Summary
Single-cell whole-genome sequencing (scWGS) enables the assessment of genome-level molecular differences between individual cells with particular relevance to genetically diverse systems like solid tumors. The application of scWGS was limited due to a dearth of accessible platforms capable of producing high-throughput profiles. We present a technique that leverages nucleosome disruption methodologies with the widely adopted 10× Genomics ATAC-seq workflow to produce scWGS profiles for high-throughput copy-number analysis without new equipment or custom reagents. We further demonstrate the use of commercially available indexed transposase complexes from ScaleBio for sample multiplexing, reducing the per-sample preparation costs. Finally, we demonstrate that sequential indexed tagmentation with an intervening nucleosome disruption step allows for the generation of both ATAC and WGS data from the same cell, producing comparable data to the unimodal assays. By exclusively utilizing accessible commercial reagents, we anticipate that these scWGS and scWGS+ATAC methods can be broadly adopted by the research community.
Keywords: single-cell genomics, cancer biology, chromatin accessibility, copy-number alterations
Graphical abstract

Highlights
-
•
Integration of single-cell WGS with widely available single-cell ATAC-seq workflows
-
•
Indexed tagmentation enables sample multiplexing for scWGS experiments
-
•
Double tagmentation using indexed transposomes produces paired ATAC and WGS profiles
Motivation
Single-cell whole-genome sequencing (scWGS) is a powerful tool for studying complex, heterogeneous biology. The motivation for this study was to establish an scWGS method that relies only on commercially available reagents and instrumentation to make the technology accessible to the widest possible segment of the research community.
Queitsch et al. report a method for paired, single-cell whole-genome sequencing and ATAC-seq using kits on the widely used 10× Genomics Chromium platform. The method comprises an effective and accessible strategy for examining genomic alterations and tumor subclonal structures, enabling tumor evolution studies that can inform therapeutic development.
Introduction
Single-cell whole-genome sequencing (scWGS) has proven invaluable for the identification of tumor subpopulations and in advancing our understanding of clonal dynamics and tumor evolution,1,2,3,4,5,6 as well as in the study of somatic genomic copy-number alterations in healthy tissues, including the brain.7 Despite the value of scWGS, there is a dearth of commercially available options that provide enough cell throughput (i.e., power) to fully catalog and characterize clonal populations within a tumor sample. This can be particularly problematic when rare, possibly therapy-resistant, subpopulations are present within a tumor that may elude detection using existing methodologies. To compound these challenges, assay availability was further reduced when the widely available 10× Genomics Chromium permanently discontinued its scWGS product.
We previously developed several technologies that leverage in situ disruption of nucleosomes such that nuclei remain intact and can be processed through in situ tagmentation methods typically used for scATAC-seq. The nucleosome disruption, which is performed by fixation and then treatment with a detergent, enables genome-wide access by the Tn5 transposase, thus producing random coverage across the entire genome. In prior technologies, combinatorial barcoding was used to enable single-cell profiling8; however, these methods require boutique reagents in the form of at least 96 tagmentation complexes with unique DNA barcodes, making the technology difficult to access for the majority of potential users. Here, we leverage similar nucleosome disruption methods, in situ tagmentation and cell barcoding, using off-the-shelf kits, with the widely accessible Chromium instrument from 10× Genomics. We further leverage commercially available indexed tagmentation reagents (24-plex), designed for use in scATAC sample multiplexing, to achieve multiplexing of scWGS samples in individual lanes of a Chromium chip. The same indexed tagmentation reagents lend themselves to a novel double-tagmentation technique that enables the production of both ATAC and WGS data from the same single cells that can additionally capitalize on the aforementioned sample multiplexing capabilities.
Results
scWGS using the 10× Chromium scATAC workflow
To achieve high-throughput scWGS using off-the-shelf reagents, we reasoned that the nucleosome disruption technology that we had developed for combinatorial-indexing-based scWGS could be deployed within the scATAC-seq technology workflow that is commercially available from 10× Genomics (Figure 1A). Key to this workflow is the successful disruption of nucleosomes to enable genome-wide access for the Tn5 transposase to fragment DNA and append adapters (tagmentation) and that the nuclei remain intact during the process for subsequent processing using droplet fluidics to index each library fragment with a corresponding cell barcode.
Figure 1.
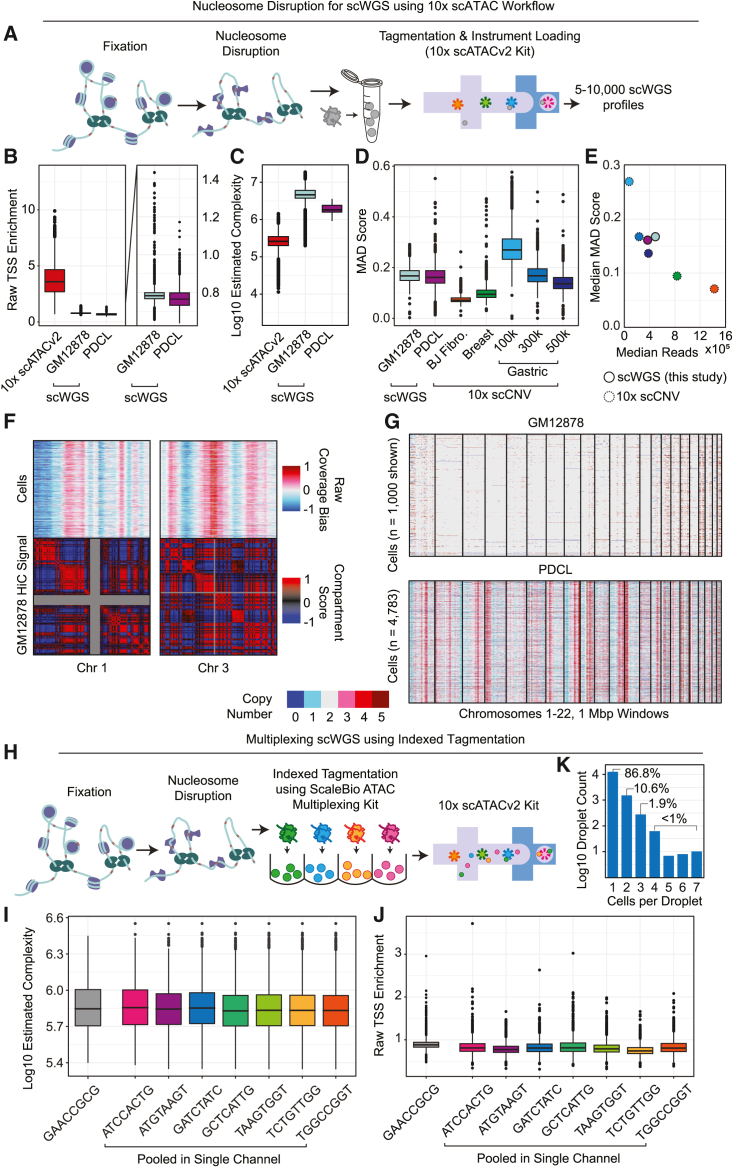
Single-cell WGS using the 10× Genomics scATAC kit
(A) Workflow diagram.
(B) Raw TSS enrichment for each condition. PDCL, patient-derived cell line. Boxes indicate 25th and 75th percentile, whiskers represent 90th and 10th percentile. These boxplot settings are used for all subsequent figures.
(C) Estimated library complexity.
(D) Comparison of MAD scores to assess coverage biases between our technique and the discontinued 10× scCNV kit.
(E) Median MAD scores as a function of median read depth. Colors are as in (D).
(F) Association between raw coverage bias and HiC compartments.
(G) RIDDLER copy-number calls for GM12878 and PDCL samples.
(H) Multiplexing workflow diagram.
(I) Estimated library complexity.
(J) Raw TSS enrichment for each indexed sample.
(K) Cells per droplet assessed by the number of unique tagmentation barcodes associated with each droplet barcode.
We previously described two versions of the nucleosome disruption technique: lithium-assisted nucleosome disruption (LAND) and a method that relies on crosslinking followed by sodium dodecyl sulfate detergent treatment (xSDS).9 We found that LAND, which leverages the chaotropic salt, lithium diiodosalycylate (LIS), was highly variable in its ability to retain nuclear integrity, often resulting in the rupture of all nuclei, likely due to the salt readily crashing out of solution at room temperature, resulting in variable effective concentrations.9 We therefore focused on the xSDS approach, which we previously optimized for other combinatorial-indexing-based technologies and have applied to a variety of tissues.8 For the proof of principle, we employed the control lymphoblastic cell line GM12878. Half of the cells were preprocessed for nucleosome disruption, and the control half was undisturbed, prior to being carried through the standard 10× Genomics scATACv2 workflow. Subsequently, we nucleosome-disrupted cells from a patient-derived pancreatic cancer cell line, PDCL1 (patient-derived cell line 1), and assayed them in the same manner. Sequenced libraries were taken through barcode deconvolution and cell calling based on a unique read threshold and assessed for key metrics (STAR Methods).
We first assessed the ability of our nucleosome disruption technique to ablate the chromatin accessibility signal by assessing transcription start site enrichment (TSSe), which is a standard measurement for ATAC-seq signal evaluation. As with our previous nucleosome disruption techniques for whole-genome as well as DNA-methylation techniques,8,10 the TSSe was reduced to below 1 for both the GM12878 control line (median = 0.783092) and the PDCL (median = 0.685304) using a raw TSSe value that calculates the ratio of reads within a 500 bp window centered on TSSs over reads within 250 bp windows ±1 kbp TSSs, which produces a TSSe ≤1 for shotgun WGS data8 (Figure 1B; STAR Methods). This is in contrast to the scATAC control, which exhibited a median raw TSSe of 6.02, which is consistent with high-quality scATAC preparations using this calculation method.8 We next assessed the complexity of each library by estimating the total possible unique observed sequence reads if libraries were sequenced to saturation (STAR Methods), enabling a direct comparison that is not impacted by variation in raw sequencing depth. This revealed a high-complexity library for the scATAC control, resulting in a median estimated unique read count of 260,846. The scWGS preparations produced a higher complexity due to the ability to tagment and sequence the entire genome, with median estimates of 4,603,746 and 1,950,670 for the GM12878 control and the PDCL, respectively (Figure 1C). The reduced complexity of the aneuploid PDCL may be due to the increased DNA content, which alters the effective ratio of tagmentation complexes and DNA present, or possibly, the increased gDNA within the nucleus reduces the ability of tagmentation complexes to penetrate throughout the entire volume.
Copy-number calling of scWGS data typically involves binning the genome into windows and assessing the read counts for each cell in each of the windows followed by methods to mitigate amplification or other biases—often correlating with GC content.11,12 We assessed counts across non-overlapping 1 Mbp windows and first assessed the median absolute deviation (MAD) scores, a measurement of overall bias in WGS data,12 and compared our values to released datasets produced using the now-discontinued 10× Genomics scCNV kit, including a diploid fibroblast cell line and aneuploid breast cancer tissue sequenced to high depth and then an aneuploid gastric cancer cell line (MKN-45) sequenced at increasing levels of coverage13 (Figure 1D). At the sequencing depth we obtained for the GM12878 and PDCL experiments, we observed a slightly greater median MAD score than the 10× Genomics scCNV datasets (Figure 1E). We next assessed global coverage and noticed that the primary bias in window counts correlated with chromatin domains (Figure 1F). We suspect that this is due to the tagmentation process occurring in situ, where tagmentation efficiency is likely correlated with proximity to the nuclear periphery. We therefore included a chromatin domain as a factor in copy-number assessment using RIDDLER,14 a versatile copy-number calling tool that leverages robust Poisson regression to reliably detect outlier windows in each cell. This produced the expected copy-number neutral profiles for the GM12878 cell line and the expected copy-number aberrant profiles for the PDCL line (Figure 1G).
Multiplexing samples for scWGS using indexed tagmentation reagents
The ability to produce large-cell-count scWGS datasets shifts the primary cost burden of any given study to the sequencing depth that is required. Cell throughput from a single 10× Genomics fluidics channel can reach an excess of 10,000 cells, which results in a prohibitive sequencing cost for any individual sample for most scWGS studies. Furthermore, an assessment of the clonal diversity of most samples can typically be deconvolved with far fewer profiles.8 We therefore leveraged the scATAC multiplexing kit, commercially available from ScaleBio, which uses indexed tagmentation to preindex samples prior to loading onto the chip. This can be used for standard loading as well as “superloading,” where multiple preindexed nuclei can be processed within the same droplet.15 The classic approach is to simply load the desired number of cells of each sample into an individual fluidics channel; however, this results in greater library preparation costs, with each sample requiring its own set of reagents.
We carried out two preparations leveraging indexed tagmentation reagents using the GM12878 control cell line. The first leveraged a single tagmentation complex with 5,000 nuclei loaded into a single channel to serve as a non-multiplex control using the indexing reagents. The second pooled approximately 3,000 nuclei from each of seven indexed tagmentation reactions, which were also loaded onto a single channel (Figure 1H). After index demultiplexing (STAR Methods) the single index channel produced 3,678 cell profiles, and the multiplex channel produced 16,280, with a mean of 2,326 ± 352 cells per tagmentation index (median 2,289), with equivalent estimated complexity and TSSe (Figures 1I andFigure 2J). We then assessed the distribution of cell counts across droplets, which revealed that the large majority (86.8%) of droplets contained a single cell, with the remaining droplets containing two (10.6%) or more (2.6%) preindexed cells (Figure 1K).
Figure 2.
Double tagmentation for paired ATAC+WGS
(A) Workflow diagram.
(B) Raw TSS enrichment for each modality for each sample.
(C) Estimated library complexity.
(D) Log10 unique reads for each cell for ATAC versus WGS modalities.
Double tagmentation with indexed complexes enables scWGS + scATAC from the same cell
The ability to perform indexed tagmentation and deconvolution within a single droplet raises the possibility of leveraging multiple indexes per cell to encode separate properties. We reasoned that separate indexed tagmentation complexes could be used in succession to profile both chromatin accessibility and genomic DNA from the same cells and developed a double tagmentation (dTag) workflow (Figure 2A). Nuclei are first isolated according to scATAC workflows and carried through tagmentation across several indexed tagmentation complexes using the commercially available scATAC multiplexing kit. Importantly, ATAC tagmentation is performed prior to cell fixation, as in the standard scATAC workflow, thus avoiding any potential biases associated with tagmenting fixed DNA. Replicate wells are then pooled to achieve enough nuclei to perform fixation and nucleosome disruption followed by a second round of tagmentation with a second, distinct tagmentation index. dTag nuclei can then be pooled with other samples that leverage other sets of tagmentation indexes and loaded onto a 10× Genomics scATAC channel for droplet encapsulation and bead barcoding followed by cleanup, PCR amplification, and sequencing.
We carried out an initial preparation containing only GM12878 cells that were tagmented using a single index for the ATAC component across multiple reactions and then a single index for the whole-genome component. After the second tagmentation, approximately 2% of the post-nucleosome-disrupted nuclei recovered, all of which were loaded onto a single fluidics channel. After sequencing and index deconvolution, we assessed the TSSe for each matched component of each cell, which produced a medians of 3.75 for the ATAC component and 0.89 for the WGS component (Figure 2B), matching what we observed with single-modality assays (medians of 6.02 and 0.78 for ATAC and WGS, respectively). Similarly, the estimated total unique library molecules also reflected the balance of fewer ATAC reads compared to WGS at medians of 23,676 and 1,652,516, respectively (Figure 2C).
We next assessed the ability to multiplex samples using distinct index sets within the same scATAC fluidics channel by processing both GM12878 and K562, a human chronic myelogenous leukemia cell line, in parallel. To improve post-WGS tagmentation nuclei survival, we also assessed increased formaldehyde fixation conditions prior to nucleosome disruption after the initial ATAC tagmentation reaction (1% and 1.25% for multiplex 1 and multiplex 2, respectively, versus 0.75% for the initial experiment). For each ATAC tagmentation, we leveraged 5 sets of indexes for each sample, which were then pooled for fixation, nucleosome disruption, and WGS tagmentation. The increased formaldehyde concentration resulted in improved nuclei survival rates (approximately 5-fold for 1% and ∼2-fold for 1.25% fixation), though they were still lower than WGS tagmentation alone. For each preparation, we loaded nuclei, pooling the same fixation conditions for the GM12878 and K562 preparations within each channel. Loaded nuclei counts were limited due to the costs associated with sequencing the WGS component, targeting approximately 3,000 total nuclei loaded per channel. Passing cell profiles were comparable between conditions for each cell line, with GM12878 producing 165 and 169 cells and K562 producing 332 and 392 cells for the 1% and 1.25% fixation conditions, respectively (STAR Methods). Yields were notably lower than expectations based on the loaded count; however, a clear and distinct population of cell barcodes that contained a high count of WGS and ATAC reads was observed (Figure 2D), suggesting that the low cell count was due to an underestimation of the loaded concentration as opposed to a high failure rate.
Consistent with previous preparations, WGS reads associated with each sample produced a raw TSSe centered near 1, indicating ablation of the chromatin accessibility signal (Figure 2B). Each preparation also produced comparable estimated total unique library molecules with a median over 1 million for each GM12878 preparation (1,684,065 and 2,797,486) and somewhat less for the K562 cells (714,419 and 507,716) (Figure 2C), which is also consistent with our scWGS standalone datasets that produced reduced coverage for cells that contain greater amounts of gDNA. Notably, for both TSSe and library complexity, the fixation percentage did not appear to have any impact, making the higher nuclei yields of the 1% fixation condition favorable overall.
To evaluate the WGS portion of the libraries, we reasoned that inclusion of the ATAC reads for each cell is warranted for genomic copy-number calling, as WGS-alone assays do not exclude accessible regions, they just do not enrich for them. Using the combined data, we first assessed the MAD score coverage uniformity producing values that fell within the same range (median < 0.3; Figure 3A) as the standalone scWGS datasets and the 10× Genomics scWGS datasets using the discontinued kit (Figures 1D and 1E). We next evaluated how many cells were required to achieve genome-wide physical coverage by randomly sampling cells and aggregating reads to achieve the mean fold coverage of the genome and the percentage of the genome covered (Figure 3B). Consistent with complexity observations, coverage was reduced for aneuploid cells (PDCL and K562) versus the diploid GM12878 cell line. For GM12878 conditions other than our initial dTag singleplex experiment prior to optimization, cell counts required to reach 5-fold genome coverage, which is sufficient for cluster-based genotyping, was 20 versus 70. Between GM12878 conditions, the standalone produced the highest fold coverage and was able to achieve 30-fold coverage at 80 cells, which is considered sufficient for de novo variant calling.
Figure 3.
dTag WGS performance
(A) MAD scores for dTag scWGS data.
(B) Accumulated fold coverage (top) and percentage of genome covered (bottom) for random subsampled cell increments.
(C) RIDDLER copy-number profiles for dTag scWGS cells.
(D) Distribution of copy-number calls for GM12878 compared between scWGS and dTag scWGS methods. Copy number = 2 is highlighted.
Using the combined data for each sample, we then performed copy-number calling using RIDDLER as detailed previously (Figure 3C). Visually, increased noise was observed for the dTag GM12878 copy-number profiles when compared to the standalone assay (Figure 1G). To assess this in greater detail, we calculated the distribution of copy-number calls against each integer for the dTag experiments compared to the GM12878 standalone scWGS assay (Figure 3D). This assessment revealed the proportions of copy number 2 calls were 91.7%, 90.8%, and 94.6% for the dTag multiplex 1 and 2 and dTag singleplex experiments, respectively. This represents a decreased proportion when compared to the 95.9% for the standalone scWGS workflow and the theoretical expectation of 100%. The discrepant windows tended to fall in regions that exhibited chromatin domain signal bias, such as the q-arm of chromosome 1, suggesting that increased noise in the dTag scWGS dataset may reduce the efficacy of the bias reduction performed by RIDDLER.
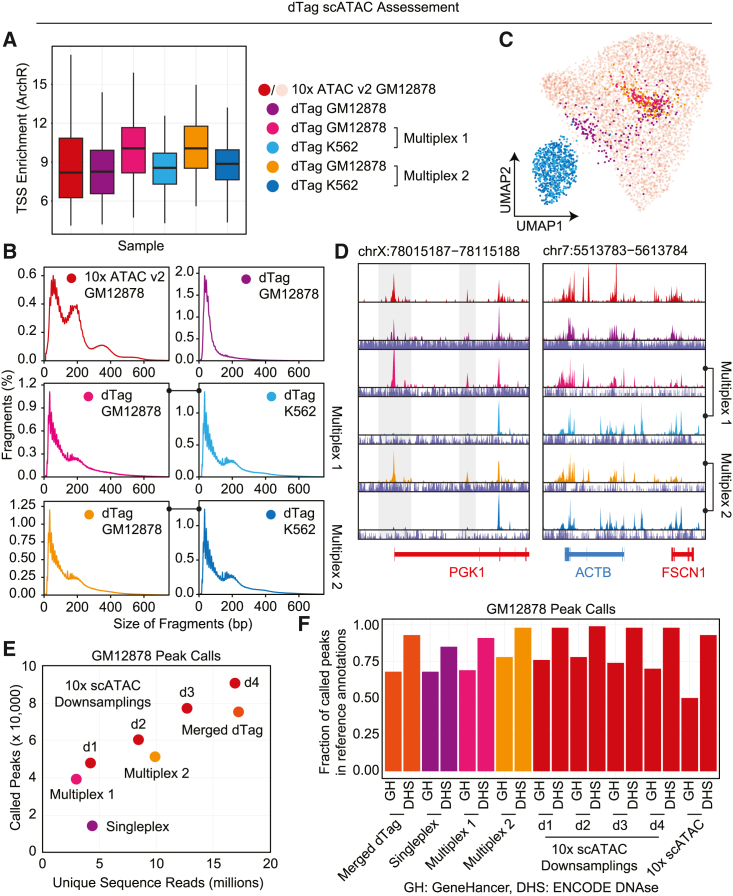
Next, we evaluated the ATAC component of the dTag preparations by producing fragment files and evaluating performance alongside the GM12878 scATAC control preparation. TSSe as assessed using the ArchR software suite revealed comparable enrichment across conditions with a slight increase in enrichment for the dTag preparations16 (Figure 4A). This observation may be due to the increased proportion of subnucleosomal fragments that occurs due to the second tagmentation for WGS that likely disrupts these fragments, leaving only the end tagmentation events from the initial ATAC reaction (Figure 4B). Despite the difference in fragment size, the two cell types produced distinct profiles that separated cleanly on a uniform manifold approximation and projection (UMAP) with GM12878 cells from both the unimodal assay and dTag conditions co-embedding with one another (Figure 4C). When examining genomic signal tracks, clear ATAC peaks were observed across each experiment with distinct differences between the GM12878 and K562 cell lines (Figure 4D).
Figure 4.
dTag ATAC performance
(A) ArchR TSS enrichment.
(B) ATAC modality fragment size distribution.
(C) UMAP of standalone scATAC (light red) and dTag scATAC conditions. Respective cell lines group together.
(D) ATAC coverage tracks for housekeeping genes shows the same patterning for GM12878 for standalone and dTag assays, with clear differences in the K562 line highlighted. Below each track is the corresponding scWGS track.
(E) Called peaks as a function of unique sequence reads for the dTag scATAC conditions as well as four downsampled datasets from the scATAC standalone assay.
(F) Percentage of peak calls in GM12878 scATAC standalone and dTag datasets compared to GeneHancer (GH) annotated regions and ENCODE DNase hypersensitivity sites (DHSs).
Finally, we assessed the ability to call peaks on pooled cell profiles from our dTag experiments and the standalone 10× scATAC dataset. Given that peak calling power correlates strongly with depth of sequencing, we downsampled the standalone 10× scATAC dataset to several increments covering a comparable range to our dTag scATAC datasets and performed peak calling on each sample as well as the merged reads from all dTag scATAC datasets. This produced slightly higher peak call counts using the standalone assay versus the dTag multiomic workflow (Figure 4E), with a similar trend of increased peak calls based on the total reads used in the peak calling. To further explore these calls, we assessed the percentage of overlap of peaks with public annotations of putative regulatory elements, including GeneHancer (GH) and ENCODE DNase hypersensitivity sites (DHSs) (Figure 4F). This produced comparable percentages for the multiplex 2 dataset versus the 10× scATAC standalone conditions with comparable coverage and a reduced percentage for other dTag conditions.
Discussion
In this work, we detail the adaptation of our previously described nucleosome disruption technologies to enable the acquisition of scWGS data using off-the-shelf kits for the widely adopted 10× Genomics Chromium instrument. These techniques seek to fill the gap in high-throughput scWGS with the goal of profiling copy-number alterations without the need to purchase new instrumentation, depend on bespoke enzymes, or resort to custom sequencing chemistry. The primary motivation was demonstrably achieved via alterations to the standard 10× Genomics workflows using readily available reagents. The scWGS data produced are of high complexity, producing sufficient read depth for copy-number calling genome-wide, and we expect that the ease of access to the technology will make it an appealing option for studies that rely on copy-number assessment for tumor clonal structure and evolution analysis.
We next utilized a commercially available scATAC multiplexing kit (ScaleBio) to enable one to load multiple samples in a single chip lane without a loss in the quality of the data. This flexibility allows many more samples to be queried on a full chip; or alternatively, allows for a rationed consumption of 10× barcoding reagents driving down library preparation costs. The improved flexibility granted by preindexing also laid the foundation for the dTag protocol to achieve ATAC plus WGS from the same cells. The dTag workflow leverages an initial round of tagmentation with indexed Tn5 followed by nucleosome disruption and then a second tagmentation using differently indexed Tn5 complexes. We also demonstrate that separate samples can be multiplexed in the same 10× Genomics instrument channel if separate sets of indexes are utilized. Notably, the scWGS data generated by the dTag method matches the quality and complexity of the scWGS protocol we present, and the scATAC component matches closely to standalone 10× Genomics scATAC data. Taken together, dTag, while using exclusively commercially available reagents, promises researchers the ability to assess heterogeneous samples for copy-number alternations in individual cells and, additionally, to use accessibility information to evaluate the putative functional impact on the epigenome within copy-number-altered loci.
Limitations of the study
The scWGS standalone assay is capable of delivering genome-wide coverage sufficient for copy-number calling to enable tumor subclone identification; however, we do observe a bias in raw coverage that correlates with broad chromosome compartments. We believe that this bias is due to the high-level structure of chromatin in the nucleus with so-called “A” and “B” compartments that are positioned in either the nuclear interior or the nuclear periphery, respectively, and that the Tn5 enzyme has variable penetration into the core of the nucleus, resulting in the observed coverage bias. Fortunately, the broad chromatin compartment structure has been shown to be consistent across cell types17 and can be readily accounted for during the copy-number calling process. The other limitation to the scWGS approach is that it does not provide sufficient coverage for de novo single-nucleotide variant calling due to the lack of a genome amplification step—something that is shared across all direct tagmentation scWGS techniques,18 restricting its application to copy-number assessment or the genotyping of variants identified in an aggregate dataset or variant calling within copy-number-defined clusters.
The primary challenge with the dTag technique is establishing ideal fixation and nucleosome disruption conditions to maximize nuclei recovery after the second tagmentation reaction. We noticed improved nuclei yields with a higher formaldehyde percentage without a reduction in data quality; however, the majority (∼90%) of nuclei were lost at this step, suggesting that additional optimization will be required for samples that do not have at least ∼1 million nuclei to start with—something that is not a problem with most tumor tissue specimens but can be tricky when handling biopsy-derived tissue that is often portioned for multiple assays. Furthermore, the low cell yield makes it challenging to achieve sufficient genome coverage for de novo variant calling on identified clusters, restricting utility to samples where paired bulk exome or whole-genome data can be used to produce a reference variant set against which dTag cell clusters can be genotyped.
STAR★Methods
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Chemicals, peptides, and recombinant proteins | ||
| HEPES, pH 7.5 | Sigma-Aldrich | H4034 |
| MgCl2 | Sigma-Aldrich | M8226 |
| NaCl | Fisher Scientific | M-11624 |
| IGEPAL | Sigma-Aldrich | I8896 |
| Tween 20 | Sigma-Aldrich | P7949 |
| Formaldehyde | Fisher Scientific | PI28906 |
| Glycine | Sigma-Aldrich | G8898-500G |
| UltraPure Sodium Dodecyl Sulfate | Invitrogen | 15525–017 |
| D-(+)-Glucosamine hydrochloride | Sigma-Aldrich | G1414-100G |
| Critical commercial assays | ||
| Chromium Next GEM Single Cell ATAC Kit v2 | 10x Genomics | PN-1000390 |
| Next GEM Chip H Single Cell Kit | 10x Genomics | PN-1000162 |
| Single Index Kit N, Set A, 96 | 10x Genomics | PN-1000212 |
| Single Cell ATAC Gel Beads v2 | 10x Genomics | PN-2000210 |
| scATAC Pre-Indexing Kit | ScaleBio | N/A |
| Qubit 1x dsDNA HS Assay Kit | Invitrogen | Q33231 |
| High Sensitivity D1000 ScreenTape | Agilent | 5067–5584 |
| High Sensitivity D1000 Sample Buffer | Agilent | 5067–5603 |
| NextSeq2000 Kits | Illumina | 20046811, 20046812 |
| Deposited data | ||
| Raw and analyzed data | This paper |
https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc = GSE243430 |
| Human Reference Genome NCBI Build 37, GRCCh37 | Genome Reference Consortium | http://www.ncbi.nlm.nih.gov/projects/genome/assembly/grc/human/ |
| 10x Genomics scCNV kit released datasets | 10x Genomics13 | 10xgenomics.com/resources/ datasets |
| Experimental models: Cell lines | ||
| K562 | Tanner Lab | N/A |
| GM12878 | Coriell Institute | Cat. No. GM12878 |
| Patient-Derived Pancreatic Cancer Cell Line | Rosie Sears Lab (OHSU) | ST-00024058 |
| Software and algorithms | ||
| Unidex | Adey Lab | https://github.com/adeylab/unidex |
| bwa-mem (v0.7.15-r1140) | Li and Durbin19 | https://github.com/lh3/bwa |
| Scitools | Adey Lab | https://github.com/adeylab/scitools |
| RIDDLER | Gurkan Lab | https://github.com/yardimcilab/RIDDLER |
| ArchR | Granja et al.16 | https://www.archrproject.com/ |
| Other | ||
| Cytiva HyClone RPMI 1640 Media | Fisher Scientific | SH3002701 |
| Gibco DMEM Media, high glucose | ThermoFisher Scientific | 11960044 |
| Glutamax 100x | Life Technologies | 35050061 |
| HyClone Bovine Calf Serum, heat inactivated | VWR | SH30073.04HI |
| Penicillin/Streptomycin (10,000 U/ml) | Life Technologies | 15-140-122 |
| TrypLE Express Enzyme | ThermoFisher Scientific | 12604013 |
Resource availability
Lead contact
Requests for protocols and reagents can be directed to Andrew Adey (adey@ohsu.edu).
Materials availability
Materials used in this study are commercially available
Experimental model and subject details
Preparation of cell lines
We cultured GM12878 and K562 in Roswell Park Memorial Institute Medium, supplemented with 1% additional glutamine, 1% penicillin/streptavidin, and 10% calf bovine serum. Standard humidified incubator conditions were kept at 37°C with 5% CO2. We cultured the adherent Neuro2A cells in Dulbecco’s Modified Eagle’s Medium and dissociated them with TrypLE Express Enzyme, as required for cell passage. Otherwise, the culture conditions remained identical.
Preparation of PDCL
The PDCL (ST-00024058) was established from a dissociated human pancreatic ductal adenocarcinoma tumor metastasis and cultured for continuous propagation in culture medium containing ROCK inhibitor (Y-276320).20 The sample was deidentified and anonymized and is part of a study overseen and approved by the OHSU Institutional Review Board. Briefly, 100,000 viable, disaggregated tumor cells were plated to a 13mm diameter, collagen-coated well (Gibco, A11428–02) and passaged while subconfluent until reaching 85% confluence on a 10cm diameter dish, which was designated as passage 1. DNA was extracted from a third passage to validate by whole exome sequencing that the mutation profile in the PDCL matched the patient tumor DNA profile, including KrasG12D and TP53R175H mutations (detected at 45% and 100% frequency, respectively, in the PDCL). The PDCL also exhibited morphologies consistent with epithelial tumor cells and abundant KRT expression was detected by immunocytofluorescence using the monoclonal antibodies AE1/AE3, C-11, and Cam5.2. The tumor originated from a deidentified 60-year-old male, under the oversight and with the approval of the OHSU Institutional Review Board.
Method details
Nuclei Isolation
To isolate the nuclei, we collected a 2 mL suspension of cells in media. We spun the suspension down (5 min, 500xg, 4C) to remove the media supernatant and resuspended it in 1 mL of NIB-Hepes buffer (10 mM HEPES, pH 7.5, 3 mM MgCl2, 10 mM NaCl, 0.1% IGEPAL (v/v), 0.1% Tween 20 (v/v)). We then incubated the cell suspension for 5 min on ice; and subsequently, we spun down the sample (5 min, 500xg, 4C). We then resuspended the pellet in 1 mL NIB-Hepes, spun again, and resuspended in 1 mL once more before quantification.
Nucleosome disruption
We diluted the samples to 1 million nuclei per mL in NIB-H buffer and then added 46.9 μL 16% formaldehyde (final concentration is ∼0.75% formaldehyde). After pipette mixing, we fixed the sample at room temperature over 10 min, on an orbital shaker set to 50rpm. We next added 46.9 μL 2.5M glycine to quench the reaction, incubated for 5 min on ice, and then spun the suspension down (5 min, 500xg, 4°C). We next resuspended the sample in 970 μL NIB-H and then added 30 μL 10% SDS. We incubated the nuclei for 20 min at 37°C in this solution. We carefully spun down the samples (5 min, 500xg, 10°C), as the SDS can precipitate and taint the pellet if left cold for too long. Nuclei were then resuspended in 1 mL NIB-H buffer and quantified.
10x tagmentation and barcoding for scWGS
For the single cell whole genome sequencing protocol leveraging the 10x transposition chemistry, we took the preprocessed, nucleosome disrupted nuclei and diluted them in 10x Genomics 20x Nuclei Buffer to the desirable targeted cell recovery. We then proceeded with the 10x Genomics Single Cell ATAC v2 Kit according to their published protocol.
ScaleBio tagmentation and 10x barcoding for scWGS
For the scWGS protocol leveraging the ScaleBio transposition chemistry, we resuspended the preprocessed nuclei in PBS Buffer with BSA, such that there were 10,000 or 20,000 nuclei per μL. We also added 30 mM D-glucosamine to the PBS/BSA buffer, which will amount to final concentration of 10 mM during the tagmentation reaction, to improve the recovery of nuclei after the tagmentation.21 In a 96-well plate, we added 5 μL of diluted nuclei to 5 μL of ETB3, prior to adding 5 μL of the indexed Scale TSM. We mixed the plate by shaking at 1400 rpm for a minute, briefly spun it down at 500xg, and then we incubated the plate at 37°C for 1 h on a thermocycler. The lid should be set to 47°C. In the interim, we thawed the ScaleBio wash and loading buffers on ice.
After the tagmentation was completed, we removed the plate from the thermocycler, briefly spun the plate down at 500xg, and incubated it on ice for 5 min. We pooled the samples in an Eppendorf tube and filled the remaining volume with the ScaleBio wash buffer. For subsequent steps, using a spinning bucket centrifuge aids in retention of the nuclei pellet. We spun down the samples (3 min, 500xg, 4°C), discarded the supernatant, added another 1.5 mL of wash buffer, and spun again. We again removed all the supernatant and resuspended the pellet in 25 μL of ScaleBio’s loading buffer. We quantified the nuclei and added the desired amount to be loaded in 15 μL loading buffer on the 10x Chromium. At this point multiple samples with unique Tn5 indexes can be pooled in the 15 μL of loading buffer, to be run on a single lane on the Chromium Chip for multiplexing.
With the quantified, transposed nuclei in hand, we continued to Step 2 (Gem Generation and Barcoding) of the 10x Genomics Single Cell ATAC v2 Kit protocol. We mixed our 15 μL of nuclei with 60 μL of the described 10x Master Mix and proceeded from here on out exactly as described by the 10x protocol. The one subsequent deviation is during Step 4.1.c (Sample Index PCR), where instead of using the provided Sample Index N, Set A Reagent – we used a ScaleBio S700 index primer compatible with the ScaleBio tagmentation.
ScaleBio tagmentation and 10x barcoding for DoubleTag
For the double tagmentation protocol leveraging the ScaleBio transposition chemistry, we proceeded through the ScaleBio Tagmentation approach described above on isolated, but not nucleosome disrupted, nuclei. After tagmentation, we washed twice with the ScaleBio wash buffer. We next resuspended the nuclei in NIB-H buffer and perform the nucleosome disruption protocol. Following nucleosome disruption, we repeated the ScaleBio tagmentation – being sure to use differently indexed Tn5 so that the ATAC and WGS fragments can be distinguished. After the second tagmentation, and subsequent the washing and resuspension in the loading buffer, we quantified the nuclei and added the desired amount to be loaded in 15 μL loading buffer to the 10x Chromium. As before, we continued from Step 2 (Gem Generation and Barcoding) of the 10x Genomics Single Cell ATAC v2 Kit protocol, after mixing the 15 μL with 60 μL of the 10x Master Mix and loading the chip. Again, we would go on to use the ScaleBio S700 index primers in lieu of the 10x Sample Index N during Step 4.1.c (Sample Index PCR).
Library quantification & sequencing
We cleaned library preparations as described in the 10x Genomics Single Cell ATAC v2 Kit. We quantified generated libraries via the Qubit dsDNA High Sensitivity assay (Thermo Fisher Q32851). We then confirmed the molarity of the DNA via the Agilent Tapestation 4150 D500 tape (Agilent 5067–5592), with preparations diluted to 2 ng/μL based on the Qubit data if necessary. We sequenced library preparations using standard chemistry on the Illumina NextSeq2000 for 650p.m. using a P2-100 or P2-200 flow cell (Illumina Inc. 20046811, 20046812). ScaleBio tagmented libraries were sequenced as paired-end with 85 cycles for read 1, 125 cycles for read 2, 8 cycles for index 1, and 16 cycles for index 2. Standard 10x tagmentation libraries were sequenced as paired-end with 50 cycles for read 1 and read 2, but 8 cycles for index 1, and 16 index 2. Several libraries, for which we required great sequencing depth, were also sequenced on a NovaSeq S2 flowcell following manufacturer’s instructions (Illumina Inc. 20028315).
Quantification and statistical analysis
Computational analysis
Primary sequence data processing
Sequence reads were demultiplexed using unidex (https://github.com/adeylab/unidex) which matches barcode regions to a whitelist, allowing for a hamming distance of 1 from the 10x bead barcode and index read 2, and then a hamming distance of 2 for index read 1 (sample index) or the first 8bp of read 2 which serves as the tagmentation index when indexed tagmentation is performed. For indexed tagmentation experiments the 8 bp index was trimmed, along with the next 20 bp of mosaic end sequence. Read names were then replaced with the error-corrected barcode combination. Demultiplexed and barcode-matched reads were then aligned to the human reference genome hg38 using bwa-mem (v0.7.15-r1140), then PCR-duplicate removed in a barcode-aware manner using scitools rmdup (https://github.com/adeylab/scitools) and then filtered to only contain reads in cell barcodes reaching a minimum unique read count as determined by the inflection point on a knee plot.
MAD scores
Median Absolute Deviation (MAD) scores were calculated as previously described8,12:
Where is the raw read count for the cell of the bin. is the cell-specific scaling factor (total reads), and is a bin-specific scaling factor (total reads in bin across all cells).
Coverage accumulation
Coverage accumulation was carried out by randomly sampling cells without replacement and aggregating the coverage assuming paired 150 bp sequence reads, and only considering the physical coverage of the read if the insert size was less than 300 bp (i.e., when read pairs overlap, it is collapsed to a coverage of 1x for the overlapping bases). 150 bp was chosen as it represents what a typical sequencing instrument can produce if genomic coverage is the goal (as opposed to shorter reads if only copy number calling is desired).
CNV detection using RIDDLER
CNV detection of cells was performed using the RIDDLER framework.14 Reads were binned at 1 Mbp resolution, creating a matrix of cells by windows used as input. Zero values in the matrix were assessed as potential dropout events, treating each cell as a sample of reads over all windows. The probability of these windows having zero reads in the absence of CNVs was estimated, based on the total reads in the cell and average window distribution across all cells. Windows with a high estimated probability (0.95 or higher) of being dropout events were removed from the input data. This typically only occurred in low coverage windows of lower coverage cells.
Robust Poisson regression, detailed in Cantoni & Ronchetti (2001)22 and implemented in the R package robustbase, was used to model the expected distribution of reads per window across cells. The use of robust regression as opposed to least-squares regression reduces the influence of outliers, making them easier to disentangle from the data. As covariates for the regression model, we used mappability, AB compartment, and median coverage. Mappability was scored using 50mer alignability,22 AB compartment scores were taken as the ENCODE GM12878 Hi-C 5 kbp genome compartments,23 and median coverage was computed using the reference cell line GM12878. Each covariate was averaged within each window to match the input dimensions, and the same covariates were used for each cell and each experiment. Regression models were computed for each chromosome individually, and used to predict an expected read count for each window.
CNVs were detected by comparing the observed window reads in each cell to the expected reads predicted by the model. Using the model prediction as the expected value, p values were computed for each window using a negative binomial distribution. Dispersion values for the negative binomial distribution were estimated for each chromosome using the GM12878 cell line, to establish a baseline of expected variation. Window p-values were combined with neighboring windows up to 3 Mbps away to create aggregate p-values using an empirical Bates distribution. These aggregate p-values were then FDR corrected (Benjamini Hochberg 1995),24 and thresholded for significance at a value of 0.025 for each tail. Windows that passed this threshold on the upper tail are labeled as CNV gains, and those that pass on the lower tail are labeled as CNV losses.
To assign copy numbers (0, 1, 2, 3, 4, or 5+) to each window, we computed the log likelihood of the observed reads using a corresponding multiplier of the model fitted reads (0, 0.5, 1, 1.5, 2, and 2.5 respectively) as the expected value. Since having an expected value of 0 produces errors in the negative binomial distribution, we instead used an expected value multiplier of 0.1 to assess the likelihood of the copy number 0 label. The copy number with the highest log likelihood was then assigned to the window.
Analysis of scATAC data
Aligned and duplicate-removed bam files were converted into fragment files using sinto 0.9.0 and then loaded into ArchR15 to generate arrow files for each ATAC fragment file which were then compiled into an ArchR project. Iterative LSI, harmony integration, and UMAP projections were performed using default parameters. Track plots were generated by selecting known housekeeping genes that are expected to be active in all cell lines.
Acknowledgments
This work was supported by an NCI IMAT R33 (R33CA269015) to A.C.A. and an NIH Ruth L Kirschstein T32 Fellowship (5T32GM142619-02) to K.Q.
Author contributions
A.C.A. conceptualized the method. A.C.A., B.L.O., and K.Q. designed experiments; K.Q. performed all experiments with assistance from B.L.O.; R.V.N., G.G.Y., and T.W.M. devised RIDDLER; T.W.M. performed copy-number calling analysis; A.C.A., B.L.O., R.V.N., and K.Q. performed analyses; and J.L.M., D.K., C.L., G.B.M., and R.C.S. managed the banked patient-derived samples.
Declaration of interests
A.C.A. is an author on a patent that covers one or more aspects of the nucleosome disruption technology utilized here. This potential conflict is managed by the OHSU office of research integrity.
Inclusion and diversity
One or more of the authors of this paper self-identifies as an underrepresented ethnic minority in their field of research or within their geographical location. One or more of the authors of this paper self-identifies as a gender minority in their field of research. One or more of the authors of this paper self-identifies as a member of the LGBTQIA+ community.
Published: November 1, 2023
Data and code availability
-
•
Data are accessible through the NCBI Gene Expression Omnibus (GEO) under accession GEO: GSE243430.
-
•
This study does not report original code.
-
•
Any additional information needed to reanalyze the data reported in this paper is available from the lead contact by request.
References
- 1.Gerstung M., Jolly C., Leshchiner I., Dentro S.C., Gonzalez S., Rosebrock D., Mitchell T.J., Rubanova Y., Anur P., Yu K., et al. The evolutionary history of 2,658 cancers. Nature. 2020;578:7793. doi: 10.1038/s41586-019-1907-7. 122–128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Baslan T., Kendall J., Rodgers L., Cox H., Riggs M., Stepansky A., Troge J., Ravi K., Esposito D., Lakshmi B., et al. Genome wide copy number analysis of single cells. Nat. Protoc. 2012;7:1024–1041. doi: 10.1038/NPROT.2012.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Kim C., Gao R., Sei E., Brandt R., Hartman J., Hatschek T., Crosetto N., Foukakis T., Navin N.E. Chemoresistance Evolution in Triple-Negative Breast Cancer Delineated by Single-Cell Sequencing. Cell. 2018;173:879–893.e13. doi: 10.1016/J.CELL.2018.03.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Navin N., Kendall J., Troge J., Andrews P., Rodgers L., McIndoo J., Cook K., Stepansky A., Levy D., Esposito D., et al. Tumour evolution inferred by single-cell sequencing. Nature. 2011;472:90–94. doi: 10.1038/NATURE09807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Wang Y., Waters J., Leung M.L., Unruh A., Roh W., Shi X., Chen K., Scheet P., Vattathil S., Liang H., et al. Clonal Evolution in Breast Cancer Revealed by Single Nucleus Genome Sequencing. Nature. 2014;512:155–160. doi: 10.1038/NATURE13600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Danilenko M., Zaka M., Keeling C., Crosier S., Lyman S., Finetti M., Williamson D., Hussain R., Coxhead J., Zhou P., et al. Single-cell DNA sequencing identifies risk-associated clonal complexity and evolutionary trajectories in childhood medulloblastoma development. Acta Neuropathol. 2022;144:565–578. doi: 10.1007/S00401-022-02464-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Cai X., Evrony G.D., Lehmann H.S., Elhosary P.C., Mehta B.K., Poduri A., Walsh C.A. Single-Cell, Genome-wide Sequencing Identifies Clonal Somatic Copy-Number Variation in the Human Brain. Cell Rep. 2014;8:1280–1289. doi: 10.1016/j.celrep.2014.07.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Mulqueen R.M., Pokholok D., O’Connell B.L., Thornton C.A., Zhang F., O’Roak B.J., Link J., Yardımcı G.G., Sears R.C., Steemers F.J., Adey A.C. High-content single-cell combinatorial indexing. Nat. Biotechnol. 2021;39:1574–1580. doi: 10.1038/S41587-021-00962-Z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Vitak S.A., Torkenczy K.A., Rosenkrantz J.L., Fields A.J., Christiansen L., Wong M.H., Carbone L., Steemers F.J., Adey A. Sequencing thousands of single-cell genomes with combinatorial indexing. Nat. Methods. 2017;14:302–308. doi: 10.1038/nmeth.4154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Nichols R.V., O’Connell B.L., Mulqueen R.M., Thomas J., Woodfin A.R., Acharya S., Mandel G., Pokholok D., Steemers F.J., Adey A.C. High-throughput robust single-cell DNA methylation profiling with sciMETv2. Nat. Commun. 2022;13:7627. doi: 10.1038/s41467-022-35374-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Jiang Y., Wang R., Urrutia E., Anastopoulos I.N., Nathanson K.L., Zhang N.R. CODEX2: full-spectrum copy number variation detection by high-throughput DNA sequencing. Genome Biol. 2018;19:202. doi: 10.1186/S13059-018-1578-Y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Wang R., Lin D.Y., Jiang Y. SCOPE: A Normalization and Copy-Number Estimation Method for Single-Cell DNA Sequencing. Cell Syst. 2020;10:445–452.e6. doi: 10.1016/J.CELS.2020.03.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.10x Genomics 10xgenomics.com/resources/datasets
- 14.Moore T.W., Galip Gurkan Yardımcı Robust CNV detection using single-cell ATAC-seq. bioRxiv. 2023 doi: 10.1101/2023.10.04.560975. Preprint at. 2023.10.04.560975. [DOI] [Google Scholar]
- 15.Lareau C.A., Duarte F.M., Chew J.G., Kartha V.K., Burkett Z.D., Kohlway A.S., Pokholok D., Aryee M.J., Steemers F.J., Lebofsky R., Buenrostro J.D. Droplet-based combinatorial indexing for massive-scale single-cell chromatin accessibility. Nat. Biotechnol. 2019;37:916–924. doi: 10.1038/s41587-019-0147-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Granja J.M., Corces M.R., Pierce S.E., Bagdatli S.T., Choudhry H., Chang H.Y., Greenleaf W.J. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 2021;53:403–411. doi: 10.1038/s41588-021-00790-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Yardımcı G.G., Ozadam H., Sauria M.E.G., Ursu O., Yan K.K., Yang T., Chakraborty A., Kaul A., Lajoie B.R., Song F., et al. Measuring the reproducibility and quality of Hi-C data. Genome Biol. 2019;20:57. doi: 10.1186/S13059-019-1658-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Minussi D.C., Nicholson M.D., Ye H., Davis A., Wang K., Baker T., Tarabichi M., Sei E., Du H., Rabbani M., et al. Breast Tumors Maintain a Reservoir of Subclonal Diversity During Expansion. Nature. 2021;592:302–308. doi: 10.1038/S41586-021-03357-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Li H., Durbin R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics. 2009;25:1754–1760. doi: 10.1093/BIOINFORMATICS/BTP324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Liu X., Krawczyk E., Suprynowicz F.A., Palechor-Ceron N., Yuan H., Dakic A., Simic V., Zheng Y.L., Sripadhan P., Chen C., et al. Conditional reprogramming and long-term expansion of normal and tumor cells from human biospecimens. Nat. Protoc. 2017;12:439–451. doi: 10.1038/nprot.2016.174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.O’Connell B.L., Nichols R.V., Pokholok D., Thomas J., Acharya S.N., Nishida A., Thornton C.A., Co M., Fields A.J., Steemers F.J., Adey A.C. Atlas-scale single-cell chromatin accessibility using nanowell-based combinatorial indexing. Genome Res. 2023;33:208–217. doi: 10.1101/GR.276655.122/-/DC1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Derrien T., Estellé J., Marco Sola S., Knowles D.G., Raineri E., Guigó R., Ribeca P. Fast Computation and Applications of Genome Mappability. PLoS One. 2012;7 doi: 10.1371/JOURNAL.PONE.0030377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Rao S.S.P., Huntley M.H., Durand N.C., Stamenova E.K., Bochkov I.D., Robinson J.T., Sanborn A.L., Machol I., Omer A.D., Lander E.S., Aiden E.L. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell. 2014;159:1665–1680. doi: 10.1016/j.cell.2014.11.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Benjamini Y., Hochberg Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. Roy. Stat. Soc. B. 1995;57:289–300. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
-
•
Data are accessible through the NCBI Gene Expression Omnibus (GEO) under accession GEO: GSE243430.
-
•
This study does not report original code.
-
•
Any additional information needed to reanalyze the data reported in this paper is available from the lead contact by request.