Abstract
Optimizing the metabolic pathways of microbial cell factories is essential for establishing viable biotechnological production processes. However, due to the limited understanding of the complex setup of cellular machinery, building efficient microbial cell factories remains tedious and time-consuming. Machine learning (ML), a powerful tool capable of identifying patterns within large datasets, has been used to analyze biological datasets generated using various high-throughput technologies to build data-driven models for complex bioprocesses. In addition, ML can also be integrated with Design–Build–Test–Learn to accelerate development. This review focuses on recent ML applications in genome-scale metabolic model construction, multistep pathway optimization, rate-limiting enzyme engineering, and gene regulatory element designing. In addition, we have discussed some limitations of these methods as well as potential solutions.
Keywords: Machine learning, Metabolic pathway optimization, Active learning, Bayesian optimization, Mechanism model, Data-driven model
1. Introduction
Microbiological fermentation is a green and sustainable approach to produce chemicals, materials, fuels, food, and pharmaceuticals. It can also contribute to solving the global energy crisis and environmental problems [1]. However, natural microbials are rarely suitable for directly producing desired chemicals on an industrial scale. To overcome this obstacle, metabolic pathway optimization technologies, including genetic interventions, have been exploited to develop highly efficient microbial cell factories by redistributing the carbon metabolic flow toward desired metabolites [2].
Significant progress has been made in metabolic pathway optimization over the past few decades. For example, replacement of a promoter in Escherichia coli BL21 led to a 4.2-fold increase in 2,3-butanediol production (73.8 g/L) in fed-batch fermentation compared with that obtained in a previous study [3]. In another case, rubusoside was de novo biosynthesized in Saccharomyces cerevisiae using a systematic engineering strategy, and the titer reached 1368.6 mg/L [4]. Moreover, implementation of a global transcriptional machinery engineering strategy led to a 114% increase in L-tyrosine production in E. coli P2 in large-scale fermentation [5]. Despite notable applications of metabolic pathway optimization technologies, some potential problems may hinder further development. For instance, due to incomplete understanding of the relationship between the phenotype and genotype of target cells, the conventional trial-and-error approach is often adopted [6]. In addition, identifying and testing each combination of different pieces of the genome takes time and effort, and this can significantly hinder the construction and application of microbial cell factories [7]. Moreover, despite rapid development in DNA editing technology, the incredible potential of genome-scale engineering has not been fully exploited because conventional research has focused on the redirection of carbon flux in a limited number of metabolic pathways [8], [9], [10].
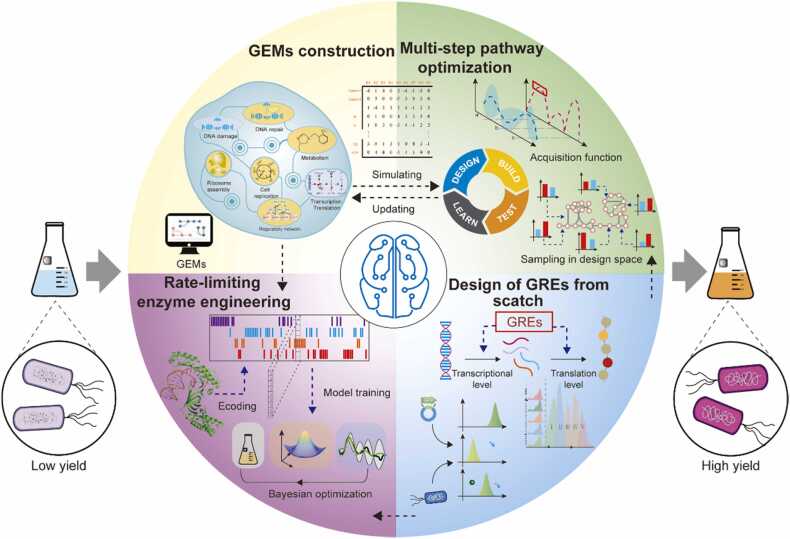
Machine learning (ML), a subdiscipline of artificial intelligence, attempts to imitate how the human brain learns and interacts using computers that implement learning algorithms [11], [12], [13]. ML-enabled systems can extract knowledge from previously collected data and use this knowledge to build simulation models. ML has recently been applied in optimizing metabolic pathways due to its excellent modeling ability. On one hand, ML has often replaced conventional statistical methods, including linear regression and partial least squares-based statistical modeling, to build models used to identify features within biological datasets. For instance, DeepEC was developed to predict enzyme commission (EC) numbers using a protein sequence as an input [14]. On the other hand, ML can also be integrated into Design–Build–Test–Learn (DBTL) cycles to explore design space more effectively. For example, Zhou et al. proposed an ML-assisted tool to determine the optimal combination of enzyme expression levels [14]; Greenhalgh et al. proposed an ML-based workflow to improve the performance of rate-limiting enzymes [15]. Based on this, the present study reviews the recent applications of ML in genome-scale metabolic model (GEM) construction, multistep pathway optimization, rate-limiting enzyme engineering, and gene regulatory element (GRE) designing, which constitute the critical frameworks of metabolic pathway optimization. In addition, we have discussed some limitations of these methods as well as potential solutions.
2. Application of ML in genome-scale metabolic model (GEM) construction
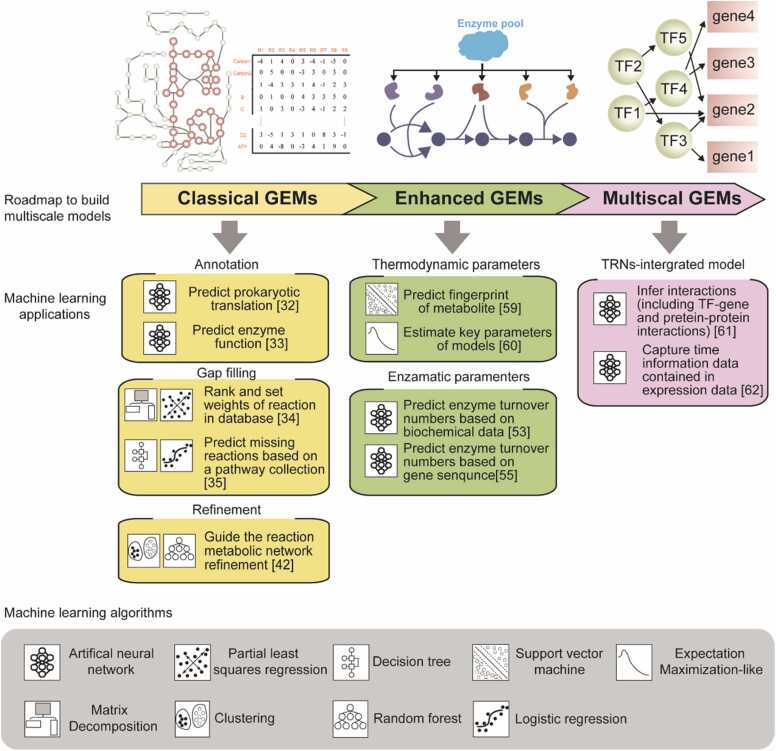
GEMs computationally describe the metabolic networks of organisms using gene-protein-reaction (GPR) rules [16], which aim to understand the authentic relationships between genotypes and phenotypes [17]. Classical GEMs comprise the mathematical metabolic network and the objective function, both of which are subject to stoichiometric constraints in flux balance analysis (FBA) [18], [19]. However, applying purely stoichiometric constraints usually results in an underdetermined system with infinite solutions [20]. To achieve more accurate prediction of metabolic flux distribution, novel modeling concepts that include more cellular processes have been proposed, such as enzyme-constrained genome-scale metabolic models (ecGEMs) [21], macromolecular expression models [18], and whole-cell models [22]. Despite considerable advancement in the modeling concept, the deficiencies in quantitative mechanistic representations of our knowledge of molecular processes have limited the development of advanced GEMs. Since its recent emergence as a key data-driven method, ML has been employed to quantitatively depict various subcellular processes (Fig. 1).
Fig. 1.
Machine learning (ML) applications for genome-scale metabolic model (GEM) construction. There are three main categories of ML applications for GEM construction: classical GEMs, enhanced GEMs, and multiscale models. Typical applications are listed in the figure according to the adopted ML algorithm and primary task.
2.1. Classical GEMs
Since the first GEM of Haemophilus influenzae was reported in 1999, several classical GEMs have been constructed for approximately 6239 organisms [23]. Moreover, some classical GEMs for industrial organisms have been upgraded several times as the understanding of GPR relations has been updated; these industrial organisms include S. cerevisiae, E. coli [18], Bacillus subtilis [24], and Corynebacterium glutamicum [25], [26]. Although classical GEMs only consider the stoichiometric constraints of the metabolic network [22], they have offered many valuable suggestions for metabolic pathway optimization over the past decade [27], [28], [29]. Moreover, the construction workflow of some advanced GEMs, including thermodynamic, enzymatic, and kinetic constraint models, requires classic GEMs to offer high-quality metabolic networks. Thus, building high-quality classical GEMs remains critical in the GEM field [17], [30], [31].
Drafts of classical GEMs are always constructed using the annotation results for the whole genome [32]. Thus, employing genome annotation tools with excellent performance is crucial for developing high-quality metabolic networks. To improve the accuracy of the genome annotation step, various ML algorithms are applied to exploit the information in the target genome. For example, to determine minor variations in annotated gene regions between different prokaryote (sub)-species, a novel neural network named DeepRibo was proposed to precisely delineate and annotate expressed genes using features extracted from ribosome profiles and binding site sequence patterns [33]. In addition to precisely locating expressed genes, high-quality and high-throughput prediction of enzyme functions is essential for genome annotation. DeepEC, a deep learning-based computational framework, was developed to predict EC numbers of protein sequences with high precision and in a high-throughput manner [34]. For the development of DeepEC, three convolutional neural networks were integrated into a single engine that could predict EC numbers.
After construction of the draft model, gaps usually exist due to incomplete knowledge of the metabolic network [35]. To ensure adequate GEM quality, it is essential to analyze the metabolic network and fill gaps within it. Recently, several studies have focused on improving the accuracy and efficiency of the gap-filling process. For example, Tolutola et al. proposed an ML-based strategy named BoostGAPFILL [35], which leverages ML methodologies and constraint-based models to generate hypotheses for gap-filling and model refinement. More specifically, reactions that fill gaps are constrained by metabolite patterns in the incomplete network. BoostGAPFILL shows> 60% precision and recall. Similarly, in another study, a series of ML algorithms, including decision trees and logistic regression [36], was applied to identify missing reactions and enzymes in the model. By collecting 123 pathway features from 5610 pathway instances, these ML methods exhibited greater interpretability and extensibility than other pathway prediction algorithms.
Model refinement is also essential to verify gap-filling solutions following the construction of a draft model. In recent years, several automatic construction tools, such as CarveMe [37], Merlin [38], ModelSEED [39], and Pathway Tools [40], have been proposed due to the massive increase in biochemical knowledge and computational annotation of genomes [41]. However, curation and refinement of metabolic networks in GEMs are still rare [42]. Recently, an ML-based method, called automated metabolic model ensemble-driven uncertainty elimination using statistical learning, has shown that ML can reduce the workload of curation work for draft GEMs. In their research, iterative gap-filling was initially performed to produce random gap-filling solutions [43]. Moreover, supervised and unsupervised learning were integrated to determine the uncertainty of the model. This study suggested that combining ML strategies can improve the efficiency of tedious yet essential manual refinement and curation steps.
2.2. Enhanced and multiscale models
Compared with classical GEMs, multiscale models consist of multiple heterogeneous models or networks of different cellular layers [22]. Typical examples of multiscale models include thermodynamic constraint GEMs [43], [44], enzymatic constraint GEMs [21], [45], and multiomics-integrated GEMs [46]. Each of these has been widely used in biological discoveries, extensive data analysis, and metabolic engineering [22]. Recently, various ML strategies have been applied to construct heterogeneous models and networks. For example, expectation maximization (EM)-like algorithms are used to estimate critical parameters in metabolite identification models [47], [48], and RF is applied to predict enzyme turnover numbers. Introducing ML algorithms into multiscale models can improve model quality effectively and expand model network dimensionality [49].
Traditional model analysis approaches, such as FBA, search for an optimal growth rate that is constrained only by metabolic network stoichiometric constraints and uptake rates [20]. To further enhance the simulation capabilities of GEMs, the cost of expressing enzymes within metabolic networks is an essential constraint to be considered while constructing ecGEMs [50]. By adding this enzyme constraint, ecGEMs can accurately simulate maximum growth ability, metabolic shifts, and proteome allocations of various enzymes [51]. ecGEMs are constructed using genome-scale enzyme turnover numbers (kcats), each of which defines a reaction’s maximum chemical conversion rate. Nevertheless, the scope of the kcats dataset is far from the genome scale because kcats are usually measured via low-throughput assays in vitro [52]. In addition, there is a vast difference between kcats in vivo and in vitro due to incomplete saturation, posttranslational modifications, and allosteric regulation [53]. To alleviate this problem, Heckmann et al. proposed a ML method to predict kcats [50]. EC numbers, molecular weight, in silico flux predictions, and assay conditions were integrated to predict kcats under both in vivo and in vitro conditions. Finally, improved forecasts of proteome allocation were achieved by applying ML models to parameterize GEMs. In addition, to further improve prediction accuracy, the in silico flux predictions were replaced with 13C fluxomics data to estimate kcats in vivo [54]. Moreover, whether the maximum value of kcats is robust to genetic perturbations was tested by performing gene knockout experiments and adaptive laboratory evolution. The results indicated that maximum kcats values in vivo are stable. However, despite improvement in predicting kcats in vivo, features such as average metabolic flux and catalytic sites obtained from protein structure are typically too complex to obtain from nonmodel organisms. To this end, Lee et al. developed a deep learning approach (DLKcat) to predict kcats from only substrate structure and protein sequence data. Using this method, kcats profiles for 343 yeast/fungi species were predicted. Furthermore, an automatic Bayesian-based pipeline was proposed in their study, which enabled the automatic reconstruction of 343 ecGEMs of yeast [55].
In addition, calculating the Gibbs energy of reactions is an essential step in constructing thermodynamic constraints for multiscale models [56]. However, metabolite identification is a significant challenge in metabolomics due to the number and diversity of molecules. Metabolites can be rapidly identified via fingerprint identification. Nevertheless, many metabolites require more accurate fingerprints. To this end, ML has been employed in recent studies for the prediction of metabolite fingerprints. FingerID, a classical method, has been proposed to predict corresponding fingerprints from a mass spectrometry (MS) set with supervised ML [57]. A support vector machine (SVM) selects fingerprints with integral mass and probability product kernels. FingerID is mainly based on information derived from individual peaks present in the spectra. However, the relationships between different peaks have also been used to predict fingerprints. To further increase model predictive power, CSI: FingerID, an extended version of FingerID, was proposed by combining MS spectra with a corresponding fragmentation tree [58]. CSI: FingerID exhibited better predictive accuracy than FingerID in predicting the fingerprints of metabolites. Nevertheless, the optimization of hyperparameters becomes more complex since more inputs are required in CSI: FingerID. Moreover, kernel-based methods are not desirable for dealing with MS spectra since the spectrum comprises only a few peaks. To mitigate these limitations, a new algorithm named SIMPLE was proposed [59]. SIMPLE is more efficient and interpretable in predicting metabolite fingerprints. SIMPLE is a generalized additive model that captures information from individual peak and peak interactions. Compared with kernel-based methods, an obvious advantage of SIMPLE is its prediction speed. Moreover, the performance of SIMPLE is correlated with several peaks in the spectrum, whereas that of kernel-based methods explicitly depends on the size of the training dataset.
Transcriptional regulatory networks (TRNs) explain complex life phenotypes at the genomic level of organisms under different environments [60]. Integrating TRNs and GEMs can enable a more comprehensive understanding of metabolic regulation and stress response [22]. Several conventional methods, such as Pearson correlation, have been employed to infer TRNs. However, these methods require multilevel biological data to accurately predict TRNs [61]. Recently, a supervised ML strategy, CNN for coexpression (CNNC), was proposed to infer gene relationship [62]. In their study, image representation was used to replace conventional information. More specifically, the model was trained with negative and positive examples of a specific domain, and the prediction could be either binary or multinomial. However, an important feature, time information, is ignored by the CNNC model. Therefore, a hybrid deep learning framework for gene regulatory network inference from single-cell transcriptomic data (DGRNS) was developed to capture time information in expression data [62]. DGRNS provides a supervised ML method that can extract both statistical and time-related features. In its workflow, DGRNS first performs a series of pretreatments and constructs correlation vectors that represent gene expression features [17], and this approach therefore improves the accuracy of the inference of TRNs.
Although ML has many successful applications in the construction of GEMs, some potential problems may hamper the further application of ML in GEMs. For example, advanced ML tools are yet to be integrated into conventional construction frameworks. Moreover, constructing a whole-cell model by coupling all mechanism models is challenging because it is laborious to establish mechanistic models for all subcellular processes.
3. ML-guided multistep pathway optimization
A major task of metabolic pathway optimization is to identify the optimal combination of multiple gene expression levels within a pathway. It is a promising strategy to fully explore the potential genetic design space using high-throughput (HTP) technologies [63]. However, the performance of HTP screening depends on accurate and rapid detection methods, which are only available for some biochemicals [64]. Moreover, searching the entire genetic design space is a resource-intensive strategy, which may incur high costs. To this end, many computational approaches have been used to understand the metabolic regulation processes of microorganisms and identify genetic interventions necessary to achieve a desired phenotype [65]. Nevertheless, due to the complexity of biological systems, mechanistic models, such as GEMs, and kinetic models can only offer suggestions for optimizing metabolic pathways. Optimal genetic interventions remain to be identified using conventional trial-and-error approaches [66]. To overcome these challenges, ML and statistical methods have been used to explore the genetic design space more effectively.
3.1. Active learning
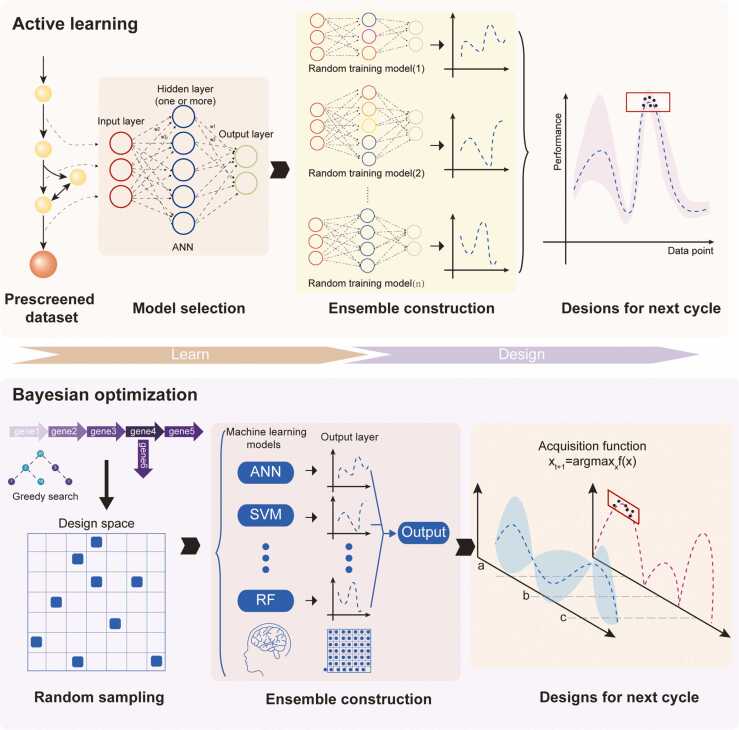
Active learning reduces resource overhead and human effort by gradually exploring design space to improve model quality [67]. Design–Build–Test–Learn (DBTL) cycles are usually tedious and time-consuming because the learn phase of DBTL needs to be better developed [2]. To this end, active learning has been introduced to accelerate the DBTL cycle by enhancing the learning phase. For example, MiYa, an ML workflow that works in conjunction with the YeastFab [68] assembly strategy, was proposed to optimize the expression level of numerous genes by replacing their promoters [14]. As an active learning method, MiYa adopted a low-throughput experimental strategy to ensure the accuracy of experimental data. To avoid overfitting, 1000 ANN models with random initial weights were used to predict the best strain within the design space. In addition, the quality of the initial dataset has been proven to affect the accuracy of prediction. Since the products are colored, a high-quality initial training dataset was constructed via manual screening to improve prediction accuracy (Fig. 2). Similarly, Opgenorth et al. integrated ML into DBTL cycles to optimize dodecanol production in E. coli [69]. Nevertheless, the biosynthesis of dodecanol is more complicated because protein abundance within the pathway not only affects the yield but also the purity of the product. Thus, multiobjective modeling methods, including random forest, polynomial, and multilayer perceptron models, have been systematically tested and compared. After two cycles of DBTL, the yield obtained was 6-fold greater than that previously reported [58]. Furthermore, active learning has been applied in developing classification models. For example, Kumar et al. [65] proposed a strategy based on an SVM, ActiveOpt, to enhance microbial chemical production. In their strategy, data are divided into two classes according to a specific cutoff and is used to train the SVM. The SVM is then applied to find the point farthest from the hyperplane within the design space. If this point can be verified experimentally, it is entered into the dataset and the cutoff is updated.
Fig. 2.
Main strategies used to accelerate Design–Build–Test–Learn (DBTL) cycles. Two strategies can be applied to accelerate DBTL cycles: active learning and Bayesian optimization. Research using the active learning strategy often avoided overfitting by multiple modeling. In this framework, data points with higher output were considered in subsequent evaluation rounds. In contrast, research using the Bayesian strategy attempted to avoid overfitting by constructing a Bayesian ensemble model in which the tradeoff between output value and confidence could be exploited.
Although the abovementioned studies successfully integrate active learning with DBTL, several potential problems hamper further application of these methods. For example, the frameworks mentioned above adopt only one ML algorithm and may therefore be applicable in only some cases. Moreover, the design space is often too conservative due to limitations associated with experimental throughput. Furthermore, the methods listed above generally make decisions on where to evaluate in the next round based only on the output value of the predictive model. They can therefore be trapped by local optima when the confidence of the output value is ignored.
3.2. Bayesian optimization
Theoretically, active learning aims to improve a model’s performance by gradually exploring the areas of the design space with the lowest confidence. However, in reality, researchers generally prefer to use fewer resources to quickly identify optimal solutions. In addition, the accuracy of the predictive model in low output regions of design space is not critical [70].
To escape from local optima and save resources, an exploitation–exploration strategy known as Bayesian optimization has been introduced for metabolic pathway optimization [65]. For example, a new metabolic pathway optimization tool, the Illinois Biological Foundry for Advanced Biomanufacturing (iBioFAB), simultaneously considers the expected outcome of each evaluation and the confidence in the desired outcome [71]. Gaussian Process was employed to assign a mean and variance to each point within the design space. With increasing size of the training set, the mean value and confidence are gradually adjusted. Next, an acquisition function, termed expected improvement, identifies the point of optimal exploitation by quantifying the tradeoff between output value and confidence. To test the performance of iBioFAB, the lycopene production pathway was optimized by fine-tuning gene expression levels. iBioFAB successfully introduced the exploitation–exploration strategy into metabolic pathway optimization. However, a problem still exists. The prediction distribution is assumed to be Gaussian, which does not apply to all situations [70]. To extend the application scope of Bayesian optimization, an automated recommendation tool (ART) for ML approaches was proposed [72]. Compared with iBioFAB, ART integrates multiple ML algorithms using the scikit-learn toolbox. Notably, ART enables sidestepping the challenge of model selection using an ensemble model approach (Fig. 2). Parallel-tempering-based Markov chain Monte Carlo-based (MCMC) sampling is used as an acquisition function to determine exploration or exploitation in the decision-making process. Furthermore, the capabilities of ART were demonstrated on simulated datasets and experimental data from real metabolic engineering projects related to the production of renewable biofuels, hops-flavored beer without hops, fatty acids, and tryptophan [69].
Frameworks based on active learning and Bayesian optimization have significantly enhanced the ability of researchers to explore the vastness of design space. Nevertheless, some limitations still exist. For example, the identification of targets in many studies is based on accurate prior knowledge, which limits the application scope of this advanced framework. Moreover, the targets often belong to the same pathway, which may strongly restrict the power of advanced ML frameworks to optimize metabolic flux on the genome scale.
4. Application of ML in rate-limiting enzyme engineering
Natural enzymes are rarely optimal for industrial applications [73]. However, their untapped potential can be leveraged to satisfy the demand for diverse biotechnological applications and meet specific biotechnological challenges [74]. There is a relationship between the selection function (i.e., fitness) and the sequence of amino acids. This is termed as the fitness landscape and is represented as a surface in a high-dimensional space defined by the function f(sequence) = fitness [75], [76], [77]. Exploration of the protein fitness landscape is the primary task of protein engineering [78]. Nevertheless, this job is challenging because the search space grows exponentially with the number of amino acid positions considered. Moreover, functional proteins are extremely rare in the fitness landscape [79]. To identify optimal sequences within the landscape, rational protein redesign and directed evolution (DE) have been applied to protein engineering. Rational protein redesign builds a mechanistic model based on molecular dynamics simulations to predict changes in structure and protein fitness caused by specific mutations [80]. However, the success of this method depends on accurate protein structure data. In addition, full simulations of the complete protein landscape in silico are resource-intensive [81]. Compared with rational protein redesign, DE improves protein fitness by iteratively accumulating positive mutation results, independently of prior knowledge and simulation data in silico [82]. Moreover, DE can be conducted with a low screening burden because only single mutations and not combinations of mutations are assessed in each round [83]. However, this process ignores the cooperation of different mutations and can be easily trapped by local optima. Increasingly, ML algorithms have been applied to approximate the protein fitness landscape, and they require no prior physical, chemical, or biological knowledge [84].
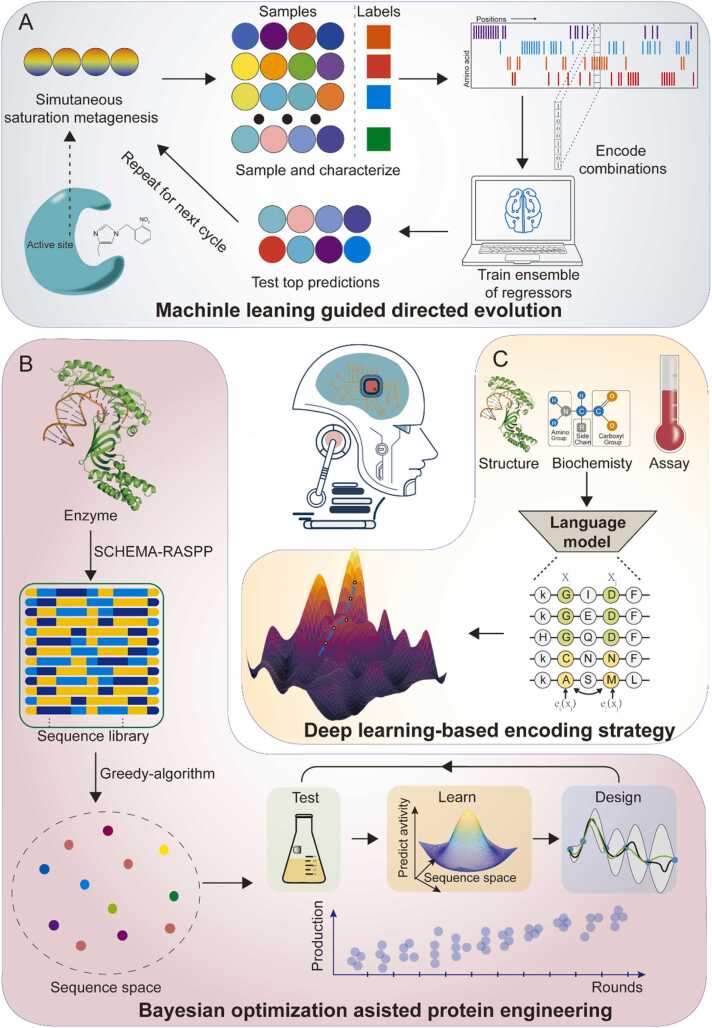
In addition to accumulating single positive mutations, the DE’s ability to explore the fitness landscape can be further enhanced via simultaneous saturation mutagenesis. ML can be employed to assist traditional strategies to explore the fitness landscape. For instance, Wu et al. proposed a ML-assisted directed protein evolution strategy (MLDE) similar to the active learning pipeline mentioned above [80]; this model can offer data points for a researcher to collect according to simulation results, and the tested data points are employed to update the model. This cycle continues until a desired engineering goal is achieved. MDLE serves as an excellent framework for successfully integrating DE with ML, and it has been found to effectively avoid some local fitness traps or long paths to the global optimum (Fig. 3A). However, MLDE still needs to consider a few design considerations.
Fig. 3.
Typical machine learning (ML) applications in rate-limiting enzyme engineering. A. ML-guided directed evolution. The ML model can offer simulated data points for researchers to collect and test, and the results of these tests generate new data that can be used to update the model. This cycle continues until a desired engineering goal is achieved. B. Deep learning-based encoding strategy. Unsupervised deep learning was used to distill the fundamental features of an individual protein, including its biophysical, structural, and evolutionary information, into a statistical summary. C. Bayesian optimization-assisted protein engineering. A target sequence gradually converges to a specific position in sequence space with an increasing number of cycles of optimization.
There are two main questions in exploring the fitness landscape through ML. (1) How to select the suitable encoding strategy? (2) How to handle low-fitness variants in the fitness landscape? Amino acid sequences must be numerically encoded for the training process [85]. Mutating amino acids to different ones (i.e., in terms of charge and molecular weight) is more likely to affect the structure and function of a protein than mutating it to residues that more closely resemble the original [86]. Thus, researchers generally include such information in ML models via an encoding step to improve the efficiency of the learning phase. Nevertheless, the one-shot encoding strategy adopted in MLDE ignores information regarding the biochemical similarities and differences between amino acids [87]. In addition, fitness landscapes tend to be enriched in zero- or extremely low-fitness variants, which provide minimal information regarding the regions of interest in a landscape. For instance, 92% of regions in the empirically determined four-site combinatorial fitness landscape (204 = 160,000) had fitness values that were below 1% of the global fitness maximum [87]. Thus, the initial sampling of the fitness landscape deserves more consideration. However, MLDE adopted a random sampling protocol to build an initial dataset, which may contain diverse sequences but may provide little information. In any case, the model’s accuracy in predicting low-fitness variants is not critical.
To alleviate these challenges, an improved version of MLDE named ftMLDE was proposed to enhance the encoding strategy and initial sampling [85]. Several alternate encoding methods, including physicochemical encoding and learning models derived from eight natural language processing models, have been evaluated [88], [89]. Furthermore, a novel strategy that leverages global sequence information derived from large sequence databases was employed to reduce uninformative holes in MLDE training sets. Compared with the original MLDE, ftMLDE represents at least a 12.2-fold improvement in searching the fitness landscape. Thus, building a high-quality model of the fitness landscape of a focal protein appears to be feasible by improving the encoding strategy. Moreover, some studies are trying to reduce the exploration of low-fitness regions of the landscape to save resources. Complex design principles are hidden in the amino acid sequence of natural proteins, and by employing ML to understand these principles, it may be feasible to appropriately predict low-fitness variants in the fitness landscape. An unsupervised deep learning model, UniRep, was recently employed to distill the fundamental features of a protein, including biophysical, structural, and evolutionary information, into a statistical summary (Fig. 3C). Moreover, unsupervised deep learning was performed on> 20 million raw amino acid sequences to distill the general features of all functions [90]. Furthermore, a novel framework called Low-N protein engineering has been developed by combining UniRep with active learning. Unlike the studies discussed above, Low-N protein engineering does not rely on high-quality training sets but realizes data dimensionality reduction by incorporating information from numerous proteins to improve the model accuracy; moreover, MCMC-based sampling can help avoid local optimization [90].
In addition, Bayesian optimization has been employed to search chimeric libraries. A recent study successfully enhanced the catalytic rate of alcohol-forming fatty acyl reductases on acyl-ACP substrates using an ML-based protein engineering strategy (Fig. 3B) [15], [91]. Initially, better enzyme performance was achieved through the rearrangement of different subunits of three enzymes from three different sources. To further explore how gene shuffling can enhance fatty alcohol production, the authors of this study constructed an extensive library of rate-limited domains using SCHEMA structure-guided recombination. Next, a Bayesian-based DBTL cycle was used to optimize protein fitness. As the number of cycles increased, the target sequence gradually converged to a specific position in sequence space. In the end, the target strains produced 50% more fatty alcohols than the parental strains.
ML has also been used to reveal the potential underlying protein scaffolds, whereas unsupervised deep learning has been used to learn the semantic grammar within protein sequences deposited in large databases and guide protein engineering. On the other hand, supervised active learning has been applied to model the protein fitness landscape through numerous cycles and identify optimal designs. These strategies can also be combined to further improve protein engineering. However, most of these strategies overemphasize the intelligence of protein engineering, which may lead to situations where we are unable to understand the underlying mechanism of protein engineering.
5. Application of ML in GRE designing
Metabolic pathway optimization is based on essential GREs, such as promoters, ribosome binding sites (RBSs), and terminators. Hence, one of the main challenges of synthetic biology is to artificially design GREs to meet specific requirements [92]. So far, many studies have attempted to build models of the relationship between sequences and the functional properties of proteins. For instance, Jensen et al. used a statistical method to explore the effect of nucleotide position on promoter intensity. Moreover, partial least squares methods were employed to analyze synthetic promoters in E. coli and B. subtilis, respectively [81]. Despite advancements in GRE modeling, these studies are hampered by small datasets, use of single-modeling approaches, and imperfect correlations [93]. Recently, ML has therefore been employed to resolve these problems.
5.1. Promoters
The de novo design of promoters, including both known and novel promoters, has also been facilitated by ML approaches [94]. This has subsequently contributed to optimizing metabolic pathways in systems metabolic engineering. Recently, an ML workflow was developed to integrate mutation-construction-screening-characterization (MCSC) engineering cycles and the XgBoost algorithm to identify relationships between promoter sequences and promoter intensity [95]. In short, a de novo synthetic promoter library was reconstructed and characterized based on high-strength constitutive promoters and broad dynamic range libraries. Next, ML algorithms were used to analyze the relationships between sequences and functions of promoters. MCSC could effectively extend the dynamic range of promoters and provide high-quality data for ML to build models [96]. However, this study was limited by a relatively small library size compared with all possible sequence combinations. In another study, the potential hidden in sequence combinations was analyzed using a generative adversarial network (GAN) to extract features from natural promoters and generate millions of new artificial sequences [97]. GAN learned the design principles of the critical regions of a promoter from the natural sequence, such as k-mer frequency, –10 and –35 motifs, and their spacing constraints. Then, a considerable part of the sequence combination space was filtered out using a GAN. Consequently, 70.8% of AI-designed promoters were experimentally demonstrated to be functional, establishing a new strategy for effectively designing brand-new functional promoters [98].
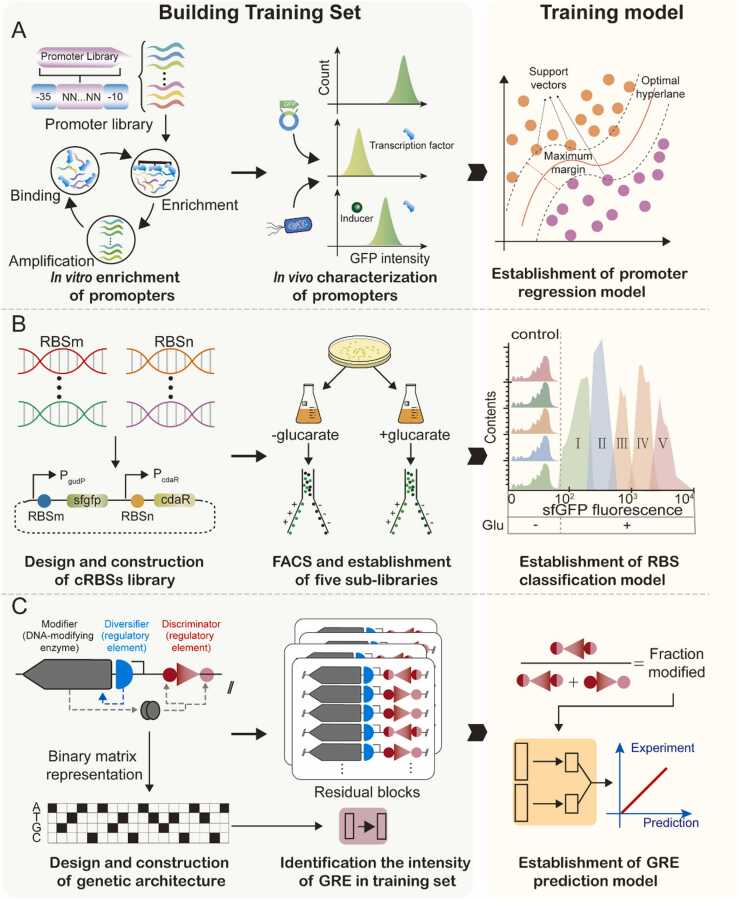
In addition to constitutive promoters, designing inducible promoters from scratch is attractive. For instance, ligand-responsive allosteric transcription factors (aTFs) are essential because they transform biochemical signals into gene expression changes [99]. Thus, accurate control of gene expression may be achieved by manipulating an aTF-regulated promoter. Recently, a novel strategy was proposed to design aTF-regulated promoters from scratch. The innovation of this work lies in two aspects: (1) a high-quality dataset was constructed based on the combination of in vivo and in vitro screening and (2) support vector regression was employed to build models to accurately predict induction ratios (Fig. 4A). Interestingly, the inducible promoters identified in this study were based on a minimal constitutive promoter.
Fig. 4.
Typical machine learning (ML) applications as used in gene regulatory element (GRE) design. These strategies are divided into two steps: generating training sets and training models. A. Establishment of a regression model for promoter intensity. A training dataset was built based on in vitro enrichment and in vivo characterization. Support vector regression was then used to build the model. B. Establishment of a classification model for ribosome binding sites (RBSs). A cross-RBS (c-RBS) library was constructed by combining RBSs and biosensor systems. Next, the total dynamic range of these c-RBSs was divided into five sub-libraries. Finally, a convolutional neural network was used to build a model to classify c-RBSs according to dynamic range. C. A large-scale DNA-based phenotypic recording strategy. A three-compartment genetic architecture was used to record GRE phenotypes. Subsequently, a deep learning model relates sequence to function.
5.2. RBSs
Promoter engineering focuses on regulating the transcriptional level or stability of RNA, i.e., steps that occur long before translation and protein folding. Unlike promoters, RBSs tune gene expression by directly adjusting translation levels and protein folding [100]. Recently, some studies have used ML to elucidate the relationship between RBS sequences and their strength. For example, a Bayesian optimization pipeline was used to improve the translation initiation rate (TIR) of RBSs through DBTL cycles [101]. This study used GPR to model the design space in the learn phase. The upper confidence bound multiarmed bandit algorithm was then applied to balance exploitation and exploration in the design phase. By testing a total of 450 RBS variants from four DBTL cycles, RBSs with high TIR values exceeded their RBS benchmark by up to 34%. However, compared with improving RBS strength, achieving a desired output at an appropriate translation level is more attractive for biosensor systems. Hence, another study attempted to identify the optimal combination of RBS sequences in a biosensor system driven by small molecule responsive transcription factors [92]. This resulted in the construction and characterization of 120,000 cross-RBSs. These biosensor systems were classified into five categories based on their dynamic range (Fig. 4B). A classification model derived from CNNs can accurately predict dynamic ranges based on RBS sequence combinations in a biosensor system by learning the dataset.
Two steps are required to understand the relationship between protein sequence and function: (1) prepare a training dataset and (2) select an ML algorithm to model the sequence space. A recent study provided these methods simultaneously [93]. An innovative HTP approach was initially developed to assign a quantitative functional readout to each specific sequence by constructing a three-component genetic architecture using the same DNA molecule. Thus, GRE sequences and their functional readouts could be read and unambiguously linked from a single sequencing read (Fig. 4C). Further, an approach termed SAPIENs has been developed to quantitatively predict RBS activity and to reliably quantify prediction uncertainty using a residual CNN. Using several million data points from measured translation data, SAPIENs successfully identified a correlation between an RBS sequence and its functional readout [93]. Furthermore, this strategy also showed excellent portability, and may therefore be employed to study the relationship between sequence and function for other GREs (Table 1).
Table 1.
ML tasks and their corresponding algorithms previously applied in metabolic pathway optimization.
| Learning style | Task | Algorithm | Task description | Reference |
|---|---|---|---|---|
| Active learning | Classification/ regression | Bayesian ensemble approach |
|
[72] |
| Bayesian optimization/ Gaussian process (GP) |
|
[71] | ||
| Regression | GP/Upper-confidence bound (UCB) |
|
[93] | |
| Supervised/ unsupervised learning | Clustering/ classification |
K-means/random forest (RF) |
|
[43] |
| Classification/ regression |
Convolutional neural networks (CNNs) |
|
[88], [89] | |
| Supervised learning | Classification | CNNs |
|
[54] |
| CNNs |
|
[61] | ||
| Recurrent neural networks (RNNs) and CNNs |
|
[22] | ||
| RF |
|
[89] | ||
| CNNs |
|
[93] | ||
| Regression | Support vector regression (SVR) |
|
[99] | |
| Artificial neural networks (ANNs) | ●1000 ANNs models are trained with random initial weights to avoid overfitting. | [14] | ||
| Support vector machine (SVM) |
|
[65] | ||
| EVOLVE |
|
[85] | ||
| Residual neural network (ResNet) | ●10 ResNets, each consisting of three residual blocks of two convolutional layers, constitute SAPIENs, which could be explored in RBS activities and sequences. | [92] | ||
| Clustering/ regression |
RF |
|
[84] | |
| Unsupervised learning | Classification | Matrix factorization |
|
[58] |
6. Summary and outlook
Various workflows and frameworks based on ML are rapidly emerging and being applied to solve different types of problems. When sufficient available data exist, ML can replace conventional statistical modeling methods to build accurate models that simulate complex nonlinear processes. For instance, various omics datasets can be fully leveraged to build data-driven models that simulate complex cellular processes. Moreover, the relationship between sequence and function can be precisely characterized by ML using corresponding HTP technologies. When there is insufficient data, ML can also be employed to effectively search design space or landscape. For example, active learning and Bayesian optimization have been applied to identify the optimal combination of gene expression levels, to guide protein engineering, and to redesign GREs. This critical review provides an overview of ML technologies that can be used to guide further metabolic pathway optimization. However, despite the advantages of ML technologies, potential challenges and research gaps may limit the further application of ML for metabolic pathway optimization.
These potential challenges and solutions in metabolic pathway optimization are as follows. First, ML has many applications in genome annotation, gap-filling, and metabolic network refinement. However, no framework or software fully integrates all advanced ML applications at the GEMs construction stage and therefore cannot provide explicit guidance for manual refining. In addition, constructing a whole-cell model using coupling mechanism models is difficult because it is laborious to establish mechanistic models for all subcellular processes. To address these problems, ML methods for model construction tasks should be integrated into a framework or software. Moreover, an active ML framework should be combined with non-ML frameworks to further improve model quality by carrying out limited experiments. Furthermore, it may also be feasible to simulate metabolic processes accurately by constructing a digital twin model. To pursue this goal, ML must be more involved with the construction of GEMs instead of estimating parameters for mechanistic models. Moreover, with advancements in multistep pathway optimization frameworks, the potential of new toolkits based on active learning and Bayesian optimization will be further explored. Nevertheless, identifying targets in most of the studies mentioned above depended to some degree on prior knowledge. Moreover, these targets often belong to the same path, which may excessively restrict the power of ML to optimize metabolic flux on the genome scale. To alleviate these challenges, mechanistic models, such as GEMs, can be integrated into the pathway optimization toolkit to provide more convincing targets. Furthermore, ML has successfully identified untapped potential beneath protein scaffolds. However, most of these strategies overemphasize the intelligence of protein engineering, which may lead to our inability to understand the underlying mechanisms involved in particular protein engineering pathways. To cope with these problems, models of protein structures should be closely combined with the other methods listed above. Through multiple rounds of DBTL, we can continuously improve our understanding of the underlying mechanisms of all enzyme interactions.
In addition, we note that studies using ML for metabolic pathway optimization are not fully integrated, which may limit its applications. For example, the targets of multistep pathway optimization can identify based on prior knowledge rather than by computational simulations. Moreover, ML-guided multistep pathway optimization and enzyme engineering have yet to be combined due to the lack of a global perspective. Furthermore, the data generated from DBTL cycles was not sufficiently leveraged to strengthen the interpretability of this kind of “black box” model. Therefore, to further explore the potential of ML for metabolic pathway optimization, GEMs should be considered a core element during construction of an ML-assisted DBTL cycle (Fig. 5). According to our simulation results, GEMs can provide insight into target identification during multistep pathway optimization. Moreover, fluxomics and protein allocation data generated by simulating large and complex networks can be considered as additional input for active learning or Bayesian optimization to improve the performance of predictive models. Meanwhile, the data generated by DBTL cycles can be employed to update GEMs. Subsequently, the updated GEM will identify the highest-priority enzymes that need to be produced using enzyme cost and flux analysis. Coupling multistep pathway optimization and rate-limiting enzyme engineering with the interpretability of GEMs can therefore help optimize metabolic flux on a system level. This will provide global insights into metabolic pathway optimization. Furthermore, this framework can be further upgraded by adopting automated processes that use robots and high-throughput systems.
Fig. 5.
Prospective tasks for machine learning technologies in metabolic pathway optimization.
CRediT authorship contribution statement
Yang Cheng: Conceptualization, Investigation, Writing − original draft. Xinyu Bi: Supervision, Investigation, Writing − original draft. Yameng Xu: Supervision, Writing − review & editing. Yanfeng Liu: Supervision, Project administration, Writing − review & editing. Jianghua Li: Writing − review & editing. Guocheng Du: Writing − review & editing. Xueqin Lv: Supervision, Project administration, Writing − review & editing. Long Liu: Funding acquisition, Writing − review & editing.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgments
This work was financially supported by the Key Research and Development Program of China (2020YFA0908300), the National Natural Science Foundation of China (32070085), and the Fundamental Research Funds for the Central Universities (JUSRP622004, JUSRP222007).
References
- 1.Liu Y., Nielsen J. Recent trends in metabolic engineering of microbial chemical factories. Curr Opin Biotechnol. 2019;60:188–197. doi: 10.1016/j.copbio.2019.05.010. [DOI] [PubMed] [Google Scholar]
- 2.Nielsen J., Keasling J.D. Engineering cellular metabolism. Cell. 2016;164:1185–1197. doi: 10.1016/J.CELL.2016.02.004. [DOI] [PubMed] [Google Scholar]
- 3.Zhou P., Ye L., Xie W., Lv X., Yu H. Highly efficient biosynthesis of astaxanthin in Saccharomyces cerevisiae by integration and tuning of algal crtZ and bkt. Appl Microbiol Biotechnol. 2015;99:8419–8428. doi: 10.1007/s00253-015-6791-y. [DOI] [PubMed] [Google Scholar]
- 4.Xu Y., Wang X., Zhang C., Zhou X., Xu X., Han L., et al. De novo biosynthesis of rubusoside and rebaudiosides in engineered yeasts. Nat Commun. 2022;13:3040. doi: 10.1038/s41467-022-30826-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Santos C.N.S., Xiao W., Stephanopoulos G. Rational, combinatorial, and genomic approaches for engineering L-tyrosine production in Escherichia coli. Proc Natl Acad Sci. 2012;109:13538–13543. doi: 10.1073/pnas.1206346109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Patra P., B.R.D., Kundu P., Das M., Ghosh A. Recent advances in machine learning applications in metabolic engineering. Biotechnol Adv. 2023;62 doi: 10.1016/j.biotechadv.2022.108069. [DOI] [PubMed] [Google Scholar]
- 7.Hodgman C.E., Jewett M.C. Cell-free synthetic biology: Thinking outside the cell. Metab Eng. 2012;14:261–269. doi: 10.1016/j.ymben.2011.09.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Lawson C.E., Harcombe W.R., Hatzenpichler R., Lindemann S.R., Löffler F.E., O’Malley M.A., et al. Common principles and best practices for engineering microbiomes. Nat Rev Microbiol. 2019;17:725–741. doi: 10.1038/s41579-019-0255-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Islam M.R., Tudryn G., Bucinell R., Schadler L., Picu R.C. Publisher Correction: Morphology and mechanics of fungal mycelium. Sci Rep. 2018;8:4206. doi: 10.1038/s41598-018-20637-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Hastings A., Byers J.E., Crooks J.A., Cuddington K., Jones C.G., Lambrinos J.G., et al. Ecosystem engineering in space and time. Ecol Lett. 2007;10:153–164. doi: 10.1111/j.1461-0248.2006.00997.x. [DOI] [PubMed] [Google Scholar]
- 11.Ma E.J., Siirola E., Moore C., Kummer A., Stoeckli M., Faller M., et al. Machine-directed evolution of an imine reductase for activity and stereoselectivity. ACS Catal. 2021;11:12433–12445. doi: 10.1021/acscatal.1c02786. [DOI] [Google Scholar]
- 12.Sakr G.E., Mokbel M., Darwich A., Khneisser M.N., Hadi A. Comparing deep learning and support vector machines for autonomous waste sorting. 2016 IEEE International Multidisciplinary Conference on Engineering Technology (IMCET), IEEE; 2016, p. 207–212. https://doi.org/10.1109/IMCET.2016.7777453.
- 13.Eitzinger S., Asif A., Watters K.E., Iavarone A.T., Knott G.J., Doudna J.A., et al. Machine learning predicts new anti-CRISPR proteins. Nucleic Acids Res. 2020;48:4698–4708. doi: 10.1093/nar/gkaa219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Zhou Y., Li G., Dong J., Xing X., Dai J., Zhang C. MiYA, an efficient machine-learning workflow in conjunction with the YeastFab assembly strategy for combinatorial optimization of heterologous metabolic pathways in Saccharomyces cerevisiae. Metab Eng. 2018;47:294–302. doi: 10.1016/j.ymben.2018.03.020. [DOI] [PubMed] [Google Scholar]
- 15.Greenhalgh J.C., Fahlberg S.A., Pfleger B.F., Romero P.A. Machine learning-guided acyl-ACP reductase engineering for improved in vivo fatty alcohol production. Nat Commun. 2021;12:5825. doi: 10.1038/s41467-021-25831-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Camacho D.M., Collins K.M., Powers R.K., Costello J.C., Collins J.J. Next-generation machine learning for biological networks. Cell. 2018;173:1581–1592. doi: 10.1016/j.cell.2018.05.015. [DOI] [PubMed] [Google Scholar]
- 17.Rana P., Berry C., Ghosh P., Fong S.S. Recent advances on constraint-based models by integrating machine learning. Curr Opin Biotechnol. 2020;64:85–91. doi: 10.1016/j.copbio.2019.11.007. [DOI] [PubMed] [Google Scholar]
- 18.Fang X., Lloyd C.J., Palsson B.O. Reconstructing organisms in silico: genome-scale models and their emerging applications. Nat Rev Microbiol. 2020;18:731–743. doi: 10.1038/s41579-020-00440-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Orth J.D., Thiele I., Palsson B.Ø. What is flux balance analysis. Nat Biotechnol. 2010;28:245–248. doi: 10.1038/nbt.1614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Zampieri G., Vijayakumar S., Yaneske E., Angione C. Machine and deep learning meet genome-scale metabolic modeling. PLoS Comput Biol. 2019;15 doi: 10.1371/journal.pcbi.1007084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Sánchez B.J., Zhang C., Nilsson A., Lahtvee P., Kerkhoven E.J., Nielsen J. Improving the phenotype predictions of a yeast genome‐scale metabolic model by incorporating enzymatic constraints. Mol Syst Biol. 2017;13:935. doi: 10.15252/msb.20167411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Lu H., Kerkhoven E.J., Nielsen J. Multiscale models quantifying yeast physiology: towards a whole-cell model. Trends Biotechnol. 2022;40:291–305. doi: 10.1016/j.tibtech.2021.06.010. [DOI] [PubMed] [Google Scholar]
- 23.Gu C., Kim G.B., Kim W.J., Kim H.U., Lee S.Y. Current status and applications of genome-scale metabolic models. Genome Biol. 2019;20:121. doi: 10.1186/s13059-019-1730-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Kocabaş P., Çalık P., Çalık G., Özdamar T.H. Analyses of extracellular protein production in Bacillus subtilis – I: Genome-scale metabolic model reconstruction based on updated gene-enzyme-reaction data. Biochem Eng J. 2017;127:229–241. doi: 10.1016/j.bej.2017.07.005. [DOI] [Google Scholar]
- 25.Feierabend M., Renz A., Zelle E., Nöh K., Wiechert W., Dräger A. High-Quality Genome-Scale Reconstruction of Corynebacterium glutamicum ATCC 13032. Front Microbiol. 2021:12. doi: 10.3389/fmicb.2021.750206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zhang Y., Cai J., Shang X., Wang B., Liu S., Chai X., et al. A new genome-scale metabolic model of Corynebacterium glutamicum and its application. Biotechnol Biofuels. 2017;10:169. doi: 10.1186/s13068-017-0856-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Becker J., Zelder O., Häfner S., Schröder H., Wittmann C. From zero to hero—Design-based systems metabolic engineering of Corynebacterium glutamicum for l-lysine production. Metab Eng. 2011;13:159–168. doi: 10.1016/j.ymben.2011.01.003. [DOI] [PubMed] [Google Scholar]
- 28.Chemler J.A., Fowler Z.L., McHugh K.P., Koffas M.A.G. Improving NADPH availability for natural product biosynthesis in Escherichia coli by metabolic engineering. Metab Eng. 2010;12:96–104. doi: 10.1016/j.ymben.2009.07.003. [DOI] [PubMed] [Google Scholar]
- 29.Qian Z.-G., Xia X.-X., Lee S.Y. Metabolic engineering of Escherichia coli for the production of cadaverine: A five carbon diamine. Biotechnol Bioeng. 2011;108:93–103. doi: 10.1002/bit.22918. [DOI] [PubMed] [Google Scholar]
- 30.Kim G.B., Kim W.J., Kim H.U., Lee S.Y. Machine learning applications in systems metabolic engineering. Curr Opin Biotechnol. 2020;64:1–9. doi: 10.1016/j.copbio.2019.08.010. [DOI] [PubMed] [Google Scholar]
- 31.Lawson C.E., Martí J.M., Radivojevic T., Jonnalagadda S.V.R., Gentz R., Hillson N.J., et al. Machine learning for metabolic engineering: A review. Metab Eng. 2021;63:34–60. doi: 10.1016/j.ymben.2020.10.005. [DOI] [PubMed] [Google Scholar]
- 32.Thiele I., Palsson B.Ø. A protocol for generating a high-quality genome-scale metabolic reconstruction. Nat Protoc. 2010;5:93–121. doi: 10.1038/nprot.2009.203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Clauwaert J., Menschaert G., Waegeman W. DeepRibo: a neural network for precise gene annotation of prokaryotes by combining ribosome profiling signal and binding site patterns. Nucleic Acids Res. 2019;47 doi: 10.1093/nar/gkz061. –e36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Ryu J.Y., Kim H.U., Lee S.Y. Deep learning enables high-quality and high-throughput prediction of enzyme commission numbers. Proceedings of the National Academy of Sciences, 2019;116:13996–14001. https://doi.org/10.1073/pnas.1821905116. [DOI] [PMC free article] [PubMed]
- 35.Pan S., Reed J.L. Advances in gap-filling genome-scale metabolic models and model-driven experiments lead to novel metabolic discoveries. Curr Opin Biotechnol. 2018;51:103–108. doi: 10.1016/j.copbio.2017.12.012. [DOI] [PubMed] [Google Scholar]
- 36.Dale J.M., Popescu L., Karp P.D. Machine learning methods for metabolic pathway prediction. BMC Bioinforma. 2010;11:15. doi: 10.1186/1471-2105-11-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Machado D., Andrejev S., Tramontano M., Patil K.R. Fast automated reconstruction of genome-scale metabolic models for microbial species and communities. Nucleic Acids Res. 2018;46:7542–7553. doi: 10.1093/nar/gky537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Dias O., Rocha M., Ferreira E.C., Rocha I. Reconstr High-Qual Large-Scale Metab Models merlin. 2018:1–36. doi: 10.1007/978-1-4939-7528-0_1. [DOI] [PubMed] [Google Scholar]
- 39.Henry C.S., DeJongh M., Best A.A., Frybarger P.M., Linsay B., Stevens R.L. High-throughput generation, optimization and analysis of genome-scale metabolic models. Nat Biotechnol. 2010;28:977–982. doi: 10.1038/nbt.1672. [DOI] [PubMed] [Google Scholar]
- 40.Karp P.D., Latendresse M., Paley S.M., Krummenacker M., Ong Q.D., Billington R., et al. Pathway Tools version 19.0 update: software for pathway/genome informatics and systems biology. Brief Bioinform. 2016;17:877–890. doi: 10.1093/bib/bbv079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Zimmermann J., Kaleta C., Waschina S. gapseq: informed prediction of bacterial metabolic pathways and reconstruction of accurate metabolic models. Genome Biol. 2021;22:81. doi: 10.1186/s13059-021-02295-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Machado D., Andrejev S., Tramontano M., Patil K.R. Fast automated reconstruction of genome-scale metabolic models for microbial species and communities. Nucleic Acids Res. 2018;46:7542–7553. doi: 10.1093/nar/gky537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Biggs M.B., Papin J.A. Managing uncertainty in metabolic network structure and improving predictions using EnsembleFBA. PLoS Comput Biol. 2017;13 doi: 10.1371/journal.pcbi.1005413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Oftadeh O., Salvy P., Masid M., Curvat M., Miskovic L., Hatzimanikatis V. A genome-scale metabolic model of Saccharomyces cerevisiae that integrates expression constraints and reaction thermodynamics. Nat Commun. 2021;12:4790. doi: 10.1038/s41467-021-25158-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Österberg L., Domenzain I., Münch J., Nielsen J., Hohmann S., Cvijovic M. A novel yeast hybrid modeling framework integrating Boolean and enzyme-constrained networks enables exploration of the interplay between signaling and metabolism. PLoS Comput Biol. 2021;17 doi: 10.1371/journal.pcbi.1008891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Wang Z., Danziger S.A., Heavner B.D., Ma S., Smith J.J., Li S., et al. Combining inferred regulatory and reconstructed metabolic networks enhances phenotype prediction in yeast. PLoS Comput Biol. 2017;13 doi: 10.1371/journal.pcbi.1005489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Allen F., Greiner R., Wishart D. Competitive fragmentation modeling of ESI-MS/MS spectra for putative metabolite identification. Metabolomics. 2015;11:98–110. doi: 10.1007/s11306-014-0676-4. [DOI] [Google Scholar]
- 48.Nguyen D.H., Nguyen C.H., Mamitsuka H. Recent advances and prospects of computational methods for metabolite identification: a review with emphasis on machine learning approaches. Brief Bioinform. 2019;20:2028–2043. doi: 10.1093/bib/bby066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Bi X., Liu Y., Li J., Du G., Lv X., Liu L. Construction of multiscale genome-scale metabolic models: frameworks and challenges. Biomolecules. 2022;12:721. doi: 10.3390/biom12050721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Heckmann D., Lloyd C.J., Mih N., Ha Y., Zielinski D.C., Haiman Z.B., et al. Machine learning applied to enzyme turnover numbers reveals protein structural correlates and improves metabolic models. Nat Commun. 2018;9:5252. doi: 10.1038/s41467-018-07652-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Chen Y., Nielsen J. Mathematical modeling of proteome constraints within metabolism. Curr Opin Syst Biol. 2021;25:50–56. doi: 10.1016/j.coisb.2021.03.003. [DOI] [Google Scholar]
- 52.Nilsson A., Nielsen J., Palsson B.O. Metabolic models of protein allocation call for the kinetome. Cell Syst. 2017;5:538–541. doi: 10.1016/j.cels.2017.11.013. [DOI] [PubMed] [Google Scholar]
- 53.Davidi D., Noor E., Liebermeister W., Bar-Even A., Flamholz A., Tummler K., et al. Global characterization of in vivo enzyme catalytic rates and their correspondence to in vitro kcat measurements. Proc Natl Acad Sci. 2016;113:3401–3406. doi: 10.1073/pnas.1514240113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Heckmann D., Campeau A., Lloyd C.J., Phaneuf P.V., Hefner Y., Carrillo-Terrazas M., et al. Kinetic profiling of metabolic specialists demonstrates stability and consistency of in vivo enzyme turnover numbers. Proc Natl Acad Sci. 2020;117:23182–23190. doi: 10.1073/pnas.2001562117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Li F., Yuan L., Lu H., Li G., Chen Y., Engqvist M.K.M., et al. Deep learning-based kcat prediction enables improved enzyme-constrained model reconstruction. Nat Catal. 2022;5:662–672. doi: 10.1038/s41929-022-00798-z. [DOI] [Google Scholar]
- 56.Flamholz A., Noor E., Bar-Even A., Milo R. eQuilibrator--the biochemical thermodynamics calculator. Nucleic Acids Res. 2012;40:D770–D775. doi: 10.1093/nar/gkr874. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Heinonen M., Shen H., Zamboni N., Rousu J. Metabolite identification and molecular fingerprint prediction through machine learning. Bioinformatics. 2012;28:2333–2341. doi: 10.1093/bioinformatics/bts437. [DOI] [PubMed] [Google Scholar]
- 58.Dührkop K., Shen H., Meusel M., Rousu J., Böcker S. Searching molecular structure databases with tandem mass spectra using CSI:FingerID. Proc Natl Acad Sci. 2015;112:12580–12585. doi: 10.1073/pnas.1509788112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Nguyen D.H., Nguyen C.H., Mamitsuka H. SIMPLE: Sparse Interaction Model over Peaks of moLEcules for fast, interpretable metabolite identification from tandem mass spectra. Bioinformatics. 2018;34:i323–i332. doi: 10.1093/bioinformatics/bty252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Kwon M.S., Lee B.T., Lee S.Y., Kim H.U. Modeling regulatory networks using machine learning for systems metabolic engineering. Curr Opin Biotechnol. 2020;65:163–170. doi: 10.1016/j.copbio.2020.02.014. [DOI] [PubMed] [Google Scholar]
- 61.Zhao M., He W., Tang J., Zou Q., Guo F. A hybrid deep learning framework for gene regulatory network inference from single-cell transcriptomic data. Brief Bioinform. 2022:23. doi: 10.1093/bib/bbab568. [DOI] [PubMed] [Google Scholar]
- 62.Yuan Y., Bar-Joseph Z. Deep learning for inferring gene relationships from single-cell expression data. Proc Natl Acad Sci. 2019;116:27151–27158. doi: 10.1073/pnas.1911536116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Bottoms S., Dickinson Q., McGee M., Hinchman L., Higbee A., Hebert A., et al. Chemical genomic guided engineering of gamma-valerolactone tolerant yeast. Micro Cell Fact. 2018;17:5. doi: 10.1186/s12934-017-0848-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Skerker J.M., Leon D., Price M.N., Mar J.S., Tarjan D.R., Wetmore K.M., et al. Dissecting a complex chemical stress: chemogenomic profiling of plant hydrolysates. Mol Syst Biol. 2013;9:674. doi: 10.1038/msb.2013.30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Kumar P., Adamczyk P.A., Zhang X., Andrade R.B., Romero P.A., Ramanathan P., et al. Active and machine learning-based approaches to rapidly enhance microbial chemical production. Metab Eng. 2021;67:216–226. doi: 10.1016/j.ymben.2021.06.009. [DOI] [PubMed] [Google Scholar]
- 66.Chandrasekaran S., Price N.D. Probabilistic integrative modeling of genome-scale metabolic and regulatory networks in Escherichia coli and Mycobacterium tuberculosis. Proc Natl Acad Sci. 2010;107:17845–17850. doi: 10.1073/pnas.1005139107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Ascher S., Wang X., Watson I., Sloan W., You S. Interpretable machine learning to model biomass and waste gasification. Bioresour Technol. 2022;364 doi: 10.1016/j.biortech.2022.128062. [DOI] [PubMed] [Google Scholar]
- 68.Yuan T., Guo Y., Dong J., Li T., Zhou T., Sun K., et al. Construction, characterization and application of a genome-wide promoter library in Saccharomyces cerevisiae. Front Chem Sci Eng. 2017;11:107–116. doi: 10.1007/s11705-017-1621-7. [DOI] [Google Scholar]
- 69.Opgenorth P., Costello Z., Okada T., Goyal G., Chen Y., Gin J., et al. Lessons from Two Design–Build–Test–Learn Cycles of Dodecanol Production in Escherichia coli Aided by Machine Learning. ACS Synth Biol. 2019;8:1337–1351. doi: 10.1021/acssynbio.9b00020. [DOI] [PubMed] [Google Scholar]
- 70.Kushner H.J. A new method of locating the maximum point of an arbitrary multipeak curve in the presence of noise. J Basic Eng. 1964;86:97–106. doi: 10.1115/1.3653121. [DOI] [Google Scholar]
- 71.HamediRad M., Chao R., Weisberg S., Lian J., Sinha S., Zhao H. Towards a fully automated algorithm driven platform for biosystems design. Nat Commun. 2019;10:5150. doi: 10.1038/s41467-019-13189-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Radivojević T., Costello Z., Workman K., Garcia Martin H. A machine learning Automated Recommendation Tool for synthetic biology. Nat Commun. 2020;11:4879. doi: 10.1038/s41467-020-18008-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Hu R., Fu L., Chen Y., Chen J., Qiao Y., Si T. Protein engineering via Bayesian optimization-guided evolutionary algorithm and robotic experiments. Brief Bioinform. 2023:24. doi: 10.1093/bib/bbac570. [DOI] [PubMed] [Google Scholar]
- 74.Mazurenko S., Prokop Z., Damborsky J. Machine learning in enzyme engineering. ACS Catal. 2020;10:1210–1223. doi: 10.1021/acscatal.9b04321. [DOI] [Google Scholar]
- 75.Jumper J., Evans R., Pritzel A., Green T., Figurnov M., Ronneberger O., et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021;596:583–589. doi: 10.1038/s41586-021-03819-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Yang K.K., Wu Z., Arnold F.H. Machine-learning-guided directed evolution for protein engineering. Nat Methods. 2019;16:687–694. doi: 10.1038/s41592-019-0496-6. [DOI] [PubMed] [Google Scholar]
- 77.Bryant P., Pozzati G., Zhu W., Shenoy A., Kundrotas P., Elofsson A. Predicting the structure of large protein complexes using AlphaFold and Monte Carlo tree search. Nat Commun. 2022;13:6028. doi: 10.1038/s41467-022-33729-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Cui Y., Sun J., Wu B. Computational enzyme redesign: large jumps in function. Trends Chem. 2022;4:409–419. doi: 10.1016/j.trechm.2022.03.001. [DOI] [Google Scholar]
- 79.Lovelock S.L., Crawshaw R., Basler S., Levy C., Baker D., Hilvert D., et al. The road to fully programmable protein catalysis. Nature. 2022;606:49–58. doi: 10.1038/s41586-022-04456-z. [DOI] [PubMed] [Google Scholar]
- 80.Wu Z., Kan S.B.J., Lewis R.D., Wittmann B.J., Arnold F.H. Machine learning-assisted directed protein evolution with combinatorial libraries. Proc Natl Acad Sci. 2019;116:8852–8858. doi: 10.1073/pnas.1901979116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Fowler D.M., Araya C.L., Fleishman S.J., Kellogg E.H., Stephany J.J., Baker D., et al. High-resolution mapping of protein sequence-function relationships. Nat Methods. 2010;7:741–746. doi: 10.1038/nmeth.1492. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Goldsmith M., Tawfik D.S. Enzyme engineering: reaching the maximal catalytic efficiency peak. Curr Opin Struct Biol. 2017;47:140–150. doi: 10.1016/j.sbi.2017.09.002. [DOI] [PubMed] [Google Scholar]
- 83.Romero P.A., Arnold F.H. Exploring protein fitness landscapes by directed evolution. Nat Rev Mol Cell Biol. 2009;10:866–876. doi: 10.1038/nrm2805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Borkowski O., Koch M., Zettor A., Pandi A., Batista A.C., Soudier P., et al. Large scale active-learning-guided exploration for in vitro protein production optimization. Nat Commun. 2020;11:1872. doi: 10.1038/s41467-020-15798-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Wittmann B.J., Yue Y., Arnold F.H. Informed training set design enables efficient machine learning-assisted directed protein evolution. Cell Syst. 2021;12:1026–1045.e7. doi: 10.1016/j.cels.2021.07.008. [DOI] [PubMed] [Google Scholar]
- 86.Romero P.A., Krause A., Arnold F.H. Navigating the protein fitness landscape with Gaussian processes. Proc Natl Acad Sci. 2013:110. doi: 10.1073/pnas.1215251110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Wu N.C., Dai L., Olson C.A., Lloyd-Smith J.O., Sun R. Adaptation in protein fitness landscapes is facilitated by indirect paths. Elife. 2016:5. doi: 10.7554/eLife.16965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Georgiev A.G. Interpretable numerical descriptors of amino acid space. J Comput Biol. 2009;16:703–723. doi: 10.1089/cmb.2008.0173. [DOI] [PubMed] [Google Scholar]
- 89.Rives A., Meier J., Sercu T., Goyal S., Lin Z., Liu J., et al. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Proc Natl Acad Sci. 2021:118. doi: 10.1073/pnas.2016239118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Biswas S., Khimulya G., Alley E.C., Esvelt K.M., Church G.M. Low-N protein engineering with data-efficient deep learning. Nat Methods. 2021;18:389–396. doi: 10.1038/s41592-021-01100-y. [DOI] [PubMed] [Google Scholar]
- 91.Silberg J.J., Endelman J.B., Arnold F.H. SCHEMA-Guide Protein Recomb. 2004:35–42. doi: 10.1016/S0076-6879(04)88004-2. [DOI] [PubMed] [Google Scholar]
- 92.Ding N., Yuan Z., Zhang X., Chen J., Zhou S., Deng Y. Programmable cross-ribosome-binding sites to fine-tune the dynamic range of transcription factor-based biosensor. Nucleic Acids Res. 2020;48:10602–10613. doi: 10.1093/nar/gkaa786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Höllerer S., Papaxanthos L., Gumpinger A.C., Fischer K., Beisel C., Borgwardt K., et al. Large-scale DNA-based phenotypic recording and deep learning enable highly accurate sequence-function mapping. Nat Commun. 2020:11. doi: 10.1038/s41467-020-17222-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Gilman J., Singleton C., Tennant R.K., James P., Howard T.P., Lux T., et al. Rapid, Heuristic Discovery and Design of Promoter Collections in Non-Model Microbes for Industrial Applications. ACS Synth Biol. 2019;8:1175–1186. doi: 10.1021/acssynbio.9b00061. [DOI] [PubMed] [Google Scholar]
- 95.Zhao M., Yuan Z., Wu L., Zhou S., Deng Y. Precise Prediction of Promoter Strength Based on a De Novo Synthetic Promoter Library Coupled with Machine Learning. ACS Synth Biol. 2022;11:92–102. doi: 10.1021/acssynbio.1c00117. [DOI] [PubMed] [Google Scholar]
- 96.Chen T., Guestrin C.X.G.Boost. Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, New York, NY, USA: ACM; 2016, p. 785–794. https://doi.org/10.1145/2939672.2939785.
- 97.Creswell A., White T., Dumoulin V., Arulkumaran K., Sengupta B., Bharath A.A. Generative adversarial networks: an overview. IEEE Signal Process Mag. 2018;35:53–65. doi: 10.1109/MSP.2017.2765202. [DOI] [Google Scholar]
- 98.Wang Y., Wang H., Wei L., Li S., Liu L., Wang X. Synthetic promoter design in Escherichia coli based on a deep generative network. Nucleic Acids Res. 2020;48:6403–6412. doi: 10.1093/nar/gkaa325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Liu X., Gupta S.T.P., Bhimsaria D., Reed J.L., Rodríguez-Martínez J.A., Ansari A.Z., et al. De novo design of programmable inducible promoters. Nucleic Acids Res. 2019;47:10452–10463. doi: 10.1093/nar/gkz772. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Groher A.-C., Jager S., Schneider C., Groher F., Hamacher K., Suess B. Tuning the performance of synthetic riboswitches using machine learning. ACS Synth Biol. 2019;8:34–44. doi: 10.1021/acssynbio.8b00207. [DOI] [PubMed] [Google Scholar]
- 101.Zhang M., Holowko M.B., Hayman Zumpe H., Ong C.S. Machine learning guided batched design of a bacterial ribosome binding site. ACS Synth Biol. 2022;11:2314–2326. doi: 10.1021/acssynbio.2c00015. [DOI] [PMC free article] [PubMed] [Google Scholar]