Abstract
The liver is the largest glandular organ in the body and has a unique distribution of cells and biomolecules. However, the treatment outcome of end-stage liver disease is extremely poor. Single-cell sequencing is a new advanced and powerful technique for identifying rare cell populations and biomolecules by analyzing the characteristics of gene expression between individual cells. These cells and biomolecules might be used as potential targets for immunotherapy of liver diseases and contribute to the development of precise individualized treatment. Compared to whole-tissue RNA sequencing, single-cell RNA sequencing (scRNA-seq) or other single-cell histological techniques have solved the problem of cell population heterogeneity and characterize molecular changes associated with liver diseases with higher accuracy and resolution. In this review, we comprehensively summarized single-cell approaches including transcriptomic, spatial transcriptomic, immunomic, proteomic, epigenomic, and multiomic technologies, and described their application in liver physiology and pathology. We also discussed advanced techniques and recent studies in the field of single-cell; our review might provide new insights into the pathophysiological mechanisms of the liver to achieve precise and individualized treatment of liver diseases.
Keywords: Single-cell technology, Normal liver, Chronic liver disease, Hepatocellular carcinoma, Acute-on-chronic liver failure
Graphical abstract
Introduction
Liver diseases have become a global health burden in recent decades. For example, cirrhosis causes 1.16 million deaths, and liver cancer causes 0.788 million deaths annually, making them the 11th and 16th most common causes of death.1 More than one-fifth of the Chinese population is affected by liver diseases, particularly hepatitis B viral infections, while cases of nonalcoholic hepatitis and autoimmune hepatitis are increasing.2 Persistent chronic inflammation in the liver is closely associated with cirrhosis and ultimately leads to liver cancer or liver failure, and the main cause of acute liver failure in developing countries is viral hepatitis.3,4 However, effective treatment options for liver diseases are so limited that liver transplantation is regarded as the only viable solution, but its demand fairly exceeds the supply.1 Therefore, better tools are urgently required to elucidate the pathological mechanism and development of liver diseases with a clearer perspective. Their application may contribute to precise, personalized therapy and a decrease in mortality.
Each cell is unique for it occupies an exclusive position in space, carries distinct errors in its copied genome and is subject to programmed and induced changes in gene expression. Yet traditional sequencing is performed on tissue samples or cell populations, in which biological differences between cells can be obscured by averaging or mistaken for technical noise. In contrast, single-cell sequencing technology can isolate several thousand single cells at one time and has the advantage of detecting heterogeneity among individual cells then depicting cellular profiles, which can help us identify the rare populations of cells that have derived significance for disease progression with unprecedented high resolution.5 The workflow of single-cell sequencing experiments and analyses generally includes single-cell suspension preparation, single-cell sorting, amplification and library preparation, high-throughput sequencing, and bioinformatics analysis (Fig. 1). Tang et al.6 proposed the first single-cell transcriptome sequencing (scRNA-seq) technology in 2009. Since then, the single-cell field has advanced rapidly, and several new approaches have been developed, including genome, transcriptome, proteome, spatial transcriptome, epigenome. Single-cell sequencing can not only analyze the heterogeneity of cells with the same phenotype but also obtain the genetic information of hard-to-culture microorganisms and valuable clinical samples.
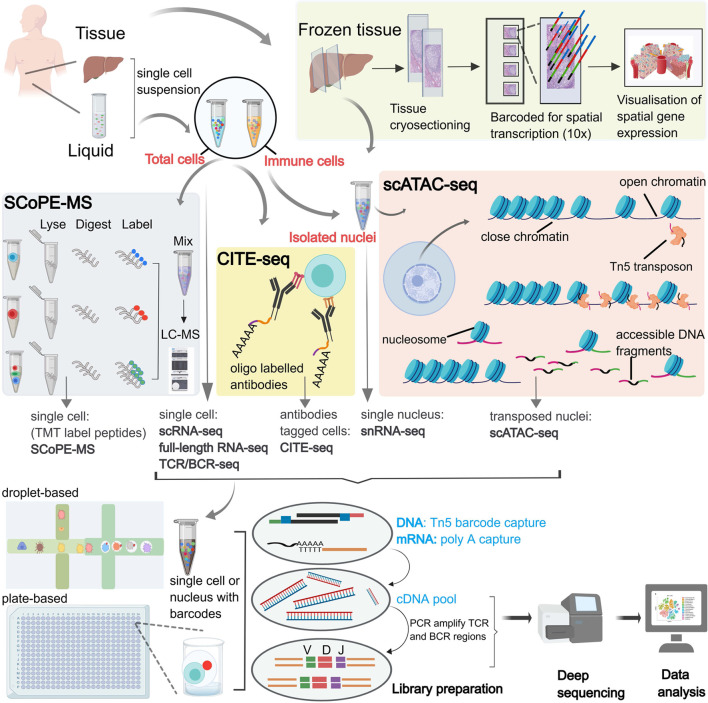
Fig. 1. Flow diagram of different single-cell approaches in clinical liver samples.
Single-cell suspension preparation: single-cell comes from fresh samples including liquid (peripheral blood, ascites) and tissue (liver cancer, adjacent tumor, normal liver) by dissociation; frozen samples can be used to snRNA-seq and ST. ST: frozen liver tissue samples are sectioned, fixed, and permeated, then transcripts are barcoded according to their location based on a matrix of spots. These barcodes are then used to show the spatial gene signatures across the tissue section. SCoPE-MS: surviving single-cell is lysed by ultrasound, proteins are extracted from the lysate then trypsinized, and peptides from multiple samples are identified and quantified using TMTs. CITE-seq: a technology consisting of mRNA and tagged antibody sequencing, which enables the simultaneous determination of unbiased transcriptional profiles and protein markers based on oligo-tagged antibody detection in a cell. ScATAC-seq: a technique for studying open chromatin regions with epigenetic and gene regulatory roles at the single-cell level. Tn5 transposase is used to cleave accessible DNA regions of open chromatin in the nuclear suspension of sorted cells, followed by post-PCR sequencing using tags of known sequence. Cell sorting: the platforms used for cell sorting are mainly based on microfluidic or cellular plate technologies, both of which allow for unbiased quantification of gene expression with the aid of unique molecular identifier and cell label technology, where mRNA expression profiles of TCR and BCR can be determined in T and B cells for adaptive immune response analysis. 10×,10× Genomics; BCR, B-cell receptor; CITE-seq, cellular indexing of transcriptome and epitopes by sequencing; PCR, polymerase chain reaction; scATAC-seq, single-cell assay for transposase-accessible chromatin with high throughput sequencing; SCoPE-MS, single-cell proteomics by mass spectrometry; scRNA-seq, single-cell RNA sequencing; snRNA-seq, single nuclear sequencing; ST, spatial transcriptome technology; TMTs, tandem mass spectrometry tags; TCR, T-cell receptor.
In this article, we review the current single-cell approaches and discussed their key principles along with their advantages and disadvantages to understand which method is more suitable under what kind of circumstances (Table 1).7–23 We also discuss the novel application of those techniques in studies on liver physiology and pathology and the application prospects and technical improvements in single-cell technology to find effective individualized treatment strategies for patients with liver diseases and reduce the overall morbidity and mortality related to liver diseases.
Table 1. Summary of single-cell analysis methods.
| Single-cell histology | Application | Single-cell sequencing | Advantages | Disadvantages | Ref. |
|---|---|---|---|---|---|
| Transcriptome | Cell type discovery and characterization in diseases; cell development trajectory and plasticity | Smart-seq | Full-length coverage across transcripts | Unstable; transcript bias | 20 |
| Smart-seq2 | Captures rare genes; commercial off-the-shelf SMARTer kit | Can not analyze nonpolyA RNAs; nonstrand specificity | 8 | ||
| Drop-seq | High throughput; low cost | Low sensitivity | 21 | ||
| 10× Genomics Chromium | High throughput; high sensitivity; short cycle time; low technical noise | High cost | 22 | ||
| snRNA-seq | Allow for frozen samples; relatively simple to operate | Few genes detected; cellular preference | 7 | ||
| Spatial transcriptome | Spatial distribution of heterogeneous cells and spatial patterns of gene expression in different tissues; tumor microenvironment | 10× Visium | Commercial platform is mature and stable; widely used; with supporting verification platform | Low spatial resolution; non-single cell level | 9 |
| GeoMx DSP | Compatible with FFPE samples and fresh frozen samples; for in-depth analysis of local tissues | Low throughput | 10 | ||
| DBiT-seq | High spatial resolution; co-localization of mRNA and protein; simplified workflow | Inability to readily identify cell types | 11 | ||
| Proteome | Disease heterogeneity and treatment resistance; cancer immunotherapy; high throughput drug screening | scWestern | High specificity; | Low protein flux | 23 |
| SCoPE-MS | Accurately differentiate the proteome of different cells | Low protein flux Complex workflow | 12 | ||
| nanoPOTS | Microprotein identification; high quantitative accuracy and protein coverage | 13 | |||
| Epigenome | Genetic characteristics of disease; deeper understanding of gene regulation from open chromatin; infer development trajectories and identify regulatory elements | scCGI-seq | High accuracy; simplified workflow | Low coverage | 14 |
| CUT & RUN technique | Low sample requirement; high resolution and accuracy; commercial kits | Highly antibody specificity-dependent | 17 | ||
| scATAC-seq | High resolution, sensitivity, and accuracy | Low recovery of DNA fragments | 15 | ||
| scChIP-seq | High throughput, high coverage | Highly antibody-dependent | 16 | ||
| Multiomics | Integrated analysis different histological techniques to characterize the state of gene regulation more comprehensively within cells and reveal the mechanisms of disease regulation | scTrio-seq | Simultaneous analysis of genome,transcriptome, and epigenome | Low throughput | 18 |
| CITE-seq | Simultaneous analysis of transcriptome and protein expression; high throughput | Only surface proteins can be measured | 19 |
CITE-seq, cellular indexing of transcriptomes and epitopes by sequencing; CUT & RUN, cleavage under targets and release using nuclease; FFPE, formalin-fixed and paraffin embedded; nanoPOTS, nanodroplet processing in one pot for trace samples; scATAC-seq, single-cell assay for transposase-accessible chromatin using sequencing; scCGI-seq, single-cell CpG island methylation sequencing; scChIP-seq, single-cell chromatin immune-precipitation sequencing; scCOOL-seq, single-cell chromatin overall omic-scale landscape sequencing; SCoPE-MS, single-cell proteomics by mass spectrometry; scTrio-seq, single-cell triple omics sequencing; scWestern, single-cell western blotting; snRNA-seq, single-nucleus RNA sequencing.
An overview of single-cell technologies
scRNA-seq
scRNA-seq measures gene expression at the single-cell level and reveals heterogeneity of gene expression in individual cells or homologous cell types, its core principle refers to tag-based identification of single cells. A unique cell tag attached to a barcode is added to each cell when reverse transcription is performed prior to RNA sequencing. Therefore, even if all cells are mixed and sequenced, RNA fragments carrying the same barcode can be considered as coming from the same cell. By building a library once, we can measure the information of tens of thousands of single cells (Fig. 1). A challenge in conducting research using scRNA-seq is that high quality suspensions based on fresh tissue are not available for all tissues. In contrast, single-nucleus RNA sequencing (snRNA-seq)7 is not limited by dissociation conditions and can be used for analyzing cell and gene expression of primary tissues obtained from frozen samples. Thus, this process complements scRNA-seq, but the nuclear sequencing depth is low. Some studies have compared the differences in sample type, study purpose, and target cell population between scRNA-seq and snRNA-seq for various tissues and diseases.24–26
In addition, immune cell receptor sequencing, which focuses on immune T and B cells, can comprehensively resolve T-cell receptor/B-cell receptor (TCR/BCR) gene rearrangement base sequences and the abundance of each sequence to reveal the mechanism of T/B cell-mediated humoral immune response. Single-cell analysis was first used for examining immune cells around 2013 and early 2014; specifically, researchers used scRNA-seq to generate immune cell profiles from tissues.27,28 Recent advancements in scRNA-seq have enabled cellular transcriptome data and immune repertoire information to be combined for the coverage of full-length mRNA,29 which can be used to determine the functional changes and interactions of subsets of immune cells during pathogen invasion and the infiltration landscape of T cells in the tumor microenvironment.30,31 The most widely used transcriptome sequencing technologies in the market today are droplet-based 10× Genomics32 and Smart-seq28 a single-cell full-length transcriptome sequencing technology (Table 1).
Spatial transcriptome technology (ST)
Compared with bulk RNA-seq and scRNA-seq, ST can be used to analyze the information of tissue sections in space without cell enzymatic hydrolysis and maximum restoration of the spatial specificity of gene expression in individual cells in human organs.33 It has advanced considerably in the past 30 years, initially, transcriptome technology could be used to analyze only a small number of genes, but now it can be used for analyzing the complete transcriptome. From the early technologies, such as single-molecule fluorescence in situ hybridization (smFISH)34 and laser capture microdissection (LCM), to the current technologies, such as 10× Visium,9 GeoMx DSP,10 and DBiT-seq (Table 1).11 More details on ST can be found in a recent review.35 Among all ST, for now, the 10× Visium ST is the most widely used.
Single-cell proteomic analysis
Qualitative and quantitative analysis of the single-cell proteome can avoid processing information on the expression level changes caused by post transcriptional and post translational modifications and efficaciously obtain information on the regulatory nature of cells. No tools are available for analyzing the whole proteome at the single-cell level, but various single-cell proteome analysis methods are available in different subsegments, including methods based on antibody-based strategies, mass spectrometry (SCoPE-MS),12 Proseek multiplex technology based on polymerase chain reaction (commonly known as PCR) immunoassay,36 and others. Recently, nanodroplet processing in one pot for trace samples (nanoPOTS) microsample processing13 combined with LCM and mass spectrometry found that in mouse uterine tissue sections prepared for blastocyst implantation, quantitative cell type-specific images of >2,000 proteins could be generated with a high spatial resolution.37 Protein profiling characterized by captured ion mobility has led to the development of four-dimensional (4D) proteomics, which can perform high-depth quantitative analysis 4D proteomics quantification (4D-LFQ, 4D-DIA), and 4D protein modification omics.38,39
Single-cell epigenomic analysis
Cellular epigenetic heterogeneity at the single-cell level can be used to explain the pathological regulatory mechanisms of diseases in the absence of altered genetic information. The detection of DNA methylation in single cells was first conducted in 2015 when Farlik et al.40 developed genome-wide bisulfite sequencing, which was used to analyze genome-wide methylation sites at the single-cell level, but required many DNA samples. Currently, methylation sequencing targeting genomic regions with high CpG content (scCGI-seq)14 and DNA methylation analysis using indexing methods (sci-MET)41 at the single-cell level are also available. There are significant differences in chromatin accessibility between cells, which often indicate gene regulation functions. For analyzing open chromatin conformations with loose features at the single-cell level, sensitive enzymes, such as the Tn5 transposase-based single-cell assay for transposase-accessible chromatin using sequencing (also referred to as scATAC-seq)15 and the DNase I-based single-cell DNase sequencing (also referred to as scDNase-seq)42 are required. The latter can detect a much wider range of chromatin, up to 350,000 reads/cell. To determine the link between the diversity of histone modification sites and changes in the chromatin state, single-cell histone modification, single-cell chromatin immune-precipitation sequencing (also referred to as scChIP-seq), was used in 2015.16 Subsequently, the cleavage under targets and release using nuclease (CUT & RUN) technique, indexing single-cell immunocleavage (also referred to as iscChICseq), and antibody-guided chromatin tagmentation (also referred to as ACT-seq)17,43,44 were developed to overcome the low throughput characteristics of cells. More information on single-cell epigenetic techniques is summarized in a previous review.45
Single-cell multiomics analysis
In the near future, with the help of single-cell multiomics analysis, the genomic information of cells and the regulation of transcription and translation in a cell can be presented in a comprehensive manner. In 2016, Hou et al.18 developed single-cell triple omics sequencing (also referred to as scTrio-seq), and were the first to simultaneously achieve high-throughput sequencing of genomic copy number variation, DNA methylome, and transcriptome.18 Cellular indexing of transcriptomes and epitopes by sequencing (also referred to as CITE-seq) using oligonucleotide-tagged antibodies can be applied to target cell surface proteins and analyze differences between protein and transcript levels in individual cells.19 Presently, multimodal analysis has greatly improved our ability to determine cellular states. The single-cell approaches described above has been applied in the field of hepatology.
Application of findings from single-cell approaches in liver physiology and pathology
The liver is the largest digestive gland and detoxification organ in the human body and participates in the synthesis and decomposition of various substances.46 The incidence of hepatitis B virus (HBV) related to end-stage liver disease have become the main causes of mortality due to liver diseases in China,47 and early in vitro and in vivo studies have found that hepatitis B virus X protein (HBx) can induce malignant transformation and carcinogenesis in hepatocytes.48 In 2019, integrated proteomic genomics identified the key signaling and metabolic pathways in HBV-associated hepatocellular carcinoma (HCC), where mutant CTNNB1-associated ALDOA phosphorylation was identified to be involved in HCC metabolic reprogramming, and metabolic alterations may be the most important factor for advanced cancer and poor clinical outcomes.49 Studies using single-cell technology have provided valuable information on the characteristics of single cells and their gene-expression profile of normal liver and liver diseases from multiple perspectives (Table 2).50–62 As Wang et al.63 provided a comprehensive single-cell immune profile of HBV-infected patients, revealing that increased Treg and Tex cells are associated with the degree of liver damage, and that immune cell interactions networks may promote persistent HBV infection and liver histopathology. Thus, HBV infection is a dynamic process, and a deeper understanding of cellular interactions and the changes of immune microenvironment will help develop more effective therapeutic targets and biomarkers for immunotherapy in chronic hepatitis B (CHB) patients, thereby slowing down the occurrence of end-stage liver disease (Fig. 2).
Table 2. Overview of recent studies on human liver disease at the single-cell level.
| Liver diseases | Samples | Patients | Methods | Synopsis | Year | Ref. |
|---|---|---|---|---|---|---|
| Hepatitis | Huh7.5-NTCP cells | none | scRNA-seq(Illumina HiSeq PE150); CRISPR-Cas9 | Rare transcripts from HBV-infected hepatocytes are preferentially enriched; HBV infection did not significantly alter gene expression but treatment with IFN-α altered gene expression in Huh7.5-NTCP cells | 2021 | 50 |
| Peripheral blood | 12 | scATAC-seq | Identified 12 leukocytic clusters corresponding to five cell types; specific cell type of CHB was B-0 and T-3 | 2022 | 51 | |
| NASH liver tissue | 10 | scRNA-seq (10×) | Cell type-specific transcriptional features of NASH; two distinct populations of activated hepatic stellate cells associated with fibrosis levels | 2022 | 56 | |
| Liver fibrosis / Cirrhosis | Healthy, cirrhotic liver tissue; peripheral blood | 10 | scRNA-seq (10×) | Characterizing and comparing the functional diversity of liver biopsy cells from human fibrotic and normal livers; identified scar-associated TREM2+CD9+macrophages, ACKR1+ and PLVAP+ endothelial cells | 2019 | 52 |
| Healthy, HBV-related cirrhosis, alcoholic cirrhosis liver tissue | 15 | scRNA-seq (Illumina HiSeq X) | Different immune profiles between alcoholic and HBV cirrhotic patients; Galectin-9 may be a potential therapeutic target for alcoholic cirrhosis treatment | 2023 | 53 | |
| ACLF | Healthy, cirrhosis, ACLF liver tissue | 25 | scRNA-seq (Illumina HiSeq X) | Revealed the state of hepatic lymphatic endothelial cells in hepatitis B virus-associated ACLF; monocytes/macrophages infiltrate into ACLF | 2022 | 54 |
| HCC | HCC tumor tissue | 8 | scRNA-seq (10×) | Cellular heterogeneity in HCC; intertumoral heterogeneity higher than intratumor heterogeneity; immunocyte characteristics | 2021 | 57 |
| HCC tumor tissue | 1 | scRNA-seq (SMART- seq) | Heterogeneity of CSCs in human HCC; identified CSCs as CD24+/CD133+/EpCAM+/CD45- cells | 2018 | 58 | |
| HCC tumor tissue; normal liver tissue; peripheral blood | 8 | scRNA-seq (Illumina HiSeq) | Characteristics of immune cell in HCC; revealed specific cytokines/compartmental factors | 2019 | 59 | |
| HCC, iCCA tumor tissue | 7 | scRNA-seq (10×) | Similar developmental trajectories in the same tumor, but heterogeneity in stemness gene expression | 2022 | 60 | |
| HCC Tumor tissue; normal liver tissue; peripheral blood | 6 | scRNA-seq (Illumina HiSeq) | Identified 11 populations of T cells in HCC; TME; exhausted CD8+T cells and Tregs preferentially enriched in HCC | 2017 | 55 | |
| HCC tumor tissue; normal liver tissue | 18 | scRNA-seq (10×) | Immune characteristics in early-relapse HCC; early-relapse tumors have less regulatory T cells, DCs and increased infiltrating CD8+T cells and higher expressed CD161+CD8+T cell compared to the primary tumors | 2021 | 61 | |
| (HCC, iCCA, HH, CHC, ASC, SAR, SLC) Tumor tissue; normal liver tissue; peripheral blood | 124 | scRNA-seq (Illumina HiSeq) | Identified six tumor-associated neutrophils, among them CCL4+ TAN and PD-L1+ TAN could promote tumor growth | 2022 | 62 |
ACLF, acute-on-chronic liver failure; ACKR1, atypical chemokine receptor 1; ASC, adenosquamous carcinoma; CCL4, C-C motif chemokine ligand 4; CHB, chronic hepatitis B; CHC, combined hepatocellular-cholangiocarcinoma; CSCs, cancer stem cells; DCs, dendritic cells; EpCAM, epithelial cell adhesion molecule; HBV, hepatitis B virus; HCC, hepatocellular carcinoma; HH, hepatic hemangioma; iCCA, intrahepatic cholangiocarcinoma; IFN-α, interferon-α; NASH, nonalcoholic steatohepatitis; PD-L1, programmed cell death-ligand 1; PLVAP, plasmalemma vesicle-associated protein; SAR, sarcomatoid carcinoma; scATAC-seq, single-cell assay for transposase-accessible chromatin with high throughput sequencing; scRNA-seq, single-cell transcriptome sequencing; SLC, secondary liver cancer; TAN, tumors-associated neutrophil; TME, tumor microenvironment; TREM2, triggering receptor expressed on myeloid cells-2.
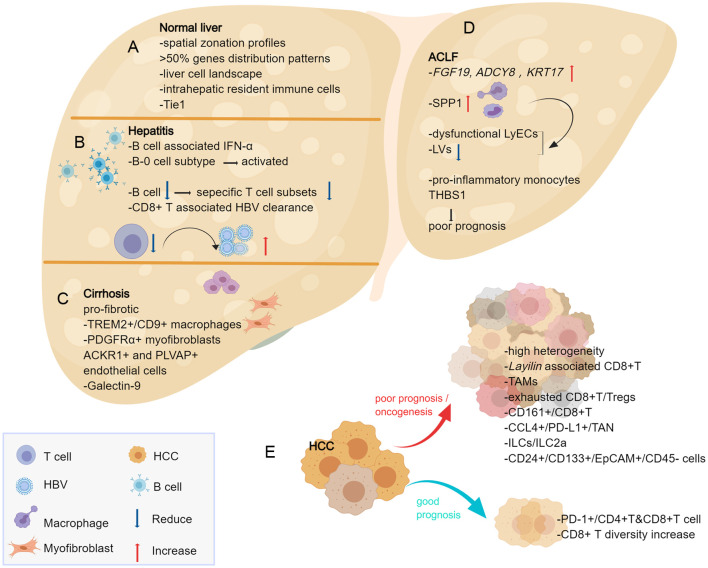
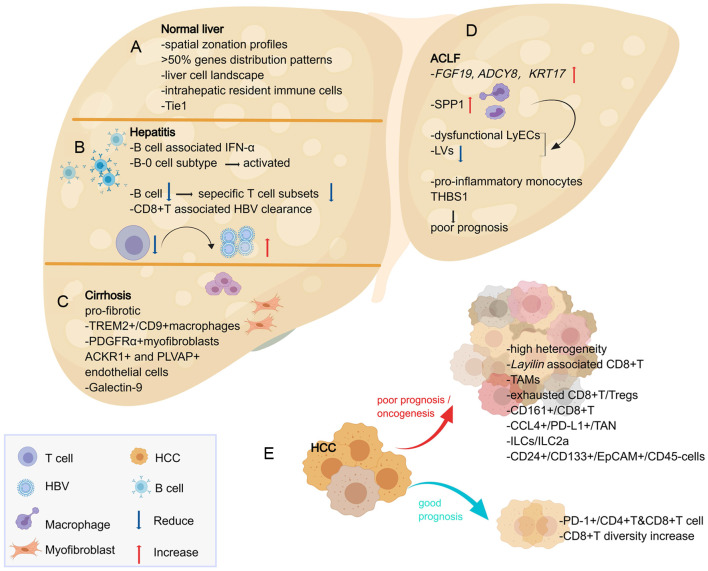
Fig. 2. Insights provided by single-cell approaches into the normal liver and the process of chronic liver disease.
(A) scRNA-seq with ST can reveal spatial zonation profiles of gene expression in the normal liver and a detailed cell landscape of the human liver, showing that 50% of liver genes have partitioning rules. Intrahepatic resident immune cells are demonstrated their unique distribution and functions. The receptor tyrosine kinase Tie1 is the key to regulate vascular activity signals and liver regeneration. (B) The most important feature of the CHB process is immune dysregulation, with CD8+T at the center of the process; B cell reduction impairs specific T-cell subsets and decreases the clearance of HBV and the decreased cell population in hepatitis B samples results in poor IFN-α response but activated B-0 subpopulation. (C) TREM2+/CD9+macrophages and PDGFRα+collagen-producing myofibroblasts subpopulations involve in fibrosis process; ACKR1+ and PLVAP+ endothelial cells are found to accumulate in the fibrotic ecotone and participate in immune cell migration; Galectin-9 on Kupffer cells is associated with specific immunity and could be a potential therapeutic target for cirrhosis treatment. (D) ACLF samples show high expression of the genes FGF19, ADCY8, KRT17 and SPP1 secreted by macrophages resulting in reduced LVs and dysfunctional LyECs. The pro-inflammatory monocyte subtype THBS1 may reflect ACLF progression and poor prognosis. (E) HCC with different prognosis shows differential immune cell microenvironment changes, with increasing PD-1+/CD4+T & CD8+T cells and CD8+T diversity suggesting a good prognosis; high heterogeneity in HCC as well as the increasing subtypes of TAMs, exhausted CD8+T/Tregs, CD161+/CD8+T, CCL4+/PD-L1+/TAN, ILCs/ILC2a, CD24+/CD133+/EpCAM+/CD45- cells suggesting a poor prognosis. The Layilin gene is functionally associated with tumor-characteristic CD8+T cells and may serve as a new target for immunotherapy in HCC. ACLF, acute-on-chronic liver failure; ACKR1, atypical chemokine receptor 1; CHB, chronic hepatitis B; EpCAM, epithelial cell adhesion molecule; HBV, hepatitis B virus; HCC, hepatocellular carcinoma; IFN-α, interferon-α; LVs, lymphatic vessels; LyECs, lymphatic vessel endothelial cells; PD-1, programmed cell death-1; PD-L1, programmed cell death-ligand 1; PDGFRα, recombinant human platelet-derived growth factor-α; PLVAP, plasmalemma vesicle-associated protein; scRNA-seq, single-cell transcriptome sequencing; SPP1, secretory phosphoprotein 1; ST, spatial transcriptome technology; TAMs, tumor-associated macrophages; TAN, tumors-associated neutrophil; THBS1, thrombospondin-1; Tie 1, tyrosine kinase with immunoglobulin like and EGF like domains 1; TREM2, triggering receptor expressed on myeloid cells-2.
Normal liver
scRNA-seq can be used to construct the complete cellular map of the normal liver based on the information of human liver regarding zonation, development, regeneration, and immune cell heterogeneity. In 2017, Halpern et al.64 performed scRNA-seq to measure the entire transcriptome of thousands of mouse hepatocytes, combining it with marker gene function by smFISH to determine the positional coordinate characteristic of liver lobe positions. The researchers found that about 50% of liver genes had partitioning rules and obtained the spatial zonation profiles of the liver.64 In 2019, a group of researchers performed scRNA-seq to analyze approximately 10,000 cells in normal liver tissue from nine human donors and constructed a detailed cellular landscape of the human liver,47 which covered almost all important liver cell types, including hepatocytes, major liver metabolic cells, vascular endothelial cells, liver macrophages, and other immune cell types. A subsequent study integrated and analyzed scRNA-seq datasets from five independent studies of healthy livers from individuals of different sexes and ages and developed an interactively accessible online browser that could be used to access information on the expression of the gene of interest for each cell (sub) type from 28 healthy livers.65 ScRNA-seq combined with ST has provided a comprehensive understanding of the differentiation of all major cell types in the human fetal/embryonic liver, including the function of fetal liver hematopoietic stem cells.66,67 Such information can be used for developing effective strategies for treating liver diseases and enhancing liver regeneration. Other than that the developmental trajectories of various cell types in the human liver at the single-cell level have been described.68 In 2021, Inverso et al.69 combined transcriptomics and proteomics to analyze and obtain high-resolution data on liver vascular endothelial cells and their hepatic vascular signaling regulation. They found that the receptor tyrosine kinase Tie1 (a protein activation region) regulates vascular activity signals and strongly influences the maintenance of liver regeneration.
Immune cell proteomic analysis associated with liver damage and regeneration were developed later.70,71 In 2018, MacParland et al.72 isolated 8,444 parenchymal cells from five healthy human livers and then performed scRNA-seq analysis to obtain the macrophage population distribution of the human liver and mapped the human liver cell landscape. In addition, researchers also used comprehensive transcriptomes of unbiased scRNA-seq to study liver-resident immune cells (LrICs).73 That study showed the tissue distribution, gene expression, and functional modules of specific immune cell populations (natural killer cells, T cells, B cells, and plasma cells) in human liver and presented a landscape of LrICs to explain how the liver promotes immune defense and tolerance in vivo. Liver sinusoidal endothelial cells can function as host immune defenses by coordinating the proper distribution of intrahepatic resident immune cells.74 The integrated information on the liver and immunity by single-cell analysis can promote the development of new immunotherapies and targeted therapies, such as programmed cell death-1 (PD-1) for antitumor treatment.75 Thus, a complete intrahepatic immune signature is essential for understanding the development of liver disease and conditions under which intrahepatic compartmentalization extends to immune cells (Fig. 2).
Hepatitis B
Hepatitis, for various reasons, can lead to adverse outcomes, including cirrhosis, liver failure, or HCC. Among them, for viral hepatitis B, which is very common in China, which affects 257 million people worldwide, a suitable functional cure is lacking, even though effective viral suppression regimens have been developed.76,77 A study used scRNA-seq to analyze crosstalk between B cell and T cell subpopulations and blood samples from interferon-α (IFN-α)-treated CHB patients.78 The results showed that the immune profile of B cells differed across patients, and the effect of IFN-α treatment also differed. Based on the antigen-presenting and costimulatory functions of B cells in cellular immunity, B cell depletion was found to impair functional T-cell subsets, such as specific CD8+T cells, leading to the ineffective clearance of HBV.78 The ability of different subgroups of CD8+T cells to control the virus in patients with chronic viral hepatitis is closely related to the prognosis of the disease.79,80 The CRISPR-Cas9 technology integrated with scRNA-seq analysis can be combined to interpret host gene expression in HBV-infected hepatocytes, including sequence information of low-abundance genes.50 Interestingly, Mederacke et al.81 analyzed the TCR repertoire of CD4+T, CD8+T, and regulatory T cells from peripheral blood and the transplanted livers of liver transplant recipients and showed that their TCR pools were nearly distinct, and the activated T cells might be released by the graft in an acute cellular response to rejection. The latest study on the application of scATAC-seq in the pathophysiology of hepatitis B infection reported specific immune cell populations and chromatin-accessible specific molecules associated with the HBV infection through the detection of peripheral blood mononuclear cells (PBMCs); for example, the B-0 subgroup was found to be over activated in CHB patients.51
Liver fibrosis and cirrhosis
Liver fibrosis involves complex interactions between multiple mesenchymal cells located in scarred areas. These cells include mesenchymal cells, immune cells, vascular endothelial cells, etc. To determine the specific mechanisms of human liver fibrosis, Ramachandran et al.52 used scRNA-seq to analyze the cellular diversity of more than 105 human cells in normal and fibrotic livers of humans and found that TREM2+/CD9+macrophages and PDGFRα+collagen-producing myofibroblast subpopulations were involved in fibrosis. Additionally, profibrotic atypical chemokine receptor 1 (referred to herein as ACKR1) and plasmalemma vesicle-associated protein (referred to herein as PLVAP) endothelial cells were found to accumulate in the fibrotic ecotone and participate in the migration of immune cells, which present several profibrotic signaling pathways through receptor and ligand interactions. Such interactions, combined with newly identified fibrogenic markers, facilitate the development of rational therapeutic targets.
Cirrhosis is a major factor associated with mortality in nonalcoholic steatohepatitis (NASH). The application of scRNA-seq by Schwabe et al.82 provided new insights into the cellular networks involved in the development of NASH fibrosis. The cellular heterogeneity and intercellular transitions involved in the injurious fibrotic response are becoming clearer at the single-cell level.83 To investigate the immune characteristics of cirrhosis due to different causes, a recent study in China performed scRNA-seq on 15 liver specimens (five each from healthy controls, alcoholic cirrhotic (ALC) patients, and HBV-related cirrhotic patients) and found that the proportion of intrahepatic monocytes and macrophages was higher in the ALC group compared to that in the HBV group, with CD5L+Kupffer cells being the predominant subpopulation in ALC. The proportion of T and B cells was significantly lower, which might be related to the expression of Galectin-9 in Kupffer cells.53 Therefore, the immune profile of ALC patients differs from that of HBV cirrhotic patients, and Galectin-9 might be a therapeutic target for treating ALC.
Acute-on-chronic liver failure (ACLF)
Liver failure occurs due to serious liver damage caused by various factors, especially cirrhosis, which leads to severe impairment or loss of liver function. Thus, a comprehensive and systematic understanding of liver failure and the development of new treatment strategies are extremely important. In 2022, a study on the hepatic inflammatory milieu in ACLF found that hepatic lymphatic vessels (LVs) and lymphatic vessel endothelial cells (LyECs) might be new therapeutic targets for treating ACLF.54 That study used scRNA-seq to analyze nonparenchymal cells from three groups of human liver tissue (healthy participants and cirrhotic and ACLF patients) and found the development of apoptotic and dysfunctional LyECs caused by the release of secretory phosphoprotein 1 (also known as SPP1) from infiltrating monocytes/macrophages in the liver. The study also found that the number of LVs was significantly lower in the ACLF group than that in the cirrhotic group. A study on HBV- associated ACLF combined RNA-seq and differential gene analysis and identified key genes associated with immunometabolic disorders, including FGF19, ADCY8, and KRT17. The expression of these genes was significantly upregulated during the progression of ACLF, and thus, they might be sensitive biomarkers for HBV-associated ACLF.84 A transcriptomic study of PBMCs suggested that significant immune dysregulation leads to the development of ACLF from CHB.85 In a recent study, researchers performed scRNA-seq in 17,310 circulating monocytes from healthy & ACLF patients and found five monocyte subpopulations, including pro-inflammatory monocytes, CD16 monocytes, HLA monocytes, megakaryocyte-like monocytes, and natural killer-like monocytes; thrombospondin-1 (THBS1) from pro-inflammatory monocytes characterized specific types of cells and transcriptional signatures, and its production was speculated to reflect the progression of ACLF and a poor prognosis. A comparison between the ACLF survival group and the death group showed that inflammation-related cytokines were significantly higher in the ACLF death group. The results of that study indicated that a detailed analysis of the characteristic monocyte subpopulations and relevant proinflammatory molecular markers might provide therapeutic strategies to arrest the progression of ACLF.86 Shen et al.87 investigated the dynamics of T cells in ACLF longitudinally by collecting PBMCs from five HBV-ACLF patients and sorting total CD4+T and CD8+T cells. They performed unbiased high-throughput sequencing to analyze the TCRβ complementarity-determining regions 3 (also referred to as CDR3) and found that the diversity of the T cell pool decreased significantly as HBV-ACLF progressed. There was also a significant clonal expansion of CD8+T cells but not CD4+T cells. Thus, the diversity of the CD8+T-TCRβ pool might be a predictor of ACLF prognosis. To summarize, single-cell studies assisted by functional experiments might help in understanding the immunological aspect of liver failure and reveal new methods for treating this disease.
HBV-associated liver cancer
Tumorigenesis is a complex pathological process characterized by intratumoral and intertumor heterogeneity and changes in the immune microenvironment.88 HCC is the most prevalent liver tumor due to HBV infection. Here, we summarized the studies based on scRNA-seq related to human HCC in the last five years (Table 2). Tumor heterogeneity and the complexity of the immune microenvironment associated with HCC are the main reasons for its drug resistance and low clinical cure rate.89 To elucidate the immune landscape of HCC, Zheng et al.55 defined 11 functionally characterized T cell subsets and their characteristic genes by combining TCR-seq and scRNA-seq. The gene known as Layilin is functionally associated with tumor-characteristic CD8+T cells and might serve as a new target for immunotherapy in HCC. Wu et al.90 constructed a complete spatial ecological landscape of tumor nodules in a comprehensive genome-wide transcriptome analysis of primary HCC cells, including normal liver tissue, tumor margins, and tumor regions. They found that the tumor envelope might influence the spatial distribution and growth of tumor cells, and HCC heterogeneity can even exist in a single tumor nodule, which might contribute to the ineffectiveness of drug therapy for HCC. Tumor marginal cells are most responsive to their local microenvironment and are highly aggressive and metastatic.91 In 2022, the relationship between the immunophenotype of human liver-resident lymphocytes and HCC was determined by scRNA-seq.92 The researchers found that the changes in the HCC cell number and subpopulation were associated with the dysregulation of innate lymphoid cells (ILCs); seven characteristic ILCs were identified, including inducible T cell costimulator (ICOS)+ ILC2a, which were enriched in HCC and suggested a poor prognosis.
The accurate identification of specific cell subtypes and their targeted molecules is conducive to the diagnosis and treatment of liver diseases.93,94 For example, in the study of intrahepatic cholangiocarcinoma (iCCA), the tagging function of proteomics to complement other histological examinations was used to successfully identify three distinct functional molecular subtypes, of which POE+/C1QB+tumor-associated macrophages (commonly referred to as TAMs) represented a chronic inflammatory marker that might be developed as an immunotherapeutic target for iCCA.95 In a study on the association between HBV and iCCA, a proteomic analysis of the link between cells from HBV-iCCA patients found that a decrease in intercellular adhesion was more likely to lead to an increase in the epithelial-mesenchymal transition signaling.96 Additionally, abnormal methylation was found to be a marker for the progression of cirrhosis to HCC, and altered epigenetic modifications were found to affect chromatin structure and function in HCC patients. For example, the viral protein HBx was found to be an epigenetic regulator, playing an important role in HBV-associated HCC.97–99
Regarding metastatic liver cancer, an adequate understanding of the immune status of the tumor is needed to detect immune checkpoint suppression. By combining transcriptomic, epigenomic, and proteomic data from five HCC cell lines, Wang et al.100 showed that the metastatic potential of HCC is related to the mesenchymal state of tissues and the proliferation ability of cells. They also found that the hypoxic subtype increases the metastatic ability of cancer cells. The combination of multiomics can help identify the regulatory factors of tumor growth and metastasis and reveal the multifaceted nature of tumor cells and their interaction with the immune microenvironment, which might be used to find therapeutic targets.101,102 Yang et al.103 combined transcriptome and whole exome sequencing in primary and metastatic HCC and identified an immunosuppressive target known as SLC2A1, which acts by regulating SPP1+macrophages and T cells. To summarize, single-cell technologies provide valuable information for the diagnosis and treatment of liver cancers, and some of these strategies might be implemented soon in the clinic.
Challenges and future perspectives
The rapid advancement in single-cell sequencing has promoted the identification of the gene structure and gene expression state of single cells at extremely high resolution. With scRNA-seq having a key role, newly developed single-cell approaches have combined genome, epigenome, proteome, spatial transcriptome, and even multiomics to demonstrate the heterogeneity between cells and the changes in the microenvironment of cell populations. Such information might be used for the individualized diagnosis and treatment of diseases and to develop better precision medicine.104 By focusing on the progression of CHB to end-stage liver disease, our single-cell perspective on the mechanism of disease development can help identify relevant sensitive biomarkers and, thus, arrest disease progression on time.
Advances in technology and analytical techniques have encouraged further development in the field of single-cell technology, but research in this field is still at an early and has not been clinically accessible because of many challenges. First, single-cell studies require fresh liver tissue samples, and the availability and dissociation of samples greatly limit the ability to expand the spectrum of liver diseases.105 Although snRNA-seq can reduce the effects of enzymatic hydrolysis and mechanical pressure and is suitable for sequencing frozen samples and hard-to-dissociate samples,24,26,106 its sequencing depth and cell diversity are limited. Thus, snRNA-seq might not be suitable for studying immune cells. In 2023, Yu et al.107 developed a new technology known as “scONE-seq”, which can simultaneously sequence the DNA and RNA of single cells in frozen and fresh tissues. This technology was used for identifying one kind of cancer cell in the human brain with genomic mutations and near-normal transcriptomes. Another challenge involves the complexity of data analysis; specifically, defining quality control criteria, removing technical artifacts, and interpreting results are performed by a few technical staff members. Additionally, open source software tools are unsuitable for most clinical analyses. Therefore, in the future, data visualization and computational methods might be performed by artificial intelligence, which can be designed to analyze complex datasets quickly, thus allowing the networked dissemination of the results of single-cell analysis to a wider field. In addition, the current single-cell sequencing data visualization mainly applies principal component analysis, t-distributed stochastic neighbor embedding, and uniform manifold approximation and projection dimensionality reduction methods to project the cell population into two-dimensional space,108 but in fact, it is difficult for each dimension to correspond to the last biological structure after dimensionality reduction, which will lead to the loss of spatial information in a certain direction of the cell population, so mapping cells to three-dimensional space one by one is the future improvement of single-cell sequencing. Recently, in order to visualize different dataset types in a unified spatial framework, Voytek109 used a pure data approach of neurological mapping to study neuroscience, which opens new doors for other fields to study the spatial three-dimensional mapping of cells. More computational tools and related methods for ST data analysis and visualization are summarized in this review.110 It must be mentioned that the cost of sequencing is an important factor. Therefore, addressing common issues related to single-cell sequencing technologies, such as reducing sequencing costs, simplifying the sequencing process, enhancing sensitivity, and improving resolution, are areas that need to be focused on to further elucidate the molecular mechanisms of the development and progression of different liver diseases.
Novel single-cell techniques combined with multimodal, integrated analytical approaches have expanded the perspective of analysis at different levels within a cell. For example, the newly described technique for studying histone modifications in single cells, known as single-cell chromatin immune cleavage and unmixing sequencing (or scChIX-seq),111 overcomes the limitation that only one histone modification can be studied in a cell. This technology combines and compares the modification patterns of different histones in a cell to generate corresponding maps. The spatial information of gene expression with high resolution provides accurate information for analyzing the native environment of the lesion. The newly described Spatial Tumor Cell Estimator (commonly referred to as SpaCET)112 was used in 2023 to identify different types of immune and stromal cells from information on the spatial abundance of tumor cells. Thus, it can be used to study the role of immune cells in the progression of tumors. In addition, full-length transcriptome sequencing technology represented by single-molecule real-time sequencing can be used to obtain the full-length information of transcripts without splicing. Such information can be used to accurately identify gene mutations and find new mutants quickly.112 Surprisingly, the recently developed droplet-based technology detecting single nucleus information (also referred to as snRandom-seq) allows to capture full-length total RNAs with random primers and detect those noncoding RNA without polyA tails and nascent RNAs.113
The application of single-cell approaches for identifying and assessing HCC is the most prominent use of this technology. However, HCC is often the end-stage manifestation of chronic liver disease, and its prognosis is extremely poor; also, only a few patients are suitable for liver transplantation.114 In China, CHB, cirrhosis-HCC are part of a typical axis of chronic liver disease. Advancements in single-cell technologies are needed for detecting abnormal biomolecules in the early stages of chronic liver disease, especially in the early stages of hepatitis, to arrest the progression of the disease. Some of the latest studies show promise. For example, studies have shown that core transcription factors and pre-mRNA splicing regulators of the hepatocyte phenotype can predict the regression of chronic liver disease.115 Single-cell analysis in the liver involving parenchymal cells, immune cells, and stromal cells has elucidated the pathological mechanisms of viral infections, parasites, cancers, and other liver injuries.74 A more in-depth single-cell analysis of macrophages from patients with liver metastasis was performed to identify two markers associated with cancer.116 In the future, single-cell techniques should be combined with general clinical examinations, and such techniques should be applied beyond basic research to analyze the pathology and physiology of liver disease in patients with different characteristics. Such steps can facilitate precise and individualized treatment of patients with liver disease.
Conclusions
With scRNA-seq as a core sequencing technology, single-cell approaches have yielded significant discoveries in the physiology and pathology of the liver at an unprecedented high resolution. We comprehensively discuss the application and findings of the single-cell approach in the progression of CHB to liver cancer or liver failure. These continuously optimized single-cell techniques provide invaluable tools for exploring subpopulations of liver cells and developing new biomarkers. At present, experience in single-cell technology is most extensive in HCC, and we look forward to increased use in other liver diseases, reduced cost, and simplifying the procedures. One day, single-cell technology will become a basic tool for clinical diagnosis, achieving precise treatment of patients with liver disease.
Abbreviations
- 4D
four-dimensional
- 10×
10× Genomics
- ACLF
acute-on-chronic liver failure
- ACKR1
atypical chemokine receptor 1
- ALC
alcoholic cirrhotic
- CCL4
C-C motif chemokine ligand 4
- CHB
chronic hepatitis B
- CHC
combined hepatocellular-cholangiocarcinoma
- CITE-seq
cellular indexing of transcriptomes and epitopes by sequencing
- CSCs
cancer stem cells
- CUT & RUN
cleavage under targets and release using nuclease
- DCs
dendritic cells
- EpCAM
epithelial cell adhesion molecule
- FFPE
formalin-fixed and paraffin embedded
- HBV
hepatitis B virus
- HCC
hepatocellular carcinoma
- HH
hepatic hemangioma
- iCCA
intrahepatic cholangiocarcinoma
- IFN-α
interferon-α
- ILCs
innate lymphoid cells
- LVs
lymphatic vessels
- LyECs
lymphatic vessel endothelial cells
- nanoPOTS
nanodroplet processing in one pot for trace samples
- NASH
nonalcoholic steatohepatitis
- PBMCs
peripheral blood mononuclear cells
- PD-1
programmed cell death-1
- PDGFRα
recombinant human platelet-derived growth factor-α
- PD-L1
programmed cell death-ligand 1
- PLVAP
plasmalemma vesicle-associated protein
- SAR
sarcomatoid carcinoma
- scATAC-seq
single-cell assay for transposase-accessible chromatin using sequencing
- scCGI-seq
single-cell CpG island methylation sequencing
- scChIP-seq
single-cell chromatin immune-precipitation sequencing
- scChIX-seq
single-cell chromatin immune cleavage and unmixing sequencing
- scCOOL-seq
single-cell chromatin overall omic-scale landscape sequencing
- scDNase-seq
single-cell DNase sequencing
- sci-MET
single-cell combinatorial indexing for methylation analysis
- SCoPE-MS
single-cell proteomics by mass spectrometry
- scRNA-seq
single-cell RNA sequencing
- scTrio-seq
single-cell triple omics sequencing
- scWestern
single-cell western blotting
- SLC
secondary liver cancer
- smFISH
single-molecule fluorescence in situ hybridization
- snRNA-seq
single-nucleus RNA sequencing
- SPP1
secretory phosphoprotein 1
- ST
spatial transcriptome technology
- TAMs
tumor-associated macrophages
- TAN
tumors-associated neutrophil
- THBS1
thrombospondin-1
- Tie 1
tyrosine kinase with immunoglobulin like and EGF like domains 1
- TREM2
triggering receptor expressed on myeloid cells-2
References
- 1.Asrani SK, Devarbhavi H, Eaton J, Kamath PS. Burden of liver diseases in the world. J Hepatol. 2019;70(1):151–171. doi: 10.1016/j.jhep.2018.09.014. [DOI] [PubMed] [Google Scholar]
- 2.Xiao J, Wang F, Wong NK, He J, Zhang R, Sun R, et al. Global liver disease burdens and research trends: Analysis from a Chinese perspective. J Hepatol. 2019;71(1):212–221. doi: 10.1016/j.jhep.2019.03.004. [DOI] [PubMed] [Google Scholar]
- 3.Kisseleva T, Brenner D. Molecular and cellular mechanisms of liver fibrosis and its regression. Nat Rev Gastroenterol Hepatol. 2021;18(3):151–166. doi: 10.1038/s41575-020-00372-7. [DOI] [PubMed] [Google Scholar]
- 4.Stravitz RT, Lee WM. Acute liver failure. Lancet. 2019;394(10201):869–881. doi: 10.1016/s0140-6736(19)31894-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Wen L, Tang F. Boosting the power of single-cell analysis. Nat Biotechnol. 2018;36(5):408–409. doi: 10.1038/nbt.4131. [DOI] [PubMed] [Google Scholar]
- 6.Tang F, Barbacioru C, Wang Y, Nordman E, Lee C, Xu N, et al. mRNA-Seq whole-transcriptome analysis of a single cell. Nat Methods. 2009;6(5):377–382. doi: 10.1038/nmeth.1315. [DOI] [PubMed] [Google Scholar]
- 7.Grindberg RV, Yee-Greenbaum JL, McConnell MJ, Novotny M, O’Shaughnessy AL, Lambert GM, et al. RNA-sequencing from single nuclei. Proc Natl Acad Sci U S A. 2013;110(49):19802–19807. doi: 10.1073/pnas.1319700110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Picelli S, Faridani OR, Björklund AK, Winberg G, Sagasser S, Sandberg R. Full-length RNA-seq from single cells using Smart-seq2. Nat Protoc. 2014;9(1):171–181. doi: 10.1038/nprot.2014.006. [DOI] [PubMed] [Google Scholar]
- 9.Maynard KR, Collado-Torres L, Weber LM, Uytingco C, Barry BK, Williams SR, et al. Transcriptome-scale spatial gene expression in the human dorsolateral prefrontal cortex. Nat Neurosci. 2021;24(3):425–436. doi: 10.1038/s41593-020-00787-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Van TM, Blank CU. A user’s perspective on GeoMx(TM) digital spatial profiling. Immunooncol Technol. 2019;1:11–18. doi: 10.1016/j.iotech.2019.05.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Liu Y, Yang M, Deng Y, Su G, Enninful A, Guo CC, et al. High-Spatial-Resolution Multi-Omics Sequencing via Deterministic Barcoding in Tissue. Cell. 2020;183(6):1665–1681.e18. doi: 10.1016/j.cell.2020.10.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Budnik B, Levy E, Harmange G, Slavov N. SCoPE-MS: mass spectrometry of single mammalian cells quantifies proteome heterogeneity during cell differentiation. Genome Biol. 2018;19(1):161. doi: 10.1186/s13059-018-1547-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ouldali H, Sarthak K, Ensslen T, Piguet F, Manivet P, Pelta J, et al. Electrical recognition of the twenty proteinogenic amino acids using an aerolysin nanopore. Nat Biotechnol. 2020;38(2):176–181. doi: 10.1038/s41587-019-0345-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Han L, Wu HJ, Zhu H, Kim KY, Marjani SL, Riester M, et al. Bisulfite-independent analysis of CpG island methylation enables genome-scale stratification of single cells. Nucleic Acids Res. 2017;45(10):e77. doi: 10.1093/nar/gkx026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Chen X, Miragaia RJ, Natarajan KN, Teichmann SA. A rapid and robust method for single cell chromatin accessibility profiling. Nat Commun. 2018;9(1):5345. doi: 10.1038/s41467-018-07771-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Rotem A, Ram O, Shoresh N, Sperling RA, Goren A, Weitz DA, et al. Single-cell ChIP-seq reveals cell subpopulations defined by chromatin state. Nat Biotechnol. 2015;33(11):1165–1172. doi: 10.1038/nbt.3383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Bartosovic M, Kabbe M, Castelo-Branco G. Single-cell CUT&Tag profiles histone modifications and transcription factors in complex tissues. Nat Biotechnol. 2021;39(7):825–835. doi: 10.1038/s41587-021-00869-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Hou Y, Guo H, Cao C, Li X, Hu B, Zhu P, et al. Single-cell triple omics sequencing reveals genetic, epigenetic, and transcriptomic heterogeneity in hepatocellular carcinomas. Cell Res. 2016;26(3):304–319. doi: 10.1038/cr.2016.23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Stoeckius M, Hafemeister C, Stephenson W, Houck-Loomis B, Chattopadhyay PK, Swerdlow H, et al. Simultaneous epitope and transcriptome measurement in single cells. Nat Methods. 2017;14(9):865–868. doi: 10.1038/nmeth.4380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Goetz JJ, Trimarchi JM. Transcriptome sequencing of single cells with Smart-Seq. Nat Biotechnol. 2012;30(8):763–765. doi: 10.1038/nbt.2325. [DOI] [PubMed] [Google Scholar]
- 21.Macosko EZ, Basu A, Satija R, Nemesh J, Shekhar K, Goldman M, et al. Highly Parallel Genome-wide Expression Profiling of Individual Cells Using Nanoliter Droplets. Cell. 2015;161(5):1202–1214. doi: 10.1016/j.cell.2015.05.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Gao C, Zhang M, Chen L. The Comparison of Two Single-cell Sequencing Platforms: BD Rhapsody and 10× Genomics Chromium. Curr Genomics. 2020;21(8):602–609. doi: 10.2174/1389202921999200625220812. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Quadri SM. Single-cell western blotting. Methods Mol Biol. 2015;1312:455–464. doi: 10.1007/978-1-4939-2694-7_46. [DOI] [PubMed] [Google Scholar]
- 24.Wu H, Kirita Y, Donnelly EL, Humphreys BD. Advantages of Single-Nucleus over Single-Cell RNA Sequencing of Adult Kidney: Rare Cell Types and Novel Cell States Revealed in Fibrosis. J Am Soc Nephrol. 2019;30(1):23–32. doi: 10.1681/asn.2018090912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Slyper M, Porter CBM, Ashenberg O, Waldman J, Drokhlyansky E, Wakiro I, et al. Author Correction: A single-cell and single-nucleus RNA-Seq toolbox for fresh and frozen human tumors. Nat Med. 2020;26(8):1307. doi: 10.1038/s41591-020-0976-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ding J, Adiconis X, Simmons SK, Kowalczyk MS, Hession CC, Marjanovic ND, et al. Systematic comparison of single-cell and single-nucleus RNA-sequencing methods. Nat Biotechnol. 2020;38(6):737–746. doi: 10.1038/s41587-020-0465-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Shalek AK, Satija R, Adiconis X, Gertner RS, Gaublomme JT, Raychowdhury R, et al. Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells. Nature. 2013;498(7453):236–240. doi: 10.1038/nature12172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Jaitin DA, Kenigsberg E, Keren-Shaul H, Elefant N, Paul F, Zaretsky I, et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science. 2014;343(6172):776–779. doi: 10.1126/science.1247651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Stubbington MJT, Lönnberg T, Proserpio V, Clare S, Speak AO, Dougan G, et al. T cell fate and clonality inference from single-cell transcriptomes. Nat Methods. 2016;13(4):329–332. doi: 10.1038/nmeth.3800. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Lyu M, Wang S, Gao K, Wang L, Zhu X, Liu Y, et al. Dissecting the Landscape of Activated CMV-Stimulated CD4+ T Cells in Humans by Linking Single-Cell RNA-Seq With T-Cell Receptor Sequencing. Front Immunol. 2021;12:779961. doi: 10.3389/fimmu.2021.779961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Zheng L, Qin S, Si W, Wang A, Xing B, Gao R, et al. Pan-cancer single-cell landscape of tumor-infiltrating T cells. Science. 2021;374(6574):abe6474. doi: 10.1126/science.abe6474. [DOI] [PubMed] [Google Scholar]
- 32.Ziegenhain C, Vieth B, Parekh S, Reinius B, Guillaumet-Adkins A, Smets M, et al. Comparative Analysis of Single-Cell RNA Sequencing Methods. Mol Cell. 2017;65(4):631–643.e4. doi: 10.1016/j.molcel.2017.01.023. [DOI] [PubMed] [Google Scholar]
- 33.Marx V. Method of the Year: spatially resolved transcriptomics. Nat Methods. 2021;18(1):9–14. doi: 10.1038/s41592-020-01033-y. [DOI] [PubMed] [Google Scholar]
- 34.Kwon S. Single-molecule fluorescence in situ hybridization: quantitative imaging of single RNA molecules. BMB Rep. 2013;46(2):65–72. doi: 10.5483/bmbrep.2013.46.2.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Piñeiro AJ, Houser AE, Ji AL. Research Techniques Made Simple: Spatial Transcriptomics. J Invest Dermatol. 2022;142(4):993–1001.e1. doi: 10.1016/j.jid.2021.12.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Darmanis S, Gallant CJ, Marinescu VD, Niklasson M, Segerman A, Flamourakis G, et al. Simultaneous Multiplexed Measurement of RNA and Proteins in Single Cells. Cell Rep. 2016;14(2):380–389. doi: 10.1016/j.celrep.2015.12.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Piehowski PD, Zhu Y, Bramer LM, Stratton KG, Zhao R, Orton DJ, et al. Automated mass spectrometry imaging of over 2000 proteins from tissue sections at 100-µm spatial resolution. Nat Commun. 2020;11(1):8. doi: 10.1038/s41467-019-13858-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Deng Y, Xie M, Li Q, Xu X, Ou W, Zhang Y, et al. Targeting Mitochondria-Inflammation Circuit by β-Hydroxybutyrate Mitigates HFpEF. Circ Res. 2021;128(2):232–245. doi: 10.1161/circresaha.120.317933. [DOI] [PubMed] [Google Scholar]
- 39.Wu M, Chen Y, Xia H, Wang C, Tan CY, Cai X, et al. Transcriptional and proteomic insights into the host response in fatal COVID-19 cases. Proc Natl Acad Sci U S A. 2020;117(45):28336–28343. doi: 10.1073/pnas.2018030117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Farlik M, Sheffield NC, Nuzzo A, Datlinger P, Schönegger A, Klughammer J, et al. Single-cell DNA methylome sequencing and bioinformatic inference of epigenomic cell-state dynamics. Cell Rep. 2015;10(8):1386–1397. doi: 10.1016/j.celrep.2015.02.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Mulqueen RM, Pokholok D, Norberg SJ, Torkenczy KA, Fields AJ, Sun D, et al. Highly scalable generation of DNA methylation profiles in single cells. Nat Biotechnol. 2018;36(5):428–431. doi: 10.1038/nbt.4112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Jin W, Tang Q, Wan M, Cui K, Zhang Y, Ren G, et al. Genome-wide detection of DNase I hypersensitive sites in single cells and FFPE tissue samples. Nature. 2015;528(7580):142–146. doi: 10.1038/nature15740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Carter B, Ku WL, Kang JY, Hu G, Perrie J, Tang Q, et al. Mapping histone modifications in low cell number and single cells using antibody-guided chromatin tagmentation (ACT-seq) Nat Commun. 2019;10(1):3747. doi: 10.1038/s41467-019-11559-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Ku WL, Pan L, Cao Y, Gao W, Zhao K. Profiling single-cell histone modifications using indexing chromatin immunocleavage sequencing. Genome Res. 2021;31(10):1831–1842. doi: 10.1101/gr.260893.120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Preissl S, Gaulton KJ, Ren B. Characterizing cis-regulatory elements using single-cell epigenomics. Nat Rev Genet. 2023;24(1):21–43. doi: 10.1038/s41576-022-00509-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Ben-Moshe S, Itzkovitz S. Spatial heterogeneity in the mammalian liver. Nat Rev Gastroenterol Hepatol. 2019;16(7):395–410. doi: 10.1038/s41575-019-0134-x. [DOI] [PubMed] [Google Scholar]
- 47.Aizarani N, Saviano A, Sagar, Mailly L, Durand S, Herman JS, et al. A human liver cell atlas reveals heterogeneity and epithelial progenitors. Nature. 2019;572(7768):199–204. doi: 10.1038/s41586-019-1373-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Zhang T, Zhang J, You X, Liu Q, Du Y, Gao Y, et al. Hepatitis B virus X protein modulates oncogene Yes-associated protein by CREB to promote growth of hepatoma cells. Hepatology. 2012;56(6):2051–2059. doi: 10.1002/hep.25899. [DOI] [PubMed] [Google Scholar]
- 49.Gao Q, Zhu H, Dong L, Shi W, Chen R, Song Z, et al. Integrated Proteogenomic Characterization of HBV-Related Hepatocellular Carcinoma. Cell. 2019;179(2):561–577.e22. doi: 10.1016/j.cell.2019.08.052. [DOI] [PubMed] [Google Scholar]
- 50.Le C, Liu Y, López-Orozco J, Joyce MA, Le XC, Tyrrell DL. CRISPR Technique Incorporated with Single-Cell RNA Sequencing for Studying Hepatitis B Infection. Anal Chem. 2021;93(31):10756–10761. doi: 10.1021/acs.analchem.1c02227. [DOI] [PubMed] [Google Scholar]
- 51.Xu H, Yu H, Zheng F, Zhang C, Cai W, Zhang X, et al. Analyzing the gene regulatory network in hepatitis B patients by single-cell ATAC sequencing. Clin Rheumatol. 2022;41(11):3513–3524. doi: 10.1007/s10067-022-06310-z. [DOI] [PubMed] [Google Scholar]
- 52.Ramachandran P, Dobie R, Wilson-Kanamori JR, Dora EF, Henderson BEP, Luu NT, et al. Resolving the fibrotic niche of human liver cirrhosis at single-cell level. Nature. 2019;575(7783):512–518. doi: 10.1038/s41586-019-1631-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Zhang P, Li H, Peng B, Zhang Y, Liu K, Cheng K, et al. Single-cell RNA transcriptomics reveals differences in the immune status of alcoholic and hepatitis B virus-related liver cirrhosis. Front Endocrinol (Lausanne) 2023;14:1132085. doi: 10.3389/fendo.2023.1132085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Zhang P, Li H, Zhou C, Liu K, Peng B, She X, et al. Single-Cell RNA Transcriptomics Reveals the State of Hepatic Lymphatic Endothelial Cells in Hepatitis B Virus-Related Acute-on-Chronic Liver Failure. J Clin Med. 2022;11(10):2910. doi: 10.3390/jcm11102910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Zheng C, Zheng L, Yoo JK, Guo H, Zhang Y, Guo X, et al. Landscape of Infiltrating T Cells in Liver Cancer Revealed by Single-Cell Sequencing. Cell. 2017;169(7):1342–1356.e16. doi: 10.1016/j.cell.2017.05.035. [DOI] [PubMed] [Google Scholar]
- 56.Fred RG, Steen Pedersen J, Thompson JJ, Lee J, Timshel PN, Stender S, et al. Single-cell transcriptome and cell type-specific molecular pathways of human non-alcoholic steatohepatitis. Sci Rep. 2022;12(1):13484. doi: 10.1038/s41598-022-16754-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Ho DW, Tsui YM, Chan LK, Sze KM, Zhang X, Cheu JW, et al. Single-cell RNA sequencing shows the immunosuppressive landscape and tumor heterogeneity of HBV-associated hepatocellular carcinoma. Nat Commun. 2021;12(1):3684. doi: 10.1038/s41467-021-24010-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Zheng H, Pomyen Y, Hernandez MO, Li C, Livak F, Tang W, et al. Single-cell analysis reveals cancer stem cell heterogeneity in hepatocellular carcinoma. Hepatology. 2018;68(1):127–140. doi: 10.1002/hep.29778. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Zhang Q, Lou Y, Yang J, Wang J, Feng J, Zhao Y, et al. Integrated multiomic analysis reveals comprehensive tumour heterogeneity and novel immunophenotypic classification in hepatocellular carcinomas. Gut. 2019;68(11):2019–2031. doi: 10.1136/gutjnl-2019-318912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Ma L, Heinrich S, Wang L, Keggenhoff FL, Khatib S, Forgues M, et al. Multiregional single-cell dissection of tumor and immune cells reveals stable lock-and-key features in liver cancer. Nat Commun. 2022;13(1):7533. doi: 10.1038/s41467-022-35291-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Sun Y, Wu L, Zhong Y, Zhou K, Hou Y, Wang Z, et al. Single-cell landscape of the ecosystem in early-relapse hepatocellular carcinoma. Cell. 2021;184(2):404–421.e16. doi: 10.1016/j.cell.2020.11.041. [DOI] [PubMed] [Google Scholar]
- 62.Xue R, Zhang Q, Cao Q, Kong R, Xiang X, Liu H, et al. Liver tumour immune microenvironment subtypes and neutrophil heterogeneity. Nature. 2022;612(7938):141–147. doi: 10.1038/s41586-022-05400-x. [DOI] [PubMed] [Google Scholar]
- 63.Zhang C, Li J, Cheng Y, Meng F, Song JW, Fan X, et al. Single-cell RNA sequencing reveals intrahepatic and peripheral immune characteristics related to disease phases in HBV-infected patients. Gut. 2023;72(1):153–167. doi: 10.1136/gutjnl-2021-325915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Halpern KB, Shenhav R, Matcovitch-Natan O, Toth B, Lemze D, Golan M, et al. Single-cell spatial reconstruction reveals global division of labour in the mammalian liver. Nature. 2017;542(7641):352–356. doi: 10.1038/nature21065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Brancale J, Vilarinho S. A single cell gene expression atlas of 28 human livers. J Hepatol. 2021;75(1):219–220. doi: 10.1016/j.jhep.2021.03.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Hou X, Yang Y, Li P, Zeng Z, Hu W, Zhe R, et al. Integrating Spatial Transcriptomics and Single-Cell RNA-seq Reveals the Gene Expression Profling of the Human Embryonic Liver. Front Cell Dev Biol. 2021;9:652408. doi: 10.3389/fcell.2021.652408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Gao S, Shi Q, Zhang Y, Liang G, Kang Z, Huang B, et al. Identification of HSC/MPP expansion units in fetal liver by single-cell spatiotemporal transcriptomics. Cell Res. 2022;32(1):38–53. doi: 10.1038/s41422-021-00540-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Wesley BT, Ross ADB, Muraro D, Miao Z, Saxton S, Tomaz RA, et al. Single-cell atlas of human liver development reveals pathways directing hepatic cell fates. Nat Cell Biol. 2022;24(10):1487–1498. doi: 10.1038/s41556-022-00989-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Inverso D, Shi J, Lee KH, Jakab M, Ben-Moshe S, Kulkarni SR, et al. A spatial vascular transcriptomic, proteomic, and phosphoproteomic atlas unveils an angiocrine Tie-Wnt signaling axis in the liver. Dev Cell. 2021;56(11):1677–1693.e10. doi: 10.1016/j.devcel.2021.05.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Ölander M, Wiśniewski JR, Artursson P. Cell-type-resolved proteomic analysis of the human liver. Liver Int. 2020;40(7):1770–1780. doi: 10.1111/liv.14452. [DOI] [PubMed] [Google Scholar]
- 71.Guilliams M, Bonnardel J, Haest B, Vanderborght B, Wagner C, Remmerie A, et al. Spatial proteogenomics reveals distinct and evolutionarily conserved hepatic macrophage niches. Cell. 2022;185(2):379–396.e38. doi: 10.1016/j.cell.2021.12.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.MacParland SA, Liu JC, Ma XZ, Innes BT, Bartczak AM, Gage BK, et al. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat Commun. 2018;9(1):4383. doi: 10.1038/s41467-018-06318-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Zhao J, Zhang S, Liu Y, He X, Qu M, Xu G, et al. Single-cell RNA sequencing reveals the heterogeneity of liver-resident immune cells in human. Cell Discov. 2020;6:22. doi: 10.1038/s41421-020-0157-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Ray K. Zones of immune defence in the liver. Nat Rev Gastroenterol Hepatol. 2021;18(2):81. doi: 10.1038/s41575-020-00403-3. [DOI] [PubMed] [Google Scholar]
- 75.Bukhari S, Henick BS, Winchester RJ, Lerrer S, Adam K, Gartshteyn Y, et al. Single-cell RNA sequencing reveals distinct T cell populations in immune-related adverse events of checkpoint inhibitors. Cell Rep Med. 2023;4(1):100868. doi: 10.1016/j.xcrm.2022.100868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Nguyen MH, Wong G, Gane E, Kao JH, Dusheiko G. Hepatitis B Virus: Advances in Prevention, Diagnosis, and Therapy. Clin Microbiol Rev. 2020;33(2):e00046–19. doi: 10.1128/CMR.00046-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Wang D, Fu B, Wei H. Advances in Immunotherapy for Hepatitis B. Pathogens. 2022;11(10):1116. doi: 10.3390/pathogens11101116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Zhong S, Li Q, Wen C, Li Y, Zhou Y, Jin Z, et al. Interferon α facilitates anti-HBV cellular immune response in a B cell-dependent manner. Antiviral Res. 2022;207:105420. doi: 10.1016/j.antiviral.2022.105420. [DOI] [PubMed] [Google Scholar]
- 79.Wen C, Dong Z, Wang Y, Ye G, Ma Y, Yi X, et al. CTLA4(+)CD4(+)CXCR5(-)FOXP3(+) T cells associate with unfavorable outcome in patients with chronic HBV infection. BMC Immunol. 2023;24(1):3. doi: 10.1186/s12865-022-00537-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Li Y, Tang L, Guo L, Chen C, Gu S, Zhou Y, et al. CXCL13-mediated recruitment of intrahepatic CXCR5(+)CD8(+) T cells favors viral control in chronic HBV infection. J Hepatol. 2020;72(3):420–430. doi: 10.1016/j.jhep.2019.09.031. [DOI] [PubMed] [Google Scholar]
- 81.Mederacke YS, Nienen M, Jarek M, Geffers R, Hupa-Breier K, Babel N, et al. T cell receptor repertoires within liver allografts are different to those in the peripheral blood. J Hepatol. 2021;74(5):1167–1175. doi: 10.1016/j.jhep.2020.12.014. [DOI] [PubMed] [Google Scholar]
- 82.Schwabe RF, Tabas I, Pajvani UB. Mechanisms of Fibrosis Development in Nonalcoholic Steatohepatitis. Gastroenterology. 2020;158(7):1913–1928. doi: 10.1053/j.gastro.2019.11.311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Terkelsen MK, Bendixen SM, Hansen D, Scott EAH, Moeller AF, Nielsen R, et al. Transcriptional Dynamics of Hepatic Sinusoid-Associated Cells After Liver Injury. Hepatology. 2020;72(6):2119–2133. doi: 10.1002/hep.31215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Yang L, Zhen L, Li Z, Zhu S, Xu W, Luo Q, et al. Human liver tissue transcriptomics revealed immunometabolic disturbances and related biomarkers in hepatitis B virus-related acute-on-chronic liver failure. Front Microbiol. 2022;13:1080484. doi: 10.3389/fmicb.2022.1080484. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Li J, Liang X, Jiang J, Yang L, Xin J, Shi D, et al. PBMC transcriptomics identifies immune-metabolism disorder during the development of HBV-ACLF. Gut. 2022;71(1):163–175. doi: 10.1136/gutjnl-2020-323395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Yao J, Liu T, Zhao Q, Ji Y, Bai J, Wang H, et al. Genetic landscape and immune mechanism of monocytes associated with the progression of acute-on-chronic liver failure. Hepatol Int. 2023;17(3):676–688. doi: 10.1007/s12072-022-10472-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Shen G, Sun S, Huang J, Deng H, Xu Y, Wang Z, et al. Dynamic changes of T cell receptor repertoires in patients with hepatitis B virus-related acute-on-chronic liver failure. Hepatol Int. 2020;14(1):47–56. doi: 10.1007/s12072-019-10008-x. [DOI] [PubMed] [Google Scholar]
- 88.Jerby-Arnon L, Shah P, Cuoco MS, Rodman C, Su MJ, Melms JC, et al. A Cancer Cell Program Promotes T Cell Exclusion and Resistance to Checkpoint Blockade. Cell. 2018;175(4):984–997.e24. doi: 10.1016/j.cell.2018.09.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Aliya S, Lee H, Alhammadi M, Umapathi R, Huh YS. An Overview on Single-Cell Technology for Hepatocellular Carcinoma Diagnosis. Int J Mol Sci. 2022;23(3):1402. doi: 10.3390/ijms23031402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Wu R, Guo W, Qiu X, Wang S, Sui C, Lian Q, et al. Comprehensive analysis of spatial architecture in primary liver cancer. Sci Adv. 2021;7(51):eabg3750. doi: 10.1126/sciadv.abg3750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Smit MM, Feller KJ, You L, Storteboom J, Begce Y, Beerens C, et al. Spatially Annotated Single Cell Sequencing for Unraveling Intratumor Heterogeneity. Front Bioeng Biotechnol. 2022;10:829509. doi: 10.3389/fbioe.2022.829509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.He Y, Luo J, Zhang G, Jin Y, Wang N, Lu J, et al. Single-cell profiling of human CD127(+) innate lymphoid cells reveals diverse immune phenotypes in hepatocellular carcinoma. Hepatology. 2022;76(4):1013–1029. doi: 10.1002/hep.32444. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Wernberg CW, Ravnskjaer K, Lauridsen MM, Thiele M. The Role of Diagnostic Biomarkers, Omics Strategies, and Single-Cell Sequencing for Nonalcoholic Fatty Liver Disease in Severely Obese Patients. J Clin Med. 2021;10(5):930. doi: 10.3390/jcm10050930. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Liu Z, Wang Y, Yao Y, Fang Z, Miao QR, Ye M. Quantitative proteomic and phosphoproteomic studies reveal novel 5-fluorouracil resistant targets in hepatocellular carcinoma. J Proteomics. 2019;208:103501. doi: 10.1016/j.jprot.2019.103501. [DOI] [PubMed] [Google Scholar]
- 95.Bao X, Li Q, Chen J, Chen D, Ye C, Dai X, et al. Molecular Subgroups of Intrahepatic Cholangiocarcinoma Discovered by Single-Cell RNA Sequencing-Assisted Multiomics Analysis. Cancer Immunol Res. 2022;10(7):811–828. doi: 10.1158/2326-6066.Cir-21-1101. [DOI] [PubMed] [Google Scholar]
- 96.Shen Y, Xu S, Ye C, Li Q, Chen R, Wu W, et al. Proteomic and single-cell landscape reveals novel pathogenic mechanisms of HBV-infected intrahepatic cholangiocarcinoma. iScience. 2023;26(2):106003. doi: 10.1016/j.isci.2023.106003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Nagaraju GP, Dariya B, Kasa P, Peela S, El-Rayes BF. Epigenetics in hepatocellular carcinoma. Semin Cancer Biol. 2022;86(Pt 3):622–632. doi: 10.1016/j.semcancer.2021.07.017. [DOI] [PubMed] [Google Scholar]
- 98.Wolinska E, Skrzypczak M. Epigenetic Changes Affecting the Development of Hepatocellular Carcinoma. Cancers (Basel) 2021;13(16):4237. doi: 10.3390/cancers13164237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Taniai M. Alcohol and hepatocarcinogenesis. Clin Mol Hepatol. 2020;26(4):736–741. doi: 10.3350/cmh.2020.0203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Wang S, Xie J, Zou X, Pan T, Yu Q, Zhuang Z, et al. Single-cell multiomics reveals heterogeneous cell states linked to metastatic potential in liver cancer cell lines. iScience. 2022;25(3):103857. doi: 10.1016/j.isci.2022.103857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Chen S, Teichmann SA. Completing the cancer jigsaw puzzle with single-cell multiomics. Nat Cancer. 2021;2(12):1260–1262. doi: 10.1038/s43018-021-00306-5. [DOI] [PubMed] [Google Scholar]
- 102.Wang H, Jiang W, Wang H, Wei Z, Li H, Yan H, et al. Identification of Mutation Landscape and Immune Cell Component for Liver Hepatocellular Carcinoma Highlights Potential Therapeutic Targets and Prognostic Markers. Front Genet. 2021;12:737965. doi: 10.3389/fgene.2021.737965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Yang S, Qian L, Li Z, Li Y, Bai J, Zheng B, et al. Integrated Multi-Omics Landscape of Liver Metastases. Gastroenterology. 2023;164(3):407–423.e17. doi: 10.1053/j.gastro.2022.11.029. [DOI] [PubMed] [Google Scholar]
- 104.Gohil SH, Iorgulescu JB, Braun DA, Keskin DB, Livak KJ. Applying high-dimensional single-cell technologies to the analysis of cancer immunotherapy. Nat Rev Clin Oncol. 2021;18(4):244–256. doi: 10.1038/s41571-020-00449-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.van den Brink SC, Sage F, Vértesy Á, Spanjaard B, Peterson-Maduro J, Baron CS, et al. Single-cell sequencing reveals dissociation-induced gene expression in tissue subpopulations. Nat Methods. 2017;14(10):935–936. doi: 10.1038/nmeth.4437. [DOI] [PubMed] [Google Scholar]
- 106.Slyper M, Porter CBM, Ashenberg O, Waldman J, Drokhlyansky E, Wakiro I, et al. A single-cell and single-nucleus RNA-Seq toolbox for fresh and frozen human tumors. Nat Med. 2020;26(5):792–802. doi: 10.1038/s41591-020-0844-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Yu L, Wang X, Mu Q, Tam SST, Loi DSC, Chan AKY, et al. scONE-seq: A single-cell multi-omics method enables simultaneous dissection of phenotype and genotype heterogeneity from frozen tumors. Sci Adv. 2023;9(1):eabp8901. doi: 10.1126/sciadv.abp8901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Kulkarni A, Anderson AG, Merullo DP, Konopka G. Beyond bulk: a review of single cell transcriptomics methodologies and applications. Curr Opin Biotechnol. 2019;58:129–136. doi: 10.1016/j.copbio.2019.03.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Voytek B. The data science future of neuroscience theory. Nat Methods. 2022;19(11):1349–1350. doi: 10.1038/s41592-022-01630-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Waylen LN, Nim HT, Martelotto LG, Ramialison M. From whole-mount to single-cell spatial assessment of gene expression in 3D. Commun Biol. 2020;3(1):602. doi: 10.1038/s42003-020-01341-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Yeung J, Florescu M, Zeller P, de Barbanson BA, Wellenstein MD, van Oudenaarden A. scChIX-seq infers dynamic relationships between histone modifications in single cells. Nat Biotechnol. 2023;41(6):813–823. doi: 10.1038/s41587-022-01560-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Ru B, Huang J, Zhang Y, Aldape K, Jiang P. Estimation of cell lineages in tumors from spatial transcriptomics data. Nat Commun. 2023;14(1):568. doi: 10.1038/s41467-023-36062-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Xu Z, Zhang T, Chen H, Zhu Y, Lv Y, Zhang S, et al. High-throughput single nucleus total RNA sequencing of formalin-fixed paraffin-embedded tissues by snRandom-seq. Nat Commun. 2023;14(1):2734. doi: 10.1038/s41467-023-38409-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Acharya C, Bajaj JS. Chronic Liver Diseases and the Microbiome-Translating Our Knowledge of Gut Microbiota to Management of Chronic Liver Disease. Gastroenterology. 2021;160(2):556–572. doi: 10.1053/j.gastro.2020.10.056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Berasain C, Arechederra M, Argemí J, Fernández-Barrena MG, Avila MA. Loss of liver function in chronic liver disease: An identity crisis. J Hepatol. 2023;78(2):401–414. doi: 10.1016/j.jhep.2022.09.001. [DOI] [PubMed] [Google Scholar]
- 116.Cortese N, Carriero R, Barbagallo M, Putignano AR, Costa G, Giavazzi F, et al. High-Resolution Analysis of Mononuclear Phagocytes Reveals GPNMB as a Prognostic Marker in Human Colorectal Liver Metastasis. Cancer Immunol Res. 2023;11(4):405–420. doi: 10.1158/2326-6066.Cir-22-0462. [DOI] [PMC free article] [PubMed] [Google Scholar]