Abstract
We present a genome assembly from an individual male Anopheles moucheti (the malaria mosquito; Arthropoda; Insecta; Diptera; Culicidae), from a wild population in Cameroon. The genome sequence is 271 megabases in span. The majority of the assembly is scaffolded into three chromosomal pseudomolecules with the X sex chromosome assembled. The complete mitochondrial genome was also assembled and is 15.5 kilobases in length.
Keywords: Anopheles moucheti, African malaria mosquito, genome sequence, chromosomal
Species taxonomy
Animalia; Arthropoda; Insecta; Diptera; Culicidae; Anophelinae; Anopheles; Anopheles moucheti; Evans, 1925 (NCBI txid:186751).
Background
Anopheles moucheti moucheti (hereafter An. moucheti) has the greatest geographical distribution within the Moucheti group, which includes An. bervoetsi and An. nigeriensis 1, 2 . These species can only be distinguished by slight morphological characters 3 and / or using molecular tools 4, 5 . An. moucheti is widely distributed across forested areas of West and Central Africa, and the persistence of this species is linked to its ability to lay eggs in lentic streams and rivers where its larvae develop preferentially and predominate over other malaria vectors 1, 6 . An. moucheti females usually feed indoors and exhibit a preference for feeding on humans 7 although the species can also be found in forested areas of Gabon, away from any human presence 8, 9 . An. moucheti is among the most important vectors of human malaria in the equatorial rain forest of Africa 1, 10 . It is present in a large number of countries, from Guinea to Kenya 11 , and it has been found naturally infected with Plasmodium parasites across Central African countries, sustaining year-round transmission 1 . For instance, it can show infective biting rates up to 300 bites per person per year in villages located along slow moving rivers 7, 10 . Moreover, An. moucheti has been recently incriminated for contributing to the origin of malaria in humans, being a potential bridge vector for malaria parasites from non-human primates to humans 8, 9 .
The main genetic work carried out on An. moucheti has been focused on differentiation between members of the group. To this end, molecular markers have been developed that are based on the sequence variation in the ribosomal internal transcribed spacer ITS1 region 5 . Microsatellite loci have also been developed showing a low level of genetic differentiation among populations in Cameroon 5, 12 . Several polymorphic inversions have been detected in An. moucheti, although their role in local adaptation or speciation is not yet understood 13 . Therefore, genomic characterization of this neglected but major malaria vector, An. moucheti, is critical for developing sustainable control strategies for malaria elimination in Central Africa.
The genome of the African malaria mosquito, Anopheles moucheti, was sequenced as part of the Anopheles Reference Genomes Project (PRJEB51690). Here we present a chromosomally complete genome sequence for Anopheles moucheti, based on a single male specimen from Ebogo, Cameroon.
Genome sequence report
The genome was sequenced from a single male Anopheles moucheti collected from Ebogo, Cameroon (3.384, 11.466). A total of 41-fold coverage in Pacific Biosciences single-molecule HiFi long reads (N50 13.064 kb) were generated. Primary assembly contigs were scaffolded with chromosome conformation Hi-C data from a sibling male mosquito. Manual assembly curation corrected 48 missing joins or misjoins and removed 3 haplotypic duplications, reducing the scaffold number by 7.8% and reducing assembly size by 1.8%.
The final assembly has a total length of 271 Mb in 345 sequence scaffolds with a scaffold N50 of 92.649 Mb ( Table 1). 84.7% of the assembly sequence was assigned to three chromosomal-level scaffolds, representing two autosomes (numbered and oriented against the An. gambiae PEST strain assembly AgamP3 ( 14; GCF_000005575.2) with chromosome arms homologies for Myzomyia series taken from 15), and the X sex chromosome ( Figure 1– Figure 6; Table 2).
Figure 1. Snail plot summary of assembly statistics for Anopheles moucheti assembly idAnoMoucSN_F20_07.

The main plot is divided into 1,000 size-ordered bins around the circumference with each bin representing 0.1% of the 271,332,225 bp assembly. The distribution of sequence lengths is shown in dark grey with the plot radius scaled to the longest sequence present in the assembly (103,511,506 bp, shown in red). Orange and pale-orange arcs show the N50 and N90 sequence lengths (92,649,126 and 283,162 bp), respectively. The pale grey spiral shows the cumulative sequence count on a log scale with white scale lines showing successive orders of magnitude. The blue and pale-blue area around the outside of the plot shows the distribution of GC, AT and N percentages in the same bins as the inner plot. A summary of complete, fragmented, duplicated and missing BUSCO genes in the diptera_odb10 set is shown in the top right. An interactive version of this figure is available at https://blobtoolkit.genomehubs.org/view/idAnoMoucSN_F20_07/dataset/CALSEA01/snail.
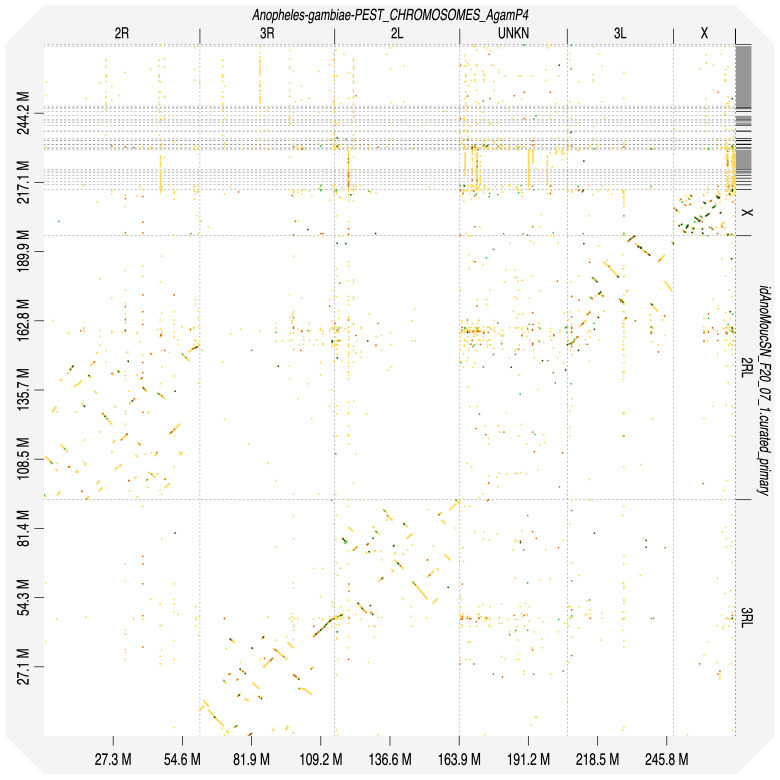
Figure 6. Alignment dotplot between genome assemblies of An. moucheti (idAnoMoucSN_F20_07) and An. gambiae, AgamP4 (PEST).
Visualised in DGenies. Chromosome arms correspondence (moucheti-gambiae): 2R-2R, 2L-3L, 3R-3R, 3L-2L in agreement with 15.
Table 1. Genome data for Anopheles moucheti, idAnoMoucSN_F20_07.
| Project accession data | |
|---|---|
| Assembly identifier | idAnoMoucSN_F20_07 |
| Species | Anopheles moucheti |
| Specimen | idAnoMoucSN-F20_07 |
| NCBI taxonomy ID | 186751 |
| BioProject | PRJEB53257 |
| BioSample ID | ERS10527373 |
| Isolate information | male, whole organism |
| Raw data accessions | |
| PacificBiosciences SEQUEL II | ERR9439507 |
| Hi-C Illumina | ERR9356830 |
| PolyA RNA-Seq Illumina | ERR9356831, ERR9356832 |
| Genome assembly | |
| Assembly accession | GCA_943734755 |
| Accession of alternate haplotype | GCA_943734785 |
| Span (Mb) | 271.317 |
| Number of contigs | 406 |
| Contig N50 length (Mb) | 4.744 |
| Number of scaffolds | 345 |
| Scaffold N50 length (Mb) | 92.649 |
| Longest scaffold (Mb) | 103.512 |
| BUSCO * genome score | C:97.6%[S:97.3%,D:0.3%],
F:0.7%,M:2.0%,n:3285 |
* BUSCO scores based on the diptera_odb10 BUSCO set using BUSCO 5.3.2. C=complete [S=single copy, D=duplicated], F=fragmented, M=missing, n=number of orthologues in comparison. A full set of BUSCO scores is available at https://blobtoolkit.genomehubs.org/view/idAnoMoucSN_F20_07/dataset/CALSEA01/busco.
Figure 2. Blob plot of base coverage in idAnoMoucSN-F20_07 PacBio HiFi reads against GC proportion for An. moucheti assembly idAnoMoucSN_F20_07.
Chromosomes are coloured by phylum. Circles are sized in proportion to chromosome length. Histograms show the distribution of chromosome length sum along each axis. An interactive version of this figure is available at https://blobtoolkit.genomehubs.org/view/idAnoMoucSN_F20_07/dataset/CALSEA01/blob.
Figure 3. Cumulative chromosome length for An. moucheti assembly idAnoMoucSN_F20_07.
The grey line shows cumulative length for all chromosomes. Coloured lines show cumulative lengths of chromosomes assigned to each phylum using the buscogenes taxrule. The interactive version of this figure is available at https://blobtoolkit.genomehubs.org/view/idAnoMoucSN_F20_07/dataset/CALSEA01/cumulative.
Figure 4. Genome assembly of An. moucheti, idAnoMoucSN_F20_07: Hi-C contact map.
Visualised in HiGlass. Chromosomes are arranged in size order from left to right and top to bottom. X chromosome signal is lower as individuals used for both PacBio and HiC are males. The interactive Hi-C map can be viewed at https://genome-note-higlass.tol.sanger.ac.uk/l/?d=GZ6DShODRcSMNgw7a0UIfQ.
Figure 5. Distribution plot of repeat masked sequence proportion against genomic position for chromosomes of An. moucheti, idAnoMoucSN_F20_07.
1Mb windows with assembly gaps highlighted by larger blue circles, size is proportional to total size of gaps in the window (max 7800bp per window).
Table 2. Chromosomal pseudomolecules in the genome assembly of An. moucheti, idAnoMoucSN_F20_07.
| INSDC
accession |
Chromosome | Size (Mb) | Count | Gaps |
|---|---|---|---|---|
| OX030915.1 | 2RL | 103.512 | 1 | 30 |
| OX030916.1 | 3RL | 92.649 | 1 | 15 |
| OX030917.1 | X | 18.119 | 1 | 6 |
| OX030918.1 | MT | 0.016 | 1 | 0 |
| X Unlocalised | 15.547 | 98 | 0 | |
| Unplaced | 41.489 | 244 | 10 |
The assembly has a BUSCO 5.3.2 16 completeness of 97.6% using the diptera_odb10 reference set. While not fully phased, the assembly deposited is of one haplotype. Contigs corresponding to the second haplotype have also been deposited.
Methods
Sample acquisition and nucleic acid extraction
Anopheles moucheti offspring were reared from a wild caught gravid female collected from Ebogo, Cameroon (3.384, 11.466) by Sandrine Eveline Nsango. A single male idAnoMoucSN-F20_07 was used for Pacific BioSciences, a single sibling male idAnoMoucSN-F20_09 was used for Arima Hi-C.
For high molecular weight (HMW) DNA extraction one whole insect (idAnoMoucSN-F20_07) was disrupted by manual grinding with a blue plastic pestle in Qiagen MagAttract lysis buffer 17 and then extracted using the Qiagen MagAttract HMW DNA extraction kit with two minor modifications including halving volumes recommended by the manufacturer due to small sample size ( Anopheles mosquitoes typically weigh 2–3 mg) and running two elution steps of 100 μl each to increase DNA yield. The quality of the DNA was evaluated using an Agilent FemtoPulse to ensure that most DNA molecules were larger than 30 kb, and preferably > 100 kb. In general, single mosquito extractions ranged in total estimated DNA yield from 200 ng to 800 ng, with an average yield of 500 ng. Low molecular weight DNA was removed using 0.8X AMpure XP purification. DNA was sheared to an average fragment size of 12–20 kb using a Diagenode Megaruptor 3 at speeds ranging from 27 to 30. Sheared DNA was purified using AMPure PB beads with a 1.8X ratio of beads to sample to remove the shorter fragments and concentrate the DNA sample. The concentration and quality of the sheared and purified DNA was assessed using a Nanodrop spectrophotometer and Qubit Fluorometer with the Qubit dsDNA High Sensitivity Assay kit. Fragment size distribution was evaluated by running the sheared and cleaned sample on the FemtoPulse system once more. The median DNA fragment size for Anopheles mosquitoes was 15 kb and the median yield of sheared DNA was 200 ng, with samples typically losing about 50% of the original estimated DNA quantity through the process of shearing and purification.
For Hi-C data generation, a sibling male mosquito specimen (idAnoMoucSN-F20_09) was used as input material for the Arima V2 Kit according to the manufacturer’s instructions for animal tissue. This approach of using a sibling was followed in order to enable all material from a single specimen to contribute to the PacBio data generation given we were not always able to meet the minimum suggested guidance of starting with > 300 ng of HMW DNA from a specimen. Samples proceeded to the Illumina library prep stage even if they were suboptimal (too little tissue) going into the Arima reaction.
To assist with annotation, which will be made available through VectorBase 18 in due course, RNA was extracted from separate whole unrelated lab-reared male and wild-caught female insect specimens (idAnoMoucSN-F4_01, idAnoMoucSN-F7_x) using TRIzol, according to the manufacturer’s instructions. RNA was then eluted in 50 μl RNAse-free water, and its concentration was assessed using a Nanodrop spectrophotometer and Qubit Fluorometer using the Qubit RNA Broad-Range (BR) Assay kit. Analysis of the integrity of the RNA was done using Agilent RNA 6000 Pico Kit and Eukaryotic Total RNA assay. Samples were not always ideally preserved for RNA, so qualities varied but all were sequenced anyway.
Sequencing
We prepared libraries as per the PacBio procedure and checklist for SMRTbell Libraries using Express TPK 2.0 with low DNA input. Every library was barcoded to support multiplexing. Final library yields ranged from 20 ng to 100 ng, representing only about 25% of the input sheared DNA. Libraries from two specimens were typically multiplexed on a single 8M SMRT Cell. Sequencing complexes were made using Sequencing Primer v4 and DNA Polymerase v2.0. Sequencing was carried out on the Sequel II system with 24-hour run time and 2-hour pre-extension. For Hi-C data generation, following the Arima Hi-C V2 reaction, samples were processed through Library Preparation using a NEB Next Ultra II DNA Library Prep Kit and sequenced aiming for 100x depth. RNA libraries were created using the directional NEB Ultra II stranded kit. Sequencing was performed by the Scientific Operations core at the Wellcome Sanger Institute on Pacific Biosciences SEQUEL II (HiFi), Illumina NovaSeq 6000 (10X and Hi-C), or Illumina HiSeq 4000 (RNAseq) instruments.
Genome assembly
Assembly was carried out with Hifiasm 19 ; haplotypic duplications were identified and removed with purge_dups 20 . The assembly was then scaffolded with Hi-C data 21 using SALSA2 22 . The assembly was checked for contamination as described previously 23 . Manual curation was performed using gEVAL 24 , HiGlass 25 and Pretext 26 . The mitochondrial genome was assembled using MitoHiFi 27 , which performs annotation using MitoFinder 28 . The genome was analysed and BUSCO scores were generated within the BlobToolKit environment 29 . Synteny analysis was performed with D-GENIES 30 and minimap2 31 . Table 3 contains a list of all software tool versions used, where appropriate.
Table 3. Software tools used.
Ethics/compliance issues
The genetic resources accessed and utilised under this project were done so in accordance with the UK ABS legislation (Nagoya Protocol (Compliance) (Amendment) (EU Exit) Regulations 2018 (SI 2018/1393)) and the national ABS legislation within the country of origin, where applicable.
Funding Statement
This work was supported by a Bill & Melinda Gates Foundation Award (INV-009760) to ML. The Wellcome Sanger Institute is funded by the Wellcome Trust (206194), which supports ML. SAM is supported by the Wellcome Trust Grant (WT207492). Field collection was supported by an ANR grant (ANR-18-CE35-0002-01 –WILDING) to DA.
The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
[version 1; peer review: 2 approved]
Data availability
European Nucleotide Archive: Anopheles moucheti genome assembly, idAnoMoucSN_F20_07. Accession number PRJEB53257; https://identifiers.org/bioproject/PRJEB53257.
The genome sequence is released openly for reuse. The Anopheles moucheti genome sequencing initiative is part of the Anopheles Reference Genomes project PRJEB51690. All raw sequence data and the assembly have been deposited in INSDC databases. Raw data and assembly accession identifiers are reported in Table 1.
Author information
Members of the Sanger Scientific Operations: DNA Pipelines collective are listed here: https://doi.org/10.5281/zenodo.4790455.
References
- 1. Antonio-Nkondjio C, Simard F: Highlights on Anopheles nili and Anopheles moucheti, Malaria Vectors in Africa.In: Manguin S editor. Anopheles Mosquitoes: New Insights into Malaria Vectors. Rijeka (HR): InTech;2013. [PubMed] [Google Scholar]
- 2. Harbach RE, Wilkerson RC: The insupportable validity of mosquito subspecies (Diptera: Culicidae) and their exclusion from culicid classification. Zootaxa. 2023;5303(1):1–184. 10.11646/zootaxa.5303.1.1 [DOI] [PubMed] [Google Scholar]
- 3. Gillies MT, De Meillon B: The anophelinae of Africa south of the Sahara (Ethiopian zoogeographical region).South African Institute for Medical Research;1968. Reference Source [Google Scholar]
- 4. Antonio-Nkondjio C, Ndo C, Awono-Ambene P, et al. : Population genetic structure of the malaria vector Anopheles moucheti in south Cameroon forest region. Acta Trop. 2007;101(1):61–68. 10.1016/j.actatropica.2006.12.004 [DOI] [PubMed] [Google Scholar]
- 5. Kengne P, Antonio-Nkondjio C, Awono-Ambene HP, et al. : Molecular differentiation of three closely related members of the mosquito species complex, Anopheles moucheti, by mitochondrial and ribosomal DNA polymorphism. Med Vet Entomol. 2007;21(2):177–182. 10.1111/j.1365-2915.2007.00681.x [DOI] [PubMed] [Google Scholar]
- 6. Ayala D, Costantini C, Ose K, et al. : Habitat suitability and ecological niche profile of major malaria vectors in Cameroon. Malar J. 2009;8: 307. 10.1186/1475-2875-8-307 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Antonio-Nkondjio C, Kerah CH, Simard F, et al. : Complexity of the malaria vectorial system in Cameroon: contribution of secondary vectors to malaria transmission. J Med Entomol. 2006;43(6):1215–1221. 10.1603/0022-2585(2006)43[1215:cotmvs]2.0.co;2 [DOI] [PubMed] [Google Scholar]
- 8. Paupy C, Makanga B, Ollomo B, et al. : Anopheles moucheti and Anopheles vinckei are candidate vectors of ape Plasmodium parasites, including Plasmodium praefalciparum in Gabon. PLoS One. 2013;8(2): e57294. 10.1371/journal.pone.0057294 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Makanga B, Yangari P, Rahola N, et al. : Ape malaria transmission and potential for ape-to-human transfers in Africa. Proc Natl Acad Sci U S A. 2016;113(19):5329–5334. 10.1073/pnas.1603008113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Mouchet J: Biodiversité du paludisme dans le monde.John Libbey Eurotext;2004. Reference Source [Google Scholar]
- 11. Irish SR, Kyalo D, Snow RW, et al. : Updated list of Anopheles species (Diptera: Culicidae) by country in the Afrotropical Region and associated islands. Zootaxa. 2020;4747(3): zootaxa.4747.3.1. 10.11646/zootaxa.4747.3.1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Annan Z, Durand P, Ayala FJ, et al. : Population genetic structure of Plasmodium falciparum in the two main African vectors, Anopheles gambiae and Anopheles funestus. Proc Natl Acad Sci U S A. 2007;104(19):7987–7992. 10.1073/pnas.0702715104 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Sharakhova MV, Antonio-Nkondjio C, Xia A, et al. : Polymorphic chromosomal inversions in Anopheles moucheti, a major malaria vector in Central Africa. Med Vet Entomol. 2014;28(3):337–340. 10.1111/mve.12037 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Sharakhova MV, Hammond MP, Lobo NF, et al. : Update of the Anopheles gambiae PEST genome assembly. Genome Biol. 2007;8(1): R5. 10.1186/gb-2007-8-1-r5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Green CA, Hunt RH: Interpretation of variation in ovarian polytene chromosomes of Anopheles Funestus Giles, A. Parensis Gillies, and A. Aruni? Genetica. 1980;51:187–195. 10.1007/BF00121610 [DOI] [Google Scholar]
- 16. Simão FA, Waterhouse RM, Ioannidis P, et al. : BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics. 2015;31(19):3210–3212. 10.1093/bioinformatics/btv351 [DOI] [PubMed] [Google Scholar]
- 17. Teltscher F, Johnson H, Lawniczak M: Manual extraction of High Molecular Weight DNA from single mosquitoes using the Qiagen MagAttract HMW DNA kit. 2023; [cited 2 Oct 2023]. 10.17504/protocols.io.n92ldp6ool5b/v1 [DOI] [Google Scholar]
- 18. Giraldo-Calderón GI, Harb OS, Kelly SA, et al. : VectorBase.org updates: bioinformatic resources for invertebrate vectors of human pathogens and related organisms. Curr Opin Insect Sci. 2022;50: 100860. 10.1016/j.cois.2021.11.008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Cheng H, Concepcion GT, Feng X, et al. : Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat Methods. 2021;18(2):170–175. 10.1038/s41592-020-01056-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Guan D, McCarthy SA, Wood J, et al. : Identifying and removing haplotypic duplication in primary genome assemblies. Bioinformatics. 2020;36(9):2896–2898. 10.1093/bioinformatics/btaa025 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Rao SSP, Huntley MH, Durand NC, et al. : A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell. 2014;159(7):1665–1680. 10.1016/j.cell.2014.11.021 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Ghurye J, Rhie A, Walenz BP, et al. : Integrating Hi-C links with assembly graphs for chromosome-scale assembly. PLoS Comput Biol. 2019;15(8): e1007273. 10.1371/journal.pcbi.1007273 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Howe K, Chow W, Collins J, et al. : Significantly improving the quality of genome assemblies through curation. GigaScience. Oxford University Press,2021;10(1): giaa153. 10.1093/gigascience/giaa153 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Chow W, Brugger K, Caccamo M, et al. : gEVAL — a web-based browser for evaluating genome assemblies. Bioinformatics. 2016;32(16):2508–10. 10.1093/bioinformatics/btw159 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Kerpedjiev P, Abdennur N, Lekschas F, et al. : HiGlass: web-based visual exploration and analysis of genome interaction maps. Genome Biol. 2018;19(1): 125. 10.1186/s13059-018-1486-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. PretextView: OpenGL Powered Pretext Contact Map Viewer.Github; [Accessed 19 October 2022]. Reference Source
- 27. Uliano-Silva M, Nunes JGRN, Krasheninnikova K, et al. : marcelauliano/MitoHiFi: mitohifi_v2.0..2021. 10.5281/zenodo.5205678 [DOI] [Google Scholar]
- 28. Allio R, Schomaker‐Bastos A, Romiguier J, et al. : MitoFinder: Efficient automated large‐scale extraction of mitogenomic data in target enrichment phylogenomics. Mol Ecol Resour. 2020;20(4):892–905. 10.1111/1755-0998.13160 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Challis R, Richards E, Rajan J, et al. : BlobToolKit - interactive quality assessment of genome assemblies. G3 (Bethesda). 2020;10(4):1361–1374. 10.1534/g3.119.400908 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Cabanettes F, Klopp C: D-GENIES: dot plot large genomes in an interactive, efficient and simple way. PeerJ. 2018;6: e4958. 10.7717/peerj.4958 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Li H: Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics. 2018;34(18):3094–3100. 10.1093/bioinformatics/bty191 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Challis R, Richards E, Rajan J, et al. : BlobToolKit - interactive quality assessment of genome assemblies. G3 (Bethesda). 2020;10(4):1361–1374. 10.1534/g3.119.400908 [DOI] [PMC free article] [PubMed] [Google Scholar]