Summary
Fruit ripening and disease resistance are two essential biological processes for quality formation and maintenance. DNA methylation, in the form of 5‐methylcytosine (5mC), has been elucidated to modulate fruit ripening, but its role in regulating fruit disease resistance remains poorly understood. In this study, we show that mutation of SlDML2, the DNA demethylase gene essential for fruit ripening, affects multiple developmental processes of tomato besides fruit ripening, including seed germination, leaf length and width and flower branching. Intriguingly, loss of SlDML2 function decreased the resistance of tomato fruits against the necrotrophic fungal pathogen Botrytis cinerea. Comparative transcriptomic analysis revealed an obvious transcriptome reprogramming caused by SlDML2 mutation during B. cinerea invasion. Among the thousands of differentially expressed genes, SlβCA3 encoding a β‐carbonic anhydrase and SlFAD3 encoding a ω‐3 fatty acid desaturase were demonstrated to be transcriptionally activated by SlDML2‐mediated DNA demethylation and positively regulate tomato resistance to B. cinerea probably in the same genetic pathway with SlDML2. We further show that the pericarp tissue surrounding B. cinerea infection exhibited a delay in ripening with singnificant decrease in expression of ripening genes that are targeted by SlDML2 and increase in expression of SlβCA3 and SlFAD3. Taken together, our results uncover an essential layer of gene regulation mediated by DNA methylation upon B. cinerea infection and raise the possible that the DNA demethylase gene SlDML2, as a multifunctional gene, participates in modulating the trade‐off between fruit ripening and disease resistance.
Keywords: DNA methylation, SlDML2, defence response, transcriptome reprogramming, Botrytis cinerea, tomato
Introduction
Botrytis cinerea, the pathogenic agent of grey mould, annually causes dramatical agriculture losses due to its widespread and necrotrophic infection characteristics (Williamson et al., 2007; Zhang et al., 2021). More than 200 economically important crop species are susceptible to B. cinerea worldwide (Elad et al., 2007; Liu et al., 2021). B. cinerea generally secretes diverse toxic substances to kill the host cells for acquiring adequate nutrition during the pathogenic process (Cheung et al., 2020). Utilization of chemical fungicides represents an efficient strategy at present for controlling fungal diseases, but their overuse harbours potential threats to human health (Jankowska et al., 2016). Therefore, it is currently imperative to explore more secure alternatives to effectively control grey mould, and the information concerning the molecular mechanisms of plant disease resistance to B. cinerea will provide us new perspectives.
Tomato, one of the most important horticultural crops, is easily accessible for B. cinerea invasion, especially the fruits at the postharvest stage that undergo continuous ripening process (Cantu et al., 2008, 2009; Prusky et al., 2013). The interaction between tomato fruits and B. cinerea is generally utilized as a model pathosystem for dissecting the defence mechanisms employed by host cells (Min et al., 2020; Silva et al., 2023; Wang et al., 2017b). Accumulating evidences suggest that tomato cuticles and cell walls are the important physical barriers, and phytoalexins, such as phenolics and flavonoids, act as antimicrobial chemical components to defence B. cinerea infection (Xu et al., 2022). Phytohormones jasmonic acid (JA) and methyl jasmonate (MJ), the endogenous signal molecules, also play a critical role in regulating tomato resistance to B. cinerea disease, partially by inducing the biosynthesis of phytoalexins or activating the transcription of downstream defence genes (Huang et al., 2022; Reyes‐Diaz et al., 2016; Shu et al., 2021). In addition, the resistance of tomato fruits towards B. cinerea is dramatically influenced by fruit maturity, and one direct evidence is that ripe red fruits are more susceptible to B. cinerea than unripe green fruits (Cantu et al., 2009; Prusky et al., 2013; Silva et al., 2023). This diverse susceptibility is generally thought to be self‐interested, since the fruits need to defence external invasions for surviving until seed maturation, and then spread mature seeds with the aid of enhanced fungal infection (Cantu et al., 2009). Thus, fruit ripening and disease resistance are two tightly interconnected biological processes, although the mechanisms underlying their interplays remain elusive. Understanding in the molecular link between fruit ripening and disease resistance has great potential for controlling fungal invasion and maintaining fruit quality.
DNA methylation, in the form of 5‐methylcytosine (5mC), is a broad‐spectrum epigenetic modification, mainly occurring in sequences of gene promoters, gene bodies and transposable elements (Bartels et al., 2018). DNA methylation modification generally facilitates transcriptional inactivation through remodelling chromatin structure or regulating the accessibility of transcriptional regulators to associated gene sequences (Zhang et al., 2018). DNA methylation is subjected to the synergistic action of DNA methyltransferases and demethylases, thereby exhibits a dynamic status in vivo and participates in the regulation of many important biological processes, including plant stress resistance, genome management, and developmental process (Bartels et al., 2018; Chen et al., 2020; Tang et al., 2020). More recently, DNA methylation was also demonstrated to regulate plant–pathogen interactions in agricultural and horticultural crops, including Magnaporthe oryzae‐infected rice, Fusarium graminearum‐infected maize, Xanthomonas phaseoli‐infected cassava and Fusarium. oxysporum‐infected banana (Deng et al., 2017; Luo et al., 2016; Tirnaz and Batley, 2019; Veley et al., 2023; Wang et al., 2017a). In addition, DNA methylation functionally modulates the disease resistance of woody plant mulberry towards B. cinerea, and the DNA methyltransferase gene MnMET1 plays a negative regulatory role in B. cinerea invasion through impeding the transcription of defence genes by DNA hypermethylation (Xin et al., 2021). However, whether DNA methylation participates in regulating the disease resistance of horticultural crops including tomato to B. cinerea has not been defined. Moreover, the trade‐off between DNA methylation‐mediated disease resistance and fruit ripening remains to be determined.
In tomato, there are four DEMETER‐like (DML) DNA demethylase genes, i.e. SlDML1‐SlDML4 (Liu et al., 2015). SlDML2, the homologue of Arabidopsis Repressor of Silencing 1 (AtROS1) encoding a 5mC DNA glycosylase, has been well defined as a pivotal ripening gene for its significant functions in facilitating the expression of ripening‐related genes through active DNA demethylation (Gao et al., 2022; Lang et al., 2017; Li et al., 2020; Zhou et al., 2019). In the present study, we investigated the functional role of SlDML2 in the pathogenic process of B. cinerea and found that fruits and leaves of the sldml2 mutants generated by CRISPR/Cas9 gene editing system exhibited increased susceptibility to B. cinerea compared with those of the wild‐type. The comparative transcriptomic analysis between wild‐type and sldml2 mutant fruits after B. cinerea inoculation revealed that SlDML2 is required for the expression of plenty of defence‐related genes, including the CO2‐binding protein gene SlβCA3 encoding a β‐carbonic anhydrase and the JA biosynthetic gene SlFAD3 encoding a ω‐3 fatty acid desaturase (Domínguez et al., 2010; Hu et al., 2021; Yu et al., 2009). We provided evidences that both SlβCA3 and SlFAD3 undergo SlDML2‐mediated transcriptional activation and play a positive role against B. cinerea invasion probably in the same genetic pathway with SlDML2. Furthermore, we found that B. cinerea infection causes a delay in fruit ripening in the pericarp tissue surrounding disease regions, accompanied by the decreased expression of ripening genes targeted by SlDML2 and the increased expression of SlβCA3 and SlFAD3. Our findings uncover the dual roles of SlDML2 in regulating fruit ripening and defence response to fungal pathogen and highlight the possible function of SlDML2 as a molecular link between ripening and disease resistance in fruit.
Results
SlDML2 modulates multiple developmental processes besides fruit ripening
Tomato DNA demethylase SlDML2 has been considered primarily to modulate fruit ripening through activating 5mC demethylation in numerous ripening genes (Lang et al., 2017; Liu et al., 2015). Recently, emerging evidences from DNA methylomes indicate that SlDML2 potentially targets thousands of genes relevant to distinct biological processes, rather than merely those of fruit ripening (Gao et al., 2022; Lang et al., 2017). To further explore the physiological roles of SlDML2, we first generated its loss‐of‐function mutant by CRISPR/Cas9‐mediated gene editing. Three single guide RNAs (sgRNAs) with different target sequences (i.e., T1, T2 and T3) were designed to specifically target the first exon of SlDML2 (Figure 1a). The sgRNAs were introduced into the binary vector pYLCRISPR/Cas9Pubi‐H that contains a Cas9 expression cassette, and then transformed into leaf explants of wild‐type tomato (Solanum lycopersicum cv. Ailsa Craig) by using Agrobacterium infection (Fillatti et al., 1987). Following the standard tissue culture, a total of six transgenic lines were obtained. Among them, three homozygous mutant lines that carry 2‐bp or 5‐bp deletion were identified through genotyping mediated by direct sequencing of PCR products (Figure 1a). Both the 2‐bp and 5‐bp deletion mutations, which were predicted to cause premature stop codons in the first exon of SlDML2, occurred in the DNA sequence targeted by the first sgRNA. The three homozygous mutants, namely sldml2‐3, sldml2‐4 and sldml2‐5, were predicted to produce short peptides with a length of 30–40 amino acids, respectively (Figure S1).
Figure 1.
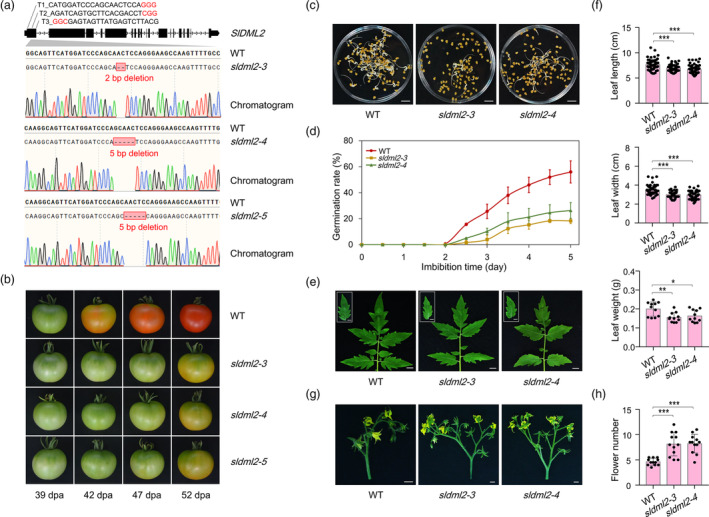
SlDML2 participates in modulating multiple developmental processes. (a) Genotyping of sldml2‐3, sldml2‐4, and sldml2‐5 mutants generated by CRISPR/Cas9‐mediated gene editing. Three single guide RNAs (sgRNAs) containing different target sequences (T1, T2, and T3) were designed to specifically target the first exon of SlDML2. The red letters indicate the protospacer adjacent motif (PAM). The sldml2‐3 mutant contains a homozygous 2‐bp deletion, and the sldml2‐4 and sldml2‐5 mutants contain a homozygous 5‐bp deletion caused by target T1. (b) Phenotypes of fruit ripening in sldml2 mutants. Representative photographs of the wild‐type (WT), sldml2‐3, sldml2‐4, and sldml2‐5 fruits at 39, 42, 47, and 52 dpa are shown. dpa, days post‐anthesis. (c) Representative photographs and (d) germination rates of the WT, sldml2‐3, and sldml2‐4 seeds. For each germination experiment, one‐hundred mature seeds were cultured with deionized water under long‐day conditions (16 h light/8 h dark, 25 °C). The gemmiparous seeds were photographed at the fourth day after imbibition. Data are presented as mean ± standard deviation (n = 3). (e) Representative photographs and (f) length, width, and weight of the WT, sldml2‐3, and sldml2‐4 leaves. Six‐week‐old leaves were collected, photographed, and measured. (g) Representative photographs and (h) flower number of the WT, sldml2‐3, and sldml2‐4 inflorescences. The first inflorescences were collected and photographed, and the flower number in each inflorescence was measured. In (c), (e), and (g), scale bar = 1 cm. In (f and h), asterisks indicate significant differences (*P < 0.05, **P < 0.01, ***P < 0.001; Student's t test).
To determine the ripening phenotype, fruits of the wild‐type and sldml2 mutants (sldml2‐3, sldml2‐4 and sldml2‐5) at the second generation were harvested at 39, 42, 47 and 52 days post‐anthesis (dpa), when the wild‐type fruits reach to mature green (MG), breaker (Br), orange ripe (OR) and red ripe (RR) stages, respectively. Compared to the wild‐type, the sldml2 mutants showed a similar and obvious delay in ripening, especially at 52 dpa, when fruits of the wild‐type exhibited a homogenous red surface, whereas those of the sldml2 mutants were only just starting to change colour (Figure 1b). This ripening phenotype is consistent with that of previously reported sldml2 mutants in Micro‐Tom background (Lang et al., 2017), both indicating that SlDML2 is necessary for normal fruit ripening. Fruits of the sldml2 mutants displayed a homogenous red colour at approximately 90 dpa, similar to those of the wild‐type (Figure S2), suggesting that fruits of the sldml2 mutants harbour the capacity to reach red ripe.
We next sought to determine whether SlDML2 functions in other developmental processes besides fruit ripening. As expected, tomato seeds, leaves and flowers exhibited abnormal developmental phenotypes under the disruption of SlDML2 gene. The germination rate of seeds from the sldml2 mutants decreased obviously under long‐day conditions (16 h light/8 h dark, 25 °C) compared to those from the wild‐type (Figure 1c,d), although no significant difference was detected in seed length, width and weight (Figure S3a,b). The leaves of sldml2 mutants were smaller than those of the wild‐type with shorter length and width, and lighter weight (Figure 1e,f). The flower branching of sldml2 mutants had more floral organs than those of the wild‐type (Figure 1g,h), indicating a floriferous phenomenon caused by SlDML2 mutation. Quantitative RT‐PCR analysis showed that SlDML2 gene was highly expressed in tomato seeds, leaves and flowers besides the fruits (Figure S4). These results revealed the functional diversity of SlDML2 in modulating multiple developmental processes in tomato.
SlDML2 positively regulates resistance of tomato fruits to fungal pathogens B. cinerea
Having observed the developmental phenotypes, we next investigated whether SlDML2 participates in modulating tomato resistance to B. cinerea invasion, which is generally known to cause serious yield and quality declines in horticultural crops including tomato (Blanco‐Ulate et al., 2013; Tian et al., 2016). Fruits of the wild‐type and sldml2 mutants harvested at 39 and 90 dpa were inoculated with B. cinerea stain B05.10, and the lesion diameters were measured at the second to third day post‐inoculation (dpi) when obvious soft rot symptoms appeared. It was shown that green and red fruits of the sldml2 mutants (at 39 and 90 dpa, respectively) exhibited significantly larger lesion diameters than the wild‐type (Figure 2a–d). We concurrently detected the leaf susceptibility to B. cinerea infection and found that leaves of the sldml2 mutants also displayed larger lesion diameters by comparison with the wild‐type (Figure 2e,f). These results indicate that the disease resistance was impaired in the sldml2 mutants, suggesting that SlDML2 positively regulates the resistance response of tomato to B. cinerea.
Figure 2.
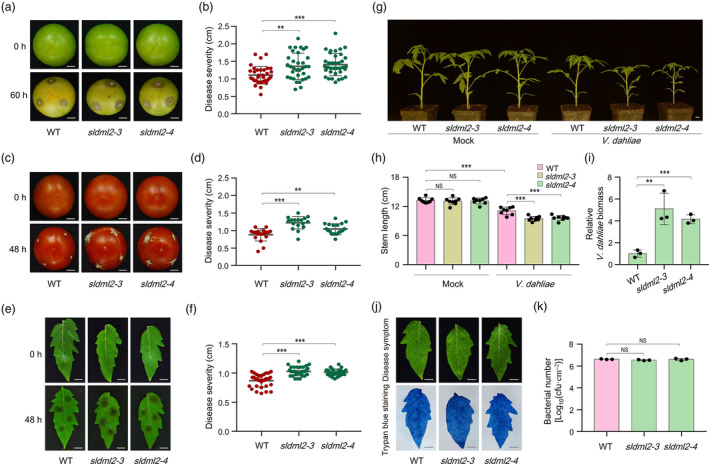
The sldml2 mutants exhibited decreased resistance to fungal pathogens B. cinerea and V. dahliae. (a) Disease symptoms and (b) lesion diameters on the detached wild‐type (WT), sldml2‐3, and sldml2‐4 fruits inoculated with B. cinerea at 39 days post‐anthesis (dpa). The disease symptom was observed after 60 h. (c) Disease symptoms and (d) lesion diameters on the detached WT, sldml2‐3, and sldml2‐4 fruits inoculated with B. cinerea at 90 dpa. The disease symptom was observed after 48 h. (e) Disease symptoms and (f) lesion diameters on 4‐week‐old detached leaves of WT, sldml2‐3, and sldml2‐4 inoculated with B. cinerea. The disease symptom was observed after 48 h. (g) Representative photographs and (h) stem lengths of the WT, sldml2‐3, and sldml2‐4 seedlings that were either inoculated with V. dahliae or mock‐inoculated for 18 days. (i) Relative V. dahliae biomass in stems of the WT, sldml2‐3, and sldml2‐4 seedlings that were inoculated with V. dahliae for 18 days. The amounts of V. dahliae GADPH gene were determined by quantitative RT‐PCR analysis, and the tomato RuBisCo gene was used as an internal control. (j) Disease symptoms and trypan blue staining and (k) bacterial biomass in WT, sldml2‐3, and sldml2‐4 leaves that were inoculated with Pst DC3000 for 3 days. In (a), (c), (e), (g), and (j), scale bar = 1 cm. In (b, d, f, h, i and k), asterisks indicate significant differences (**P < 0.01, ***P < 0.001; Student's t test). NS, no significance.
We next asked, besides the necrotrophic pathogen B. cinerea, whether SlDML2 regulates tomato resistance to biotrophic pathogens, such as Verticillium dahliae and Pseudomonas syringae pv. tomato (Pst) DC3000. V. dahliae is a soil‐borne fungal pathogen responsible for severe losses of biomass, yield and quality in plenty of economically important crops including tomato and cotton (Zhang et al., 2023), while P. syringae represents a model biotrophic bacterial pathogen that has been extensively investigated in tomato and Arabidopsis (Xin and He, 2013). Three‐week‐old tomato seedlings of wild‐type and sldml2 mutants that display non‐differential stem lengths were inoculated with V. dahliae and the disease response was observed at 18 days post‐inoculation (dpi) (Figure S5a,b). Compared to the mock group, the V. dahliae‐infected tomato seedlings exhibited a dwarf phenotype in both wild‐type and sldml2 mutants, implying an obvious growth inhibition caused by V. dahliae invasion (Figure 2g,h). Importantly, the seedlings of V. dahliae‐infected sldml2 mutants displayed significantly shorter stem length compared to the V. dahliae‐infected wild‐type, indicating a more serious growth inhibition after V. dahliae invasion (Figure 2g,h). Detection of V. dahliae biomass by relative quantification of V. dahliae GADPH gene in tomato stems revealed an obvious increase in GADPH expression (~4–6‐fold) in the sldml2 mutants compared to the wild‐type (Figure 2i). These results support that SlDML2 also positively regulates tomato resistance to the biotrophic pathogen V. dahliae.
To explore whether SlDML2 regulates the resistance of tomato to Pst DC3000, 4‐week‐old leaves of the wild‐type and sldml2 mutants were inoculated with Pst DC3000, and the disease response was measured at 3 dpi. Similar disease symptom and cell death were observed in the wild‐type and sldml2 mutant leaves (Figure 2j). Bacterial growth assay demonstrated no significant difference in bacterial number between leaves of the wild‐type and sldml2 mutants (Figure 2k), suggesting that SlDML2 is dispensable for tomato resistance to the biotrophic bacterial pathogen Pst DC3000. Together, these data suggest that SlDML2 confers resistance of tomato to the necrotrophic fungal pathogen B. cinerea and biotrophic fungal pathogen V. dahliae, but not to the biotrophic bacterial pathogen Pst DC3000.
SlDML2 mutation causes transcriptome reprogramming in tomato fruits during B. cinerea infection
To decipher the molecular basis underlying SlDML2‐mediated defence response of tomato fruits to B. cinerea, a comparative transcriptomic analysis was performed using B. cinerea‐infected tomato fruits of the sldml2‐3 mutant and wild‐type at 39 dpa. High Pearson correlation coefficients were observed between biological replicates, representing the excellent repeatability (Figure S6). A total of 2310 differentially expressed genes (DEGs; fold change ≥2 and adjusted P value <0.05) were identified, of which 1431 were up‐regulated and 879 were down‐regulated in fruits of the sldml2‐3 mutant compared to the wild‐type (Figure 3a,b; Tables S1 and S2). This indicates an obvious transcriptome reprogramming caused by SlDML2 mutation in the process of B. cinerea infection.
Figure 3.

SlDML2 mutation causes transcriptome reprogramming during B. cinerea infection. (a) Volcano plot showing up‐regulated (red) and down‐regulated (blue) genes in the sldml2‐3 mutant compared to the wild‐type (WT). (b) Heat map of differentially expressed genes in the sldml2‐3 mutant compared with the wild‐type. FPKM, fragments per kilobase of exon per million mapped fragments. (c) Gene ontology (GO) enrichment for the differentially expressed genes in the sldml2‐3 mutant compared with the WT. The top 20 catalogues in biological process with the most significant P value were shown. (d) KEGG analysis for the differentially expressed genes in the sldml2‐3 mutant compared with the WT. The top 20 pathways with the most significant P value were shown. The differentially expressed genes were analysed by three independent RNA‐seq experiments in fruits of the WT and sldml2‐3 mutant after B. cinerea inoculation for 48 h.
Gene ontology (GO) enrichment analysis revealed that those DEGs were highly enriched in items related to defence response to biotic and abiotic stresses (Figure 3c; Table S3). Importantly, several items associated with fungal pathogen infection are included in the top 20 catalogues with the most significant P value, including ‘defense response to fungus’, ‘response to chitin’, ‘response to wounding’, ‘response to oxidative stress’ and ‘cell wall organization’ (Figure 3c). KEGG analysis revealed that the DEGs are involved in multiple pathways, in addition to those closely associated with metabolism (Figure 3d). Among them, the ‘plant–pathogen interaction’ pathway contains the largest number of genes with high significance (Figure 3d). These data uncovered the involvement of SlDML2 in the transcriptional regulation of defence genes related to tomato disease resistance.
SlDML2‐mediated DNA demethylation facilitates SlβCA3 expression
We subsequently focused on genes with significantly lower expression level in B. cinerea‐infected sldml2‐3 fruits for the reason that loss‐of‐function of SlDML2 is likely to induce DNA hypermethylation, which is generally thought to cause transcriptional inactivation. Comparison of those down‐regulated genes (Table S2) with genes containing hypermethylated DMR (differentially methylated region) identified in the sldml2 mutants (Lang et al., 2017) revealed that, among the 879 down‐regulated genes, 528 (~60%) were hypermethylated under SlDML2 disruption (Table S4), suggesting a critical transcription regulation of those genes by SlDML2‐mediated DNA demethylation.
Among the down‐regulated genes identified in the sldml2‐3 mutant during B. cinerea invasion, a gene encoding the β‐type carbonic anhydrase, namely SlβCA3 (Solyc02g067750) (Table S2), was previously reported to positively regulate tomato resistance to the virulent pathogen Pst DC3000 (Hu et al., 2021). SlβCA3 gene exhibited lower transcription level in fruits or leaves of the sldml2‐3 and sldml2‐4 mutants than the wild‐type after B. cinerea inoculation for 2 days, as revealed by quantitative RT‐PCR (Figure 4a,b). The transcription level of SlβCA3 was also obviously decreased in fruits of the sldml2 mutants under normal developmental process (Figure S7a). These results indicate that SlDML2 positively regulates SlβCA3 expression.
Figure 4.

SlβCA3 is involved in the defence response to B. cinerea. (a) Transcription levels of the tomato β‐type carbonic anhydrase gene SlβCA3 in fruits of the wild‐type (WT) and sldml2 mutants after B. cinerea inoculation for 48 h. (b) Transcription levels of the SlβCA3 gene in leaves of the WT and sldml2 mutants after B. cinerea inoculation for 48 h. (c) Transcription levels of the SlβCA3 gene in WT fruits with or without B. cinerea inoculation at the indicated time. (d) Transcription levels of the SlβCA3 gene in WT leaves with or without B. cinerea inoculation at the indicated time. In (a–d), the SlβCA3 transcription levels were determined by quantitative RT‐PCR analysis, and the tomato SlUBI3 gene was used as an internal control. Asterisks indicate significant differences (*P < 0.05, **P < 0.01, ***P < 0.001; Student's t test). NS, no significance. (e) Transient expression of SlβCA3‐HA protein in N. benthamiana leaves. Total protein was extracted at indicated time after agroinfiltration and then submitted to immunoblot with anti‐HA antibody. Equal loading was confirmed by using the tomato Actin as an internal control. (f) B. cinerea disease symptoms and lesion diameters on detached N. benthamiana leaves with or without the co‐expression of SlβCA3‐HA. Asterisks indicate significant differences (**P < 0.01; Student's t test). Scale bar = 1 cm.
We next investigated whether SlβCA3 functions in modulating B. cinerea disease resistance. SlβCA3 expression was strongly induced in fruits and leaves of the wild‐type tomato following inoculation with B. cinerea (Figure 4c,d), suggesting it is a B. cinerea‐responsive gene. We then transiently expressed SlβCA3 protein in Nicotiana benthamiana leaves and inoculated B. cinerea at the 36 h post‐agroinfiltration when the protein was substantially expressed (Figure 4e). Analysis of the disease symptoms at the second day post‐inoculation showed that the lesion diameters were significantly decreased upon the expression of SlβCA3 protein (Figure 4f), indicating that SlβCA3 has the ability to restrict B. cinerea invasion. We also evaluated the function of SlβCA3 in modulating resistance to V. dahliae and Pst DC3000 and found that SlβCA3 showed no effect on V. dahliae (Figure S8), but conferred resistance to Pst DC3000 (Figure S9a) as previously reported (Hu et al., 2021).
According to the published DNA methylomes in sldml2 mutants (Lang et al., 2017), no DMR exists in the SlβCA3 promoter region (~2000 bp upstream of the start codon) when SlDML2 was disrupted (Figure S10a). By contrast, an obvious DMR was identified in the first intron of SlβCA3, in which the 5mC levels of 16 representative cytosines were obviously increased in the sldml2 mutant (Figure 5a). Bisulphite sequencing revealed that 11 of the 16 representative cytosines displayed higher 5mC levels in fruits of the sldml2 mutants (sldml2‐3 and sldml2‐4) than the wild‐type (Figure 5b), confirming the existence of DMR in the intronic region. The increase in 5mC levels of the intronic DMR was also observed in the sldml2 mutants (sldml2‐3 and sldml2‐4) after B. cinerea infection as revealed by McrBC‐PCR assay (Figure 5c). ChIP‐qPCR assay using the anti‐SlDML2 polyclonal antibody showed that SlDML2 protein can bind to the intronic DMR of SlβCA3 gene (Figure 5d), indicating that SlβCA3 is the direct target of SlDML2, which mediates 5mC level of SlβCA3 in this region.
Figure 5.

SlβCA3 is transcriptionally regulated by SlDML2‐mediated DNA demethylation. (a) DNA methylation (5mC) level of the SlβCA3 gene body in fruits of the wild‐type (WT) and sldml2 mutant at indicated developmental stages. The 5mC level was analysed by using the published DNA methylome database (Lang et al., 2017). Each vertical bar represents an 5mC and the bar height indicates 5mC level. The differentially methylated region (DMR) in the first intron of SlβCA3 gene was indicated by a shadow box. 5mC levels of representative cytosines in the DMR were shown with pie charts, and the numbers indicate the positions relative to the start codon. dpa, days post‐anthesis. (b) Bisulphite sequencing showing 5mC levels of the representative cytosines shown in (a) in fruits of the wild‐type and sldml2 mutants (sldml2‐3 and sldml2‐4) at 46 dpa. (c) 5mC level of the DMR of SlβCA3 gene in fruits of the WT and sldml2 mutants after B. cinerea inoculation for 48 h. (d) ChIP‐qPCR assay showing that SlDML2 binds to the DMR in the first intron of SlBCA3 gene. The wild‐type fruits at 42 dpa were subjected to the immunoprecipitation with the anti‐SlDML2 polyclonal antibody, and the pre‐immune IgG was used as a control. The first intron structure of SlβCA3 is shown, and the red line indicates the region for PCR. (e) Transcription levels of the SlβCA3 gene in N. benthamiana leaves with or without the co‐expression of SlDML2‐HA. The sequence of SlβCA3 gene body containing all exons and introns was cloned into the dual‐luciferase reporter vector for transcription activity assay under the driving of its native promoter. The renilla luciferase gene RLUC was used as an internal control. (f) 5mC level of the DMR of SlβCA3 gene in N. benthamiana leaves with or without the co‐expression of SlDML2‐HA. (g) Transcription levels of the mutated SlβCA3 gene under the drive of its native promoter in N. benthamiana leaves with or without the co‐expression of SlDML2‐HA. The representative cytosines shown in (a) were mutated to adenine or thymine. NS, no significance. In (c) and (f), 5mC level was revealed by McrBC‐PCR assay. A total of 0.4 μg genomic DNA was digested by McrBC enzyme with GTP (+GTP), or without GTP (−GTP) as a negative control. An unmethylated region in the promoter of tomato PME gene was used as an internal control. In (d and e), asterisks indicate significant differences (**P < 0.01; Student's t test).
We then adopted a dual‐luciferase reporter system to determine whether SlβCA3 is transcriptionally regulated by SlDML2‐mediated DNA demethylation (Han et al., 2016). The sequence fragment of SlβCA3 promoter was cloned into the dual‐luciferase reporter vector pGreen II‐0800‐LUC to drive the expression of promoterless firefly luciferase (FLUC) reporter gene (Figure S10b). The transcription level of FLUC gene was relatively calculated by normalizing against the renilla luciferase (RLUC) reference gene driven by the CaMV 35S promoter. Quantitative RT‐PCR analysis showed that the FLUC transcription level was not increased under the expression of SlDML2‐HA fusion protein (Figure S10c,d), which is consistent with the observation that no DMR occurred in the SlβCA3 promoter region under SlDML2 disruption, suggesting that the SlβCA3 promoter is indeed free from SlDML2‐mediated DNA demethylation.
We then asked whether SlDML2 regulates SlβCA3 transcription through modulating 5mC level in the intronic DMR. The sequence of SlβCA3 gene body containing all exons and introns was cloned into the dual‐luciferase reporter vector to replace the FLUC gene sequence for transcription activity assay under the driving of its native promoter. The SlβCA3 transcription level increased significantly when SlDML2 was co‐expressed (Figure 5e), concomitant with a decline in 5mC level of the intronic DMR (Figure 5f). By contrast, SlDML2 co‐expression did not induce a significant increase in SlβCA3 transcription when those representative cytosines in the intronic DMR were mutated to adenine or thymine (Figure 5g), indicating that SlβCA3 could be transcriptionally regulated by SlDML2‐mediated DNA demethylation.
Taken together, these data suggest that SlβCA3, which could be transcriptionally activated by SlDML2 through active DNA demethylation, functions as a defence gene towards B. cinerea invasion, thus plays an important role in SlDML2‐mediated tomato resistance to B. cinerea.
SlDML2‐mediated DNA demethylation regulates the expression of JA biosynthetic genes
Previous studies have shown that the phytohormone jasmonic acid (JA) plays an essential role in activating plant defence responses against fungal pathogens (Du et al., 2017). During B. cinerea infection, enhanced JA biosynthesis and signalling transduction induce a series of defence responses, which finally confers increased resistance of tomato fruits to B. cinerea (Huang et al., 2022; Martel et al., 2015; Reyes‐Diaz et al., 2016; Shu et al., 2021). To investigate whether JA pathway is involved in the elevated susceptibility of sldml2 mutants to B. cinerea infection, the changes in transcription levels of JA biosynthetic genes were analysed based on our RNA‐seq data. JA biosynthesis is synergistically controlled by multiple gene families that encode distinct biosynthetic enzymes (Figure S11a). Heat map showed that some genes within those families exhibited lower transcription level in the sldml2‐3 mutant than the wild‐type during B. cinerea invasion, including the ω‐3 desaturase 3 (SlFAD3) (Domínguez et al., 2010; Yu et al., 2009), the lipoxygenase B/C/D (SlLOXB/C/D) (Zhu‐Salzman et al., 2008), the allene oxide synthase 1 (SlAOS1) (Sivasankar et al., 2000), the allene oxide cyclase (SlAOC) (Ziegler et al., 2000) and the 12‐oxophytodienoate reductase 3 (SlOPR3) (Strassner et al., 2002) (Figure S11a). The differential gene expression was further confirmed by quantitative RT‐PCR analysis (Figure S11b), indicating the involvement of SlDML2 in regulating the expression of JA biosynthetic genes.
Since the JA biosynthetic gene SlFAD3, which encodes one of the ω‐3 fatty acid desaturases that catalyse the conversion of linoleic acid (18:2) to linolenic acid (18:3) (Domínguez et al., 2010; Yu et al., 2009), exhibited a dramatically decreased expression when SlDML2 was mutated (Figure S11b), we speculate that SlFAD3 may be directly regulated by SlDML2‐mediated DNA demethylation and involved in the elevated susceptibility of sldml2 mutants to B. cinerea infection. Compared to the wild‐type, the SlFAD3 transcription level was obviously lower in fruits and leaves of the sldml2‐3 and sldml2‐4 mutants after B. cinerea infection (Figure 6a,b). Moreover, the expression of SlFAD3 decreased obviously in fruits of the sldml2 mutants under normal developmental process (Figure S7b), indicating that SlDML2 positively regulates SlFAD3 expression.
Figure 6.

SlFAD3 is transcriptionally regulated by SlDML2‐mediated DNA demethylation. (a) Transcription levels of the tomato ω‐3 fatty acid desaturase gene SlFAD3 in fruits of the wild‐type (WT) and sldml2 mutants after B. cinerea inoculation for 48 h. (b) Transcription levels of the SlFAD3 gene in leaves of the WT and sldml2 mutants after B. cinerea inoculation for 48 h. (c) Transient expression of SlFAD3‐HA protein in N. benthamiana leaves. Total protein was extracted at indicated time after agroinfiltration and then submitted to immunoblot with anti‐HA antibody. Equal loading was confirmed by using the tomato Actin as an internal control. (d) B. cinerea disease symptoms and lesion diameters on detached N. benthamiana leaves with or without the co‐expression of SlFAD3‐HA. Scale bar = 1 cm. (e) DNA methylation (5mC) levels of the SlFAD3 promoter in fruits of the WT and sldml2 mutant at indicated developmental stages. The 5mC level was analysed by using the published DNA methylome database (Lang et al., 2017). Each vertical bar represents a 5mC and the bar height indicates 5mC level. The differentially methylated region (DMR) in the SlFAD3 promoter was indicated by a shadow box. 5mC levels of the representative cytosines in the DMR were shown with pie charts, and the numbers indicate the positions relative to the start codon. dpa, days post‐anthesis. (f) Bisulphite sequencing showing 5mC levels of the representative cytosines shown in (e) in fruits of the wild‐type and sldml2 mutants (sldml2‐3 and sldml2‐4) at 46 dpa. (g) 5mC level of the DMR of SlFAD3 promoter in fruits of the WT and sldml2 mutants after B. cinerea inoculation for 48 h. (h) ChIP‐qPCR assay showing that SlDML2 binds to the DMR in the promoter of SlFAD3. The wild‐type fruits at 42 dpa were subjected to the immunoprecipitation with the anti‐SlDML2 polyclonal antibody, and the pre‐immune IgG was used as a control. The promoter structure of SlFAD3 is shown, and the red line indicates the region for PCR. (i) Relative firefly luciferase (FLUC) activity derived by the SlFAD3 promoter in N. benthamiana leaves with or without the co‐expression of SlDML2‐HA. The representative image was shown. The FLUC activity was normalized against the renilla luciferase (RLUC) activity, followed by normalization against the control. (j) Transcription levels of the FLUC gene under the drive of SlFAD3 promoter in N. benthamiana leaves with or without the co‐expression of SlDML2‐HA. The RLUC gene was used as an internal control. (k) 5mC level of the DMR of SlFAD3 promoter in N. benthamiana leaves with or without the co‐expression of SlDML2‐HA. (l) Transcription levels of the FLUC gene under the drive of mutated SlFAD3 promoter in N. benthamiana leaves with or without the co‐expression of SlDML2‐HA. The representative cytosines shown in (e) were mutated to adenine or thymine. Ns, no significance. In (g) and (k), total 5mC level was revealed by McrBC‐PCR assay. A total of 0.4 μg genomic DNA was digested by McrBC enzyme with GTP (+GTP), or without GTP (−GTP) as a negative control. An unmethylated region in the promoter of tomato PME gene was used as an internal control. In (a, b, d, h–j), asterisks indicate significant differences (*P < 0.05, ***P < 0.001; Student's t test).
We next investigated whether SlFAD3 functions in modulating B. cinerea disease resistance using the N. benthamiana expression system. SlFAD3 protein was transiently expressed in N. benthamiana leaves, and the B. cinerea was inoculated at the 36 h post‐agroinfiltration when the protein was obviously expressed (Figure 6c). Observation of the disease symptoms at the second day post‐inoculation revealed that the expression of SlFAD3 protein conferred the N. benthamiana leaves a significantly increased resistance to B. cinerea (Figure 6d), indicating that SlFAD3 harbours the capability of defensing B. cinerea invasion. We also analyzed the function of SlFAD3 in regulating resistance to V. dahliae (Figure S8) and Pst DC3000 (Figure S9b) but found that SlFAD3 was dispensable for resistance to both pathogens.
An obvious DMR was identified in the promoter region, rather than the gene body, of SlFAD3 in the sldml2 mutants, according to the published methylomes (Lang et al., 2017), in which the 5mC levels of 16 representative cytosines are obviously increased in the sldml2 mutant compared to the wild‐type (Figure 6e). Bisulphite sequencing showed that 13 of the 16 representative cytosines exhibited higher 5mC levels in fruits of the sldml2 mutants (sldml2‐3 and sldml2‐4) than the wild‐type, confirming that DMR exists in the promoter region of SlFAD3 (Figure 6f). The DMR also displayed higher 5mC level in fruits of the sldml2‐3 and sldml2‐4 mutants than those of the wild‐type during B. cinerea invasion (Figure 6g). ChIP‐qPCR assay revealed that SlDML2 protein can bind to the DMR in the promoter region of SlFAD3 gene (Figure 6h), indicating that SlFAD3 is the direct target of SlDML2, which mediates 5mC level of SlFAD3 in the promoter.
To determine whether SlFAD3 is transcriptionally regulated by SlDML2‐mediated DNA demethylation, the sequence fragment of SlFAD3 promoter (~2000 bp upstream of the start codon) was cloned into the dual‐luciferase reporter vector to drive the expression of FLUC reporter gene. The relative FLUC activity and FLUC transcription level were significantly increased when SlDML2 was co‐expressed with the reporter vector (Figure 6i,j), concomitant with a decline in 5mC level in the DMR of SlFAD3 promoter as determined by McrBC‐PCR assay (Figure 6k). Importantly, SlDML2 co‐expression only induced a slight increase in FLUC transcription (~1.3‐fold) when those representative cytosines in the DMR were mutated to adenine or thymine (Figure 6l). These results indicate that SlFAD3 could be transcriptionally regulated by SlDML2‐mediated DNA demethylation.
Collectively, these data suggest that SlFAD3, which could be transcriptionally activated by SlDML2 through active DNA demethylation, functions as a defence gene towards B. cinerea invasion and, thus plays a critical role in SlDML2‐mediated tomato resistance to B. cinerea.
SlβCA3 and SlFAD3 functions probably in the same genetic pathway with SlDML2 to regulate tomato resistance to B. cinerea
To further verify the roles of SlβCA3 and SlFAD3 in SlDML2‐mediated tomato resistance to B. cinerea, we silenced SlβCA3 or SlFAD3 gene in the sldml2 mutant (sldml2‐3) background by using virus‐induced gene silencing (VIGS), and then investigated whether this led to a decrease in resistance of the sldml2 mutant to B. cinerea. Quantitative RT‐PCR analysis showed that the transcription levels of SlβCA3 and SlFAD3 in the VIGS materials (sldml2/TRV2‐SlβCA3 and sldml2/TRV2‐SlFAD3) were decreased to almost 10% and 25% of the control group (sldml2/TRV2), respectively (Figure 7a). Meanwhile, RNA fragments transcribed from the virus vectors pTRV1, pTRV2, pTRV2‐SlβCA3 and pTRV2‐SlFAD3 were obviously detected in corresponding samples (Figure 7b). These results indicate that SlβCA3 and SlFAD3 gene were successfully silenced in the sldml2 mutant. The leaves of the sldml2 mutant displayed larger lesion diameters compared to the wild‐type, and the symptom severity was further increased in leaves of the sldml2/TRV2‐SlβCA3 or sldml2/TRV2‐SlFAD3 plants (Figure 7c,d), indicating an obviously additive effect on B. cinerea susceptibility induced by gene silencing of SlβCA3 or SlFAD3. Furthermore, we also overexpressed SlβCA3 or SlFAD3 gene in the sldml2 mutant and analysed its resistance to B. cinerea. Quantitative RT‐PCR and immunoblot assays showed that the two genes were successfully overexpressed in leaves of the sldml2 mutant (Figure 7e,f). Compared to the sldml2 mutant, the two overexpression materials (sldml2/35S pro :SlβCA3 and sldml2/35S pro :SlFAD3) exhibited significantly smaller lesion diameters, which were equally to those of the wild‐type, indicating that overexpression of SlβCA3 or SlFAD3 in the sldml2 mutant can recover the resistance to B. cinerea (Figure 7g,h). Taken together, these data suggest that SlβCA3 and SlFAD3 may regulate tomato resistance to B. cinerea in the same genetic pathway with SlDML2.
Figure 7.

SlDML2 functions in the same genetic pathway with SlβCA3 and SlFAD3 to regulate tomato resistance to B. cinerea. (a) Transcription levels of SlβCA3 and SlFAD3 in leaves of the sldml2 mutant after virus‐induced gene silencing (VIGS), as determined by quantitative RT‐PCR. The sldml2 mutant infiltrated with the empty vector pTRV2 was used as the control group. The tomato Actin gene was used as an internal control. (b) PCR amplification showing that the virus vectors pTRV1, pTRV2, pTRV2‐SlβCA3, and pTRV2‐SlFAD3 were successfully expressed in leaves of the wild‐type (WT) or sldml2 mutant. (c) Disease symptoms and (d) lesion diameters on leaves of the WT/TRV2, sldml2/TRV2, sldml2/TRV2‐SlβCA3, and sldml2/TRV2‐SlFAD3 inoculated with B. cinerea for 40 h. (e) Transcription levels of SlβCA3 and SlFAD3 in leaves of the sldml2 mutant after overexpression, as determined by quantitative RT‐PCR. The sldml2 mutant infiltrated with the empty vector pCambia1302‐HA was used as the control groups. The tomato Actin gene was used as an internal control. (f) Western blot showing that SlβCA3‐HA and SlFAD3‐HA fusion proteins were successfully expressed in leaves of sldml2/35S pro :SlβCA3 And sldml2/35S pro :SlFAD3. Total protein was extracted and submitted to immunoblot with anti‐HA antibody. Equal loading was confirmed by using the tomato Actin as an internal control. (g) Disease symptoms and (h) lesion diameters on leaves of the WT, sldml2, sldml2/35S pro :SlβCA3, And sldml2/35S pro :SlFAD3 Inoculated with B. cinerea for 40 h. In (a, d, e, and h), asterisks indicate significant differences (*P < 0.05, **P < 0.01, ***P < 0.001; Student's t test). NS, no significance. In (c and g), scale bar = 1 cm.
B. cinerea invasion causes a ripening delay in the pericarp tissues surrounding the infection region
At the fourth day after B. cinerea infection, we observed that the pericarp of wild‐type tomato fruits primarily exhibited a homogeneous orange colour, likely due to continual postharvest ripening. However, the pericarp tissue around B. cinerea disease region displayed green colour of different degree (Figure S12a), indicating a regional ripening delay caused by B. cinerea infection. We defined the pericarp tissue with a 5‐mm thickness surrounding disease regions as the response region of B. cinerea infection, and the pericarp tissue with a 5‐mm thickness around the response region as the control region (Figure 8a). Compared to the control region, the response region contained significantly decreased lycopene content (Figure 8b). Furthermore, transcription levels of several known ripening genes were obviously lower in the response region than in the control region (Figure 8c), and this decrease in gene expression was observed even in fruits that did not exhibit a macroscopic ripening delay in the response region (Figure S12b). These results suggest that B. cinerea invasion causes a ripening delay in the pericarp tissues surrounding the infection region.
Figure 8.

B. cinerea invasion causes a delay in ripening of pericarp tissues around the disease region. (a) Representative photograph of wild‐type tomato fruits after B. cinerea inoculation for 4 days. The pericarp tissue with a 5‐mm thickness surrounding disease regions was collected as “response region”, and the pericarp tissue with a 5‐mm thickness around the response region was collected as “control region”. (b) Lycopene content of the response region and control region. (c) Transcription levels of ripening‐related genes in the response region and control region as determined by quantitative RT‐PCR analysis. The tomato SlUBI3 gene was used as an internal control. PG2a, polygalacturonase 2a; EXP1, expansion 1; RIN, ripening inhibitor; CNR, colourless nonripening; NOR, nonripening; NR, never ripe; PSY1, phytoene synthase 1; PDS, phytoene desaturase. (d) Transcription levels of SlβCA3 and SlFAD3 in the response region and control region as determined by quantitative RT‐PCR analysis. The tomato SlUBI3 gene was used as an internal control. In (b–d), asterisks indicate significant differences (*P < 0.05, **P < 0.01, ***P < 0.001; Student's t test). (e) Model for SlDML2‐mediated regulation of tomato resistance to B. cinerea. SlDML2 transcriptionally activates the expression of defence‐related genes SlβCA3 and SlFAD3 through active DNA demethylation in the gene body and promoter region, respectively, thus positively regulating tomato fruit resistance to B. cinerea.
Considering that ripe tomato fruits are more susceptible to B. cinerea than unripe green fruits (Cantu et al., 2009; Prusky et al., 2013; Silva et al., 2023), we referred to the delayed ripening under B. cinerea invasion as a self‐protection mechanism of tomato fruits. It is possible that tomato fruit pericarp around disease region tend to abandon normal ripening process for obtaining increased defence capability in face with B. cinerea infection, thus achieving the balance between ripening and disease resistance. The ripening genes exhibiting decreased transcription level in the response region appeared to be the targets of SlDML2 (Figure 8c) (Lang et al., 2017). By contrast, the defence genes SlβCA3 and SlFAD3 exhibited significantly increased expression in the response region compared with the control region (Figure 8d). Accordingly, it is possible that SlDML2 preferentially activates the expression of defence genes, rather than those ripening genes, to confer tomato fruits increased resistance as far as possible and plays an essential role in mediating the trade‐off between fruit ripening and disease resistance during the pathogenic process of B. cinerea. Notably, B. cinerea invasion induced a decrease in SlDML2 mRNA level in the response region compared with the control region (Figure S13), suggesting that the delay in ripening of the pericarp tissue surrounding B. cinerea infection and the decrease in expression of ripening genes targeted by SlDML2 might be caused by the decreased expression of SlDML2 in this region.
Discussion
The DNA demethylase gene SlDML2 functions as a multifunctional gene
DEMETER/DEMETER‐like DNA demethylases contain the conserved domain that non‐specifically recognizes DNA sequence, which is generally thought to confer the DNA demethylases broad targets and diversified functions (Mok et al., 2010). Indeed, Arabidopsis DEMETER DNA demethylase gene AtDME has been demonstrated to modulate many aspects of sporophytic growth and development, including seed germination, root hair growth and de novo shoot formation (Kim et al., 2021). Moreover, the Arabidopsis DNA demethylase gene AtROS1, the ortholog of AtDME, participates in regulating the development of stomatal lineage cells, seed dormancy and the resistance responses to Hyaloperonospora arabidopsidis through active DNA demethylation (Yamamuro et al., 2014). However, SlDML2, the tomato homologue of AtROS1, has only been characterized as a pivotal ripening gene so far (Gao et al., 2022; Lang et al., 2017; Liu et al., 2015; Zhou et al., 2019). In this study, we generated the sldml2 homozygous mutants using CRIPSR/Cas9 gene‐editing system and found that sldml2 mutants exhibited obvious development defects in seeds, leaves and flowers, in addition to the delay in fruit ripening (Figure 1). We also reported here that SlDML2 positively regulates tomato resistance to the fungal pathogens B. cinerea and V. dahliae (Figure 2). Thus, our study extends the molecular function of SlDML2 to plant organ development and disease resistance and uncovers the SlDML2 as a multifunctional gene.
SlDML2‐mediated DNA demethylation facilitates the expression of defence‐related genes
Previous studies have suggested that SlDML2‐mediated DNA demethylation promotes the expression of hundreds of ripening genes, and thus facilitates tomato fruit ripening (Gao et al., 2022; Lang et al., 2017; Li et al., 2020; Zhou et al., 2019). In this study, we found that SlDML2 mutation weakened tomato resistance to B. cinerea and decreased the expression of plenty of defence‐related genes during B. cinerea invasion (Figure 2; Table S3). Accordingly, it is possible that SlDML2‐mediated DNA demethylation facilitates tomato resistance to B. cinerea by activating the transcription of plenty of defence‐related genes, similar to the ripening regulation that involves numerous ripening genes. To decipher the molecular basis underlying SlDML2‐mediated disease resistance, we focused on the β‐type carbonic anhydrase gene SlβCA3 for the reason that it has been identified as an important defence gene against the virulent pathogen Pst DC3000 (Hu et al., 2021), and the JA biosynthetic gene SlFAD3 because of the functional importance of JA pathway in defensing B. cinerea infection (Huang et al., 2022; Reyes‐Diaz et al., 2016; Shu et al., 2021). Although the two defence genes were revealed to be transcriptionally regulated by SlDML2 through active DNA demethylation and play an important role in the defence response against B. cinerea (Figures 4, 5, 6), it is possible that SlDML2 also targets other defence‐related genes, such as those in JA biosynthesis and signalling transduction pathway, to modulate tomato resistance to B. cinerea.
DNA methylation occurs in distinct sequences of genome, including promoters, gene bodies, transposable elements and intergenic regions (Tang et al., 2016; Zhang et al., 2018). Although SlDML2 was previously demonstrated to preferentially target promoters, transposable elements and intergenic regions, like AtROS1 in Arabidopsis (Tang et al., 2016), there still exist approximately 10% of SlDML2 targets located in gene body regions (Lang et al., 2017). In contrast to the role of promoter methylation in gene expression, the role of SlDML2‐targeted methylation of gene bodies in gene regulation is less characterized. In Arabidopsis, gene body methylation in the CG context exhibited a positive correlation with gene expression (Schmitz et al., 2013; Zilberman et al., 2007), while those in the CHG and CHH contexts tend to negatively regulate gene expression (You et al., 2012). The function in gene expression regulation mediated by gene body methylation was also revealed in other plant species, such as rice (Wang et al., 2017c), tomato (González et al., 2011) and common bean (Richard et al., 2018). In this study, we found that SlDML2‐mediated DNA demethylation in the first intron of SlβCA3 gene body facilitated its transcription (Figure 5), indicating a positive regulation of SlDML2 in gene expression by 5mC removal of the intronic regions. However, the underlying molecular basis is currently elusive. It is possible that SlDML2‐mediated intronic DNA demethylation regulates gene expression by associating with multiple events. For example, DNA demethylation in intron‐exon boundaries may impact alternative splicing, thereby modulating gene expression (Regulski et al., 2013).
SlDML2 may be an essential factor in modulating the trade‐off between fruit ripening and disease resistance
Fruit ripening and disease resistance are two tightly interconnected processes, as fleshy fruit exhibits increased susceptibility to B. cinerea with the ripening (Cantu et al., 2009), and B. cinerea infection induces the early‐ripe of unripe fruits (Silva et al., 2023). It is further supported by the cases that some ripening genes participate in regulating tomato resistance to B. cinerea, such as the vacuolar protease gene SlVPE3 (Wang et al., 2017b), the cell wall‐modifying gene SlPG and SlEXP1 (Cantu et al., 2008; Perini et al., 2017), and the transcription factor gene SlNOR (Cantu et al., 2009). In this study, the involvement of the pivotal ripening gene SlDML2 in tomato resistance to B. cinerea invasion was revealed (Figure 2), which indicates the functional importance of SlDML2 in simultaneously regulating fruit ripening and disease resistance.
B. cinerea infection is generally thought to accelerate tomato fruit ripening, partially through inducing ethylene synthesis and promoting respiration rate (Cristescu et al., 2002; Silva et al., 2023). This ripening‐promoting effect mainly refers to the whole unripe fruit. Indeed, we found that B. cinerea infection induced the pericarp tissues surrounding disease regions a delay in ripening (Figure 8a–c). Tomato fruits tend to abandon normal ripening process for obtaining enhanced defence capability when threated by continuous B. cinerea infection, known as the trade‐off between fruit ripening and disease resistance, as tomato fruits commonly exhibits increased susceptibility to B. cinerea with the progress of maturity. Therefore, the regional ripening inhibitory we observed could be an important self‐protection mechanism for tomato fruits under B. cinerea invasion.
A previous comparative transcriptomic analysis indicates that SlDML2 mutation decreased the expression of hundreds of ripening‐related genes during tomato fruit ripening, including those regulating pigment synthesis, flavonoid synthesis and fruit softening (Lang et al., 2017). As a comparison, our comparative transcriptomic analysis between tomato fruits of the wild‐type and sldml2 mutants during the pathogenic process of B. cinerea revealed that the down‐regulated genes in the sldml2 mutant fruit were most significantly enriched in defence response to biotic and abiotic stresses, including ‘defense response to fungus’, ‘response to chitin’, ‘response to wounding’, ‘response to oxidative stress’ and ‘cell wall organization’ (Table S3). It implicates that SlDML2 may preferentially target defence‐related genes to promote their expressions during B. cinerea infection, rather than those ripening genes. Consistent with this inference, several SlDML2‐targeted ripening genes exhibited decreased transcription levels in the pericarp tissue surrounding B. cinerea infection compared with the control (Figure 8c), whereas the expression of defence‐related genes SlβCA3 and SlFAD3 increased obviously (Figure 8d). Thus, we speculate that SlDML2 may participate in modulating the trade‐off between fruit ripening and disease resistance during B. cinerea invasion, although the underlying mechanisms remain to be further investigated.
In summary, our data revealed SlDML2‐mediated regulatory mechanism in tomato resistance against the fungal pathogen B. cinerea, wherein two defence‐related genes SlβCA3 and SlFAD3 were focused and proved to play important roles (Figure 8e). In addition, we reported here that SlDML2 participates in modulating multiple developmental processes besides fruit ripening. Thus, we defined the DNA demethylase gene SlDML2 as a multifunctional gene, rather than merely a ripening gene. This study provided us new perspectives concerning the physiological function of SlDML2 and facilitated the potential application of this essential regulatory gene in the future improvement of tomato.
Materials and methods
Plant materials and culture conditions
Seeds of wild‐type tomato (Solanum lycopersicum cv. Ailsa Craig) were obtained from Tomato genetics resource center (TGRC, https://tgrc.ucdavis.edu/policy.aspx). The sldml2 mutants in Ailsa Craig background were produced by CRISPR/Cas9 gene‐editing as described below. Tomato seedlings were cultured in a greenhouse with 25 °C, 60%–80% relative humidity and a 16/8 h light/dark photoperiod. Tomato flowers were tagged at anthesis to determine the ripening stages of fruits. For the ripening phenotype observation, fruits of the wild‐type and sldml2 mutants were harvested at 39, 42, 47 and 52 days post‐anthesis (dpa), when the wild‐type fruits reach to mature green (MG), breaker (Br), orange ripe (OR) and red ripe (RR) stages, respectively.
CRISPR/Cas9‐mediated gene editing
CRISPR/Cas9‐mediated gene editing was performed as the described (Ma et al., 2015). Briefly, three specific targets were designed using CRISPR‐P 2.0 (http://crispr.hzau.edu.cn/CRISPR2/) and then inserted into sgRNA expression cassettes containing the AtU3b, AtU6‐1 and AtU6‐29 promoters, respectively. The constructed sgRNA expression cassettes were introduced into the pYLCRISPR/Cas9Pubi‐H binary vector with the Golden Gate method (Engler and Marillonnet, 2014). The resulting vector was transformed into Agrobacterium tumefaciens strain GV3101. The Agrobacterium was cultured in Luria‐Bertani (LB) liquid medium containing 50 μg mL−1 kanamycin, 50 μg mL−1 gentamycin and 50 μg ml−1 rifampicin at 28 °C for 24 h and then utilized to infect the cotyledons of wild‐type tomato at a final OD600 of 0.5. Following the standard tissue culture described by Fillatti et al. (1987), transgenic seedlings were obtained, and the genotypes were analysed by PCR detection with primers flanking the targets. The homozygous seedlings were remained and cultured for subsequent studies. Sequences of primers used for PCR amplification are listed in Table S5.
B. cinerea culture condition and disease symptom assay
B. cinerea strain B05.10 was cultured on potato dextrose agar (PDA) medium at 22 °C and a 16/8 h light/dark photoperiod for 10 days. Then, the B. cinerea was collected in 1/2 potato dextrose broth (PDB) medium, followed by vortex for 30 s. The spores were separated from mycelium by filtration through two layers of sterile gauze, diluted in 1/2 PDB medium to a final concentration of 5 × 105 spores per millilitre, and finally used for inoculation. For the disease symptom assay in tomato, 5 μL spore suspension of B. cinerea was pipetted onto pre‐wound fruits at 39 and 90 dpa or 4‐week‐old detached leaves. For the disease symptom assay in N. benthamiana leaves, 5 μL spore suspension of B. cinerea was pipetted onto detached N. benthamiana leaves. The inoculated fruits and leaves were placed into enclosed plastic crates for maintaining high‐relative humidity, and then incubated in a growth room with 23 °C and a 16/8 h light/dark photoperiod. Disease lesion diameters were measured with a ruler and analysed by the crossing method (Li et al., 2022). The experiment was performed with three independent biological replicates.
RNA‐seq and data analysis
RNA‐seq was performed in B. cinerea‐infected fruits of the wild‐type and sldml2‐3 mutant. Total RNAs were extracted from the 5 mm‐width pericarp tissue surrounding disease regions with hot‐phenol according to the method of Moore et al. (2005). RNA quantity and purity were analysed by Bioanalyzer 2100 (Agilent, G2939A) and RNA 6000 Nano LabChip kit (Agilent, 5067‐1511), respectively. About 2 μg high‐quality RNA samples with an RNA integrity (RIN) number >7.0 were employed to library construction using NEBNext ultra RNA library prepare kit (NEB, E7530), according to the standard protocols. The constructed libraries were subsequently sequenced on an Illumina Novaseq™ 6000 sequencer with a paired‐end read length of 150 bp. The sequencing contains three independent biological replicates, and each RNA sample was extracted from at least six inoculated tomato fruits to avoid individual difference.
Raw sequencing reads containing adapters or low‐quality bases were filtered by Cutadapt (Version 1.9) (Martin, 2011) to obtain clean reads. The clean reads were aligned to the tomato build_SL3.0 reference genome (ftp://ftp.solgenomics.net/tomato_genome/) using HISAT2 package (Version 2.2.1) (Kim et al., 2019). The uniquely mapped reads were assembled by StringTie (Version 2.1.6) with default parameters (Pertea et al., 2016). Then, the gene expression value FPKM (fragment per kilobase of transcript per million mapped reads) was estimated and calculated by StringTie and ballgown (Pertea et al., 2016). Comparative transcriptomic analysis was performed by DESeq2 (Love et al., 2014), and genes with an adjusted P value <0.05 and FPKM fold change ≥2 were considered as differentially expressed genes. GO enrichment was performed on Gene Ontology Consortium (http://www.geneontology.org/) and only statistically significant results with a Bonferroni‐corrected P value <0.05 were remained. KEGG (Kyoto Encyclopedia of Genes and Genomes) enrichment was carried out on the KEGG Database (https://www.kegg.jp/kegg/) with default parameters.
Quantitative RT‐PCR analysis
Total RNAs (2 μg) were reverse transcribed into cDNAs using the HiScript® III RT SuperMix for qPCR kit (Vazyme, R323‐01). The synthetic cDNAs were then employed as templates for PCR amplification with the ChemQ universal SYBR qPCR master mix (Vazyme, Q711‐02‐AA) and a StepOne plus real‐time PCR system (Applied Biosystems). The PCR program (20 μL) was as follows: 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 s and 60 °C for 30 s. Transcription level was relatively calculated by normalizing to reference genes using the cycle threshold (CT) 2(−ΔCT) method (Schmittgen and Livak, 2008). Sequences of primers used for PCR amplification are listed in Table S5.
Details of the other methods are provided in Methods S1–S11.
Funding
This work was supported by the National Natural Science Foundation of China (31925035 and 32202557), the China Postdocral Science Foundation (2022M723371) and the Shandong Provincial Key Research and Development Program (2022TZXD0023).
Conflict of interest statement
The authors declare no conflicts of interest.
Author contributions
G.Q. and L.Z. conceived and designed the experiments. S.T. and W.W. provided critical discussions. L.Z., G.G. and X.L. performed the experiments and analysed the data. L.Z. and G.Q. wrote the manuscript.
Supporting information
Figure S1 Genotypes and predicted peptides of the SlDML2 locus in sldml2 mutants generated by CRISPR/Cas9‐mediated gene editing.
Figure S2 Fruits of sldml2 mutants can turn red at the final ripe stage.
Figure S3 SlDML2 mutation does not cause significant difference in seed appearance.
Figure S4 SlDML2 gene expression in various tomato organs as determined by quantitative RT‐PCR.
Figure S5 SlDML2 mutation does not cause significant difference in length of seedling stem.
Figure S6 Pearson correlation coefficients between RNA‐seq samples.
Figure S7 SlDML2 mutation impairs the gene expression of SlβCA3 and SlFAD3.
Figure S8 SlβCA3 and SlFAD3 show no effect on tomato resistance to V. dahliae.
Figure S9 Roles of SlβCA3 and SlFAD3 in regulating resistance to Pst DC3000.
Figure S10 SlDML2 mutation does not cause differential 5mC modification in the SlβCA3 promoter.
Figure S11 SlDML2 mutation disturbs the expression of JA biosynthetic genes.
Figure S12 B. cinerea invasion causes a delay in ripening of tomato pericarp tissues around the disease region.
Figure S13 Influence of B. cinerea invasion on SlDML2 gene expression.
Method S1 V. dahliae culture condition and disease symptom assay.
Method S2 Pst DC3000 culture condition and disease symptom assay.
Method S3 5mC assay by bisulphite sequencing.
Method S4 ChIP‐qPCR assay.
Method S5 McrBC‐PCR assay.
Method S6 Protein extraction and western blot.
Method S7 VIGS.
Method S8 Transient overexpression.
Method S9 Lycopene content measurement.
Method S10 Transcription activity assay.
Method S11 Data analysis.
Table S1 Up‐regulated genes in the fruit of sldml2‐3 mutant during the pathogenic process of B. cinerea compared with the wild‐type.
Table S2 Down‐regulated genes in the fruit of sldml2‐3 mutant during the pathogenic process of B. cinerea compared with the wild‐type.
Table S3 Gene ontology (GO) enrichment analysis of differentially expressed genes in the fruit of sldml2‐3 mutant during the pathogenic process of B. cinerea compared with the wild‐type.
Table S4 Down‐regulated genes in the fruit of sldml2‐3 mutant that was previously reported to harbour hypermethylated DMRs when the SlDML2 was disrupted.
Table S5 A summary of primer informations.
Acknowledgements
We thank Yaoguang Liu from South China Agriculture University for providing the binary vector pYLCRISPR/Cas9Pubi‐H. We thank Daqi Fu from China Agriculture University for providing the virus vectors pTRV1 and pTRV2. We also thank Zhaosheng Kong from Institute of Microbiology, Chinese Academy of Sciences for providing the V. dahliae strain V592.
Data availability statement
The raw sequencing data from the RNA‐seq assay have been deposited in the Gene Expression Omnibus database under the accession number GSE223774. All the other support data are included in the article or the supplemental files.
References
- Bartels, A. , Han, Q. , Nair, P. , Stacey, L. , Gaynier, H. , Mosley, M. , Huang, Q. et al. (2018) Dynamic DNA methylation in plant growth and development. Int. J. Mol. Sci. 19, 2144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blanco‐Ulate, B. , Vincenti, E. , Powell, A.L. and Cantu, D. (2013) Tomato transcriptome and mutant analyses suggest a role for plant stress hormones in the interaction between fruit and Botrytis cinerea . Front. Plant Sci. 4, 142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cantu, D. , Blanco‐Ulate, B. , Yang, L. , Labavitch, J.M. , Bennett, A.B. and Powell, A.L. (2009) Ripening‐regulated susceptibility of tomato fruit to Botrytis cinerea requires NOR but not RIN or ethylene. Plant Physiol. 150, 1434–1449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cantu, D. , Vicente, A.R. , Greve, L.C. , Dewey, F.M. , Bennett, A.B. , Labavitch, J.M. and Powell, A.L. (2008) The intersection between cell wall disassembly, ripening, and fruit susceptibility to Botrytis cinerea . Proc. Natl. Acad. Sci. USA, 105, 859–864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen, T. , Qin, G. and Tian, S. (2020) Regulatory network of fruit ripening: Current understanding and future challenges. New Phytol. 228, 1219–1226. [DOI] [PubMed] [Google Scholar]
- Cheung, N. , Tian, L. , Liu, X. and Li, X. (2020) The destructive fungal pathogen Botrytis cinerea‐insights from genes studied with mutant analysis. Pathogens, 9, 923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cristescu, S.M. , De Martinis, D. , Te Lintel Hekkert, S. , Parker, D.H. and Harren, F.J. (2002) Ethylene production by Botrytis cinerea in vitro and in tomatoes. Appl. Environ. Microbiol. 68, 5342–5350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deng, Y. , Zhai, K. , Xie, Z. , Yang, D. , Zhu, X. , Liu, J. , Wang, X. et al. (2017) Epigenetic regulation of antagonistic receptors confers rice blast resistance with yield balance. Science, 355, 962–965. [DOI] [PubMed] [Google Scholar]
- Domínguez, T. , Hernández, M.L. , Pennycooke, J.C. , Jiménez, P. , Martínez‐Rivas, J.M. , Sanz, C. , Stockinger, E.J. et al. (2010) Increasing omega‐3 desaturase expression in tomato results in altered aroma profile and enhanced resistance to cold stress. Plant Physiol. 153, 655–665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Du, M. , Zhao, J. , Tzeng, D.T.W. , Liu, Y. , Deng, L. , Yang, T. , Zhai, Q. et al. (2017) MYC2 orchestrates a hierarchical transcriptional cascade that regulates jasmonate‐mediated plant immunity in tomato. Plant Cell, 29, 1883–1906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elad, Y. , Williamson, B. , Tudzynski, P. and Delen, N. (2007) Botrytis Spp. and Diseases they Cause in Agricultural Systems‐an Introduction Botrytis: biology, Pathology and Control. London: Springer. [Google Scholar]
- Engler, C. and Marillonnet, S. (2014) Golden Gate cloning. Methods Mol. Biol. 1116, 119–131. [DOI] [PubMed] [Google Scholar]
- Fillatti, J.J. , Kiser, J. , Rose, R. and Comai, L. (1987) Efficient transfer of a glyphosate tolerance gene into tomato using a binary Agrobacterium tumefaciens vector. Nat. Biotechnol. 5, 726–730. [Google Scholar]
- Gao, Y. , Lin, Y. , Xu, M. , Bian, H. , Zhang, C. , Wang, J. , Wang, H. et al. (2022) The role and interaction between transcription factor NAC‐NOR and DNA demethylase SlDML2 in the biosynthesis of tomato fruit flavor volatiles. New Phytol. 235, 1913–1926. [DOI] [PubMed] [Google Scholar]
- González, R.M. , Ricardi, M.M. and Iusem, N.D. (2011) Atypical epigenetic mark in an atypical location: Cytosine methylation at asymmetric (CNN) sites within the body of a non‐repetitive tomato gene. BMC Plant Biol. 11, 94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han, Y. , Kuang, J. , Chen, J. , Liu, X. , Xiao, Y. , Fu, C. , Wang, J. et al. (2016) Banana transcription factor MaERF11 recruits histone deacetylase MaHDA1 and represses the expression of MaACO1 and expansins during fruit ripening. Plant Physiol. 171, 1070–1084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hu, Z. , Ma, Q. , Foyer, C.H. , Lei, C. , Choi, H.W. , Zheng, C. , Li, J. et al. (2021) High CO2‐ and pathogen‐driven expression of the carbonic anhydrase βCA3 confers basal immunity in tomato. New Phytol. 229, 2827–2843. [DOI] [PubMed] [Google Scholar]
- Huang, H. , Zhao, W. , Li, C. , Qiao, H. , Song, S. , Yang, R. , Sun, L. et al. (2022) SlVQ15 interacts with jasmonate‐ZIM domain proteins and SlWRKY31 to regulate defense response in tomato. Plant Physiol. 190, 828–842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jankowska, M. , Kaczynski, P. , Hrynko, I. and Lozowicka, B. (2016) Dissipation of six fungicides in greenhouse‐grown tomatoes with processing and health risk. Environ. Sci. Pollut. Res. Int. 23, 11885–11900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim, D. , Paggi, J.M. , Park, C. , Bennett, C. and Salzberg, S.L. (2019) Graph‐based genome alignment and genotyping with HISAT2 and HISAT‐genotype. Nat. Biotechnol. 37, 907–915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim, S. , Park, J.S. , Lee, J. , Lee, K.K. , Park, O.S. , Choi, H.S. , Seo, P.J. et al. (2021) The DME demethylase regulates sporophyte gene expression, cell proliferation, differentiation, and meristem resurrection. Proc. Natl. Acad. Sci. USA, 118, e2026806118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lang, Z. , Wang, Y. , Tang, K. , Tang, D. , Datsenka, T. , Cheng, J. , Zhang, Y. et al. (2017) Critical roles of DNA demethylation in the activation of ripening‐induced genes and inhibition of ripening‐repressed genes in tomato fruit. Proc. Natl. Acad. Sci. USA, 114, E4511–E4519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li, G. , Wang, Y. , Zhang, Z. , Chen, Y. and Tian, S. (2022) Mushroom alcohol controls gray mold caused by Botrytis cinerea in harvested fruit via activating the genes involved in jasmonic acid signaling pathway. Postharvest Biol. Technol. 186, 111843. [Google Scholar]
- Li, Z. , Jiang, G. , Liu, X. , Ding, X. , Zhang, D. , Wang, X. , Zhou, Y. et al. (2020) Histone demethylase SlJMJ6 promotes fruit ripening by removing H3K27 methylation of ripening‐related genes in tomato. New Phytol. 227, 1138–1156. [DOI] [PubMed] [Google Scholar]
- Liu, M. , Zhang, Z. , Xu, Z. , Wang, L. , Chen, C. and Ren, Z. (2021) Overexpression of SlMYB75 enhances resistance to Botrytis cinerea and prolongs fruit storage life in tomato. Plant Cell Rep. 40, 43–58. [DOI] [PubMed] [Google Scholar]
- Liu, R. , How‐Kit, A. , Stammitti, L. , Teyssier, E. , Rolin, D. , Mortain‐Bertrand, A. , Halle, S. et al. (2015) A DEMETER‐like DNA demethylase governs tomato fruit ripening. Proc. Natl. Acad. Sci. USA, 112, 10804–10809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Love, M.I. , Huber, W. and Anders, S. (2014) Moderated estimation of fold change and dispersion for RNA‐seq data with DESeq2. Genome Biol. 15, 550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luo, J. , Pan, X. , Peng, T. , Chen, Y. , Zhao, H. , Mu, L. , Peng, Y. et al. (2016) DNA methylation patterns of banana leaves in response to Fusarium oxysporum f. sp. cubense tropical race 4 . J. Integr. Agr. 15, 2736–2744. [Google Scholar]
- Ma, X. , Zhang, Q. , Zhu, Q. , Liu, W. , Chen, Y. , Qiu, R. , Wang, B. et al. (2015) A robust CRISPR/Cas9 system for convenient, high‐efficiency multiplex genome editing in monocot and dicot plants. Mol. Plant, 8, 1274–1284. [DOI] [PubMed] [Google Scholar]
- Martel, C. , Zhurov, V. , Navarro, M. , Martinez, M. , Cazaux, M. , Auger, P. , Migeon, A. et al. (2015) Tomato whole genome transcriptional response to tetranychus urticae identifies divergence of spider mite‐induced responses between tomato and Arabidopsis. Mol. Plant Microbe Interact. 28, 343–361. [DOI] [PubMed] [Google Scholar]
- Martin, M. (2011) Cutadapt removes adapter sequences from high‐throughput sequencing reads. EMBnet J. 17, 10–12. [Google Scholar]
- Min, D. , Li, F. , Cui, X. , Zhou, J. , Li, J. , Ai, W. , Shu, P. et al. (2020) SlMYC2 are required for methyl jasmonate‐induced tomato fruit resistance to Botrytis cinerea . Food Chem. 310, 125901. [DOI] [PubMed] [Google Scholar]
- Mok, Y.G. , Uzawa, R. , Lee, J. , Weiner, G.M. , Eichman, B.F. , Fischer, R.L. and Huh, J.H. (2010) Domain structure of the DEMETER 5‐methylcytosine DNA glycosylase. Proc. Natl. Acad. Sci. USA, 107, 19225–19230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moore, S. , Payton, P. , Wright, M. , Tanksley, S. and Giovannoni, J. (2005) Utilization of tomato microarrays for comparative gene expression analysis in the Solanaceae. J. Exp. Bot. 56, 2885–2895. [DOI] [PubMed] [Google Scholar]
- Perini, M.A. , Sin, I.N. , Villarreal, N.M. , Marina, M. , Powell, A.L. , Martinez, G.A. and Civello, P.M. (2017) Overexpression of the carbohydrate binding module from Solanum lycopersicum expansin 1 (Sl‐EXP1) modifies tomato fruit firmness and Botrytis cinerea susceptibility. Plant Physiol. Biochem. 113, 122–132. [DOI] [PubMed] [Google Scholar]
- Pertea, M. , Kim, D. , Pertea, G.M. , Leek, J.T. and Salzberg, S.L. (2016) Transcript‐level expression analysis of RNA‐seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prusky, D. , Alkan, N. , Mengiste, T. and Fluhr, R. (2013) Quiescent and necrotrophic lifestyle choice during postharvest disease development. Annu. Rev. Phytopathol. 51, 155–176. [DOI] [PubMed] [Google Scholar]
- Regulski, M. , Lu, Z. , Kendall, J. , Donoghue, M.T. , Reinders, J. , Llaca, V. , Deschamps, S. et al. (2013) The maize methylome influences mRNA splice sites and reveals widespread paramutation‐like switches guided by small RNA. Genome Res. 23, 1651–1662. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reyes‐Diaz, M. , Lobos, T. , Cardemil, L. , Nunes‐Nesi, A. , Retamales, J. , Jaakola, L. , Alberdi, M. et al. (2016) Methyl jasmonate: An alternative for improving the quality and health properties of fresh fruits. Molecules, 21, 567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richard, M.M.S. , Gratias, A. , Thareau, V. , Kim, K.D. , Balzergue, S. , Joets, J. , Jackson, S.A. et al. (2018) Genomic and epigenomic immunity in common bean: The unusual features of NB‐LRR gene family. DNA Res. 25, 161–172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmittgen, T.D. and Livak, K.J. (2008) Analyzing real‐time PCR data by the comparative C(T) method. Nat. Protoc. 3, 1101–1108. [DOI] [PubMed] [Google Scholar]
- Schmitz, R.J. , Schultz, M.D. , Urich, M.A. , Nery, J.R. , Pelizzola, M. , Libiger, O. , Alix, A. et al. (2013) Patterns of population epigenomic diversity. Nature, 495, 193–198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shu, P. , Zhang, S. , Li, Y. , Wang, X. , Yao, L. , Sheng, J. and Shen, L. (2021) Over‐expression of SlWRKY46 in tomato plants increases susceptibility to Botrytis cinerea by modulating ROS homeostasis and SA and JA signaling pathways. Plant Physiol. Biochem. 166, 1–9. [DOI] [PubMed] [Google Scholar]
- Silva, C. , Adaskaveg, J.A. , Mesquida‐Pesci, S.D. , Ortega‐Salazar, I.B. , Pattathil, S. , Zhang, L. , Hahn, M.G. et al. (2023) Botrytis cinerea infection accelerates ripening and cell wall disassembly to promote disease in tomato fruit. Plant Physiol. 191, 575–590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sivasankar, S. , Sheldrick, B. and Rothstein, S.J. (2000) Expression of allene oxide synthase determines defense gene activation in tomato. Plant Physiol. 122, 1335–1342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strassner, J. , Schaller, F. , Frick, U.B. , Howe, G.A. , Weiler, E.W. , Amrhein, N. , Macheroux, P. et al. (2002) Characterization and cDNA‐microarray expression analysis of 12‐oxophytodienoate reductases reveals differential roles for octadecanoid biosynthesis in the local versus the systemic wound response. Plant J. 32, 585–601. [DOI] [PubMed] [Google Scholar]
- Tang, D. , Gallusci, P. and Lang, Z. (2020) Fruit development and epigenetic modifications. New Phytol. 228, 839–844. [DOI] [PubMed] [Google Scholar]
- Tang, K. , Lang, Z. , Zhang, H. and Zhu, J.K. (2016) The DNA demethylase ROS1 targets genomic regions with distinct chromatin modifications. Nat. Plants, 2, 16169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tian, S. , Torres, R. , Ballester, A.R. , Li, B. , Vilanova, L. and González‐Candelas, L. (2016) Molecular aspects in pathogen‐fruit interactions: virulence and resistance. Postharvest Biol. Technol. 122, 11–21. [Google Scholar]
- Tirnaz, S. and Batley, J. (2019) DNA methylation: toward crop disease resistance improvement. Trends Plant Sci. 24, 1137–1150. [DOI] [PubMed] [Google Scholar]
- Veley, K.M. , Elliott, K. , Jensen, G. , Zhong, Z. , Feng, S. , Yoder, M. , Gilbert, K.B. et al. (2023) Improving cassava bacterial blight resistance by editing the epigenome. Nat. Commun. 14, 85. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang, C. , Yang, Q. , Wang, W. , Li, Y. , Guo, Y. , Zhang, D. , Ma, X. et al. (2017a) A transposon‐directed epigenetic change in ZmCCT underlies quantitative resistance to Gibberella stalk rot in maize. New Phytol. 215, 1503–1515. [DOI] [PubMed] [Google Scholar]
- Wang, W. , Cai, J. , Wang, P. , Tian, S. and Qin, G. (2017b) Post‐transcriptional regulation of fruit ripening and disease resistance in tomato by the vacuolar protease SlVPE3. Genome Biol. 18, 47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang, X. , Zhang, Z. , Fu, T. , Hu, L. , Xu, C. , Gong, L. , Wendel, J.F. et al. (2017c) Gene‐body CG methylation and divergent expression of duplicate genes in rice. Sci. Rep. 7, 2675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williamson, B. , Tudzynski, B. , Tudzynski, P. and van Kan, J.A. (2007) Botrytis cinerea: The cause of grey mould disease. Mol. Plant Pathol. 8, 561–580. [DOI] [PubMed] [Google Scholar]
- Xin, X. and He, S. (2013) Pseudomonas syringae pv. tomato DC3000: a model pathogen for probing disease susceptibility and hormone signaling in plants. Annu. Rev. Phytopathol. 51, 473–498. [DOI] [PubMed] [Google Scholar]
- Xin, Y. , Ma, B. , Zeng, Q. , He, W. , Qin, M. and He, N. (2021) Dynamic changes in transposable element and gene methylation in mulberry (Morus notabilis) in response to Botrytis cinerea . Hortic. Res. 8, 154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu, X. , Chen, Y. , Li, B. , Zhang, Z. , Qin, G. , Chen, T. and Tian, S. (2022) Molecular mechanisms underlying multi‐level defense responses of horticultural crops to fungal pathogens. Hortic. Res. 9, uhac066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamamuro, C. , Miki, D. , Zheng, Z. , Ma, J. , Wang, J. , Yang, Z. , Dong, J. et al. (2014) Overproduction of stomatal lineage cells in Arabidopsis mutants defective in active DNA demethylation. Nat. Commun. 5, 4062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- You, W. , Tyczewska, A. , Spencer, M. , Daxinger, L. , Schmid, M.W. , Grossniklaus, U. , Simon, S.A. et al. (2012) Atypical DNA methylation of genes encoding cysteine‐rich peptides in Arabidopsis thaliana . BMC Plant Biol. 12, 51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu, C. , Wang, H. , Yang, S. , Tang, X. , Duan, M. and Meng, Q. (2009) Overexpression of endoplasmic reticulum omega‐3 fatty acid desaturase gene improves chilling tolerance in tomato. Plant Physiol. Biochem. 47, 1102–1112. [DOI] [PubMed] [Google Scholar]
- Zhang, H. , Lang, Z. and Zhu, J.K. (2018) Dynamics and function of DNA methylation in plants. Nat. Rev. Mol. Cell Biol. 19, 489–506. [DOI] [PubMed] [Google Scholar]
- Zhang, Y. , Zhang, Y. , Ge, X. , Yuan, Y. , Jin, Y. , Wang, Y. , Zhao, L. et al. (2023) Genome‐wide association analysis reveals a novel pathway mediated by a dual‐TIR domain protein for pathogen resistance in cotton. Genome Biol. 24, 111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang, Z. , He, C. , Chen, Y. , Li, B. and Tian, S. (2021) DNA methyltransferases regulate pathogenicity of Botrytis cinerea to horticultural crops. J. Fungi (Basel), 7, 659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou, L. , Tian, S. and Qin, G. (2019) RNA methylomes reveal the m6A‐mediated regulation of DNA demethylase gene SlDML2 in tomato fruit ripening. Genome Biol. 20, 156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu‐Salzman, K. , Luthe, D.S. and Felton, G.W. (2008) Arthropod‐inducible proteins: Broad spectrum defenses against multiple herbivores. Plant Physiol. 146, 852–858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ziegler, J. , Stenzel, I. , Hause, B. , Maucher, H. , Hamberg, M. , Grimm, R. , Ganal, M. et al. (2000) Molecular cloning of allene oxide cyclase. The enzyme establishing the stereochemistry of octadecanoids and jasmonates. J. Biol. Chem. 275, 19132–19138. [DOI] [PubMed] [Google Scholar]
- Zilberman, D. , Gehring, M. , Tran, R.K. , Ballinger, T. and Henikoff, S. (2007) Genome‐wide analysis of Arabidopsis thaliana DNA methylation uncovers an interdependence between methylation and transcription. Nat. Genet. 39, 61–69. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figure S1 Genotypes and predicted peptides of the SlDML2 locus in sldml2 mutants generated by CRISPR/Cas9‐mediated gene editing.
Figure S2 Fruits of sldml2 mutants can turn red at the final ripe stage.
Figure S3 SlDML2 mutation does not cause significant difference in seed appearance.
Figure S4 SlDML2 gene expression in various tomato organs as determined by quantitative RT‐PCR.
Figure S5 SlDML2 mutation does not cause significant difference in length of seedling stem.
Figure S6 Pearson correlation coefficients between RNA‐seq samples.
Figure S7 SlDML2 mutation impairs the gene expression of SlβCA3 and SlFAD3.
Figure S8 SlβCA3 and SlFAD3 show no effect on tomato resistance to V. dahliae.
Figure S9 Roles of SlβCA3 and SlFAD3 in regulating resistance to Pst DC3000.
Figure S10 SlDML2 mutation does not cause differential 5mC modification in the SlβCA3 promoter.
Figure S11 SlDML2 mutation disturbs the expression of JA biosynthetic genes.
Figure S12 B. cinerea invasion causes a delay in ripening of tomato pericarp tissues around the disease region.
Figure S13 Influence of B. cinerea invasion on SlDML2 gene expression.
Method S1 V. dahliae culture condition and disease symptom assay.
Method S2 Pst DC3000 culture condition and disease symptom assay.
Method S3 5mC assay by bisulphite sequencing.
Method S4 ChIP‐qPCR assay.
Method S5 McrBC‐PCR assay.
Method S6 Protein extraction and western blot.
Method S7 VIGS.
Method S8 Transient overexpression.
Method S9 Lycopene content measurement.
Method S10 Transcription activity assay.
Method S11 Data analysis.
Table S1 Up‐regulated genes in the fruit of sldml2‐3 mutant during the pathogenic process of B. cinerea compared with the wild‐type.
Table S2 Down‐regulated genes in the fruit of sldml2‐3 mutant during the pathogenic process of B. cinerea compared with the wild‐type.
Table S3 Gene ontology (GO) enrichment analysis of differentially expressed genes in the fruit of sldml2‐3 mutant during the pathogenic process of B. cinerea compared with the wild‐type.
Table S4 Down‐regulated genes in the fruit of sldml2‐3 mutant that was previously reported to harbour hypermethylated DMRs when the SlDML2 was disrupted.
Table S5 A summary of primer informations.
Data Availability Statement
The raw sequencing data from the RNA‐seq assay have been deposited in the Gene Expression Omnibus database under the accession number GSE223774. All the other support data are included in the article or the supplemental files.


