotide variants (SNVs) and 134 small insertions or deletions (indels), before filtering. These variants were then filtered using several metrics, including total coverage, variant allele frequency (VAF), and consensus across somatic variant callers (STAR Methods). 10,427 variants remained after filtering, of which 10,407 were SNVs and 20 were indels, showing that the MCB6C cell line has a high SNV burden (approximately 4.17 mutations per Mb) consistent with a mutagen induced tumor model. We then characterized the clonality of the cell line by examination of the VAF distribution (STAR Methods).21 This distribution appeared to be centered close to 50%, with a median VAF of ∼ 48.0%, and had a near-normal distribution (Figure 2A; Table S1B). However, there was a small group of low VAF (i.e., VAF of 20% or less) variants detected. Out of the 2,128 variants used to assess clonality based on VAF distribution, 31 variants had low VAF values. These variants could represent a subclone within the cell line.
Figure 2.
Bulk DNA sequencing shows that the MCB6C cell line has a high mutation burden, normal ploidy, and a stable genome
(A) Distribution of variant allele frequencies (VAFs) for the set of somatic SNVs used to assess clonality. Only variants with 100× normal and tumor coverage, normal VAF = 0%, tumor VAF >5%, and no overlap of CNV regions were used. SNVs were detected using matched tumor and normal whole exome sequencing of the MCB6C cell line. Red line indicates overall median VAF (∼48.0%).
(B) Visualization of copy number variants detected using matched tumor and normal whole genome sequencing of the MCB6C cell line. Copy number calling was performed using CNVkit with 100k bin size. Dots represent read count differences in bins. Solid lines represent segments identified by CNVkit using circular binary segmentation. The red dotted line corresponds to a log2 copy ratio of 0.32. Bins and segments falling above this line are classified as copy gains. The blue dotted line corresponds to a log2 copy ratio of −0.32. Bins and segments falling below this line are classified as copy losses. Three chromosomes exhibited large segments of copy number alteration: chr2 (copy gain of ∼86.9 Mb), chr11 (copy gain of ∼41.2 Mb), and chr12 (copy loss of ∼43.0 Mb). See also Tables S1B and S1D.
Looking at individual somatic mutations, we confirmed three driver mutations (Kras G12D, Trp53 T122K, and Kdm6a H1146Y) for the MCB6C cell line, which were previously reported from analysis of bulk whole transcriptome sequencing (RNA-seq).19 Along with these three mutations, we identified 31 additional mutations across 20 previously reported driver genes in human bladder cancer, including a second Trp53 mutation (a splice donor variant) and a second missense Kdm6a mutation (Table S1C, STAR Methods).22 This set also included mutations in Atm (S1884T), Fat1 (two missense, one stop-gained mutations), Kmt2a (H1067Q), and Kmt2c (one splice region mutation), which have each been shown to harbor mutations in over 10% of bladder cancers, although none of the specific mutations identified appear to have been previously reported in bladder cancer.3 In addition to the stop-gained and splice region mutations identified in Atm and Kmt2c, respectively, two additional stop-gained mutations (one in Birc6 and one in Rnf213) and three additional splice region variants (one in Birc6, one in Brca2, and one in Sf3b1) were identified. Finally, a mutation in Sf3b1 (E873K), which was identified as a possible driver of a similar mouse urothelial carcinoma cell line, but was not previously detected in MCB6C using RNA-seq, was detected using WES.19
In addition to calling SNVs and indels, we called copy number variants using the WGS data (STAR Methods). These results indicated that the MCB6C cell line has a relatively stable genome with only a few larger regions of copy number alteration consisting of copy gains on chromosomes 2 and 11 and a single copy loss on chromosome 12 (Figure 2B; Table S1D). Together, these results indicated that the MCB6C cell line has high SNV burden and low CNV burden. Previous research has shown that metastatic urothelial carcinoma patients with high SNV/low CNV tumor profiles may benefit more from ICB therapy. While high SNV/low CNV status has been associated with greater chance of response, the utility of SNV and CNV status is still being evaluated as a possible predictor of treatment response in bladder cancer.23
scRNA-seq was generated for over 64,000 cells, with over 59,000 cells passing filtering
10x Genomics 5′ single cell gene expression sequencing (scRNA-seq) was performed for five biological replicates, with each replicate consisting of three conditions: untreated tumor, treated tumor, and tumor treated after CD4+ T cell depletion (Figure 1A). In total, fifteen samples were sequenced, generating ∼8.3 billion reads across 64,049 cells (Table S2A). In addition to gene expression sequencing, 10x Genomics V(D)J B cell receptor (BCR) and T cell receptor (TCR) sequencing was also performed for all fifteen samples (Figure 1B; Tables S2B and S2C).
Before analyzing the scRNA-seq data, we aggregated all three conditions for each replicate and performed basic filtering on each of the five replicates to remove cells that appeared to be low quality based on mitochondrial gene expression per cell, detected gene count per cell, and/or total UMI count per cell. Briefly, cells expressing high percentages of mitochondrial genes, cells with low gene counts, and cells with high UMI counts were removed (Table S3, STAR Methods). Ultimately, 4,708 cells across all fifteen samples were removed, with 59,341 cells remaining.
scRNA-seq allows identification of lymphocyte, myeloid, and stromal cell populations in the tumor microenvironment
After completing basic filtering of cells, we used SingleR with the ImmGen dataset to assign fine label cell types to all remaining cells from each replicate.24,25,26 We then further filtered the set of remaining cells, removing all cells marked as “pruned” by SingleR. “Pruned cells” are those cells that have received poor-quality cell type assignments, potentially because of underlying poor quality of the cell itself. Once we removed all pruned cells, we were left with 57,818 cells total across all conditions and all replicates, which we aggregated into a single gene-barcode matrix for downstream analysis.
Next, we performed manual curation of SingleR’s fine label cell type assignments to group fine labels of the same broad cell type and to identify subtypes within certain broad cell types, e.g., to identify naive CD4 and CD8 T cells within the broader CD4 and CD8 T cell populations (STAR Methods). We also confirmed the general accuracy of the cell type assignments. First, for several cell types, we picked one reported marker for each cell type (e.g., Cd79a for B cells, Epcam for epithelial cells, and Col3a1 for fibroblasts) and compared the expression of each marker in the cell type expected to express it (based on the SingleR cell type assignment) versus all other cell types (Figures S1A–S1D; Table S4). These plots confirmed that the expected cell types generally showed more common and higher expression of their markers than non-expected cell types. Additionally, cells identified as expressing BCR sequences (i.e., cells that are likely B cells) or TCR sequences (i.e., cells that are likely T or NK cells) were compared to cells labeled as either B or T/NK cells, respectively, according to their gene expression signatures. These results showed that approximately 92.8% of cells identified as expressing BCR sequences were labeled as B cells by SingleR and 98.9% of cells identified as expressing TCR sequences were labeled as some type of T or NK cell, i.e., CD4, CD8, NK, NKT, Tgd, or Treg cells (Figures S1E and S1F; Table S4).
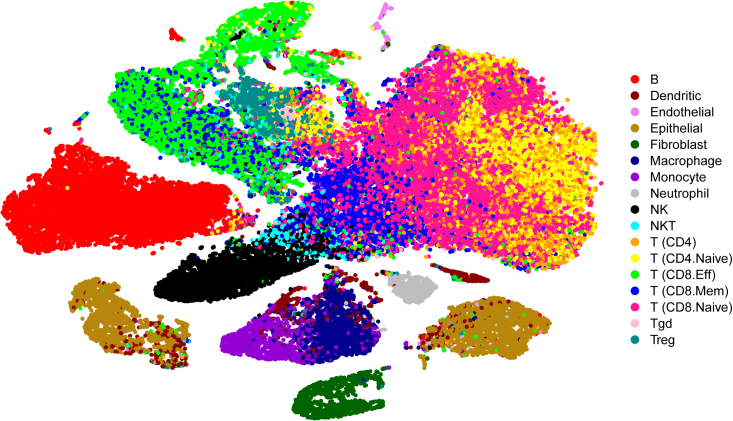
Before beginning any additional analysis, we also filtered out lowly expressed genes. For a gene to pass filtering, we required the gene to be detected in two or more cells in each replicate, with a UMI count of at least two in each cell. After filtering genes based on these criteria, we were left with 11,398 genes. Finally, we generated a tSNE projection for the aggregated dataset and colored cells by their manually curated cell type labels (Figure 3; Table S4, STAR Methods). This tSNE projection suggested that epithelial cells formed two distinct clusters, indicating there may be two transcriptionally distinct populations of epithelial cells within the dataset (discussed extensively in the following section).
Figure 3.
scRNA-seq allows identification of lymphocyte, myeloid, and stromal cell populations in the tumor microenvironment
tSNE projection of the aggregated dataset containing 57,818 cells, across all replicates and conditions, that passed filtering and were not “pruned” by SingleR. Cells were clustered using PCs = 20. Cells are colored by manually curated SingleR cell types. See also Figure S1 and Table S4.
After cell typing was completed, we also explored the BCR and TCR sequencing results. This analysis revealed little evidence of clonotype expansion of BCRs or TCRs across any of the conditions or replicates. For the TCR analysis, we excluded samples containing less than 1,500 TCR+ cells after filtering. We found that the majority of samples with less than 1,500 cells did not appear to have had their naive T cell populations captured during sequencing, which could lead to a skewed appearance of clonal expansion among the T cell populations that were captured (Figures S2A and S2B). For the remaining samples, there did not appear to be any dominant clonotypes detected (Figure S3A). The most commonly observed clonotype in any sample was present in only 41 cells and the vast majority of clonotypes were present in only one cell (Figure S3B). While no samples had evidence of dominant clonotypes, several samples did show evidence of cells with modestly expanded clonotypes (i.e., clonotypes detected in more than one cell) clustering together on the tSNE projection (Figure S4A). These clonotypes were largely found to be expressed in CD8 effector T cells, which showed evidence of co-expression of some exhaustion markers (Figures S4B, S5A, and S5B). For BCR analysis, most samples (11 of 15) had less than 1,000 BCR+ cells that passed filtering (Figure S6A). For samples where more than 1,000 cells passed filtering, there was little evidence of clonal expansion (Figure S6B).
Somatic variation can be used to identify tumor cell populations with high confidence
Since we expected tumor tissue to be epithelial, we expected that the epithelial populations identified by SingleR would correspond to the tumor cell populations. To verify this expectation, we classified cells as tumor or non-tumor based on the presence or absence of somatic mutations as follows.
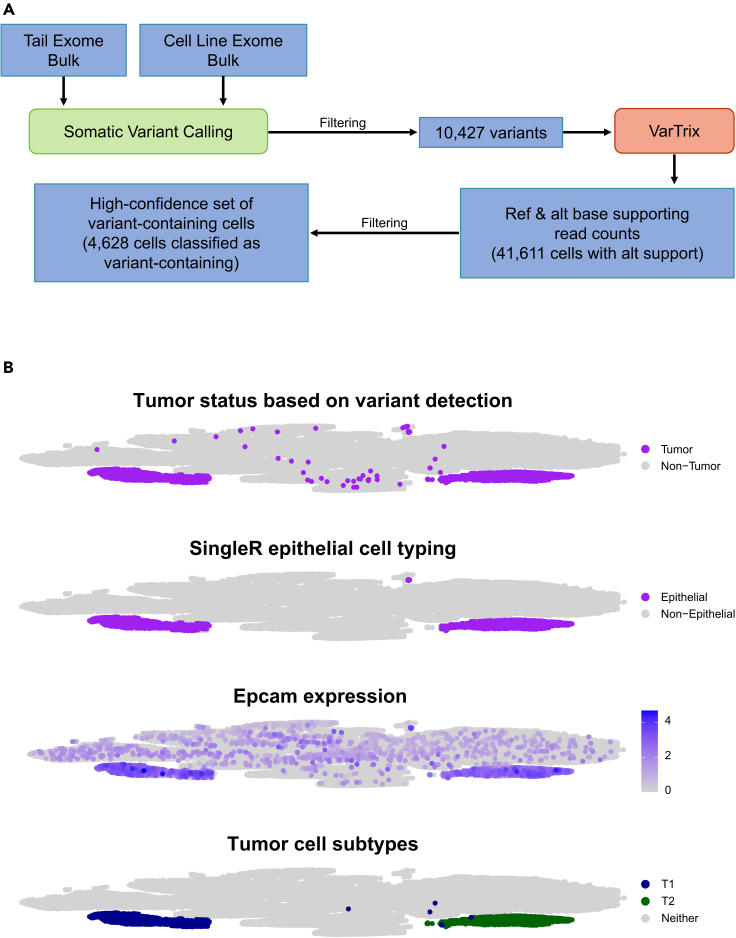
With the 10,427 somatic variants identified from WES, we used VarTrix to detect supporting reads for the reference and alternate alleles at each variant position in each individual cell in the aggregated dataset (Figure 4A). To identify a high confidence set of variant-containing cells, we required a cell to have at least two variant positions with greater than 20× total coverage, greater than five reads supporting the alternate allele, and a VAF over 10%. Using these criteria, we classified 4,628 cells as somatic variant-containing cells.
Figure 4.
Somatic variation can be used to identify tumor cell populations with high confidence
(A) Workflow for detecting somatic variation in scRNA-seq data to identify variant-containing cells.
(B) tSNE projections showing the classification of tumor cells based on variant detection, epithelial cell typing from SingleR, Epcam expression, and labeling of tumor subpopulations (T1 and T2) based on clustering, respectively. See also Table S5.
These variant-containing cells largely formed two distinct clusters on the tSNE projection, which heavily overlapped the two clusters identified as epithelial clusters using SingleR’s cell type labels (Figure 4B; Table S5). Since we expected the tumor tissue to be epithelial tissue, this extensive overlap appeared to confirm that variant-positive status could be used to identify tumor cells with high confidence. Additionally, we compared the overlap of variant-positive cells, cells that were assigned as epithelial cells by SingleR, and cells that were expressing Epcam, a marker of epithelial tissue. While the two variant-positive, epithelial-typed clusters showed high, widespread expression of Epcam as expected, there was also expression of Epcam detected across numerous other clusters (Figure 4B). These results indicated that variant status could be used to distinguish Epcam+ cells that are epithelial tumor cells from cells that appear to be Epcam+, but are not likely to be tumor cells (i.e., variant-negative, non-epithelial labeled cells) and may have simply been contaminated by ambient Epcam RNA.
Tumor cell populations show evidence of two distinct subpopulations
After confirming which cells and clusters corresponded to tumor cell populations, we investigated why tumor cells appeared to form two distinct clusters, which we labeled T1 and T2 (Figure 4B). To further explore whether these populations truly represented distinct tumor cell populations, we separated the tumor clusters from the rest of the aggregated dataset and reclustered them (STAR Methods). The tSNE projection again revealed distinct clustering of each of the two subpopulations (Figure 5A). We then assigned relative differentiation scores to each cell using CytoTRACE.27 We also performed differential expression analysis comparing the T1 subpopulation, containing all three conditions, to the T2 subpopulation, also containing all three conditions (STAR Methods).
Figure 5.
Tumor cell populations show evidence of two distinct subpopulations
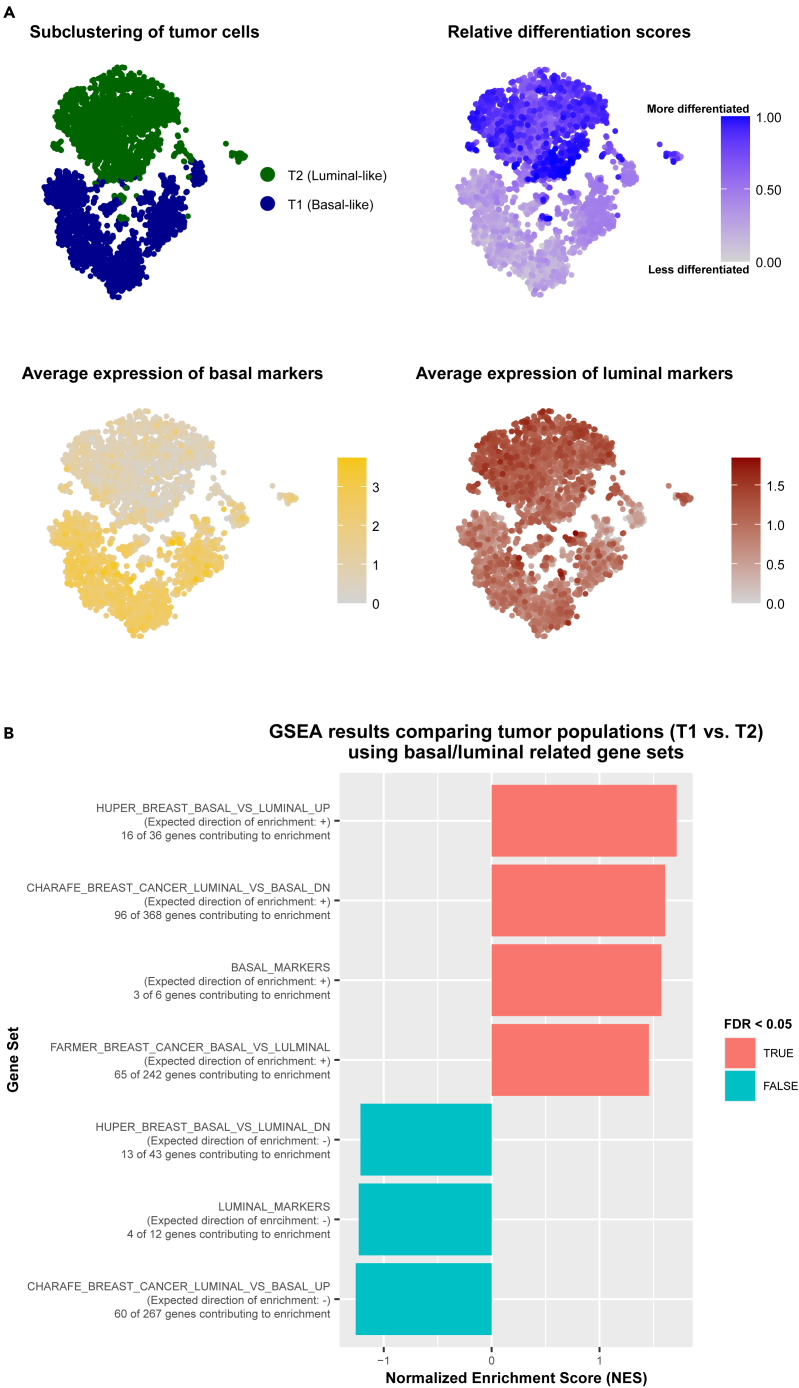
(A) tSNE projections of tumor cell populations showing subpopulation labels, differentiation scores (1 - CytoTRACE scores), average expression of basal bladder cancer markers, and average expression of luminal bladder cancer markers. Differentiation scores indicate the relative differentiation states of each cell within the full tumor cell population. Differentiation scores close to 1.00 indicate cells are relatively more differentiated. Differentiation scores close to 0.00 indicate cells are relatively less differentiated. Cells were clustered using PCs = 20.
(B) Bar plot showing the normalized enrichment scores (NES) for GSEA of gene sets related to basal and luminal bladder and breast cancer gene expression. Bars are colored by their FDR q-value status. Salmon pink indicates a significant FDR q-value (<0.05). Blue indicates a non-significant FDR q-value (>0.05). See also Tables S6A and S6B.
The relative differentiation scores revealed that the T2 cells largely corresponded to the most highly differentiated cells, while the T1 cells appeared to form two groups of cells—one which corresponded to the least differentiated cells and one which corresponded to slightly more differentiated, but still relatively lowly differentiated cells (Figure 5A; Table S6A). As previous literature has established that luminal bladder cancers display characteristics of greater differentiation than basal bladder cancers, we next wanted to determine if these two subpopulations might have different basal-like and luminal-like expression patterns.3
First, we calculated the average expression of reported basal (Cd44, Krt14, Krt5, Krt16, Krt6a) and luminal (Cd24a, Erbb2, Erbb3, Foxa1, Gata3, Gpx2, Krt18, Krt19, Krt7, Krt8, Upk1a) bladder cancer markers in each tumor cell and overlaid the values on the tSNE projection of the tumor populations (Figure 5A; Table S6A).28,29 These results indicated that the less differentiated cells (the T1 population) had stronger and more widespread expression of basal markers than the more differentiated cells (the T2 population), consistent with the expectation that basal-like bladder cancer cells would be less differentiated than luminal-like bladder cancer cells (Figure 5A). Both populations showed expression of luminal markers. However, the less differentiated T1 population appeared to have lower levels of luminal marker expression overall, suggesting that the T1 population could represent a population with more basal-like expression, while the T2 population could represent a population with more luminal-like expression.
Next, after generating differential expression analysis results for comparing the full basal-like population (T1) to the full luminal-like population (T2), we performed gene set enrichment analysis (GSEA) using gene sets generated by comparing basal and luminal breast cancers, which have been shown to have highly similar expression profiles to basal and luminal bladder cancers (STAR Methods).30 We also included two gene sets that we generated from reported lists of basal and luminal bladder cancer markers, respectively.28,29 These results indicated that upregulated genes in the T1 population were significantly enriched (FDR <0.05) for basal bladder cancer markers. These upregulated genes were also significantly enriched for genes that were found to be upregulated in basal breast cancers compared to luminal breast cancers (Figure 5B; Table S6B). Both of these observations were consistent with the T1 population being more basal-like than the T2 population.
By contrast, genes that were downregulated in the T1 population showed enrichment of luminal bladder cancer markers as well as genes that were found to be downregulated in basal breast cancers compared to luminal breast cancers. While this enrichment was not significant at an FDR cutoff of 0.05, the direction of enrichment was consistent with the observation that the T2 population appeared to have stronger expression of luminal markers than the T1 population (Figure 5B; Table S6B). These results further indicated that there were two distinct subpopulations of tumor cells within the full tumor cell population, i.e., a population with more basal-like characteristics and a population with more luminal-like characteristics.
Overrepresentation and gene set enrichment analysis identify IFN-g response as a commonly perturbed gene set across immune and tumor cell types upon ICB treatment
After assigning cell types and subtypes, where appropriate, to all cells, we explored how each individual cell type was responding to ICB treatment. To do this, we performed differential expression analysis comparing each possible pair of conditions within each cell type (STAR Methods). We then used the results of these differential expression analyses to perform overrepresentation analysis and GSEA.
Overrepresentation analysis showed that the top 5 most commonly overrepresented hallmark gene sets across cell types and comparisons were related to immune response (Table S7A). The IFN-g response gene set was the second most commonly overrepresented gene set (Figure 6A). Given that prior research suggested that IFN-g within the TME may be an important mediator of treatment response, we chose to explore the IFN-g response gene set further.19
Figure 6.
Overrepresentation and gene set enrichment analysis identify IFN-g response as a commonly perturbed gene set across immune and tumor cell types upon ICB treatment
(A) Heatmap showing the overrepresentation status of MSigDB’s hallmark IFN-g response gene set across each pairwise comparison of conditions in each cell type. A green square indicates IFN-g response genes were significantly overrepresented for the given comparison and cell type. A gray square indicates IFN-g response genes were not significantly overrepresented for the given comparison and cell type.
(B) Sum of fold changes for hallmark IFN-g response genes for each pairwise comparison of conditions in endothelial cells. Positive values indicate that IFN-g response genes skew toward upregulation in the first condition of a given comparison.
(C) Fold changes for the top 10 genes contributing to gene set enrichment of MSigDB’s hallmark IFN-g response gene set for each pairwise comparison of conditions in endothelial cells. ICB = combined PD-1/CTLA-4 immune checkpoint blockade treatment, ICBΔT = combined PD-1/CTLA-4 immune checkpoint blockade treatment received after CD4+ T cell depletion. See also Figures S7A and S7B and Table S7.
Since the overrepresentation analysis did not include information about the directionality or magnitude of overrepresentation, we generated a quantitative metric that captured both these aspects. Specifically, we summed the average log2 fold changes reported by Seurat for each detected gene in the IFN-g response gene set. With this method, a positive value indicates that genes from the gene set skew toward upregulation in the first condition of a given comparison in a given cell type, while a negative value indicates that genes skew toward downregulation. These sums indicated that ICB-treated endothelial cells experience upregulation of IFN-g response genes when compared to both untreated endothelial cells and endothelial cells treated after CD4+ T cell depletion (Figure 6B). These sums also suggested that untreated endothelial cells experience upregulation of IFN-g response genes when compared to endothelial cells treated after CD4+ T cell depletion. Endothelial cells appeared to be the only non-tumor cell type that experienced upregulation across all three comparisons (Figure S7A; Table S7B).
To explore enrichment of up- and downregulated genes more formally, we performed ranked GSEA, using average log2 fold changes as the ranking metric and MSigDB’s hallmark gene sets as the test set (STAR Methods). Similar to the results seen with the overrepresentation analysis, we found that the IFN-g response gene set was commonly enriched across multiple cell types. Furthermore, the enrichment results followed similar patterns to those seen using the “sum of fold changes” metric. When looking at the ICB treated condition versus both the control and CD4+ T cell depleted conditions, endothelial cells showed significant enrichment of upregulated IFN-g response genes (Figure S7B). Additionally, endothelial cells were the only cell type to show significant positive enrichment across all three comparisons (Figure S7B). Examination of the top genes contributing to enrichment in each comparison within endothelial cells indicated that upregulation of chemokines, such as Cxcl9 and Cxcl10, and adhesion molecules, such as Vcam1 and Icam1, was common across comparisons (Figure 6C; Tables S7C–S7E). Since IFN-g signaling in endothelial cells has been suggested to play multiple roles in the tumor immune response, but its role in ICB treatment response has not been documented, we examined the role of IFN-g signaling in endothelial cells further.31,32
Functional analysis confirms endothelial cells are a principal target of IFN-g and a key mediator of treatment response
To test the role of IFN-g signaling in endothelial cells in response to ICB treatment, we generated a mouse model system where IFNgR1 could be knocked out specifically in endothelial cells with tamoxifen treatment by crossing CDH5-ERT2-Cre+ mice with IFNgR1 flox/flox (f/f) mice. Using flow cytometry, we confirmed IFNgR1 expression was significantly reduced in CD31+ endothelial cells from mice in the knockout conditions compared to intact mice lacking the Cre-expressing allele (Figures S8A, S9A, and S9B).
After establishing this model system, we compared tumor growth in IFNgR1 intact mice with and without ICB treatment to tumor growth in endothelial IFNgR1 knockout mice with and without ICB treatment (STAR Methods). This comparison revealed that ICB treated knockout mice had tumor growth patterns nearly identical to untreated intact mice, demonstrating that significantly reducing IFNgR1 expression in endothelial cells negated the anti-tumor effects of ICB treatment (Figure 7A; Table S8A). Thus, IFN-g response in endothelial cells is necessary for an effective ICB treatment response. Furthermore, untreated tumors in the knockout mice grew more quickly than untreated tumors in the intact mice (Figure 7A; Table S8A). These findings are analogous to previously reported findings which showed that CD4+ T cell depletion in the MCB6C model not only prevented ICB induced tumor rejection, but also led to increased tumor growth even in the absence of ICB treatment, indicating that a basal level of T cell activity restrains tumor growth.19 Similarly, the findings presented here indicated that basal levels of IFN-g signaling in endothelial cells restrained tumor growth and upregulation of IFN-g activity in endothelial cells was necessary for tumor rejection upon ICB treatment.
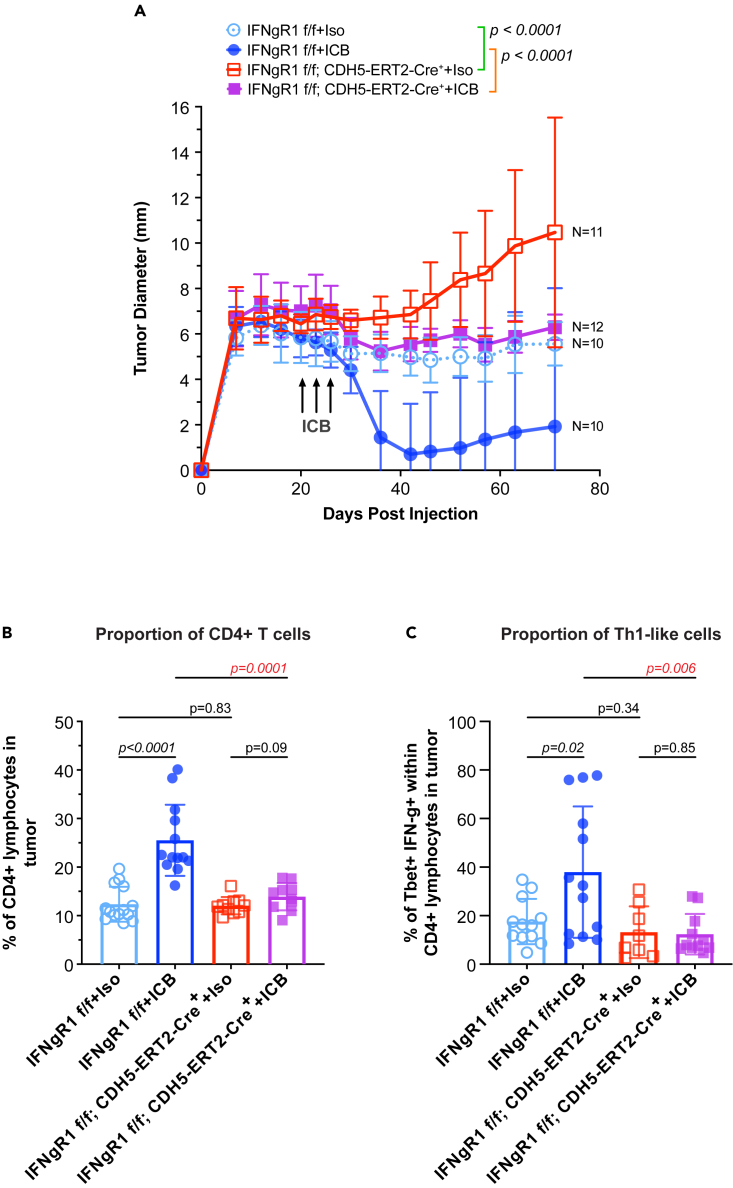
Figure 7.
Functional analysis confirms endothelial cells are a principal target of IFN-g and a key mediator of treatment response
(A) Tumor diameter measurements for IFNgR1 intact and endothelial IFNgR1 knockout mice with and without ICB treatment over time (pre- and post-treatment). For all comparisons, a 2-way ANOVA for repeated measures was performed. P-values <0.05 were considered significant. Error bars represent one standard deviation.
(B) Bar graphs displaying the percentage of CD4+ lymphocytes in the tumor microenvironment across the same four conditions as (A).
(C) Bar graphs displaying the percentage of Tbet+, IFNg+ cells detected within the CD4+ T lymphocyte population across the same four conditions as (A). For all bar graphs, bar height indicates the average percentage across all mice from the given condition. Each point represents the percentage for an individual mouse. For all comparisons, a two-tailed, unpaired Student's t-test was performed. P-values <0.05 were considered significant. Error bars represent one standard deviation. f/f = flox/flox, ICB = combined PD-1/CTLA-4 immune checkpoint blockade treatment, Iso = rat IgG2a and mouse IgG2b isotype control. See also Figures S8–S10 and Table S8.
Flow cytometric analysis indicated that ICB treatment induced recruitment of CD4+ T lymphocytes to the TME in intact mice, consistent with previous work (Figures 7B and S8B).19 However, in mice where IFNgR1 had been knocked out in endothelial cells, this recruitment of CD4+ T lymphocytes after treatment was negated (Figure 7B; Table S8B). Furthermore, recruitment of Tbet+, IFN-g+, CD4+ T lymphocytes (i.e., Th1-like cells) seen after ICB treatment was no longer seen in the knockout condition (Figure 7C; Table S8C). Similar to previously reported analysis of the MCB6C model, this analysis showed no significant change in the proportion of CD8+ T cells seen before or after ICB treatment in either intact or knockout mice (Figure S10).19 These results further indicated that IFN-g signaling in endothelial cells is a key mediator of treatment response and that it underlies recruitment of CD4+ effector T cells in the TME.
Discussion
To explore mechanisms of response to combined PD-1/CTLA-4 ICB treatment of bladder cancer in individual cell types, we generated scRNA-seq from a mouse model of urothelial carcinoma. The three sample conditions used in this study were untreated tumor, combined PD-1/CTLA-4 ICB treated tumor, and tumor that received combined ICB treatment after CD4+ T cell depletion. In total, we performed scRNA-seq on fifteen samples (five per each condition) and captured over 57,000 cells that passed filtering and were aggregated into a single dataset for downstream analysis. Within the aggregated dataset, we identified numerous lymphocyte, myeloid, and stromal cell populations. Clustering and visualization of the data revealed two distinct epithelial clusters, which we confirmed corresponded to tumor cell populations based on expression of somatic variants and, more specifically, appeared to correspond to distinct basal-like and luminal-like subpopulations.
After identifying cell types present within the aggregated dataset, we used differential expression, overrepresentation, and GSEA to explore how individual cell types were responding to treatment. This analysis showed that IFN-g response was commonly perturbed with treatment across multiple cell types, including endothelial cells. Multiple clinical trials exploring human bladder cancer have identified IFN-g pathway activity as being correlated with increased benefit from ICB treatment.12,13,33 Previous work in the MCB6C model established that IFN-g activity is necessary for ICB treatment response.19 While previous research of tumor immunosurveillance models has shown that IFN-g signaling can act through both tumor cell intrinsic and extrinsic mechanisms, the role of IFN-g and its key target cells in ICB treatment response has not been completely defined.31,34,35
While previous work excluded IFN-g activity in tumor cells as having an essential role in treatment response in the MCB6C model, we had not previously evaluated its role in endothelial cells. Here, we establish endothelial cells as a key target of IFN-g activity and further show that loss of IFN-g signaling in endothelial cells impairs recruitment of IFN-g producing CD4+ T cells to the TME. Notably, Cxcl9, Cxcl10, Vcam1, and Icam1, which are mediators of T cell trafficking, were among the most upregulated IFN-g response genes in endothelial cells following ICB treatment, suggesting that a key role of IFN-g activity in endothelial cells may be to enable recruitment of T cells to the TME.
We hypothesize a feedforward model in which ICB treatment induces IFN-g production from CD4+ T cells, which in turn leads to further recruitment of CD4+ T cells to the TME via upregulation of chemoattractant molecules in endothelial cells. However, other roles of IFN-g signaling in endothelial cells could also contribute to treatment response. For example, IFN-g signaling in endothelial cells could induce tumor ischemia or impact vascular permeability, as shown by previous studies.31,32,36 Ultimately, these results showed that IFN-g response in endothelial cells is a key mediator of treatment response and suggested that strategies which selectively induce IFN-g signaling in endothelial cells in the TME could favorably impact response to ICB treatment as well as other T cell based therapies.
Limitations of the study
While these findings support the role of IFN-g signaling in endothelial cells as a key node in treatment response, there are limitations to this analysis. In particular, effective treatment response involves a cascade of events which are still not fully defined. For example, the mechanisms by which T cells in the TME actually kill tumor cells are not elucidated in this system. Likewise, the mechanisms by which endothelial cells recruit T cells to the TME have not been fully explored. Additionally, the immune microenvironment arising from subcutaneous injection of a bladder cancer cell line could differ significantly from the immune microenvironment of a bladder cancer grown in bladder tissue. Ultimately, further analysis will be needed to verify and fully characterize the mechanisms underlying effective ICB treatment response. Nevertheless, these results underscore the power of scRNA-seq analysis to inform hypotheses that, when coupled with mouse modeling, can help identify cell-type specific signaling nodes that are key to generating an effective immune response.
STAR★Methods
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| InVivoMAb anti-mouse PD-1 (CD279) | BioXcell | Cat# BE0146, clone RMP1-14; RRID:AB_10949053 |
| InVivoMAb anti-mouse CTLA-4 (CD152) | BioXcell | Cat# BE0164, clone 9D9; RRID:AB_10949609 |
| InVivoMAb rat IgG2a isotype control, anti-trinitrophenol | BioXcell | Cat# BE0089, clone 2A3; RRID:AB_1107769 |
| InVivoMAb mouse IgG2b isotype control, unknown specificity | BioXcell | Cat# BE0086, clone MPC-11; RRID:AB_1107791 |
| InVivoMAb anti-mouse CD4 | BioXcell | Cat# BE0003-1, clone GK1.5; RRID:AB_1107636 |
| InVivoMAb rat IgG2b isotype control, anti-keyhole limpet hemocyanin | BioXcell | Cat# BE0090, clone LTF-2; RRID:AB_1107780 |
| Brilliant Violet 510™ anti-mouse CD45 Antibody | BioLegend | Cat# 103137, clone 30-F11; RRID:AB_2561392 |
| PE/Dazzle™ 594 anti-mouse CD3ε Antibody | BioLegend | Cat# 100347, clone 145-2C11; RRID:AB_2564028 |
| FITC anti-mouse CD4 Antibody | BioLegend | Cat# 116003, clone RM4-4; RRID:AB_313688 |
| PerCP/Cyanine5.5 anti-mouse CD4 Antibody | BioLegend | Cat# 116011, clone RM4-4; RRID:AB_2563022 |
| PE/Cyanine7 anti-mouse CD8α Antibody | BioLegend | Cat# 100721, clone 53-6.7; RRID:AB_312760 |
| Alexa Fluor® 700 anti-mouse CD8α Antibody | BioLegend | Cat# 100729, clone 53-6.7; RRID:AB_493702 |
| Brilliant Violet 421™ anti-mouse CD19 Antibody | BioLegend | Cat# 115537, clone 6D5; RRID:AB_10895761 |
| APC anti-mouse/human CD11b Antibody | BioLegend | Cat# 101211, clone M1/70; RRID:AB_312794 |
| PE/Cyanine7 anti-mouse CD11c Antibody | BioLegend | Cat# 117317, clone N418; RRID:AB_493569 |
| Alexa Fluor® 647 anti-mouse CD326 (Ep-CAM) Antibody | BioLegend | Cat# 118211, clone G8.8; RRID:AB_1134104 |
| PerCP/Cyanine5.5 anti-mouse CD326 (Ep-CAM) Antibody | BioLegend | Cat# 118219, clone G8.8; RRID:AB_2098647 |
| Brilliant Violet 421™ anti-mouse/human CD44 Antibody | BioLegend | Cat# 103039, clone IM7; RRID:AB_10895752 |
| PE anti-mouse CD62L Antibody | BioLegend | Cat# 104407, clone MEL-14; RRID:AB_313094 |
| PE/Cyanine7 anti-mouse Ly-6C Antibody | BioLegend | Cat# 128015, clone HK1.4; RRID:AB_1732087 |
| PerCP/Cyanine5.5 anti-mouse Ly-6G Antibody | BioLegend | Cat# 127615, clone 1A8; RRID:AB_1877272 |
| PE anti-mouse Siglec-F Antibody | BD Biosciences | Cat# 552126, clone E50-2440; RRID:AB_394341 |
| APC/Cyanine7 anti-mouse CD335 (NKp46) Antibody | BioLegend | Cat# 137645, clone 29A1.4; RRID:AB_2876479 |
| Alexa Fluor™ 700 anti-Foxp3 Antibody | eBioscience | Cat# 56-5773-80, clone FJK-16s; RRID:AB_469950 |
| PE/Dazzle™ 594 anti-T-bet Antibody | BioLegend | Cat# 644827, clone 4B10; RID:AB_2565676 |
| APC/Cyanine7 anti-mouse IFN-γ Antibody | BioLegend | Cat# 505849, clone XMG1.2; RRID:AB_2616697 |
| Alexa Fluor® 647 anti-Ki-67 Antibody | BD Biosciences | Cat# 561126 clone B56; RRID:AB_10611874 |
| Biotin anti-mouse CD119 (IFN-γ R α chain) Antibody | BioLegend | Cat# 112803, clone 2E2; RRID:AB_2123476 |
| PE Streptavidin | BioLegend | Cat# 405203 |
| Purified anti-mouse CD16/32 Antibody | BioLegend | Cat# 101301, clone 93; RRID:AB_312800 |
| Critical commercial assays | ||
| Automated Kapa HYPER PCR-free Kit | Roche | Cat# 7962371001 – KK8505 |
| SureSelect DNA - Mouse All Exon V1 | Agilent | N/A |
| Chromium Next GEM Single Cell 5' Kit v2 | 10x Genomics | Cat# PN-1000263 |
| Chromium Single Cell V(D)J Enrichment Kit, Mouse B Cell | 10x Genomics | Cat# PN-1000072 |
| Chromium Single Cell V(D)J Enrichment Kit, Mouse T Cell | 10x Genomics | Cat# PN-1000071 |
| Deposited data | ||
| WGS, WES, scRNA-seq data | This paper | BioProject: PRJNA934380 |
| Experimental models: Cell lines | ||
| MCB6C | Vivek Arora Lab | N/A |
| Experimental models: Organisms/strains | ||
| C57BL/6 mice | Taconic Biosciences | B6NTac |
| CDH5-ERT2-Cre+/IFNgR1flox/flox mice | Vivek Arora Lab | N/A |
| Software and algorithms | ||
| CellRanger (v5.0) | 10x Genomics | https://support.10xgenomics.com/single-cell-gene-expression/software/pipelines/5.0/what-is-cell-ranger |
| CNVkit (v0.9.8) | Talevich et al.37 | https://cnvkit.readthedocs.io/en/v0.9.8/ |
| Seurat (v4.0.1) | Hao et al.38 | https://satijalab.org/seurat/ |
| SingleR (v1.4.1) | Aran et al.24 | https://github.com/dviraran/SingleR |
| CytoTRACE (v0.3.3) | Gulati et al.27 | https://cytotrace.stanford.edu/ |
| clusterProfiler (v3.18.1) | Yu et al.39 | https://bioconductor.org/packages/release/bioc/html/clusterProfiler.html |
| VarTrix (v1.1.3) | 10x Genomics | https://github.com/10XGenomics/vartrix |
| Prism (v8.3.0) | GraphPad | https://www.graphpad.com/ |
Resource availability
Lead contact
Further information and requests for resources should be directed to and will be fulfilled by the lead contact, Malachi Griffith (mgriffit@wustl.edu).
Materials availability
This study did not generate any new materials.
Experimental model and study participant details
Mice used for MCB6C experiments
All animal experiments were carried out according to the guidelines of the American Association for Laboratory Animal Science under a protocol approved by the Institutional Animal Care and Use Committee at Washington University and performed in Association for Assessment and Accreditation of Laboratory Animal Care International (AAALAC)-accredited specific pathogen-free facilities at Washington University School of Medicine in St. Louis. Forty-five 5- to 6-week-old Black 6 (B6NTac) male mice were purchased from Taconic Biosciences and were allowed to acclimate for a week before in vivo experiments were performed. The maximal tumor size/burden permitted by our institutional review board is 15% of body weight (combined burden if more than one mass present) and mean tumor diameter = or >20 mm in adult mice (∼25 g). The maximal tumor size/burden permitted by our institutional review board was not exceeded.
CDH5-ERT2-Cre+, IFNgR1 flox/flox (f/f) mice
C57BL/6-Tg(Cdh5-cre/ERT2)1Rha mice were originally generated by Dr. Ralf H. Adams and purchased from Taconic Biosciences then bred with C57BL/6N-Ifngr1tm1.1Rds/J (IFNgR1flox/flox) mice that were obtained from Dr. Robert Schreiber at Washington University School of Medicine to generate CDH5-ERT2-Cre+/IFNgR1flox/flox offspring.
Method details
Bulk DNA sequencing, alignment, and analysis
Whole genome sequencing (WGS) libraries were constructed from genomic DNA isolated from an MCB6C cell line sample and a black 6 (B6NTac) matched normal tail sample using Automated Kapa HYPER PCR free preparation kits (catalog #7962371001 – KK8505) and sequenced on the Illumina NovaSeq 6000 platform. WGS reads were aligned to the GRCm38 reference genome using BWA-MEM. Copy number variant calling was performed using the CNVkit (v0.9.8) batch pipeline with a target bin size of 100,000 bp.37 Whole exome sequencing (WES) libraries were constructed and sequenced similarly to the WGS experiment following hybrid capture selection with the hybrid reagent SureSelect DNA - Mouse All Exon V1 (Agilent). WES reads were aligned to the GRCm38 reference genome using BWA-MEM. Somatic variant calling was performed using common workflow language pipelines provided by the McDonnell Genome Institute (https://github.com/genome/analysis-workflows). Somatic variants were called with Pindel, VarScan, Mutect, and Strelka and combined.40,41,42,43,44 Variants were then filtered based on the criteria of being called by at least two variant callers, normal coverage > 30X, tumor coverage > 30X, normal VAF < 5%, and tumor VAF > 5%. To explore clonality based on VAF distribution, variants were filtered more stringently based on the criteria of being called by at least two variant callers, having normal and tumor coverage >= 100X, normal VAF = 0%, tumor VAF > 5%, and no overlap with any regions of copy number alteration (i.e. any regions with abs(log2 copy ratio) > 0.32). Additionally, variants with VAF < 30% were manually reviewed to remove possible false variants caused by artifacts such as sequencing errors or misalignments.
Identifying possible driver mutations in WES
After filtering somatic variants, a subset of possible driver mutation positions was determined by further filtering the set of somatic mutations down to mutations found in genes that have been previously reported to harbor driver mutations in human bladder cancer (https://www.intogen.org/search?cancer=BLCA). Human gene names were converted to homologous mouse gene names using the Mouse Genome Informatics human and mouse homology report with mammalian phenotype IDs (https://www.informatics.jax.org/homology.shtml). Each mutation was manually reviewed against mutations reported in ProteinPaint (https://proteinpaint.stjude.org/), IntOGen, and Cancer Hotspots (https://www.cancerhotspots.org/#/home).
Mouse bladder organoid culture for injection
One previously archived frozen vial of singly suspended MCB6C organoid was thawed at least 2 weeks before mouse injection and expanded weekly in culture at least 2 times. For MCB6C organoid culture expansion, growth factor reduced Matrigel was thawed on ice for minimally 1.5 hours. Pelleted MCB6C cells were washed and resuspended in 1 ml of Advanced DMEM/F12+++ medium (Advanced DMEM/F12 medium [Gibco, catalog #12634010] supplemented with 1% penicillin/streptomycin, 1% HEPEs, and Glutamax) and cell concentration was determined by automated cell counter. To establish organoid culture, 50 μl Matrigel tabs with 10,000 cells/tab were generated and plated on 6-well suspension culture plates, 6 tabs wells. Tabs were incubated at 37°C for 15 min until Matrigel was hardened, returned to tissue culture incubator, and cultured with mouse bladder organoid medium (MBO medium - Advanced DMEM/F12+++ medium supplemented with EGF, A-83-01, Noggin, R-Spondin, N-Acetly-L-cysteine, and Nicotinamide). Organoids were replenished with fresh MBO medium every 3–4 days and also one day before mouse injection.
Mouse injection with MCB6C organoid cells
A single cell suspension of MCB6C organoid was generated by TrypLE Express (Gibco, catalog #12605010) digestion organoid Matrigel tabs at 37°C for 15 min. After digestion, pelleted cells were washed and resuspended in PBS to determine cell concentration. After cell concentration was adjusted to 20 million/ml in PBS, organoid cells were mixed with growth factor reduced Matrigel at 1:1 ratio before being injected subcutaneously into the left flank of the mouse (1 million/100 μl cells each mouse). Tumor development was monitored using digital calipers to assess the length, width, and depth of each tumor. For ICB, each mouse was injected intraperitoneally with 250 μg anti-PD1 (BioXcell, catalog #BE0146, clone #RMP1-14) and 200 μg anti-CTLA-4 (BioXcell, catalog #BE0164, clone 9D9) day 9 and 12 after organoid implantation. For isotype controls, each mouse was injected with 250 μg rat IgG2a (BioXcell, catalog #BE0089, clone 2A3) and 200 μg IgG2b (BioXcell, catalog #BE0086, clone #MPC-11). For CD4+ T cell depletion, each mouse was injected with 250 μg anti-CD4 (BioXcell, catalog #BE0003-1, clone #GK1.5) day 0 and 7 after organoid depletion. Rat IgG2b (BioXcell, catalog #BE0090, clone #LTF-2) was used as an isotype control for anti-CD4.
Harvesting tumors for scRNA-seq, BCR-seq, and TCR-seq
Based on 10x Genomics Demonstrated Protocols, 14 days after organoid implantation, tumors were dissected from euthanized mice, cut into small pieces of ∼2–4 mm3, and further processed into dead-cell depleted single cell suspension following manufacturer’s protocol using Tumor Dissociation Kit and MACS Dead Cell removal Kit (Miltenyi Biotec). Briefly, tumor tissue pieces were transferred to gentleMACS C tube containing enzyme mix before loading onto a gentleMACS Octo Dissociator with Heaters for tissue digestion at 37°C for 80 min. After tissue dissociation was completed, cell suspension was transferred to a new 50 ml conical tube, and supernatant was removed after centrifugation. Cell pellet was resuspended in RPMI 1640 medium, filtered through a prewetted 70 μM cell filter, strained, pelleted, and resuspended in red cell lysis buffer and incubated on ice for 10 min. After adding the wash buffer, the cell suspension was pelted and resuspended in the wash buffer. To remove dead cells, Dead Cell Removal Microbeads were added to resuspend cell pellet (100 μl beads per 107 cells) using a wide-bore pipette tip. After incubation for 15 min at room temperature, the cell-microbead mixture was applied onto a MS column. Dead cells remained in the column and the effluent represented to the live cell fraction. The percentage of viable cells was determined by an automated cell counter. Dead cell removal was repeated if the percentage of viable cells did not reach above 90%. Two rounds of centrifugation/resuspension were carefully performed for two rounds in 1xPBS/0.04% BSA using a wide-bore tip. To submit cell samples for single-cell RNA-seq analysis, cell concentration was determined accurately by sampling cell suspension twice and counting each sampling twice and adjusted to 1167 cells/μl. 40 μl of each cell suspension was submitted to the Genome Technology Access Center/McDonnell Genome Institute (GTAC/MGI) for single-cell RNA-seq analysis using the 5’v2 library kit (10x Genomics catalog #PN-1000263) with BCR and TCR V(D)J enrichment kits (10x Genomics catalog #PN-1000072 and #PN-1000071, respectively). cDNA generation and TCR/BCR enrichment were performed according to the Chromium Single Cell V(D)J Reagent Kits User Guide (CG000086 Rev L). The libraries were sequenced on the S4 300 cycle kit flow (2x151 paired end reads) using the XP workflow as outlined by Illumina. FASTQ outputs were generated.
Alignment, filtering, and clustering of scRNA-seq
Alignment and gene expression quantification were performed with CellRanger count (v5.0, default parameters). Gene-barcode matrices were then imported into Seurat for filtering cells, QC, clustering, etc.38 To filter suspected dying cells, cells were clustered before filtering to identify cells appearing to cluster based on high mitochondrial gene expression (i.e. the percentage of UMIs per cell mapping to mitochondrial genes). The cutoff for mitochondrial gene expression was based on the percentage that captured the majority of these cells. A cutoff of 12.5% was used across all replicates. Doublets were filtered based on high UMI expression and CellRanger’s reported doublet rate (0.9% per 1000 cells), with the top 0.9% of cells removed from each condition in each replicate. Cutoffs for filtering cells with low feature detection were determined by assigning cell types to each cell using the CellMatch method, identifying cells that did not have enough features for their cell type to be predicted, and calculating the average number of features detected in those “non-predicted” cells. CellMatch infers cell types by training a nearest-neighbors algorithm on published expression data. Spearman correlation is used as the distance metric. CellMatch’s cell typing inference is unsupervised and infers cell types in a marker free manner.45 After filtered cells were removed, gene expression values for each gene in the remaining cells were normalized and scaled and variable genes were selected using Seurat with default settings. Principal component (PC) analysis was then performed using the top 2,000 variable genes and npcs = 20. Clustering of cells was performed using 20 PCs and resolution = 0.7. Finally, dimensionality reduction and visualization were performed using Seurat’s tSNE function. B cell and T cell receptors were assembled and identified using the 10x Genomics CellRanger V(D)J pipeline (v5.0, default parameters).
Assigning cell types using SingleR
Cell types for each cell were annotated with SingleR using expression profiles from the ImmGen dataset (https://www.immgen.org/).24,25 Cell types were manually simplified to B cell (B), CD4+ T cell (CD4), naive CD4+ T cell (CD4.Naive), naive CD8+ T cell (CD8.Naive), CD8+ effector T cell (CD8.Eff), CD8+ memory T cell (CD8.Mem), dendritic cell, endothelial cell, epithelial cell, fibroblast, macrophage, monocyte, neutrophil, natural killer cell (NK), natural killer T cell (NKT), gamma delta T cell (Tgd), and regulatory T cell (Treg).
Analysis of tumor cell subtypes
To explore tumor cell subpopulations, we labeled tumor cells as either T1 or T2 based on cluster number. Cells that were assigned to the tumor clusters (i.e. clusters 10 and 14) on the left side of the tSNE projection were labeled as T1 tumor cells and cells assigned to the tumor cluster (i.e cluster 7) on the right side were labeled as T2 tumor cells. After assigning these labels, we separated the T1 and T2 cell populations from all other cell populations. We then scaled and normalized gene expression and selected variable genes using Seurat’s default methods. Principal component (PC) analysis was then performed using the top 2,000 variable genes and npcs = 20. Clustering of cells was performed using 20 PCs and resolution = 0.7. Finally, dimensionality reduction and visualization were performed using Seurat’s tSNE function. We then assigned differentiation scores to each cell using CytoTRACE (https://cytotrace.stanford.edu/) and calculated the differentiation score as 1 - the CytoTRACE score.27
DE, overrepresentation, and GSEA in scRNA-seq
All differential expression analyses were performed using Seurat’s FindMarkers function with the Wilcoxon Rank Sum method. P-values were adjusted using Benjamini-Hochberg multiple test correction. All reference gene sets used for overrepresentation analysis and gene set enrichment analysis (GSEA) were from MSigDB with the exception of the basal and luminal marker gene sets, which were generated ad-hoc from published lists of basal and luminal bladder cancer markers.46,47 For all overrepresentation analysis, results were generated using the enricher function from the clusterProfiler package in R.39 For comparisons of conditions within each cell type, input gene lists for overrepresentation analysis were generated by taking all genes with adjusted p-value < 0.05 and fold change value > ∼1.2 (i.e. abs(log2FC) > 0.26). GSEA results were generated using UC San Diego and Broad Institute’s GSEA software to run GSEAPreranked with genes ranked by the average log2 fold changes reported by Seurat.
Deletion of IFNgR1 in endothelium by tamoxifen
Tamoxifen (Alfa Aesar, catalog #J63509) was dissolved in corn oil (MilliporeSigma, catalog #C8267) at the concentration of 20 mg/ml in a 37°C shaker overnight one day before the treatment began and kept at 4°C during the 5-day treatment.
Subcutaneous engraftment of MCB6C organoids
Tumor experiments were performed following methods established previously with modifications.19 To improve the engraftment and growth of the organoid cells on mice, Matrigel with high protein concentration (Corning, catalog #354262) was used instead of growth factor reduced Matrigel (Corning, catalog #356231). After organoids were expanded in culture for > 2 weeks and subsequently harvested by TrypLE Express (Gibco, catalog #12605010) treatment, organoid cells were resuspended in 3:1 PBS/high protein concentration Matrigel (instead of 1:1 PBS/growth factor reduced Matrigel) at 10 million cells/ml. 1 million/100 μl of cell/Matrigel mix was subcutaneously injected into the left flank of the mouse, which was performed one week after the completion of Tamoxifen treatment. Tumor growth was monitored twice a week using digital calipers. The mean of long and short diameters was used for tumor growth curves. For ICB treatment, mice were injected with 250 μg/mouse αPD-1 (BioXcell, catalog #BE0146, clone RMP1-14) and 200 μg/mouse αCTLA-4 (BioXcell, catalog #BE0164, clone 9D9) i.p. every 3 days from day 15 to 18 after organoid implantation for short term studies, and from day 15 through day 21 from long term studies. 250 μg/mouse rat IgG2a (BioXcell, catalog #BE0089, clone 2A3) and 200 μg/mouse IgG2b (BioXcell, catalog #BE0086, clone MPC-11) were used as isotype controls.
Antibodies
The following antibodies were used for flow cytometry: Brilliant Violet 510™ anti-mouse CD45 Antibody (BioLegend, catalog #103137, clone 30-F11), PE/Dazzle™ 594 anti-mouse CD3ε Antibody (BioLegend, catalog #100347, clone 145-2C11), FITC anti-mouse CD4 Antibody (BioLegend, catalog #116003, clone RM4-4), PerCP/Cyanine5.5 anti-mouse CD4 Antibody (BioLegend, catalog #116011, clone RM4-4), PE/Cyanine7 anti-mouse CD8α Antibody (BioLegend, catalog #100721, clone 53-6.7), Alexa Fluor® 700 anti-mouse CD8α Antibody (BioLegend, catalog #100729, clone 53-6.7), Brilliant Violet 421™ anti-mouse CD19 Antibody (BioLegend, catalog #115537, clone 6D5), APC anti-mouse/human CD11b Antibody (BioLegend, catalog #101211, clone M1/70), PE/Cyanine7 anti-mouse CD11c Antibody (BioLegend, catalog #117317, clone N418), Alexa Fluor® 647 anti-mouse CD326 (Ep-CAM) Antibody (BioLegend, catalog #118211, clone G8.8), PerCP/Cyanine5.5 anti-mouse CD326 (Ep-CAM) Antibody (BioLegend, catalog #118219, clone G8.8), Brilliant Violet 421™ anti-mouse/human CD44 Antibody (BioLegend, catalog #103039, clone IM7), PE anti-mouse CD62L Antibody (BioLegend, catalog #104407, clone MEL-14), PE/Cyanine7 anti-mouse Ly-6C Antibody (BioLegend, catalog #128015, clone HK1.4), PerCP/Cyanine5.5 anti-mouse Ly-6G Antibody (BioLegend, catalog #127615, clone 1A8), PE anti-mouse Siglec-F Antibody (BD Biosciences, catalog #552126, clone E50-2440), APC/Cyanine7 anti-mouse CD335 (NKp46) Antibody (BioLegend, catalog #137645, clone 29A1.4), Alexa Fluor™ 700 anti-Foxp3 Antibody (eBioscience, catalog #56-5773-80, clone FJK-16s), PE/Dazzle™ 594 anti-T-bet Antibody (BioLegend, catalog #644827, clone 4B10), APC/Cyanine7 anti-mouse IFN-γ Antibody (BioLegend, catalog #505849, clone XMG1.2), Alexa Fluor® 647 anti-Ki-67 Antibody (BD Biosciences, catalog #561126 clone B56), Biotin anti-mouse CD119 (IFN-γ R α chain) Antibody (BioLegend, catalog #112803, clone 2E2), PE Streptavidin (BioLegend, catalog #405203).
Flow cytometry
To determine the cellular composition of the tumor, tumors were isolated, minced into small pieces, and digested for 1 hour in DMEM media (MilliporeSigma, catalog #D5796) containing 100 μg/ml Collagenase type IA (Gibco, catalog #17101015), 100 μg/ml Dispase II (MilliporeSigma, catalog #D4693) and 50 U/ml of DNase I (Worthington Biochemical, catalog #LS002006). Cells were washed in ice-cold PBS with 3% FCS and 2 mM EDTA (FACS buffer) and filtered over 70-μm nylon mesh. After red blood cell lysis with ACK solution (Gibco, catalog #A1049201), cells were stained with a Zombie NIR Fixable Viability kit (BioLegend, catalog #423105) for dead cell exclusion followed by Fc-receptor blocking with purified mouse CD16/32 antibody (BioLegend, catalog #101301, clone 93). After cell surface marker staining with fluorescent-conjugated antibodies, cells were fixed and permeabilized using a Foxp3/transcription factor staining kit (eBioscience, catalog #00-5523-00) and intracellularly stained with fluorescent-conjugated antibodies. Flow cytometric data were acquired by Cytek-upgraded 10-color FACScan cytometers at Washington University Siteman Cancer Center Cell Sorting Core facility and analyzed by FlowJo 10 (TreeStar).
Quantification and statistical analysis
Statistical analyses for IFNgR1 knockout experiments were performed using Prism 8.3.0 (GraphPad). For all tumor growth curve comparisons, a 2-way ANOVA for repeated measures was used. For all other comparisons, an unpaired Student’s t test was used. All tests were 2-tailed. P-values of less than 0.05 were considered significant.
Acknowledgments
Malachi Griffith was supported by the National Human Genome Research Institute (NHGRI) of the National Institutes of Health (NIH) under Award Number R00HG007940. Malachi Griffith and Obi Griffith were supported by the NIH National Cancer Institute (NCI) under Award Numbers U01CA209936, U01CA231844, U24CA237719, and U01CA248235. Malachi Griffith was supported by the V Foundation for Cancer Research under Award Number V2018-007. Vivek Arora was supported by the Department of Defense Career Development Award W81XWH-17-1-0562, the Rabushka Bladder Cancer Research Fund, and the Clinical Investigator Award from the Damon Runyon Foundation. We thank Dr. Robert Schreiber at Washington University School of Medicine for gifting us C57BL/6N-Ifngr1tm1.1Rds/J (IFNgR1flox/flox) mice. We thank Dr. Kian H. Lim for providing infrastructure support for mouse experiments. We thank Joshua F. McMichael for designing the graphical abstract.
Author contributions
Conceptualization V.K.A., M.G., and O.L.G.; Methodology T.H.P.C. and J.K.B.; Formal Analysis S.L.F., B.F., H.S., M.M., and Z.L.S.; Investigation T.H.P.C.; Writing - Original Draft S.L.F.; Writing - Review and Editing V.K.A., M.G., O.L.G., T.H.P.C., and S.L.F.; Supervision V.K.A., M.G., and O.L.G.; Funding Acquisition V.K.A., M.G., and O.L.G.
Declaration of interests
V.K.A. currently serves as an employee of Bristol Myers Squibb and has stock options in the company. J.K.B. currently serves as an employee of Pfizer Inc.
Inclusion and diversity
We support inclusive, diverse, and equitable conduct of research.
Published: September 15, 2023
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.isci.2023.107937.
Contributor Information
Obi L. Griffith, Email: obigriffith@wustl.edu.
Vivek K. Arora, Email: arorav@wustl.edu.
Malachi Griffith, Email: mgriffit@wustl.edu.
Supplemental information
Data and code availability
-
•
Raw whole genome, exome, and single-cell RNA sequencing data have been deposited in the Sequence Read Archive (SRA) and are publicly available as of the date of publication. Accession numbers are listed in the key resources table.
-
•
This paper does not report original code.
-
•
Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
References
- 1.Bladder cancer statistics WCRF International. 2022. https://www.wcrf.org/cancer-trends/bladder-cancer-statistics/
- 2.Compérat E., Amin M.B., Cathomas R., Choudhury A., De Santis M., Kamat A., Stenzl A., Thoeny H.C., Witjes J.A. Current best practice for bladder cancer: a narrative review of diagnostics and treatments. Lancet. 2022;400:1712–1721. doi: 10.1016/S0140-6736(22)01188-6. [DOI] [PubMed] [Google Scholar]
- 3.Robertson A.G., Kim J., Al-Ahmadie H., Bellmunt J., Guo G., Cherniack A.D., Hinoue T., Laird P.W., Hoadley K.A., Akbani R., et al. Comprehensive Molecular Characterization of Muscle-Invasive Bladder Cancer. Cell. 2017;171:540–556.e25. doi: 10.1016/j.cell.2017.09.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Survival Rates for Bladder Cancer. https://www.cancer.org/cancer/bladder-cancer/detection-diagnosis-staging/survival-rates.html
- 5.Lenis A.T., Lec P.M., Chamie K. Bladder Cancer: A Review. JAMA. 2020;324:1980–1991. doi: 10.1001/jama.2020.17598. [DOI] [PubMed] [Google Scholar]
- 6.Muscle-invasive and Metastatic Bladder Cancer Uroweb - European Association of Urology. https://uroweb.org/guidelines/muscle-invasive-and-metastatic-bladder-cancer/chapter/disease-management
- 7.Rhea L.P., Aragon-Ching J.B. Advances and Controversies With Checkpoint Inhibitors in Bladder Cancer. Clin. Med. Insights Oncol. 2021;15 doi: 10.1177/11795549211044963. 11795549211044963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Lopez-Beltran A., Cimadamore A., Blanca A., Massari F., Vau N., Scarpelli M., Cheng L., Montironi R. Immune Checkpoint Inhibitors for the Treatment of Bladder Cancer. Cancers. 2021;13:131. doi: 10.3390/cancers13010131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Suzman D.L., Agrawal S., Ning Y.-M., Maher V.E., Fernandes L.L., Karuri S., Tang S., Sridhara R., Schroeder J., Goldberg K.B., et al. FDA Approval Summary: Atezolizumab or Pembrolizumab for the Treatment of Patients with Advanced Urothelial Carcinoma Ineligible for Cisplatin-Containing Chemotherapy. Oncol. 2019;24:563–569. doi: 10.1634/theoncologist.2018-0084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Albisinni S., Martinez Chanza N., Aoun F., Diamand R., Mjaess G., Azzo J.-M., Esperto F., Bellmunt J., Roumeguère T., DE Nunzio C. Immune checkpoint inhibitors for BCG-resistant NMIBC: the dawn of a new era. Minerva Urol. Nephrol. 2021;73:292–298. doi: 10.23736/S2724-6051.21.04309-5. [DOI] [PubMed] [Google Scholar]
- 11.Kartolo A., Kassouf W., Vera-Badillo F.E. Adjuvant Immune Checkpoint Inhibition in Muscle-invasive Bladder Cancer: Is It Ready for Prime Time? Eur. Urol. 2021;80:679–681. doi: 10.1016/j.eururo.2021.07.019. [DOI] [PubMed] [Google Scholar]
- 12.Sharma P., Retz M., Siefker-Radtke A., Baron A., Necchi A., Bedke J., Plimack E.R., Vaena D., Grimm M.-O., Bracarda S., et al. Nivolumab in metastatic urothelial carcinoma after platinum therapy (CheckMate 275): a multicentre, single-arm, phase 2 trial. Lancet Oncol. 2017;18:312–322. doi: 10.1016/S1470-2045(17)30065-7. [DOI] [PubMed] [Google Scholar]
- 13.Rosenberg J.E., Hoffman-Censits J., Powles T., van der Heijden M.S., Balar A.V., Necchi A., Dawson N., O’Donnell P.H., Balmanoukian A., Loriot Y., et al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet. 2016;387:1909–1920. doi: 10.1016/S0140-6736(16)00561-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kohli K., Pillarisetty V.G., Kim T.S. Key chemokines direct migration of immune cells in solid tumors. Cancer Gene Ther. 2022;29:10–21. doi: 10.1038/s41417-021-00303-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gao J., Navai N., Alhalabi O., Siefker-Radtke A., Campbell M.T., Tidwell R.S., Guo C.C., Kamat A.M., Matin S.F., Araujo J.C., et al. Neoadjuvant PD-L1 plus CTLA-4 blockade in patients with cisplatin-ineligible operable high-risk urothelial carcinoma. Nat. Med. 2020;26:1845–1851. doi: 10.1038/s41591-020-1086-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Sharma P., Bono P., Kim J.W., Spiliopoulou P., Calvo E., Pillai R.N., Ott P.A., De Braud F.G., Morse M.A., Le D.T., et al. Efficacy and safety of nivolumab monotherapy in metastatic urothelial cancer (mUC): Results from the phase I/II CheckMate 032 study. J. Clin. Orthod. 2016;34:4501. doi: 10.1200/JCO.2016.34.15_suppl.4501. [DOI] [Google Scholar]
- 17.van Dijk N., Gil-Jimenez A., Silina K., Hendricksen K., Smit L.A., de Feijter J.M., van Montfoort M.L., van Rooijen C., Peters D., Broeks A., et al. Preoperative ipilimumab plus nivolumab in locoregionally advanced urothelial cancer: the NABUCCO trial. Nat. Med. 2020;26:1839–1844. doi: 10.1038/s41591-020-1085-z. [DOI] [PubMed] [Google Scholar]
- 18.Roviello G., Catalano M., Santi R., Palmieri V.E., Vannini G., Galli I.C., Buttitta E., Villari D., Rossi V., Nesi G. Immune Checkpoint Inhibitors in Urothelial Bladder Cancer: State of the Art and Future Perspectives. Cancers. 2021;13:4411. doi: 10.3390/cancers13174411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Sato Y., Bolzenius J.K., Eteleeb A.M., Su X., Maher C.A., Sehn J.K., Arora V.K. CD4+ T cells induce rejection of urothelial tumors after immune checkpoint blockade. JCI Insight. 2018;3:e121062. doi: 10.1172/jci.insight.121062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Oh D.Y., Kwek S.S., Raju S.S., Li T., McCarthy E., Chow E., Aran D., Ilano A., Pai C.-C.S., Rancan C., et al. Intratumoral CD4+ T Cells Mediate Anti-tumor Cytotoxicity in Human Bladder Cancer. Cell. 2020;181:1612–1625.e13. doi: 10.1016/j.cell.2020.05.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Zhang X., Lv D., Zhang Y., Liu Q., Li Z. Clonal evolution of acute myeloid leukemia highlighted by latest genome sequencing studies. Oncotarget. 2016;7:58586–58594. doi: 10.18632/oncotarget.10850. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Martínez-Jiménez F., Muiños F., Sentís I., Deu-Pons J., Reyes-Salazar I., Arnedo-Pac C., Mularoni L., Pich O., Bonet J., Kranas H., et al. A compendium of mutational cancer driver genes. Nat. Rev. Cancer. 2020;20:555–572. doi: 10.1038/s41568-020-0290-x. [DOI] [PubMed] [Google Scholar]
- 23.Roviello G., Catalano M., Nobili S., Santi R., Mini E., Nesi G. Focus on Biochemical and Clinical Predictors of Response to Immune Checkpoint Inhibitors in Metastatic Urothelial Carcinoma: Where Do We Stand? Int. J. Mol. Sci. 2020;21:7935. doi: 10.3390/ijms21217935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Aran D., Looney A.P., Liu L., Wu E., Fong V., Hsu A., Chak S., Naikawadi R.P., Wolters P.J., Abate A.R., et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 2019;20:163–172. doi: 10.1038/s41590-018-0276-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Heng T.S.P., Painter M.W., Immunological Genome Project Consortium The Immunological Genome Project: networks of gene expression in immune cells. Nat. Immunol. 2008;9:1091–1094. doi: 10.1038/ni1008-1091. [DOI] [PubMed] [Google Scholar]
- 26.Shay T., Kang J. Immunological Genome Project and systems immunology. Trends Immunol. 2013;34:602–609. doi: 10.1016/j.it.2013.03.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Gulati G.S., Sikandar S.S., Wesche D.J., Manjunath A., Bharadwaj A., Berger M.J., Ilagan F., Kuo A.H., Hsieh R.W., Cai S., et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science. 2020;367:405–411. doi: 10.1126/science.aax0249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Guo C.C., Bondaruk J., Yao H., Wang Z., Zhang L., Lee S., Lee J.-G., Cogdell D., Zhang M., Yang G., et al. Assessment of Luminal and Basal Phenotypes in Bladder Cancer. Sci. Rep. 2020;10:9743. doi: 10.1038/s41598-020-66747-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Choi W., Czerniak B., Ochoa A., Su X., Siefker-Radtke A., Dinney C., McConkey D.J. Intrinsic basal and luminal subtypes of muscle-invasive bladder cancer. Nat. Rev. Urol. 2014;11:400–410. doi: 10.1038/nrurol.2014.129. [DOI] [PubMed] [Google Scholar]
- 30.Dadhania V., Zhang M., Zhang L., Bondaruk J., Majewski T., Siefker-Radtke A., Guo C.C., Dinney C., Cogdell D.E., Zhang S., et al. Meta-Analysis of the Luminal and Basal Subtypes of Bladder Cancer and the Identification of Signature Immunohistochemical Markers for Clinical Use. EBioMedicine. 2016;12:105–117. doi: 10.1016/j.ebiom.2016.08.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kammertoens T., Friese C., Arina A., Idel C., Briesemeister D., Rothe M., Ivanov A., Szymborska A., Patone G., Kunz S., et al. Tumour ischaemia by interferon-γ resembles physiological blood vessel regression. Nature. 2017;545:98–102. doi: 10.1038/nature22311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Ni L., Lu J. Interferon gamma in cancer immunotherapy. Cancer Med. 2018;7:4509–4516. doi: 10.1002/cam4.1700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Sakatani T., Kita Y., Fujimoto M., Sano T., Hamada A., Nakamura K., Takada H., Goto T., Sawada A., Akamatsu S., Kobayashi T. IFN-Gamma Expression in the Tumor Microenvironment and CD8-Positive Tumor-Infiltrating Lymphocytes as Prognostic Markers in Urothelial Cancer Patients Receiving Pembrolizumab. Cancers. 2022;14:263. doi: 10.3390/cancers14020263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Kaplan D.H., Shankaran V., Dighe A.S., Stockert E., Aguet M., Old L.J., Schreiber R.D. Demonstration of an interferon γ-dependent tumor surveillance system in immunocompetent mice. Proc. Natl. Acad. Sci. USA. 1998;95:7556–7561. doi: 10.1073/pnas.95.13.7556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Alspach E., Lussier D.M., Schreiber R.D. Interferon γ and Its Important Roles in Promoting and Inhibiting Spontaneous and Therapeutic Cancer Immunity. Cold Spring Harb. Perspect. Biol. 2019;11:a028480. doi: 10.1101/cshperspect.a028480. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Chrobak I., Lenna S., Stawski L., Trojanowska M. Interferon-γ promotes vascular remodeling in human microvascular endothelial cells by upregulating endothelin (ET)-1 and transforming growth factor (TGF) β2. J. Cell. Physiol. 2013;228:1774–1783. doi: 10.1002/jcp.24337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Talevich E., Shain A.H., Botton T., Bastian B.C. CNVkit: Genome-Wide Copy Number Detection and Visualization from Targeted DNA Sequencing. PLoS Comput. Biol. 2016;12:e1004873. doi: 10.1371/journal.pcbi.1004873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Hao Y., Hao S., Andersen-Nissen E., Mauck W.M., 3rd, Zheng S., Butler A., Lee M.J., Wilk A.J., Darby C., Zager M., et al. Integrated analysis of multimodal single-cell data. Cell. 2021;184:3573–3587.e29. doi: 10.1016/j.cell.2021.04.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Yu G., Wang L.-G., Han Y., He Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16:284–287. doi: 10.1089/omi.2011.0118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Ye K., Schulz M.H., Long Q., Apweiler R., Ning Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics. 2009;25:2865–2871. doi: 10.1093/bioinformatics/btp394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Koboldt D.C., Zhang Q., Larson D.E., Shen D., McLellan M.D., Lin L., Miller C.A., Mardis E.R., Ding L., Wilson R.K. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 2012;22:568–576. doi: 10.1101/gr.129684.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Cibulskis K., Lawrence M.S., Carter S.L., Sivachenko A., Jaffe D., Sougnez C., Gabriel S., Meyerson M., Lander E.S., Getz G. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 2013;31:213–219. doi: 10.1038/nbt.2514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Kim S., Scheffler K., Halpern A.L., Bekritsky M.A., Noh E., Källberg M., Chen X., Kim Y., Beyter D., Krusche P., Saunders C.T. Strelka2: fast and accurate calling of germline and somatic variants. Nat. Methods. 2018;15:591–594. doi: 10.1038/s41592-018-0051-x. [DOI] [PubMed] [Google Scholar]
- 44.Griffith M., Griffith O.L., Smith S.M., Ramu A., Callaway M.B., Brummett A.M., Kiwala M.J., Coffman A.C., Regier A.A., Oberkfell B.J., et al. Genome Modeling System: A Knowledge Management Platform for Genomics. PLoS Comput. Biol. 2015;11:e1004274. doi: 10.1371/journal.pcbi.1004274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Petti A.A., Williams S.R., Miller C.A., Fiddes I.T., Srivatsan S.N., Chen D.Y., Fronick C.C., Fulton R.S., Church D.M., Ley T.J. A general approach for detecting expressed mutations in AML cells using single cell RNA-sequencing. Nat. Commun. 2019;10:3660. doi: 10.1038/s41467-019-11591-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Subramanian A., Tamayo P., Mootha V.K., Mukherjee S., Ebert B.L., Gillette M.A., Paulovich A., Pomeroy S.L., Golub T.R., Lander E.S., Mesirov J.P. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA. 2005;102:15545–15550. doi: 10.1073/pnas.0506580102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Liberzon A., Birger C., Thorvaldsdóttir H., Ghandi M., Mesirov J.P., Tamayo P. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 2015;1:417–425. doi: 10.1016/j.cels.2015.12.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
-
•
Raw whole genome, exome, and single-cell RNA sequencing data have been deposited in the Sequence Read Archive (SRA) and are publicly available as of the date of publication. Accession numbers are listed in the key resources table.
-
•
This paper does not report original code.
-
•
Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.