Abstract
Genes essential for Salmonella typhimurium invasion have been localized to Salmonella pathogenicity island 1 (SPI1) on the chromosome. However, it is clear that other genes are required for the invasion process. Mutations that abolish the SPI1 invasion type III secretion system do not significantly reduce invasion into Chinese hamster ovary tissue culture cells. Two invasion defective mutants were isolated by screening 2,500 Tn10dTc insertion mutants of S. typhimurium in the tissue culture invasion assay. One of the invasion mutants, SVM167, has an insertion between centisomes 24.5 and 25.5 in an operon homologous to the ipgDEF operon of the Shigella flexneri and Shigella sonnei virulence plasmid. A second mutant, SVM168, has an insertion in an IS3-type element with homology to the Salmonella enteritidis IS1351 element and Yersinia enterocolitica IS1400 element from a high-pathogenicity island. Further characterization of SVM167 showed that culture supernatants from this mutant lack a previously uncharacterized protein that is also missing from culture supernatants of a SPI1 mutant, suggesting it can be secreted by the SPI1 type III secretion system. In addition, transcription of this operon, sigDE (Salmonella invasion gene), is dependent on the presence of sirA, an activator of hilA expression. HilA activates transcription of several of the SPI1 genes but does not appear to have a major role in activation of transcription from the sigDE promoter.
In order for salmonellae to cause disease (enteric fever or gastroenteritis) via an oral route of infection, the bacteria must penetrate the epithelial cells of the intestine (10, 33, 54, 69). Several studies have shown that salmonellae preferentially attach and enter phagocytic M cells of the Peyer’s patches (33, 37, 56). However, studies have also shown that bacteria can be found in nonphagocytic enterocytes and can also cause a systemic infection when the small intestine is bypassed (10). Much research has focused on the process of invading host cells, as this ability appears to be an important aspect of pathogenesis (for reviews, see references 17, 18, and 21). Various tissue culture cell lines, including HEp-2, Henle-407, and Chinese hamster ovary (CHO), have been used as model systems to investigate this process. Upon contact with intestinal or cultured epithelial cells, salmonellae induce a dramatic ruffling of the cell surface caused by cytoskeletal rearrangements, facilitating the engulfment of bacteria (19, 20). Mutants that are not able to actively invade host cells are noticeably attenuated in intragastrically, but not intraperitoneally, infected mice (22). Invasion requires the expression of many genes, including those encoding transcriptional regulators (2, 4, 5, 38, 44, 49, 55), components of a supramolecular secretory apparatus (13, 14, 22, 23, 26, 38), and potential effector proteins secreted by this apparatus (12, 13, 32, 34, 39, 40, 73). The majority of the known invasion genes map between centisomes 62 and 64 of the Salmonella chromosome, a region known as Salmonella pathogenicity island 1 (SPI1). These genes include the inv, hil, prg, org, spa, and sip/ssp genes (for reviews, see references 21 and 43).
Various techniques, including complementation of a naturally occurring attenuated isolate with a genomic cosmid library (22), mutagenesis with a transposon containing a strong outward facing promoter to find hyperinvasive mutants (44), and identification of anaerobically induced genes (36), have been used to identify invasion genes. Recently, signature-tagged transposon mutagenesis was used to isolate attenuated Salmonella typhimurium from intraperitoneally infected mice (30). Several loci were found to affect survival in the mouse, but most notable was the characterization of Salmonella pathogenicity island 2 (SPI2) (31, 64). This region had also been identified in another study examining sequences of the Salmonella chromosome which are not found in Escherichia coli and appear to be necessary for macrophage survival (53). Although it appears that the main function of the SPI2-encoded proteins is survival in the host, certain mutations in this region also have a measurable effect on invasion in vitro (31).
Despite the numerous creative techniques used to find invasion genes, none of these strategies has saturated the Salmonella chromosome; each study has failed to reidentify all known invasion genes. In addition, although SPI1 mutants are significantly defective for invasion of the intestinal epithelium, they are not completely attenuated in mice, suggesting that bacteria can still reach the deeper tissues in the mouse model after oral inoculation (22). These observations strongly suggest the existence of additional, unidentified invasion genes.
MATERIALS AND METHODS
Bacterial strains, bacteriophage, and plasmids.
Table 1 provides descriptions of the bacterial strains, bacteriophage, and plasmids used in this study. Chemical transformation and electroporation of plasmids into bacteria were carried out as previously described (61). Plasmids that were manipulated in E. coli were passaged through a restriction-minus (res) S. typhimurium LT2 strain (LB5000) prior to transformation or electroporation into other S. typhimurium strains.
TABLE 1.
Bacterial strains, plasmids, and bacteriophage used in this work
| Strain, plasmid, or phage | Genotype | Source or referencea |
|---|---|---|
| S. typhimurium | ||
| 14028s | Wild type | ATCC |
| SVM167 | sigE::Tn10dTc | This work |
| SVM168 | IS1351::Tn10dTc | This work |
| SVM173 | invA::Ωcat | This work; J. Galan |
| SVM169 | Same as SVM173 with a linked Tn10dTc insertion | This work |
| SVM174 | sigE::Tn10dTc invA::Ωcat | This work |
| SVM175 | IS1351::Tn10dTc invA::Ωcat | This work |
| EG9527 | mgtCB9232::MudJ (Knr) | E. Groisman |
| VV302 | SL1344 ΔhilA | 4 |
| SVM214 | ΔhilA | 4; this work |
| CJ010 | sirA::Tn10dTc phoN::Tn10dCm | 35 |
| CS015 | phoP::Tn10dCm | 49 |
| SVM226 | ssaJ::mTn5 | 31 |
| SVM227 | ssaT::mTn5 | 31 |
| SVM255 | 4-bp deletion in sigD | This work |
| SA536 | LT2 HfrK6 (origin min 76) | K. E. Sanderson (SGSC) |
| SA975 | LT2 HfrK13 (origin min 78) | K. E. Sanderson (SGSC) |
| TT10423 | LT2 proAB47/F′ pro+ lac+ zzf-1831::Tn10dTc | 15 |
| SB154 | SL1344 invA::Ωcat | Jorge Galan |
| LB5000 | LT2 flaA66 metA22 trp-2 rpsL xyl-401 ilv-452 leu res mod+ | B. Stocker |
| E. coli DH5α | F− p80dlacZΔM15 Δ(lacZYA-argF)U169 deoR recA1 endA1 hsdR17 (rK− mK+) phoA supE44 λ−thi-1 gyrA96 relA1 | Gibco, BRL |
| Plasmids | ||
| pHG329 | Apr; medium-copy-number cloning vector | 66 |
| pNK2881 | Apr; Ptac-tnpA ats-1 ats-2 | 41 |
| pRS415 and pRS528 | lacZYA transcriptional reporter fusion vectors | 65 |
| pWSK29 and pWKS30 | Apr; low-copy-number cloning vectors | 70 |
| pWSK129 and pWKS130 | Knr; low-copy-number cloning vectors | 70 |
| pMAK705 | Cmr; temperature-sensitive origin of replication vector used for gene replacement | 29 |
| pCJ13 | Apr; sirA uvrC in pWKS30 | 35 |
| pGP1-2 | Knr; λ pL-T7 RNA polymerase | 68 |
| pVV214 | Apr; hilA in pACYC177 | 4 |
| pHH10 | Apr; 4.1-kb EcoRI sigDE fragment cloned into pHG329 | This work |
| pHH20 | Apr; 3.2-kb BamHI sigDE fragment cloned into pWKS30 | This work |
| pHH21 | Apr; 0.9-kb EcoRI/PstI sigDE promoter and partial coding sequences in pRS415 | This work |
| pHH22 | Apr; same as pHH20 but in opposite orientation (BamHI/EcoRI fragment) | This work |
| pHH25 | Knr; 2.2-kb sirA PstI fragment from pCJ13 in pWSK129 | This work; 35 |
| pHH26 | Apr; same as pHH22 but with deleted PstI site in sigD | This work |
| pHH31 | Cmr; 1.7-kb KpnI fragment from pHH26 encompassing the sigD deletion cloned into pMAK705 | This work |
| Bacteriophage P22HT int | High-frequency transducing phage | 45 |
ATCC, American Type Culture Collection; SGSC, Salmonella Genetics Stock Centre.
Growth conditions.
Salmonella and E. coli strains were grown in Luria-Bertani (LB) Miller broth (Difco) at 37°C with aeration on a roller drum or without aeration in standing cultures, depending on the assay. Minimal medium was made with M63 salts, glucose, magnesium sulfate, and vitamin B1 (48). Evans-Blue uranine (EBU) agar and Bochner-Maloy media were made as previously described (45). Antibiotics were used at the following final concentrations: ampicillin, 100 μg/ml; tetracycline, 15 μg/ml; kanamycin, 25 μg/ml; erythromycin, 150 μg/ml; and chloramphenicol, 25 μg/ml. For detection of β-galactosidase activity, solid medium was supplemented with 5-bromo-4-chloro-3-indolyl-β-d-galactoside (X-Gal) at 40 μg/ml; when necessary, isopropylthio-β-d-galactoside (IPTG) was used at 100 μM. P22HT int lysates were harvested and used for transductions as previously described (45).
Transposon mutagenesis.
To create random Tn10dTc (41) insertions in the S. typhimurium chromosome, pNK2881 (ampicillin resistant [Apr]), containing a transposase gene (tnpA), was transformed into the wild-type parent strain 14028s. A P22 HT int lysate was prepared from TT10423 (15), a strain which carries a Tn10dTc on an F′ element. This lysate was used to transduce Tn10dTc into 14028s (pNK2881); transductions were performed as previously described (45) and plated on minimal (M63) salts agar supplemented with tetracycline. Transductants were pooled (ca. 1,000 per pool) and inoculated with P22 HT int broth (LB, E salts, bacteriophage P22HT int) (45) to make a lysate of these transductants. These lysates were then used to transduce the Tn10dTc insertions into SVM169 (invA::Ωcat). Transductants were plated on minimal medium as described above. Individual transductants were inoculated into 200 μl of LB-tetracycline-chloramphenicol-EGTA broth in each well of 96-well microtiter plates and incubated overnight without shaking at 37°C, and 10 μl of each was used for qualitative invasion assays. Before further characterization, putative mutants were plated on EBU-chloramphenicol-tetracycline agar plates to purify bacteria away from phage (45). Once free of phage, bacteria were cross-streaked against P22 HT int to determine if they were P22 sensitive (true transductants) or resistant (pseudolysogens). The motility phenotype was determined by stabbing bacteria into 0.25% LB-agar plates. The Tn10dTc insertions were separated from the invA mutation by transduction into the wild-type background and purified as described above.
pHH10, encoding sigDE, was mutagenized with TnMax2 as previously described (28). Plasmids with insertions in sigD were transformed into strain 14028s, and culture supernatant proteins were examined as described below (data not shown).
Construction of a sigD mutant.
To construct the sigD mutant, SVM255, pMAK705 (chloramphenicol resistant [Cmr]), with a temperature-sensitive origin of replication, was used as previously described (29). At 44°C, pMAK705 derivatives form cointegrates, but at 30°C, these cointegrates resolve from the chromosome as replicating plasmids. A 1.7-kb KpnI fragment from pHH26 carrying a partial sigD fragment with a 4-bp deletion within the coding sequence was cloned into pMAK705, creating pHH31. This plasmid was electroporated into and purified from LB5000 before being electroporated into wild-type S. typhimurium 14028s. After several shifts in temperature between 30 and 44°C, 40 colonies were purified, cured of the plasmid, and screened in the invasion assay. Approximately 50% of the tested clones were reduced for invasion. Culture supernatants from three of the invasion mutants were analyzed and found to lack the same protein absent from SVM167 (sigE::Tn10dTc). Southern analysis using a probe to sigD confirmed the exchange of the disrupted sigD allele with the wild-type allele on the chromosome; this was easily detected because the 4-bp deletion in sigD removed a PstI site (data not shown).
Construction of isogenic invA and hilA mutants.
SVM173 was constructed by transducing an invA::Ωcat mutation (courtesy of Jorge Galan) from strain SB154 into our parent strain, 14028s. The original hilA deletion mutant, VV302 (courtesy of Catherine Lee), was made in S. typhimurium SL1344 (4) and does not contain an associated selectable marker. To transfer the hilA deletion into 14028s, a P22HT int lysate was prepared from VV302 and used to transduce the deletion into SVM169, a 14028s derivative which contains the invA::Ωcat mutation linked to an unmapped Tn10dTc insertion. Transductants were plated on Bochner-Maloy medium, which selects against tetracycline resistance (Tcr). SVM169 is Tcr; thus, only transductants which have lost the Tn10dTc marker could grow. Because the wild-type hilA, invA::Ωcat and Tn10dTc were all linked, some Tcs transductants were expected to have replaced the hilA+ and invA::Ωcat with ΔhilA and invA+ sequences. Forty-four Tcs transductants were purified on EBU plates and confirmed to be P22 sensitive and phage free. All 44 were also Cms sensitive, indicating that the invA::Ωcat insertion was replaced with a wild-type invA copy. Four spontaneous Tcs colonies (from a control plate which had bacteria only, no phage) had also been tested on chloramphenicol plates, and all were resistant as expected. Invasion assays in the HEp-2 cell line were done with the control strains (wild type and VV302) and 44 putative 14028sΔhilA strains; all except the wild type were equally reduced for invasion (data not shown). Southern analysis (61) was performed on chromosomal DNA purified from four of the new ΔhilA strains, VV302, and 14028s. An NsiI fragment containing hilA from pVV214 (4) was used as a probe to confirm the deletion in the 14028s ΔhilA derivatives (data not shown).
Plasmid constructions.
All relevant plasmids used in this study are described in Table 1. The junctions between the transposon insertions and chromosomal sequences were cloned as follows. Chromosomal DNA preparations isolated from SVM167 and SVM168 were digested with EcoRI, separated on a 0.7% Tris-acetate-EDTA-agarose gel, and transferred to nitrocellulose filters (61). Because the tetAR genes are lethal to E. coli in multicopy (41), the cloning of random fragments and selection for Tcr was not possible; EcoRI was chosen since it has a site within the Tcr genes. The filters were probed with a [32P]dATP-labeled BglII fragment from pNK81 encoding the tetAR locus (41) to determine the approximate sizes of the junction fragments that needed to be cloned. Size-fractionated fragments of chromosomal DNA isolated from the mutants were cloned into pHG329 (66), and transformants were screened by colony hybridization using the same tetAR probe. Once cloned, the sequences were used to probe subgenomic libraries for wild-type sequences. For SVM167, pHH10 was isolated and found to have a 4.1-kb EcoRI fragment carrying the wild-type sigDE locus.
For complementation studies, a 3.2-kb BamHI fragment from pHH10 was subcloned into the low-copy-number cloning vector pWKS30 (70). For T7 expression, a 3.2-kb EcoRI/BamHI sigDE fragment from pHH20 was cloned into pWSK29 (the sister plasmid to pWSK30 with the polylinker in the opposite orientation), creating pHH22. To create pHH26, a small deletion was made in sigD in pHH22 as follows. pHH22 was digested with PstI to completion and then incubated with Deep Vent DNA polymerase (New England Biolabs), Deep Vent DNA polymerase buffer, and 200 μM deoxynucleoside triphosphates at 72°C for 20 min as described by the manufacturer. These fragments were recircularized with ligase and recovered. DNA sequence analysis showed that a 4-bp deletion was created as expected (data not shown). sigDE was expressed in the T7 expression system as previously described (3, 68). Sodium dodecyl sulfate (SDS)-polyacrylamide (10%) gels were prepared as previously described (42), dried under vacuum, and exposed to X-ray film for 1 and 16 h (3).
To create the reporter plasmid pHH21, a 1-kb PstI fragment from pHH10 was cloned into pHG329 to create pHH15. This fragment contained sequences starting about 408 bp upstream of the putative start codon of sigD. From pHH15, the sigDE promoter region was isolated as an 1.0-kb EcoRI fragment and was cloned into pRS528, a lacZYA transcriptional fusion reporter vector (65), to create pHH17. Although dark blue colonies on X-Gal were selected, sequence analysis revealed the sigDE promoter was in the wrong orientation relative to lacZYA, probably because of the presence of an activated or constitutively expressed divergent promoter in pHH17. This construct was used to directionally clone the sigDE promoter region into the sister transcriptional fusion vector, which has a polylinker in the opposite orientation of pRS528 (pRS415). The resulting plasmid with a sigD::lacZYA transcriptional fusion was named pHH21.
Tissue culture attachment and invasion assays.
CHO and HEp-2 cells were maintained and passaged as recommended by the American Type Culture Collection. For tissue culture invasion (TCI) assays, 2 × 105 cells/ml were seeded in Falcon 24-well tissue culture plates to obtain about 90% confluent monolayers on the following day. For bacterial cultures, single colonies were inoculated into 2 ml of medium and grown for 18 h without shaking; 5 μl (107 to 108 CFU) was used per well of tissue culture cells. Trays were centrifuged for 10 min at 1,000 rpm in a tabletop swing-bucket centrifuge and incubated for 1 h at 37°C with 5% CO2. For the mutants isolated in this study, centrifugation was ultimately found not to make a difference. Monolayers were then washed three times with phosphate-buffered saline (PBS) for tissue culture and then incubated for another 90 min in the presence of RPMI 1640 (Cellgro) with gentamicin at 100 μg/ml to kill extracellular bacteria. The monolayers were then washed with PBS as described previously (67), lysed with double-distilled water for 20 min, diluted, and plated on selective LB-agar plates to determine the number of intracellular bacteria. Quantitative TCI assay results are calculated as follows: percent invasion = 100 × (number of bacteria resistant to gentamicin/total number of bacteria added). Attachment assays were performed in the same manner but with five washes after the first incubation and omission of the gentamicin killing step.
Mapping and transduction analysis.
The Tn10dTc insertion in SVM167 was mapped by using a set of MudP22 mapping strains (6, 45). SVM168 could not be mapped by the MudP22 mapping phage and was therefore mapped with a set of Hfr (high-frequency recombination) strains (Salmonella Genetics Stock Centre). Cultures of each Hfr strain were mixed with equal volumes of SVM168 containing invA::Ωcat (in order to select against the donor Hfr strain), diluted, and plated onto Bochner-Maloy medium (45). Two of ten Hfr strains representing regions of the chromosome not represented by the MudP22 mapping phage (SA536 and SA975, with origins of transfer at centisomes 76 and 78, respectively) were able to recombine out the Tcr marker from SVM168.
Analysis of culture supernatants.
Cultures were grown in 2 ml of LB broth with antibiotics for 18 h, without aeration, and harvested. Bacteria were pelleted at 8,000 × g for 15 min, and the supernatants were immediately transferred to clean tubes. The supernatants were filtered through 0.45-μm-pore-size Tuffryn (low-protein-binding) Gelman Acrodiscs, and proteins were precipitated with cold trichloroacetic acid (TCA) at a final concentration of 10%. The proteins were collected by centrifugation at 8,000 × g at 4°C and resuspended in 1 ml of cold acetone. These mixtures were centrifuged for 10 min at 8,000 rpm at 4°C, and pellets were resuspended in 30 μl of PBS. Sample buffer with β-mercaptoethanol (61) was added to samples, the samples were boiled for 5 min, and proteins were separated by SDS-polyacrylamide gel electrophoresis (PAGE) (7.5%) (42). Proteins were visualized with silver stain (8). Western analysis using antibodies to SipA and SipB was performed with the Amersham Life Science ECL Western blotting detection system.
Enzyme assays.
β-Galactosidase assays were performed and values were calculated as previously described (48).
Sequence analysis.
The majority of the sequencing reactions were performed by Joan Strange (University of Montana, Missoula) and Bio S&T (Quebec, Quebec, Canada). For reactions done in the lab, Sequenase 2.0 (United States Biochemical) was used, and labeled fragments were separated in a 6% acrylamide gel (Sequagel-6; National Diagnostics). Sequence analyses (homologies, mapping, etc.) were performed with the Wisconsin Sequence Analysis Package (Genetics Computer Group, Inc.).
Nucleotide sequence accession number.
The nucleotide sequence of sigDE has been submitted to GenBank and assigned the accession no. AF021817.
RESULTS
Identification of invasion loci.
To find novel S. typhimurium invasion genes, two issues were addressed: (i) reidentification of known invasion genes and (ii) redundancy of Salmonella invasion systems (21). The ability to quickly map or clone mutations was necessary to eliminate regions that have previously been identified (i.e., SPI1). Tn10dTc was chosen because the Tcr (tetAR) marker could be exploited by rapid MudP22 transduction mapping (6, 45). This would allow us to eliminate mutants with insertions in regions under intense study, i.e., SPI1. In addition, Tn10dTc does not encode a transposase and therefore would be fairly stable upon insertion into the chromosome.
The possible existence of multiple invasion systems was another issue addressed. Since SPI1 mutants were still invasive in CHO cells, albeit not at wild-type levels (data not shown), and able to cause systemic disease in vivo, it was likely that at least one other invasion system might be present in Salmonella; single-mutation phenotypes in one invasion system might be masked by the presence of a second invasion system. Thus, we screened for novel invasion genes in an SPI1− mutant background. Wild-type S. typhimurium 14028s was mutagenized with Tn10dTc (41); Tn10dTc was transduced by P22 int into 14028s containing pNK2881, a plasmid which provides transposase for Tn10 in trans (41). Transductants were plated on minimal medium to eliminate auxotrophic mutants, thus reducing the number of mutants to be screened in the TCI assay. Twenty-five transductions were done, resulting in several hundred to more than 1,000 transductants per plate; transductants on each plate were pooled. Aliquots from these pools were used to prepare P22 int lysates, which were then used to transduce the Tn10dTc insertions into SVM169 (invA::Ωcat). Transductants were plated on minimal medium with tetracycline and chloramphenicol to select for Tn10dTc insertions and the invA::Ωcat mutation.
Approximately 2,500 Tcr transductants from the 25 independent transduction pools plated on minimal medium were screened in the TCI assay for reduced invasion into CHO tissue culture cells. The Tn10dTc insertions from putative mutants were separated from the invA::Ωcat mutation by transducing the Tcr marker into 14028s (wild type). Tcr transductants were purified and assayed in both CHO and HEp-2 (human laryngeal epithelium) tissue culture cells. Mutants which appeared reduced for invasion were also tested for motility in 0.25% LB agar; several motility mutants were found and omitted from further study. Two motile, prototrophic mutants, SVM167 and SVM168 (both invA+), were 5- to 10-fold less invasive than the wild type in both HEp-2 and CHO cell lines (Table 2). Both mutants were wild type for attachment to the tissue culture cells (data not shown). Chromosomal DNA was isolated from these mutants and analyzed by Southern hybridization using tetAR genes as a probe (data not shown). Based on these data, DNA fragments containing the junctions between the transposon and the chromosome were isolated. The junction sequences were cloned from each mutant and used to screen subgenomic libraries for the wild-type loci. Clones containing wild-type DNA which hybridized to junction DNA were sequenced.
TABLE 2.
Characterization of novel invasion mutants
| Strain | % Invasiona (% of wild-type level)
|
Map position | Homology to region of insertionc | |
|---|---|---|---|---|
| CHO | HEp-2 | |||
| 14028s (wild type) | 2.2 ± 0.2 (100) | 5.3 ± 0.3 (100) | ||
| SVM167 | 0.2 ± 0.2 (9) | 0.4 ± 0.02 (7.5) | ∼25.5b | Shigella ipgDE(F) |
| SVM168 | 0.5 ± 0.6 (22) | 1.8 ± 0.07 (34) | ∼78d | S. enteritidis IS1351 ORFB; Y. enterocolitica high-pathogenicity island ORFB |
Percentage of intracellular bacteria ± range after bacteria were allowed to invade during a 1-h incubation; duplicate assays performed. Values are representative of several assays done in this study.
Map (centisome) position of the Tn10dTc insertions was determined by MudP22 transduction mapping (SVM167). The MudP22 element in strain TT15240 (putA1019::MudP) cotransduced with the Tn10dTc insertion in SVM167.
Homologies are at the amino acid sequence level.
The insertion in SVM168 was mapped by using Hfr strains with origins of transfer at 76 and 78 min of the LT2 chromosome.
SVM168 appears to have an insertion at about centisome 78 in an IS3 element homologous to Salmonella enteritidis IS1351 (A. P. Burnens, direct submission to GenBank) and Yersinia enterocolitica high-pathogenicity island (HPI) IS1400 (9) elements. These insertion sequence elements contain open reading frame A (ORFA) and ORFB sequences, and SVM168 has an insertion in the ORFB homolog. The possibility that SVM168 maps to a putative third pathogenicity island, SPI3 (7), was tested. A marker in mgtCB of SPI3, which is located at centisome 82.5, could not be cotransduced with the mutation in SVM168, suggesting that the mutation in SVM168 was outside this region.
SVM167 was determined to have an insertion in a homolog of the Shigella flexneri and Shigella sonnei virulence plasmid encoded gene ipgE of the ipgDEF operon (1). In Shigella, ipgDEF was characterized due to its location between the ipa and mxi virulence plasmid loci (46, 62); however, this operon has no observable virulence function for Shigella either in in vitro invasion assays or in Serény tests in guinea pigs (1). The Salmonella operon contains homologs to ipgD and ipgE but not ipgF. Interestingly, a homolog of ipgF, iagB, has been identified downstream of hilA in SPI1 and in an unmapped location in Salmonella typhi (51). As in Shigella, iagB has no known virulence phenotype (4). Unlike the Shigella genes, the Salmonella homologs, sigD and -E, are not located between what are believed to be homologs of the ipa and mxi virulence genes at SPI1 (27); instead, sigDE maps between centisomes 24.5 and 25.5. The predicted amino acid sequence of sigD (62 kDa) is 59% similar and 41% identical to that of IpgD (60 kDa), and SigE (13 kDa) is 57% similar and 29% identical to IpgE (14 kDa) (Fig. 1). Recently, a sigD homolog (sopB) in Salmonella dublin was identified; however, a sigE homolog was not found (24). SopB did not appear to have a role in invasion in either in vitro or in vivo invasion assays in the S. dublin study.
FIG. 1.
Amino acid alignment of S. typhimurium SigD and SigE with S. flexneri IpgD and IpgE, respectively. SigD (62 kDa) is 59% similar and 41% identical to IpgD (60 kDa), and SigE (13 kDa) is 57% similar and 29% identical to IpgE (14 kDa). The Wisconsin sequence analysis package (Genetics Computer Group) was used to identify homologies to sigDE.
Complementation of the SVM167 invasion defect by sigDE.
The wild-type copy of sigDE on a 4.1-kb EcoRI fragment (defined by Southern analysis described above) was cloned into pHG329, a pBR322-based plasmid of medium copy number (66), creating pHH10 (Fig. 2). This plasmid was transformed into SVM167 but did not reproducibly restore invasion into tissue culture cells. It was possible that the invasion phenotype of SVM167 was affected by having the genes in a multicopy plasmid. A slightly smaller fragment containing sigDE was subcloned into pWKS30, a low-copy-number cloning vector (about six copies per cell) (70). The resulting plasmid, pHH20 (Fig. 2), was transformed into both the wild-type and sigE strains. Invasion by SVM167 harboring pHH20 increased fourfold and was similar to that of the wild type; however, 100% complementation was never observed (Table 3). This is often the case in complementation studies of invasion mutants (13, 14, 23, 26, 35). It is also possible that the insertion in sigE affects downstream genes. However, the nearest ORF is more than 500 bp downstream of sigE, and there is a putative transcriptional terminator between sigE and the downstream ORF.
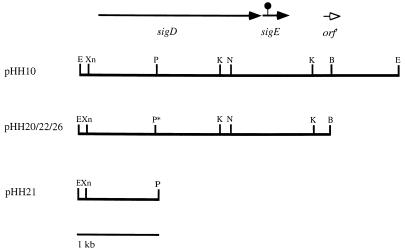
FIG. 2.
Restriction map of sigDE+ plasmids and the transcriptional reporter plasmid pHH21. pHH10 is a pHG329-based, medium-copy-number plasmid. pHH20 (pWKS30-based, low-copy-number plasmid) contains a 3.2-kb BamHI fragment from pHH10. The sigDE fragment from pHH20 was cloned into the pWKS30 sister cloning vector, pWSK29, to obtain a clone with the insert in the opposite orientation with respect to the T7 promoter. The coding regions of sigD and sigE are indicated by the closed arrows. The original Tn10dTc insertion in sigE is represented by the black circle. An uncharacterized partial ORF (orf′) is indicated by the open arrowhead. pHH26 is pHH22 with a deleted PstI (p*) site (see Materials and Methods). pHH21 contains an EcoRI/PstI fragment containing about 0.9 kb of sequence from sigDE fused to lacZYA. Restriction enzymes: B, BamHI; E, EcoRI; P, PstI; Xn, XmnI; K, KpnI; N, NdeI.
TABLE 3.
Complementation of SVM167
| Strain | CHO
|
HEp-2
|
||
|---|---|---|---|---|
| % Invasiona | % of wild-type level | % Invasion | % of wild-type level | |
| 14028s (wild type) | 4.8 ± 0.3 | 100 | 5.3 ± 0.3 | 100 |
| 14028s (pWKS30) | 2.3 ± 0.0 | 100 | 5.7 ± 0.8 | 100 |
| 14028s (pHH20) | 2.2 ± 0.05 | 100 | 3.3 ± 0.2 | 100 |
| 14028s (pHH26) | 4.5 ± 0.15 | 100 | 7.8 ± 0.1 | 100 |
| SVM167 (sigE::Tn10dTc) | 0.3 ± 0.07 | 6.3 | 0.53 ± 0.06 | 10 |
| SVM167 (pWKS30) | 0.2 ± 0.02 | 8.7 | 0.4 ± 0.07 | 7 |
| SVM167 (pHH20) | 0.8 ± 0.05 | 36 | 1.6 ± 0.3 | 49 |
| SVM167 (pHH26) | 1.5 ± 0.2 | 33 | 2.3 ± 0.02 | 35 |
Percentage of intracellular bacteria ± range after allowing bacteria to invade during a 1-h incubation; duplicate assays were performed. Values are representative of several assays done in this study.
T7 expression of sigDE.
To identify the protein products of sigDE, sigDE was introduced into a T7 expression system (68) to specifically express sigD and sigE. The clone used to complement invasion, pHH20, contained a T7 promoter but was in the wrong orientation relative to sigDE. Therefore, sigDE from pHH20 was directionally cloned downstream of the T7 promoter in pWSK29, the sister vector to pWKS30 (70) used to construct pHH20. The resulting plasmid, pHH22, was transformed into an E. coli strain harboring pGP1-2, which encodes T7 polymerase under the control of a temperature-sensitive λ repressor protein (68). Induction and expression of the plasmid-encoded proteins were done in the presence of rifampin and a mixture of [35S]methionine and [35S]cysteine (3, 68). One of the proteins expressed, most likely SigD, migrated at about 77 kDa, as observed in the culture supernatant gels (see below), strongly suggesting that it was the same protein (Fig. 3). SigD is predicted to have a molecular size of about 62 kDa, not the observed 77 kDa. This discrepancy in protein migration was apparently due to a departure from the typical bisacrylamide concentration used in our protein gels. When a standard SDS-polyacrylamide gel (30:0.8, acrylamide to bisacrylamide) (42) was used, SigD migrated more closely to the predicted size of 62 kDa (data not shown); however, in these gels, SigD was not well separated from other proteins. Fortuitously, the acrylamide gels used in this work had a 30:1.6 ratio of acrylamide to bisacrylamide, which optimally resolved SigD from other proteins.
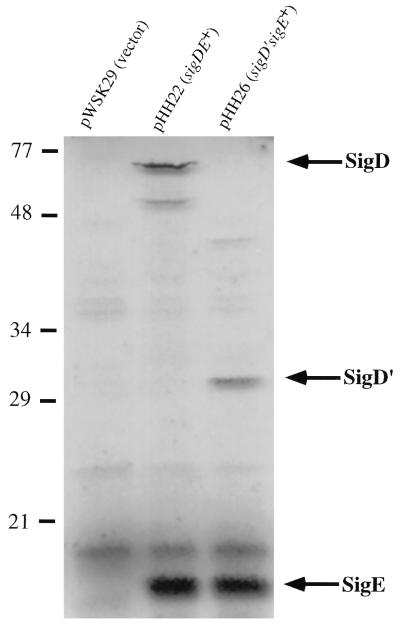
FIG. 3.
Expression of sigDE from a T7 promoter in E. coli. Expression of genes from pWSK29 (vector) (lane 1), pHH22 (sigDE+) (lane 2), and pHH26 (PstI site deleted from sigD) (lane 3) was done in DH5α harboring pGP1-2. pGP1-2 contains a T7 RNA polymerase gene under the control of the λ pL promoter and a gene which encodes a temperature-sensitive λ repressor protein (cI857). Proteins were labeled with [35S]methionine and [35S]cysteine after the addition of rifamipin. Sizes are indicated in kilodaltons.
To further demonstrate that the 77-kDa protein was SigD, a small deletion was made in sigD, resulting in a truncated protein, SigD′. pHH22 was digested with PstI (unique in sigD and pHH22), treated with Deep Vent DNA polymerase (New England Biolabs) to remove the 3′ overhangs, and ligated. This plasmid, pHH26, contains a 4-bp deletion in sigD, resulting in a frameshift mutation. SigD′ was expressed from pHH26 (Fig. 3); however, SigD′ migrated at about 29 kDa rather than the predicted 22 kDa, consistent with the observation that the full-length SigD (62 kDa, predicted size) appears to be larger in the higher-percentage bisacrylamide gels.
SigE was also synthesized in the T7 expression system and appeared to be about 13 kDa, as predicted by sequence analysis (Fig. 3). The SigE band was more intense than SigD in these gels; SigD is predicted to have 16 cysteines and methionines, while SigE is predicted to have only 5. Nucleotide sequence analysis revealed that sigE has a consensus Shine-Dalgarno ribosome-binding site whereas sigD has no conspicuous ribosome-binding site, suggesting that SigE may be translated more efficiently than SigD.
Analysis of culture supernatants.
IpgD of Shigella (1) and several proteins required for Salmonella invasion appear to be secreted (12, 13, 32, 34, 39, 40, 73), suggesting that SigD may also be secreted. Culture supernatant profiles of the sigE mutant and several control strains were examined. Supernatants from nonaerated (inducing conditions) (16) overnight cultures were harvested and filtered; proteins in the filtrates were TCA precipitated from wild-type, invA (SPI1−), sigE, and sigDE+ plasmid-carrying sigE strains. Proteins were separated by SDS-PAGE and visualized by silver staining (Fig. 4). The sigE mutant was missing one band at about 77 kDa. Interestingly, the invA mutant was also missing the 77-kDa protein in addition to the previously identified Sip/Ssp proteins, suggesting that under these conditions this 77-kDa protein is translocated by the SPI1 type III secretion system. In addition, absence of the 77-kDa protein did not significantly affect the secretion of the Sip/Ssp proteins. Presence of this protein in the supernatant was restored in the sigE mutant harboring pHH10 (sigDE+) (Fig. 4, lane 5). Consistent with these results, transposon insertions in the sigD coding region of pHH10 eliminated this protein from the supernatant (data not shown).
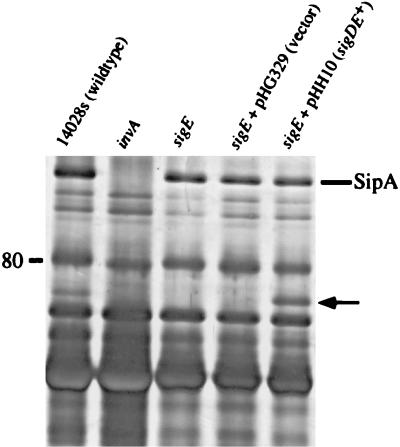
FIG. 4.
Supernatant proteins from wild-type (lane 1), SVM169 (invA SPI1−) (lane 2), SVM167 (sigE) (lane 3), SVM167 (pHG329) (lane 4), and SVM167 (pHH10) (lane 5) S. typhimurium strains. Cultures were grown without aeration overnight (18 h). Culture supernatants were TCA (10%, final concentration) precipitated and separated by SDS-PAGE (7.5% gel). Proteins were fixed and visualized silver stain. Position of the size marker is shown in kilodaltons on the left. The location of SipA was determined by Western analysis using polyclonal antibodies to SipA (data not shown). The putative location of SigD is indicated by the arrow.
Further support for the hypothesis that the 77-kDa protein is SigD came from the analysis of a sigD mutant. The sigD allele encoding the truncated SigD from pHH26 was exchanged with the wild-type allele on the chromosome. SVM255 was equally defective for invasion into both CHO and HEp-2 cells as the sigE mutant SVM167 (data not shown). T7 expression data (Fig. 3) suggests that SigE may be expressed in wild-type amounts in the sigD mutant. The sigD disruption strain lacked the same protein from culture supernatants as SVM167 (Fig. 5). As with SVM167, transformation of SVM255 with a plasmid encoding sigDE restored the 77-kDa protein to culture supernatants (Fig. 5). Together, these data suggest that the 77-kDa protein missing from the supernatant of SVM167 is SigD.
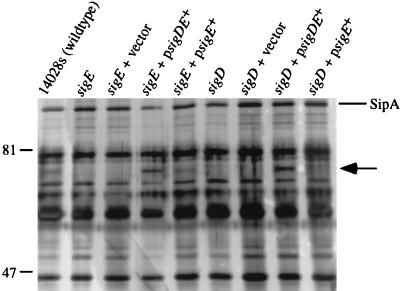
FIG. 5.
Complementation of SigD secretion by SVM167 (sigE) and SVM255 (sigD). Cultures were grown and proteins were precipitated as described for Fig. 4. Shown are results for supernatant proteins from wild-type (lane 1), SVM167 (sigE) (lane 2), SVM167 (pWKS30) (vector) (lane 3), SVM167 (pHH20) (sigDE+) (lane 4), SVM167 (pHH26) (sigE+) (lane 5), SVM255 (sigD) (lane 6), SVM255 (pWKS30) (lane 7), SVM255 (pHH20) (lane 8), and SVM255 (pHH26) (lane 9). Positions of size standards are indicated in kilodaltons on the left. The position of SipA is indicated on the right. The putative location of SigD is indicated by the arrow.
The possibility that SigD is secreted by the SPI2-encoded type III secretion system was also tested (31, 53, 64). A mutation in one of the proposed SPI2 secretion apparatus genes, ssaT, has been shown to have reduced levels of SipC in its supernatant (31). SipC is a SPI1-secreted protein and is thought to be at least one of the effectors for invasion (12, 40). Culture supernatant proteins from the ssaT mutant were harvested and visualized as described above. Wild-type amounts of SigD were still present in the ssaT mutant (data not shown). Moreover, wild-type amounts of SipC were also present. It is notable that Hensel and coworkers also reported that in a small proportion of their experiments, SipC was still found (31). Thus, under these conditions the SPI2 type III secretion system was not required for SigD secretion.
Complementation of SVM167 with sigE+.
In the T7 expression system, SigE was produced by E. coli harboring pHH26 (sigD′sigE+) in amounts similar to that produced by the pHH22 strain (sigDE+). If SVM167 could still produce but not secrete SigD, it was possible that pHH26 could restore secretion of SigD and complement the invasion defect. To test this, pHH26 was transformed into SVM167 and 14028s, and the resulting strains were assayed for invasion in both CHO and HEp-2 cell lines. pHH26 was able to complement the sigE mutation in SVM167 to the same degree as pHH20 (Table 3).
To determine if pHH26 could restore the secretion of wild-type SigD (chromosomal copy of sigD is wild type) or secrete a truncated SigD′, proteins from culture supernatants were precipitated and examined on an SDS-polyacrylamide gel as previously described. Secretion of SigD from SVM167 carrying pHH26 was restored (Fig. 5), suggesting that SigD was still being produced but retained by the mutant. Furthermore, pHH26 did not restore secretion of SigD in the sigD mutant (Fig. 5). It is notable that in several independent assays, there appeared to be less SigD in supernatant preparations from SVM167 and SVM255 that harbored pHH26 than in samples from a strain which contained pHH10 (medium-copy-number clone of sigDE+) or pHH20 (low-copy-number clone of sigDE+). A novel, truncated form of SigD was never observed in these gels.
sirA-dependent regulation of sigDE.
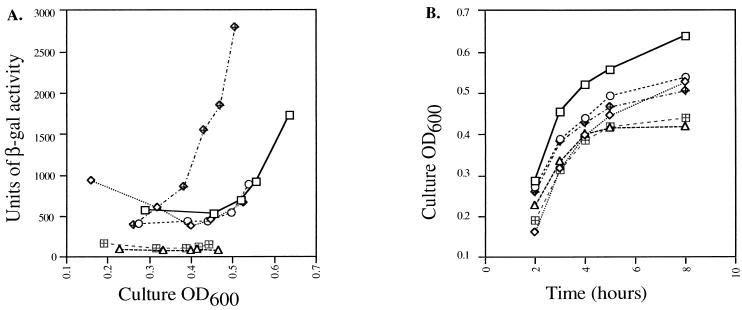
It is clear that sigDE plays a role in invasion and that SigD may be secreted through the SPI1 secretion system. Many of the known invasion genes at SPI1 appear to be coordinately regulated. Therefore, the possibility that expression of sigDE was also coordinated with that of other invasion genes was considered. The current model of SPI1 invasion gene regulation suggests that HilA activates the expression of several invasion genes, including that of another regulator, InvF (5, 38). Transcription of hilA is activated by SirA (35) but repressed by PhoP, a response regulator of virulence and housekeeping genes (5, 25, 49). To determine if any of the known invasion gene regulators affected expression of sigDE, a plasmid with a lacZYA fusion to sigD, pHH21, was made. β-Galactosidase activity was measured in isogenic hilA (4), sirA (35), and phoP (50) mutants containing pHH21. β-Galactosidase activity was also measured in SVM167 containing the reporter plasmid (data not shown). Overnight cultures were subcultured 1:100, and samples were assayed for β-galactosidase activity over a time course from 2 to 24 h. Expression from the sigDE promoter region was clearly activated in late log to early stationary phase in both anaerobic (inducing condition) (Fig. 6A) and aerobic (data not shown) cultures of all of the strains except the sirA mutant, suggesting that sigDE expression is dependent on SirA. When the sirA strain was transformed with a low-copy-number clone containing only sirA (35), activation of lacZYA expression was restored above wild-type levels. Curiously, in nonaerated cultures, the hilA, sirA, and phoP mutants reached nearly their maximum optical densities at 600 nm at about 8 h, unlike wild-type S. typhimurium or E. coli (Fig. 6B). Nevertheless, induction of β-galactosidase was consistently observed at about the same cell culture density or growth phase and at the same amounts in the wild type and the hilA and phoP mutants. However, due to the multicopy nature of the reporter plasmid used, it is conceivable that HilA and PhoP have subtle effects on regulation that were not detected under the conditions tested.
FIG. 6.
Effect of regulatory mutations on sigD-lacZYA expression. (A) lacZYA expression from sigD in plasmid pHH21 plotted versus culture density. Expression from this plasmid was measured in strains 14028s (wild type) (squares), SVM214 (hilA) (diamonds), CS015 (phoP) (circles), CJ010 (sirA) (triangles), CJ010 (pWSK129) (vector) (hatched squares), and CJ010 (pHH25) (sirA+) (hatched diamonds). β-gal, β-galactosidase. (B) Growth curves of cultures in panel A. Overnight cultures were subcultured 1:100 and grown without aeration; 1-ml samples were taken to determine cell optical density at 600 nm (OD600) at time points between 2 and 24 h. The same samples were then used to measure β-galactosidase activity in these strains. The 24-h time point was omitted from these graphs since the mutant culture densities were virtually saturated by 8 h (β-galactosidase activity was also maximal for all strains at this time).
The possibility that the Salmonella SirA can activate transcription of the sigD-lacZYA construct in E. coli was tested. SirA, encoded on a low-copy-number plasmid, did not activate transcription of sigDE in E. coli. This result was not unexpected because E. coli has a SirA homolog, UvrY (96% identity to SirA), that was also unable to activate transcription (data not shown). This finding suggests that other factors required for regulation of sigDE might be absent in E. coli. However, it is not known whether the E. coli strain used for this experiment (DH5α) actually expresses UvrY (35, 52, 63).
DISCUSSION
Only 2 insertion mutants of 2,500 were found to retain motility and be reduced for invasion in this study. Of course, because the SPI1 invasion system had been inactivated, finding mutations in this region of the chromosome was not anticipated. In this screen, a novel invasion locus homologous to the Shigella ipgDEF operon was identified by transposon mutagenesis and direct screening for invasion mutants in the TCI assay. A Tn10dTc insertion was found in the S. typhimurium homolog of ipgE which was not well characterized in the Shigella study (1). Complementation for invasion of the Salmonella mutant was accomplished by both sigDE+ and sigE+ in trans. A mutation in sigD was also introduced onto the S. typhimurium chromosome and found to have the same phenotype as the sigE mutant. Failure to complement a sigD mutant with sigE alone suggests that both SigD and SigE are required. In Shigella, IpgD appeared to be secreted into culture supernatants, and an IpgF-PhoA fusion protein was found in the periplasmic space (1). Mutations in either of these genes did not affect invasion or virulence in the guinea pig model of infection; it was proposed that ipgDEF function might be observed only within the natural primate host (1). In S. typhimurium, SigD, like the IpgD homolog, appeared to be in culture supernatants. Unlike in Shigella, an ipgF homolog was not found in this region, although a homolog, iagB, has been found in SPI1 of S. typhimurium (4) and in an unmapped region of S. typhi (51). Recently, a homolog of SigD, SopB (Salmonella outer protein B; 98% identity) was identified in S. dublin by reverse genetics (24). In S. dublin, SopB appears to be translocated into eucaryotic cells and to promote an inflammatory response in infected calf ileal loops. Interestingly, SopB did not appear to be necessary for the invasion of S. dublin into HeLa cells or calf intestinal cells. It will be necessary to assess the role of sigDE in S. typhimurium pathogenesis in the mouse model of infection and to determine whether the primary function of SigD and SigE is for invasion or the stimulation of the immune response.
SigD appears to be secreted through the SPI1-encoded type III secretion apparatus and requires SigE for this process to occur. Several other proteins secreted by the SPI1-encoded apparatus that are required for Salmonella invasion into cultured epithelial cells have been identified. It has been proposed that these proteins act by stimulating endocytosis of the bacteria by the host cell (bacterium-mediated endocytosis or BME) (21, 57). How these proteins stimulate the host cells has yet to be determined, but it seems that many proteins may be involved in this process.
Although SigE has not been detected in our analysis of secreted proteins, it is possible that it is secreted, perhaps in conjunction with SigD, as IpaB and IpaC are cosecreted from Shigella (47). Another possibility for the role of SigE is that it may target SigD to the SPI1 type III secretion apparatus. In a recent study of Y. enterocolitica, YopE was shown to be secreted through a single type III secretion apparatus via one of two independent signals; one involves the recognition of the first 15 amino acids of YopE by the secretion system, and the second is the YopE-SycE complex, which is proposed to be recognized by or targeted to the type III secretion machinery (11).
It is unlikely that SigE is a component of the SPI1 secretion apparatus since the absence of SigE does not affect secretion of the Sip/Ssp proteins. It is possible, however, that SigE acts as a chaperone for stability or secretion of SigD, as precedence for this has been found in other type III secretion systems in pathogenic bacteria. In one scenario, SigE may ensure the stability of the SigD polypeptide prior to secretion; a sigE mutation would in this case result in the rapid degradation of SigD in the cytoplasm. In Yersinia, SycE (YerA) appears to prevent the degradation of YopE, an antiphagocytic cytotoxin, prior to secretion (59, 60, 71). Similarly, the IpaB and IpaC invasion proteins in Shigella are rapidly degraded in the absence of the IpgC chaperone (47). In Shigella and other organisms, it has been proposed that inappropriate association of proteins such as IpaB with IpaC within the cytoplasmic compartment might result in unstable structures; these would be subject to proteolysis. IpgC, therefore, was proposed to prevent association of IpaB with IpaC prior to secretion, thereby protecting them from degradation (47). In addition, IpaB and IpaC individually could possibly misfold in the cytoplasm in the absence of IpgC to produce aberrant structures subject to proteolysis.
SigE has some similarities to these and other putative chaperones in that it is small (13 kDa), has a predicted acidic isoelectric point (pI = 3.99), and may have an amphipathic alpha helix at its C terminus (72). In addition, T7 expression of sigDE suggested that SigE is translated in higher amounts than SigD. It has been proposed that in Yersinia, SycE exists in the bacterial cytoplasm as a homodimer and binds to YopE in this form (71). Perhaps SigE also exists as a multimer and must therefore be translated in higher amounts in order to accommodate SigD molecules.
Several studies have reported that mutations in the SPI1 secretion apparatus result in the absence of up to 25 proteins in culture supernatants (34, 39, 40). Only a handful of these proteins has been characterized and found to be encoded in SPI1. An unique aspect of the sigDE locus is that although the secretion of SigD relies on proteins encoded in SPI1 at centisome 63, it is encoded near centisome 25. It is certainly possible, if not probable, that there are other invasion or virulence factors whose genes, like sigDE, map to locations outside of SPI1 and are secreted.
Another feature of sigDE is that its regulation is dependent on SirA, a regulator of SPI1 genes. SirA has at least two homologs, UvrY and GacA from E. coli and Pseudomonas sp., respectively (35, 52, 58, 63). Both uvrY and gacA are encoded immediately upstream of uvrC or its homolog and appear to be transcribed with uvrC as a single message in their respective organisms. UvrC is believed to have an excision repair function, and mutants have an increased sensitivity to UV light damage (52, 63). The function of UvrY in E. coli has yet to be determined. In Pseudomonas, GacA appears to be a response regulator for the LemA sensor kinase (58). Mutations in either gacA or lemA result in a decreased pathology in a plant model of infection, suggesting decreased expression of virulence factors in these strains. In Salmonella, a sensor kinase partner for SirA has not yet been identified. However, previous work has identified at least two different loci which complement a sirA mutation (35). Although SirA is believed to be a DNA binding protein based on sequence analysis, it is possible that SirA itself does not bind the sigDE promoter. In Salmonella, it appears that several regulators activate or repress expression of the SPI1-encoded genes, among them genes encoding additional regulators. It is possible that SirA may activate transcription of another regulator which in turn regulates sigDE expression. This may indeed be the case since the E. coli SirA homolog (UvrY) could not activate transcription of sigDE in E. coli. This observation suggests that there is a factor in Salmonella but absent in E. coli that is necessary for sigDE expression; however, it is not known whether UvrY is expressed in the E. coli strain used in this study.
Why is the phenotype of a mutation in a gene whose product is secreted by the SPI1 secretion system more dramatic when in an SPI1− background? The original mutation was identified by screening a pool of Tn10dTc insertions in an SPI1− background. The reduced invasion phenotype of the double mutant (invA sigE) compared to invA alone in both CHO and HEp-2 cell lines was consistently reproducible early in this study. It is conceivable that there exist other invasion systems that require the expression of sigD and/or sigE that are independent of SPI1. SigD secretion through another secretion apparatus could be triggered in a SPI1 mutant under conditions that we have not been able to reproduce. The functions of the identified secreted proteins of S. typhimurium have not been characterized, but it is tempting to suggest that specific proteins may be required for invasion of particular cell types or tissues. Perhaps certain invasion proteins are produced or secreted only upon contact with a specific target cell type. It has been observed that SipC, a SPI1-secreted protein, was absent from culture supernatants of certain SPI2 mutants, suggesting that an interaction between the two type III secretion systems might exist (31). Although SigD secretion from bacteria cultured in the laboratory is not affected by a defective SPI2 apparatus, it is not inconceivable that SPI2 or yet another type III secretion apparatus may exist that secretes SigD from bacteria associated with host cells.
ACKNOWLEDGMENTS
We thank Andrew Darwin and Glenn Young for critical review of the manuscript. We also thank Jorge Galan, Eduardo Groisman, David Holden, Sidney Kushner, Catherine Lee, Samuel Miller, and Kenneth Sanderson for strains used in this work. Jorge Galan also graciously provided antibodies to SipA and InvJ. We especially thank Stanley Maloy and Valley Stewart for strains and technical advice.
This work was supported by the National Institutes of Health grant AI01230 and AI31403 to V.L.M.
REFERENCES
- 1.Allaoui A, Ménard R, Sansonetti P J, Parsot C. Characterization of the Shigella flexneri ipgD and ipgF genes, which are located in the proximal part of the mxi locus. Infect Immun. 1993;61:1707–1714. doi: 10.1128/iai.61.5.1707-1714.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Alpuche-Arande C M, Swanson J A, Loomis W P, Miller S I. Salmonella typhimurium activates virulence gene transcription within acidified macrophage phagosomes. Proc Natl Acad Sci USA. 1992;89:10079–10083. doi: 10.1073/pnas.89.21.10079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ausubel F M, Brent R, Kingston R E, Moore D D, Seidman J G, Smith J A, Struhl K, editors. Short protocols in molecular biology. 2nd ed. New York, N.Y: Greene Publishing Associates and John Wiley & Sons; 1992. [Google Scholar]
- 4.Bajaj V, Hwang C, Lee C A. hilA is a novel ompR/toxR family member that activates the expression of Salmonella typhimurium invasion genes. Mol Microbiol. 1995;18:715–727. doi: 10.1111/j.1365-2958.1995.mmi_18040715.x. [DOI] [PubMed] [Google Scholar]
- 5.Bajaj V, Lucas R L, Hwang C, Lee C A. Co-ordinate regulation of Salmonella typhimurium invasion genes by environmental and regulatory factors is mediated by control of hilA expression. Mol Microbiol. 1996;22:703–714. doi: 10.1046/j.1365-2958.1996.d01-1718.x. [DOI] [PubMed] [Google Scholar]
- 6.Benson N R, Goldman B S. Rapid mapping in Salmonella typhimurium with Mud-P22 prophages. J Bacteriol. 1992;174:1673–1681. doi: 10.1128/jb.174.5.1673-1681.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Blanc-Potard A, Groisman E A. The Salmonella selC locus contains a pathogenicity island mediating intramacrophage survival. EMBO J. 1997;16:5376–5385. doi: 10.1093/emboj/16.17.5376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Blum H, Beier H, Gross H J. Improved silver staining of plant proteins, RNA and DNA in polyacrylamide gels. Electrophoresis. 1987;8:93–99. [Google Scholar]
- 9.Carniel E, Guilvout I, Prentice M. Characterization of a large chromosomal “high-pathogenicity island” in biotype 1B Yersinia enterocolitica. J Bacteriol. 1996;178:6743–6751. doi: 10.1128/jb.178.23.6743-6751.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Carter P B, Collins F M. The route of enteric infection in normal mice. J Exp Med. 1974;139:1189–1203. doi: 10.1084/jem.139.5.1189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Cheng L W, Anderson D M, Schneewind O. Two independent type III secretion mechanisms for YopE in Yersinia enterocolitica. Mol Microbiol. 1997;24:757–765. doi: 10.1046/j.1365-2958.1997.3831750.x. [DOI] [PubMed] [Google Scholar]
- 12.Collazo C M, Galan J E. The invasion-associated type III system of Salmonella typhimurium directs the translocation of the Sip proteins into the host cell. Mol Microbiol. 1997;24:747–756. doi: 10.1046/j.1365-2958.1997.3781740.x. [DOI] [PubMed] [Google Scholar]
- 13.Collazo C M, Zierler M K, Galan J E. Functional analysis of the Salmonella typhimurium invasion genes invI and invJ and identification of a target of the protein secretion apparatus encoded in the inv locus. Mol Microbiol. 1995;15:25–38. doi: 10.1111/j.1365-2958.1995.tb02218.x. [DOI] [PubMed] [Google Scholar]
- 14.Eichelberg K, Ginocchio C C, Galan J E. Molecular and functional characterization of the Salmonella typhimurium invasion genes invB and invC: homology of InvC to the F0F1 ATPase family of proteins. J Bacteriol. 1994;176:4501–4510. doi: 10.1128/jb.176.15.4501-4510.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Elliot T, Roth J R. Characterization of Tn10d-Cm: a transposition-defective Tn10 specifying chloramphenicol resistance. Mol Gen Genet. 1988;213:332–337. doi: 10.1007/BF00339599. [DOI] [PubMed] [Google Scholar]
- 16.Ernst R K, Dombroski D M, Merrick J M. Anaerobiosis, type 1 fimbriae, and growth phase are factors that affect invasion of HEp-2 cells by Salmonella typhimurium. Infect Immun. 1990;58:2014–2016. doi: 10.1128/iai.58.6.2014-2016.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Falkow S, Isberg R R, Portnoy D A. The interaction of bacteria with mammalian cells. Annu Rev Cell Biol. 1992;8:333–363. doi: 10.1146/annurev.cb.08.110192.002001. [DOI] [PubMed] [Google Scholar]
- 18.Finlay B B, Falkow S. A comparison of microbial invasion strategies of Salmonella, Shigella and Yersinia species. Bacteria-Host Cell Interact. 1988;64:227–243. [Google Scholar]
- 19.Finlay B B, Ruschkowski S, Dedhar S. Cytoskeletal rearrangements accompanying Salmonella entry into epithelial cells. J Cell Sci. 1991;99:283–296. doi: 10.1242/jcs.99.2.283. [DOI] [PubMed] [Google Scholar]
- 20.Francis C L, Ryan T A, Jones B D, Smith S J, Falkow S. Ruffles induced by Salmonella and other stimuli direct macropinocytosis of bacteria. Nature. 1993;364:639–642. doi: 10.1038/364639a0. [DOI] [PubMed] [Google Scholar]
- 21.Galan J E. Molecular genetic bases of Salmonella entry into host cells. Mol Microbiol. 1996;20:263–271. doi: 10.1111/j.1365-2958.1996.tb02615.x. [DOI] [PubMed] [Google Scholar]
- 22.Galan J E, Curtiss R., III Cloning and molecular characterization of genes whose products allow Salmonella typhimurium to penetrate tissue culture cells. Proc Natl Acad Sci USA. 1989;86:6383–6387. doi: 10.1073/pnas.86.16.6383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Galan J E, Ginocchio C, Costeas P. Molecular and functional characterization of the Salmonella invasion gene invA: homology of InvA to members of a new protein family. J Bacteriol. 1992;174:4338–4349. doi: 10.1128/jb.174.13.4338-4349.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Galyov E E, Wood M W, Rosqvist R, Mullan P B, Watson P R, Hedges S, Wallis T S. A secreted effector protein of Salmonella dublin is translocated into eucaryotic cells and mediates inflammation and fluid secretion in infected ileal mucosa. Mol Microbiol. 1997;25:903–912. doi: 10.1111/j.1365-2958.1997.mmi525.x. [DOI] [PubMed] [Google Scholar]
- 25.Garcia-Vescovi E, Soncini F C, Groisman E G. Mg2+ as an extracellular signal: environmental regulation of Salmonella virulence. Cell. 1996;84:165–174. doi: 10.1016/s0092-8674(00)81003-x. [DOI] [PubMed] [Google Scholar]
- 26.Ginocchio C, Pace J, Galan J E. Identification and molecular characterization of a Salmonella typhimurium gene involved in triggering the internalization of salmonellae into cultured epithelial cells. Proc Natl Acad Sci USA. 1992;89:5976–5980. doi: 10.1073/pnas.89.13.5976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Groisman E A, Ochman H. Cognate gene clusters govern invasion of host epithelial cells by Salmonella typhimurium and Shigella flexneri. EMBO J. 1993;12:3779–3787. doi: 10.1002/j.1460-2075.1993.tb06056.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Haas R, Kahrs A F, Facius D, Allmeier H, Schmitt R, Meyer T F. TnMax—a versatile mini-transposon for the analysis of cloned genes and shuttle mutagenesis. Gene. 1993;130:23–31. doi: 10.1016/0378-1119(93)90342-z. [DOI] [PubMed] [Google Scholar]
- 29.Hamilton C M, Aldea M, Washburn B K, Babitzke P, Kushner S R. New method for generating deletions and gene replacements in Escherichia coli. J Bacteriol. 1989;171:4617–4622. doi: 10.1128/jb.171.9.4617-4622.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Hensel M, Shea J E, Gleeson C, Jones M D, Dalton E, Holden D W. Simultaneous identification of bacterial virulence genes by negative selection. Science. 1995;269:400–403. doi: 10.1126/science.7618105. [DOI] [PubMed] [Google Scholar]
- 31.Hensel M, Shea J E, Raupach B, Monack D, Falkow S, Gleeson C, Kubo T, Holden D W. Functional analysis of ssaJ and the ssaK/U operon, 13 genes encoding components of the type III secretion apparatus of Salmonella pathogenicity island 2. Mol Microbiol. 1997;24:155–167. doi: 10.1046/j.1365-2958.1997.3271699.x. [DOI] [PubMed] [Google Scholar]
- 32.Hermant D, Ménard R, Arricau N, Parsot C, Popoff M Y. Functional conservation of the Salmonella and Shigella effectors of entry into epithelial cells. Mol Microbiol. 1995;17:781–789. doi: 10.1111/j.1365-2958.1995.mmi_17040781.x. [DOI] [PubMed] [Google Scholar]
- 33.Hohmann A W, Schmidt G, Rowley D. Intestinal colonization and virulence of Salmonella in mice. Infect Immun. 1978;22:763–770. doi: 10.1128/iai.22.3.763-770.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Hueck C J, Hantman M J, Bajaj V, Johnston C, Lee C A, Miller S I. Salmonella typhimurium secreted invasion determinants are homologous to Shigella Ipa proteins. Mol Microbiol. 1995;18:479–490. doi: 10.1111/j.1365-2958.1995.mmi_18030479.x. [DOI] [PubMed] [Google Scholar]
- 35.Johnston C, Pegues D A, Hueck C J, Lee C A, Miller S I. Transcriptional activation of Salmonella typhimurium invasion genes by a member of the phosphorylated response-regulator superfamily. Mol Microbiol. 1996;22:715–727. doi: 10.1046/j.1365-2958.1996.d01-1719.x. [DOI] [PubMed] [Google Scholar]
- 36.Jones B D, Falkow S. Identification and characterization of a Salmonella typhimurium oxygen-regulated gene required for bacterial internalization. Infect Immun. 1994;62:3745–3752. doi: 10.1128/iai.62.9.3745-3752.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Jones B D, Ghori N, Falkow S. Salmonella typhimurium initiates murine infection by penetrating and destroying the M cells of the Peyer’s patches. J Exp Med. 1994;180:15–23. doi: 10.1084/jem.180.1.15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kaniga K, Bossio J C, Galan J E. The Salmonella typhimurium invasion genes invF and invG encode homologues of the AraC and PulD family of proteins. Mol Microbiol. 1994;13:555–568. doi: 10.1111/j.1365-2958.1994.tb00450.x. [DOI] [PubMed] [Google Scholar]
- 39.Kaniga K, Trollinger D, Galan J E. Identification of two targets of the type III protein secretion system encoded by the inv and spa loci of Salmonella typhimurium that have homology to the Shigella IpaD and IpaA proteins. J Bacteriol. 1995;177:7078–7085. doi: 10.1128/jb.177.24.7078-7085.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Kaniga K, Tucker S, Trollinger D, Galan J E. Homologs of the Shigella IpaB and IpaC invasins are required for Salmonella typhimurium entry into cultured epithelial cells. J Bacteriol. 1995;177:3965–3971. doi: 10.1128/jb.177.14.3965-3971.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kleckner N, Bender J, Gottesman S. Uses of transposons with emphasis on Tn10. Methods Enzymol. 1991;204:139–180. doi: 10.1016/0076-6879(91)04009-d. [DOI] [PubMed] [Google Scholar]
- 42.Laemmli U K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970;227:680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- 43.Lee C A. Type III secretion systems: machines to deliver bacterial proteins into eukaryotic cells? Trends Microbiol. 1997;5:148–156. doi: 10.1016/S0966-842X(97)01029-9. [DOI] [PubMed] [Google Scholar]
- 44.Lee C A, Jones B D, Falkow S. Identification of a Salmonella typhimurium invasion locus by selection for hyperinvasive mutants. Proc Natl Acad Sci USA. 1992;89:1847–1851. doi: 10.1073/pnas.89.5.1847. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Maloy S R, Stewart V J, Taylor R K. Genetic analysis of pathogenic bacteria. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory Press; 1996. [Google Scholar]
- 46.Maurelli A T, Baudry B, d’Hauteville H, Hale T L, Sansonnetti P J. Cloning of plasmid DNA sequences involved in invasion of HeLa cells by Shigella flexneri. Infect Immun. 1985;49:164–171. doi: 10.1128/iai.49.1.164-171.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Ménard R, Sansonetti P, Parsot C, Vasselon T. Extracellular association and cytoplasmic partitioning of the IpaB and IpaC invasins of S. flexneri. Cell. 1994;79:515–525. doi: 10.1016/0092-8674(94)90260-7. [DOI] [PubMed] [Google Scholar]
- 48.Miller J H. Experiments in molecular genetics. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory Press; 1972. [Google Scholar]
- 49.Miller S I, Kukral A M, Mekalanos J J. A two-component regulatory system (phoP phoQ) controls Salmonella typhimurium virulence. Proc Natl Acad Sci USA. 1989;86:5054–5058. doi: 10.1073/pnas.86.13.5054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Miller S I, Mekalanos J J. Constitutive expression of the phoP regulon attenuates Salmonella virulence and survival within macrophages. J Bacteriol. 1990;172:2485–2490. doi: 10.1128/jb.172.5.2485-2490.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Miras I, Hermant D, Arricau N, Popoff M Y. Nucleotide sequence of iagA and iagB genes involved in invasion of HeLa cells by Salmonella enterica subsp. enterica ser. Typhi. Res Microbiol. 1995;146:17–20. doi: 10.1016/0923-2508(96)80267-1. [DOI] [PubMed] [Google Scholar]
- 52.Moolenar G F, van Sluis C A, Backendorf C, van de Putte P. Regulation of the Escherichia coli excision repair gene uvrC. Overlap between the uvrC structural gene and the region coding for a 24 kD protein. Nucleic Acids Res. 1987;15:4273–4289. doi: 10.1093/nar/15.10.4273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Ochman H, Soncini F C, Solomon F, Groisman E A. Identification of a pathogenicity island required for Salmonella survival in host cells. Proc Natl Acad Sci USA. 1996;93:7800–7804. doi: 10.1073/pnas.93.15.7800. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Ozawa A, Goto J, Ito Y, Shibata H. Histopathological and biochemical responses of germfree and conventional mice with Salmonella infection. In: Heneghan J B, editor. Germfree research, biological effect of gnotobiotic environments. New York, N.Y: Academic Press; 1973. p. 325. [Google Scholar]
- 55.Pegues D A, Hantman M J, Behlau I, Miller S I. PhoP/PhoQ transcriptional repression of Salmonella typhimurium invasion genes: evidence for a role in protein secretion. Mol Microbiol. 1995;17:169–181. doi: 10.1111/j.1365-2958.1995.mmi_17010169.x. [DOI] [PubMed] [Google Scholar]
- 56.Penheiter K L, Mathur N, Giles D, Fahlen T, Jones B D. Non-invasive Salmonella typhimurium mutants are avirulent because of an inability to enter and destroy M cells of ileal Peyer’s patches. Mol Microbiol. 1997;24:697–709. doi: 10.1046/j.1365-2958.1997.3741745.x. [DOI] [PubMed] [Google Scholar]
- 57.Portnoy D A, Smith G A. Devious devices of Salmonella. Nature. 1992;357:536–537. doi: 10.1038/357536a0. [DOI] [PubMed] [Google Scholar]
- 58.Rich J J, Kinscherf T G, Kitten T, Willis D K. Genetic evidence that the gacA gene encodes the cognate response regulator for the lemA sensor in Pseudomonas syringae. J Bacteriol. 1994;176:7468–7475. doi: 10.1128/jb.176.24.7468-7475.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Rosqvist R, Forsberg A, Wolf-Watz H. Intracellular targetting of the Yersinia YopE cytotoxin in mammalian cells induces actin microfilament disruption. Infect Immun. 1991;59:4562–4569. doi: 10.1128/iai.59.12.4562-4569.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Rosqvist R, Håkansson S, Forsberg A, Wolf-Watz H. Functional conservation of the secretion and translocation machinery for virulence proteins of yersiniae, salmonellae and shigellae. EMBO J. 1995;14:4187–4195. doi: 10.1002/j.1460-2075.1995.tb00092.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Sambrook J, Fritsch E F, Maniatis T. Molecular cloning: a laboratory manual. 2nd ed. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory Press; 1989. [Google Scholar]
- 62.Sasakawa C, Kamata K, Sakai T, Makino S, Yamada M, Okada N, Yoshikawa M. Virulence-associated genetic regions comprising 31 kilobases of the 230-kilobase plasmid in Shigella flexneri 2a. J Bacteriol. 1988;170:2480–2484. doi: 10.1128/jb.170.6.2480-2484.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Sharma S, Stark T F, Beattie W G, Moses R E. Multiple control elements for the uvrC gene unit of Escherichia coli. Nucleic Acids Res. 1986;14:2301–2318. doi: 10.1093/nar/14.5.2301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Shea J E, Hensel M, Gleeson C, Holden D W. Identification of a virulence locus encoding a second type III secretion system in Salmonella typhimurium. Proc Natl Acad Sci USA. 1996;93:2593–2597. doi: 10.1073/pnas.93.6.2593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Simons R W, Houman F, Kleckner N. Improved single and multicopy lac-based cloning vectors for protein and operon fusions. Gene. 1987;53:85–96. doi: 10.1016/0378-1119(87)90095-3. [DOI] [PubMed] [Google Scholar]
- 66.Stewart G S A B, Lubinsky-Mink S, Jackson C G, Cassel A, Kuhn J. pHG165: a pBR322 copy number derivative of pUC8 for cloning and expression. Plasmid. 1986;15:172–181. doi: 10.1016/0147-619x(86)90035-1. [DOI] [PubMed] [Google Scholar]
- 67.Stone B J, Garcia C M, Badger J L, Hassett T, Smith R I F, Miller V L. Identification of novel loci affecting entry of Salmonella enteritidis into eukaryotic cells. J Bacteriol. 1992;174:3945–3952. doi: 10.1128/jb.174.12.3945-3952.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Tabor S, Richardson C C. A bacteriophage T7 RNA polymerase/promoter system for controlled exclusive expression of specific genes. Proc Natl Acad Sci USA. 1985;82:1074–1078. doi: 10.1073/pnas.82.4.1074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Takeuchi A. Electron microscope studies of experimental Salmonella infection. Am J Pathol. 1967;50:109–119. [PMC free article] [PubMed] [Google Scholar]
- 70.Wang R F, Kushner S R. Construction of versatile low-copy-number vectors for cloning, sequencing and gene expression in Escherichia coli. Gene. 1991;100:195–199. [PubMed] [Google Scholar]
- 71.Wattiau P, Cornelis G R. SycE, a chaperone-like protein of Yersinia enterocolitica involved in the secretion of YopE. Mol Microbiol. 1993;8:123–131. doi: 10.1111/j.1365-2958.1993.tb01209.x. [DOI] [PubMed] [Google Scholar]
- 72.Wattiau P, Woestyn S, Cornelis G R. MicroReview: customized secretion chaperones in pathogenic bacteria. Mol Microbiol. 1996;20:255–262. doi: 10.1111/j.1365-2958.1996.tb02614.x. [DOI] [PubMed] [Google Scholar]
- 73.Wood M W, Rosqvist R, Mullan P B, Edwards M H, Galyov E E. SopE, a secreted protein of Salmonella dublin, is translocated into the target eukaryotic cell via a sip-dependent mechanism and promotes bacterial entry. Mol Microbiol. 1996;22:327–338. doi: 10.1046/j.1365-2958.1996.00116.x. [DOI] [PubMed] [Google Scholar]