Abstract
Vancomycin-resistant enterococci acquire high-level resistance to glycopeptide antibiotics through the synthesis of peptidoglycan terminating in d-alanyl-d-lactate. A key enzyme in this process is a d-alanyl-d-alanine ligase homologue, VanA or VanB, which preferentially catalyzes the synthesis of the depsipeptide d-alanyl-d-lactate. We report the overexpression, purification, and enzymatic characterization of DdlN, a VanA and VanB homologue encoded by a gene of the vancomycin-producing organism Amycolatopsis orientalis C329.2. Evaluation of kinetic parameters for the synthesis of peptides and depsipeptides revealed a close relationship between VanA and DdlN in that depsipeptide formation was kinetically preferred at physiologic pH; however, the DdlN enzyme demonstrated a narrower substrate specificity and commensurately increased affinity for d-lactate in the C-terminal position over VanA. The results of these functional experiments also reinforce the results of previous studies that demonstrated that glycopeptide resistance enzymes from glycopeptide-producing bacteria are potential sources of resistance enzymes in clinically relevant bacteria.
The origin of antibiotic resistance determinants is of significant interest for several reasons, including the prediction of the emergence and spread of resistance patterns, the design of new antimicrobial agents, and the identification of potential reservoirs for resistance elements. Antibiotic resistance can occur either through spontaneous mutation in the target or by the acquisition of external genetic elements such as plasmids or transposons which carry resistance genes (7). The origins of these acquired genes are varied, but it has long been recognized that potential reservoirs are antibiotic-producing organisms which naturally harbor antibiotic resistance genes to protect themselves from the actions of toxic compounds (6).
High-level resistance to glycopeptide antibiotics such as vancomycin and teicoplanin in vancomycin-resistant enterococci (VRE) is conferred by the presence of three genes, vanH, vanA (or vanB), and vanX, which, along with auxiliary genes necessary for inducible gene expression, are found on transposons integrated into plasmids or the bacterial genome (1, 20). These three genes are essential to resistance and serve to change the C-terminal peptide portion of the peptidoglycan layer from d-alanyl-d-alanine (d-Ala-d-Ala) to d-alanyl-d-lactate (d-Ala-d-Lac). This change results in the loss of a critical hydrogen bond between vancomycin and the d-Ala-d-Ala terminus and in a 1,000-fold decrease in binding affinity between the antibiotic and the peptidoglycan layer, which is the basis for the bactericidal action of this class of compounds (5). The vanH gene encodes a d-lactate dehydrogenase which provides the requisite d-Lac (3, 5), while the vanX gene encodes a highly specific dd-peptidase which cleaves only d-Ala-d-Ala produced endogenously while leaving d-Ala-d-Lac intact (19, 21). The final gene, vanA or vanB, encodes an ATP-dependent d-Ala-d-Lac ligase (4, 8, 10). This enzyme has sequence homology with the chromosomal d-Ala-d-Ala ligases, which are essential for peptidoglycan synthesis but which generally lack the ability to synthesize d-Ala-d-Lac (9).
We have recently cloned vanH, vanA, and vanX homologues from two glycopeptide antibiotic-synthesizing organisms: Amycolatopsis orientalis C329.2, which produces vancomycin, and Streptomyces toyocaensis NRRL 15009, which produces A47934 (14). In addition, the vanH-vanA-vanX gene cluster was identified in several other glycopeptide producers. We have also demonstrated that the VanA homologue from S. toyocaensis NRRL 15009 can synthesize d-Ala-d-Lac in vitro and in the glycopeptide-sensitive host Streptomyces lividans (15, 16). We now report the expression of the A. orientalis C329.2 VanA homologue DdlN in Escherichia coli, its purification, and its enzymatic characterization. These data reinforce the striking similarity between vancomycin resistance elements in VRE and glycopeptide-producing organisms and support the possibility of a common origin for these enzymes.
Expression, purification, and specificity of DdlN.
DdlN was overexpressed in E. coli under the control of the bacteriophage T7 promoter. The construct gave good yields of highly purified enzyme following a four-step purification procedure (Table 1; Fig. 1). Like other dd-ligases, DdlN behaved like a dimer in solution (not shown).
TABLE 1.
Purification of DdlN from E. coli BL21 (DE3)/pETDdlN
| Sample | Protein (mg) | Activity (nmol/min) | Sp act (nmol/ min/mg) | Recovery (%) | Purification (fold) |
|---|---|---|---|---|---|
| Lysate | 124 | 843 | 6.82 | 100 | |
| Ammonium sulfate (20–50% saturation) | 67.6 | 780 | 11.5 | 92 | 1.7 |
| Sephacryl S200 | 11.6 | 825 | 71.4 | 98 | 11 |
| Q Sepharose | 2.8 | 742 | 265 | 88 | 39 |
| Phenyl Superose | 0.4 | 299 | 748 | 35 | 110 |
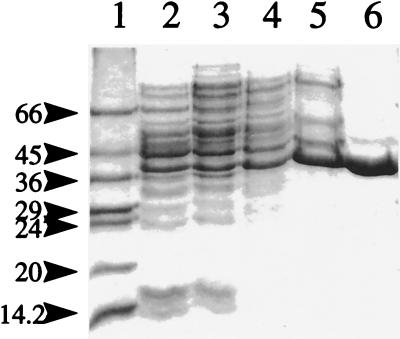
FIG. 1.
Purification of DdlN from E. coli BL21 (DE3)/pETDdlN. Proteins were separated on an SDS–11% polyacrylamide gel and stained with Coomassie blue. Lane 1, molecular mass markers (masses are noted at the left in kilodaltons); lane 2, whole-cell lysate; lane 3, ammonium sulfate fraction (20 to 50% saturation); lane 4, Sephacryl S200; lane 5, Q Sepharose; lane 6, phenyl Superose.
The amino acid substrate specificity of DdlN was assessed by incubation of 14C-d-Ala with all 20 common amino acids in the d configuration. Purified DdlN catalyzed the synthesis of d-Ala-d-Ala in addition to that of several other mixed dipeptides, including d-Ala-d-Met and d-Ala-d-Phe (Fig. 2). Thus, DdlN exhibits a substrate specificity which is similar to that of VanA (4), with the capacity to synthesize not only d-Ala-d-Ala but also mixed dipeptides with bulky side chains in the C-terminal position.
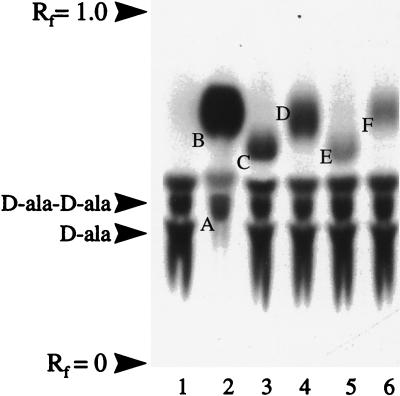
FIG. 2.
Substrate specificity of DdlN. Autoradiogram from thin-layer chromatography analysis of DdlN substrate specificity. All reaction mixtures contained 2.5 mM d-Ala and 1 mM ATP, and the radiolabel was 14C-d-Ala, except where noted. Lane 1, d-Ala; lane 2, d-Lac with 14C-d-Lac label; lane 3, d,l-methionine; lane 4, dl-phenylalanine; lane 5, d-Hbut; lane 6, d-hydroxyvalerate. Letters indicate the following: A, d-Ala-d-Lac; B, d-Lac; C, d-Ala-d-Met; D, d-Ala-d-Phe; E, d-Ala-d-Hbut; F, d-Ala-d-hydroxyvalerate.
Importantly, DdlN is a depsipeptide synthase with the ability to synthesize d-Ala-d-Lac, d-Ala-d-hydroxybutyrate (Hbut), and d-Ala-d-hydroxyvalerate (Fig. 2). However, unlike VanA (5), d-hydroxycaproate and d-phenyllactate are not substrates (not shown). Thus, DdlN is a broad-spectrum d-Ala-d-X ligase with depsipeptide synthase activity.
Characterization of d-Ala-d-X ligase activity.
Following the initial assessment of the specificity of the enzyme, several substrates were selected for quantitative analysis by evaluation of their steady-state kinetic parameters (Table 2). DdlN has two amino acid (or hydroxy acid) Km values. Steady-state kinetic plots indicated that, like other dd-ligases, the N-terminal Km (Km1) was significantly lower (higher specificity) than the C-terminal Km (Km2). Since the former value is expected to be independent of the C-terminal substrate, only Km2 values were determined and are reported here.
TABLE 2.
Characterization of steady-state parameters of DdlN and VanA
| Ligase | Substrate | Km2 (mM) | kcat (min−1) | kcat/Km2 (M−1 s−1) |
|---|---|---|---|---|
| DdlN | d-Ala | 21 ± 2 | 229 ± 7 | 1.8 × 102 |
| d-Lac | 0.4 ± 0.05 | 55 ± 1 | 2.3 × 103 | |
| d-Hbut | 2.5 ± 0.3 | 32 ± 2 | 2.1 × 102 | |
| ATPa | 1.2 ± 0.2 | 71 ± 5 | 0.98 × 102 | |
| DdlMb | d-Ala | 166 ± 27 | ||
| d-Lac | 1.08 ± 0.10 | |||
| VanAc | d-Ala | 38 | 295 | 1.3 × 102 |
| d-Lac | 7.1 | 94 | 2.2 × 102 | |
| d-Hbut | 0.60 | 108 | 3.0 × 103 |
DdlN showed good d-Ala-d-Ala ligase activity but with a very high and physiologically questionable Km2 (21 mM). On the other hand, d-Ala-d-Lac synthesis was excellent, with a 4-fold decrease in kcat, compared to d-Ala-d-Ala synthesis, which was offset by a 52-fold drop in Km that resulted in a >12-fold increase in specificity (kcat/Km2). d-Hbut was also a good substrate, with a kcat/Km2 comparable to that of d-Ala.
Steady-state kinetic parameters for d-Ala-d-X formation showed trends similar to those found with both VanA and DdlN. For example, the kcat values between VanA and DdlN were virtually the same for most substrates. There were significant differences, however. For instance, while the Km2 values for d-Ala were very high for all three enzymes, DdlN does have greater affinity for d-Ala, with a 1.8- and 7.9-fold lower Km2 than those of VanA and DdlM, respectively. Additionally, the Km2 for d-Lac was 17.8- and 2.7-fold lower than those for VanA and DdlM. Thus, DdlN has a more restrictive specificity for the C-terminal residue than VanA, which is compensated for by a higher affinity for the critical substrate d-Lac.
pH dependence of peptide versus that of depsipeptide synthesis activity.
The partitioning of the syntheses of d-Ala-d-Ala and d-Ala-d-Hbut in VanA and other depsipeptide-competent dd-ligases has been shown to be pH dependent (17). Determination of the pH dependence of DdlN in synthesizing peptide versus depsipeptide (Fig. 3) directly paralleled the results obtained with VanA in similar experiments. At lower pHs (<7), d-Ala-d-Hbut synthesis predominates and is exclusive at a pH of <6 (Fig. 3). At pH 7.5, levels of synthesis of d-Ala-d-Hbut and d-Ala-d-Ala are relatively equal, while at a pH greater than 8, the capacity to synthesize peptide overtakes the capacity to synthesize depsipeptide, although the latter is never abolished.
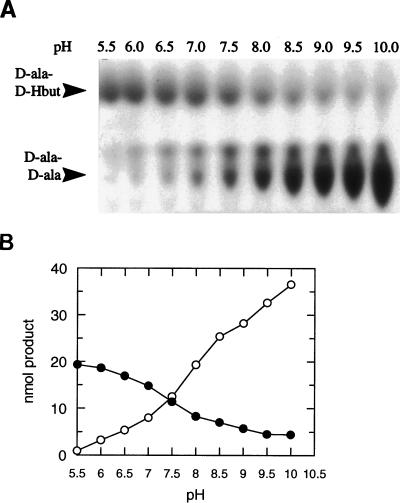
FIG. 3.
pH dependence of partitioning of the syntheses of peptide and depsipeptide by DdlN. (A) Autoradiogram of a thin-layer chromatography separation of the products of reaction mixtures containing 14C-D-Ala, unlabeled D-Ala, and d-Hbut. (B) Quantification of reaction products following phosphorimage analysis. Filled circles, D-Ala-d-Hbut; open circles, D-Ala-D-Ala.
The partitioning of the formation of peptide versus depsipeptide as a function of pH by DdlM is comparable to that by VanA and depsipeptide-competent mutants of DdlB (17), which show essentially exclusively depsipeptide formation at lower pHs and increasing peptide formation as the pH increases. This implies a potential role for the protonated ammonium group of d-Ala2 in second-substrate recognition and suggests a mechanism for the discrimination between d-Ala and d-Lac at physiologic pH. The structural basis for this distinction remains obscure for DdlB and VanA or DdlN.
Concluding remarks.
Resistance to vancomycin and other glycopeptides is mediated through the synthesis of a peptidoglycan which does not terminate with the canonical d-Ala-d-Ala dipeptide. Thus, enterococci which exhibit the VanC phenotype, which consists of low-level, noninducible resistance to vancomycin only, have peptidoglycan terminating in d-Ala-d-Ser (19). On the other hand, bacteria which are constitutively resistant to high concentrations of glycopeptides, such as lactic acid bacteria and VRE exhibiting the VanA or VanB phenotype (high-level inducible resistance to vancomycin), incorporate the depsipeptide d-Ala-d-Lac into their cell walls (2, 12, 13). The enzymes responsible for the intracellular synthesis of d-Ala-d-Lac not surprisingly have significant amino acid sequence similarity with d-Ala-d-Ala ligases, which are responsible for d-Ala-d-Ala synthesis in all bacteria with a cell wall (9).
The d-Ala-d-Lac synthases can be subdivided into two groups based on sequence homology: those found in the constitutively resistant lactic acid bacteria and those found in glycopeptide-producing organisms and VanA or VanB VRE (9, 14). The former have more similarity with exclusive d-Ala-d-Ala ligases. Indeed, single point mutations in d-Ala-d-Ala ligases which yield sequences more similar to those of lactic acid bacterium d-Ala-d-Lac ligases are sufficient to induce significant depsipeptide synthase activity in these enzymes (17). Similarly, mutational studies of the d-Ala-d-Lac ligase from Leuconostoc mesenteroides have demonstrated that the converse also holds (18). On the other hand, the molecular basis for depsipeptide synthesis by the VanA or VanB ligases is unknown, in large part due to the lack of protein structural information on which to base mutational studies, unlike the situation with d-Ala-d-Ala ligases, where the E. coli DdlB structure serves as a template for mechanistic research (11).
Significantly, a major difference in the VanA or VanB ligases and other dd-ligases lies in the amino acid sequence of the ω-loop region, which closes off the active site of DdlB (11) and has been shown to contribute amino acid residues with the capacity to control the syntheses of d-Ala-d-Ala and d-Ala-d-Lac, notably, Tyr216 (17, 18). Until recently, the VanA and VanB ligases were exceptional in amino acid structure and had no known homologues. The sequencing of resistance genes from glycopeptide-producing bacteria has uncovered enzymes with >60% homology to VanA or VanB and which are virtually superimposable in the critical ω-loop region (14, 15). One of these, DdlM from S. toyocaensis NRRL 15009, has been shown to have d-Ala-d-Lac ligase ability (15, 16), although no rigorous analysis of this activity has been performed. The results presented here demonstrate that DdlN from the vancomycin producer A. orientalis C329.2 not only is a d-Ala-d-Lac ligase but also has significant functional homology with VanA. It is not known at present if, like S. toyocaensis NRRL 15009 (16), A. orientalis C329.2 also possess a d-Ala-d-Ala-exclusive ligase, though the presence of a vanX gene (14) suggests that it may.
These studies demonstrate that DdlN cloned from a vancomycin-producing bacterium is a d-Ala-d-Lac ligase which has not only amino acid sequence homology with the dd-ligases from VRE but also functional homology. Thus, VanA, VanB, DdlN, and DdlM have likely evolved from similar origins. The fact that a vanH-vanA-vanX gene cluster can be found in other glycopeptide producers as well (14) suggests that the genes now found in VRE may have originated in glycopeptide-producing bacteria. Our finding that overexpressed, purified, DdlN shows many enzymatic characteristics similar (though not identical) to those of VanA suggests that the genes from glycopeptide-producing bacteria can be important in elucidating biochemical and protein structural aspects of the VRE proteins.
Acknowledgments
This research was supported by an operating grant from the Natural Sciences and Engineering Research Council of Canada (NSERC), by a Medical Research Council of Canada (MRC) graduate studentship to C.G.M., and by an MRC Scholar Award to G.D.W.
REFERENCES
- 1.Arthur M, Courvalin P. Genetics and mechanism of glycopeptide resistance in enterococci. Antimicrob Agents Chemother. 1993;37:1563–1571. doi: 10.1128/aac.37.8.1563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Arthur M, Molinas C, Bugg T D H, Wright G D, Courvalin P. Evidence for in vivo incorporation of d-lactate into peptidoglycan precursors of vancomycin-resistant enterococci. Antimicrob Agents Chemother. 1992;36:867–869. doi: 10.1128/aac.36.4.867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Arthur M, Molinas C, Dutka-Malen S, Courvalin P. Structural relationship between the vancomycin resistance protein VanH and 2-hydroxycarboxylic acid dehydrogenases. Gene. 1991;103:133–134. doi: 10.1016/0378-1119(91)90405-z. [DOI] [PubMed] [Google Scholar]
- 4.Bugg T D, Dutka-Malen S, Arthur M, Courvalin P, Walsh C T. Identification of vancomycin resistance protein VanA as a D-alanine:D-alanine ligase of altered substrate specificity. Biochemistry. 1991;30:2017–2021. doi: 10.1021/bi00222a002. [DOI] [PubMed] [Google Scholar]
- 5.Bugg T D H, Wright G D, Dutka-Malen S, Arthur M, Courvalin P, Walsh C T. Molecular basis for vancomycin resistance in Enterococcus faecium BM4147: biosynthesis of a depsipeptide peptidoglycan precursor by vancomycin resistance proteins VanH and VanA. Biochemistry. 1991;30:10408–10415. doi: 10.1021/bi00107a007. [DOI] [PubMed] [Google Scholar]
- 6.Cundliffe E. How antibiotic-producing organisms avoid suicide. Annu Rev Microbiol. 1989;43:207–233. doi: 10.1146/annurev.mi.43.100189.001231. [DOI] [PubMed] [Google Scholar]
- 7.Davies J, Webb V. Antibiotic resistance in bacteria. In: Krause R M, editor. Emerging infections. San Diego, Calif: Academic Press; 1998. pp. 239–273. [Google Scholar]
- 8.Dutka-Malen S, Molinas C, Arthur M, Courvalin P. The VANA glycopeptide resistance protein is related to D-alanyl-D-alanine ligase cell wall biosynthesis enzymes. Mol Gen Genet. 1990;224:364–372. doi: 10.1007/BF00262430. [DOI] [PubMed] [Google Scholar]
- 9.Evers S, Casadewall B, Charles M, Dutka-Malen S, Galimand M, Courvalin P. Evolution of structure and substrate specificity in D-alanine:D-alanine ligases and related enzymes. J Mol Evol. 1996;42:706–712. doi: 10.1007/BF02338803. [DOI] [PubMed] [Google Scholar]
- 10.Evers S, Sahm D F, Courvalin P. The vanB gene of vancomycin-resistant Enterococcus faecalis V583 is structurally related to genes encoding D-Ala:D-Ala ligases and glycopeptide-resistance proteins VanA and VanC. Gene. 1993;124:143–144. doi: 10.1016/0378-1119(93)90779-3. [DOI] [PubMed] [Google Scholar]
- 11.Fan C, Moews P C, Walsh C T, Knox J R. Vancomycin resistance: structure of D-alanine:D-alanine ligase at 2.3 Å resolution. Science. 1994;266:439–443. doi: 10.1126/science.7939684. [DOI] [PubMed] [Google Scholar]
- 12.Handwerger S, Pucci M J, Volk K J, Liu J, Lee M S. The cytoplasmic peptidoglycan precursor of vancomycin-resistant Enterococcus faecalis terminates in lactate. J Bacteriol. 1992;174:5982–5984. doi: 10.1128/jb.174.18.5982-5984.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Handwerger S, Pucci M J, Volk K J, Liu J, Lee M S. Vancomycin-resistant Leuconostoc mesenteroides and Lactobacillus casei synthesize cytoplasmic peptidoglycan precursors that terminate in lactate. J Bacteriol. 1994;176:260–264. doi: 10.1128/jb.176.1.260-264.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Marshall C G, Lessard I A D, Park I-S, Wright G D. Glycopeptide antibiotic resistance in glycopeptide-producing organisms. Antimicrob Agents Chemother. 1998;42:2215–2220. doi: 10.1128/aac.42.9.2215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Marshall C G, Broadhead G, Leskiw B, Wright G D. D-Ala-D-Ala ligases from glycopeptide antibiotic-producing organisms are highly homologous to the enterococcal vancomycin-resistance ligases VanA and VanB. Proc Natl Acad Sci USA. 1997;94:6480–6483. doi: 10.1073/pnas.94.12.6480. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Marshall C G, Wright G D. The glycopeptide antibiotic producer Streptomyces toyocaensis NRRL 15009 has both D-alanyl-D-alanine and D-alanyl-D-lactate ligases. FEMS Microbiol Lett. 1997;157:295–299. doi: 10.1111/j.1574-6968.1997.tb12788.x. [DOI] [PubMed] [Google Scholar]
- 17.Park I S, Lin C H, Walsh C T. Gain of D-alanyl-D-lactate or D-lactyl-D-alanine synthetase activities in three active-site mutants of the Escherichia coli D-alanyl-D-alanine ligase B. Biochemistry. 1996;35:10464–10471. doi: 10.1021/bi9603128. [DOI] [PubMed] [Google Scholar]
- 18.Park I S, Walsh C T. D-Alanyl-D-lactate and D-alanyl-D-alanine synthesis by D-alanyl-D-alanine ligase from vancomycin-resistant Leuconostoc mesenteroides. Effects of a phenylalanine 261 to tyrosine mutation. J Biol Chem. 1997;272:9210–9214. doi: 10.1074/jbc.272.14.9210. [DOI] [PubMed] [Google Scholar]
- 19.Reynolds P E, Depardieu F, Dutka-Malen S, Arthur M, Courvalin P. Glycopeptide resistance mediated by enterococcal transposon Tn1546 requires production of VanX for hydrolysis of D-alanyl-D-alanine. Mol Microbiol. 1994;13:1065–1070. doi: 10.1111/j.1365-2958.1994.tb00497.x. [DOI] [PubMed] [Google Scholar]
- 20.Walsh C T, Fisher S L, Park I-S, Prahalad M, Wu Z. Bacterial resistance to vancomycin: five genes and one missing hydrogen bond tell the story. Chem Biol. 1996;3:21–28. doi: 10.1016/s1074-5521(96)90079-4. [DOI] [PubMed] [Google Scholar]
- 21.Wu Z, Wright G D, Walsh C T. Overexpression, purification, and characterization of VanX, a D-D-dipeptidase which is essential for vancomycin resistance in Enterococcus faecium BM4147. Biochemistry. 1995;34:2455–2463. doi: 10.1021/bi00008a008. [DOI] [PubMed] [Google Scholar]