Abstract
X chromosome inactivation silences one of two X chromosomes in the mammalian female cell and is controlled by a binary switch that involves interactions between Xist and Tsix, a sense-antisense pair of noncoding genes. On the future active X chromosome, Tsix expression suppresses Xist upregulation, while on the future inactive X chromosome, Tsix repression is required for Xist-mediated chromosome silencing. Thus, understanding the binary switch mechanism depends on ascertaining how Tsix expression is regulated. Here we have taken an unbiased approach toward identifying Tsix regulatory elements within the X chromosome inactivation center. First, we defined the major Tsix promoter and found that it cannot fully recapitulate the developmental dynamics of Tsix expression, indicating a requirement for additional regulatory elements. We then delineated two enhancers, one classical enhancer mapping upstream of Tsix and a bipartite enhancer that flanks the major Tsix promoter. These experiments revealed the intergenic transcription element Xite as an enhancer of Tsix and the repeat element DXPas34 as a component of the bipartite enhancer. Each enhancer contains DNase I-hypersensitive sites and appears to confer developmental specificity to Tsix expression. Characterization of these enhancers will facilitate the identification of trans-acting regulatory factors for X chromosome counting and choice.
In mammals, the expression of X-linked genes is equalized in males and females by the transcriptional inactivation of one X chromosome in the female (2, 25). During embryogenesis, random X chromosome inactivation (XCI) occurs as the epiblast lineage differentiates and is regulated by counting and choice mechanisms, such that one randomly selected X chromosome remains active in each cell (33). Appropriate developmental regulation of XCI requires the X chromosome inactivation center (Xic), a cis-acting locus on the X chromosome that functions as a binary switch between the on and off transcriptional states (34). Molecular and genetic analyses of the Xic have focused on two genes, Xist and Tsix, that are integral to its function. Xist produces a large noncoding RNA that triggers X chromosome inactivation and coats the inactive X chromosome (6, 7, 27, 31), while Tsix, the antisense partner of Xist, antagonizes Xist expression and negatively regulates X chromosome inactivation (11, 21, 22, 35). Thus, Tsix works genetically upstream of Xist, and the opposing interplay between these two genes determines the pattern of XCI in the female cell.
Many aspects of Tsix-Xist dynamics remain poorly understood. Because the suppression and induction of Xist are regulated by Tsix, understanding how XCI is regulated in turn requires an understanding of how Tsix expression is controlled. During cell differentiation, Tsix expression is silenced asynchronously on the active and inactive X chromosomes (Xa and Xi, respectively) (21, 22). On the Xi chromosome, the loss of Tsix expression permits the upregulation of Xist and chromosome inactivation in cis. On the Xa chromosome, the persistence of Tsix expression blocks Xist upregulation and preserves the activity of the chromosome, with Tsix expression continuing until just after Xist expression is silenced. Loss-of-function mutations that reduce, abolish, or truncate Tsix transcription cause Xist upregulation and XCI on the linked X chromosome (9, 22, 24, 28, 35, 37). Conversely, gain-of-function alterations that augment the strength and duration of Tsix expression from one allele prevent Xist upregulation and XCI in cis (24, 38). Thus, the appropriate temporal and allelic regulation of Tsix expression is necessary to specify the transcriptional fate of each X chromosome by suppressing Xist activity on one X chromosome and permitting its upregulation on the other. In this manner, the asymmetric profile of Tsix expression underlies the epigenetic choice of which X chromosome is inactivated.
How is Tsix expression controlled developmentally and allelically? Several elements were recently implicated. One study suggested that a regulatable chromatin insulator, CTCF, acts as a transcriptional switch for Xist and Tsix (8). CTCF binding occurs around DXPas34 (10), a repeat sequence downstream of the Tsix transcription start site at a location between the promoters of Tsix and Xist. This genetic arrangement is reminiscent of that observed at the autosomally imprinted locus H19/Igf2, where CTCF insulators are thought to regulate the accessibility of H19 and Igf2 on reciprocal chromosomes to shared enhancers (4, 14, 16, 17). In the Xic, however, such enhancers have not yet been reported. Relevant to this information, another recent study identified Xite, an intergenic region that is located upstream of Tsix and that harbors strong DNase I-hypersensitive sites (DHS) and several intergenic transcripts (30). Because deleting Xite results in the downregulation of Tsix at the onset of XCI and a skewed pattern of XCI, Xite is a candidate regulator of Tsix expression. Its mechanism of action is presently unknown. Finally, knockout studies of a 65-kb region (9, 29) that includes Tsix and Xite suggested the presence of X chromosome counting elements that are distinct from the 5′ ends of Tsix and Xite. Such elements are also candidate regulators of Tsix expression. Thus, there is considerable physical overlap among the various candidate regulatory elements of Tsix. However, although these processes are likely to be coupled, the molecular links have remained unclear.
Here we have taken a systematic and unbiased approach toward identifying regulatory sequences in the region downstream of Xist. We found that the major Tsix promoter is insufficient to confer appropriate developmental control, implicating additional elements in transcriptional regulation. Several enhancer-like sequences were identified within the Tsix and Xite gene bodies. As discussed below, the data suggest that the combination of these elements confers transcriptional regulation to Tsix. Because they reside in domains implicated in counting and choice, we speculate that counting and choice elements will be identified as transcriptional regulators of Tsix.
MATERIALS AND METHODS
RNase protection.
Total RNA was isolated with Trizol (Invitrogen). RNase protection was performed essentially as described previously (1). For Tsix riboprobes, pNS2 and pNS3 were cut with BamHI and transcribed with SP6 polymerase and [α-32P]CTP. An actin template (Clontech) was transcribed with T3 polymerase. For hybridizations, 2.5 × 105 cpm of probe and 20 μg of RNA were used. A radiolabeled DNA sequencing reaction served as a size standard.
Cell culture and transfection.
Male J1, female 16.7, and transgenic 116.6 embryonic stem (ES) cells and their culture conditions have been described elsewhere (21-23). For transfection of undifferentiated ES cells, cells were passaged ∼1:2.5 on gelatin-treated 24-well plates and cultured overnight. On the next day, cells were transfected with Lipofectamine 2000 (Invitrogen) according to the manufacturer's protocol. Briefly, transfection was performed with serum-containing antibiotic-free medium, 3 μl of Lipofectamine 2000, and 1 μg of DNA (100 ng of pCS2+cβgal [see below] and 900 ng of luciferase vector) per well. Transfection for 12 to 18 h yielded efficiencies of ∼50 to 90% for J1 cells and ∼10 to 20% for 16.7 cells.
To induce the differentiation of 16.7 ES cells, cells were aggregated into embryoid bodies, grown in the absence of leukemia inhibitory factor (LIF) for 4 days in suspension cultures, and grown for an additional 5 to 11 days on gelatin-treated plates. On the day prior to transfection, cells were trypsinized, used to seed each well of a 24-well plate at 105 cells, and cultured overnight. Transfection was performed with serum-containing antibiotic- and LIF-free medium, 2 μl of Lipofectamine 2000, and 1 μg of DNA (100 ng of pCS2+cβgal and 900 ng of luciferase vector) per well.
NIH 3T3 cells and immortalized female fibroblasts were grown in Dulbecco minimal essential medium containing 10% fetal bovine serum. For transfection of NIH 3T3 cells, 105 cells were used to seed each well of a 24-well plate. On the next day, cells were transfected in a manner similar to that used for ES cells with 3 μl of Lipofectamine 2000 and 2 μg of DNA (200 ng of pCS2+cβgal and 1.8 μg of luciferase vector) per well. For immortalized female fibroblasts, 0.5 × 105 cells were used to seed each well of a 24-well plate and then were transfected with 3 μl of Lipofectamine 2000 and 1 μg of DNA (100 ng of pCS2+cβgal and 900 ng of luciferase vector) per well.
In each experiment, samples were transfected in duplicate wells.
Luciferase and β-galactosidase assays.
Media were aspirated, and cells were lysed by the addition of 100 μl of Glo lysis buffer (Promega) and incubation at room temperature for 5 to 15 min. For the luciferase assay, 80 μl of lysate was mixed with 80 μl of Steady-Glo reagent (Promega) in a 96-well black OptiPlate-96F (Packard). The plate was incubated at room temperature for 5 min and covered with TopSeal-A sealing film (Packard). Luminescence was measured with a Packard TopCount-NXT instrument (3-min precount delay, 2-s count time).
For the β-galactosidase assay, 20 μl of lysate was diluted with 15 μl of Glo lysis buffer in a flat-bottom 96-well clear polystyrene plate. A total of 35 μl of ice-cold 2× β-galactosidase assay buffer (200 mM sodium phosphate [pH 7.3], 2 mM MgCl2, 100 mM β-mercaptoethanol, 1.33 mg of o-nitrophenyl-β-d-galactopyranoside/ml) was added, and the plate was incubated at 37°C in a humidified chamber. The reaction was monitored visually and terminated with 130 μl of 1 M Tris base. The absorbance was measured at 405 nm with a multiwell reader (Coulter) and corrected against the value for a water blank.
To control for transfection efficiency in each well, luciferase activity was normalized to β-galactosidase activity. Values for duplicate wells were averaged, and the mean was subsequently normalized to the value for a reference promoter to enable comparison across multiple experiments. For each plasmid, luciferase activity was measured in two to six independent experiments.
Plasmids.
pCS2+cβgal was described previously (39). Plasmids generated for this study are described in Table 1. Except where specifically noted otherwise, luciferase activity was assayed with pNS11, a kan/neo-marked promoterless luciferase vector derived from pGL3-Basic (Promega) as follows. A PvuII-ClaI fragment containing the herpes simplex virus thymidine kinase (TK) promoter (−196 to +18 bp [−196/+18] fragment; C to T at +13) was ligated in a three-piece reaction with 260-bp ClaI-PvuII and 3.5-kb PvuII fragments from pEGFP-1 (Clontech). In the resultant vector, pNS9, the TK promoter replaces the simian virus 40 origin-promoter-enhancer of pEGFP-1. pNS9 was subsequently cut with AflII, blunted, ligated to a BamHI linker, cut with BglII, blunted, and ligated to a NotI linker; the 3-kb NotI-BamHI fragment was liberated and ligated to a 2.2-kb NotI-BamHI fragment from pGL3-Basic, yielding pNS11. In pNS185, the BamHI site of pNS11 has been replaced with a BssHII site.
TABLE 1.
Plasmids generated in this studya
| Figure | Plasmid | Insert (5′ terminus-3′ terminus) | Backbone | Description |
|---|---|---|---|---|
| 1C | pNS2 | Tsix −44/+237 (BamHI-MscI) | pGEM7 | Riboprobe template |
| 1C | pNS3 | Tsix −44/+144 (BamHI-EcoRV) | pGEM7 | Riboprobe template |
| 1D | pNS11 | kan/neo promoterless luciferase vector | ||
| 1D | pNS58 | Herpes simplex virus TK promoter −198/+18; C to T at +13 | pNS11 | Herpes simplex virus TK promoter |
| 1D | pNS65 | Tsix −557/+176 (KpnI-SmaI) | pNS11 | 733-bp major promoter |
| 1D | pNS94 | Tsix −17271/−16131 (NheI-NheI) | pNS11 | 1.1-kb minor promoter |
| 2A | pNS78 | Tsix −457/+176 (AhdI-SmaI) | pNS11 | 5′ Truncation of major promoter |
| 2A | pNS79 | Tsix −279/+176 (BlpI-SmaI) | pNS11 | 5′ Truncation of major promoter |
| 2A | pNS80 | Tsix −160/+176 (BanII-SmaI) | pNS11 | 5′ Truncation of major promoter |
| 2A | pNS81 | Tsix −82/+176 (FspI-SmaI) | pNS11 | 5′ Truncation of major promoter |
| 2A | pNS82 | Tsix −44/+176 (BamHI-SmaI) | pNS11 | 5′ Truncation of major promoter |
| 2A | pNS83 | Tsix −24/+176 (Bsu36I-SmaI) | pNS11 | 5′ Truncation of major promoter |
| 2A | pNS84 | Tsix +31/+176 (BsgI-SmaI) | pNS11 | 5′ Truncation of major promoter |
| 2A | pNS85 | Tsix +67/+176 (BsiHKAI-SmaI) | pNS11 | 5′ Truncation of major promoter |
| 2A | pNS86 | Tsix +95/+176 (BspMI-SmaI) | pNS11 | 5′ Truncation of major promoter |
| 2A | pNS87 | Tsix +114/+176 (NarI-SmaI) | pNS11 | 5′ Truncation of major promoter |
| 2B | pNS66 | Tsix −557/+144 (KpnI-EcoRV) | pNS11 | 3′ Truncation of major promoter |
| 2B | pNS67 | Tsix −557/+116 (KpnI-NarI) | pNS11 | 3′ Truncation of major promoter |
| 2B | pNS69 | Tsix −557/+63 (KpnI-BsiHKAI) | pNS11 | 3′ Truncation of major promoter |
| 2B | pNS70 | Tsix −557/+29 (KpnI-BsgI) | pNS11 | 3′ Truncation of major promoter |
| 2B | pNS71 | Tsix −557/−21 (KpnI-Bsu36I) | pNS11 | 3′ Truncation of major promoter |
| 2B | pNS72 | Tsix −557/−40 (KpnI-BamHI) | pNS11 | 3′ Truncation of major promoter |
| 2B | pNS73 | Tsix −557/−82 (KpnI-FspI) | pNS11 | 3′ Truncation of major promoter |
| 2B | pNS74 | Tsix −557/−164 (KpnI-BanII) | pNS11 | 3′ Truncation of major promoter |
| 2B | pNS75 | Tsix −557/−276 (KpnI-BlpI) | pNS11 | 3′ Truncation of major promoter |
| 2C | pNS215 | Tsix −160/+116 (BanII-NarI) | pNS11 | 5′ and 3′ Truncations of major promoter |
| 3 | pNS64 | Tsix −1420/+116 (SacI-NarI) | pNS11 | −1.4 kb to promoter |
| 3 | pNS63 | Tsix −2407/+116 (PmlI-NarI) | pNS11 | −2.4 kb to promoter |
| 3 | pNS62 | Tsix −3498/+116 (NheI-NarI) | pNS11 | −3.5 kb to promoter |
| 3 | pNS61 | Tsix −4880/+116 (NciI-NarI) | pNS11 | −4.9 kb to promoter |
| 3 | pNS60 | Tsix −5920/+116 (BamHI-NarI) | pNS11 | −5.9 kb to promoter |
| 3 | pNS96 | Tsix −9009/+116 (StuI-NarI) | pNS11 | −9.0 kb to promoter |
| 3 | pNS228 | λ 17053/18556 (KpnI-KpnI) | pNS65 | λ 1.5 kb + Tsix promoter |
| 3 | pNS229 | λ 39355/41275 (EcoRV-EcoRV) | pNS65 | λ 1.9 kb + Tsix promoter |
| 3 | pNS231 | λ 18560/21271 (KpnI-EcoRV) | pNS65 | λ 2.7 kb + Tsix promoter |
| 3 | pNS232 | λ 45829/652 (EcoRV-EcoRV) | pNS65 | λ 3.3 kb + Tsix promoter |
| 3 | pNS234 | λ 2087/6683 (EcoRV-EcoRV) | pNS65 | λ 4.6 kb + Tsix promoter |
| 3 | pNS235 | λ 33589/28214 (EcoRV-EcoRV) | pNS65 | λ 5.4 kb + Tsix promoter |
| 3 | pNS236 | λ 34503/27973 (BamHI-BamHI) | pNS65 | λ 6.5 kb + Tsix promoter |
| 3 | pNS237 | λ 17058/5505 (KpnI-BamHI) | pNS65 | λ 11.5 kb + Tsix promoter |
| 3 | pNS113 | Tsix −12045*/−6535 (ND-AvrII) | pNS65 | (−12 kb to −6.5 kb) + Tsix promoter |
| 3 | pNS116 | Tsix −15105/−10229 (XhoI-XhoI) | pNS65 | (−15.1 kb to −10.2 kb) + Tsix promoter |
| 3 | pNS114 | Tsix −18196/−13656 (KpnI-KpnI) | pNS65 | (−18.2 kb to −13.7 kb) + Tsix promoter |
| 3 | pNS117 | Tsix −23351/−17267 (ClaI-NheI) | pNS65 | (−23.4 kb to −17.3 kb) + Tsix promoter |
| 3 | pNS115 | Tsix −27322/−21489 (XbaI-XbaI) | pNS65 | (−27.3 kb to −21.5 kb) + Tsix promoter |
| 3 | pNS123 | Tsix −6535/−12045* (AvrII-ND) | pNS65 | (−6.5 kb to −12 kb) + Tsix promoter |
| 3 | pNS109 | Tsix −13656/−18196 (KpnI-KpnI) | pNS65 | (−13.7 kb to −18.2 kb) + Tsix promoter |
| 4A | pNS134 | Tsix −11329/−6535 (ScaI-AvrII) | pNS65 | 5′ Deletion of proximal enhancer + Tsix promoter |
| 4A | pNS133 | Tsix −10234/−6535 (XhoI-AvrII) | pNS65 | 5′ Deletion of proximal enhancer + Tsix promoter |
| 4A | pNS132 | Tsix −9009/−6535 (StuI-AvrII) | pNS65 | 5′ Deletion of proximal enhancer + Tsix promoter |
| 4A | pNS129 | Tsix −12045*/−9010 (ND-StuI) | pNS65 | 3′ Deletion of proximal enhancer + Tsix promoter |
| 4A | pNS130 | Tsix −12045*/−10229 (ND-XhoI) | pNS65 | 3′ Deletion of proximal enhancer + Tsix promoter |
| 4A | pNS131 | Tsix −12045*/−11330 (ND-ScaI) | pNS65 | 3′ Deletion of proximal enhancer + Tsix promoter |
| 4A | pNS135 | Tsix −10234/−9010 (XhoI-StuI) | pNS65 | 1.2-kb enhancer + Tsix promoter |
| 4A | pNS142 | Tsix −9010/−10234 (StuI-XhoI) | pNS65 | 1.2-kb enhancer (reverse) + Tsix promoter |
| 4B | pNS248 | Tsix −10234/+116 (XhoI-NarI) | pNS11 | −10.2 kb to promoter |
| 4C | pNS155 | Tsix −10234/−9010 (XhoI-StuI) | pNS11 | 1.2-kb enhancer |
| 4C | pNS156 | Tsix −9010/−10234 (StuI-XhoI) | pNS11 | 1.2-kb enhancer (reverse) |
| 4F | pNS258 | Xist P1 promoter (−540/+9) | pNS11 | Xist P1 promoter |
| 4F | pNS262 | Tsix −10234/−9010 (XhoI-StuI) | pNS258 | 1.2-kb enhancer + Xist P1 promoter |
| 4F | pNS263 | Tsix −9010/−10234 (StuI-XhoI) | pNS258 | 1.2-kb enhancer (reverse) + Xist P1 promoter |
| 4F | pNS259 | Xist P2 promoter (−601/+21) | pNS11 | Xist P2 promoter |
| 4F | pNS264 | Tsix −10234/−9010 (XhoI-StuI) | pNS259 | 1.2-kb enhancer + Xist P2 promoter |
| 4F | pNS265 | Tsix −9010/−10234 (StuI-XhoI) | pNS259 | 1.2-kb enhancer (reverse) + Xist P2 promoter |
| 5A | pNS106 | Tsix −4880/+116 (NciI-NarI) placed 5′ to luciferase gene, Tsix +145/+724 (EcoRV-AgeI) placed 3′ to luciferase gene | pNS11 | Combined Tsix upstream and downstream sequences |
| 5A | pNS126 | Tsix −4880/+116 (NciI-NarI) placed 5′ to luciferase gene, Tsix +145/+1867 (EcoRV-SalI) placed 3′ to luciferase gene | pNS11 | Combined Tsix upstream and downstream sequences |
| 5A | pNS127 | Tsix −4880/+116 (NciI-NarI) placed 5′ to luciferase gene, Tsix +145/+4049 (EcoRV-HindIII) placed 3′ to luciferase gene | pNS11 | Combined Tsix upstream and downstream sequences |
| 5A | pNS107 | Tsix −9009/+116 (StuI-NarI) placed 5′ to luciferase gene, Tsix +145/+724 (EcoRV-AgeI) placed 3′ to luciferase gene | pNS11 | Combined Tsix upstream and downstream sequences |
| 5A | pNS108 | Tsix −9009/+116 (StuI-NarI) placed 5′ to luciferase gene, Tsix +145/+1867 (EcoRV-SalI) placed 3′ to luciferase gene | pNS11 | Combined Tsix upstream and downstream sequences |
| 5A | pNS128 | Tsix −9009/+116 (StuI-NarI) placed 5′ to luciferase gene, Tsix +145/+4049 (EcoRV-HindIII) placed 3′ to luciferase gene | pNS11 | Combined Tsix upstream and downstream sequences |
| 5B | pNS255 | Tsix −9009/+116 (StuI-NarI) placed 5′ to luciferase gene, Tsix +721/+4049 (AgeI-HindIII) placed 3′ to luciferase gene | pNS11 | Combined Tsix upstream and downstream sequences |
| 5B | pNS256 | Tsix −9009/+116 (StuI-NarI) placed 5′ to luciferase gene, Tsix +1863/+4049 (SalI-HindIII) placed 3′ to luciferase gene | pNS11 | Combined Tsix upstream and downstream sequences |
| 5C | pNS254 | λ 27479/36895 and Tsix −557/+116 (KpnI-NarI) placed 5′ to luciferase gene, Tsix +145/+4049 (EcoRV-HindIII) placed 3′ to luciferase gene | pNS11 | λ substitution, Tsix promoter, and downstream sequences |
| 5C | pNS251 | Tsix −9009/+116 (StuI-NarI) placed 5′ to luciferase gene, | pNS11 | Tsix upstream sequences and λ substitution |
| λ 18555/22350 placed 3′ to luciferase gene | ||||
| 5D | pNS190 | Tsix +145/+4049 (EcoRV-HindIII) and Tsix −557/+144 (KpnI-EcoRV) placed 5′ to luciferase gene, Tsix −9009/−557 (StuI-KpnI) placed 3′ to luciferase gene | pNS185 | Swap of Tsix upstream and downstream sequences |
| 5D | pNS195 | Tsix +145/+4049 (EcoRV-HindIII) and Tsix −557/+144 (KpnI-EcoRV) placed 5′ to luciferase gene, Tsix −4880/−557 (NciI-KpnI) placed 3′ to luciferase gene | pNS185 | Swap of Tsix upstream and downstream sequences |
| 5F | pNS249 | Tsix −10234/+116 (XhoI-NarI) placed 5′ to luciferase gene, Tsix +145/+4049 (EcoRV-HindIII) placed 3′ to luciferase gene | pNS11 | Combined Tsix upstream and downstream sequences |
Plasmids are listed in the order in which they first appear in the figures, from left to right and top to bottom. The orientation of λ inserts is not known. An asterisk indicates that the terminus is approximate. For Tsix sequences, +1 corresponds to base 77675 of the sequence assigned GenBank accession no. X99946. Construction details and plasmid sequences are available on request. ND, end not determined.
DHS mapping.
DHS mapping was performed essentially as described previously (13). Briefly, 108 trypsinized cells were washed with phosphate-buffered saline, pelleted, and resuspended in 10 ml of ice-cold nuclear isolation buffer (NIB) (0.32 M sucrose, 3 mM CaCl2, 2 mM magnesium acetate, 10 mM Tris-HCl [pH 7.5], 0.1 mM EDTA, 1 mM dithiothreitol). A 1/20 volume of NIB-0.3% NP-40 was added, and the suspension was subjected to Dounce homogenization 15 times with pestle B. Nuclei were centrifuged at 400 × g and resuspended in ice-cold NIB at 0.2 mg/ml, as determined by total nucleic acid content. For ES cells, DNase I (Worthington) was added to aliquots at final concentrations of 0 to 10 μg/ml, and samples were incubated at 37°C for 2 min. For fibroblasts, the final concentrations of DNase I were 0 to 80 μg/ml, and samples were incubated for 5 min. Digestions were terminated by the addition of an equal volume of 2× stop solution (1% sodium dodecyl sulfate, 0.6 M NaCl, 20 mM Tris-HCl [pH 7.5], 10 mM EDTA, 0.2 mg of proteinase K/ml), and samples were incubated overnight at 37°C. DNA was precipitated with 1.25 M ammonium acetate, resuspended, and digested for Southern analysis.
For Southern blotting, HindIII- and BamHI-digested DNAs were probed with a 0.5-kb BamHI-AvrII fragment (bases 141031 to 141540 of the sequence assigned GenBank accession no. AJ421479) and a 0.5-kb BamHI-MscI fragment (bases 146400 to 146909 of the sequence assigned GenBank accession no. AJ421479), respectively.
RESULTS
The major Tsix promoter cannot recapitulate developmental regulation.
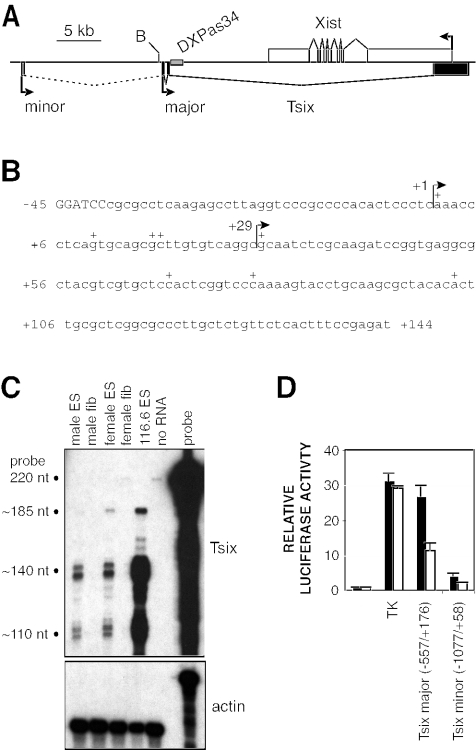
Two CpG-rich transcription initiation sites of Tsix have been described previously, with major and minor promoters located, respectively, 12 and 28 kb downstream of Xist (Fig. 1A) (21, 35, 36). Because several different 5′ termini have been described at the major initiation site (Fig. 1B), we performed RNase protection to determine the precise location of the major Tsix promoter. With a 220-nucleotide (nt) Tsix probe spanning the 5′ termini, two clusters of protected bands specific to ES cells were observed at nt 140 and 110, corresponding, respectively, to initiation sites 45 and 75 bp downstream of a BamHI site (Fig. 1B and C). A less abundant, fully protected band of 185 nt was also observed, likely corresponding to transcription originating from the minor Tsix promoter and other upstream positions (Fig. 1C) (11, 30, 35, 36). Similar results were obtained with a second, overlapping Tsix probe (data not shown). These data define within the major promoter two transcription initiation sites that coincide with previously described 5′ termini located 46 bp (36) and 75 bp (35) downstream of the BamHI site (Fig. 1B). We designate these termini bp +1 and bp +29, respectively.
FIG. 1.
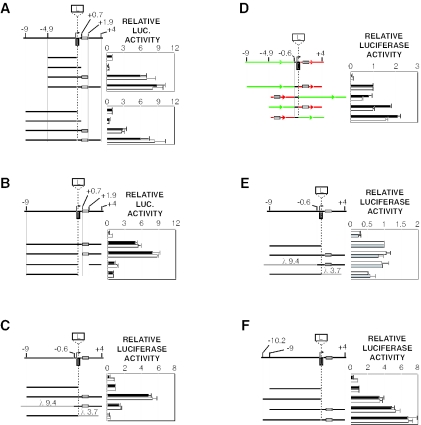
Functional delineation of the major Tsix promoter. (A) Map of the Xist/Tsix locus. White boxes, Xist exons; black boxes, Tsix exons; broken line, less abundant Tsix transcripts; grey rectangle, DXPas34 repeat array; B, BamHI. (B) Sequence proximal to the major initiation site. The 5′ terminus is the BamHI site in panel A. Previously described 5′ RACE termini are indicated by plus signs (21, 35, 36; D. Warshawsky, unpublished data). Arrows denote start sites defined by RNase protection analysis (see panel C). (C) RNase protection analysis at the major Tsix initiation site. Results obtained with Tsix (top panel) and actin (bottom panel) probes are shown. The Tsix riboprobe contains a 31-nt 5′ leader sequence fused to a 189-nt Tsix sequence antisense to that shown in panel B. 116.6 ES cells bear ∼25 copies of the Xic (23). (D) Activities of the major and minor Tsix promoters. Mean luciferase activities are shown for transiently transfected male ES cells (black bars) and female ES cells (white bars). Error bars denote one standard error. The activity of the backbone vector was set to 1. The major promoter is numbered as in panel B; the minor promoter is numbered with respect to the minor start site (35).
To functionally define the major promoter, we cloned a −557/+176 fragment upstream of a promoterless luciferase reporter. In transiently transfected ES cells, this fragment provided luciferase expression comparable to that of a control herpes simplex virus TK promoter (Fig. 1D). The minor upstream Tsix promoter was found to be five- to sixfold less active when assayed in parallel (Fig. 1D), consistent with the lower abundance of upstream Tsix transcripts (Fig. 1C) (11, 30, 35, 36). Because the upstream promoter produces a small fraction of Tsix transcripts and its removal causes no obvious consequences for XCI (30, 35), our subsequent experiments focused exclusively on the major promoter.
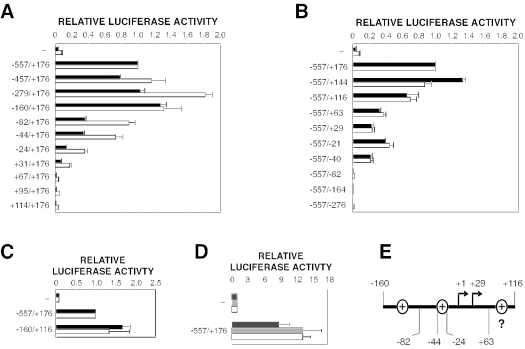
To define the sequences that confer full promoter activity, we next generated 5′ and 3′ deletion derivatives of the −557/+176 region and compared their activities to that of the parental fragment. As measured in ES cells, 5′ truncations as far as bp −160 did not decrease transcriptional activity (Fig. 2A). Further truncations sharply reduced activity, with discrete reductions suggesting the presence of positive elements between bp −160 and −82 and between bp −44 and −24 that are distinct from the two initiation sites (Fig. 2A and E). For deletions at the 3′ terminus, fragments truncated at bp +144 or +116 showed full or slightly reduced promoter activity, respectively (Fig. 2B). More severe 3′ truncations elicited discrete reductions in activity, consistent with the loss of a positive regulatory element between bp +63 and +116 (Fig. 2E) or with alterations in transcript stability or translation efficiency. As predicted by this deletion analysis, a −160/+116 fragment that combined both 5′ and 3′ deletions was sufficient for full promoter activity (Fig. 2C).
FIG. 2.
The major Tsix promoter is constitutively active and is contained within a 276-bp region. (A to C) Activities of Tsix promoter deletion derivatives. Mean luciferase activities are shown for transiently transfected male ES cells (black bars) and female ES cells (white bars). Error bars denote one standard error. The activity of the +557/+176 fragment was set to 1. (D) The major Tsix promoter is constitutively active. Mean luciferase activities are shown for transiently transfected immortalized female mouse fibroblasts (dark grey bars), male NIH 3T3 fibroblasts (light grey bars), and differentiated female ES cells (days 11 to 17) (white bars). Error bars denote one standard error. The activity of the backbone vector was set to 1. (E) Anatomy of the major Tsix promoter. Inferred locations of positive elements are indicated by ovals within denoted sequence intervals.
Tsix is expressed in undifferentiated embryonic cells and is silenced during differentiation. Because this property is a key criterion for judging whether isolated regulatory elements faithfully recapitulate Tsix expression, we compared the activities of the promoter in undifferentiated ES cells, which express Tsix, and in immortalized female mouse fibroblasts and male NIH 3T3 fibroblasts, which do not (Fig. 1C and data not shown) (21). Additionally, we assayed the promoter in female ES cells that had been allowed to differentiate for 11 to 17 days in the absence of LIF, a time at which Tsix expression has been silenced in most cells (38). In each of these assays, the Tsix promoter was active (Fig. 2D). Thus, the Tsix promoter is constitutively active and does not recapitulate the developmental dynamics of Tsix expression. We concluded that additional cis regulatory elements must be present in Xic and endeavored to identify them.
Identification of a Tsix enhancer that maps within Xite.
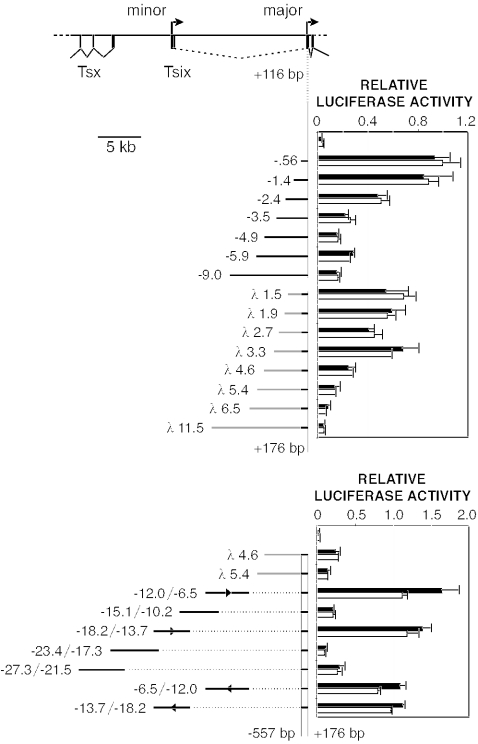
To determine whether transcriptional control elements reside upstream of the major Tsix promoter, we performed two experiments. First, we examined whether contiguous 5′-flanking sequences extending up to kb −9 could modify promoter activity (Fig. 3, top panel). Because transfection efficiency is related inversely to plasmid size (18), we also assayed control plasmids containing equivalent amounts of heterologous sequences from bacteriophage λ. When controlled for insert size, none of the Tsix fragments up to kb −9 significantly enhanced or repressed the Tsix promoter in transiently transfected ES cells. These results indicate that upstream Tsix sequences extending to −9 kb are by themselves unable to modify promoter activity.
FIG. 3.
Identification of enhancing activities in the Xite region. Mean luciferase activities are shown for transiently transfected male ES cells (black bars) and female ES cells (white bars). Error bars denote one standard error. The activity of the Tsix promoter (−557/+176; not shown) was set to 1. A map of the Tsix upstream region is shown at the top, with terminal Tsx exons at the left. (Upper panel) Assay of contiguous Tsix sequences extending from bp +116 to kb −9; λ sequences are coupled to the Tsix promoter (−557/+176). (Lower panel) Assay of distal sequences extending from kb −6.5 to −27.3. The orientations of selected fragments are indicated.
Second, we examined whether sequences beyond kb −9 might contain Tsix regulatory elements. Five overlapping fragments extending from kb −6.5 to kb −27.3 were cloned upstream of the Tsix promoter-luciferase cassette in their natural orientation and assayed in ES cells. Two of these fragments, the first extending from kb −12 to kb −6.5 and the second extending from kb −18.2 to kb −13.7, stimulated luciferase activity 4- to 12-fold relative to the results obtained with similarly sized control plasmids (Fig. 3, bottom panel). The stimulatory effects were orientation independent (Fig. 3), suggesting the presence of transcriptional enhancers in these regions.
To carry out finer mapping, we generated 5′ and 3′ deletion derivatives for both fragments and assayed their activities in ES cells. Deletion analysis of fragment from kb −18.2 to −13.7 (−18.2/−13.7 fragment) defined a 1.1-kb MluI-XhoI segment that was required for promoter stimulation (data not shown). This segment overlaps the minor upstream promoter defined previously (30, 35). However, because genetic analyses have shown no apparent function for this region in either random or imprinted X chromosome inactivation in vivo (30, 35), we chose not to pursue its analysis further.
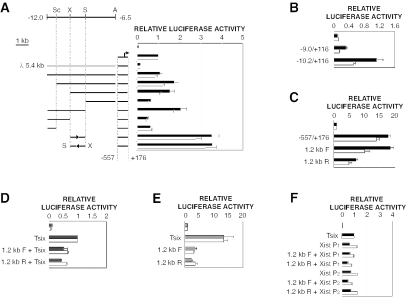
Deletion analysis of the proximal element defined a 1.2-kb XhoI-StuI segment between kb −10.2 and −9.0 that was both necessary and sufficient for Tsix promoter stimulation, yielding a 2.5- to 3.5-fold stimulatory effect in male and female ES cells in an orientation-independent manner (Fig. 4A). In these experiments, the tested fragments were juxtaposed to the Tsix promoter, a synthetic configuration lacking intervening sequences. To test whether the 1.2-kb segment was able to stimulate the promoter in a more natural context, we assayed the entire upstream region from kb −10.2 to the promoter. In ES cells, this fragment stimulated the promoter approximately 3.5-fold relative to the results obtained with a similarly sized fragment extending to kb −9 (Fig. 4B). Thus, the 1.2-kb element was able to stimulate the Tsix promoter not only when placed immediately upstream but also from a significant distance in its natural context. Thus, we conclude that the −10.2/−9.0 region is able to function as a classical transcriptional enhancer of Tsix.
FIG. 4.
Xite contains a 1.2-kb enhancer that is developmentally specific. (A) Deletion analysis of the upstream enhancer. Sc, ScaI; X, XhoI; S, StuI; A, AvrII. The orientations of selected fragments are indicated. Mean luciferase activities are shown for transiently transfected male ES cells (black bars) and female ES cells (white bars). Error bars denote one standard error. The activity of the Tsix promoter (−557/+176) was set to 1. (B) The 1.2-kb enhancer stimulates the Tsix promoter from its natural context. The activity of the Tsix promoter (−557/+116; not shown) was set to 1. (C) Bidirectional promoter activities of the 1.2-kb element. In this and subsequent panels, the forward (F) orientation of the 1.2-kb element is that in which the downstream (kb −9) terminus abuts the luciferase reporter; in the reverse (R) orientation, the upstream (kb −10.2) terminus abuts the reporter. The activity of the backbone vector was set to 1. (D) The 1.2-kb enhancer is unable to stimulate the Tsix promoter (−557/+176) in differentiated cell types. Mean luciferase activities are shown for immortalized female fibroblasts (grey bars) and differentiated female ES cells (days 11 to 17) (white bars). The activity of the Tsix promoter (−557/+176) was set to 1. (E) Bidirectional promoter activities are reduced in differentiated cell types. Mean luciferase activities are shown for transiently transfected NIH 3T3 fibroblasts (grey bars) and differentiated female ES cells (days 11 to 17) (white bars). The activity of the backbone vector was set to 1. (F) The enhancer is unable to stimulate the Xist promoter in undifferentiated ES cells. The 1.2-kb enhancer was assayed in the indicated orientation upstream of the Xist P1 promoter (−540/+9) or the Xist P2 promoter (−601/+21). Mean luciferase activities are shown for transiently transfected male ES cells (black bars) and female ES cells (white bars). The activity of the Tsix promoter (−557/+176) was set to 1.
The position of the enhancer between kb −10.2 and −9.0 places it within the Xite locus (30). Indeed, Xite contains multiple DHS, a hallmark of enhancers, and has been shown to positively regulate Tsix expression. The 1.2-kb enhancer coincides with the most prominent DHS in the region and with initiation sites for the intergenic Xite transcript (30). In the course of the present analysis, we found that the 1.2-kb region contains bidirectional promoter activities. When joined to a promoterless luciferase reporter in its natural orientation, the 1.2-kb region provided luciferase expression at a level comparable to that provided by the major Tsix promoter (Fig. 4C). Promoter activity was slightly weaker in the reverse orientation (Fig. 4C). Given the bidirectional promoter activities, one potential explanation for the enhancing effect observed earlier is that the 1.2-kb region simply adds to luciferase expression through its promoter activity and a mechanism of transcriptional readthrough. However, we do not believe this to be the case, as the Xite promoter would provide a maximum twofold increase in expression, significantly lower than the observed level of stimulation. Furthermore, potential readthrough transcripts from the 1.2-kb region are unlikely to increase luciferase expression when initiated at a significant distance from the reporter (Fig. 4B). Therefore, we conclude that the 1.2-kb region contains an enhancer that is functionally distinct from any inherent promoter activity.
The 1.2-kb Xite enhancer exhibits proper developmental regulation and is specific for Tsix.
To test whether the Xite enhancer stimulates Tsix expression in a developmentally specific manner, we compared its activities in undifferentiated ES cells, where Tsix is active, and in differentiated ES cells and fibroblasts, where Tsix expression has been reduced or extinguished. In contrast to its ability to stimulate the Tsix promoter in undifferentiated ES cells (Fig. 4A and B), the Xite enhancer was inactive in male NIH 3T3 fibroblasts, immortalized female fibroblasts, and female ES cells that had been allowed to differentiate for over 11 days (Fig. 4D and data not shown). This profile of enhancer activity correlates with the presence of a DHS within the 1.2-kb Xite enhancer specifically in undifferentiated cells (30). Interestingly, the bidirectional promoter activities of this element were also specific to undifferentiated ES cells (Fig. 4C and E), consistent with the loss of intergenic Xite transcripts upon cell differentiation (30). Thus, both enhancer and promoter activities were specific to undifferentiated ES cells, suggesting that the 1.2-kb region mediates many of the properties of Xite and contributes to the developmental regulation of Tsix expression in vivo.
One proposal for the control of Xist and Tsix expression invokes an enhancer that is shared by both genes and that is regulated in an allele-specific manner by an intervening chromatin insulator (8). To determine whether the Xite enhancer might be shared by Tsix and Xist, we tested whether it could stimulate the Xist promoter in undifferentiated ES cells and found that it induced neither the Xist P1 nor the Xist P2 promoter (15) when placed upstream in either orientation (Fig. 4F). Because Xist expression increases significantly in differentiated female cells after the onset of XCI, we also tested somatic cells and female ES cells that had been allowed to differentiate for 11 to 17 days. However, neither immortalized female fibroblasts nor differentiated ES cells exhibited any stimulation of the P1 and P2 promoters when the Xite enhancer was placed upstream in either orientation (data not shown). Thus, the Xite enhancer acts specifically on Tsix and has no apparent effect on the Xist promoter in the context of these experiments.
Identification of a bipartite enhancer in Tsix with developmental specificity.
To identify additional regulatory elements, we screened for stimulatory sequences downstream of the Tsix promoter. Some attention has been focused on the repeat array DXPas34 (10) and a number of flanking CTCF-binding sites (8). To test the idea that DXPas34 might be a Tsix control element (8, 11), we examined whether it could stimulate the Tsix promoter in various contexts. When placed upstream of the Tsix promoter, various fragments containing DXPas34 alone were unable to confer stimulation in either orientation (data not shown). When placed 3′ of a luciferase reporter driven by the Tsix promoter, DXPas34 conferred a low level of stimulation, but only in some vector backbones (data not shown). These findings indicated that DXPas34 by itself has little to no stimulatory capability.
We next considered whether DXPas34 might regulate Tsix expression in a context-specific fashion that requires additional sequences. To test this idea, we assayed various combinations of upstream and downstream sequences (relative to the Tsix start site) for their ability to stimulate the promoter. As described earlier, upstream sequences extending to kb −4.9 or −9 were by themselves unable to augment promoter activity significantly (Fig. 3, top panel). However, combinations of these upstream fragments with downstream segments terminating at kb +1.9 or +4 elicited potent luciferase stimulation, ranging from three- to ninefold, relative to that obtained with the upstream fragments in isolation (Fig. 5A). When each upstream fragment was combined with a downstream sequence extending to kb +0.7, stimulation was not observed, indicating that DXPas34 was required for this effect. Inclusion of additional sequences from kb +1.9 to +4.0 further increased stimulation, especially in the context of upstream sequences extending to kb −9. Although the 6.8-kb region extending from kb −4.9 to the DXPas34 3′ terminus at kb +1.9 appeared to provide the bulk of this stimulation, the apparent contributions of distal upstream and downstream sequences suggested that as many as four distinct elements within a 13-kb region might mediate this effect. Thus, despite the inability of these elements to stimulate the promoter individually, combinations of these elements could stimulate Tsix expression in a highly cooperative manner.
FIG. 5.
A bipartite enhancer flanks the Tsix promoter and is developmentally specific. In sequence schematics, alternatively spliced exons 2′ and 3 are omitted for clarity; L, luciferase cassette. DXPas34 is shown as a rectangle between kb +0.7 and +1.9. For all panels except E, mean luciferase activities are shown for transiently transfected male ES cells (black bars) and female ES cells (white bars). For all panels except D, the activity of upstream sequences in isolation was set to 1. Error bars denote one standard error. (A) Synergistic stimulation of the Tsix promoter by combinations of upstream and downstream sequences. (B) DXPas34 is required for stimulation. (C) Stimulation depends specifically on Tsix sequences. Bacteriophage λ sequences are labeled. (D) Swap of upstream and downstream sequences preserves stimulation. Orientations of upstream (green) and downstream (red) sequences are indicated. The activity of the native configuration was set to 1. The backbone was pNS185. (E) Combinatorial stimulation is absent in differentiated cells. Mean luciferase activities are shown for transiently transfected differentiated female ES cells (days 11 to 17) (lower bar in each set) and male NIH 3T3 fibroblasts (upper bar in each set). (F) Stimulation of the promoter by the 1.2-kb enhancer and more proximal upstream and downstream regions.
In all contexts, DXPas34 is essential for this cooperative stimulation. While the +0.1/+0.7 region (including Tsix exons 2 and 3) was dispensable, extending the deletion further to include DXPas34 caused a loss of stimulation. Moreover, although sequences between kb +1.9 and +4 provided additional stimulation (Fig. 5A), they were unable to do so in the absence of DXPas34 (Fig. 5B). We also found that when upstream or downstream sequences were replaced with stuffer fragments from bacteriophage λ, stimulation was severely reduced (Fig. 5C). Thus, the regulatory region has a bipartite structure, with an upstream region that contains sequences extending to kb −9.0 and a downstream region that contains DXPas34 and additional sequences extending to kb +4.0.
To probe the context dependence of these interactions further, we assayed the consequences of swapping upstream and downstream sequences with respect to the Tsix promoter (Fig. 5D). Significantly, such swapping did not affect cooperative stimulation of the Tsix promoter for most substitutions (Fig. 5D and data not shown). These results demonstrate that combinations of upstream and downstream sequences can interact in a modular fashion and, like classical enhancers, tolerate orientation and position changes. DXpas34 appears to have a central role in mediating the cooperative interactions of upstream and downstream regions within the enhancer.
Because the downstream sequences were placed 3′ of the reporter in these assays, one potential concern is that the stimulation attributed to a bipartite enhancer might instead result from posttranscriptional alterations, such as alternative splicing or polyadenylation. Several experiments argued against this possibility. The removal of Tsix exons and splice donors within the downstream region did not affect cooperative stimulation (Fig. 5B). Furthermore, Northern blotting and 3′ rapid amplification of cDNA ends (RACE) indicated that, in all instances, the sizes and 3′ termini of luciferase messages were indistinguishable from those produced by a control simian virus 40 promoter-enhancer (data not shown). Thus, we conclude that the bipartite enhancer stimulates Tsix expression at the transcriptional level rather than through alterations of splicing or polyadenylation.
To determine whether the stimulation provided by the bipartite enhancer is developmentally specific, we assayed its activity in NIH 3T3 fibroblasts and in differentiated female ES cells. In both backgrounds, the enhancer did not stimulate the Tsix promoter (Fig. 5E). Additionally, replacement of upstream or downstream sequences with heterologous bacteriophage sequences had little to no effect on luciferase activities in these differentiated cell types. Thus, as with the Xite enhancer, the stimulation conferred by the bipartite enhancer was specific to undifferentiated ES cells. We suggest that, together with the Xite enhancer, the bipartite enhancer contributes to the developmental specificity of Tsix transcription.
We next examined whether the bipartite enhancer might add to the effect of the Xite enhancer in regulating Tsix (Fig. 5F). Indeed, the presence of both the Xite enhancer and the bipartite enhancer augmented luciferase expression, yielding ∼40% greater stimulation than the bipartite enhancer alone (compare −10.2/+4.0 and −9.0/+4.0). This finding is consistent with the ideas that the Xite enhancer and the bipartite enhancer can function in parallel and that these elements work together to control both the level and the proper developmental timing of Tsix expression in vivo.
Tsix enhancers are associated with developmentally specific DHS.
In vivo, transcriptional enhancers are often associated with an open chromatin conformation. DHS mapping in the Xite region previously revealed several DHS specific to undifferentiated ES cells, with a strong DHS coinciding with the 1.2-kb enhancer and several weaker ones located in close promixity, approximately −7.7 and −8.8 kb from the major Tsix promoter (30). Thus, both the Xite enhancer and the 5′ portion of the bipartite enhancer are associated with developmentally specific DHS. To determine whether the remainder of the bipartite enhancer also lies in specialized chromatin, we performed DHS mapping for the −5.9/+4.0 region.
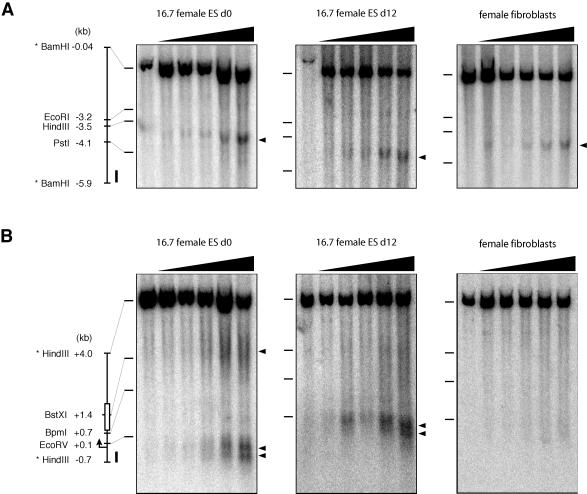
We found several novel DHS in this region. First, within the upstream half of the bipartite enhancer, a single DHS was observed approximately −3.7 to −3.6 kb upstream of the Tsix promoter in both undifferentiated male and female ES cells (Fig. 6A and data not shown). This DHS was also present in differentiating ES cells and in fibroblasts, indicating that it is constitutive. Second, within the −0.7/+4.0 region, analysis of undifferentiated female and male ES cells revealed three DHS. Two were observed in a prominent doublet upstream of the EcoRV site at bp +144, corresponding to the two start sites of the major Tsix promoter (Fig. 6B and data not shown). This doublet was also present in differentiating ES cells, where Tsix expression is reduced, but was absent in primary fibroblasts, where Tsix expression is silenced. Thus, hypersensitivity at the Tsix promoter correlates with the pattern of expression during differentiation.
FIG. 6.
DHS mapping in the bipartite enhancer. Sequence schematics of analyzed regions are shown at the left. The positions of DXPas34 (white rectangle), the major Tsix promoter, and hybridization probes (black rectangles) are indicated. Mobilities of control DNA fragments terminating at the indicated restriction sites are shown to the left of each panel by horizontal tick marks. Positions of hypersensitive sites are shown to the right of each panel by arrowheads. From left to right, the DNase concentrations used for ES cells were 0, 0.5, 1, 2.5, 5, and 10 μg/ml and those used for fibroblasts were 0, 5, 10, 20, 40, and 80 μg/ml. (A) DHS mapping from −5.9 to −0.04 kb. (B) DHS mapping from −0.7 to +4.0 kb.
The third DHS was located downstream of the promoter at kb +1.4, approximately within or just downstream of DXPas34. The band corresponding to this DHS was consistently diffuse in female ES cells, perhaps because of the length polymorphism in DXPas34 between the 129 allele and the Mus. castaneus allele (3). Consistent with this interpretation, the DHS band in male ES cells (carrying only the 129 X chromosome) was consistently more distinct (data not shown). The DHS was observed in undifferentiated female and male ES cells, was markedly reduced in differentiated ES cells, and was undetectable in fibroblasts (Fig. 6B and data not shown). These results indicate that the downstream half of the bipartite enhancer is associated with specialized chromatin that changes dynamically with cell differentiation and the onset of XCI. Interestingly, the DHS at DXPas34 was lost before that of the promoter (Fig. 6B), supporting a model in which the bipartite enhancer regulates the activity of the Tsix promoter in vivo.
Taken together with previous results (30), these findings map eight DHS in the Tsix-Xite region from kb +4.0 to −12.0. While two map to the major Tsix promoter, the remainder correspond to regions that are associated with transcriptional enhancer elements. All but one of these DHS display developmentally specific changes that correlate with the pattern of Tsix expression during differentiation. We propose that the dynamic changes in Tsix transcription are regulated by the coordinate action of enhancer elements that span the Xite and Tsix loci.
DISCUSSION
Novel transcriptional switches for Tsix.
The initiation of XCI in female cells is controlled by antagonistic interactions between Xist, the trigger for chromosomal silencing, and Tsix, its antisense regulator. During cell differentiation, this molecular interplay leads to opposite fates on the two X chromosomes: on the Xa chromosome, the persistent expression of Tsix prevents the spread of Xist RNA, and on the Xi chromosome, the extinction of Tsix expression allows Xist RNA to initiate the formation of heterochromatin. Thus, the control of XCI depends upon the mechanisms that regulate Tsix expression during differentiation. In the present work, we have identified the first enhancers at the X chromosome inactivation center (Fig. 7A). These enhancers constitute novel activities which act as developmentally appropriate transcriptional switches for Tsix.
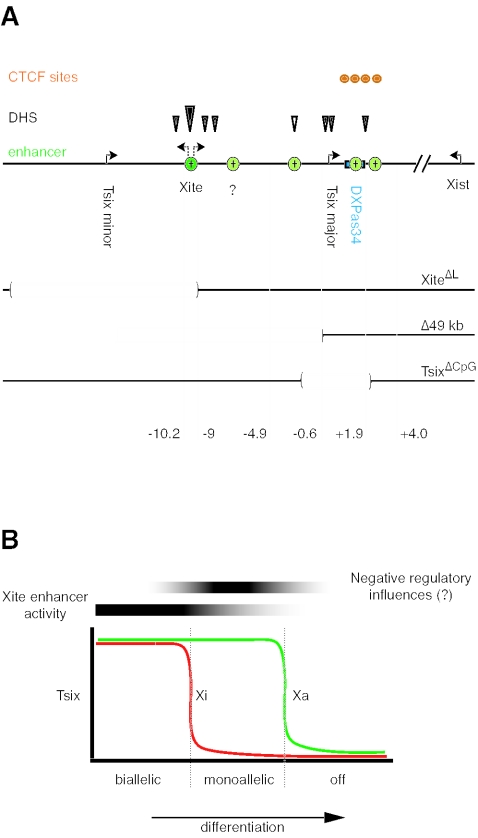
FIG. 7.
Tsix control elements and Tsix binary switch model. (A) Summary of Tsix transcriptional regulatory sequences known to date. Embryonic specific stimulatory elements are shown in the context of genes, functional elements, and selected deletions in Xic. The novel enhancers are shown in green: dark green, 1.2-kb enhancer; light green, bipartite enhancer with multiple elements therein. The exact position of the light green element marked by a question mark is not known. Circles, CTCF elements; closed arrowheads, ES-specific hypersensitive sites; open arrowheads, constitutive DHS; largest arrowhead, strongest DHS. (B) Model for the binary switch through positive and negative control of Tsix expression. In this model, the Xite enhancer promotes the persistence of Tsix expression in cis but does not exert a major influence on the quantitative level of Tsix expression. The activity of the Xite enhancer is lost gradually as differentiation proceeds and, in conjunction with the induction of unidentified negative regulatory influences, yields the asynchronous silencing of Tsix expression (21).
One enhancer maps within Xite and appears to behave as a classical enhancer. The second lies at the 5′ end of Tsix and possesses a bipartite structure. While each is a self-sufficient enhancer unit, the combination of the two results in greater stimulation of the Tsix promoter. DHS mapping reveals that the 16-kb domain from kb −12.0 to +4.0 around the Tsix promoter is replete with DHS, suggesting the occurrence of a specialized chromatin structure. Therefore, we propose that Tsix is coordinately regulated by enhancer activities in this 16-kb domain. These enhancers work together to establish proper kinetics for Tsix expression and reciprocal behavior of its two alleles—that is, persistence of Tsix on the future Xa chromosome and extinction on the future Xi chromosome.
A potential mechanism of enhancer action is promotion of the persistence of Tsix expression.
At what level might these enhancers work on Tsix? Several studies of the Xite and Tsix loci provide clues to their molecular function. A 12-kb deletion of Xite that encompasses the 1.2-kb enhancer does not significantly affect Tsix expression in undifferentiated ES cells but causes the premature extinction of Tsix expression in cis during cell differentiation and the onset of XCI (30). This analysis suggests important mechanistic attributes of the Xite enhancer. First, Tsix expression does not require the Xite enhancer in undifferentiated ES cells, despite the fact that the enhancer appears to be active in these cells, as indicated by transfection and DHS analysis. Second, the Xite enhancer functions to promote the persistence of Tsix expression rather than to increase the absolute level of Tsix expression. Together, these findings suggest that the environment for Tsix expression becomes progressively more restrictive as differentiation proceeds, with the Xite enhancer functioning to counteract these restrictive influences and promote the monoallelic persistence of Tsix expression in cis (Fig. 7B).
Although specific knockouts of the bipartite enhancer have not been made, several deletions have included DXPas34 and downstream portions of the enhancer (19, 22, 35). Although these deletions include the major Tsix promoter, they cause skewed XCI, with a bias toward inactivating the mutated X chromosome, a phenotype that is broadly consistent with the proposed role of DXPas34 and downstream elements in positively regulating Tsix expression. In a 49-kb knockout in which the upstream half of the bipartite enhancer and all of Xite are deleted (29), XCI is also biased in favor of silencing the mutated X chromosome, supporting a role for both regions in regulating Tsix expression. Significantly, the 49-kb deletion displays a more severe bias in the skewing of XCI than does the deletion of Xite alone, suggesting that the Xite and bipartite enhancers each contribute to promoting the persistence of Tsix expression. Clearly, however, knockouts of specific elements in the bipartite enhancer, including DXPas34, are required to confirm this function and to determine whether the bipartite enhancer functions in a manner similar to that of the Xite enhancer.
Activator versus insulator models.
The identification of CTCF-binding sites within and around DXPas34 has suggested two potential (but not mutually exclusive) mechanisms of action (8). One model of Tsix-Xist regulation invokes enhancer competition, in which the two genes compete for access to a shared enhancer (8). In this model, differential access on the two X chromosomes is determined by a chromatin insulator (CTCF), whose allele-specific binding to Tsix on the future Xa chromosome blocks the shared enhancer from the Xist promoter. Here we tested whether the 1.2-kb Xite enhancer could be the hypothetical shared enhancer and found that it could not stimulate Xist promoters in transient assays, suggesting that the 1.2-kb Xite enhancer might be specific for Tsix, at least in the context of these experiments. With other DHS in the region, it is possible that the 1.2-kb region (which spans just one DHS) is not sufficient to recapitulate full enhancer function and that, through the inclusion of additional sequences, the Xite enhancer actually stimulates Xist as well as Tsix. It is also possible that the shared enhancer lies elsewhere in Xic.
The location and structure of the bipartite enhancer at the 5′ end of Tsix make it unlikely to be the shared enhancer. While mammalian enhancers tend to be intronic or distinct from the structural genes that they stimulate, the Tsix enhancer embodies about 9 kb of sequence around the promoter itself. Most curiously, the downstream half of the bipartite enhancer actually overlaps the proposed CTCF insulator domain in Tsix (Fig. 7A). This finding raises several points. First, in the insulator-mediated enhancer competition model, the insulator must lie between the promoters of the two competing genes and the enhancer must lie on the other side of either promoter (8). Thus, the placement of the bipartite enhancer around the Tsix promoter violates the requirement for distal placement, making it an improbable shared enhancer. Second, the overlap of CTCF-binding sites in Tsix poses a certain quandary: what is the functional consequence of juxtaposing enhancer and insulator activities? In Tsix, CTCF-binding sites are dispersed across the region from kb +0.7 to +4.0, with DXPas34 itself harboring some of these elements. As proposed previously for Tsix, CTCF might have both activating and insulating properties (8)—a scenario similar to what has been postulated for CTCF action at H19 (12). The juxtaposition of an enhancer and an insulator has also been described for the β-globin locus (5). At Tsix, this configuration might be responsible for some observed orientation-dependent effects. For example, while DXPas34 works well as a chromatin insulator in one orientation, it has a relatively weaker effect in the opposite orientation (8). Juxtaposition of insulator and enhancer sequences might impart directional character to enhancer action. This architecture might be integral to the mechanism of Tsix control—a possibility to be tested in the future.
A second model of Tsix-Xist regulation envisions CTCF as a direct transcriptional activator of Tsix rather than as a chromatin insulator between Tsix and Xist (8). In other systems, CTCF apparently has direct activating potential (40, 41). The data accrued here are consistent with such a model for XCI regulation. Indeed, we have shown that the bipartite element (in which the CTCF-binding sites reside) is a transcriptional enhancer of Tsix. Given its context dependence, this enhancer appears to be highly specific for Tsix. Given the similar Tsix specificity of the Xite enhancer, we propose that the two enhancers might actually work synergistically to bolster Tsix expression. Furthermore, the activator or enhancer must work allele specifically so that stimulation takes place only on the future Xa chromosome. The activator model is also consistent with other lines of evidence. For example, deletions which encompass one or both enhancers (9, 28-30) compromise Tsix expression and favor Xist expression in cis. Importantly, however, while the data here are consistent with an activator model, the activator and insulator models are not mutually exclusive. Both mechanisms might be required to set up the critical pattern of cis regulation between Tsix and Xist.
The binary switch, X chromosome counting, and epigenetic choice.
With the delineation of activating sequences, our present work addresses how Tsix expression is preserved on the future Xa chromosome. However, the constitutive activity of the major Tsix promoter indicates that cis-acting silencers or other repressive mechanisms must also exist (Fig. 7B). This notion is further supported by the fact that Tsix silencing is obligatory for Xist upregulation on the future Xi chromosome (24, 38). Clearly, to gain a full understanding of Tsix regulation, future work must address how Tsix is turned off on the Xi chromosome. We suggest that enhancer action on the future Xa chromosome, together with negative regulatory influences, comprises the binary switch for X chromosome activity.
The molecular bases of X chromosome counting and choice remain two major goals of ongoing research. X chromosome counting elements have been shown to reside in a 37-kb region downstream of Xist that includes Tsix and Xite (29). In particular, a comparison of various deletion mutants has further suggested that counting elements are located within a discontinuous 20-kb sequence (29) that overlaps the bipartite enhancer. Thus, the bipartite enhancer may function in X chromosome counting. Similarly, elements involved in X chromosome choice have been argued to reside in Xite (30), in DXPas34 and the region flanking the major Tsix promoter (20, 22, 37), and in Xist itself (26, 32). Indeed, we believe that counting, choice, and the Tsix binary switch are tightly coupled events and predict that the counting and choice mechanisms will be revealed through further dissection of the enhancers identified here.
Acknowledgments
We thank the members of the Lee Laboratory for comments on the manuscript and discussions.
This work was supported by Howard Hughes Medical Institute (HHMI) and Ryan Foundation predoctoral fellowships (N.S.), the MGH Fund for Medical Discovery (R.K.R), and the National Institutes of Health and Pew Scholars Program (J.T.L). J.T.L. is an assistant investigator of the HHMI.
REFERENCES
- 1.Ausubel, F. M., et al. (ed.). 1995. Current protocols in molecular biology. John Wiley & Sons, Inc., New York, N.Y.
- 2.Avner, P., and E. Heard. 2001. X-chromosome inactivation: counting, choice and initiation. Nat. Rev. Genet. 2:59-67. [DOI] [PubMed] [Google Scholar]
- 3.Avner, P., M. Prissette, D. Arnaud, B. Courtier, C. Cecchi, and E. Heard. 1998. Molecular correlates of the murine Xce locus. Genet. Res. 72:217-224. [DOI] [PubMed] [Google Scholar]
- 4.Bell, A. C., and G. Felsenfeld. 2000. Methylation of a CTCF-dependent boundary controls imprinted expression of the Igf2 gene. Nature 405:482-485. [DOI] [PubMed] [Google Scholar]
- 5.Bell, A. C., A. G. West, and G. Felsenfeld. 1999. The protein CTCF is required for the enhancer blocking activity of vertebrate insulators. Cell 98:387-396. [DOI] [PubMed] [Google Scholar]
- 6.Brockdorff, N., A. Ashworth, G. F. Kay, V. M. McCabe, D. P. Norris, P. J. Cooper, S. Swift, and S. Rastan. 1992. The product of the mouse Xist gene is a 15 kb inactive X-specific transcript containing no conserved ORF and located in the nucleus. Cell 71:515-526. [DOI] [PubMed] [Google Scholar]
- 7.Brown, C. J., B. D. Hendrich, J. L. Rupert, R. G. Lafreniere, Y. Xing, J. Lawrence, and H. F. Willard. 1992. The human XIST gene: analysis of a 17 kb inactive X-specific RNA that contains conserved repeats and is highly localized within the nucleus. Cell 71:527-542. [DOI] [PubMed] [Google Scholar]
- 8.Chao, W., K. D. Huynh, R. J. Spencer, L. S. Davidow, and J. T. Lee. 2002. CTCF, a candidate trans-acting factor for X-inactivation choice. Science 295:345-347. [DOI] [PubMed] [Google Scholar]
- 9.Clerc, P., and P. Avner. 1998. Role of the region 3′ to Xist exon 6 in the counting process of X-chromosome inactivation. Nat. Genet. 19:249-253. [DOI] [PubMed] [Google Scholar]
- 10.Courtier, B., E. Heard, and P. Avner. 1995. Xce haplotypes show modified methylation in a region of the active X chromosome lying 3′ to Xist. Proc. Natl. Acad. Sci. USA 92:3531-3535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Debrand, E., C. Chureau, D. Arnaud, P. Avner, and E. Heard. 1999. Functional analysis of the DXPas34 locus, a 3′ regulator of Xist expression. Mol. Cell. Biol. 19:8513-8525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Engel, N., A. G. West, G. Felsenfeld, and M. S. Bartolomei. 2004. Antagonism between DNA hypermethylation and enhancer-blocking activity at the H19 DMD is uncovered by CpG mutations. Nat. Genet. 36:883-888. [DOI] [PubMed] [Google Scholar]
- 13.Forrester, W. C., E. Epner, M. C. Driscoll, T. Enver, M. Brice, T. Papayannopoulou, and M. Groudine. 1990. A deletion of the human beta-globin locus activation region causes a major alteration in chromatin structure and replication across the entire beta-globin locus. Genes Dev. 4:1637-1649. [DOI] [PubMed] [Google Scholar]
- 14.Hark, A. T., C. J. Schoenherr, D. J. Katz, R. S. Ingram, J. M. Levorse, and S. M. Tilghman. 2000. CTCF mediates methylation-sensitive enhancer-blocking activity at the H19/Igf2 locus. Nature 405:486-489. [DOI] [PubMed] [Google Scholar]
- 15.Johnston, C. M., T. B. Nesterova, E. J. Formstone, A. E. Newall, S. M. Duthie, S. A. Sheardown, and N. Brockdorff. 1998. Developmentally regulated Xist promoter switch mediates initiation of X inactivation. Cell 94:809-817. [DOI] [PubMed] [Google Scholar]
- 16.Kanduri, C., C. Holmgren, M. Pilartz, G. Franklin, M. Kanduri, L. Liu, V. Ginjala, E. Ulleras, R. Mattsson, and R. Ohlsson. 2000. The 5′ flank of mouse H19 in an unusual chromatin conformation unidirectionally blocks enhancer-promoter communication. Curr. Biol. 10:449-457. [DOI] [PubMed] [Google Scholar]
- 17.Kanduri, C., V. Pant, D. Loukinov, E. Pugacheva, C. F. Qi, A. Wolffe, R. Ohlsson, and V. V. Lobanenkov. 2000. Functional association of CTCF with the insulator upstream of the H19 gene is parent of origin-specific and methylation-sensitive. Curr. Biol. 10:853-856. [DOI] [PubMed] [Google Scholar]
- 18.Kreiss, P., B. Cameron, R. Rangara, P. Mailhe, O. Aguerre-Charriol, M. Airiau, D. Scherman, J. Crouzet, and B. Pitard. 1999. Plasmid DNA size does not affect the physicochemical properties of lipoplexes but modulates gene transfer efficiency. Nucleic Acids Res. 27:3792-3798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Lee, J. T. 2000. Disruption of imprinted X inactivation by parent-of-origin effects at Tsix. Cell 103:17-27. [DOI] [PubMed] [Google Scholar]
- 20.Lee, J. T. 2002. Homozygous Tsix mutant mice reveal a sex-ratio distortion and revert to random X-inactivation. Nat. Genet. 32:195-200. [DOI] [PubMed] [Google Scholar]
- 21.Lee, J. T., L. S. Davidow, and D. Warshawsky. 1999. Tsix, a gene antisense to Xist at the X-inactivation centre. Nat. Genet. 21:400-404. [DOI] [PubMed] [Google Scholar]
- 22.Lee, J. T., and N. Lu. 1999. Targeted mutagenesis of Tsix leads to nonrandom X inactivation. Cell 99:47-57. [DOI] [PubMed] [Google Scholar]
- 23.Lee, J. T., W. M. Strauss, J. A. Dausman, and R. Jaenisch. 1996. A 450 kb transgene displays properties of the mammalian X-inactivation center. Cell 86:83-94. [DOI] [PubMed] [Google Scholar]
- 24.Luikenhuis, S., A. Wutz, and R. Jaenisch. 2001. Antisense transcription through the Xist locus mediates Tsix function in embryonic stem cells. Mol. Cell. Biol. 21:8512-8520. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Lyon, M. F. 1961. Gene action in the X-chromosome of the mouse (Mus musculus L.). Nature 190:372-373. [DOI] [PubMed] [Google Scholar]
- 26.Marahrens, Y., J. Loring, and R. Jaenisch. 1998. Role of the Xist gene in X chromosome choosing. Cell 92:657-664. [DOI] [PubMed] [Google Scholar]
- 27.Marahrens, Y., B. Panning, J. Dausman, W. Strauss, and R. Jaenisch. 1997. Xist-deficient mice are defective in dosage compensation but not spermatogenesis. Genes Dev. 11:156-166. [DOI] [PubMed] [Google Scholar]
- 28.Morey, C., D. Arnaud, P. Avner, and P. Clerc. 2001. Tsix-mediated repression of Xist accumulation is not sufficient for normal random X inactivation. Hum. Mol. Genet. 10:1403-1411. [DOI] [PubMed] [Google Scholar]
- 29.Morey, C., P. Navarro, E. Debrand, P. Avner, C. Rougeulle, and P. Clerc. 2004. The region 3′ to Xist mediates X chromosome counting and H3 Lys-4 dimethylation within the Xist gene. EMBO J. 23:594-604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Ogawa, Y., and J. T. Lee. 2003. Xite, X-inactivation intergenic transcription elements that regulate the probability of choice. Mol. Cell 11:731-743. [DOI] [PubMed] [Google Scholar]
- 31.Penny, G. D., G. F. Kay, S. A. Sheardown, S. Rastan, and N. Brockdorff. 1996. Requirement for Xist in X chromosome inactivation. Nature 379:131-137. [DOI] [PubMed] [Google Scholar]
- 32.Plenge, R. M., R. A. Stevenson, H. A. Lubs, C. E. Schwartz, and H. F. Willard. 2002. Skewed X-chromosome inactivation is a common feature of X-linked mental retardation disorders. Am. J. Hum. Genet. 71:168-173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Rastan, S. 1983. Non-random X-chromosome inactivation in mouse X-autosome translocation embryos—location of the inactivation centre. J. Embryol. Exp. Morphol. 78:1-22. [PubMed] [Google Scholar]
- 34.Rastan, S., and E. J. Robertson. 1985. X-chromosome deletions in embryo-derived (EK) cell lines associated with lack of X-chromosome inactivation. J. Embryol. Exp. Morphol. 90:379-388. [PubMed] [Google Scholar]
- 35.Sado, T., Z. Wang, H. Sasaki, and E. Li. 2001. Regulation of imprinted X-chromosome inactivation in mice by Tsix. Development 128:1275-1286. [DOI] [PubMed] [Google Scholar]
- 36.Shibata, S., and J. T. Lee. 2003. Characterization and quantitation of differential Tsix transcripts: implications for Tsix function. Hum. Mol. Genet. 12:125-136. [DOI] [PubMed] [Google Scholar]
- 37.Shibata, S., and J. T. Lee. 2004. Tsix RNA- versus transcription-based mechanisms in Xist repression and epigenetic choice. Curr. Biol. 14:1747-1754. [DOI] [PubMed] [Google Scholar]
- 38.Stavropoulos, N., N. Lu, and J. T. Lee. 2001. A functional role for Tsix transcription in blocking Xist RNA accumulation but not in X-chromosome choice. Proc. Natl. Acad. Sci. USA 98:10232-10237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Turner, D. L., and H. Weintraub. 1994. Expression of achaete-scute homolog 3 in Xenopus embryos converts ectodermal cells to a neural fate. Genes Dev. 8:1434-1447. [DOI] [PubMed] [Google Scholar]
- 40.Vostrov, A. A., and W. W. Quitschke. 1997. The zinc finger protein CTCF binds to the APBbeta domain of the amyloid beta-protein precursor promoter. Evidence for a role in transcriptional activation. J. Biol. Chem. 272:33353-33359. [DOI] [PubMed] [Google Scholar]
- 41.Yang, Y., W. W. Quitschke, A. A. Vostrov, and G. J. Brewer. 1999. CTCF is essential for up-regulating expression from the amyloid precursor protein promoter during differentiation of primary hippocampal neurons. J. Neurochem. 73:2286-2298. [DOI] [PubMed] [Google Scholar]