Abstract
Background
Fatty acid metabolism (FAM) is closely connected with tumorigenesis as well as disease progression and affects the efficacy of platinum-based drugs. Exploring biomarkers related to FAM in bladder cancer (BLCA) is essential to improve cancer prognosis.
Methods
High-throughput sequencing data from The Cancer Genome Atlas (TCGA) were bioinformatically resolved to identify molecular subtypes of fatty acid metabolic profiles in BLCA using coherent clustering analysis. Based on fatty acid metabolic profile, a prognostic model was created using COX and LASSO COX models. CIBERSORT, Estimation of STromal and Immune cells in MAlignant Tumours using Expression (ESTIMATE), MCP-Count, and single sample gene set enrichment analysis (ssGSEA) were used to assess the differences in tumor microenvironment (TME) among different molecular subtypes, prognostic groups. Kaplan-Meier (K-M) survival curve was plotted to assess patients’ prognosis. Receiver operating characteristic curve (ROC) and the clinical prognostic value of prognostic models was evaluated by the Nomogram.
Results
Three molecular subtypes (FAMC1, FAMC2, FAMC3) of fatty acid metabolic patterns were determined. FAMC1 showed significant prognostic advantage with immunoreactivity. Five key prognostic FAMGs were identified and RiskScore was developed. We found that patients with low RiskScore showed significantly better immune microenvironment status, survival and response to immunotherapy. Similarly, both Nomogram and RiskScore demonstrated excellent prognostic value.
Conclusions
In conclusion, our study showed that the RiskScore was closely related to the clinical traits of BLCA patients. The RiskScore may provide essential clinical guidance for predicting prognosis and treatment response in bladder cancer.
Keywords: TCGA, BLCA, Fatty acid metabolism, Prognosis, Immunotherapy, Nomogram
Abbreviations
- BLCA
Bladder cancer
- CNV
copy number variation
- DSS
disease-specific survival
- ESTIMATE
Estimation of STromal and Immune cells in MAlignant Tumour tissues using Expression data
- FAMGs
fatty acid metabolism genes
- GEO
Gene Expression Omnibus
- GO
Gene Ontology
- K-M
Kaplan-Meier
- KEGG
Kyoto Encyclopedia of Genes and Genomes
- LRFS
local recurrence-free survival
- MeFS
metastasis-free survival
- MSigDB
Molecular Signatures Database
- ssGSEA
single sample gene set enrichment analysis
- TME
tumor microenvironment
- TIDE
Tumor Immune Dysfunction and Exclusion
- TCGA
The Cancer Genome Atlas
1. Introduction
Bladder cancer (BLCA) is a male urologic malignancy with high mortality and recurrence rates [1]. Clinical evidence proved that the current treatments for BLCA include chemotherapy, total body resection surgery and emerging immunotherapy, among which platinum-based chemotherapy and surgical resection are the most effective options for BLCA treatment [2]. However, high recurrence results in a low 5-year overall survival rates of only 23%–48 % [3,4], but platinum-based chemotherapy is highly aggressive [2]. Due to the current lack of universal prognostic biomarkers for BLCA, urologists could not accurately determine the prognostic risk of patients as well as develop treatment plans [5]. Therefore, there was an urgent demand for effective and novel prognostic biomarkers [[6], [7], [8]].
Fatty acid metabolism (FAM) is an important source of energy, cellular components, and signaling molecules in cellular life activities, and it also played an essential function in cancer [9]. Numerous studies already demonstrated that dysregulated FAM in tumor microenvironment (TME) influenced tumor progression [10,11]. For example, regulatory T cells (Treg) are affected by FA in TME, inducing suppressive TME and angiogenesis to promote tumor metastasis [12]. Notably, a recent study showed that dysregulated FAM in TME also influences tumor cell chemoresistance via FA levels to disrupt cellular uptake of chemotherapeutic agents [13], which can be extremely detrimental to some BLCA patients who could only receive chemotherapy. Therefore, it is crucial to further investigate the molecular mechanisms of FAM in BLCA as well as the correlation between FA-related genes and chemotherapy resistance in BLCA.
In this study, we explored the molecular mutational landscape of genes involved in FAM in BLCA patients and identified molecular subtypes of BLCA with different levels of FAM in BLCA. Finally, a robust prognostic risk assessment system with Nomogram was constructed based on key prognostic genes to assess the potential clinical value in BLCA patients. In addition, we evaluated immune checkpoints, immunotherapy response, and chemotherapy sensitivity in patients with different molecular subtypes, prognostic risk, and this study was dedicated to provide new prognostic insights for BLCA.
2. Materials and methods
2.1. Dataset and filtering
The high-throughput sequencing dataset TCGA-BLCA of BLCA and somatic mutation data were accessed in TCGA (https://portal.gdc.cancer.gov/) database. The GSE32894 cohort was from in the Gene Expression Omnibus (GEO, https://www.ncbi.nlm.nih.gov/geo/) database. TCGA-BLCA was used as the training cohort as well as GSE32894 as the validation cohort in this study. Samples was processed in SangerBox (http://sangerbox.com/home.html) database, from which samples without follow-up information as well as Gene symbol were removed [14]. After processing, 403 tumor samples were stored in the TCGA-BLCA cohort while 316 samples were in the GSE32894 cohort. The HALLMARK_FATTY_ACID_METABOLISM gene set was sourced from the Molecular Signatures Database (MSigDB, https://www.gsea-msigdb.org/gsea/msigdb/index.jsp), from which the FAM genes (FAMGs) were extracted.
2.2. Identification of fatty acid molecular subtypes
Univariate COX model analysis was conducted with FAMGs in the TCGA-BLCA cohort was to determine FAMGs affecting BLCA prognosis (p < 0.05). Based on the FAMGs related with BLCA prognosis, consistent cluster analysis was performed to determine molecular subtypes of fatty acid profiles in BLCA following a previous study [15]. The “pam” algorithm and “spearman” were considered as a measurement distance, and 500 bootstraps were conducted, and 80 % of the patients in the training set randomly selected for each bootstraps. The optimal number of clusters was determined between 2 and 10 according to the consistency matrix and consistency cumulative distribution function to finally decide the molecular subtypes for the samples.
2.3. Biological pathway analysis in subtype groups
In order to explore some of the different active biological pathways in molecular subtypes. We collected h.all.v7.5.1.entrez.gmt gene sets in the MSigDB database and performed gene set enrichment analysis based on these gene sets [16,17]. Significantly enriched biological pathways were compared between FAMC1_VS_other (FAMC2 and FAMC3), FAMC2_VS_other (FAMC1 and FAMC3), FAMC3_VS_other (FAMC1 and FAMC2), respectively, in the TCGA-BLCA cohort. Pathways with FDR <0.05 were considered to be significantly enriched in the subtype.
2.4. Differential analysis and developing a protein-protein interaction network
Differential analysis was performed using the limma code package [18] in different fatty acid molecular subtypes to determine the differentially expressed FAMGs. Under the parameter of FDR<0.05 & |log2FC|>1, Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis was performed in enrichment analysis of differentially expressed FAMGs by the clusterProfiler code package [19]. FAM pathway activity assessment was achieved by single sample gene set enrichment analysis (ssGSEA) by the GSVA package [20] to explore the pathway activity changes, which meant FAM score. Differential analyses were conducted among fatty acid molecular subtypes (FDR <0.05 & |log2FC| > 1). Based on the identified differentially expressed genes (DEGs), the protein-protein interaction network (PPI network) of DEGs was created in the STRING (https://cn.string-db.org/, version 11.5) database, a confidence score of score >0.7 was chosen and after removing all isolated and partially disconnected nodes, the PPI network was developed. The PPI network was visualized by Cytoscape software (version 3.9.1), of which the MCODE clustering algorithm was used for sub-network screening [21]. In addition, the genes in the MCODE clustering were also analyzed for GO KEGG annotation, which was implemented by the clusterProfiler code package to explore their biological functions and meanings. GO terms and KEGG pathways with FDR<0.05 were considered to be significantly enriched for genes in MCODE clustering.
2.5. Construction of BLCA prognostic risk model
Taking DEGs in PPI networks as the research target, univariate COX, LASSO COX, and multivariate COX models were conducted in the TCGA-BLCA cohort to screen important prognostic factor. Based on those prognostic factors, we constructed a RiskScore. The prognostic risk of BLCA patients was calculated by formula (1):
| (1) |
where i referred to the level of gene expression and β was the Cox regression coefficient of a gene. High and low RiskScore groups were divided based on the median RiskScore. K-M survival curves were used for prognostic analysis. RiskScore function was validated in the validation set GSE32894 cohort.
2.6. Gene set enrichment analysis (GSEA)
Pathway analysis was performed using GSEA to analyze biological processes pathways in different molecular subtypes and RiskScore groups. Candidate gene sets were conducted, HALLMARK gene set was captured for GSEA.
2.7. Analysis of immune cell infiltration score in TME
Relative abundance of 22 immune cells in TME was quantified by CIBERSORT algorithm. ESTIMATE, MCP-Count, and ssGSEA were utilized to calculate immune cell scores [[22], [23], [24], [25]].
2.8. Analysis of immunotherapy response
To investigate the treatment response of patients in different molecular subtypes and RiskScore groups after receiving immunotherapy, the Tumor Immune Dysfunction and Exclusion (TIDE: http://tide.dfci.harvard.edu/login/) scores of BLCA patients were obtained at TIDE website. The T-cell-inflamed GEP-related signature gene set was accessed from Ayers et al. to calculate the T-cell-inflamed GEP Score for patients in the RiskScore groups by ssGSEA [26].
2.9. Chemotherapy drug sensitivity analysis
The IC50 values of eight chemotherapeutic agents were calculated by ridge regression using the R-code package pRRophetic [27] to determine the chemotherapeutic agents applicable to patients in distinct RiskScore groups.
2.10. Nomogram construction
Univariate and multivariate COX models were conducted to determine the factors significantly affecting BLCA prognosis by RiskScore, and a Nomogram was constructed. The prediction results of the two were assessed by Calibrationcurve, Decision curve and ROC curve.
3. Results
3.1. Mutation landscape of prognostic FAM genes
Initially, univariate COX model analysis of FAM genes (FAMGs) in TCGA-BLCA was conducted, obtaining 30 prognosis-related FAMGs (FDR <0.05 & |log2FC| > 1). Mutation landscape of 30 FAMGs were shown in Supplementary Fig. 1A, we noted that FASN, PTPRG, CYP4A22, HAO2 and HMGCS2 were the top 5 frequently mutated genes, of which most mutation was missense mutation. Then, it could be viewed that 30 prognosis-related FAMGs showed low copy number variation (CNV) (Supplementary Fig. 1 1B). Further comparison on the expression of 30 prognosis-related FAMGs across the CNV-diploid group, CNV-amplification group, and CNV-deletion group showed that 18 FAMGs were remarkably higher expressed in the CNV-amplification group than the CNV-deletion group (Supplementary Fig. 1C, p < 0.05).
3.2. Molecular subtypes of 30 prognosis-related FAMGs based on consistent clustering
We clustered BLCA samples in TCGA-BLCA cohort according to the expression of 30 prognosis-related FAMGs using consistent clustering algorithm. When k = 3, samples clustering had the highest consistency (Fig. 1A–C). Therefore, three FAM molecular subtypes were defined, namely, FAMC1, FAMC2, FAMC3. K-M plot showed that BLCA patients in the FAMC1 had the most favorable prognosis (Fig. 1D). Moreover, FAMC1 exhibited extremely high FAM score (Fig. 1E). Finally, the expression levels of 30 prognosis-related FAMGs and the clinicopathological information of the patients in FAMC1-3 were presented in heatmap. It could be clearly observed that most FAMGs were highly expressed in FAMC2 and FAMC3, which also had poorer prognosis (Fig. 1F). There were distinct differences in Grade, Stage, M. Stage, T. Stage, and Status in FAMC1-3, and we found that most patients in FAMC1 were at early stages of cancer with more survived cases (Fig. 1F).
Fig. 1.
Identification of molecular subtypes associated with fatty acid metabolism in BLCA (A) CDF curves of TCGA-BLCA cohort samples when k = 2 to k = 10. (B) CDF Delta area curves of TCGA-BLCA cohort samples when k = 2 to k = 10. The maximum inflection point of the slope of the curve occurred at K = 3. (C) Heat map of sample clustering at consensus k = 3, Clustering of samples in the three subtypes was relatively well. (D) K-M survival curves of FAMC1-3 patients. (E) FATTY_ACID_METABOLISM score in FAMC1-3 groups. (F) 30 prognosis-related FAMGs expression heat map as well as clinical characteristics statistics. ns p > 0.05; *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001.
3.3. Distinctions of immune cells and checkpoint genes in BLCA
A total of infiltration scores of TME for 22 various immune cells in BLCA were counted using CIBERSORT website. Remarkable differences in some immune cell types between subtypes were detected (Fig. 2A). ssGSEA, MCP-Count immunoassays showed significantly higher immune cell scores and expression of immune-related gene sets in FAMC2 and FAMC3 than in FAMC1 (Fig. 2B). We also assessed the immune cell and stromal cell scores in TME of BLCA using ESTIMATE method, which could significantly find higher StromalScore, ImmuneScore, ESTIMATEScore in FAMC3 (Fig. 2C). Furthermore, the highest Angiogenesis score was detected in FAMC3 patients (Fig. 3D). Finally, we also analyzed the immunotherapeutic response of patients in the TCGA-BLCA cohort in FAMC1-3 and the differences in immune checkpoint gene expression. FAMC1-3 demonstrated a sequential increase in the expression of the immune checkpoint genes CD80, PDCD1LG2, HAVCR2, CD28, CTLA4, and PDCD1 (Fig. 2E), indicating that TME immunosuppression was most severe in patients with FAMC3. These findings were further confirmed by the results of the TIDE score, where we observed that the FAMC3 had significantly higher Exclusion score, TIDE score, and Dysfunction score than in the other two subtypes (Fig. 2F), and immunotherapy gain in FAMC3 patients was also worse than other two subtypes (Fig. 2G). These results indicated that cancer patients with FAMC3 subtype were more prone to occur immune escape, which in turn caused distal metastasis to promote malignant progression of BLCA.
Fig. 2.
Immunological characteristics among subtypes in the TCGA-BLCA cohort. (A) CIBERSORT results. (B) ssGSEA and MCP-Count results. (C) ESTIMATE results. (D) Angiogenesis score. (E) Immune checkpoint expression levels. (F) TIDE score. (G) TIDE immune response differences among FAMC1-3 subtypes. ns p > 0.05; *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001.
Fig. 3.
RiskScore construction and validation. (A) Trajectory of each independent variable with lambda. (B) Confidence interval of lambda under tenfold cross-validation. (C) Forest plot of multivariate COX model. (D) K-M survival curves of high and low RiskScore subgroups in TCGA-BLCA cohort. (E) ROC curves of TCGA-BLCA cohort. (F) ROC curves of high and low RiskScore subgroups in TCGA-BLCA cohort Score subgroup subgroup patients' survival status. (G) K-M survival curves of high and low RiskScore subgroup in GSE32894 cohort. (H) ROC curves of GSE32894 cohort. (I) Survival status of high and low RiskScore subgroup subgroup patients in GSE32894 cohort.
3.4. Identification of FAMGs in molecular subtypes and PPI network analysis
By differential analysis of FAMC1-3 subtypes, we identified 1451 FAMGs in FAMC1, 465 FAMGs in FAMC2, and 1047 FAMGs in FAMC3. A total of 1786 FAMGs were obtained to develop PPI networks by taking the intersection of FAMGs in the three groups. Next, we constructed PPI networks using the STRING database according to the confidence score >0.7 as the threshold, followed by visualization in Cytoscape 3.9.1 software. We found 1159 nodes in the PPI network. Module-based network analysis was performed using cytoscape mainly by MCODE algorithm to find tightly connected protein clusters in the target network, and here we selected module 1 with the highest score for functional analysis (Supplementary Fig. 2A). The KEGG results showed that the FAMGs in module 1 were mainly enriched in the Allograft rejection, ECM-receptor interaction, and Cytokine-cytokine receptor interaction pathways (Supplementary Fig. 2B). Process module was mainly enriched in cellular response to biotic stimulus, cell chemotaxis, and extracellular matrix organization entries (Supplementary Fig. 2C). In Cellular Component module, they were mainly enriched in MHC class II protein complex, MHC protein complex, and lumenal side of endoplasmic reticulum membrane (Supplementary Fig. 2D). The Molecular Function module was mainly enriched in the chemokine activity, chemokine receptor binding entries (Supplementary Fig. 2E).
3.5. GSVA results in FAMC1-3 subtypes of BLCA
Next, we analyzed the biological pathway differences in FAMC1-3 subtypes and observed that differentially expressed genes in FAMC1 were mainly enriched in ANGIOGENESIS, E2F_TARGETS, EPITHELIAL_MESENCHYMAL_TRANSITION, INTERFERON_ALPHA_RESPONSE, G2M_CHECKPOINT entries (Supplementary Fig. 3A). The differentially expressed genes in FAMC2 were mainly enriched in OXIDATIVE_PHOSPHORYLATION, E2F_TARGETS, INTERFERON_ALPHA_RESPONSE, INTERFERON_GAMMA_RESPONSE, MYC_TARGETS_V1 entries (Supplementary Fig. 3B). The differentially expressed genes in FAMC3 were mainly enriched in INFLAMM_RESPONSE, RESPONSE, INFLAMMATORY_RESPONSE, INTERFERON_ALPHA_RESPONSE, EPITHELIAL_MESENCHYMAL_TRANSITION, GAMMA_RESPONSE. E2F_TARGETS, G2M_CHECKPOINT entries (Supplementary Fig. 3C). Next, we analyzed the pathway differences activated in the FAMC1-3 isoforms using the GSVA method. Most pathways were inhibited in FAMC1, while most of them were activated in FAMC2 and FAMC3 (Supplementary Fig. 3D), and the activated pathways in FAMC2 and FAMC3 were mainly associated with tumor metastasis.
3.6. RiskScore construction and validation
Effective prognostic markers are crucial for BLCA prognostic evaluation. We performed univariate COX model analysis of FAMGs in Module 1 and identified 55 FAMGs showing a high prognostic impact on BLCA (p < 0.05). Next, we streamlined the model by reducing the number of genes by the LASSO COX model. Under 10-fold cross-validation, the model was the optimal at lambda = 0.0309 when the model contained 13 FAMGs, including COL4A2, COL6A1, LAMA2, ADAMTSL4, P4HA2, EGR1, ADAMTS15, CTSV, HLA__F, SRC, CTSE, CTLA4, CD3D (Fig. 3A–B). Finally, multivariate COX model selected SRC, CTLA4, CTSE, LAMA2, and ADAMTSL4 as the most important BLCA prognostic markers from the 13 FAMGs, and we defined a FAM prognostic scoring system for BLCA, RiskScore = −0.171*SRC-0.303*CTLA4- 0.092*CTSE+0.29*LAMA2+0.12*ADAMTSL4 (Fig. 3C).
Then, we calculated the RiskScore for all the samples in the TCGA-BLCA cohort and divided the samples into high RiskScore and low RiskScore groups based on the median RiskScore value. In the TCGA-BLCA cohort, the K-M curves showed a significant difference in prognosis between the high RiskScore and low RiskScore groups, with patients in the low RiskScore group having a better prognosis (Fig. 3D). The ROC curves showed that the AUC values of RiskScore predicting 1-year, 3-year, and 5-year survival in BLCA patients were 0.71, 0.68, and 0.65, respectively (Fig. 3E). Next, we counted the survival status of patients in different RiskScore groups, and found that a higher proportion of cases died in the high RiskScore group (Fig. 3F). It can be seen that RiskScore performed better in the training set TCGA-BLCA cohort and can predict the prognosis of BLCA patients. Then, we validated the RiskScore in the validation set GSE32894 cohort. The K-M curve results were consistent with the trend of TCGA-BLCA cohort, and the prognosis of patients with low RiskScore was better (Fig. 3G). The AUC values at 1, 3, and 5 years were 0.8, 0.84, and 0.86, respectively (Fig. 3H). Similarly, there were more death cases in the high RiskScore group (Fig. 3I). These above results suggested that the RiskScore in this study had clinical significance and can be used to predict the prognosis of BLCA patients.
3.7. Association between clinical characteristics of BLCA and RiskScore
Difference comparison in clinicopathological characteristics between RiskScore subgroups in the TCGA cohort showed significant differences between Stage and Grade in RiskScore, and more patients with high clinical grade in the high RiskScore group (Supplementary Figs. 4A–E). Also, we compared the differences in RiskScore between different clinicopathological subgroups, and found that the RiskScore increased with higher clinical grade in BLCA patients (Supplementary Figs. 4F–J). Finally, we compared survival differences between patients in RiskScore subgroups in Age, gender, Stage, Grade, and TNM. Stage subgroups, and found that patients with low RiskScore had better prognosis in all subgroups (Supplementary Fig. 4K). These results indicated that RiskScore possessed equally strong prognostic predictive power in different clinical subgroups, and that RiskScore was reliable in predicting the prognostic trend of BLCA patients.
3.8. Correlation between RiskScore and BLCA treatment response
The potential correlation between RiskScore and TME of BLCA was analyzed. TME activity of BLCA was assessed using the CIBERSORT algorithm, we found that 11 immune cell infiltration scores differed in the RiskScore grouping (Fig. 4A). RiskScore was significantly correlated with T cells CD4 memory resting, Macrophages M0, Macrophages M2, Neutrophils positive correlation and significant negative correlation with Plasma cells and T cells CD8 (Fig. 4B). Next, we calculated the ssGSEA scores of 29 TME signatures in BLCA, and the results showed that RiskScore was significantly and positively correlated with the signatures related to Angiogenesis, Fibroblasts, EMT signature, Proliferation rate, and Anti-tumor immune infiltration rate, but significantly negatively correlated with tumor immune infiltration rate (Fig. 4C–D). In addition, we also found a higher Angiogenesis score in the high RiskScore group (Fig. 4E).
Fig. 4.
Correlation between RiskScore and BLCA treatment response. (A) CIBERSORT results of patients in the RiskScore subgroup in the TCGA-BLCA cohort. (B) Correlation between RiskScore and immune cell infiltration score, realization indicates positive correlation, dashed line indicates negative correlation. (C) 29 TME signature scores. (D) Correlation between RiskScore and TME signature. (E) Angiogenesis score. (F) T-cell-inflamed GEP score. (G) TIDE score. (H) IC50 values of chemotherapeutic agents.
Based on a computerized predictive algorithm and using TCGA-BLCA sample expression profile data as the object of analysis, we predicted the sensitivity of BLCA samples to immunotherapy. We found that the T-cell-inflamed GEP score was significantly higher in low RiskScore patients than in high FAM_ Score patients (Fig. 4F). The TIDE score showed that patients with high RiskScore had higher TIDE score and were more prone to immune escape (Fig. 4G). Finally, we analyzed the correlation between RiskScore and chemotherapeutic agents, and the results showed that patients with high RiskScore were more sensitive to WH-4-023, TGX221, XMD8-85, and Midostaurin treatment, while those with low RiskScore were more sensitive to Rapamycin, Z-LLNle-CHO, A 443654, QS11 treatment (Fig. 4H). Overall, based on the results predicted by the computer algorithm, our study illustrated that RiskScore also had clinical significance in predicting the direction of BLCA treatment response.
3.9. GSEA results in RiskScore groups of BLCA
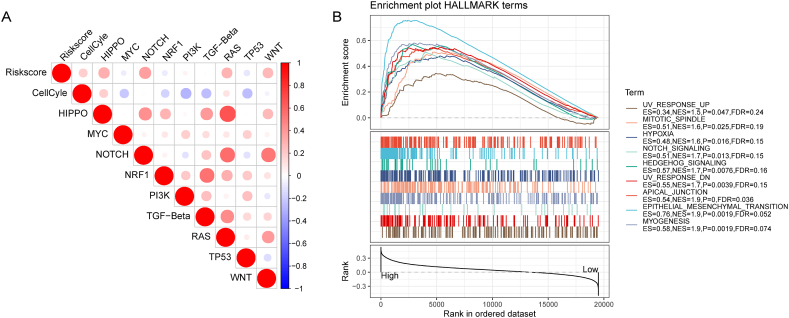
To investigate the correlation between RiskScore and pathways related to cancer development, we calculated the enrichment scores of 10 pathways related to cancer development by ssGSEA algorithm, and we found that RiskScore was associated with RAS, NRF1, Cell Cyle, HIPPO, TGF- Beta, PI3K, NOTCH, MYC, WNT, and TP53 pathways were all positively correlated (Fig. 5A). Our findings indicated that as the RiskScore increased, the risk of cancer development or malignant progression was higher. Next, the GSEA results for the RiskScore grouping showed that the high RiskScore groups of UV_RESPONSE_UP, MITOTIC_SPINDLE, HYPOXIA, NOTCH_SIGNALING, HEDGEHOG_SIGNALING, UV_RESPONSE_DN APICAL_JUNCTION, EPITHELIAL_MESENCHYMAL_TRANSITION, and MYOGENESIS pathways were significantly enriched in the high RiskScore group (Fig. 5B).
Fig. 5.
GSEA results in RiskScore groups of BLCA. (A) Heat map of correlation between RiskScore and cancer-related pathways. (B) GSEA results.
3.10. Nomogram construction
Univariate and multivariate Cox model analysis of RiskScore and clinical characteristics revealed that RiskScore and Age, Stage were significant prognostic factors (Fig. 6A–B). As shown in Fig. 6C, we integrated RiskScore with other clinicopathological traits to produce a Nomogram that quantified the risk assessment and survival chance of patients. The results showed that RiskScore had the strongest influence on the survival prediction. We further evaluated the prediction accuracy of the model using calibration curve, as displayed in Fig. 6D, the predicted calibration curves at 1-, 3-, and 5-years calibration points nearly overlapped with the standard curve, indicating an accurate prediction by the Nomogram. Decision curve was plotted for assessing the model reliability of the model, and the advantages of both Riskscore and nomogram were significantly greater than the extreme curves (Fig. 6E). Both RiskScore and nomogram were highly robust in comparison with other clinicopathological traits (Fig. 6F–H).
Fig. 6.
Nomogram construction. (A–B) Forest plot of single-factor and multi-factor COX models of RiskScore and clinical characteristics. (C) Columnar plot. (D) Calibration curve. (E) Decision curve. (F–G) The ROCcurves of a variety of clinical features for overall survival (OS) at 1-, 3-, and 5 years.
4. Discussion
In recent years, researchers have identified that dysregulation of FAM in the tumor microenvironment affects cancer progression, treatment, postoperative recurrence, and distant metastasis [28]. Accumulating evidence suggested that FAM affects numerous life activities such as cell membrane formation in cancer cells and signaling molecule transduction in tumor progression [29]. It was concluded that cancer cells endogenously produce lipids by capturing extracellular free fatty acid molecules or intracellularly through lipogenic pathways, and that excessive accumulation of fatty acids or lipids in the cytoplasm causes dysregulation of FAM and enhances cellular stress [29]. In this study, we performed a comprehensive systematic bioinformatics analysis of genes related to FAM in BLCA and distinguished different molecular subtypes (FAMC1, FAMC2, FAMC3) based on fatty acid metabolic activity for BLCA. And the RiskScore for assessing the prognosis of BLCA patients was systematically constructed to comprehensively assess its clinical value.
We found that immune cell scores increased sequentially in FAMC1, FAMC2, and FAMC3. First, we noticed that CD8 T cells, Type 1 T helper cell in FAMC3 clearly showed active status. High CD8 T cells abundance and high Type 1 T helper cell abundance were hallmarks of the inflammatory tumor microenvironment [30]. It suggested that FAMC3 may present an inflammatory tumor microenvironment phenomenon. Further, we also observed that immune checkpoint (CD80, PDCD1LG2, HAVCR2, CD28, CTLA4, and PDCD1) were significantly high-expressed in FAMC3. Combined with the TIDE results, these findings suggested that patients with FAMC3 subtype had the highest probability of immune escape. The study confirmed that high-expressed immune checkpoints can create a suppressive immune environment to allow tumor cells to circumvent the normal organismal immune function of immune cells [31]. CD28, CTLA4, and PDCD1 are associated with activated T cells and Tregs [32]. The study also confirmed that high immune checkpoint gene expression contributed to an inflammatory immune microenvironment [30]. This was consistent with our results that the expression levels of immune checkpoint genes were progressively higher in FAMC1, FAMC2, and FAMC3. It indicated that FAMC1 was a non-inflammatory immune microenvironment subtype and FAMC3 was an inflammatory immune microenvironment subtype. Overall, the identification of FAM molecular subtypes complements the pathological findings of BLCA and is expected to play a crucial role in identifying patients with immunosuppressive microenvironment.
Finally, the RiskScore developed in this study contained 5 reliable FAM genes, namely SRC, CTLA4, CTSE, LAMA2, and ADAMTSL4. RiskScore can not only predict the prognosis of BLCA patients, but also the immunotherapy response, and chemotherapy drug sensitivity. And the results have been further validated in independent datasets. Several studies were conducted to confirm the critical role of these FAMRGs in cancer. For example, SRC is the first oncogene identified, and SRC is associated with multiple immune cells and signaling pathways in TME, positively affecting cancer progression, treatment resistance [33]. CTLA4 is the most common immune checkpoint gene, and previous studies demonstrated that blocking CTLA4 responses increases T cell immune responses, suppresses TGF-β secretion, and enhances IFN-γsecretion [34]. A study by Chou et al. demonstrated that high expression of CTSE could be a predictor of disease-specific survival (DSS), metastasis-free survival (MeFS) and local recurrence-free survival (LRFS) after colon cancer patients receiving preoperative synchronous radiotherapy [35]. Liang et al. used an in vitro nude mouse model of lung cancer metastasis and observed that Mex3a downregulates LAMA2 expression in tumor cells and activates PI3K/AKT pathway to promote lung adenocarcinoma metastasis [36]. ADAMTSL4, a secreted glycoprotein, is a novel immune-related biomarker for primary glioblastoma multiforme [37]. FAMRGs contained in the RiskScore were associated with cancer progression signaling pathways, however, these FAMRGs have been less studied in BLCA, and most of the existing studies focus on the remaining cancer species. Their key roles remain to be further explored. Overall, these findings may provide new insights for elucidating the dysregulation of FAM in TME of BLCA.
To conclude, our study demonstrated that fatty acid dysregulation influenced BLCA progression. Furthermore, the RiskScore composed of FAMRGs was an important prognostic indicator of BLCA and a therapeutic assessment parameter. Although the RiskScore showed excellent robustness, subsequent clinical trials with larger samples in multiple centers are required for in-depth evaluation.
5. Conclusion
In conclusion, our study established a dependable prediction model based on the composition of FAMRGs. The RiskScore was a reliable prognostic indicator for BLCA, and the Nomogram composed of the RiskScore was a highly applicable clinical tool. In addition, RiskScore had a strong potential clinical value in predicting treatment response in BLCA patients. The current findings supported the potential significance of key genes in FAM as potential therapeutic targets for BLCA.
Ethics approval and consent to participate
Informed consent was not required for this study because.
Consent for publication
None.
Funding
This study was funded by Tianjin Key Medical Discipline (Specialty) Construction Project(TJYXZDXK-009A.
Availability of data and material
The datasets analyzed in this study are available in the GSE datasets (http://www.ncbi.nlm.nih.gov/geo), [GSE32894] at [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE32894]. The code used in this study can be obtained from https://github.com/xchen1212/BLCA_Fatty_acid.git.
CRediT authorship contribution statement
Xusheng Chen: Data curation, Investigation, Resources, Supervision, Writing – original draft, Writing – review & editing. Zhenting Zhang: Investigation, Resources, Supervision, Visualization, Writing – original draft. Wenfeng Liao: Investigation, Resources, Supervision, Writing – review & editing. Yujie Zhao: Formal analysis, Project administration, Supervision, Visualization.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgements
None.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.heliyon.2023.e22768.
Appendix ASupplementary data
The following are the supplementary data to this article:
References
- 1.Sung H., et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71(3):209–249. doi: 10.3322/caac.21660. [DOI] [PubMed] [Google Scholar]
- 2.Seiler R., et al. Impact of molecular subtypes in muscle-invasive bladder cancer on predicting response and survival after neoadjuvant chemotherapy. Eur. Urol. 2017;72(4):544–554. doi: 10.1016/j.eururo.2017.03.030. [DOI] [PubMed] [Google Scholar]
- 3.Kuroki L., Guntupalli S.R. Treatment of epithelial ovarian cancer. BMJ. 2020;371:m3773. doi: 10.1136/bmj.m3773. [DOI] [PubMed] [Google Scholar]
- 4.Lee J.Y., et al. Changes in ovarian cancer survival during the 20 years before the era of targeted therapy. BMC Cancer. 2018;18(1):601. doi: 10.1186/s12885-018-4498-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Witjes J.A., et al. European association of urology guidelines on muscle-invasive and metastatic bladder cancer: summary of the 2020 guidelines. Eur. Urol. 2021;79(1):82–104. doi: 10.1016/j.eururo.2020.03.055. [DOI] [PubMed] [Google Scholar]
- 6.Baladehi R.F., et al. The effect of oncogene proteins of human papillomaviruses on apoptosis pathways in prostate cancer. Oncologie. 2022;24(2) [Google Scholar]
- 7.Liu C., et al. Ferroptosis's role in genitourinary system cancer. Oncologie. 2022;24(4) [Google Scholar]
- 8.Wang D., Chen H., Hu Y. Polarized autologous Macrophages (PAM) can Be a tumor vaccine. Oncologie. 2022;24(3) [Google Scholar]
- 9.Koundouros N., Poulogiannis G. Reprogramming of fatty acid metabolism in cancer. Br. J. Cancer. 2020;122(1):4–22. doi: 10.1038/s41416-019-0650-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Milo I., et al. The immune system profoundly restricts intratumor genetic heterogeneity. Sci Immunol. 2018;3(29) doi: 10.1126/sciimmunol.aat1435. [DOI] [PubMed] [Google Scholar]
- 11.Li X., et al. Navigating metabolic pathways to enhance antitumour immunity and immunotherapy. Nat. Rev. Clin. Oncol. 2019;16(7):425–441. doi: 10.1038/s41571-019-0203-7. [DOI] [PubMed] [Google Scholar]
- 12.Lan H.R., et al. Role of immune regulatory cells in breast cancer: foe or friend? Int Immunopharmacol. 2021;96 doi: 10.1016/j.intimp.2021.107627. [DOI] [PubMed] [Google Scholar]
- 13.Hoy A.J., Nagarajan S.R., Butler L.M. Tumour fatty acid metabolism in the context of therapy resistance and obesity. Nat. Rev. Cancer. 2021;21(12):753–766. doi: 10.1038/s41568-021-00388-4. [DOI] [PubMed] [Google Scholar]
- 14.Shen W., et al. Sangerbox: a comprehensive, interaction-friendly clinical bioinformatics analysis platform. iMeta. 2022;1(3):e36. doi: 10.1002/imt2.36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Wilkerson M.D., Hayes D.N. ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics. 2010;26(12):1572–1573. doi: 10.1093/bioinformatics/btq170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Liberzon A., et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 2015;1(6):417–425. doi: 10.1016/j.cels.2015.12.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Liberzon, A., et al., The Molecular Signatures Database (MSigDB) Hallmark Gene Set Collection. (2405-4712 (Print)).. [DOI] [PMC free article] [PubMed]
- 18.Ritchie M.E., et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43(7):e47. doi: 10.1093/nar/gkv007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Yu G., et al. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16(5):284–287. doi: 10.1089/omi.2011.0118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Hänzelmann S., Castelo R., Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinf. 2013;14:7. doi: 10.1186/1471-2105-14-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Shannon P., et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13(11):2498–2504. doi: 10.1101/gr.1239303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Chen B., et al. Profiling tumor infiltrating immune cells with CIBERSORT. Methods Mol. Biol. 2018;1711:243–259. doi: 10.1007/978-1-4939-7493-1_12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Yoshihara K., et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 2013;4:2612. doi: 10.1038/ncomms3612. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Barbie D.A., et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature. 2009;462(7269):108–112. doi: 10.1038/nature08460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Becht E., et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol. 2016;17(1):218. doi: 10.1186/s13059-016-1070-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ayers M., et al. IFN-gamma-related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Invest. 2017;127(8):2930–2940. doi: 10.1172/JCI91190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Geeleher P., Cox N., Huang R.S. pRRophetic: an R package for prediction of clinical chemotherapeutic response from tumor gene expression levels. PLoS One. 2014;9(9) doi: 10.1371/journal.pone.0107468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Corn K.C., Windham M.A., Rafat M. Lipids in the tumor microenvironment: from cancer progression to treatment. Prog. Lipid Res. 2020;80 doi: 10.1016/j.plipres.2020.101055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Currie E., et al. Cellular fatty acid metabolism and cancer. Cell Metab. 2013;18(2):153–161. doi: 10.1016/j.cmet.2013.05.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Gajewski, T.F., et al., Cancer Immunotherapy Targets Based on Understanding the T Cell-Inflamed versus Non-T Cell-Inflamed Tumor Microenvironment. (65-2598 (Print)).. [DOI] [PMC free article] [PubMed]
- 31.Li B., Chan H.L., Chen P. Immune checkpoint inhibitors: basics and challenges. Curr. Med. Chem. 2019;26(17):3009–3025. doi: 10.2174/0929867324666170804143706. [DOI] [PubMed] [Google Scholar]
- 32.Ding C., et al. Characterization of the fatty acid metabolism in colorectal cancer to guide clinical therapy. Mol Ther Oncolytics. 2021;20:532–544. doi: 10.1016/j.omto.2021.02.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Caner A., Asik E., Ozpolat B. SRC signaling in cancer and tumor microenvironment. Adv. Exp. Med. Biol. 2021;1270:57–71. doi: 10.1007/978-3-030-47189-7_4. [DOI] [PubMed] [Google Scholar]
- 34.Zheng F., et al. Cancer stem cell vaccination with PD-L1 and CTLA-4 blockades enhances the eradication of melanoma stem cells in a mouse tumor model. J. Immunother. 2018;41(8):361–368. doi: 10.1097/CJI.0000000000000242. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Chou C.L., et al. CTSE overexpression is an adverse prognostic factor for survival among rectal cancer patients receiving CCRT. Life. 2021;11(7) doi: 10.3390/life11070646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Liang J., et al. Mex3a interacts with LAMA2 to promote lung adenocarcinoma metastasis via PI3K/AKT pathway. Cell Death Dis. 2020;11(8):614. doi: 10.1038/s41419-020-02858-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Zhao Z., et al. ADAMTSL4, a secreted glycoprotein, is a novel immune-related biomarker for primary glioblastoma multiforme. Dis. Markers. 2019;2019 doi: 10.1155/2019/1802620. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The datasets analyzed in this study are available in the GSE datasets (http://www.ncbi.nlm.nih.gov/geo), [GSE32894] at [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE32894]. The code used in this study can be obtained from https://github.com/xchen1212/BLCA_Fatty_acid.git.