Visual Abstract
Keywords: kidney development, proximal tubule, transcription factors, transcriptional profiling
Abstract
Significance Statement
HNF4 genes promote proximal tubule differentiation in mice, but their function in human nephrogenesis is not fully defined. This study uses human pluripotent stem cell (PSC)–derived kidney organoids as a model to investigate HNF4A and HNF4G functions. The loss of HNF4A, but not HNF4G, impaired reabsorption-related molecule expression and microvilli formation in human proximal tubules. Cleavage under targets and release using nuclease (CUT&RUN) sequencing and CRISPR-mediated transcriptional activation (CRISPRa) further confirm that HNF4A directly regulates its target genes. Human kidney organoids provide a good model for studying transcriptional regulation in human kidney development.
Background
The proximal tubule plays a major role in electrolyte homeostasis. Previous studies have shown that HNF4A regulates reabsorption-related genes and promotes proximal tubule differentiation during murine kidney development. However, the functions and gene regulatory mechanisms of HNF4 family genes in human nephrogenesis have not yet been investigated.
Methods
We generated HNF4A-knock out (KO), HNF4G-KO, and HNF4A/4G-double KO human pluripotent stem cell lines, differentiated each into kidney organoids, and used immunofluorescence analysis, electron microscopy, and RNA-seq to analyze them. We probed HNF4A-binding sites genome-wide by cleavage under targets and release using nuclease sequencing in both human adult kidneys and kidney organoid-derived proximal tubular cells. Clustered Regularly Interspaced Short Palindromic Repeats–mediated transcriptional activation validated HNF4A and HNF4G function in proximal tubules during kidney organoid differentiation.
Results
Organoids lacking HNF4A, but not HNF4G, showed reduced expression of transport-related, endocytosis-related, and brush border–related genes, as well as disorganized brush border structure in the apical lumen of the organoid proximal tubule. Cleavage under targets and release using nuclease revealed that HNF4A primarily bound promoters and enhancers of genes that were downregulated in HNF4A-KO, suggesting direct regulation. Induced expression of HNF4A or HNF4G by CRISPR-mediated transcriptional activation drove increased expression of selected target genes during kidney organoid differentiation.
Conclusions
This study reveals regulatory mechanisms of HNF4A and HNF4G during human proximal tubule differentiation. The experimental strategy can be applied more broadly to investigate transcriptional regulation in human kidney development.
Introduction
The kidney maintains homeostasis by regulating water and electrolyte balance, vitamin metabolism, and hormone production. Most of these functions are accomplished by nephron epithelia consisting of podocytes, proximal tubular (PT) cells, the loop of Henle, distal tubular cells, and collecting duct cells. PT cells play the major role in reabsorption of the filtrate, including electrolytes, amino acids, glucose, and low molecular weight proteins. Active transport in the PT requires this segment to be highly metabolically active, which also renders it susceptible to ischemic and toxic injury leading to AKI and in some cases progression to CKD.1–3
In mammalian kidney development, the nephron epithelia, including PT cells, are derived from nephron progenitor cells (NPCs).4 Wnt signals from the ureteric bud (UB) induce NPCs to undergo mesenchymal-to-epithelial transition and form the renal vesicle, C-shaped, and S-shaped body.5,6 PT progenitor cells reside in the proximal to medial segments of the S-shaped body.7 Previous studies in mouse have established that Hnf4a is required for differentiation of Cdh6-positive PT progenitors into fully differentiated LTL-high mature PT cells.8,9 Consistent with this result, patients with a point mutation in the HNF4A DNA-binding domain have autosomal dominant Fanconi syndrome,10 reflecting proximal tubule dysfunction. In addition, a recent study has shown that Hnf4a regulates a brush border–related gene program in the murine intestine, kidney, and yolk sac, whereas very low Hnf4g expression was observed in the murine kidney.11 However, the gene regulatory network governed by HNF4A and HNF4G in human nephrogenesis has not been investigated.
Kidney organoids generated from PSCs form a small three-dimensional kidney-like tissue.12,13 Protocols for inducing kidney organoids have been established by the following in vivo kidney developmental process, first directly inducing progenitor cells, NPCs, and UB; then podocytes, PT, and loop of Henle from NPCs;14–20 and collecting duct cells from UB.21–26 Previous studies used kidney organoids from gene-edited PSCs or patient-derived PSCs to model developmental abnormalities and kidney diseases, such as polycystic kidney diseases27–33 and nephrotic syndrome.34–38 Kidney organoids thus provide a good model system to study gene function and elucidate pathophysiological mechanisms during human kidney development.
In this study, we investigated the function of HNF4A and HNF4G during human nephrogenesis using kidney organoids. We generated kidney organoids from HNF4A-knock out (KO), HNF4G-KO, and HNF4A/4G-double KO human PSC lines and examined their molecular and morphologic characteristics. In addition, we tested the functional differences between HNF4A and HNF4G during human kidney organoid differentiation using a conditional CRISPRa system. These studies provide a deeper understanding of human proximal tubule development and the specific gene programs driven by the transcriptional activators HNF4A and HNF4G.
Results
HNF4A and HNF4G Are Top Putative Regulatory Transcription Factors in the Human Proximal Tubules
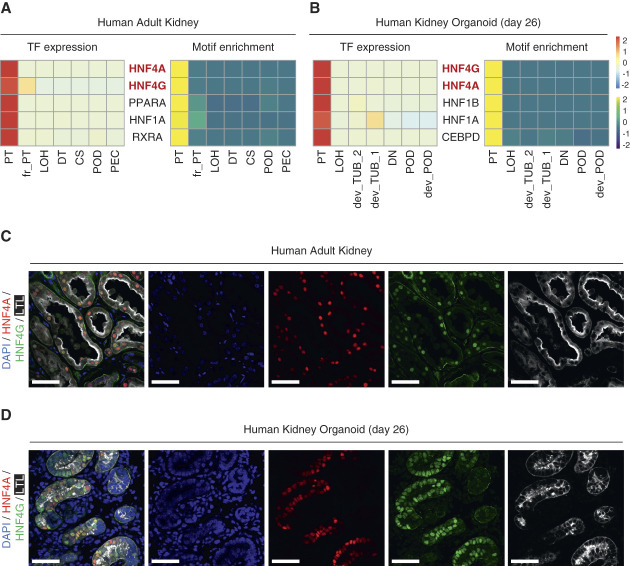
To elucidate transcription the top factors (TFs) regulating human PT, we analyzed single-cell multiome datasets that measured single-nucleus RNA sequencing and single-nucleus assay for transposase-accessible chromatin with sequencing generated from either human adult kidneys39 or human kidney organoids.40 The TFs with combined high gene expression and motif accessibility in the PT are listed (Figure 1, A and B, Supplemental Tables 1 and 2). HNF4A and HNF4G were the highest ranked TFs in both human adult kidneys and kidney organoids. Other listed transcription factors included PPARA and RXRA (Supplemental Tables 1 and 2). PPARA has been reported to regulate fatty acid metabolism in PT41 and has a protective role against PT cell injury in mouse.42,43 RXR forms heterodimers with vitamin D receptor and regulates the action of 1,25(OH)2D3 in the kidney.44,45 RXRA expression in organoid may also imply the involvement of retinoic acid signaling during PT patterning.46,47 We next confirmed HNF4A and HNF4G protein expression in the PT cells by immunostaining (Figure 1, C and D). Interestingly, in the human adult kidney, all LTL-positive PT cells expressed both HNF4A and HNF4G (Figure 1C), whereas not all PT cells in the organoids expressed HNF4A and HNF4G (Figure 1D), most likely reflecting that organoid PT cells are in variable states of maturity.
Figure 1.
HNF4A and HNF4G are top regulatory transcription factors in the human proximal tubular cells. (A, B) Heatmaps showing gene expression levels (left) and motif enrichment in ATAC peaks (right) of top listed transcription factors in the proximal tubular cells of multiome datasets generated from human adult kidneys (A) and kidney organoids (B). (C, D) Immunofluorescence images of HNF4A (red), HNF4G (green), LTL (white), and nuclear DAPI (blue) staining of human adult kidney (C) and day 26 kidney organoid (D). Scale bars indicate 50 µm. CS, connecting segment; dev_POD, developing podocyte; dev_TUB, developing tubule; DN, distal nephron; DT, distal tubule; fr_PT, failed repaired proximal tubule; LOH, Loop of Henle; PEC, parietal epithelial cells in the glomerulus; POD, podocyte; PT, proximal tubule.
Generation of Kidney Organoids from HNF4A-KO, HNF4G-KO, and HNF4A/4G-DKO Human iPSCs
To investigate the function of HNF4A and HNF4G in human nephrogenesis, we generated HNF4A-KO, HNF4G-KO, and HNF4A/4G-DKO human iPSC lines from wild-type AN1.1 human iPSCs using CRISPR gene editing. We targeted early exons common to all isoforms of the HNF4A and HNF4G genes and established two independent KO clones from single-cell colonies that harbored frame-shift alleles in the target regions. Sequencing revealed that both HNF4A-KO clones harbored compound heterozygous frame-shift mutations, whereas both HNF4G-KO clones harbored homozygous frame-shift mutations (Figure 2, A and B). Both HNF4A/4G-DKO clones harbored compound heterozygous frame-shift mutations in HNF4A gene and heterozygous frame-shift mutations in HNF4G gene (Supplemental Figure 1A). All KO clones had the same short tandem repeat polymorphisms compared with the parental AN1.1 clone (Supplemental Figure 1B) and maintained pluripotency marker expression, including OCT4, SOX2, and NANOG (Supplemental Figure 2). We next induced kidney organoids from these KO clones using the commonly used Takasato protocol15,48 because it has been shown to robustly induce PT lineage cells. The organoids from all iPSC clones showed well-epithelialized structures on day 26 (Supplemental Figure 3A). Immunostaining confirmed that all organoids contained multiple nephron epithelial cell types, including WT1-positive podocyte, LTL-positive PT, and CDH1-positive distal nephron epithelia (Figure 2C, Supplemental Figure 3B). Importantly, LTL-positive PT cells in the HNF4A-KO organoids lacked HNF4A protein, those in the HNF4G-KO organoids lacked HNF4G protein, and those in the HNF4A/4G-DKO organoids lacked both HNF4A and HNF4G protein expression (Figure 2D). The absence of either HNF4A or HNF4G did not affect the proportion or number of PT cells in the resulting organoids (Supplemental Figure 3, C and D), suggesting neither HNF4A nor HNF4G is required for early PT lineage specification during human nephrogenesis.
Figure 2.
Kidney organoid induction from HNF4A-KO, HNF4G-KO, and HNF4A/HNF4G-DKO iPSC lines. (A, B) Strategy for targeting human HNF4A (A) and HNF4G (B) genes with CRISPR gene editing to generate HNF4A-KO and HNF4G-KO iPSC lines, respectively. The sequencing at the targeted sites of the HNF4A-KO (A) and HNF4G-KO (B) iPSC lines are shown. Magenta letters and hyphens in the sequencing indicate inserted or deleted bases, respectively. (C) Immunofluorescence images of CDH1 (red, distal nephron epithelium marker), WT1 (green, podocyte marker), LTL (white, proximal tubular cell marker), and nuclear DAPI (blue) staining of day 26 human kidney organoids induced from wild-type (clone AN1.1), HNF4A-KO (clone 2F2), HNF4G-KO (clone 2D3), and HNF4A/4G-KO (clone 3) iPSC lines. Scale bars indicate 200 µm. (D) Immunofluorescence images of HNF4A (red), HNF4G (green), LTL (white), and nuclear DAPI (blue) staining of day 26 human kidney organoids induced from wild-type (clone AN1.1), HNF4A-KO (clone 2G4), HNF4G-KO (clone 2B3), and HNF4A/4G-KO (clone 5) iPSC lines. Scale bars indicate 50 µm.
HNF4A-KO and HNF4A/4G-DKO Organoids Are Characterized by Downregulated Transport, Endocytic, and Brush Border–Related Molecules
To further characterize kidney organoids lacking HNF4A or HNF4G, we performed bulk RNA-seq of day 26 organoids and compared the gene expression profiles of each organoid clone. Principal component analysis revealed that HNF4A-KO and HNF4A/4G-DKO organoids had the largest differences in the PC1 axis compared with either wild-type or HNF4G-KO organoids (Supplemental Figure 4). Consistent with its known role as a transcriptional activator, most genes were downregulated in the KO organoids. Downregulated genes in the HNF4A-KO organoids related to transport (SLC22A8, SLC22A7, and SLC16A4), endocytosis (CUBN), and lipid metabolism (APOA2 and APOM) (Figure 3A). Gene ontology enrichment analysis of the downregulated genes in the HNF4A-KO organoids contained terms, including transport, lipid metabolism, and brush border, the latter is an essential structure for reabsorption in the PT. By contrast, HNF4G-KO organoids had fewer predominantly downregulated genes (Figure 3C). Although downregulated genes in the HNF4G-KO organoids were related to lipid metabolism functions, statistical significance was smaller compared with HNF4A-KO organoids (Figure 3, B and D). Downregulated genes in the HNF4A/4G-DKO organoids were shared with those in the HNF4A-KO organoids, including transport, lipid metabolism, and brush border–related genes (Figure 3, E and F). These results suggest that HNF4A plays a dominant role in regulating PT differentiation compared with HNF4G.
Figure 3.
HNF4A-KO and HNF4A/4G-DKO organoids showed downregulation of transport-related, endocytosis-related, and brush border–related genes. (A, C, E) Volcano plots showing the differentially expressed genes in the comparison of HNF4A-KO (clone 2G4, A), HNF4G-KO (clone 2D3, C), and HNF4A/4G-DKO (clone 5, E) kidney organoids to wild-type (clone AN1.1) kidney organoids. (B, D, F) Enriched pathways from gene ontology analysis of the downregulated genes in the HNF4A-KO (clone 2G4, B), HNF4G-KO (clone 2D3, D), and HNF4A/4G-DKO (clone 5, F) kidney organoids compared with wild-type (clone AN1.1) kidney organoids. Each clone shows an average of different batches of organoids (n=6). Brown bars, biological process (BP); yellow bars, cellular component (CC); green bars, molecular function (MF).
We confirmed by immunostaining that LTL-positive PT cells in the HNF4A-KO and HNF4A/4G-DKO organoids had decreased CUBN expression, but this was not observed in the HNF4G-KO organoids (Figure 4A, Supplemental Figure 5). Furthermore, transmission electron microscopy images revealed that microvilli in the apical lumen of the PT in HNF4A-KO and HNF4A/4G-DKO organoids had wavy and disorganized structures, in contrast to the straight microvilli structure observed in WT organoids (Figure 4B). These morphological differences are consistent with the measured downregulation of brush border–related genes in our RNA-seq analysis (Figure 3, B and F). Together, these results indicate that the loss of HNF4A, but not HNF4G, impairs human PT maturation, including decreased reabsorption-related molecules and incomplete microvilli formation.
Figure 4.

HNF4A-KO and HNF4A/4G-DKO organoids showed disorganized microvilli structure in the proximal tubule. (A) Immunofluorescence images of CUBN (red), HNF4A (green), LTL (white), and nuclear DAPI (blue) staining of day 26 human kidney organoids induced from wild-type (clone AN1.1), HNF4A-KO (clone 2G4), HNF4G-KO (clone 2D3), and HNF4A/4G-KO (clone 5) iPSC lines. Scale bars indicate 50 µm. (B) Transmission electron microscopic images of day 26 human kidney organoids induced from wild-type (clone AN1.1), HNF4A-KO (clone 2G4), HNF4G-KO (clone 2D3), and HNF4A/4G-KO (clone 5) iPSC lines. Lower panels are magnified images of the yellow boxes in their corresponding upper images. Blue arrowheads indicate disorganized microvilli. Scale bars indicate 10 µm (upper panels) and 500 nm (lower panels).
HNF4A Directly Regulates CUBN and SLC16A4 in Proximal Tubules in Human Kidneys and Kidney Organoids
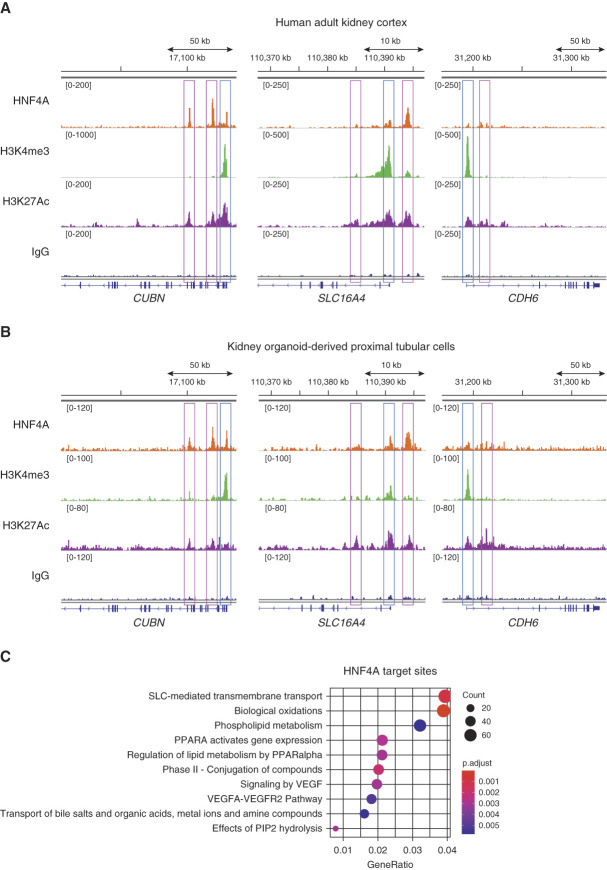
Because HNF4A significantly affects gene regulation in human PT, we next sought to more precisely define direct HNF4A target genes. To do so, we performed CUT&RUN sequencing49,50 with anti-HNF4A and antihistone modification antibodies using both whole human adult kidney cortex and sorted kidney organoid-derived PT cells. In the human adult kidney, HNF4A bound near the transcription start site and two intronic regions of the CUBN gene (Figure 5A). For SLC16A4, the monocarboxylate transporter, HNF4A bound near the transcription start site and at an upstream noncoding region (Figure 5A). For both of these genes, HNF4A-binding sites were associated with increased H3K4me3 (indicating promoter activity) and H3K27Ac (indicating promoter and enhancer activity) binding. Kidney organoid-derived PT cells largely recapitulated these HNF4A-binding patterns (Figure 5B), suggesting that HNF4A directly regulates CUBN and SLC16A4 expression by binding to the promoter and enhancer regions of these genes. By contrast, no significant HNF4A-binding peaks were observed in the CDH6 gene in either human adult kidneys or kidney organoid-derived PT cells (Figure 5, A and B), suggesting that it is not an HNF4A target, consistent with the expression of CDH6 before HNF4A during early nephron development. Furthermore, Reactome pathway analysis supported the notion that HNF4A target genes were highly enriched to SLC-mediated transmembrane transport (Figure 5C). These results indicate that HNF4A directly regulates transport-related genes responsible for critical functions in human PT.
Figure 5.
CUT&RUN sequencing confirmed direct regulation of HNF4A for transport and endocytosis-related genes in the human kidneys. (A, B) Integrative genomics viewer (IGV) images showing target sites of HNF4A (top), H3K4me3 (promoter activity, second from the top), H3K27Ac (enhancer and promoter activity, second from the bottom), and IgG (negative control, bottom) antibodies in CUT&RUN sequencing with human adult kidney cortex (A) and kidney organoid-derived proximal tubule (B) of CUBN (left), SLC16A4 (middle), and CDH6 (right) genes. Blue boxes indicate promoter regions; magenta boxes indicate enhancer regions. (C) Enriched pathways from Reactome pathway analysis of the HNF4A-binding peaks detected in the CUT&RUN analysis of human adult kidney cortex.
Conditional Activation of HNF4A and HNF4G in Kidney Organoids Using CRISPR Activation
To provide direct evidence of HNF4A or HNF4G target gene activation, we used the CRISPRa system in which the transcriptional activator VP64-p65-Rta is fused to nuclease-null Cas9 (dCAS9-VPR) plasmid51 with a gRNA-containing plasmid targeting the HNF4A or HNF4G transcriptional start site (Supplemental Figure 6A). We tested multiple gRNAs and chose the most efficient gRNAs for activation in HEK293T cells (Supplemental Figure 6B). Among three candidate gRNAs, HNF4A gRNA_2 showed approximately 800-fold upregulation of HNF4A (Supplemental Figure 6C). Compared with HNF4A, gRNAs targeting HNF4G resulted in milder upregulation; however, gRNA_3 still significantly upregulated HNF4G by approximately three-fold in HEK293T cells (Supplemental Figure 6D).
Using the optimal gRNAs, we next applied CRISPRa to the kidney organoid culture system. Previous reports have shown that HNF4A is a critical regulator for hepatic cell differentiation in the mouse and from human PSCs,52–55 and we were therefore concerned that constitutive HNF4A activation would lead to hepatocyte lineage rather than the kidney. To provide temporal control of HNF4A expression during kidney organoid induction, we used a doxycycline (DOX)–inducible CRISPRa human ES cell line containing dCas9 fused to both the VPR activation domain and to EGFP into the AAVS1 safe-harbor locus.56 This cell line allows transient expression of dCAS9, VPR, and EGFP with DOX exposure. We first transduced lentiviral gRNA plasmids targeting HNF4A or HNF4G to the dCAS9-VPR hESCs and established gRNA-containing clones by puromycin selection (Figure 6A). We then induced kidney organoids from these PSCs and treated them with DOX for 5 days (days 21–26) before collection. At day 26, we confirmed EGFP expression of the DOX-treated organoids by flow cytometry analysis (Supplemental Figure 6E). HNF4A-CRISPRa organoids showed more than ten-fold upregulation of HNF4A (Figure 6B), and HNF4G-CRISPRa organoids showed about six-fold upregulation of HNF4G (Figure 6C) compared with DOX untreated organoids. Importantly, the HNF4A-CRISPRa organoids showed upregulation of CUBN and transporter-related molecules, including SLC3A1 and SLC16A4, compared with HNF4G-CRISPRa organoids (Figure 6, B–D). These results confirm that HNF4A directly regulates these target genes during human nephrogenesis.
Figure 6.
CRISPR activation using kidney organoid system confirmed transcriptional activity differences between HNF4A and HNF4G. (A) Schematic of experiment for CRISPR activation (CRISPRa) during kidney organoid differentiation. (B, C) qRT-PCR results of CRISPRa kidney organoids (day 26) targeting HNF4A (B) and HNF4G (C). n=3 biological replicates. The data are presented as mean±SEM. * P < 0.05, ** P < 0.01. (D) Immunofluorescence images of SLC3A1 (magenta), HNF4A (yellow), and nuclear DAPI (blue) staining of CRISPRa kidney organoids targeting HNF4A without DOX (left) and with DOX (right) conditions. Scale bars indicate 100 μm. hES cells, human embryonic stem cells; n.s., not significant.
Discussion
Here, we identify HNF4A and HNF4G as top candidate regulators of human PT differentiation on the basis of analysis of human adult kidney and human kidney organoid single-cell multiome datasets. Kidney organoids generated from either HNF4A-KO or HNF4G-KO human PSCs still contained LTL-positive PT, but only organoids lacking HNF4A were characterized by decreased expression of membrane transport and endocytosis-related molecules and microvilli malformation. Using CUT&RUN sequencing, we validated the direct regulation of HNF4A for select target genes. We further investigated functional differences between HNF4A and HNF4G genes using a conditional CRISPRa system and confirmed expression changes in genes regulated by HNF4A in the organoid model.
Our results suggest that the HNF4A gene regulatory network includes molecules responsible for reabsorption and electrolyte balance in human PT. These results are consistent with the previous findings of nephron-specific Hnf4a-deficient mice9 and patients with HNF4A mutation,10 indicating that HNF4A is a critical PT regulator across species. In our organoid model, HNF4A-KO kidney organoids contained LTL-positive PT cells, whereas Hnf4a-deficient postnatal (P0) murine kidneys show impaired differentiation into LTL-high PT cells.9 Given the fact that LTL-weak/LRP2-weak PT cells are present in the Hnf4a-deficient murine kidneys, we speculate that this phenotypic difference may reflect the relative immaturity of organoid cell types in that organoid PT cells are not mature enough to show strong LTL intensity. Our results both from KO organoids and CRISPRa experiments suggest that HNF4G is a weaker regulator than HNF4A, at least during kidney organoid differentiation, consistent with previous reports in C2 and HeLa cell lines.57 However, an important caveat from these studies is that our organoid model does not allow us to make conclusions about potential roles for HNF4G in fully mature PT (because organoid cell types lack full differentiation) or in response to cell injury.
To elucidate a detailed transcriptional regulatory mechanism of HNF4A, we used CUT&RUN to identify direct HNF4A-binding targets. To the best of our knowledge, this study described HNF4A-binding targets in human kidneys. HNF4A directly binds to transporters and brush border–related genes, consistent with previous reports that performed with ChIP-seq using anti-HNF4A antibody in murine kidneys.9,11 On the other hand, we could not obtain clear results for HNF4G target sites in our preliminary study using human adult kidneys. The anti-Hnf4g antibody used for ChIP-seq with murine intestine58 is no longer available. We tried two commercially available antibodies, but both had broad background peaks (data not shown).
We used a conditional CRISPRa system during organoid differentiation to enhance gene expression. CRISPRa can activate endogenous genes better than conventional overexpression.59 To the best of our knowledge, this study uses CRISPRa for the functional validation of transcription factors in kidney organoids. Our approach allowed for temporal control of gene expression using a PSC line with the Tet-ON system. This prevented endoderm–hepatocyte lineage differentiation55 and enabled successful HNF4A activation after kidney lineage commitment. A limitation is that CRISPRa in this study was not performed in a cell-specific manner, although we could confirm HNF4A protein upregulation in PT. The responsiveness to HNF4A activation may vary among cell types in kidney organoids. Future strategies to apply a conditional and cell type-specific CRISPRa system may reveal more precise regulation of HNF4A and HNF4G during human PT differentiation.
In conclusion, we elucidated the regulatory mechanisms of HNF4A and HNF4G in human kidney development using kidney organoids. The experimental approach presented in this study will facilitate using kidney organoids for studying transcriptional regulation.
Methods
Data and Materials Availability
The datasets generated and analyzed in this study, including RNA-seq and CUT&RUN sequencing, are available in GEO: GSE226441 (reviewer password: gvsvsmwilfixtqv). The sn-multiome datasets for human adult kidneys are available in GEO: GSE220289. The datasets for kidney organoid sn-multiome analysis and CUT&RUN sequencing for human adult kidney and kidney organoid-derived PT cells are available in GEO: GSE213152.
Cell Lines
AN1.1 human iPSC and H9 human ESC lines were obtained from Genome Engineering & Stem Cell Center at McDonnell Genome Institute. dCAS9-VPR human ESC line56 was made using WiCell's WA01 cell line and provided by Dr. Kevin Eggan and Dr. Lindy Barrett on MTA. All human PSC lines were maintained and expanded in a humidified 5% CO2 atmosphere at 37°C in Stem Flex medium (Thermo Fisher Scientific, A3349401) on Matrigel-coated (Corning, 354277) plate with daily media changes. Cells were passaged every 3–4 days with ReLeSR (STEMCELL Technologies, 05872). Cells were confirmed to be karyotypically normal and mycoplasma-free. Experiments were performed below passage 50. HEK293T cells (ATCC, CRL-3216) were maintained in a humidified 5% CO2 atmosphere at 37°C in Dulbecco modified Eagle medium (DMEM, Gibco; 11965092) supplemented with 10% fetal bovine serum (Gibco; 10437028) and antibiotics. HEK293T cells were passaged by treatment with 0.05% trypsin-EDTA (Thermo Fisher Scientific, 25300054).
Generation of HNF4A-KO, HNF4G-KO, and HNF4A/4G-DKO iPSC Lines
The KO iPSC lines were created from AN1.1 human iPSCs. In brief, synthetic gRNA targeting the sequence was purchased from Integrated DNA Technologies, complexed with Cas9 recombinant protein, and transfected into iPSCs. The transfected cells were then single-cell sorted into 96-well plates, and single-cell clones were identified using NGS to analyze the target site region as those harboring only out-of-frame alleles. HNF4A gRNA (5′- TACGGTGCCTCGAGCTGTGANGG -3′) was used to generate HNF4A-KO iPSCs (clones #2F2 and #2G4). HNF4G gRNA (5′- CACTGGCACAAGCTGAAGTTNGG -3′) was used to generate HNF4G-KO iPSCs (clones #2B3 and #2D3). The HNF4G gRNA was also used to generate HNF4A/4G-DKO iPSCs (clones #3 and #5) from the HNF4A-KO iPSC line (clone #2G4). The short tandem repeat analysis of KO iPSC lines was performed using next-generation sequencing on the targets to confirm cell identity to their parental line.
Kidney Organoid Induction
Kidney organoids were generated by Takasato protocol15,48 with slight modifications. In brief, on the next day of passage, PSCs were treated with 8 µM CHIR99021 (TOCRIS, 4423) in basal medium that comprised APEL2 (STEMCELL Technologies, 05275) supplemented with 1.5% Protein-Free Hybridoma Medium II (Thermo Fisher Scientific, 12040077) and 1× antibiotic–antimycotic solution (Corning, 30-004-CI) for 4 days. On day 4, medium was switched to 200 ng/ml FGF9 (R&D Systems, 273-F9-025) and 1 μg/ml heparin (Sigma-Aldrich, H4784) in the basal medium. On day 7, cells were dissociated into single cells using Accutase (Sigma-Aldrich, SF006) and diluted five-fold with the basal medium with 10 µM Y27632 (TOCRIS, 1254). In total, 2×105 cells were spun down at 300 g for 2 minutes (twice with 180° flip) to make cell aggregates in a low cell-binding tube (VWR, 490003-230) and transferred onto a 0.4 µm pore polyester membrane transwell (Corning, 3460). Pellets were incubated with 5 µM CHIR99021 in the basal medium for 1 hour and then cultured with 200 ng/ml FGF9 and 1 µg/ml heparin in the basal medium for 5 days. From day 12 to day 26, organoids were grown in the basal medium. Medium were changed every other day.
RNA Sequencing and Analysis
RNA was extracted with a RNeasy Plus Micro Kit (Qiagen, 74034), and total RNA integrity was determined using a 4150 TapeStation (Agilent). Libraries were prepared using a SMARTer cDNA synthesis kit (Takara) according to the manufacturer's instruction and then indexed, pooled, and sequenced on an Illumina NovaSeq 6000. Basecalls and demultiplexing were performed with Illumina's bcl2fastq2 software. RNA-seq reads were then aligned and quantitated to the Ensembl release 101 primary assembly with an Illumina DRAGEN Bio-IT on-premise server running version 3.9.3–8 software. The count matrices were then input to edgeR60 for data normalization and differential gene analysis. Gene ontology enrichment analysis was performed with DAVID (v6.8)61 for functional annotation of differentially expressed genes.
CUT&RUN Sequencing and Analysis
CUT&RUN assay libraries for human adult kidneys or kidney organoid-derived PT cells were generated with the CUTANA ChIC/CUT&RUN Kit (EpiCypher, 14–1048). For human adult kidney, 5.0×105 isolated nuclei were fixed with 0.5% formalin for 1 minute at room temperature, followed by addition of glycine. Kidney organoid-derived PT cells were sorted as an LTL-positive population by flow cytometry, filtered through a 40 μm cell strainer, and fixed with 0.5% formalin for 1 minute at room temperature. After inactivating unreacted aldehyde by adding glycine, 1.0×105 cells were treated with chilled lysis buffer on ice for 5 minutes to obtain nuclei. Fixed nuclei were then incubated with Concanavalin A (ConA)–conjugated paramagnetic beads and stained with 0.5 µg of antibodies. The remaining steps were performed according to the manufacturer's instructions. Library preparation was performed using the NEBNext Multiplex Oligos for Illumina (New England Biolabs, E6440S) with the manufacturer's instructions, including minor modifications indicated by CUTANA described above. CUT&RUN libraries were sequenced on a NovaSeq instrument (Illumina, 150 bp paired-end reads). FASTQ files were trimmed with Trim Galore (Cutadapt [v2.8]) and aligned with Bowtie2 (parameters: —local —very sensitive-local —no-unal —no-mixed —no-discordant —phred33 -I 10 -X 700) using hg38. Peak calling was performed using MACS2 (v2.2.7.1) with default parameters using samtools (1.9) and DeepTools (3.5.0). The bam files were converted to BigWig format with bigwigbamCoverage function in DeepTools, and the data were visualized with Interactive Genome Viewer (v2.6.3). For CUT&RUN for H3K4me3 and H3K27Ac, we used our deposited datasets (GSE213152). Genomic regions containing HNF4A-binding peaks were annotated with ChIPSeeker (v1.30.3),62 and functional enrichment analysis was performed using Reactome.63
CRISPR Activation
Small guide RNA (sgRNA) for activating HNF4A and HNF4G genes was designed using CHOPCHOP (https://chopchop.cbu.uib.no/). Sequences of the sgRNAs are presented in Supplemental Table 4. Oligonucleotides of 5′CACC-sense gRNA -3′ and 5′ AAAC-antisense gRNA -3′ were purchased from Integrated DNA Technology. The sgRNAs were inserted into downstream of the U6 promoter of the lentiGuide-Puro plasmid (a gift from Dr. Feng Zhang, Addgene, 52963). In brief, sense and antisense oligonucleotides were annealed by cooling from 95°C to 25°C for 1.5 hours. The annealed oligonucleotides were then cloned into the lentiGuide-Puro plasmid by Golden Gate Assembly with Esp3I restriction enzyme (NEB, R0734L) and T4 DNA ligase (NEB, M0202L) on a thermal cycler repeating 37°C for 5 minutes and 16°C for 5 minutes for 60 cycles, followed by transformation to NEB 5-α Competent E. coli (NEB, C2987H) as the manufacturer's instruction. The cloned lentiviral vectors were purified with mini high-speed plasmid kit (IBI Scientific; IB47102), and sgRNA insertion was confirmed with Sanger sequencing by GENEWIZ.
For transfection to HEK293T cells, Lipofectamine 3000 transfection reagent (Thermo Fisher Scientific, L3000015) was used according to the manufacturer's instructions. In brief, HEK293T cells were seeded at 2.0×105 cells per well on 12-well tissue culture plates. After 18–24 hours, cells were transfected with 1.0 µg of plasmid DNA, 2.0 µl of P3000TM enhancer reagent, and 3.0 µl of Lipofectamine 3000 reagent in Opti-MEM (Thermo Fisher Scientific, 31-985-070). At 48 hours after transfection, transfected cells were collected and evaluated using qRT-PCR.
To generate lentivirus, HEK293T cells were seeded at 3.0×106 cells on a 10-cm tissue culture dish 1 day before transfection. The cells were transfected with 9.0 µg of psPAX2 (a gift from Dr. Didier Trono, Addgene, 12260), 0.9 µg of pMD2.G (a gift from Dr. Didier Trono, Addgene, 12259), and 9.0 µg of lentiGuide-Puro plasmid with sgRNAs by Lipofectamine 3000 transfection reagent according to the manufacturer's instructions. Culture media were changed to DMEM supplemented with 30% FBS at 24 hours after transfection. Lentivirus-containing supernatants were collected 24 and 48 hours later, and they were filtered with 0.45 µm PVDF filters (CELLTREAT, 229745). The resultant supernatants were concentrated using Lenti-X concentrator (Takara Bio, 631231). The titer of the concentrated lentivirus was measured using the qPCR Lentivirus Titer Kit (Applied Biological Materials, LV900).
For transduction to PSCs, we transiently changed PSC culture method to the single-cell passage using StemFit Basic04 (AJINOMOTO, BASIC04CT) to increase transduction efficiency. dCAS9-VPR human ESCs were seeded at 2.0×104 cells per well on Matrigel-coated 12-well tissue culture plates, and MOI five of the concentrated lentiGuide-Puro lentiviruses were added during seeding. In total, 10 µM of Y27632 was also added to the medium for 48 hours after passage. After 48 hours, lentivirus-containing media was replaced to StemFit Basic04 supplemented with 0.5 µg/ml puromycin. The transduced cells were passaged at 2.6×104 cells per well on Matrigel-coated 12-well tissue culture and treated with 0.5 µg/ml puromycin for a total of 7 days. After puromycin selection, gRNA-containing dCAS9-VPR ESCs were induced to differentiation into kidney organoids as described above. From day 21–26 after the initiation of differentiation, 1 µg/ml of DOX was added to the medium in the lower chamber of transwell to activate Tet-ON system. At day 26, organoids treated with and without DOX were collected and evaluated.
Quantification and Statistical Analysis
Details of sample size for each experiment are provided in the figure legends. Data were presented as mean±SEM. The t test was used for two-group comparison. ANOVA with post hoc Bonferroni correction was used for multiple group comparison. No statistical methods were used to predetermine sample size. Experiments were not randomized, and investigators were not blinded to allocation during library preparation, experiments, or analysis.
Supplementary Material
Acknowledgments
We thank all members of the Humphreys lab for helpful discussions, the Genome Engineering & Stem Cell Center (GESC) at the McDonnell Genome Institute (MGI) at Washington University in St. Louis for creating the knock-out iPSC lines, Genome Technology Access Center (GTAC) at the MGI for performing sequencing, and Washington University Center for Cellular Imaging (WUCCI) for assisting with electron microscopy imaging. Y.Y. and B.D.H. designed the study and wrote the manuscript. Y.Y. performed experiments and analysis with contributions from Y.M., K.O., Y.Y., Y.M., K.O., J.H.M., and B.D.H. analyzed data. All authors read and approved the final manuscript.
Disclosures
B.D. Humphreys reports Consultancy: Chinook Therapeutics, Janssen, Pfizer; Ownership Interest: Chinook Therapeutics; Research Funding: Janssen, Pfizer; Honoraria: Novartis; Patents or Royalties: AG, Evotec; and Advisory or Leadership Role: Seminars in Nephrology: Editorial Board, Kidney International: Editorial Board, JCI Insight: Editorial Board, American Journal of Physiology Renal Physiology, Editorial Board, RegMed XB, Regenerative Medicine Crossing Borders: SAB, ASCI: President, Chinook Therapeutics, SAB, Member of Board of Scientific Advisors of NIDDK. J.H. Miner reports Consultancy: Bayer, Guidepoint; Research Funding: Chinook Therapeutics, Keros Therapeutics, Lung Therapeutics Inc, Myonid Therapeutics; Honoraria: AstraZeneca, Axiom Healthcare Strategies, Pfizer, Visterra; Patents or Royalties: Elsevier; Kerafast; Maze Therapeutics; Advisory or Leadership Role: American Society for Matrix Biology, President; Journal of Clinical Investigation, Consulting Editor; Kidney International, Editorial Board; Matrix Biology, Editorial Board; Matrix Biology Plus, Editorial Board; and Other Interests or Relationships: Alport Syndrome Foundation (Scientific Advisory Research Network). All remaining authors have nothing to disclose.
Funding
This work was funded by seed network grant CZF2019-002430 from the Chan Zuckerberg Initiative (to B.D.H.). Additional support was from the Japan Society for the Promotion of Science (JSPS) Overseas Research (to Y.Y.).
Author Contributions
Conceptualization: Benjamin D. Humphreys, Yasuhiro Yoshimura.
Data curation: Kohei Omachi, Yasuhiro Yoshimura.
Formal analysis: Yoshiharu Muto, Yasuhiro Yoshimura.
Funding acquisition: Benjamin D. Humphreys, Yasuhiro Yoshimura.
Investigation: Yoshiharu Muto, Kohei Omachi, Yasuhiro Yoshimura.
Resources: Yoshiharu Muto.
Software: Yasuhiro Yoshimura.
Supervision: Benjamin D. Humphreys, Jeffrey H. Miner.
Validation: Yoshiharu Muto, Kohei Omachi, Yasuhiro Yoshimura.
Writing – original draft: Benjamin D. Humphreys, Yasuhiro Yoshimura.
Writing – review & editing: Benjamin D. Humphreys, Jeffrey H. Miner, Yoshiharu Muto, Kohei Omachi, Yasuhiro Yoshimura.
Supplemental Material
This article contains the following supplemental material online at http://links.lww.com/JSN/E487.
Supplemental Figure 1. Characterization of HNF4A-KO, HNF4G-KO, and HNF4A/4G-DKO iPSC Lines.
Supplemental Figure 2. All generated KO iPSC lines maintained pluripotency marker expression.
Supplemental Figure 3. Kidney organoids induced from HNF4A-KO, HNF4G-KO, and HNF4A/4G-DKO iPSC lines contained LTL-positive proximal tubular cells.
Supplemental Figure 4. Principal component analysis (PCA) plot of RNA-seq (n=6 for each clone) according to gene expression levels.
Supplemental Figure 5. HNF4A-KO and HNF4A/4G-DKO organoids induced from another each KO iPSC clone showed CUBN downregulation.
Supplemental Figure 6. CRISPR activation for HNF4A and HNF4G genes using HEK293T cells and human kidney organoids.
Supplemental Table 1. Top ten transcription factors of the proximal tubular cell cluster in the multiome datasets (snATAC-seq and snRNA-seq) from human adult kidneys.
Supplemental Table 2. Top ten transcription factors of the proximal tubular cell cluster in the multiome datasets (snATAC-seq and snRNA-seq) from day 26 kidney organoids.
Supplemental Table 3. Antibody information used in this study.
Supplemental Table 4. Primer and gRNA sequences used in this study.
References
- 1.Takaori K, Nakamura J, Yamamoto S, Nakata H, Sato Y, Takase M. Severity and frequency of proximal tubule injury determines renal prognosis. J Am Soc Nephrol. 2016;27(8):2393–2406. doi: 10.1681/ASN.2015060647 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Kirita Y, Wu H, Uchimura K, Wilson PC, Humphreys BD. Cell profiling of mouse acute kidney injury reveals conserved cellular responses to injury. Proc Natl Acad Sci. 2020;117(27):15874–15883. doi: 10.1073/pnas.2005477117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Gerhardt LMS, Liu J, Koppitch K, Cippà PE, McMahon AP. Single-nuclear transcriptomics reveals diversity of proximal tubule cell states in a dynamic response to acute kidney injury. Proc Natl Acad Sci USA. 2021;118(27):e2026684118. doi: 10.1073/pnas.2026684118 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Kobayashi A, Valerius MT, Mugford JW, Carroll TJ, Self M, Oliver G. Six2 defines and regulates a multipotent self-renewing nephron progenitor population throughout mammalian kidney development. Cell Stem Cell. 2008;3(2):169–181. doi: 10.1016/j.stem.2008.05.020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Costantini F, Kopan R. Patterning a complex organ: branching morphogenesis and nephron segmentation in kidney development. Dev Cell. 2010;18(5):698–712. doi: 10.1016/j.devcel.2010.04.008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Schnell J, Achieng M, Lindström NO. Principles of human and mouse nephron development. Nat Rev Nephrol. 2022;18:628–642. doi: 10.1038/s41581-022-00598-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lindström NO, Sealfon R, Chen X, Parvez RK, Ransick A, De Sena Brandine G. Spatial transcriptional mapping of the human nephrogenic program. Dev Cell. 2021;56(16):2381–2398.e6. doi: 10.1016/j.devcel.2021.07.017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Marable SS, Chung E, Adam M, Potter SS, Park J-S. Hnf4a Deletion in the Mouse Kidney Phenocopies Fanconi Renotubular Syndrome. JCI Insight [Internet]. 2018. https://insight.jci.org/articles/view/97497 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Marable SS, Chung E, Park J-S. Hnf4a is required for the development of Cdh6-expressing progenitors into proximal tubules in the mouse kidney. JASN. 2020;31(11):2543–2558. doi: 10.1681/ASN.2020020184 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Hamilton AJ, Bingham C, McDonald TJ, Cook PR, Caswell RC, Weedon MN. The HNF4A R76W mutation causes atypical dominant Fanconi syndrome in addition to a β cell phenotype. J Med Genet. 2014;51(3):165–169. doi: 10.1136/jmedgenet-2013-102066 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Chen L, Luo S, Dupre A, Vasoya RP, Parthasarathy A, Aita R. The nuclear receptor HNF4 drives a brush border gene program conserved across murine intestine, kidney, and embryonic yolk sac. Nat Commun. 2021;12(1):2886. doi: 10.1038/s41467-021-22761-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Nishinakamura R. Human kidney organoids: progress and remaining challenges. Nat Rev Nephrol. 2019;15(10):613–624. doi: 10.1038/s41581-019-0176-x [DOI] [PubMed] [Google Scholar]
- 13.Little MH, Combes AN. Kidney organoids: accurate models or fortunate accidents. Genes Dev. 2019;33(19-20):1319–1345. doi: 10.1101/gad.329573.119 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Taguchi A, Kaku Y, Ohmori T, Sharmin S, Ogawa M, Sasaki H. Redefining the in vivo origin of metanephric nephron progenitors enables generation of complex kidney structures from pluripotent stem cells. Cell Stem Cell. 2014;14(1):53–67. doi: 10.1016/j.stem.2013.11.010 [DOI] [PubMed] [Google Scholar]
- 15.Takasato M, Er PX, Chiu HS, Maier B, Baillie GJ, Ferguson C. Kidney organoids from human iPS cells contain multiple lineages and model human nephrogenesis. Nature. 2015;526(7574):564–568. doi: 10.1038/nature15695 [DOI] [PubMed] [Google Scholar]
- 16.Morizane R, Lam AQ, Freedman BS, Kishi S, Valerius MT, Bonventre JV. Nephron organoids derived from human pluripotent stem cells model kidney development and injury. Nat Biotechnol. 2015;33(11):1193–1200. doi: 10.1038/nbt.3392 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Przepiorski A, Sander V, Tran T, Hollywood JA, Sorrenson B, Shih J-H. A simple bioreactor-based method to generate kidney organoids from pluripotent stem cells. Stem Cell Rep. 2018;11(2):470–484. doi: 10.1016/j.stemcr.2018.06.018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Low JH, Li P, Chew EGY, Zhou B, Suzuki K, Zhang T. Generation of human PSC-derived kidney organoids with patterned nephron segments and a de novo vascular network. Cell Stem Cell. 2019;25(3):373–387.e9. doi: 10.1016/j.stem.2019.06.009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Yoshimura Y, Taguchi A, Tanigawa S, Yatsuda J, Kamba T, Takahashi S. Manipulation of nephron-patterning signals enables selective induction of podocytes from human pluripotent stem cells. JASN. 2019;30(2):304–321. doi: 10.1681/ASN.2018070747 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Vanslambrouck JM Wilson SB Tan KS, et al. Enhanced Metanephric Specification to Functional Proximal Tubule Enables Toxicity Screening and Infectious Disease Modelling in Kidney Organoids. [Internet]. 2021. https://www.biorxiv.org/content/10.1101/2021.10.14.464320v2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Taguchi A, Nishinakamura R. Higher-order kidney organogenesis from pluripotent stem cells. Cell Stem Cell. 2017;21(6):730–746.e6. doi: 10.1016/j.stem.2017.10.011 [DOI] [PubMed] [Google Scholar]
- 22.Uchimura K, Wu H, Yoshimura Y, Humphreys BD. Human pluripotent stem cell-derived kidney organoids with improved collecting duct maturation and injury modeling. Cell Rep. 2020;33(11):108514. doi: 10.1016/j.celrep.2020.108514 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Mae S-I, Ryosaka M, Sakamoto S, Matsuse K, Nozaki A, Igami M. Expansion of human iPSC-derived ureteric bud organoids with repeated branching potential. Cell Rep. 2020;32(4):107963. doi: 10.1016/j.celrep.2020.107963 [DOI] [PubMed] [Google Scholar]
- 24.Howden SE, Wilson SB, Groenewegen E, Starks L, Forbes TA, Tan KS. Plasticity of distal nephron epithelia from human kidney organoids enables the induction of ureteric tip and stalk. Cell Stem Cell. 2021;28(4):671–684.e6. doi: 10.1016/j.stem.2020.12.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Zeng Z, Huang B, Parvez RK, Li Y, Chen J, Vonk AC. Generation of patterned kidney organoids that recapitulate the adult kidney collecting duct system from expandable ureteric bud progenitors. Nat Commun. 2021;12(1):3641. doi: 10.1038/s41467-021-23911-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Shi M, McCracken KW, Patel AB, Zhang W, Ester L, Valerius MT. Human ureteric bud organoids recapitulate branching morphogenesis and differentiate into functional collecting duct cell types. Nat Biotechnol. 2023;41(2):252–261. doi: 10.1038/s41587-022-01429-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Freedman BS, Brooks CR, Lam AQ, Fu H, Morizane R, Agrawal V. Modelling kidney disease with CRISPR-mutant kidney organoids derived from human pluripotent epiblast spheroids. Nat Commun. 2015;6(1):8715. doi: 10.1038/ncomms9715 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Cruz NM, Song X, Czerniecki SM, Gulieva RE, Churchill AJ, Kim YK. Organoid cystogenesis reveals a critical role of microenvironment in human polycystic kidney disease. Nat Mater. 2017;16(11):1112–1119. doi: 10.1038/nmat4994 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Kuraoka S, Tanigawa S, Taguchi A, Hotta A, Nakazato H, Osafune K. PKD1-Dependent renal cystogenesis in human induced pluripotent stem cell-derived ureteric bud/collecting duct organoids. JASN. 2020;31(10):2355–2371. doi: 10.1681/ASN.2020030378 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Shimizu T, Mae S-I, Araoka T, Okita K, Hotta A, Yamagata K. A novel ADPKD model using kidney organoids derived from disease-specific human iPSCs. Biochem Biophysical Res Commun. 2020;529(4):1186–1194. doi: 10.1016/j.bbrc.2020.06.141 [DOI] [PubMed] [Google Scholar]
- 31.Cruz NM, Reddy R, McFaline-Figueroa JL, Tran C, Fu H, Freedman BS. Modelling ciliopathy phenotypes in human tissues derived from pluripotent stem cells with genetically ablated cilia. Nat Biomed Eng. 2022;6(4):463–475. doi: 10.1038/s41551-022-00880-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Hiratsuka K, Miyoshi T, Kroll KT, Gupta NR, Valerius MT, Ferrante T. Organoid-on-a-chip model of human ARPKD reveals mechanosensing pathomechanisms for drug discovery. Sci Adv. 2022;8(38):eabq0866. doi: 10.1126/sciadv.abq0866 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Tran T, Song CJ, Nguyen T, Cheng S-Y, McMahon JA, Yang R. A scalable organoid model of human autosomal dominant polycystic kidney disease for disease mechanism and drug discovery. Cell Stem Cell. 2022;29(7):1083–1101.e7. doi: 10.1016/j.stem.2022.06.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Tanigawa S, Islam M, Sharmin S, Naganuma H, Yoshimura Y, Haque F. Organoids from nephrotic disease-derived iPSCs identify impaired NEPHRIN localization and slit diaphragm formation in kidney podocytes. Stem Cell Rep. 2018;11(3):727–740. doi: 10.1016/j.stemcr.2018.08.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Hale LJ, Howden SE, Phipson B, Lonsdale A, Er PX, Ghobrial I. 3D organoid-derived human glomeruli for personalised podocyte disease modelling and drug screening. Nat Commun. 2018;9(1):5167. doi: 10.1038/s41467-018-07594-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ohmori T, De S, Tanigawa S, Miike K, Islam M, Soga M. Impaired NEPHRIN localization in kidney organoids derived from nephrotic patient iPS cells. Sci Rep. 2021;11(1):3982. doi: 10.1038/s41598-021-83501-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Jansen J, van den Berge BT, van den Broek M, Maas RJ, Daviran D, Willemsen B. Human pluripotent stem cell-derived kidney organoids for personalized congenital and idiopathic nephrotic syndrome modeling. Development. 2022;149(9):dev200198. doi: 10.1242/dev.200198 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Dorison A, Ghobrial I, Graham A, Peiris T, Forbes TA, See M. Kidney organoids generated using an allelic series of NPHS2 point variants reveal distinct intracellular podocin mistrafficking. J Am Soc Nephrol. 2023;34(1):88–109. doi: 10.1681/ASN.2022060707 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Ledru N Wilson PC Muto Y, et al. Predicting Regulators of Epithelial Cell State through Regularized Regression Analysis of Single Cell Multiomic Sequencing. [Internet]. 2022. https://www.biorxiv.org/content/10.1101/2022.12.29.522232v1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Yoshimura Y, Muto Y, Ledru N, Wu H, Omachi K, Miner JH. A single-cell multiomic analysis of kidney organoid differentiation. Proc Natl Acad Sci USA. 2023;120(20):e2219699120. doi: 10.1073/pnas.2219699120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kamijo Y, Hora K, Tanaka N, Usuda N, Kiyosawa K, Nakajima T. Identification of functions of peroxisome proliferator-activated receptor α in proximal tubules. J Am Soc Nephrol. 2002;13(7):1691–1702. doi: 10.1097/01.ASN.0000018403.61042.56 [DOI] [PubMed] [Google Scholar]
- 42.Kamijo Y, Hora K, Kono K, Takahashi K, Higuchi M, Ehara T. PPARα protects proximal tubular cells from acute fatty acid toxicity. J Am Soc Nephrol. 2007;18(12):3089–3100. doi: 10.1681/ASN.2007020238 [DOI] [PubMed] [Google Scholar]
- 43.Li S, Nagothu KK, Desai V, Lee T, Branham W, Moland C. Transgenic expression of proximal tubule peroxisome proliferator–activated receptor-α in mice confers protection during acute kidney injury. Kidney Int. 2009;76(10):1049–1062. doi: 10.1038/ki.2009.330 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Sugawara A, Sanno N, Takahashi N, Osamura RY, Abe K. Retinoid X receptors in the kidney: their protein expression and functional significance. Endocrinology. 1997;138(8):3175–3180. doi: 10.1210/endo.138.8.5351 [DOI] [PubMed] [Google Scholar]
- 45.Christakos S, Dhawan P, Liu Y, Peng X, Porta A. New insights into the mechanisms of vitamin D action. J Cell Biochem. 2003;88(4):695–705. doi: 10.1002/jcb.10423 [DOI] [PubMed] [Google Scholar]
- 46.Wingert RA, Selleck R, Yu J, Song H-D, Chen Z, Song A. The cdx genes and retinoic acid control the positioning and segmentation of the zebrafish pronephros. PLOS Genet. 2007;3(10):e189. doi: 10.1371/journal.pgen.0030189 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Naylor RW, Davidson AJ. Pronephric tubule formation in zebrafish: morphogenesis and migration. Pediatr Nephrol. 2017;32(2):211–216. doi: 10.1007/s00467-016-3353-1 [DOI] [PubMed] [Google Scholar]
- 48.Takasato M, Er PX, Chiu HS, Little MH. Generation of kidney organoids from human pluripotent stem cells. Nat Protoc. 2016;11(9):1681–1692. doi: 10.1038/nprot.2016.098 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Skene PJ, Henikoff S. An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. eLife. 2017;6:e21856. doi: 10.7554/elife.21856 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Skene PJ, Henikoff JG, Henikoff S. Targeted in situ genome-wide profiling with high efficiency for low cell numbers. Nat Protoc. 2018;13(5):1006–1019. doi: 10.1038/nprot.2018.015 [DOI] [PubMed] [Google Scholar]
- 51.Chavez A, Scheiman J, Vora S, Pruitt BW, Tuttle M, Iyer PRE. Highly efficient Cas9-mediated transcriptional programming. Nat Methods. 2015;12(4):326–328. doi: 10.1038/nmeth.3312 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Hiesberger T, Bai Y, Shao X, McNally BT, Sinclair AM, Tian X. Mutation of hepatocyte nuclear factor–1β inhibits Pkhd1 gene expression and produces renal cysts in mice. J Clin Invest. 2004;113(6):814–825. doi: 10.1172/jci200420083 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Battle MA, Konopka G, Parviz F, Gaggl AL, Yang C, Sladek FM. Hepatocyte nuclear factor 4α orchestrates expression of cell adhesion proteins during the epithelial transformation of the developing liver. Proc Natl Acad Sci. 2006;103(22):8419–8424. doi: 10.1073/pnas.0600246103 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.DeLaForest A, Nagaoka M, Si-Tayeb K, Noto FK, Konopka G, Battle MA. HNF4A is essential for specification of hepatic progenitors from human pluripotent stem cells. Development. 2011;138(19):4143–4153. doi: 10.1242/dev.062547 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Yahoo N, Pournasr B, Rostamzadeh J, Fathi F. Forced expression of Hnf4a induces hepatic gene activation through directed differentiation. Biochem Biophysical Res Commun. 2016;476(4):313–318. doi: 10.1016/j.bbrc.2016.05.119 [DOI] [PubMed] [Google Scholar]
- 56.Hazelbaker DZ, Beccard A, Angelini G, Mazzucato P, Messana A, Lam D. A multiplexed gRNA piggyBac transposon system facilitates efficient induction of CRISPRi and CRISPRa in human pluripotent stem cells. Sci Rep. 2020;10(1):635. doi: 10.1038/s41598-020-57500-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Drewes T, Senkel S, Holewa B, Ryffel GU. Human hepatocyte nuclear factor 4 isoforms are encoded by distinct and differentially expressed genes. Mol Cell Biol. 1996;16(3):925–931. doi: 10.1128/mcb.16.3.925 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Chen L, Toke NH, Luo S, Vasoya RP, Fullem RL, Parthasarathy A. A reinforcing HNF4–SMAD4 feed-forward module stabilizes enterocyte identity. Nat Genet. 2019;51(5):777–785. doi: 10.1038/s41588-019-0384-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.La Russa MF, Qi LS. The new state of the art: Cas9 for gene activation and repression. Mol Cell Biol. 2015;35(22):3800–3809. doi: 10.1128/mcb.00512-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26(1):139–140. doi: 10.1093/bioinformatics/btp616 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Huang DW, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4(1):44–57. doi: 10.1038/nprot.2008.211 [DOI] [PubMed] [Google Scholar]
- 62.Yu G, Wang L-G, He Q-Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics. 2015;31(14):2382–2383. doi: 10.1093/bioinformatics/btv145 [DOI] [PubMed] [Google Scholar]
- 63.Croft D, O’Kelly G, Wu G, Haw R, Gillespie M, Matthews L. Reactome: a database of reactions, pathways and biological processes. Nucleic Acids Res. 2011;39:D691–D697. doi: 10.1093/nar/gkq1018 [DOI] [PMC free article] [PubMed] [Google Scholar]