Abstract
Sterol regulatory element-binding proteins (SREBPs) are a subfamily of basic helix-loop-helix-leucine zipper proteins that regulate lipid metabolism. We show novel evidence of the in vivo occurrence and subnuclear spatial localization of both exogenously expressed SREBP-1a and -2 homodimers and heterodimers obtained by two-photon imaging and spectroscopy fluorescence resonance energy transfer. SREBP-1a homodimers localize diffusely in the nucleus, whereas SREBP-2 homodimers and the SREBP-1a/SREBP-2 heterodimer localize predominantly to nuclear speckles or foci, with some cells showing a diffuse pattern. We also used tethered SREBP dimers to demonstrate that both homo- and heterodimeric SREBPs activate transcription in vivo. Ultrastructural analysis revealed that the punctate foci containing SREBP-2 are electron-dense nuclear bodies, similar or identical to structures containing the promyelocyte (PML) protein. Immunofluorescence studies suggest that a dynamic interplay exists between PML, as well as another component of the PML-containing nuclear body, SUMO-1, and SREBP-2 within these nuclear structures. These findings provide new insight into the overall process of transcriptional activation mediated by the SREBP family.
Cellular lipid homeostasis is, in part, maintained by the sterol regulatory element-binding protein (SREBP) family of transcription factors (9). There are three major SREBP isoforms. SREBP-1a and -1c are splice variants derived from one gene, and SREBP-2 is encoded by a separate and unlinked gene (20). SREBPs belong to the basic helix-loop-helix-leucine zipper (bHLH-LZ) family of transcription factors and are highly conserved in this dimerization domain. SREBP-1c is a truncated form of SREBP-1a, and they only differ at their extreme amino termini, where SREBP-1c lacks the first 29 amino acids from SREBP-1a and contains five unique amino acids. Studies with transgenic and knockout mice have begun to uncover the in vivo roles of the individual SREBPs (11, 17, 28, 29). Along with several reports analyzing SREBP gene activation in cultured cells (reviewed in reference 21), these studies suggest that SREBP-1 and -2 preferentially activate genes of fatty acid and cholesterol metabolism, respectively, and that SREBP-1a is a much more potent activator of gene expression than SREBP-1c is.
SREBPs are unique in that they remain inactive through sequestration in the membrane of the endoplasmic reticulum by two membrane-spanning domains. Depletion of sterols triggers their translocation to the Golgi, where a two-step proteolytic process releases the amino-terminal half from the membrane anchor, and this mature transcription factor is then translocated to the nucleus, where it activates genes involved in regulating lipid balance (9, 10).
SREBPs, like other bHLH-LZ transcription factors, dimerize through their HLH-LZ motif and bind DNA through their basic domain. The results of overexpression studies with dominant negative versions of the SREBPs have shown that SREBP-1a and -2 can dimerize with each other, although not with the other bHLH-LZ family members tested so far (24). However, the localization, putative formation, and activity of SREBP homo- versus heterodimers has not been directly evaluated or compared in any systematic way. Because both the SREBP-1 and -2 proteins are expressed in most tissues and cell types, the function of the homo- versus heterodimer is an important question. In the present studies, we investigated the occurrence and spatial distribution of exogenously expressed SREBP-1a and -2 homo- and heterodimers in vivo and demonstrated that both homo- and heterodimers are transcriptionally active in the cell. These findings indicate significant differences in the subnuclear distribution of SREBP-1a and -2 and have major implications for gene activation mediated by the SREBP family.
MATERIALS AND METHODS
Plasmid construction. (i) SREBP-green fluorescent protein (GFP) fusions.
For SREBP-1aGFP and SREBP-2GFP, full-length mature SREBP-1a (human, amino acids 1 to 490) and SREBP-2 (human, amino acids 1 to 482) were digested with BamHI and HindIII from pPacSREBP-1a and pPacSREBP-2, respectively, as described previously (2) and cloned into BglII- and HindIII-digested pEGFP-N1 (Clontech, Palo Alto, Calif.). Serial deletions of SREBP-2 were made by PCR with specific primers, digested with BglII and HindIII, and cloned into BglII- and HindIII-digested pEGFP-N1.
(ii) PML-GFP fusion.
The GFP coding sequence was generated by PCR with pEGFP-N1 as the template and oligonucleotides containing an MluI site at the 5′ end and an XbaI site at the 3′ end. The fragment was inserted into the previously described CMX-PML vector (13), which was digested with MluI and NheI at the C terminus.
(iii) SREBP fusion plasmids for fluorescence resonance energy transfer (FRET) analysis.
For SREBP-1aCFP and SREBP-2CFP, full-length mature SREBP-1a and -2 were digested with BamHI and HindIII from pPacSREBP-1a and -2, respectively, as described above and cloned into pECFP-N1 (Clontech) digested with BglII and HindIII. For SREBP-1aYFP and SREBP-2YFP, full-length mature SREBP-1a and -2 were digested with EcoRI and HindIII from CMV-SREBP-1a and -2, respectively, as previously described (16) and cloned into EcoRI- and HindIII-digested yellow fluorescent protein (YFP; Topaz; Packard BioSciences, Meriden, Conn.).
(iv) Tethered-dimer fusions.
To construct the SREBP-1a tethered-dimer fusion (designated SREBP-1a/SREBP-1a), first a double-stranded oligonucleotide containing an 18-amino-acid tether was digested with EcoRI and ligated upstream of EcoRI-digested pCMV-5 containing mature SREBP-1a (described above). Next, an Xba fragment containing the tether and SREBP-1a fusion was subcloned downstream of Xba-digested plasmid pCDNA3.1+ (Invitrogen, San Diego, Calif.) containing two copies of the Flag epitope sequence upstream of the SREBP coding sequence. The other tethered SREBP dimer fusions were made similarly. SREBP-2/SREBP-2 represents the SREBP-2 homodimer. SREBP-1a/SREBP-2 and SREBP-2/SREBP-1a represent the SREBP-1a and -2 heterodimer, with SREBP-1a at the amino terminus for the SREBP-1a/SREBP-2 fusion and SREBP-2 at the amino terminus for the SREBP-2/SREBP-1a fusion.
(v) Reporter gene plasmid.
The construction of the reporter plasmid for the wild-type low-density lipoprotein receptor (LDL-R)-encoding gene has been described previously (26).
Cell culture, microinjection, and transient-transfection assays. (i) Microinjection.
HEp-2 cells (American Type Culture Collection, Manassas, Va.) were cultured as previously described (12) in drilled 35-mm-diameter dishes with an alpha-numerically gridded coverslip (Bellco Glass, Vineland, N.J.) affixed to the bottom. Approximately 10−15 liters of DNA plasmid per cell (200 ng/μl) was introduced into the nucleus of the cell with a semiautomated microinjection system (Eppendorf Scientific, Westbury, N.Y.) and a Zeiss Axiovert microscope (Zeiss Inc., Oberkochen, Germany). For the FRET studies, DNA plasmid concentrations were kept equal for cyan fluorescent protein (CFP) and YFP to ensure equivalent expression within a cell. Cells were incubated overnight for all studies to permit expression. Each experimental condition was performed in triplicate.
(ii) Activation studies.
CV-1 cells (from Ken Cho, University of California, Irvine) were grown in Dulbecco's modified Eagle's medium (Irvine Scientific, Irvine, Calif.) with 10% fetal bovine serum and additives purchased from Invitrogen (normal medium). Cells were plated in 6-mm wells at 60,000 per well on day 0. Cells were transfected by calcium phosphate coprecipitation on day 1. On day 2 (12 to 16 h posttransfection), cells were washed three times with 1× phosphate-buffered saline (PBS) and refed with normal medium. Cells were harvested on day 3 (24 h posttransfection) with cell lysis buffer (25 mM Gly-Gly, 15 mM MgSO4, 4 mM EGTA, 0.25% Triton X-100) and analyzed for luciferase and β-galactosidase activities as described previously (2). Each experimental point was performed in duplicate and represents several independent transfections.
(iii) Dominant negative studies.
CV-1 cells were grown as described above. Cells were plated in 60-cm-diameter dishes at 125,000 per dish on day 0. On day 1, cells were transfected by calcium phosphate coprecipitation. On day 2 (16 to 18 h posttransfection), the cells were washed three times with 1× PBS and refed with normal medium. Cells were harvested by a standard freeze-thaw method as described previously (1) on day 3 and analyzed for luciferase and β-galactosidase activities as described above. The data represent the average of two separate transfection experiments, each performed in duplicate.
(iv) Tethered-dimer activation studies.
293T cells (from Craig Walsh, University of California, Irvine) were grown as described for CV-1 cells above. Cells were plated in six-well dishes at 3.5 ×105 per well on day 0. Cells were transfected by calcium phosphate coprecipitation on day 1. On day 2 (12 to 16 h posttransfection), cells were washed two times with 1× PBS and refed with normal medium containing cholesterol and 25-hydroxycholesterol (Toruloides, Inc., Newport, R.I.) at final concentrations of 12 and 1 μg/ml, respectively, to suppress endogenous SREBP activity. Cells were harvested on day 3 (30 h after refeeding) with cell lysis buffer and analyzed for luciferase and β-galactosidase activities as described above. Each experimental data point was performed in duplicate and represents several independent transfections.
(v) Western blot analysis of protein expression.
293T cells were plated on day 0 in 60-cm-diameter dishes at 6 × 105 per dish and transfected on day 1 by calcium phosphate coprecipitation. On day 2 (12 to 16 h posttransfection), the cells were washed two times with 1× PBS and refed with normal medium. Cells were harvested on day 4 as described previously (31). Extracts were assayed for β-galactosidase activity as described above. Equivalent amounts of protein, normalized to β-galactosidase values, were resolved by sodium dodecyl sulfate-polyacrylamide gel electrophoresis and visualized by immunoblotting. SREBP-1a, GFP, and non-GFP-tagged fusions were detected with an antibody, IgG2A4, to SREBP-1a (sc-13551; Santa Cruz, Santa Cruz, Calif.). SREBP-2, GFP, and non-GFP-tagged fusions were detected with an antibody, IgGID2, to SREBP-2 (American Type Culture Collection). SREBP-tethered-dimer fusions were detected with an antibody to the Flag epitope (Sigma, St. Louis, Mo.).
Antibodies and immunostaining. (i) PML and SUMO-1 immunodetection.
HEp-2 cells microinjected with SREBPGFP fusions were fixed in 3.7% formaldehyde-PBS and immunostained with an affinity-purified rabbit antibody (6) (a kind gift from J. Dyck) as previously described (12). Similarly, cells were immunostained with a mouse monoclonal antibody to SUMO-1 (Zymed Laboratories, San Francisco, Calif.).
(ii) GFP immunodetection.
HEp-2 cells microinjected with SREBP-2GFP were fixed and immunostained prior to electron microscopy analysis (19) with affinity-purified mouse anti-GFP (Clontech) at a 1:100 dilution. Primary antibodies were subsequently detected with a 1.4-nm Nanogold-conjugated secondary goat antibody and a silver enhancement process (Nanoprobes, Yaphank, N.Y.).
Imaging and microscopy. (i) Low-light fluorescence microscopy imaging.
Samples were examined with a Zeiss Axiovert 10 microscope equipped with a 100-W Hg lamp excitation source and a 100× Zeiss Plan-NEOFLUAR objective (numerical aperture = 1.2). A slow-scan cooled charge-coupled device (CCD) camera (576 by 384 pixels, model 57-180; Princeton Instruments, Trenton, N.J.) with a 16-bit-per-pixel dynamic range of data acquisition interfaced with a personal computer was attached to the microscope. GFP fluorescence was excited with a 480- to 30-nm band-pass filter and detected with a 535- to 40-nm band-pass filter. Texas Red fluorescence for PML detection was excited with a 560- to 40-nm band-pass filter and detected with a 630- to 60-nm band-pass filter. Images were acquired with IPLab, version 3.5.5 (Scanalytics, Fairfax, Va.), and converted to TIFF files. TIFF files were subsequently processed with ADOBE Photoshop, version7.0.
(ii) Two-photon imaging and spectroscopy FRET.
The two-photon imaging and spectroscopy system used for this study has been described previously (13, 32). Briefly, it consists of a mode-locked Ti:Sapphire laser (170-fs pulse width, 76-MHz repetition rate; Mira 900F; Coherent, Santa Clara, Calif.) pumped by a 5-W Verdi laser (Coherent). The beam exiting the laser is deflected into the back port of an inverted Axiovert 100 microscope (Zeiss Inc.) and scanned across the sample via a personal-computer-controlled, galvanometer-driven x-y scanner (Series 603X; Cambridge Technology, Watertown, Mass.). The beam is reflected by a short-pass 675-nm dichroic beam splitter (Chroma Technology Corp., Brattleboro, Vt.) and focused onto the sample with a 100× oil immersion microscope objective (numerical aperture = 1.45; Zeiss Inc.). The excitation wavelength was 810 nm, which has been demonstrated to minimally excite YFP (13). The average power entering the microscope is approximately 30 mW (2.5 mW at the sample site). Two-photon-excited fluorescence from the sample is epicollected, discriminated with the dichroic beam splitter, filtered by a short-pass 600-nm filter (CVI Laser Corp., Livermore, Calif.), and detected. Two-dimensional (x-y plane) images (256 by 256 pixels) are acquired at a rate of 1 frame/s (pixel dwell time of 16 μs/pixel) with a single-photon counting photomultiplier tube (Hamamatsu Corp., Bridgewater, N.J.) and cover an area of 20 by 20 μm for the 100× microscope objective. An SBG39 wide-pass (322 to 654 nm) blue-green emission filter (CVI, Albuquerque, N.Mex.) is placed in front of the photomultiplier tube. Spectra are obtained with a SpectraPro-150 spectrograph equipped with a 300-groove/mm grating blazed at 500 nm (Acton Research Corp., Acton, Mass.) and a high-dynamic-range MicroMAX:512BFT CCD camera (Princeton Instruments), which is controlled by an ST-133 controller (Princeton Instruments). The spectrograph and camera settings are personal computer controlled through commercially available software (WinSpec/32, version 2.4.6.6; Roper Scientific, Inc., Trenton, N.J.). The CCD temperature is maintained at the minimum possible temperature (−45°C) for all of the experiments to ensure a low dark-noise level. The entrance slit of the spectrograph is set to a width of 0.5 mm. The spectrum acquisition time was 60 s. Switching between imaging and spectrum acquisition is achieved by changing the position of a built-in microscope mirror. When both two-photon images and spectra are acquired from the sample, the two-photon images are acquired and stored and then the emission spectrum is immediately acquired from the same depth (z) in the sample.
Autofluorescence of the cell is negligible. Dark-noise spectra are subtracted from the acquired sample spectra. Acquired images (256 by 256 pixels) were converted to TIFF format with IPLab, version 3.5.5 (Scanalytics), with no postprocessing modifications. Spectral data were imported into Microsoft Excel 2000 (Microsoft Corp.) and plotted as the intensity (arbitrary units) versus the emission wavelength (nanometers). Because of the low signal-to-noise ratio in the acceptor-alone case, a moving average (period = 10) function was used to plot the spectra. Peak intensity ratios were statistically analyzed by separate variance t tests, and a P value was calculated.
Transmission electron microscopy.
Ultrastructural analysis of pre-embedded immunostained serial thin sections (60 nm) of single microinjected cells was performed as previously described (15) on a Tecnai 12 BioTwin electron microscope (FEI Company, Hillsboro, Oreg.).
RESULTS
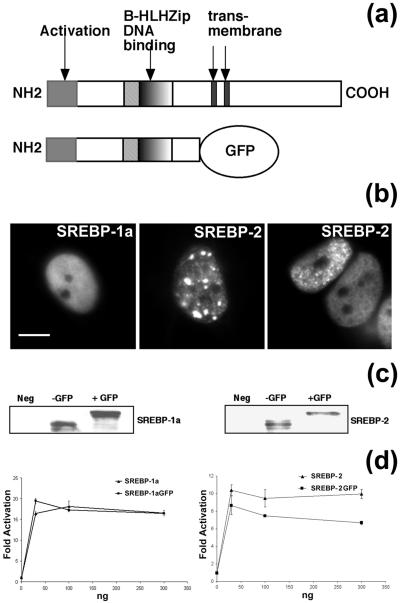
SREBP-1a and -2 are two related but distinct proteins whose overlapping functions have made it difficult to clearly define their individual roles in physiology. To characterize potential functional differences between them and monitor their potential in vivo distribution, GFP fusion proteins were generated with the mature forms (Fig. 1a) and found to constitutively localize to the nucleus. Interestingly, the two patterns of localization differed significantly. The N-terminal SREBP-1aGFP fusion construct was distributed diffusely throughout the nucleus (left part of Fig. 1b) in all of the cells analyzed (n = 136). In contrast, in the majority of cells (71%) expressing SREBP-2GFP (n = 276), the protein appeared to concentrate in subnuclear domains, exhibiting a speckled or focal pattern (center and right parts of Fig. 1b). The remaining 29% of the SREBP-2GFP-expressing cells displayed a diffuse fluorescence pattern similar to that of SREBP-1aGFP. With both expression plasmids, fluorescence was excluded from nucleoli. Western blot analysis was performed with SREBP-specific antibodies to confirm that the GFP fusion proteins were expressed at the expected size and accumulated to comparable levels as nontagged SREBP constructs (Fig. 1c). Additionally, to verify that the GFP tag did not compromise the ability of SREBP-1a or -2 to activate transcription, the activation potentials of the GFP-tagged and untagged proteins were compared with an LDL-R promoter luciferase reporter assay. When equivalent DNA amounts resulting in equal protein expression were transfected, both SREBP-1aGFP and SREBP-2GFP transactivated the LDL-R promoter as efficiently as the non-GFP-tagged versions of the proteins (Fig. 1d). Thus, there appears to be no adverse effect of the GFP tether on the expression or activity of either SREBP protein.
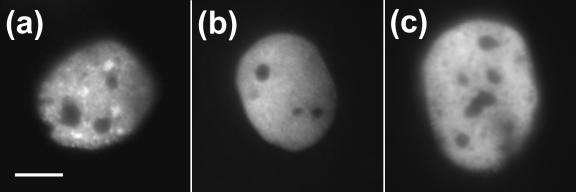
FIG. 1.
(a) Diagram of SREBP-GFP chimeras required for nuclear localization studies. (b) Micrographs of individual living cells expressing SREBP-1aGFP and SREBP-2GFP. SREBP-1a exhibits diffuse nuclear fluorescence. In contrast, SREBP-2 is localized to a speckled or focal fluorescence pattern. Scale bar, 10 μm. (c) Expression of GFP-tagged SREBP-1a and -2 in 293T cells. Immunoblot analysis with anti-SREBP-1a or SREBP-2 antibody to compare protein expression for the indicated constructs. An equivalent amount of extract from mock-transfected cells was analyzed and is designated Neg. (d) GFP-tagged SREBP-1a and -2 activate transcription of the LDL-R promoter. CV-1 cells were transfected with increasing amounts (as shown on the abscissa) of the indicated plasmids along with a constant amount of LDL-R luciferase reporter plasmid and control β-galactosidase plasmid to normalize for transfection efficiency. Luciferase activity was normalized to β-galactosidase activity, and fold activation was calculated.
The ability of expressed SREBP-2 to localize heterogeneously to discrete foci compared to the diffuse distribution of SREBP-1a represents a major distinction between the two SREBP proteins. Such morphological differences may govern the selectivity of homodimer versus heterodimer formation or localization. This, in turn, may translate to functional differences between the transcriptional activities of these proteins. To evaluate the potential homo- and heterodimerization of the two SREBPs in vivo and to determine if the localization of one of the SREBPs is altered when it is complexed as a heterodimer, two-photon imaging and spectroscopy to monitor fluorescence resonance energy transfer was used (13).
FRET.
All previous studies that have evaluated SREBP function have assumed the formation of homodimers when the monomeric expression constructs are individually expressed in cells or animals. However, these studies have all been performed with cells in which the potential for endogenous SREBPs or other bHLH proteins to dimerize with the expressed protein exists. The only evidence for potential heterodimer formation comes from the interpretation of dominant negative overexpression studies. To date, there are no previous studies that have directly evaluated the ability of SREBPs to form homo- or heterodimers inside a living cell.
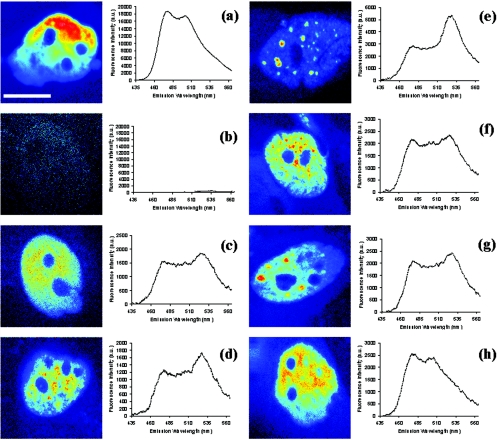
To examine the occurrence of heterodimers and homodimers between exogenously expressed SREBP-1a and -2 in vivo, we used a spectroscopic approach to monitor FRET at the subcellular level. CFP and YFP variants were used as the FRET donor-acceptor pair. Expression vectors encoding the mature domain of either SREBP-1a or SREBP-2 at the N terminus and either CFP or YFP at the C terminus were microinjected into HEp-2 cells either alone or in combination to monitor the presence of FRET. In general, SREBP-1a and -2 fusion proteins with CFP or YFP were localized to the nucleus and excluded from nucleoli. To evaluate localization and dimerization, two-photon fluorescence images were captured and the corresponding spectra of cells expressing the donor alone (SREBP-1aCFP), the acceptor alone (SREBP-2YFP), and both as homodimers (SREBP-1aCFP/SREBP-1aYFP, SREBP-2CFP/SREBP-2YFP) or as heterodimers (SREBP-1aCFP/SREBP-2YFP) were acquired with an excitation wavelength of 810 nm. Figure 2 shows images and the corresponding spectra from representative cells of each condition.
FIG. 2.
Two-photon fluorescence images and spectra of SREBP-1aCFP (a), SREBP-2YFP (b), SREBP-1aCFP/SREBP-1aYFP (c), SREBP-2CFP/SREBP-2YFP with a speckled fluorescence phenotype (d), SREBP-2CFP/SREBP-2YFP with a focal pattern (e), SREBP-1aCFP/SREBP-2YFP with a speckled fluorescence pattern (f), SREBP-1aCFP/SREBP-2YFP with a focal pattern (g), and SREBP-1aCFP/SREBP-2YFP with a diffuse fluorescence pattern (h). Scale bar, 10 μm. a.u., arbitrary units.
The two-photon-excited fluorescence image of a representative cell expressing SREBP-1aCFP alone (Fig. 2a) exhibits diffuse nuclear fluorescence that is devoid of nucleoli, as also shown in Fig. 1. The corresponding donor spectrum spans a region of 450 to 580 nm with two peaks (476 and 501 nm) (23). As expected, the fluorescence of the acceptor alone is negligible for SREBP-2YFP (Fig. 2b) since YFP has been shown to be minimally excited at 810 nm (13). When YFP is efficiently excited, a peak in the spectrum appears at 527 nm (13, 22, 23). SREBP-1a homodimer fluorescence (Fig. 2c) is confined to the nucleus in a diffuse pattern excluded from the nucleoli. The corresponding spectrum displays the characteristic donor CFP minor peak at 476 nm and major peak at 501 nm and an increased signal at 527 nm compared to the donor spectrum alone. The increase in YFP intensity compared to the individual spectra for the donor alone (Fig. 2a) and the acceptor alone (Fig. 2b) is due to the occurrence of FRET (13). To qualitatively determine the presence of FRET for the SREBP-1a homodimer pair, the ratios of the peak spectral intensities are compared. For the donor alone, the ratio of 501 to 476 nm is 0.93 and the ratio of 527 to 476 nm is 0.47. In the event of FRET, the YFP intensity value at 527 nm will increase; and thus, the ratio of 527 to 476 nm will concomitantly increase compared to that of the donor alone. The ratio of 501 to 476 nm will remain unchanged since it is characteristic of the donor's spectrum (13). For the SREBP-1a homodimer pair (Fig. 2c), the FRET ratio of 527 to 476 nm increases to 0.97, which is indicative of the occurrence of dimerization.
In agreement with Fig. 1, SREBP-2 homodimer fluorescence is confined to the nucleus and may either exhibit a diffuse pattern (not shown) or be localized to discrete domains, ranging from a speckled pattern (Fig. 2d) to larger foci (Fig. 2e). The spectra corresponding to the images in Fig. 2d and e display the characteristic donor CFP peaks at 476 and 501 nm and an increased signal at 527 nm compared to the donor spectrum alone. Similar spectra were obtained for cells exhibiting a diffuse fluorescence pattern. When fluorescence exhibits a speckled pattern (Fig. 2d), the ratio of 501 to 476 nm was 0.96 whereas the FRET ratio, 527 to 476 nm, was increased to 1.40, which is indicative of homodimer formation. When fluorescence is localized to larger discrete foci (Fig. 2e), the ratio of 501 to 476 nm is maintained at 0.96 whereas the FRET ratio, 527 to 476 nm, is increased to 1.89.
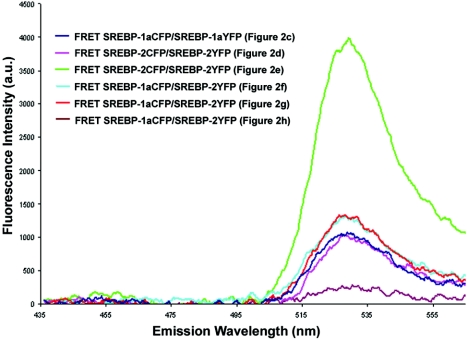
When examining SREBP-1a/SREBP-2 heterodimer interactions, three distinct patterns of fluorescence are observed. Figure 2f and g exhibit fluorescence patterns similar to that of SREBP-2 homodimer formation, where discrete nuclear domains are present. In the spectrum corresponding to the image of Fig. 2f, the ratios of 501 to 476 and 527 to 476 nm are 0.92 and 1.07, respectively. In the spectrum corresponding to the image of Fig. 2g, the ratios of 501 to 476 and 527 to 476 nm are 0.92 and 1.09, respectively. In contrast, cells exhibiting only a diffuse nuclear fluorescence pattern (Fig. 2h) had a ratio of 527 to 476 nm of 0.56. The ratio of 501 to 476 nm is consistent at 0.92. For each spectrum (Fig. 2c to h), the normalized donor spectrum (Fig. 2a) was subtracted, yielding the net FRET response shown in Fig. 3.
FIG. 3.
Net FRET responses of SREBP heterodimer and homodimer pairs. The normalized CFP spectrum (Fig. 2a) was subtracted from the corresponding individual spectra from Fig. 2c to h, resulting in the net FRET responses shown. a.u., arbitrary units.
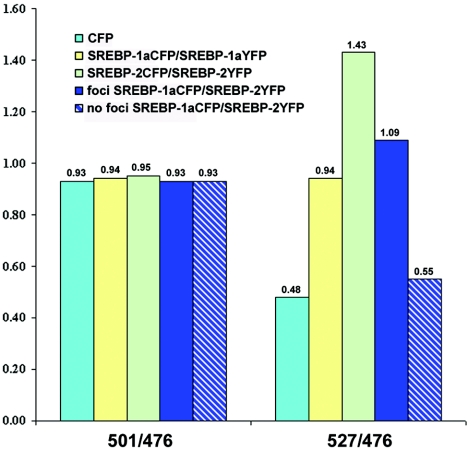
The average ratios for the donor alone and each dimer pair are reported in Fig. 4. The average ratios of 501 to 476 nm range from 0.93 to 0.95 and are not statistically significantly different, as expected, since this ratio is characteristic of the donor fluorescence. The average FRET ratio of 527 to 476 nm for the donor alone is 0.48 ± 0.004 (n = 17), which is indicative of the absence of FRET. For monitoring of homodimer formation, the average FRET ratio increases to an average of 0.94 ± 0.04 (n = 25) for SREBP-1a and 1.43 ± 0.06 (n = 23) for SREBP-2. For heterodimer pairs, the average FRET ratio is 1.09 ± 0.05 (n = 43) when fluorescence is localized to discrete nuclear domains and 0.55 ± 0.02 (n = 21) when fluorescence shows a diffuse pattern. Statistical analysis by a two-sample t test of the average FRET ratios demonstrated significant differences between the donor alone and homodimer pairs of SREBP-1a (P < 0.0001) and SREBP-2 (P < 0.0001). When comparing the donor alone to the average FRET ratio for heterodimer pairs, the P values were <0.0001 for cells exhibiting a focal pattern and 0.004 for cells without foci. More strikingly, a significant difference was observed between the average FRET ratios for heterodimers exhibiting foci and those for heterodimers that do not (P < 0.000005).
FIG. 4.
Average ratios of 501 to 476 and 527 to 476 nm. For SREBP-1a and -2 homodimers, 25 and 23 cells were analyzed, respectively. A total of 64 cells were assessed for the heterodimers. The average ratios of 501 to 476 nm for all of the cases examined ranged from 0.93 ± 0.002 to 0.96 ± 0.003. The average ratio of 527 to 476 nm for the donor alone is 0.48 ± 0.004. For homodimer formation, the FRET ratio increased to an average of 0.94 ± 0.04 for SREBP-1a and 1.43 ± 0.06 for SREBP-2. For heterodimer pairs, the average FRET ratio was 1.09 ± 0.05 when fluorescence was localized to discrete nuclear domains and 0.55 ± 0.02 when fluorescence showed a diffuse pattern.
Transcriptional activation of dimers in vivo.
We have, for the first time, demonstrated directly that exogenously expressed SREBP-1a and -2 homo- and heterodimers form in vivo with FRET. However, the FRET analysis does not provide any functional evidence about the transcriptional activation properties of the different dimer pairs. As mentioned earlier, all of the studies done to date could not unambiguously distinguish whether activation resulting from expression of a single SREBP isoform is due to a homodimer or a heterodimer formed between the expressed SREBP and another endogenous bHLH partner protein, possibly another SREBP. To determine whether the heterodimers we observed in vivo by microscopy-based techniques are functionally active and to compare their activation potential directly to that of the corresponding homodimer forms, we fused the coding sequence for individual SREBP monomers into tethered dimers. In this approach, a flexible polypeptide tether was used to preferentially link SREBP monomers into a single coding sequence, thus ensuring the preferential formation of intramolecular dimers. A similar approach was used for the E12/E47 bHLH proteins and the cJun-Fos b-Zip dimeric factors (3, 18). Two SREBP-1a or -2 monomers were joined by the tether to form the forced SREBP-1a or SREBP-2 homodimer, respectively. Similarly, an SREBP-1a monomer and an SREBP-2 monomer were joined to form the resulting forced SREBP-1a and -2 heterodimer. The SREBP heterodimer was made in both orientations. First, we evaluated the protein expression of the tethered SREBP fusions by immunoblotting with an antibody to detect the Flag epitope tag that is present at the amino terminus of all of the fusion constructs. This demonstrated that all of the fusion proteins were expressed at the expected sizes and at similar levels (Fig. 5a). We next directly compared the transcriptional activities of the homo- and heterodimers with an LDL-R promoter luciferase reporter assay. Here, the tethered SREBP heterodimers transactivated the LDL-R promoter to levels similar to those of the tethered SREBP homodimers (Fig. 5b). A more in-depth analysis of the SREBP tethered dimers has been recently published elsewhere (5) and shows that the tethered dimers bind DNA and activate transcription similarly to the monomerically expressed SREBPs. Also, in this report we show that activation by the tethered dimer is resistant to the expression of a dominant negative version of SREBP-1 that can efficiently dimerize and inhibit activation by monomerically expressed SREBPs (24).
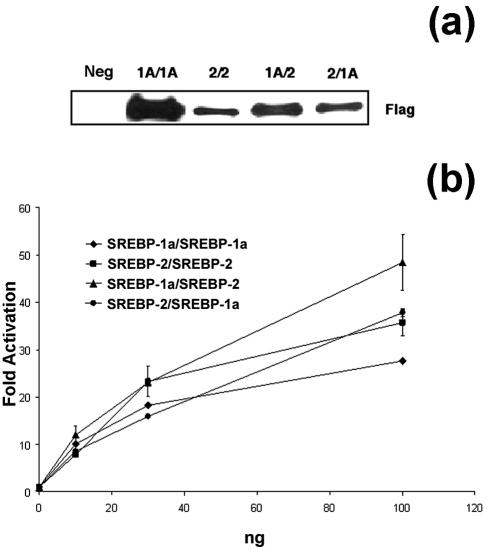
FIG. 5.
SREBP homo- and heterodimers activate transcription. (a) Expression of Flag-tagged SREBP tethered-dimer fusions in 293T cells. 293T cells were transfected with Flag-tagged SREBP tethered-dimer expression vectors. Total nuclear extracts were analyzed by immunoblotting with an antibody to the Flag epitope tag. An equivalent amount of extract from mock-transfected cells was analyzed and is designated Neg. (b) SREBP-1a and -2 tethered-dimer fusions were transfected into 293T cells in increasing amounts (as shown on the abscissa) along with a constant amount of LDL-R luciferase reporter plasmid and control β-galactosidase plasmid. Luciferase activity was normalized to β-galactosidase activity, and fold activation was calculated.
Recruitment of SREBP-1a by SREBP-2.
Because the individually expressed SREBPs were localized in different patterns, we next sought to determine if SREBP-2 could recruit SREBP-1a to a speckled or focal pattern of localization since our FRET data indicated that heterodimer protein-protein interactions were concomitant with this pattern of expression. Cells were microinjected with SREBP-1aGFP either alone or in the presence of coinjected untagged SREBP-2 (Fig. 6). Consistent with the results displayed in Fig. 1, SREBP-1aGFP was present exclusively in a diffuse nuclear pattern (Fig. 6c). In contrast, when untagged SREBP-2 was coinjected, 35% of the cells presented SREBP-1aGFP fluorescence in a speckled or focal pattern (Fig. 6a) similar to that observed in the FRET studies, whereas the remaining cells maintained a diffuse pattern of fluorescence (Fig. 6b). These results are consistent with a model in which the dimer between SREBP-1a and -2 results in redistribution of SREBP-1a into the pattern observed for SREBP-2.
FIG. 6.
Recruitment of SREBP-1a by SREBP-2. HEp-2 cells were microinjected with SREBP-1aGFP either alone or in combination with SREBP-2. Twenty-four hours later, fluorescence patterns were observed. Thirty-five percent of all cells expressing SREBP-1aGFP in the presence of SREBP-2 (n = 594) showed a speckled or focal fluorescence pattern (a), and the remaining cells showed a diffuse nuclear pattern of fluorescence (b). In contrast, 100% of all cells expressing SREBP-1aGFP alone (n = 136) exhibited a diffuse nuclear fluorescence pattern (c). Scale bar, 10 μm.
Serial deletions of SREBP-2.
In order to determine if there was a discrete region of SREBP-2 that was responsible for the speckled or focal fluorescence pattern observed with expressed SREBP-2 homodimers and heterodimers formed with SREBP-1a, serial deletions of the first 100, 150, 200, 250, 300, and 330 amino acids of SREBP-2 were fused to GFP (Fig. 7a) and their localization pattern was analyzed. Western blot analysis confirmed proper expression of the deletion mutant GFP fusion proteins (data not shown). In addition, deletion of the first 100 amino acids, which removes the SREBP transactivation domain (27), resulted in loss of transactivation (Fig. 7b). Similar results were obtained with all of the other mutant proteins (data not shown). To assess whether the truncated proteins were expressed and adopted a conformation similar to that predicted from the full-length protein, we performed dominant negative transactivation studies. We reasoned that if the bHLH-LZ DNA binding and dimerization domains of the deletion-containing proteins folded properly, they would maintain their native conformation and function as dominant negative inhibitors of the full-length SREBP-2 either by dimerizing with wild-type SREBP-2 or by binding to DNA as a homodimer and limiting access of wild-type SREBP-2 to its normal target sites in promoters. All of the deletion-containing proteins analyzed here were capable of inhibiting the activation mediated by SREBP-2 (Fig. 7c and data not shown).
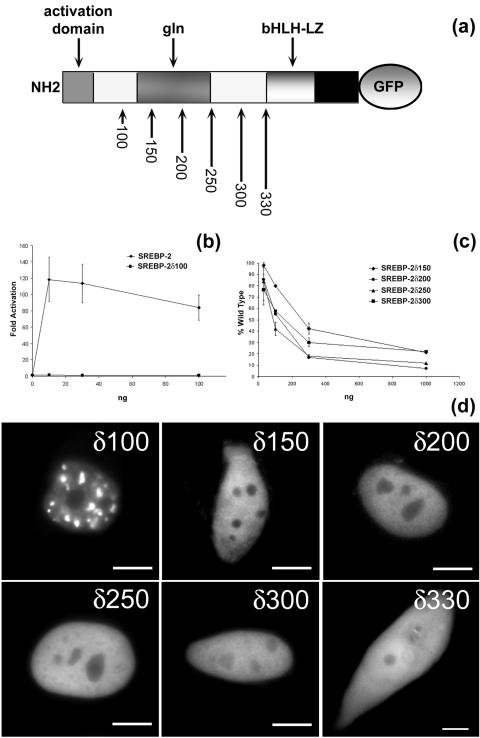
FIG. 7.
(a) SREBP-2 amino terminus deletions. (b) SREBP-2 amino terminus is required for transcriptional activity. CV-1 cells were transfected with increasing amounts of wild-type SREBP-2 or an SREBP-2 activation domain deletion construct (δ100) along with a constant amount of LDL-R luciferase reporter and control β-galactosidase plasmid, and fold activation was calculated. (c) Deletions of the SREBP-2 amino terminus result in a dominant negative inhibitor. CV-1 cells were transfected with a constant amount of wild-type SREBP-2 (30 ng) and increasing amounts of deletion-containing SREBP-2 (as shown on the abscissa). LDL-R luciferase reporter and control β-galactosidase were also transfected as for panel b. (d) Deletions of the SREBP-2 amino terminus expressed in HEp-2 cells. At 24 h postinjection, cells were fixed and imaged. Representative images for δ100GFP, δ150GFP, δ200GFP, δ250GFP, δ300GFP, and δ330GFP are shown. Scale bars, 10 μm.
Next, we compared the localization patterns of the mutant proteins to that of wild-type SREBP-2. Out of 118 cells expressing SREBP-2δ100GFP, 88% maintained a speckled or focal fluorescence pattern (Fig. 7d), which is similar to what was observed with the wild-type SREBP-2GFP construct (Fig. 1b). However, with SREBP-2δ150GFP, 92% of the cells (n = 257) displayed a diffuse fluorescence pattern in the nucleus. Similarly, for all cells expressing SREBP-2δ200GFP (n = 106), SREBP-2δ250GFP (n = 140), or SREBP-2δ300GFP (n = 149), fluorescence presented a diffuse nuclear pattern. An internal deletion of amino acids 101 to 249 also resulted in diffuse nuclear fluorescence (data not shown). Truncation of the first 330 amino acids (SREBP-2δ330GFP) resulted in cells with diffuse fluorescence in both the nucleus and the cytoplasm (n = 174). Taken together, these data indicate that deletion of amino acids 101 to 149 disrupts a domain that is responsible for localizing expressed SREBP-2 into a speckled or focal pattern and that the nuclear localization signal for SREBP-2 is disrupted when amino acids 301 to 330 are removed.
Structural characterization of foci.
An array of nuclear proteins has been shown to localize to speckles or foci within the cell, while fewer have been characterized to localize to discrete nuclear structures or physically identifiable bodies. In this regard, we noted a similarity in appearance between the SREBP-2 foci and the well-characterized PML-containing nuclear body (6, 12, 25). Thus, we wanted to evaluate whether there was a relationship between the two. First, we performed indirect immunofluorescence studies with corresponding antibodies and expression constructs to see if the SREBP-2 and PML foci were identical. In 100% of cells (n = 136) expressing SREBP-1aGFP, immunostaining for endogenous PML resulted in a normal pattern of PML localization corresponding to an average of 10 to 20 PML bodies per nucleus (Fig. 8a and b). In contrast, endogenous PML staining in cells expressing SREBP-2GFP showed a significant loss or depletion of PML staining (96%, n = 276) from the corresponding bodies (Fig. 8c and d). In some cases (4%), PML could be observed tracking out of the PML-containing nuclear bodies (Fig. 8e and f). To further determine whether expression of SREBP-2 results in loss or depletion of PML from its nuclear bodies, we coexpressed untagged SREBP-2 along with PMLGFP. This resulted in a decrease or absence of PML fluorescence within its corresponding bodies (data not shown). Similarly, cells expressing SREBP-2δ100GFP exhibited a loss or depletion of PML-containing nuclear body fluorescence (93%, n = 118) (Fig. 8g and h). In some instances (7%), colocalization of SREBP-2GFP and PML was observed (data not shown). In contrast, expression of the N-terminal deletion constructs of SREBP-2 that exhibited a diffuse pattern on their own did not alter PML localization (data not shown). In addition, similar studies with SUMO-1 to identify the PML-containing nuclear bodies resulted in the same staining patterns (data not shown). Taken together, these data suggest that SREBP-2 foci may correspond to the well-characterized PML bodies and demonstrate that there is a dynamic relationship between the nuclear body localization of SREBP-2 and components of the PML-containing nuclear body.
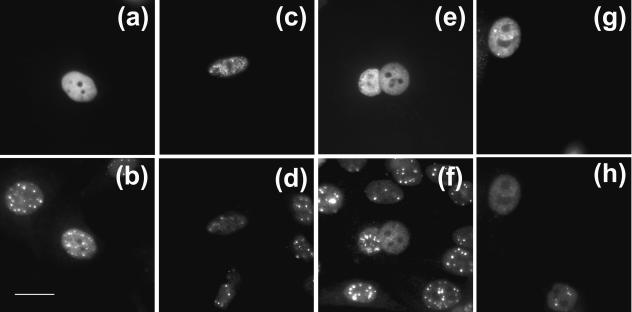
FIG. 8.
SREBP and PML localization. SREBP-1aGFP (a), SREBP-2GFP (c, e), and SREBP-2δ100GFP (g) were expressed in HEp-2 cells. At 24 h postinjection, cells were fixed and immunostained for PML. Corresponding images for PML are also presented (b, d, f, h). Scale bar, 20 μm.
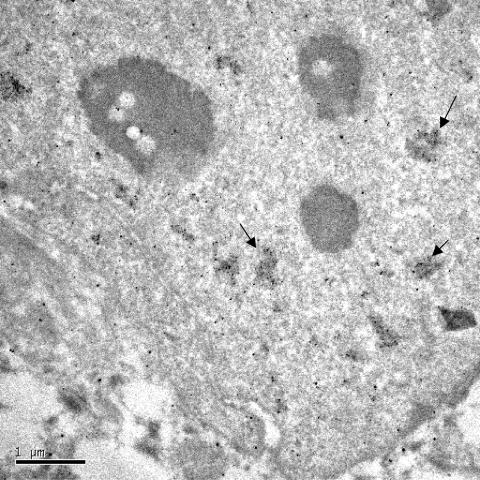
In parallel, to directly characterize the SREBP-2 foci, transmission electron microscopy was performed. Expressed SREBP-2GFP was detected by pre-embedded immunogold labeling of the GFP molecule. At the ultrastructural level, SREBP-2GFP resides in discrete electron-dense nuclear structures (Fig. 9) that are 0.5 to 1 μm in size, which is consistent with the size of the PML-containing nuclear bodies (12). Representative SREBP-2-containing bodies are indicated by arrows. In contrast to PML, which localizes to the outer shell of the body, SREBP-2 is dispersed throughout the body, similar to the coactivator CBP (12). Neighboring uninjected cells did not exhibit gold particle labeling (data not shown).
FIG. 9.
Ultrastructural analysis of SREBP-2GFP localization in HEp-2 cells by immunoelectron microscopy. Silver-enhanced Nanogold pre-embedding immunoelectron microscopy was performed with affinity-purified mouse antibodies to GFP. SREBP-2GFP was localized in a nuclear body with gold particles dispersed throughout the body. Representative SREBP-2-containing bodies are indicated by arrows.
DISCUSSION
SREBPs belong to the bHLH family of transcription factors, which are known to bind DNA and activate transcription as dimers. The dimerization properties of individual members of this family significantly influence their activity, and some members such as cMyc do not bind DNA at all as homodimers (7). Because the three major SREBP isoforms are coexpressed in most cells (9), the sum total of SREBP activity is dependent on the ratio of the individual isoforms coupled with their inherent ability to form homo- and heterodimers. SREBP-1a and -1c are encoded from spliced variants of the same gene, have identical bHLH-LZ domains, and would be expected to form homo- and heterodimers on the basis of the ratio of the two protein isoforms in the cell.
The bHLH-LZ domains of SREBP-1 and -2 are highly conserved (74% identity), and a recent report has suggested that overexpression of a dominant negative SREBP-1a protein can decrease the activity mediated by both SREBP-1a and -2 (24). However, a direct demonstration that the two proteins actually form heterodimers in cells and the activity mediated by the individual homodimers relative to that mediated by a putative heterodimer has not been directly investigated.
In the present studies, we have used novel imaging techniques including two-photon-excited imaging and spectroscopy FRET coupled with key molecular biology methods in which we connected individual SREBP isoforms together into tethered dimers (18) to analyze the subnuclear spatial localization and function of SREBP homo- and heterodimers.
These data support the in vivo occurrence and uncover distinct subnuclear spatial localization of both exogenously expressed SREBP-1a and -2 homodimers and heterodimers and provide potential insight into their differential regulation. By using two-photon imaging and spectroscopy FRET to examine protein-protein interactions in subfemtoliter volumes in vivo (13), we demonstrated that when exogenously expressed SREBP-1a homodimerizes in vivo, it is localized diffusely in the nucleus. FRET signals from similarly expressed SREBP-2 homodimers were detected both within a diffuse fluorescence pattern and in discrete nuclear domains ranging from speckles to larger foci. Importantly, SREBP-1a homodimers never exhibited a speckled or focal pattern in our studies. These techniques also provide novel evidence to directly support the existence of SREBP-1a and -2 heterodimers in vivo. Intense FRET signals from heterodimers were detected within cells exhibiting fluorescence in discrete nuclear domains. Notably, there was a significant FRET signal decrease in cells in which both SREBPs were expressed, and the fluorescence was present in a diffuse nuclear pattern, strongly suggesting that heterodimers are residing inside the discrete nuclear domains.
The FRET analysis data correlate with the different nuclear localization patterns observed for the individual GFP-tagged versions of mature SREBP-1a and -2. SREBP-1aGFP was distributed diffusely throughout the nucleus, whereas SREBP-2GFP expression was localized in subnuclear domains, exhibiting a speckled or focal pattern in the majority of cells. The FRET studies indicated that the SREBP-1a and -2 heterodimers were localized in subnuclear domains similarly to SREBP-2 homodimers, suggesting that SREBP-1a localization was influenced by the coexpression of SREBP-2. When untagged SREBP-2 was coexpressed, it also resulted in the redistribution of SREBP-1aGFP in a speckled or focal pattern. Taken together, these results indicate that SREBP-2 coexpression results in recruitment of SREBP-1aGFP into speckles or foci, indicating that SREBP-2 can affect the localization of SREBP-1a in a dominant fashion.
Our two-photon imaging and spectroscopy FRET studies indicate that both SREBP homo- and heterodimers occur in living cells, but they do not evaluate the functional significance of the homo- versus heterodimer. In order to accurately measure the activation potential of SREBP homo- and heterodimers and avoid complications from endogenous proteins, we used a tethered-dimer strategy. By linking two SREBP monomers with a flexible polypeptide spacer, we could minimize any potential dimerization with endogenous proteins, which is a significant complication when constructs expressing only the monomeric SREBPs are analyzed. We constructed tethered dimers of SREBP-1a and -2 in both orientations and analyzed their abilities to activate the LDL-R promoter in a luciferase reporter-based transactivation assay. The results show for the first time that the SREBP homo- and heterodimers activate transcription to similar levels. Similar results were obtained for other SREBP target promoters (5).
Our studies provide clear and significant new information about differential trafficking of exogenously expressed SREBP isoforms in intact cells. Because the exogenously expressed SREBP-GFP fusion proteins may display different localization patterns than the corresponding endogenously expressed proteins, it will be important to extend these observations to determine if endogenously expressed proteins behave similarly. However, these additional studies await the development of more specific antibodies and modified imaging techniques to evaluate the low levels of endogenously expressed SREBPs.
A recent report suggests that dimerization of SREBPs occurs prior to transport into the nucleus (14). When combined with our data, these results indicate that upon nuclear import, dimers are differentially directed to discrete locations, and more specifically in the case of SREBP-2, to discrete electron-dense nuclear structures. The dynamic interplay between SREBP-2 and PML/SUMO-1 trafficking suggests that they may reside in the same bodies. In agreement with previous findings, PML-containing nuclear bodies may serve as functional complexes whose constituents are dynamic, i.e., CBP (4, 12) and, as shown here, SREBP-2 dimers.
We have also demonstrated that amino acids 101 to 149 are necessary for expressed SREBP-2's ability to localize within speckles or foci. A previous report suggested that SREBP-1a and -2 were recruited to punctate foci only when p300 was also coexpressed (8). p300 interacts with the SREBP-activation domain (30), which is located at the extreme amino terminus of both SREBP-1a and -2. Our studies indicate that SREBP-2 localizes to foci independently of p300 overexpression and is independent of the amino-terminal activation domain as SREBP-2δ100GFP was efficiently localized to foci. Fluorescence cross-correlation spectroscopy would allow monitoring of the changes in complex diffusion rates and the differences in the molecular sizes of the mobile SREBP-coactivator complexes to definitively determine the participants in these complexes and where these interactions occur.
Overall, our results demonstrate that there are differences in the spatial localization of homodimers and heterodimers between exogenously expressed SREBP-1a and -2. Additionally, our data provide the basis for further studies to understand the mechanism and functional significance of this differential localization. Because the dimerization specificity for the individual bHLH proteins can have a significant impact on function, it will be important to evaluate the dimerization and activation properties of other family members with a methodology similar to the one used in the present study.
Acknowledgments
We acknowledge Bruce J. Tromberg, director of the Laser Microbeam and Medical Program; Michael W. Berns, Medical Free Electron Laser Program; and Kathryn E. Osann for statistical analysis and expert advice and Tatiana B. Krasieva, Linda Li, Katie Lee, and Marisa Magana for technical expertise.
This work was supported by the National American Heart Association (993005N to V.L.); the National Institutes of Health (HL48044 to T.O.); the National Institutes of Health Research Resource, Laser Microbeam, and Medical Program (P41R01192); and the Air Force of Scientific Research, Medical Free Electron Laser Program (FA9550-04-1-01-01).
REFERENCES
- 1.Athanikar, J. N., and T. F. Osborne. 1998. Specificity in cholesterol regulation of gene expression by coevolution of sterol regulatory DNA element and its binding protein. Proc. Natl. Acad. Sci. USA 95:4935-4940. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Athanikar, J. N., H. B. Sanchez, and T. F. Osborne. 1997. Promoter selective transcriptional synergy mediated by sterol regulatory element binding protein and Sp1: a critical role for the Btd domain of Sp1. Mol. Cell. Biol. 17:5193-5200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bakiri, L., K. Matsuo, M. Wisniewska, E. F. Wagner, and M. Yaniv. 2002. Promoter specificity and biological activity of tethered AP-1 dimers. Mol. Cell. Biol. 22:4952-4964. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Boisvert, F. M., M. J. Kruhlak, A. K. Box, M. J. Hendzel, and D. P. Bazett-Jones. 2001. The transcription coactivator CBP is a dynamic component of the promyelocytic leukemia nuclear body. J. Cell Biol. 152:1099-1106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Datta, S., and T. F. Osborne. 2005. Activation domains from both monomers contribute to transcriptional stimulation by SREBP dimers. J. Biol. Chem. 280:3338-3345. [DOI] [PubMed] [Google Scholar]
- 6.Dyck, J. A., G. G. Maul, W. H. Miller, J. D. Chen, A. Kakizuka, and R. M. Evans. 1994. A novel macromolecular structure is a target of the promelocyte-retinoic acid receptor oncoprotein. Cell 76:333-343. [DOI] [PubMed] [Google Scholar]
- 7.Eisenman, R. N. 2001. Deconstructing myc. Genes Dev. 15:2023-2030. [DOI] [PubMed] [Google Scholar]
- 8.Giandomenico, V., M. Simonsson, E. Gronroos, and J. Ericsson. 2003. Coactivator-dependent acetylation stabilizes members of the SREBP family of transcription factors. Mol. Cell. Biol. 23:2587-2599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Horton, J. D., J. L. Goldstein, and M. S. Brown. 2002. SREBPs: activators of the complete program of cholesterol and fatty acid synthesis in the liver. J. Clin. Investig. 109:1125-1131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Horton, J. D., N. A. Shah, J. A. Warrington, N. N. Anderson, S. W. Park, M. S. Brown, and J. L. Goldstein. 2003. Combined analysis of oligonucleotide microarray data from transgenic and knockout mice identifies direct SREBP target genes. Proc. Natl. Acad. Sci. USA 100:12027-12032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Horton, J. D., I. Shimomura, M. S. Brown, R. E. Hammer, J. L. Goldstein, and H. Shimano. 1998. Activation of cholesterol synthesis in preference to fatty acid synthesis in liver and adipose tissue of transgenic mice overproducing sterol regulatory element-binding protein-2. J. Clin. Investig. 101:2331-2339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.LaMorte, V. J., J. A. Dyck, R. L. Ochs, and R. M. Evans. 1998. Localization of nascent RNA and CBP with the PML-containing nuclear body. Proc. Natl. Acad. Sci. USA 95:4991-4996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.LaMorte, V. J., A. Zoumi, and B. J. Tromberg. 2003. Spectroscopic approach for monitoring two-photon excited fluorescence resonance energy transfer from homodimers at the subcellular level. J. Biomed. Optics 8:357-361. [DOI] [PubMed] [Google Scholar]
- 14.Lee, S. J., T. Sekimoto, E. Yamashita, E. Nagoshi, A. Nakagawa, N. Imamoto, M. Yoshimura, H. Sakai, K. T. Chong, T. Tsukihara, and Y. Yoneda. 2003. The structure of importin-β bound to SREBP-2: nuclear import of a transcription factor. Science 302:1571-1575. [DOI] [PubMed] [Google Scholar]
- 15.Liaw, L. H., and M. W. Berns. 1981. Electron microscope autoradiography on serial sections of preselected single living cells. J. Ultrastruct. Res. 75:187-194. [DOI] [PubMed] [Google Scholar]
- 16.Magana, M. M., and T. F. Osborne. 1996. Two tandem binding sites for sterol regulatory element binding proteins are required for sterol regulation of fatty-acid synthase promoter. J. Biol. Chem. 271:32689-32694. [DOI] [PubMed] [Google Scholar]
- 17.Matsuda, M., B. S. Korn, R. E. Hammer, Y.-A. Moon, R. Kumuro, J. D. Horton, J. L. Goldstein, M. S. Brown, and I. Shimomura. 2001. SREBP cleavage-activating protein (SCAP) is required for increased lipid synthesis in liver induced by cholesterol deprivation and insulin elevation. Genes Dev. 15:1206-1216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Neuhold, L. A., and B. Wold. 1993. HLH forced dimers: tethering MyoD to E47 generates a dominant positive myogenic factor insulated from negative regulation by Id. Cell 74:1033-1042. [DOI] [PubMed] [Google Scholar]
- 19.Ochs, R. L., T. Stein, Jr., and E. M. Tan. 1994. Coiled bodies in the nucleolus of breast cancer cells. J. Cell Sci. 107:385-399. [DOI] [PubMed] [Google Scholar]
- 20.Osborne, T. 2001. Creating a SCAP-less liver keeps SREBPs pinned in the ER membrane and prevents increased lipid synthesis in response to low cholesterol and high insulin. Genes Dev. 15:1873-1878. [DOI] [PubMed] [Google Scholar]
- 21.Osborne, T. F. 2000. Sterol regulatory element binding proteins (SREBPs): key regulators of nutritional homeostasis and insulin action. J. Biol. Chem. 275:32379-32382. [DOI] [PubMed] [Google Scholar]
- 22.Periasamy, A. 2001. Fluorescence resonance energy transfer microscopy: a minireview. J. Biomed. Optics 6:287-291. [DOI] [PubMed] [Google Scholar]
- 23.Pollok, B. A., and R. Heim. 1999. Using GFP in FRET-based applications. Trends Cell Biol. 9:57-60. [DOI] [PubMed] [Google Scholar]
- 24.Rishi, V., J. Gal, D. Krylov, J. Fridriksson, M. S. Boysen, S. Mandrup, and C. Vinson. 2004. SREBP-1 dimerization specificity maps to both the helix-loop-helix and leucine zipper domains: use of a dominant negative. J. Biol. Chem. 279:11863-11874. [DOI] [PubMed] [Google Scholar]
- 25.Ruggero, D., Z. G. Wang, and P. P. Pandolfi. 2000. The puzzling multiple lives of PML and its role in the genesis of cancer. Bioessays 22:827-835. [DOI] [PubMed] [Google Scholar]
- 26.Sanchez, H. B., L. Yieh, and T. F. Osborne. 1995. Cooperation by sterol regulatory element-binding protein and Sp1 in sterol regulation of low density lipoprotein receptor gene. J. Biol. Chem. 270:1161-1169. [DOI] [PubMed] [Google Scholar]
- 27.Sato, R., J. Yang, X. Wang, M. J. Evans, Y. K. Ho, J. L. Goldstein, and M. S. Brown. 1994. Assignment of the membrane attachment, DNA binding, and transcriptional activation domains of sterol regulatory element-binding protein-1 (SREBP-1). J. Biol. Chem. 209:17267-17273. [PubMed] [Google Scholar]
- 28.Shimano, H., J. D. Horton, R. E. Hammer, I. Shimomura, M. S. Brown, and J. L. Goldstein. 1996. Overproduction of cholesterol and fatty acids causes massive liver enlargement in transgenic mice expressing truncated SREBP-1a. J. Clin. Investig. 98:1575-1584. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Shimano, H., J. D. Horton, I. Shimomura, R. E. Hammer, M. S. Brown, and J. L. Goldstein. 1997. Isoform 1c of sterol regulatory element binding protein is less active than isoform 1a in livers of transgenic mice and cultured cells. J. Clin. Investig. 99:846-854. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Toth, J. I., S. Datta, J. N. Athanikar, L. P. Freedman, and T. F. Osborne. 2004. Selective coactivator interactions in gene activation by SREBP-1a and -1c. Mol. Cell. Biol. 24:8288-8300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Yieh, L., H. B. Sanchez, and T. F. Osborne. 1995. Domains of transcription factor Sp1 required for synergistic activation with sterol regulatory element binding protein 1 of low density lipoprotein receptor promoter. Proc. Natl. Acad. Sci. USA 92:6102-6106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Zoumi, A., A. Yeh, and B. J. Tromberg. 2002. Imaging cells and extracellular matrix in vivo by using second-harmonic generation and two-photon excited fluorescence. Proc. Natl. Acad. Sci. USA 99:11014-11019. [DOI] [PMC free article] [PubMed] [Google Scholar]