Abstract
Sterile 20–like kinases Mst1 and Mst2 (Mst1/2) and large tumor suppressor 1/2 are core kinases to mediate Hippo signaling in maintaining tissue homeostasis. We have previously demonstrated that Smad ubiquitin (Ub) regulatory factor 1 (Smurf1), a HECT-type E3 ligase, ubiquitinates and in turn destabilizes large tumor suppressor 1/2 to induce the transcriptional output of Hippo signaling. Here, we unexpectedly find that Smurf1 interacts with and polyubiquitinates Mst1/2 by virtue of K27- and K29-linked Ub chains, resulting in the proteasomal degradation of Mst1/2 and attenuation of their tumor-suppressor functions. Among the potential Ub acceptor sites on Mst1/2, K285/K282 are conserved and essential for Smurf1-induced polyubiquitination and degradation of Mst1/2 as well as transcriptional output of Hippo signaling. As a result, K285R/K282R mutation of Mst1/2 not only negates the transcriptional output of Hippo signaling but enhances the tumor-suppressor functions of Mst1/2. Together, we demonstrate that Smurf1-mediated polyubiquitination on K285/K282 of Mst1/2 destabilizes Mst1/2 to attenuate their tumor-suppressor functions. Thus, the present study identifies Smurf1-mediated ubiquitination of Mst1/2 as a hitherto uncharacterized mechanism fine-tuning the Hippo signaling pathway and may provide additional targets for therapeutic intervention of diseases associated with this important pathway.
Keywords: Smurf1, Mst1/2, ubiquitination, Hippo signaling, tumorigenesis
Hippo signaling is an evolutionarily conserved pathway that controls the organ size by regulating cell proliferation and apoptosis and regulates a variety of biological processes, such as organ development, tissue homeostasis, and tumorigenesis (1, 2, 3). Sterile 20–like kinases Mst1 and Mst2 (Mst1/2), mammalian homologs of Drosophila Hippo, are core kinases of Hippo signaling pathway and share ∼75% identical amino acid sequence (4, 5). Activation of Mst1/2 phosphorylates the large tumor suppressor 1 and 2 (Lats1/2)–Mps 1 binder (Mob1), Lats1/2–Mob1 in turn phosphorylates Yes-associated protein (YAP) and transcriptional coactivator with PDZ-binding motif (TAZ), resulting in their ubiquitination and proteasomal degradation. In contrast, inactivation of either Mst1/2 or Lats1/2–Mob1 stabilizes YAP/TAZ and induces YAP/TAZ nuclear translocation to bind to the TEA domain transcription factor (TEAD), resulting in the transcriptional output of proproliferative and prosurvival genes, such as the connective tissue growth factor (Ctgf) and cysteine-rich angiogenic inducer 61 (Cyr61) (3, 6). As a result, genetic ablation of both Mst1 and Mst2 causes the enlarged livers and spontaneous hepatocellular carcinoma (HCC) in mice (7, 8, 9).
Ubiquitination is a universal protein post-translational modification and involved in a variety of biological processes, such as inflammation, metabolism, DNA damage and repair, autophagy, and tumorigenesis (10, 11, 12). Ubiquitination is initiated by transferring ubiquitin (Ub) from an Ub-activating enzyme (E1) to an Ub-conjugating enzyme (E2) and produces a covalently linked intermediate (E2–Ub). Ub protein ligases (E3 ligases) determine the substrate specificity of ubiquitination by the covalent attachment of Ub to substrate proteins (13). Currently, there are over 600 putative E3 ligases classified into three families, homologous to E6-AP carboxy terminus (HECT), RING, and ring-between ring–ring families (14, 15). Smad Ub regulatory factor 1 (Smurf1), a HECT-type E3 ligase, is initially believed to regulate Smad1/5 protein stability in the transforming growth factor-β and bone morphogenic protein signaling pathways (16). Smurf1 ubiquitinates a variety of substrates that are involved in the cell proliferation and differentiation, cell stemness, chromatin organization and dynamics, DNA damage and repair, genomic integrity, gene expression, and cell migration and invasion (17, 18). All these events are inextricably linked to the tumorigenesis. Smurf1 is also highly expressed in a variety of tumors and functions as a tumor promoter by ubiquitinating tumor suppressors (19, 20).
We have previously demonstrated that Smurf1 ubiquitinates Lats1/2 to induce the transcriptional output of Hippo (21). Here, we have further identified that Smurf1 ubiquitinates Mst1/2 predominantly on K285/K282 to promote their proteasomal degradation and thereby attenuates their tumor-suppressor functions. Thus, the present study has uncovered Smurf1 in conjunction with Mst1/2 ubiquitination as a hitherto uncharacterized mechanism controlling Hippo signaling.
Results
Smurf1 targets Mst1/2 to regulate the transcriptional output of Hippo signaling
Knockdown of dSmurf1 (Drosophila homolog of Smurf1/2) stabilizes Wts (Drosophila homolog of Lats1/2) and increases the phosphorylation levels of Yki (Drosophila homolog of YAP and TAZ) and ultimately regulates Hippo signaling transduction. In contrast, overexpression of dSmurf1 has no effect on modulating Wts stability (22). In order to determine whether Smurf1 regulates Hippo signaling in mammalian cells, we performed Tead4-luciferase reporter and quantitative PCR (qPCR) analyses in 293T cells. Knockdown of Smurf1 significantly decreased the Tead4-luciferase reporter activities as well as the mRNA levels of Ctgf and Cyr61, target genes of Hippo signaling pathway (Fig. 1, A and B). In contrast, overexpression of Smurf1 in 293T cells unexpectedly and robustly increased the Tead4-luciferase activities and the mRNA levels of Ctgf and Cyr61 (Fig. 1, C and D), which appears to be at odds with the result in Drosophila (22).
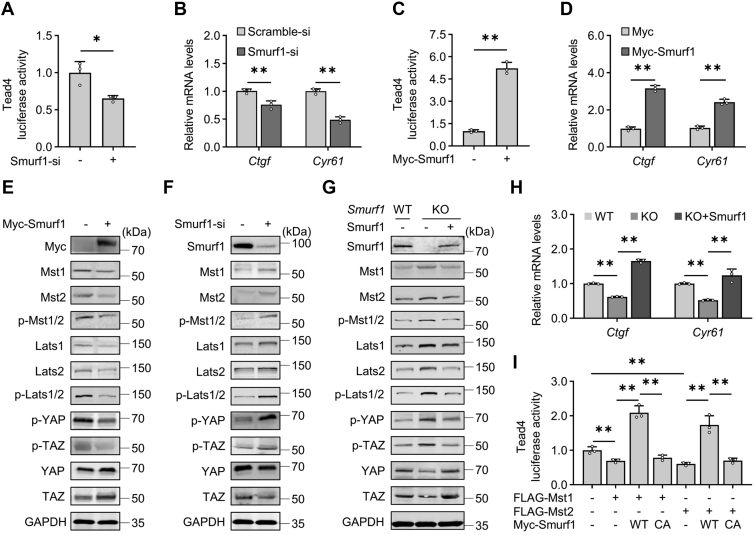
Figure 1.
Smurf1 induces the transcriptional output of Hippo signaling pathway by targeting Mst1/2.A–D, Tead4-luciferase reporter assays and qPCR analyses for mRNA levels of Ctgf and Cyr61 in 293T cells after transfection with scramble-/Smurf1-siRNA or vector-/Smurf1-expressing construct for 48 h. E and F, Western blotting analyses in 293T cells after transfection with the indicated siRNAs or constructs for 24 h. G and H, Western blotting analyses and qPCR analyses in WT or Smurf1-ablated (KO) MEFs after infection with control (−) or Smurf1-expressing lentivirus (+) for 48 h. I, Tead4-luciferase reporter assays in 293T cells after transfection with or without FLAG-Mst1/2, WT Myc-Smurf1, or inactive form of Myc-Smurf1 (CA) for 48 h. Numerical data were presented as mean ± SD. n = 3. One-way ANOVA and Tukey–Kramer multiple comparisons or Student’s t test. ∗p < 0.05 and ∗∗p < 0.01. CA, C699A; Ctgf, connective tissue growth factor; Cyr61, cysteine-rich angiogenic inducer 61; MEF, mouse embryonic fibroblast; Mst1/2, Mst1 and Mst2; qPCR, quantitative PCR; Smurf1, Smad ubiquitin regulatory factor 1; TEAD, TEA domain transcription factor.
To determine the exact role of Smurf1 in the regulation of Hippo signaling, we next performed Western blotting analyses in 293T cells. Overexpression of Smurf1 robustly and moderately negated the protein levels of p-Lats1/2, p-YAP, p-TAZ, and of Mst1/2, p-Mst1/2, Lats1/2, respectively, whereas it noticeably enhanced the protein levels of YAP and TAZ (Fig. 1E). In contrast, Smurf1 siRNA robustly and moderately induced the protein levels of p-Lats1/2, p-YAP, p-TAZ, and of Mst1/2, p-Mst1/2, Lats1/2, respectively, whereas it considerably reduced the protein levels of YAP and TAZ (Fig. 1F). To confirm the universality, we cultured primary mouse embryonic fibroblasts (MEFs) from WT and Smurf1−/− (KO) embryos and infected them with or without Smurf1-expressing lentiviruses. Western blotting analyses indicated that ablation of Smurf1 exhibited more robustly than Smurf1 siRNA in regulating the expression of these key components of Hippo signaling pathway, whereas lentiviral overexpression of Smurf1 completely restored the effects of Smurf1 ablation in MEFs (Fig. 1G). In line with these observations, qPCR analyses revealed that Smurf1 ablation in MEFs significantly decreased the mRNA levels of Ctgf and Cyr61, whereas lentiviral overexpression of Smurf1 not only restored the Smurf1 ablation–negated mRNA levels but also increased the basal mRNA levels (Fig. 1H).
Either upregulation (overexpression) or downregulation (knockdown/KO) of Smurf1 expression affects the Mst1/2 and p-Mst1/2 protein levels to the same extent (Fig. 1, E–G); this finding suggests that Smurf1-mediating Mst1/2 protein level changes cause the corresponding changes in p-Mst1/2 protein levels. Notably, either upregulation or downregulation of Smurf1 expression affects p-Lats1/2 levels more profoundly than Lats1/2 levels (Fig. 1, E–G). As Mst1/2 lie on the upstream of Lats1/2 (3, 6) and we have previously demonstrated that Smurf1 ubiquitinates and destabilizes Lats1/2 to decrease the p-Lats1/2 levels (21), the corresponding p-Lats1/2 level changes in response to Smurf1 could be resulted from Smurf1-mediating regulation of both Mst1/2 and Lats1/2. This notion was supported further by the Tead4-luciferase reporter assays, which showed overexpression of Smurf1 but not its inactive mutant (C699A [CA]) (23) not only restored the Mst1/2 negated Tead4-luciferase activities but also increased the basal levels of them (Fig. 1I). Thus, Smurf1 induces the transcriptional output of Hippo signaling by targeting not only Lats1/2 but also Mst1/2.
Smurf1 induces the proteasomal degradation of Mst1/2
To determine the potential role of Smurf1 in destabilizing Mst1/2, we overexpressed or knocked down Smurf1 in 293T cells. Overexpression of Smurf1 did not affect the mRNA levels of Mst1/2 but decreased the exogenous and endogenous protein levels of Mst1/2 in a dose-dependent manner (Figs. 2, A–C and S1A). Whereas the E3 ligase inactive mutant (CA) of Smurf1 had no effect on Mst1/2 protein levels (Fig. 2, D and E), suggesting that E3 ligase activity of Smurf1 is required for negating Mst1/2 expression. To investigate whether endogenous Smurf1 is sufficient to regulate the abundance of Mst1/2, we knocked down Smurf1 in 293T cells and performed Western blotting analyses. As expected, Smurf1 siRNA or shRNA had no effect on mRNA levels of Mst1/2 but significantly upregulated the protein levels of Mst1/2 (Figs. 2F and S1B). However, treatment of cells with MG132, a proteosome inhibitor, but not with chloroquine, a lysosomal inhibitor, completely reversed the Smurf1’s effects on Mst1/2 protein levels (Figs. 2, G and H and S1C). To investigate whether Smurf1 downregulates Mst1/2 by reducing their stability, we performed cycloheximide (CHX) chase assays (24) and measured the degradation rate of Mst1/2. In the absence of Smurf1, the protein half-life (t1/2) values of Mst1/2 were approximately 7.5 and 7.0 h, respectively, whereas in the presence of Smurf1, the t1/2 values of Mst1/2 were approximately 3.8 and 4.0 h, respectively (Fig. 2I). In contrast, Smurf1 siRNA increased the t1/2 values of Mst1/2 by approximately 90% and 58% (3.3 and 4.7 h to 7.3 and 7.4 h), as compared with scramble siRNA, respectively (Fig. 2J). Likewise, the t1/2 values of Mst1/2 in WT MEFs were approximately 2.9 and 1.8 h, respectively, whereas in Smurf1-ablated MEFs, the t1/2 values of Mst1/2 were increased to approximately 13.8 and 6.5 h, respectively (Fig. 2K). Thus, Smurf1 destabilizes Mst1/2 by promoting their proteasomal degradation.
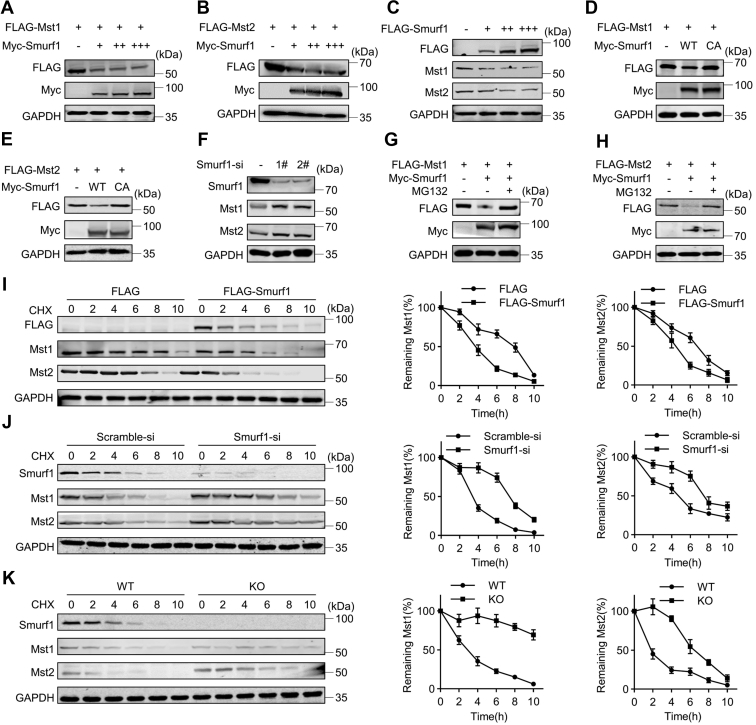
Figure 2.
Smurf1 induces Mst1/2 proteasomal degradation.A–C, Western blotting analyses in 293T cells after transfection with or without FLAG-Mst1/2 and the increasing dosages of Myc-Smurf1 for 24 h. D–F, Western blotting analyses in 293T cells after transfection with the indicated constructs or siRNAs for 48 h. G and H, Western blotting analyses in 293T cells transfected with the indicated constructs and treated with or without MG132 at 20 μM for 12 h. I and J, Mst1/2 protein half-life assays in 293T cells after transfection with indicated siRNAs or constructs and then treatment with cycloheximide (CHX) at 100 μM for the indicated time. K, Mst1/2 protein half-life assays in WT or Smurf1 KO MEFs after treatment with CHX at 100 μM for the indicated time. Numerical data were presented as mean ± SD. n = 3. MEF, mouse embryonic fibroblast; Mst1/2, Mst1 and Mst2; Smurf1, Smad ubiquitin regulatory factor 1.
Smurf1 interacts with Mst1/2 and induces their K27- and K29-linked polyubiquitination
To investigate the potential interactions between Mst1/2 and Smurf1, we performed coimmunoprecipitation in 293T cells transiently expressing FLAG-Mst1/2. The immunocomplexes precipitated by an anti-FLAG antibody but not its immunoglobulin G control contained a large amount of Smurf1 in addition to FLAG-Mst1/2 as expected (Fig. 3A). Likewise, immunofluorescence staining in 293T cells transfected with FLAG-Mst1/2 and Myc-Smurf1 consistently indicated that FLAG- and Myc-derived immunosignals were detected predominantly in the cytosol, and the apparently overlapping signals were readily observed in cytosol either (Fig. 3B). To further determine the interaction between Smurf1 and Mst1/2, we performed bioinformatics analysis by using UbiBrowser database (http://ubibrowser.ncpsb.org.cn). Smurf1 as well as Smurf2, a closely related homolog of Smurf1 (19), was most likely to interact with and ubiquitinated Mst1/2 (Fig. 3C). To explore whether Smurf2 interacts with Mst1/2 either, we performed coimmunoprecipitation in 293T cells transfected with Myc-Smurf2 and FLAG-Mst1/2. Immunocomplexes precipitated by an anti-FLAG antibody contained no Myc-Smurf2 in addition to FLAG-Mst1/2 as expected (Fig. S2, A and B). Moreover, Western blotting analyses in 293T cells transfected with FLAG-Mst1/2, and the increasing doses of Myc-Smurf2 consistently revealed that Smurf2 had no effect on Mst1/2 protein levels (Fig. S2, C and D). Thus, Smurf1 instead of Smurf2 interacts with and destabilizes Mst1/2.
Figure 3.
Smurf1 physically interacts with and ubiquitinates Mst1/2.A, coimmunoprecipitation experiments with a FLAG antibody or its immunoglobulin G (IgG) control in 293T cells transfected with FLAG-Mst1/2 for 24 h, followed by Western blotting analyses using a FLAG or Smurf1 antibody. B, immunofluorescence staining in 293T cells after transfection with Myc-Smurf1 and FLAG-Mst1/2 for 24 h. Scale bar represents 10 μm. C, prediction of E3 ubiquitin ligase-substrate by using UbiBrowser database. The predicted interactors of Mst1/2 are arranged clockwise in descending order according to the confidence score. D and E, coimmunoprecipitation experiments by using a FLAG antibody in 293T cells after transfection with FLAG-Mst1/2, Myc-Smurf1 variants, and HA-ubiquitin for 24 h. F, coimmunoprecipitation experiments by using Mst1/2 antibodies in WT or Smurf1 KO MEFs after treatment with MG132 at 20 μM for 12 h. G, in vitro ubiquitination reactions followed by Western blotting analyses. Control or FLAG-Smurf1-bound agarose was incubated with the reactive mixtures containing the purified His-Mst1/2, E1, E2, and HA-ubiquitin at 37 °C for 2 h. H and I, coimmunoprecipitation experiments in 293T cells after transfection with FLAG-Mst1/2, Myc-Smurf1, and different HA-ubiquitin variants for 24 h. HA, hemagglutinin; MEF, mouse embryonic fibroblast; Mst1/2, Mst1 and Mst2; Smurf1, Smad ubiquitin regulatory factor 1.
We next examined whether Smurf1 directly ubiquitinated Mst1/2. Coimmunoprecipitation analyses indicated that Myc-Smurf1 but not its inactive mutant (CA) significantly promoted Mst1/2 polyubiquitination in 293T cells transiently expressing Myc-Smurf1, FLAG-Mst1/2, and hemagglutinin (HA)-Ub (Fig. 3D). However, knockdown of Smurf1 or ablation of Smurf1 significantly reduced the ubiquitination levels of Mst1/2 in 293T cells or MEFs, respectively (Fig. 3, E and F). To validate that Smurf1 directly ubiquitinates Mst1/2, we performed in vitro ubiquitination reaction followed by Western blotting analyses. The apparent polyubiquitination of Mst1/2 was observed in the reactive mixture with FLAG-Smurf1 but not without FLAG-Smurf1 (Fig. 3G). To further determine the type of the polyubiquitination chain linked to Mst1/2, we generated a series of Ub mutants including K0, K6, K11, K27, K29, K33, K48, and K63. Each of them could only express a single linkage type of Ub and where K0 means that all lysine residues have been changed to arginine (R) and K6 means that all lysine residues except K6 have been changed to R (25, 26). We then performed coimmunoprecipitation assays in 293T cells transfected with FLAG-Mst1/2, Myc-Smurf1, and HA-Ub variants. Smurf1 robustly increased K27- and K29 chain–linked polyubiquitination of Mst1/2 (Fig. 3H), whereas K27R and K29R mutants largely diminished the Smurf1-induced polyubiquitination of Mst1/2 (Fig. 3I). Thus, Smurf1 induces the polyubiquitination of Mst1/2 via the atypical K27- and K29-linked Ub chains.
Smurf1 polyubiquitinates Mst1/2 on K285/K282 to destabilize them
To determine the potential lysine residues that are essential for Smurf1-induced ubiquitination and degradation of Mst1/2, mass spectrometry analysis (PhosphoSitePlus, https://www.phosphosite.org) was performed and showed that there were 18 and 15 potential Ub acceptor sites on Mst1 and Mst2, respectively (Fig. S3, A and B). We then constructed Mst1/2 deubiquitinated variants harboring mutations at the consensus lysine residues (Lys to Arg) individually and evaluated their capacity to mediate Smurf1-induced ubiquitination in 293T cells. Both K285R of Mst1 and K282R of Mst2 were apparently resistant to Smurf1-mediated ubiquitination (Fig. 4, A and B), whereas overexpression of Smurf1 but not its inactive mutant (CA) significantly negated the protein levels of WT Mst1/2 but not their K285R/K282R mutants (Fig. 4, C and D). Moreover, CHX chase assays demonstrated that K285R/K282R mutants of Mst1/2 had a significantly longer half-life than WT Mst1/2: the mutants increased the t1/2 values of Mst1/2 by approximately 100% and 170% (2 and 3.6 h to 4 and 9.7 h), respectively (Fig. 4, E and F). Consistently, qPCR analyses indicated that overexpression of K285R/K282R mutants of Mst1/2 more robustly negated the mRNA levels of Ctgf and Cyr61 than WT Mst1/2, respectively (Fig. 4, G and H). Whereas analyses of subcellular distribution revealed that K285R/K282R mutants more significantly increased and decreased the cytosolic and nuclear YAP/TAZ protein levels than WT Mst1/2, respectively (Fig. 4, I and J). Thus, K285/K282 residues of Mst1/2 are critical Ub acceptor sites for Smurf1-mediating ubiquitination and degradation of Mst1/2 in the regulation of Hippo signaling.
Figure 4.
Smurf1 polyubiquitinates Mst1/2 on K285/K282 to destabilize them.A and B, coimmunoprecipitation experiments in 293T cells after transfection with Myc-Smurf1, HA-ubiquitin, and FLAG-Mst1/2 variants for 24 h. C and D, Western blotting analyses in 293T cells after transfection with indicated plasmids for 24 h. E and F, protein half-life assays in 293T cells after transfection with indicated constructs for 24 h and treatment with cycloheximide (CHX) at 100 μM for the indicated times. G and H, qPCR analyses for mRNA levels of Ctgf and Cyr61 in 293T cells after transfection with vector, Mst1/2, or their variants for 24 h. I and J, Western blotting analyses for cytosolic and nuclear YAP/TAZ protein levels in 293T cells transfected with the indicated constructs for 24 h. Numerical data were expressed as mean ± SD. n = 3. One-way ANOVA and Tukey–Kramer multiple comparisons test. ∗p < 0.05 and ∗∗p < 0.01. Ctgf, connective tissue; growth factor; Cyr61, cysteine-rich angiogenic inducer 61; HA, hemagglutinin; Mst1/2, Mst1 and Mst2; qPCR, quantitative PCR; Smurf1, Smad ubiquitin regulatory factor 1; TAZ, transcriptional coactivator with PDZ-binding motif; YAP, Yes-associated protein.
Ubiquitination of Mst1/2 by Smurf1 attenuates their tumor-suppressor functions
Mst1/2 loss of functions are closely related to the tumorigenesis of HCC (7, 8, 27), and Smurf1 has been reported as a tumor promoter in diverse cancers, such as pancreatic cancer, gastric cancer, and lung cancer (20). To determine the potential relationship between Smurf1 and tumorigenesis of HCC, we performed the bioinformatics and statistical analyses of cancer-related gene expression from The Human Protein Atlas database and found that Smurf1 was highly expressed in the HCC (Fig. 5A). We compared the protein levels of Smurf1 and Mst1/2 in human normal hepatocyte L02 cells and human HCC HepG2 cells and found that Smurf1 and Mst1/2 protein levels were significantly higher and lower in HepG2 cells than in L02 cells, respectively (Fig. 5B). We next examine the ubiquitination levels of Mst1/2 in L02 and HepG2 cells after MG132 treatments and found that HepG2 cells underwent more robustly ubiquitination of Mst1/2 than L02 cells (Fig. 5C). Likewise, Western blotting analyses indicated that knockdown or overexpression of Smurf1 significantly increased or decreased the Mst1/2 protein levels in HepG2 cells, respectively (Fig. 5, D and E), whereas overexpression of WT Smurf1 but not its inactive mutant (CA) robustly enhanced the ubiquitination of Mst1/2 in HepG2 cells (Fig. 5F). Thus, Smurf1 induces the polyubiquitination and degradation of Mst1/2 in HepG2 HCC cells either.
Figure 5.
Ubiquitination of Mst1/2 by Smurf1 attenuates their tumor-suppressor functions.A, Smurf1 mRNA levels in normal livers and hepatocellular carcinomas, data were extracted from The Human Protein Atlas database. B, Western blotting analyses in L02 and HepG2 cells. C, coimmunoprecipitation experiments in L02 and HepG2 cells after treatment with MG132 at 20 μM for 12 h. D and E, Western blotting analyses in HepG2 cells after transfection with Smurf1-siRNA or the increasing dosages of FLAG-Smurf1 for 24 h. F, coimmunoprecipitation experiments in HepG2 cells transfected with the indicated constructs for 24 h. G–I, flow cytometry, CCK-8, and colony formation assays in Smurf1-koncked down or overexpression HepG2 cells. J, HepG2 cell xenografts expressing Smurf1-shRNA or Smurf1 at 38 days postinoculation in nude mice. K and L, Western blotting analyses and immunohistochemistry analyses with semiquantification of Smurf1 and Mst1/2 in HepG2 cell xenografts expressing Smurf1-shRNA or Smurf1 at 38 days postinoculation in nude mice. Scale bar represents 50 μm. Numerical data were presented as mean ± SD. n = 3. One-way ANOVA and Tukey–Kramer multiple comparisons or Student’s t test. ∗p < 0.05 and ∗∗p < 0.01. CCK-8, Cell Counting Kit-8; Mst1/2, Mst1 and Mst2; Smurf1, Smad ubiquitin regulatory factor 1.
To determine whether Smurf1 has a critical role in the regulation of proliferation, apoptosis, and tumorigenesis of HCCs, we generated stably Smurf1- or Smurf1-shRNA-overexpressing HepG2 cells and performed Cell Counting Kit-8 (Yeasen), flow cytometry, and colony formation assays. Smurf1 knockdown or overexpression significantly increased or decreased the cell apoptosis, respectively, whereas Smurf1 knockdown or overexpression robustly decreased or increased the proliferation and colony formation of HepG2 cells, respectively (Fig. 5, G–I). To explore the in vivo effect of Smurf1 in the tumorigenesis of HCC, we generated Smurf1-shRNA- or Smurf1-expressing HCC xenografts in nude mice and assessed their volumes on day 14 postinoculation. Knockdown or overexpression of Smurf1 significantly decreased or increased the volumes of HCC xenografts, respectively (Fig. 5J). However, knockdown or overexpression of Smurf1 in xenografts significantly induced or negated Mst1/2 protein levels in HCC xenografts, respectively, as demonstrated by Western blotting and immunohistochemistry analyses (Fig. 5, K and L). Thus, Smurf1 polyubiquitinates and degrades Mst1/2 to attenuate their tumor-suppressor functions.
Ubiquitination on K285/K282 of Mst1/2 by Smurf1 attenuates their tumor-suppressor functions
In order to investigate further the role of ubiquitination on K285/K282 of Mst1/2 in the regulation of tumorigenesis of HCC, we generated HepG2 cells stably expressing WT Mst1/2 or deubiquitinated variants of Mst1/2, Mst1/2(K285R/K282R). Overexpression of K285R/K282R variants of Mst1/2 consistently increased and decreased the cytosolic and nuclear YAP/TAZ levels, as compared with overexpression of WT Mst1/2, respectively (Fig. 6, A and B). Likewise, Cell Counting Kit-8 and colony formation assays revealed that overexpression of K285R/K282R variants of Mst1/2 significantly decreased the cell proliferation at 3 days postculture and colony formation at 14 days postculture, as compared with overexpression of WT Mst1/2, respectively (Fig. 6, C–F). Whereas overexpression of K285R/K282R variants of Mst1/2 significantly increased the cell apoptosis at 3 days postculture, as compared with overexpression of WT Mst1/2, respectively (Fig. 6, G and H). Thus, deubiquitination of Mst1/2 on K285/K282 stabilizes Mst1/2 to enhance their tumor-suppressor functions.
Figure 6.
Deubiquitination of Mst1/2 on K285/K282 enhances their tumor-suppressor functions in vitro.A and B, Western blotting analyses for cytosolic and nuclear YAP/TAZ protein levels in HepG2 cells stably expressing vector (−), WT Mst1/2, or their K285R/K282R variants. C–H, CCK-8, colony formation, and flow cytometry assays in HepG2 cells stably expressing vector, WT Mst1/2, or their K285R/K282R variants and cultured for the indicated times. Numerical data were presented as mean ± SD. n = 3. One-way ANOVA and Tukey–Kramer multiple comparisons test. ∗p < 0.05 and ∗∗p < 0.01. CCK-8, Cell Counting Kit-8; Mst1/2, Mst1 and Mst2; TAZ, transcriptional coactivator with PDZ-binding motif; YAP, Yes-associated protein.
To confirm further the role of ubiquitination on K285/K282 of Mst1/2 in the regulation of their tumor-suppressor functions in vivo, we generated stably vector-, Mst1/2(WT)-, or Mst1/2(K285R/K282R)-expressing HepG2 cell xenografts in nude mice. The volumes of HepG2 cell xenografts expressing vector, WT, or K285R/K282R were time-dependently increased within 38-day postinoculation (Fig. 7, A and B). From day 10, the volumes of WT-expressing xenografts were significantly smaller than those of vector-expressing xenografts, whereas the volumes of K285R/K282R-expressing xenografts were much smaller than those of WT-expressing xenografts (Fig. 7, A and B). On day 38, the volumes from WT-expressing xenografts were approximately 60% of those from vector-expressing xenografts; however, the volumes of K285R/K282R-expressing xenografts were 50% and 40% of those from WT-expressing xenografts, respectively (Fig. 7, C–F). Moreover, analyses of mRNA levels revealed that WT-expressing xenografts significantly decreased the Ctgf and Cyr61 mRNA levels, as compared with vector-expressing xenografts, and K285R/K282R-expressing xenografts further decreased these mRNA levels, as compared with WT-expressing xenografts (Fig. 7, G and H). The cell proliferation and apoptosis in vector-, WT-, and K285R/K282R-expressing xenografts at 38 day postinoculation were assessed by Ki-67 and TUNEL staining analyses. The TUNEL-positive cells in xenografts expressing Mst1/2(WT) were significantly more than those in xenografts expressing vector, whereas the TUNEL-positive cells in xenografts expressing Mst1/2(K285R/K282R) were much more than those in xenografts expressing Mst1/2(WT) (Fig. 7I). In contrast, Ki-67-positive cells in xenografts expressing Mst1/2(WT) were significantly less than those in xenografts expressing vector, and the Ki-67-positive cells in xenografts expressing Mst1/2(K285R/K282R) were much less than those in xenografts expressing Mst1/2(WT) (Fig. 7J). Taken together, Smurf1-mediated ubiquitination on K285/K282 of Mst1/2 attenuates their tumor-suppressor functions in HepG2 cell xenografts.
Figure 7.
Deubiquitination of Mst1/2 on K285/K282 enhances their tumor-suppressor functions in vivo. Nude mice bearing HepG2 cell xenografts stably expressing vector, Mst1/2(WT), or Mst1/2(K285R/K282R) were used for the following assays. A and B, xenograft volumes were measured every 2 days. C–F, 38 days after inoculation, the mice were sacrificed, and the xenografts were excised and measured. G and H, qPCR analyses for mRNA levels of Ctgf and Cyr61 in xenografts at 38 days postinoculation. I and J, TUNEL and immunohistochemistry staining with their semiquantification analyses in the paraffin-embedded xenografts at 38 days postinoculation. Scale bar represents 50 μm. Numerical data were presented as mean ± SD. n = 3. One-way ANOVA and Tukey–Kramer multiple comparisons test. ∗p < 0.05 and ∗∗p < 0.01. Ctgf, connective tissue; growth factor; Cyr61, cysteine-rich angiogenic inducer 61; Mst1/2, Mst1 and Mst2; qPCR, quantitative PCR.
Discussion
By biochemical, genetic, and xenograft approaches, the present study, to the best of our knowledge, is the first work uncovering that Smurf1-mediated ubiquitination on K285/K282 of Mst1/2 destabilizes Mst1/2 to attenuate their tumor-suppressor functions. We have previously demonstrated that Smurf1 ubiquitinates and destabilizes Lats1/2 to induce the transcriptional output of Hippo signaling (21). Thus, Smurf1 targets multiple acceptor sites of Hippo signaling pathway to attenuate the tumor-suppressor functions (Fig. 8).
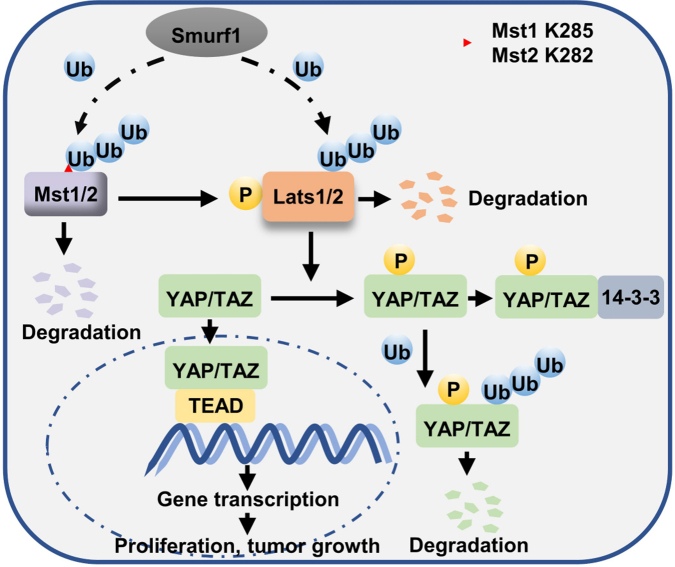
Figure 8.
The proposed model for Smurf1 targeting multiple acceptor sites (Mst1/2 and Lats1/2) of Hippo signaling pathway to attenuate the tumor-suppressor functions. Mst1/2, Mst1 and Mst2.
Our present study is consistent with previous studies that E3 Ub ligases, carboxy terminus of Hsp70-interacting protein and tumor necrosis factor receptor–associated factor 6, negatively regulate Mst1 stability, and that SCFβTrCP negatively regulates Mst2 stability through ubiquitination degradation in mammalian cells (28, 29, 30). Our finding further reveals the critical roles of Smurf1 in the regulation of ubiquitination-dependent degradation of Mst1/2 and thus expands the list of E3 Ub ligases that control the stability of Mst1/2. Mst1/2 function as the core players to Hippo signaling transduction (8, 27), and the present study demonstrates the Smurf1-mediated Mst1/2 destabilization and thus reveals a novel mechanism underlying the regulation of Hippo pathway and Hippo signaling–associated diseases.
Smurf1 is a HECT-type Ub ligase responsible for the stability of a variety of substrate proteins and thus plays an essential role in diverse biological processes, including bone homeostasis, cell cycle, apoptosis, senescence, and tumor progression (19, 31). Consistent with the present findings that Smurf1 ubiquitinates Mst1/2 to regulate Hippo signaling pathway, knockdown of dSmurf1 affects Wts protein turnover and regulates Hippo signaling transduction in Drosophila and S2 cells (22), and our previous findings have indicated that Smurf1 is required for ubiquitination of Lats1 in mammalian cells (21). In addition, the present study reveals that Smurf1 ubiquitinates Mst1/2 to promote their degradation. Thus, the overall effect of Smurf1 on the transcriptional output of Hippo signaling observed in the present study is resulted from its ubiquitination of both Mst1/2 and Lats1/2. This notion is supported by the findings that overexpression or knockdown/KO of Smurf1 produces more profound effect on the p-Lats1/2 than on the p-Mst1/2. Though we cannot readily observe the mere effect of Smurf1-mediated Mst1/2 degradation on the transcriptional output of Hippo signaling, overexpression of K285R/K282R deubiquitinated mutants of Mst1/2 does produce profound effects on the transcriptional output of Hippo signaling as well as tumor-suppressor functions. These findings thus support the notion that Smurf1-mediated destabilization of Mst1/2 contributes to the Smurf1-induced transcriptional output of Hippo signaling.
Smurf1 consists of a HECT domain at the C-terminal region, two WW domains, and a C2 domain at the N-terminal region, and HECT domain of Smurf1 is predominantly responsible for its E3 ligase activity (20, 32). The conserved catalytic active site Cys699 in HECT domain plays a central role in the formation of an intermediate thioester bond with C terminus of Ub before ubiquitination of substrates (33, 34). Though we have not identified the exact domains of Smurf1 that are involved in the polyubiquitination of Mst1/2, the inactive form of Smurf1, Smurf1(C699A), completely abolishes the effect of Smurf1 on Mst1/2 stability and transcriptional output of Hippo signaling. This finding suggests that the HECT domain of Smurf1 is essential for polyubiquitinating Mst1/2. Literatures have identified the Smurf1-induced K29- and K33-linked polyubiquitination of UV radiation resistance–associated gene protein (25) and Smurf1-mediated K29-linked polyubiquitination of axin (35, 36). In the present study, we have identified that Smurf1 induces the polyubiquitination of Mst1/2 via the atypical K27- and K29-linked Ub chains. It is well established that Smurf1 is involved in the K29-linked polyubiquitination of many protein substrates; however, we for the first time reveal that Smurf1 is also involved in the K27-linked polyubiquitination of protein substrates, and this issue needs to be further investigated in the future.
High expression of Smurf1 in tumors is positively correlated with large tumor size, vascular invasion, lymph node and distant metastasis, advanced TNM stage, and decreased survival as well (20, 37, 38, 39, 40). The Human Protein Atlas database also implicates the oncogenic role of Smurf1 in HCC, consistent with our findings that Smurf1 ubiquitinates and destabilizes Mst1/2 to attenuate their tumor-suppressor functions in HepG2 cells and their xenografts as well and with a previous study indicating that expression of Smurf1 mRNA and protein is significantly higher in HCC tissues, and Smurf1-specific siRNA promotes the apoptosis whilst inhibits the proliferation of HepG2 cells (41). Importantly, K285/K282 are determinant residues responsible for Smurf1-mediated polyubiquitination of Mst1/2, even though K285/K282 mutations are not found in Catalogue of Somatic Mutations in Cancer (COSMIC) database. However, the literatures have also acknowledged the Smurf1's dual role in cancers with some proposing a tumor suppressor role in HCC (25, 42, 43), appearing to be at odd with our findings. The discrepancy could be explained by the facts that Smurf1 also ubiquitinates and degrades diverse substrates that play the oncogenic role in HCC, such as tribbles homolog 2 and UV radiation resistance–associated gene protein (25, 42).
Overall, the present study uncovers that the Smurf1-mediated Mst1/2 polyubiquitination and subsequent destabilization fine-tune the altitude of Hippo signaling and identifies that Smurf1 targets multiple acceptor sites to control Hippo signaling–associated tumorigenesis. Given the fact that Smurf1 has been shown the advantages to become a target in the cancer chemotherapies without potential side effects caused by targeting E3 ligases (40, 44), targeting the Smurf1 is a rational approach for the therapeutic interventions of Hippo signaling–associated tumors and diseases.
Experimental procedures
Cell culture and transfection
Human embryonic kidney 293T (HEK293T) (293T), L02, and HepG2 cells were purchased from American Type Culture Collection and cultured in Dulbecco's modified Eagle's medium (Gibco) supplemented with 10% fetal bovine serum (Gibco) and 1% penicillin and streptomycin (Sigma). Primary MEFs were isolated from mouse embryos at 13.5 days of gestation and maintained in modified Eagle's medium (Gibco) containing 10% fetal bovine serum, 1% l-glutamine (Gibco), 1% nonessential amino acids (Invitrogen), and 1% sodium pyruvate (Invitrogen). All cells were grown at 37 °C in a humidified 5% CO2 incubator and routinely tested negative for mycoplasma contamination. Transient transfection was performed by using Lipofectamine 2000 (Invitrogen) for 293T cells and Hieff Trans Liposomal Transfection Reagent (Yeasen Biotechnology) for HepG2 cells according to their manufacturer’s protocols.
Plasmids and lentiviral infection
FLAG-Mst1, FLAG-Mst2, and Tead4-luciferase constructs were gifts from Dr Bin Zhao from Zhejiang University Life Science Institute (45). FLAG-Smurf1 and Myc-Smurf2 were purchased from Miaolingbio. Constructions expressing the interest genes were cloned by using specific primers. For example, to generate Myc-Smurf1, full-length Smurf1 was amplified by PCR using specific primers and inserted into the BamHI and KpnI sites of pXJ40-Myc expression plasmids (Stratagene). Point mutations were introduced by using a KOD-plus-mutagenesis kit (Toyobo) according to the manufacturer’s protocol. A series of HA-tagged Ub constructs including WT, K0, K6, K11, K27, K29, K33, K48, K63, K27R, and K29R were purchased from Youbio. siRNA targeting Smurf1 was supplied by Sangon. The lentiviral vector pSicoR-GFP (Addgene, catalog no.: 11579) was used to knock down the Smurf1 expression. Targeting sequences for shRNA were used as follows: shSmurf1, 5ʹ-CTGGAGGTTTATGAGAGGAAT-3′. The lentiviral vector pCDH-CMV-MCS-3FLAG-Puro (Miaolingbio) was used to produce viruses expressing Smurf1, Mst1/2, and their K285R/K282R mutants. All plasmids were confirmed by nucleic acid sequencing. The production of lentivirus was performed as described previously (46). Briefly, lentiviral vectors or constructs were transfected into 293T cells in combination with packaging plasmids PMDL, REV, and VSVG by using Lipofectamine 2000. The lentivirus supernatant was harvested at 48 h post-transfection and used to infect cells in the presence of 5 μg/ml polybrene (Beyotime). Selection was performed in the presence of 1 μg/ml puromycin at 48 h postinfection.
Antibodies and reagents
The following primary antibodies were used at the indicated concentrations according to the manufacturer’s instructions: anti-Mst1, anti-Mst2, anti-phospho-Mst1(Thr183)/Mst2(Thr180), anti-Lats1, anti-Lats2, anti-phospho-Lats1(Thr1079)/Lats1(Thr1041), anti-YAP, anti-phospho-YAP(S127), anti-TAZ, and anti-phospho-TAZ(S89) were purchased from Cell Signaling Technology. Anti-Ki67 was from Abcam, and anti-Smurf1, anti-Myc, anti-His, anti-Histone H3, anti-Ub, anti-α-Tubulin, and anti-rabbit/antimouse immunoglobulin G were from Huabio. Anti-GAPDH, anti-FLAG, and anti-HA were purchased from Affinity Biosciences. Anti-FLAG and Anti-Myc beads were purchased from Bimake. The second antibodies IRDye 680 and 800 were purchased from LI-COR Bioscience. Anti-mouse Alexa Fluor 488 and anti-rabbit Alexa Fluor 555 were purchased from Invitrogen. Protein A/G PLUS Agarose were purchased from Santa Cruz Biotechnology. MG132, CHX, and 4′,6-diamidino-2-phenylindole were purchased from Selleck.
Dual-luciferase reporter assays
Dual-luciferase assays were performed as described previously (47). Briefly, HEK293T cells were seeded in 24-well plates overnight to be 60∼80% confluent, and then 0.4 μg testing-constructs, 0.35 μg Tead4-luciferase construct, and 0.05 μg Renilla luciferase construct were transfected into HEK293T cells by using Lipofectamine 2000. After 48 h, cells were harvested and luciferase activities were determined by using the dual-luciferase reporter assay kit (Promega) according to the manufacturer's instruction. Tead4-luciferase reporter activities were normalized to Renilla luciferase activities, respectively.
RNA isolation and qRT–PCR assays
RNA isolation and qPCR assays were performed as described previously (48). Briefly, total RNA was isolated using a TRIzol reagent (Takara Biotechnology) according to the manufacturer’s instruction. For qPCR assays, 2 μg total RNA of the sample was used for reversely transcribed into complementary DNA by using HiScript II QRT reagent (Vazyme). The relative expression of each mRNA was calculated by 2−ΔΔCt method. Primers used in this study were as follows: Ctgf forward, 5′-CGACTGGAAGACACGTTTGG-3′ and Ctgf reverse, 5′-AGGAGGCGTTGTCATTGGTA-3′; Cyr61 forward, 5′-GACTGTGAAGATGCGGTTCC-3′ and Cyr61 reverse, 5′-CTGTAGAAGGGAAACGCTGC-3′; Smurf1 forward, 5′-AGATCCGTCTGACAGTGTTATGT-3′ and Smurf1 reverse, 5′-CCCATCCACGACAATCTTTGC-3′; Mst1 forward, 5′-TGGATTCTGGCACGATGGTTC-3′ and Mst1 reverse, 5′-GCATGGTCTCATCCCTTCTTTT-3′; Mst2 forward, 5′-CGATGTTGGAATCCGACTTGG-3′ and Mst2 reverse, 5′-GTCTTTGTACTTGTGGTGAGGTT-3′; β-actin forward, 5′-GACGATGGAGGGGCCGGACTCGTC-3′ and β-actin reverse, 5′-CAAAGACCTGTACGCCAACACAGT-3′.
Nuclear and cytoplasmic protein separation, coimmunoprecipitation, and Western blotting
Cells were washed twice by PBS (pH 7.4) and lysed in radioimmunoprecipitation lysis buffer (Beyotime) containing protease and phosphatase inhibitors (Bimake) on ice for 0.5 h. Supernatant containing total cellular proteins was collected after centrifugation. Nuclear and cytoplasmic protein extraction was performed by using nuclear and cytoplasmic protein extraction kit (Beyotime) according to the manufacturer’s instruction. Protein concentration was measured by BCA protein assay Kit (Beyotime). For coimmunoprecipitation assays, immunoprecipitates were pulled down by incubating with anti-FLAG/Myc-beads or primary antibody (with protein A/G PLUS Agarose beads) overnight at 4 °C. Certain amounts of proteins were subjected to SDS-PAGE and analyzed using the indicated antibodies after separating. National Institutes of Health Image software (ImageJ, http://rsb.info.nih.gov/ij/) was used to quantify the immunoreactive bands, and target protein-derived immunoreactive signals were normalized to their respective internal standards including GAPDH, α-tubulin, and Histone H3. Phosphoprotein-derived immunoreactive signals were normalized to their respective total proteins.
Protein half-life assays
HEK293T cells were seeded in 12-well plates overnight at 60∼80% confluency and transfected with indicated plasmids. MEFs from WT or Smurf1 KO mice were not transfected. About 36 h after transfection, cells were treated with 100 μM CHX for 0, 2, 4, 6, 8, and 10 h, respectively. Cells were then harvested for Western blotting analyses.
Immunofluorescence staining, TUNEL staining, and confocal microscopy
After transfection, cells cultured on coverslips were washed twice by PBS and fixed in 4% paraformaldehyde for 10 min and permeabilized with 0.3% Triton X-100 for 10 min. Then, cells were washed by PBS three times and incubated with 3% BSA for 30 min at room temperature, followed by staining with primary antibody overnight at 4 °C. Then, cells were washed by PBS three times and incubated with secondary antibodies conjugated with Alexa Fluor 488/555 for 1 h at room temperature. After washing with PBS three times, 4′,6-diamidino-2-phenylindole was used to counterstain the nuclei. TUNEL staining was performed with the TUNEL BrightRed apoptosis detection kit (Vazyme) using paraffin-embedded tumor tissue sections according to the manufacturer’s instruction. All images were captured by the confocal microscope Olympus FV3000 and analyzed by the FV31S-SW Viewer software.
In vitro ubiquitination assays
FLAG-Smurf1-bound or control agarose was immunoprecipitated from HEK293T cells after transfection with FLAG-Smurf1 or vector. The in vitro ubiquitination reactions were performed by incubating FLAG-Smurf1-bound or control agarose in 30 μl ubiquitination reaction buffer containing 0.5 μg purified recombinant His-Mst1 (MedChemExpress) or 0.5 μg His-Mst2 (Solarbio), 20 μg HA-Ub (R&D Systems), 1 μg Ub-activating enzyme UBE1 (E1; R&D), 1 μg E2-conjugating enzyme UbcH5c (E2; Sangon), and 2 mM ATP at 37 °C for 2 h. SDS-PAGE protein loading buffer (Beyotime) was used to terminate the reaction. The ubiquitinated Mst1/2 proteins were detected by Western blotting analyses using an anti-Ub antibody.
Cell proliferation assays
Cells were seeded in 96-well plates with 1000 cells per well, and the cell proliferation assay was performed with the Cell Counting Kit-8 every 24 h according to the manufacturer’s instruction.
Colony formation assays
Single-cell suspension including 800 cells per well was seeded in 6-well plates. Every 2 days, the cell medium was replaced by fresh growth medium. About 14 days later, cell colonies were fixed with 4% paraformaldehyde for 10 min and stained with 1% crystal violet staining solution for 5 min. Then, colonies were photographed and counted using ImageJ software.
Apoptosis assays
Apoptosis assays were performed using single-cell suspension. Cells were stained with an Annexin V-Alexa Fluor 647/PI apoptosis assay kit (Yeasen) according to the manufacturer’s instruction. Apoptosis rate was analyzed by FACS Calibur flow cytometer (BD Biosciences).
Mouse strains and xenografts
Smurf1+/− mouse strain on the C57BL/6N background was purchased from Cyagen Biosciences (serial number: KOCMP-75788-Smurf1-B6N-VA) and generated by using the CRISPR–Cas9 technique. Four-week-old male BALB/c-Nude mice (strain no.: D000521) were purchased from GemPharmatech and randomly divided into the indicated groups (eight mice/group). HepG2 cells were harvested and suspended at a density of 2.5 × 107 cells/ml with PBS and then subcutaneously inoculated into the left armpit of each mouse with 0.2 ml cell suspension. After tumor formation, tumor volume (V) of each mouse was monitored every 2 days by measuring the length (a) and width (b) with calipers and calculated with the formula “V = half a b2” (49). After 38 days, the mice were sacrificed, and the tumors were excised and measured. All mice were housed and bred in a specific pathogen-free room at Zhejiang University Animal Care Facility according to the institutional guidelines for laboratory animals. And all the procedures (protocol no.: 20180226-003) were approved by the Institutional Animal Care and Use Committee of Zhejiang University.
Bioinformatics and statistical analyses of public databases
UbiBrowser database (http://ubibrowser.bio-it.cn/ubibrowser/) predicts the human Ub ligase (E3)–substrate interaction. The Human Protein Atlas (https://www.proteinatlas.org/) provides cancer-related gene expression information by using data from The Cancer Genome Atlas and Genotype-Tissue Expression.
Statistical analysis
Numerical data were expressed as mean ± SD, and statistical analyses were performed using GraphPad Prism 8.4 software (GraphPad Software, Inc). Statistical significance was determined by one-way ANOVA and Tukey–Kramer multiple comparisons or Student’s t test. Statistical significance was assessed at levels of p <0.05 and p <0.01.
Data availability
All data generated or analyzed during this study are included in this article or are available from the corresponding author upon reasonable request.
Supporting information
This article contains supporting information.
Conflict of interest
The authors declare that they have no conflicts of interest with the contents of this article.
Acknowledgments
We thank Dr Bin Zhao from Zhejiang University for their plasmids.
Author contributions
Y. X., M. Q., and C. X. formal analysis; Y. X., M. Q., Y. H., Q. H., T. S., J. L., D. T., and H. B. investigation; Y. X., M. Q., and C. X. writing–original draft; X. H., T. S., and M. Q. B. writing–review & editing; L.-H. Z. and X. W. supervision.
Funding and additional information
This work was supported by the National Natural Science Foundation of China (grant nos. 32170841 to X. W., 31871395 to X. W., and 82100934 to X. H.) and Zhejiang Provincial Natural Science Foundation of China (grant no.: LQ22H070001 to X. H.).
Reviewed by members of the JBC Editorial Board. Edited by Donita C. Brady
Contributor Information
Ling-Hui Zeng, Email: zenglh@zucc.edu.cn.
Ximei Wu, Email: xiwu@zju.edu.cn.
Supporting information
References
- 1.Zeybek N.D., Baysal E., Bozdemir O., Buber E. Hippo signaling: a stress response pathway in stem cells. Curr. Stem Cell Res. Ther. 2021;16:824–839. doi: 10.2174/1574888X16666210712100002. [DOI] [PubMed] [Google Scholar]
- 2.Fallahi E., O'Driscoll N.A., Matallanas D. The MST/Hippo pathway and cell death: a non-canonical affair. Genes (Basel) 2016;7:28. doi: 10.3390/genes7060028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Meng Z., Moroishi T., Guan K.L. Mechanisms of Hippo pathway regulation. Genes Dev. 2016;30:1–17. doi: 10.1101/gad.274027.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Galan J.A., Avruch J. MST1/MST2 protein kinases: regulation and physiologic roles. Biochemistry. 2016;55:5507–5519. doi: 10.1021/acs.biochem.6b00763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Du X., Yu A., Tao W. The non-canonical Hippo/Mst pathway in lymphocyte development and functions. Acta Biochim. Biophys. Sin. 2015;47:60–64. doi: 10.1093/abbs/gmu112. [DOI] [PubMed] [Google Scholar]
- 6.Ma S., Meng Z., Chen R., Guan K.L. The hippo pathway: biology and pathophysiology. Annu. Rev. Biochem. 2019;88:577–604. doi: 10.1146/annurev-biochem-013118-111829. [DOI] [PubMed] [Google Scholar]
- 7.Lu L., Li Y., Kim S.M., Bossuyt W., Liu P., Qiu Q., et al. Hippo signaling is a potent in vivo growth and tumor suppressor pathway in the mammalian liver. Proc. Natl. Acad. Sci. U. S. A. 2010;107:1437–1442. doi: 10.1073/pnas.0911427107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Zhou D., Conrad C., Xia F., Park J.S., Payer B., Yin Y., et al. Mst1 and Mst2 maintain hepatocyte quiescence and suppress hepatocellular carcinoma development through inactivation of the Yap1 oncogene. Cancer Cell. 2009;16:425–438. doi: 10.1016/j.ccr.2009.09.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Zinatizadeh M.R., Miri S.R., Zarandi P.K., Chalbatani G.M., Rapôso C., Mirzaei H.R., et al. The Hippo tumor suppressor pathway (YAP/TAZ/TEAD/MST/LATS) and EGFR-RAS-RAF-MEK in cancer metastasis. Genes Dis. 2021;8:48–60. doi: 10.1016/j.gendis.2019.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Li S., Zhao J., Shang D., Kass D.J., Zhao Y. Ubiquitination and deubiquitination emerge as players in idiopathic pulmonary fibrosis pathogenesis and treatment. JCI Insight. 2018;3 doi: 10.1172/jci.insight.120362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Popovic D., Vucic D., Dikic I. Ubiquitination in disease pathogenesis and treatment. Nat. Med. 2014;20:1242–1253. doi: 10.1038/nm.3739. [DOI] [PubMed] [Google Scholar]
- 12.Shaid S., Brandts C.H., Serve H., Dikic I. Ubiquitination and selective autophagy. Cell Death Differ. 2013;20:21–30. doi: 10.1038/cdd.2012.72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Akimov V., Barrio-Hernandez I., Hansen S.V.F., Hallenborg P., Pedersen A.K., Bekker-Jensen D.B., et al. UbiSite approach for comprehensive mapping of lysine and N-terminal ubiquitination sites. Nat. Struct. Mol. Biol. 2018;25:631–640. doi: 10.1038/s41594-018-0084-y. [DOI] [PubMed] [Google Scholar]
- 14.Buetow L., Huang D.T. Structural insights into the catalysis and regulation of E3 ubiquitin ligases. Nat. Rev. Mol. Cell Biol. 2016;17:626–642. doi: 10.1038/nrm.2016.91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Morreale F.E., Walden H. Types of ubiquitin ligases. Cell. 2016;165:248. doi: 10.1016/j.cell.2016.03.003. [DOI] [PubMed] [Google Scholar]
- 16.Sapkota G., Alarcon C., Spagnoli F.M., Brivanlou A.H., Massague J. Balancing BMP signaling through integrated inputs into the Smad1 linker. Mol. Cell. 2007;25:441–454. doi: 10.1016/j.molcel.2007.01.006. [DOI] [PubMed] [Google Scholar]
- 17.Koganti P., Levy-Cohen G., Blank M. Smurfs in protein homeostasis, signaling, and cancer. Front. Oncol. 2018;8:295. doi: 10.3389/fonc.2018.00295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Jiang M., Shi L., Yang C., Ge Y., Lin L., Fan H., et al. miR-1254 inhibits cell proliferation, migration, and invasion by down-regulating Smurf1 in gastric cancer. Cell Death Dis. 2019;10:32. doi: 10.1038/s41419-018-1262-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Fu L., Cui C.P., Zhang X., Zhang L. The functions and regulation of Smurfs in cancers. Semin. Cancer Biol. 2020;67:102–116. doi: 10.1016/j.semcancer.2019.12.023. [DOI] [PubMed] [Google Scholar]
- 20.Xia Q., Li Y., Han D., Dong L. SMURF1, a promoter of tumor cell progression? Cancer Gene Ther. 2021;28:551–565. doi: 10.1038/s41417-020-00255-8. [DOI] [PubMed] [Google Scholar]
- 21.Qu M., Gong Y., Jin Y., Gao R., He Q., Xu Y., et al. HSP90beta chaperoning SMURF1-mediated LATS proteasomal degradation in the regulation of bone formation. Cell. Signal. 2023;102 doi: 10.1016/j.cellsig.2022.110523. [DOI] [PubMed] [Google Scholar]
- 22.Cao L., Wang P., Gao Y., Lin X., Wang F., Wu S. Ubiquitin E3 ligase dSmurf is essential for Wts protein turnover and Hippo signaling. Biochem. Biophys. Res. Commun. 2014;454:167–171. doi: 10.1016/j.bbrc.2014.10.058. [DOI] [PubMed] [Google Scholar]
- 23.Wang H.R., Ogunjimi A.A., Zhang Y., Ozdamar B., Bose R., Wrana J.L. Degradation of RhoA by Smurf1 ubiquitin ligase. Methods Enzymol. 2006;406:437–447. doi: 10.1016/S0076-6879(06)06032-0. [DOI] [PubMed] [Google Scholar]
- 24.Lignitto L., Arcella A., Sepe M., Rinaldi L., Delle Donne R., Gallo A., et al. Proteolysis of MOB1 by the ubiquitin ligase praja2 attenuates Hippo signalling and supports glioblastoma growth. Nat. Commun. 2013;4:1822. doi: 10.1038/ncomms2791. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Feng X., Jia Y., Zhang Y., Ma F., Zhu Y., Hong X., et al. Ubiquitination of UVRAG by SMURF1 promotes autophagosome maturation and inhibits hepatocellular carcinoma growth. Autophagy. 2019;15:1130–1149. doi: 10.1080/15548627.2019.1570063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.van Wijk S.J., Fulda S., Dikic I., Heilemann M. Visualizing ubiquitination in mammalian cells. EMBO Rep. 2019;20 doi: 10.15252/embr.201846520. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Song H., Mak K.K., Topol L., Yun K., Hu J., Garrett L., et al. Mammalian Mst1 and Mst2 kinases play essential roles in organ size control and tumor suppression. Proc. Natl. Acad. Sci. U. S. A. 2010;107:1431–1436. doi: 10.1073/pnas.0911409107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Li J.A., Kuang T., Pu N., Fang Y., Han X., Zhang L., et al. TRAF6 regulates YAP signaling by promoting the ubiquitination and degradation of MST1 in pancreatic cancer. Clin. Exper. Med. 2019;19:211–218. doi: 10.1007/s10238-018-00543-6. [DOI] [PubMed] [Google Scholar]
- 29.Xiao L., Chen D., Hu P., Wu J., Liu W., Zhao Y., et al. The c-Abl-MST1 signaling pathway mediates oxidative stress-induced neuronal cell death. J. Neurosci. 2011;31:9611–9619. doi: 10.1523/JNEUROSCI.0035-11.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Fiore A., Rodrigues A.M., Ribeiro-Filho H.V., Manucci A.C., de Freitas Ribeiro P., Botelho M.C.S., et al. Extracellular matrix stiffness regulates degradation of MST2 via SCF (betaTrCP) Biochim. Biophys. Acta Gen. Subj. 2022;1866 doi: 10.1016/j.bbagen.2022.130238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Cao Y., Zhang L. A Smurf1 tale: function and regulation of a ubiquitin ligase in multiple cellular networks. Cell. Mol. Life Sci. 2013;70:2305–2317. doi: 10.1007/s00018-012-1170-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Sangadala S., Metpally R.P., Reddy B.V. Molecular interaction between Smurf1 WW2 domain and PPXY motifs of Smad1, Smad5, and Smad6--modeling and analysis. J. Biomol. Struct. Dyn. 2007;25:11–23. [PubMed] [Google Scholar]
- 33.Cheng P.L., Lu H., Shelly M., Gao H., Poo M.M. Phosphorylation of E3 ligase Smurf1 switches its substrate preference in support of axon development. Neuron. 2011;69:231–243. doi: 10.1016/j.neuron.2010.12.021. [DOI] [PubMed] [Google Scholar]
- 34.Petrasek J., Erhartova D., Levine B. Protective effect of SMAD-specific E3 ubiquitin protein ligase 1 in alcoholic Steatohepatitis in mice. Hepatol. Commun. 2019;3:1450–1458. doi: 10.1002/hep4.1427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Fei C., He X., Xie S., Miao H., Zhou Z., Li L. Smurf1-mediated axin ubiquitination requires Smurf1 C2 domain and is cell cycle-dependent. J. Biol. Chem. 2014;289:14170–14177. doi: 10.1074/jbc.M113.536714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Fei C., Li Z., Li C., Chen Y., Chen Z., He X., et al. Smurf1-mediated Lys29-linked nonproteolytic polyubiquitination of axin negatively regulates Wnt/beta-catenin signaling. Mol. Cell. Biol. 2013;33:4095–4105. doi: 10.1128/MCB.00418-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Yu H., Li D., Zhou P., Li W. Smurf1-positive expression indicates favorable survival for resected non-small cell lung cancer patients. Int. J. Clin. Exp. Pathol. 2018;11:399–405. [PMC free article] [PubMed] [Google Scholar]
- 38.Loukopoulos P., Shibata T., Katoh H., Kokubu A., Sakamoto M., Yamazaki K., et al. Genome-wide array-based comparative genomic hybridization analysis of pancreatic adenocarcinoma: identification of genetic indicators that predict patient outcome. Cancer Sci. 2007;98:392–400. doi: 10.1111/j.1349-7006.2007.00395.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Ke M., Mo L., Li W., Zhang X., Li F., Yu H. Ubiquitin ligase SMURF1 functions as a prognostic marker and promotes growth and metastasis of clear cell renal cell carcinoma. FEBS Open Bio. 2017;7:577–586. doi: 10.1002/2211-5463.12204. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 40.Guo J., Xu G., Mao C., Wei R. Low expression of Smurf1 enhances the chemosensitivity of human colorectal cancer to gemcitabine and cisplatin in patient-derived xenograft models. Transl. Oncol. 2020;13 doi: 10.1016/j.tranon.2020.100804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Wang X., Zhang H.J., Hu Y.S., Zong Z.J., Li Y.H., Li C.Y. [The role of Smad ubiquitination regulatory factor 1 in hepatocellular carcinoma] Zhonghua Yi Xue Yi Chuan Xue Za Zhi. 2012;29:533–536. doi: 10.3760/cma.j.issn.1003-9406.2012.05.007. [DOI] [PubMed] [Google Scholar]
- 42.Wang J., Zhang Y., Weng W., Qiao Y., Ma L., Xiao W., et al. Impaired phosphorylation and ubiquitination by p70 S6 kinase (p70S6K) and Smad ubiquitination regulatory factor 1 (Smurf1) promote tribbles homolog 2 (TRIB2) stability and carcinogenic property in liver cancer. J. Biol. Chem. 2013;288:33667–33681. doi: 10.1074/jbc.M113.503292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Tang X., Chen X., Xu Y., Qiao Y., Zhang X., Wang Y., et al. CD166 positively regulates MCAM via inhibition to ubiquitin E3 ligases Smurf1 and betaTrCP through PI3K/AKT and c-Raf/MEK/ERK signaling in Bel-7402 hepatocellular carcinoma cells. Cell. Signal. 2015;27:1694–1702. doi: 10.1016/j.cellsig.2015.05.006. [DOI] [PubMed] [Google Scholar]
- 44.Masumoto K., Kitagawa M. E3 ubiquitin ligases as molecular targets in human oral cancers. Curr. Cancer Drug Targets. 2016;16:130–135. doi: 10.2174/1568009616666151112122336. [DOI] [PubMed] [Google Scholar]
- 45.Meng F., Zhou R., Wu S., Zhang Q., Jin Q., Zhou Y., et al. Mst1 shuts off cytosolic antiviral defense through IRF3 phosphorylation. Genes Dev. 2016;30:1086–1100. doi: 10.1101/gad.277533.116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Shi W., Xu C., Gong Y., Wang J., Ren Q., Yan Z., et al. RhoA/rock activation represents a new mechanism for inactivating Wnt/beta-catenin signaling in the aging-associated bone loss. Cell Regen. 2021;10:8. doi: 10.1186/s13619-020-00071-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Tang C., Mei L., Pan L., Xiong W., Zhu H., Ruan H., et al. Hedgehog signaling through GLI1 and GLI2 is required for epithelial-mesenchymal transition in human trophoblasts. Biochim. Biophys. Acta. 2015;1850:1438–1448. doi: 10.1016/j.bbagen.2015.04.005. [DOI] [PubMed] [Google Scholar]
- 48.Wu X., Iguchi T., Hirano J., Fujita I., Ueda H., Itoh N., et al. Upregulation of sodium-dependent vitamin C transporter 2 expression in adrenals increases norepinephrine production and aggravates hyperlipidemia in mice with streptozotocin-induced diabetes. Biochem. Pharmacol. 2007;74:1020–1028. doi: 10.1016/j.bcp.2007.05.024. [DOI] [PubMed] [Google Scholar]
- 49.Li R., Tang D., Zhang J., Wu J., Wang L., Dong J. The temozolomide derivative 2T-P400 inhibits glioma growth via administration route of intravenous injection. J. Neurooncol. 2014;116:25–30. doi: 10.1007/s11060-013-1255-7. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All data generated or analyzed during this study are included in this article or are available from the corresponding author upon reasonable request.